Abstract
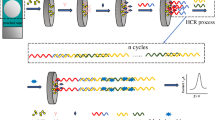
An amperometric aptasensor is reported for the electrochemical determination of the epithelial cell adhesion molecule (EpCAM). It is based on a combination of EpCAM-driven toehold-mediated DNA recycling amplification, the specific recognition of EpCAM aptamer, and its binding to EpCAM. Hairpin probe 1 (Hp1) with a toehold region was modified with a 5′-thiol group (5’-SH) and self-assembled onto the surface of a gold electrode. Upon addition of EpCAM, the probe A (a 15-mer) is liberated from the aptamer/probe A complex and then hybridizes with the toehold domain of Hp1. This results in the exposure of another toehold for further hybridizing with hairpin probe 2 (Hp2) to displace probe A in the presence of Hp2 that was labeled with the electrochemical probe Methylene Blue (MB). Subsequently, liberated probe A is hybridized again with another Hp1 to start the next round of DNA recycling amplification by reusing probe A. This leads to the formation of plenty of MB-labeled DNA strands on the electrode surface and generates an amplified current. This 1:N probe-response amplification results in ultrasensitive and specific detection of EpCAM, with a 20 pg·mL−1 detection limit. The electrode is highly stable and regenerable. It was successfully applied to the determination of EpCAM in spiked human serum, urine and saliva, and thus provides a promising tool for early clinical diagnosis.

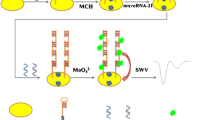
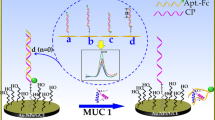
Schematic illustration of the electrochemical detection for EpCAM. The method is based on aptamer-based recognition and EpCAM-driven toehold-mediated DNA recycling amplification. Hp1: Hairpin probe 1; Hp2: Hairpin probe 2; MB: Methylene blue; MCH: 6-Mercapto-1-hexanol; EpCAM: Epithelial cell adhesion molecule.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer-associated morbidity and mortality are primarily caused by cancer metastasis which involves the dissemination of cancer cells from the primary tumor to surrounding tissues and bloodstream via blood or lymph nodes [1]. Clinical diagnosis of cancer primarily depends on imaging techniques or biopsy of diseased cells or tissues [2]. However, these methods are limited to the low sensitivity, capacity of discriminating benign and malignant lesions and diagnosis of cancer at early stage [2]. Biomarkers, including cells [3], proteins [4], nucleic acids [5] and so on, are molecules that differential expression or activity would be observed in pathologic conditions. It can be used as an indicator for evaluation of either normal or pathologic processes [6]. The epithelial cell adhesion molecule (EpCAM) is a trans-membrane glycoprotein involved in cellular signaling, proliferation, differentiation and migration. It is primarily expressed at low levels in a majority of normal epithelial tissues [7]. However, overexpression of EpCAM was found in several human carcinomas [8], malignant effusions [9], and cancer stem cells [10]. Therefore, it is considered as a cancer biomarker for cancer diagnosis, therapy and prognosis. The sensitive and specific detection of EpCAM may provide a potential reference for understanding mechanisms of tumorigenesis, tumor development and metastasis and assisting clinicians to make effective therapy strategies. Methods such as enzyme-linked immunosorbent assay (ELISA) [11], flow cytometry [9, 12], quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) [13] and immunohistochemistry (IHC) [14] were used for quantification of EpCAM or EpCAM-based circulating tumor cells (CTCs). Besides, several novel biosensor strategies have emerged for detection of EpCAM, including fluorescence resonance energy transfer-based bioassay [4, 15], microfluidic immunosensor [11, 16, 17], fluorescent biosensor [18]. Among these biosensors, electrochemical techniques were frequently adopted for EpCAM detection because of its merits of low cost, sensitivity, wide dynamic detection range and compatibility with many miniature devices including microfluidic immunosensor devices [11]. However, these microfluidic immunosensors relying on anti-EpCAM antibody were limited due to its instability and low sensitivity caused by its large size interfering with target binding ability and immune response [12]. Aptamers are single-stranded DNAs or RNAs which possesses several merits of specific recognizing and binding its targets (metal ions, small molecule, proteins and even cells), high stability and easy functionalization. It has been considered as potential alternatives to antibodies for bioanalysis [19]. Similarly, EpCAM aptamer can specifically recognize and bind EpCAM with high affinity to form aptamer-EpCAM complex, which was successfully identified by Song Yanling and its co-workers [12], providing a possibility for establishment of aptamer-based biosensors.
