Abstract
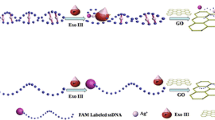
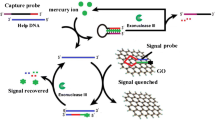
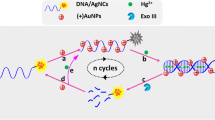
The article describes a fluorometric and sensitive assay for mercury(II) ions (Hg2+). It is based on the following scheme and experimental steps: (1) Hg2+ triggers the self-hybridization of thymine-rich ss-DNA labeled with a fluorescence tag to form a ds-DNA; (2) in the absence of Hg2+, labeled ss-DNA will be adsorbed on the surface of graphene oxide (GO) and its fluorescence is quenched; (3) the ds-DNA formed in the presence of Hg2+ is cleaved by the catalytic action of exonuclease III; (4) the cleaved labeled DNA fragments do not adsorb on the surface of GO, this resulting in an increase in fluorescence intensity. The induction of the process by Hg2+ leads to a strong amplification of fluorescence, while the fluorescence of uncleaved labeled ss-DNA is quenched because it is adsorbed on the surface of GO in the absence of Hg2+. This assay displays a detection limit of 0.1 nM (which is below the 10 nM upper limit in drinking water according to the US EPA and can be performed with 8 min.

The detection scheme is based on the finding that the fluorescence of ds-DNA formed in the presence of Hg(II) ions on the surface of graphene oxide is quenched. If the DNA is cleaved by exonuclease III, fragments will be desorbed and fluorescence pops up
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Contamination of the environment with heavy metal ions has been recognized as an important worldwide issue for decades [1, 2]. Hg2+, as one of the most ubiquitous heavy metals pollutants, is found in the atmosphere, soil, surface waters and even food [3]. According to the U.S. Environmental Protection agency (EPA), an annual total global mercury emission from natural sources as well as human activities is over 7500 tons [4]. Mercury ions can cause serious and permanent damage to the nervous system, kidney, liver, brain and other organs [5]. Because of the high toxicity of Hg2+, the EPA and the international World Health Organization (WHO) have regulated the upper limit of Hg2+ levels in drinking water as 10 nM and 30 nM, respectively [6]. Therefore, there is a need to develop efficient, facile, sensitive and selective analytical methods for determination of Hg2+.
Conventional analytical methods, such as cold-vapor atomic fluorescence spectrometry (CVAFS) [7], cold-vapor atomic absorption spectroscopy (CV-AAS) [8], inductively coupled plasma atomic emission spectrometry (ICPAES) [9], inductively coupled plasma mass spectrometry (ICP-MS) [10], have been used in the detection of Hg2+. These methods are very sensitive and selective. Nevertheless, they require complicated and expensive instruments operated by professional operators, limiting their wide application. Recently, Ono et al. reported that the Hg2+ can be specifically bind to two DNA thymine bases (T) and form stable T-T mismatches in DNA duplexes [11]. The T-Hg2+-T pair mediated by mercury ions is more stable than the natural adenine-thymine (A-T) base pair. Moreover, this T-T mismatch shows high specificity for Hg2+ against other metal ions [12]. Based on these findings, various types of Hg2+ detection assays have been developed in recent years [13–15]. Among these methods, fluorescence bioassays using labeled oligonucleotides (OND), due to its simple operation, fast response and high sensitivity, have attracted particular attention in various fields [16].
Graphene, a single-atom-thick and two-dimensional carbon nanomaterial with extraordinary electronic, mechanical and optical properties, has attracted great research interests in various fields [5, 6]. Graphene oxide (GO), an oxide form of graphene, displaying good water-solubility and flexibility for modification, has been widely applied in biological and biomedical areas [17]. In addition, GO also has some important characteristics such as highly efficient fluorescence quencher; high affinity to single-stranded DNA (ss-DNA) than double-stranded DNA (ds-DNA) or well folded ss-DNA; as well as high affinity to long ss-DNA than short ss-DNA [18]. More recently, by making use of these properties of GO, researchers have developed many fluorescent assays for Hg2+ detection [5, 6]. However, most of these methods didn’t adopt the strategy of signal amplification, leading to their detection limit above the toxic level of Hg2+ in drinking water set by the EPA (10 nM) [19].