To obtain higher biosensing performance of biosensors, aptamer-based amplification strategies were adopted for enhancing signal readout, such as hybridization chain reactions(HCR) [20], catalytic hairpin assembly(CHA) [21], strand displacement amplification (SDA) [22], rolling circle amplification (RCA) [23], enzyme-assisted signal amplification [24]. Most of these amplification strategies are triggered by toehold domain. Toehold is a short stretching single-stranded domain typically composed of 5–8 nucleotides in a partially hybridized duplex, which is used as a trigger point for initiating hybridization by base-pairing [25]. The invading strand firstly hybridizes with a toehold at one end of partially hybridized duplex and then displaces target stand. This leads to a toehold-mediated strand displacement reaction (TSDR), which can increase the kinetic rate by 106 fold [26]. In the aforementioned amplification strategies, toehold-mediated SDA has been widely applied for signal amplification because of its intriguing merits of simplicity, low cost, enzyme-free operation and highly specific self-powered dynamic self-assembly mechanism [27]. This toehold-mediated SDA includes toehold-mediated DNA recycling amplification [28] and toehold-mediated target recycling amplification [29]. However, no biosensor using aptamer-based amplification strategies has been reported for detecting EpCAM.
An ultrasensitive amperometric aptasensor is described here for electrochemical detection of EpCAM by combining EpCAM-driven toehold-mediated DNA recycling amplification and the specific recognizing and binding of EpCAM aptamer to EpCAM. Upon addition of EpCAM, probe A is firstly liberated from probe A/aptamer complex and hybridizes with toehold domain of Hp1 to open the hairpin structure of Hp1 and expose another toehold domain of Hp1. Subsequently, toehold-mediated SDA is initiated by recycling probe A when the exposed toehold domain of Hp1 further hybridizes with electro-active MB-labeled Hp2 to liberate probe A. Thus, plenty of electro-active MB-labeled Hp2 is brought in proximity to the gold electrode surface for highly efficient electron transfer and then generates the amplified current response. Thus, this electrochemical biosensor has a great potential for ultrasensitive detection of EpCAM.
Experimental section
Materials
All synthetic DNA sequences and aptamer purified by HPLC, were purchased from Sangon Biotechnology Co. Ltd. (Shanghai, China; www.sangon.com) and listed in Table S1. 6-Mercapto-1-hexanol (MCH), Tris(hydroxymethyl) aminomethane (Tris) and tris (2-carboxyethyl) phosphine hydrochloride(TCEP) were all provided by Sigma-Aldrich (St. Louis, MO, USA; www.sigmaaldrich.com). The EpCAM, bovine serum albumin(BSA), CD86 and prostate specific antigen(PSA) for verifying the specificity of the sensor were purchased from Cusabio Biotech Co. (USA; www.cusabio.com). Other reagents were all of analytical grade and used without further purification. The biological samples such as health human serum, saliva and urine were obtained from Dongfeng General Hospital. Ultrapure water used in all runs was prepared by a Millipore water purification system (18.2 MΩ•cm resistivity, Milli-Q Direct 8).
Preparation of the electrochemical biosensor
The gold working electrodes (2 mm in diameter, CH Instruments, Inc., Austin, TX) were first immersed in piranha solution at 90 °C for 5 min for chemical pretreatment before assembling of probes on its surface, and then adequately washed with ultrapure water [30, 31]. Subsequently, the mirror-like surface of gold electrode was obtained by polishing with 0.05 μm γ-alumina powder (CH Instruments, Inc.) and then sonicated with water and ethanol for 5 min, respectively, to remove residual alumina powder. After electrochemically cleaned with 0.5 M H2SO4, the pretreated gold electrodes were obtained by washing with ultrapure water and finally drying with a stream of nitrogen [25]. For the immobilization of probe onto the gold electrode surface, the 10 μM Hp1 was dissolved in 20 mM Tris–HCl buffer (containing 100 mM NaCl, 1.0 mM TCEP, pH 7.4) and incubated in the dark for 1 h to decrease the disulfide bonds of the Hp1. The prepared Hp1 Tris–HCl buffer (5 μM) was diluted to a series of the required concentration before assembling. Subsequently, 10 μL of Hp1 solution was dropped on the surface of the pretreated electrode for Hp1 self-assembly overnight at room temperature. After assembly of Hp1, the Hp1/gold electrodes were obtained by thoroughly rinsing with wash buffer (20 mM Tris-HCl, 100 mM NaCl, pH 7.4) and gently drying with nitrogen. Finally, The obtained Hp1/gold electrode was immersed into 1 mM freshly prepared MCH solution for 1 h to fill the unoccupied region and thus to inhibit the nonspecific DNA adsorption. The modified gold electrodes were finally named MCH/Hp1/AuE. Of note, the modified electrodes were rinsed with buffer (20 mM Tris-HCl, 100 mM NaCl, pH 7.4) and dried with nitrogen before detecting EpCAM.
EpCAM-driven DNA recycling amplification detection of EpCAM
Prior to EpCAM detection, the probe A was hybridized with EpCAM aptamer to form aptamer/probe A duplex. Then, Tris-HCl buffer A containing the complex of EpCAM aptamer/probe A (20 μL), Hp2 (20 μL) and various concentrations of EpCAM (10 μL) were incubated at 37 °C for 20 min. Subsequently, 10 μL of Tris-HCl buffer A was dropped on the MCH/Hp1/AuE in a hybridization oven (UVP, HB-1000) with constant temperature of 37 °C and humidity for 40 min. Subsequently, the electrodes were washed with buffer and transferred into electrochemical cells for measurements.
Electrochemical measurements
Electrochemical measurements were conducted by a CHI660D workstation (CH Instruments Inc., Shanghai, China) with a conventional three-electrode system, including a platinum wire auxiliary electrode, an Ag/AgCl reference electrode, and a modified working gold electrode. Square wave voltammetry (SWV) measurements were conducted in 20 mM Tris-HCl buffer (0.1 M NaCl, pH 7.4) by scanning the potential from −0.4 to 0 V with an amplitude of 25 mV, step potential of 4 mV and frequency of 25 Hz. The electrochemical impedance spectra (EIS) and cyclic voltammetry experiments (CV) were obtained in 0.01 M PBS containing 0.1 M KCl and 5 mM [Fe(CN) 6] 3−/4- in the frequency range from 0.1 Hz to 10 kHz, with the potential window from −0.2 to 0.6 V under a scan rate of 0.1 V·s−1 and amplitude of 5 mV, respectively.
Results and discussion
Sensor design and mechanism of signal amplification
An electrochemical biosensor for ultrasensitive detection of EpCAM was designed. The architecture of electrochemical EpCAM sensor and mechanism of its signal amplification is illustrated in Scheme 1. The Hp1 with a toehold region was labeled with 5’-SH and self-assembled on the gold electrode surface. Subsequently, the unoccupied region of the electrode was passivated with MCH to inhibit the nonspecific DNA adsorption. In this case, the rigid hairpin structures of Hp1 may maintain DNA strands in a relative upright orientation to enhance DNA recognition and hybridization efficiency [32]. Prior to EpCAM detection, the probe A was hybridized with EpCAM aptamer to form aptamer/probe A duplex. In the absence of EpCAM, the occurrence of strand displacement reaction is restrained and thus only a negligible peak current signal is generated. Upon addition of EpCAM, probe A is liberated from the aptamer/probe A duplex because of the specific binding of EpCAM to its aptamer and then hybridizes with toehold domain of Hp1 to open the hairpin structure of Hp1. This results in the exposure of another toehold domain of Hp1 for further hybridizing with Hp2 to form the Hp1-Hp2 duplex which brings electro-active reporter MB in proximity to the gold electrode surface [25]. In addition, the dissociated probe A can hybridize with another Hp1 to start next round of DNA recycling amplification reaction. Finally, plenty of MB labeled DNA strands are formed on the electrode surface and generates the amplified current response.
Electrochemical characterization
Electrochemical impedance spectroscopy (EIS) is usually used to characterize the modification process of electrochemical biosensors. A typical Nyquist plot of EIS contains a semicircle region related to electron transfer resistance at higher frequencies and a linear region at lower frequencies. The enhancement of semicircle diameter implies the enhancement of electron transfer resistance (Ret) on the surface of electrode [30]. To characterize the modification process of the gold electrode at different stages, the Nyquist plots of EIS measurements were obtained. In addition, cyclic voltammetry experiment (CV) was also conducted to further characterize the modification process. The results of EIS and CV are discussed in Electronic Supporting Materials.
The feasibility of the sensor
After successful modification of the gold electrode, the feasibility for ultrasensitive detection of EpCAM was verified via square wave voltammetry (SWV). As depicted in Fig. 1, no obvious current response is observed in the absence of EpCAM/probe duplex and Hp2 due to the lack of electro-active reporter MB on the modified electrode (curve b). When the electrode was incubated with EpCAM aptamer/probe and Hp2, the negligible current changes is exhibited due to inhibition the occurrence of the TSDR triggered by probe A in the absence of EpCAM (curve c). Interestingly, a significant enhancement of current response is exhibited when the modified electrode was incubated with Tris-HCl buffer containing EpCAM aptamer/probe A, Hp2 and EpCAM(curve a). This significant enhancement of current response proves the occurrence of DNA recycling amplification triggered by probe A and the formation of more Hp1-Hp2 duplexes, which brought more electro-active reporter MB proximity to the gold electrode surface for highly efficient electron transfer. Therefore, the aforementioned results further reveal the feasibility of the sensor for ultrasensitive detection of EpCAM.
Optimization of the assay
The following parameters were optimized: (a) concentration of Hp1; (b) concentration of EpCAM aptamer; (c) reaction time; (d) pH; (e) frequency of SWV. Respective data and Figures are given in the Electronic Supporting Materials. The following experimental conditions were found to give best results: (a) Optimal concentration of Hp1: 1.0 μM; (b) Optimal concentration of EpCAM aptamer: 40 nM; (c) Optimal reaction time: 40 min; (d) pH: 7.4; (e) Frequency of SWV: 25 Hz.
Sensitivity and specificity
Under the optimal experimental conditions, the sensitivity of the sensor for detection of EpCAM was investigated in the presence of EpCAM aptamer/probe A and Hp2. Fig. 2a shows SWV current response to EpCAM gradually increases with increasing EpCAM concentration ranging from 0 to 300 ng·mL−1, indicating a highly concentration-dependent current response for detection of EpCAM. The relationship curve of SWV current responses vs. the concentrations of EpCAM was also plotted (Fig. 2b), which exhibits good linear relationship between current response and the concentrations of EpCAM in the range from 0.1 ng·mL−1 to 20 ng·mL−1 (inset in Fig. 2b). The regression equation was expressed as i = 0.6751 + 0.4199C (R2 = 0.992), where i is the SWV current response and C is the EpCAM concentration. The detection limit was calculated to be 20 pg·mL−1 according to 3σ rule. In comparison, such a low detection limit was comparable or even superior to reported literatures listed in Table 1.
a Typical SWV current response of the modified electrode to different concentrations of EpCAM in the presence of EpCAM aptamer/probe A and Hp2 (from bottom to top: 0 to 300 ng·mL−1); bThe relationship curve of SWV current responses vs. the concentrations of EpCAM from 0 to 300 ng·mL−1. Inset of (b): linear relationship between SWV current responses and the concentrations of EpCAM in the range from 0.1 ng·mL−1 to 20 ng·mL−1. The measurements were conducted by scanning the potential from −0.4 to 0 V with an amplitude of 25 mV, step potential of 4 mV and frequency of 25 Hz
To verify the specific recognition and affinity of EpCAM aptamer to EpCAM, the specificity of the sensor for EpCAM detection was also investigated using other proteins as control, such as BSA (100 ng·mL−1), CD86 (100 ng·mL−1), PSA (100 ng·mL−1). As shown in Fig. 3, the response of the modified electrode to control proteins (100 ng·mL−1 of each control protein) exhibits negligible SWV current signal which can almost be the same as that of the blank control (n = 3). However, the significant increase of SWV current signal is observed in the presence of EpCAM (10 ng·mL−1). These results suggest the satisfactory specificity for EpCAM detection.
Stability and regenerability of the sensor
The stability and regenerability of the sensor are two critical indicators for analysis in real biological samples. To investigate the stability of the biosensor, the SWV current responses to EpCAM in the first continuous five days were measured every 24 h for 5 days after storage at 4 °C. As shown in Fig. 4a, compared with its original current response, only a negligible current response changes is observed, indicating the satisfactory stability of the designed biosensor. Moreover, we further investigated the regenerability of the sensor [33]. Prior to rinse with Tris-HCl buffer (10 min), the modified electrode was firstly dipped in 5 M urea solution at temperature of 37 °C for 5 min and then rinsed with 5 M urea solution for 2 min. The results show that the sensor can repeatedly detect EpCAM over 5 times (Fig. 4b). This indicates the good regenerability of assay.
Application to biological samples
To further verify the applicability and selectivity of the sensor system in biological samples, the biological samples were prepared by spiking EpCAM (10 ng) into 200 μL of 50% blank biological samples (human serum, urine and saliva, respectively). Then SWV measurements were separately performed. As shown in Fig. 5, compared with blank biological samples, the significant SWV current enhancement is observed in various biological samples in the presence of 10 ng·mL−1 EpCAM, respectively. In addition, the recoveries for spiking EpCAM into various biological samples were calculated and the negligible current changes (RSD are less than 9.6%) were exhibited in various biological samples with EpCAM concentration of 10 ng·mL−1 (Table 2). The above results suggest the better applicability and selectivity of the sensor system, which is due to the specific recognition and affinity of EpCAM aptamer to EpCAM and the high efficiency toehold-mediated SDA.
Conclusion
In summary, we have designed an amperometric aptasensor for ultrasensitive detection of EpCAM. The design strategy is based on the specific recognition and affinity of EpCAM aptamer to EpCAM and high specific toehold-mediated DNA recycling amplification. The designed sensor exhibits good stability, specificity and regenerability. The sensitivity was also obtained, with the detection limit low to 20 pg·mL−1. Moreover, the designed method was successfully applied for detection of EpCAM in various spiked biological samples, implying the possibility of application of this method in early clinical diagnosis. It is worthy of note that, prior to EpCAM detection, we diluted the human biological samples with the buffer to achieve the desired results. We believe that this design strategy will be a prominent potential for monitoring other biomarkers directly by altering corresponding aptamers.
References
Wang B, Zhou H, Luo Y, Tang R, Zheng S (2014) Biological interfacial engineering for metastatic cancer diagnosis and intervention. Curr Med Chem 21(22):2510–2521
Chinen AB, Guan CM, Ferrer JR, Barnaby SN, Merkel TJ, Mirkin CA (2015) Nanoparticle probes for the detection of cancer biomarkers, cells, and tissues by fluorescence. Chem Rev 115(19):10530–10574
Huhn S, Weinhold N, Nickel J, Pritsch M, Hielscher T, Hummel M, Bertsch U, Huegle-Doerr B, Vogel M, Angermund R, Hanuel M, Salwender HJ, Weisel K, Dürig J, Görner M, Kirchner H, Peter N, Graeven U, Lordick F, Hoffmann M, Reimer P, Blau IW, Jauch A, Dembowsky K, Möhler T, Wuchter P, Goldschmidt H (2017) Circulating tumor cells as a biomarker for response to therapy in multiple myeloma patients treated within the GMMG-MM5 trial. Bone Marrow Transplant 52(8):1194–1198
Shi J, Lyu J, Tian F, Yang M (2017) A fluorescence turn-on biosensor based on graphene quantum dots (GQDs) and molybdenum disulfide (MoS2) nanosheets for epithelial cell adhesion molecule (EpCAM) detection. Biosens Bioelectron 93:182–188
Alhanafy A, Shafei ME, Safan M, Elnour EA, Habib M, Rageh T, Salah El-Din A (2017) Circulating cell free DNA as a biomarker in the serum of colorectal cancer patients. Annal Oncol 28(suppl_7):vii31–vii32
Golubnitschaja O, Flammer J (2007) What are the biomarkers for glaucoma? Sur Ophthalmol 52(Suppl 2):S155–S161
Sadeghi S, Hojati Z, Tabatabaeian H (2017) Cooverexpression of EpCAM and c-myc genes in malignant breast tumours. J Genet 96(1):109–118
Zhang D, Liu X, Gao J, Sun Y, Liu T, Yan Q, Yang X (2017) The role of epithelial cell adhesion molecule N-glycosylation on apoptosis in breast cancer cells. Tumour Biol 39(3):1–8
Roca E, Lacroix R, Judicone C, Laroumagne S, Robert S, Cointe S, Muller A, Kaspi E, Roll P, Brisson AR, Tantucci C, Astoul P, Dignat-George F (2016) Detection of EpCAM-positive microparticles in pleural fluid: a new approach to mini-invasively identify patients with malignant pleural effusions. Oncotarget 7(3):3357–3366
Munz M, Baeuerle PA, Gires O (2009) The emerging role of EpCAM in cancer and stem cell signaling. Cancer Res 69(14):5627–5629
Bravo K, Ortega FG, Messina GA, Sanz MI, Fernandez-Baldo MA, Raba J (2017) Integrated bio-affinity nano-platform into a microfluidic immunosensor based on monoclonal bispecific trifunctional antibodies for the electrochemical determination of epithelial cancer biomarker. Clin Chim Acta 464:64–71
Song Y, Zhu Z, An Y, Zhang W, Zhang H, Liu D, Yu C, Duan W, Yang CJ (2013) Selection of DNA aptamers against epithelial cell adhesion molecule for cancer cell imaging and circulating tumor cell capture. Anal Chem 85(8):4141–4149
Wallwiener CW, Wallwiener M, Kurth RR, Rohm C, Neubauer H, Banys MJ, Staebler A, Schönfisch B, Meuer SC, Giese T, Fehm TN (2011) Molecular detection of breast cancer metastasis in sentinel lymph nodes by reverse transcriptase polymerase chain reaction (RT-PCR): identifying, evaluating and establishing multi-marker panels. Breast Cancer Res Treat 130(3):833–844
Phattarataratip E, Masorn M, Jarupoonphol W, Supatthanayut S, Saeoweiang P (2016) Differential expression of epithelial cell adhesion molecule in salivary gland neoplasms. Ann Diagn Pathol 24:62–67
Das P, Krull UJ (2017) Detection of a cancer biomarker protein on modified cellulose paper by fluorescence using aptamer-linked quantum dots. Analyst 142(17):3132–3135
Ortega FG, Fernández-Baldo MA, Serrano MJ, Messina GA, Lorente JA, Raba J (2015) Epithelial cancer biomarker EpCAM determination in peripheral blood samples using a microfluidic immunosensor based in silver nanoparticles as platform. Sensor Actuat B-Chem 221:248–256
Fernández-Baldo MA, Ortega FG, Pereira SV, Bertolino FA, Serrano MJ, Lorente JA, Raba J, Messina GA (2016) Nanostructured platform integrated into a microfluidic immunosensor coupled to laser-induced fluorescence for the epithelial cancer biomarker determination. Microchem J 2128:18–25
Jung YK, Woo MA, Soh HT, Park HG (2014) Aptamer-based cell imaging reagents capable of fluorescence switching. Chem Commun 50(82):12329–12332
Xu Y, Zhou W, Zhou M, Xiang Y, Yuan R, Chai Y (2015) Toehold strand displacement-driven assembly of G-quadruplex DNA for enzyme-free and non-label sensitive fluorescent detection of thrombin. Biosens Bioelectron 64:306–310
Yang D, Ning L, Gao T, Ye Z, Li G (2015) Enzyme-free dual amplification strategy for protein assay by coupling toehold-mediated DNA strand displacement reaction with hybridization chain reaction. Electrochem Commun 58:33–36
Zhang H, Wang Q, Yang X, Wang K, Li Q, Li Z, Gao L, Nie W, Zheng Y (2017) An isothermal electrochemical biosensor for the sensitive detection of microRNA based on a catalytic hairpin assembly and supersandwich amplification. Analyst 142(2):389–396
Zhang ZZ, Zhang CY (2012) Highly sensitive detection of protein with aptamer-based target-triggering two-stage amplification. Anal Chem 84(3):1623–1629
Bi S, Cui Y, Dong Y, Zhang N (2014) Target-induced self-assembly of DNA nanomachine on magnetic particle for multi-amplified biosensing of nucleic acid, protein, and cancer cell. Biosens Bioelectron 53(4):207–213
Han C, Li R, Li H, Liu S, Xu C, Wang J, Wang Y, Huang J (2017) Ultrasensitive voltammetric determination of kanamycin using a target-triggered cascade enzymatic recycling couple along with DNAzyme amplification. Microchim Acta 184(8):2941–2948
Wang S, Yang F, Jin D, Dai Q, Tu J, Liu Y, Ning Y, Zhang GJ (2017) Toehold mediated one-step conformation-switchable "signal-on" electrochemical DNA sensing enhanced with homogeneous enzymatic amplification. Anal Chem 89(10):5349–5356
Khodakov DA, Khodakova AS, Linacre A, Ellis AV (2013) Toehold-mediated nonenzymatic DNA strand displacement as a platform for DNA genotyping. J Am Chem Soc 135(15):5612–5619
Wang L, Fang L, Liu S (2015) Responsive hairpin DNA aptamer switch to program the strand displacement reaction for the enhanced electrochemical assay of ATP. Analyst 140(17):5877–5880
Meng Y, Hun X, Zhang Y, Luo X (2016) Toehold-aided DNA recycling amplification using hemin and G-quadruplex reporter DNA on magnetic beads as tags for chemiluminescent determination of riboflavin. Microchim Acta 183(11):2965–2971
Yin D, Tao Y, Tang L, Li W, Zhang Z, Li J, Xie G (2017) Cascade toehold-mediated strand displacement along with non-enzymatic target recycling amplification for the electrochemical determination of the HIV-1 related gene. Microchim Acta 184(10):3721–3728
Chen L, Sha L, Qiu Y, Wang G, Jiang H, Zhang X (2015) An amplified electrochemical aptasensor based on hybridization chain reactions and catalysis of silver nanoclusters. Nano 7(7):3300–3308
Cao Y, Zhu S, Yu J, Zhu X, Yin Y, Li G (2012) Protein detection based on small molecule-linked DNA. Anal Chem 84(10):4314–4320
Chen HG, Ren W, Jia J, Feng J, Gao ZF, Li NB, Luo HQ (2016) Fluorometric detection of mutant DNA oligonucleotide based on toehold strand displacement-driving target recycling strategy and exonuclease III-assisted suppression. Biosens Bioelectron 77:40–45
Liu Y, Tuleouva N, Ramanculov E, Revzin A (2010) Aptamer-based electrochemical biosensor for interferon gamma detection. Anal Chem 82(19):8131–8136
Acknowledgements
These works were supported by the Natural Science Foundation of Hubei Provincial Department of Education (Nos.D20172101), the Foundationfor Innovative Research Team of Hubei University of Medicine (2014CXG04), the Science and Technology Key Program of Shiyan (Nos. 16Y70, 17 K74 and 17Y49), the School Foundation for Hubei University of Medicine (No.FDFR201614) and the Key Discipline Project of Hubei University of Medicine.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The author(s) declare that they have no competing interests.
Electronic supplementary material
ESM 1
(DOCX 478 kb)
Rights and permissions
About this article
Cite this article
Chen, Q., Hu, W., Shang, B. et al. Ultrasensitive amperometric aptasensor for the epithelial cell adhesion molecule by using target-driven toehold-mediated DNA recycling amplification. Microchim Acta 185, 202 (2018). https://doi.org/10.1007/s00604-018-2739-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-018-2739-0