Signal amplification strategy based on target recycling, in which a target molecule circularly interacts with different nucleic acid-based signaling probes, is a very useful approach to improving detection sensitivity of bioassays [20, 21]. Nucleases such as Rnase H, nicking endonuclease, DNase I, duplex-specific nuclease, endonuclease IV, exonuclease III (Exo III) are often employed as the cleavage enzymes to promote the target recycling [22]. Different from the sequence-specific nicking endonuclease, Exo III, as exonuclease-assisted target recycling amplification, does not require any specific recognition sequence [23]. Exo III catalyzes the stepwise removal of mononucleotides from the 3-hydroxyl ends of dsDNA with blunt or recessed 3-terminus, while it has limited activity on ssDNA or 3-protruding termini of dsDNA [22]. In view of these features, Exo III has been widely used in the highly sensitive detection assays for DNA, RNA, protein and metal ions [22–25]. Based on these general concepts and researches, we present a highly sensitive and selective fluorescent bioassay for the detection of Hg2+ by combination of GO with Exo III-assisted signal amplification. Moreover, the fabricated sensing platform was successfully applied to the determination of Hg2+ in lake water samples.
Experimental
Apparatus
Fluorescence measurements were carried out on a Hitachi F-4600 spectrophotometer (Hitachi Co. Ltd, Japan) equipped with a xenon lamp excitation source at room temperature. The excitation was set at 495 nm and the emission spectra were collected from 510 to 600 nm. The fluorescence intensity at 518 nm was used to choose the optimal experimental conditions and evaluate the performance of the proposed sensing system. The excitation and emission slit widths were both set at 5 nm.
Materials and reagents
The Exo III was purchased from Takara Biotechnology Co. Ltd (Dalian, China, www.takara.com.cn) and DNA oligonucleotides were synthesized and purified using HPLC by Sangon Biotechnology Co. Ltd (shanghai, China, www.sangon.com). The sequence of the fluorescent dye-labeled ssDNA is 5’-CATTCTTTCTTCCCCT TGTT TGTTT-FAM-3’. GO was synthesized according to our previous report [17]. Graphite powder was oxidized by a mixture of H2SO4 and KMnO4. Water was added to end the reaction. Adequate H2O2 (30 %) was added until the color of the mixture turned to bright yellow. The mixture was washed with dilute HCl solution to remove metal ions, and then washed with water until the solution reached a neutral pH. The resulting solid was dried in air. Finally, the synthesized product was further purified by dialysis to remove the remaining metal species, and dispersed in water under sonication to get the homogeneous GO suspension (0.5 mg mL−1). Tris-hydroxymethyl aminomethane (Tris), the metal salts and all the other reagents employed were of analytical grade and used without further purification. Ultrapure water obtained from a Millipore water purification system (18.2 MΩ cm resistivity, Milli-Q Direct 8) was used in all runs.
Preparation of working solution
The GO stock solution was stable in a few days, but tended to form cluster if it was placed for a long time. Therefore, the GO stock solution should be sonicated for 1 h prior to each use. 1 μM FAM-labeled ssDNA probe stock solution was prepared with Tris buffer (20 mM, 100 mM NaCl, pH 8.2). The standard stock solution of Hg2+ was prepared by dissolving mercuric acetate with 0.1 % acetic acid. Aliquots of various concentrations of Hg2+ solutions were obtained by serial dilution of the stock solution with ultrapure water. The buffer for enzyme digestion was Tris buffer (10 mM, 10 mM KCl, 10 mM MgCl2, pH 8.0, buffer 1). The NEBuffer was always used as the assisted solution of the Exo III. But the NEBuffer contained the dithiothreitol which forms a precipitate with Hg2+. In this work, the activity of Exo III also has no obvious effect without adding the NEBuffer. Fluorescence measurements were performed in the buffer 2 which consisted of Tris buffer (20 mM, 100 mM NaCl, 2 mM MgCl2, pH 7.4). All the solutions were stored at 4 °C.
Fluorescence detection of Hg2+
Thirty nanometer FAM-labeled T-rich ss-DNA probe and a desired concentration of Hg2+ were first mixed and kept at room temperature for 2 min, followed by adding 1 unit of Exo III and buffer 1. Subsequently, the whole 200 μL solution was homogeneously mixed and incubated at 37 °C for 5 min. Finally, the reaction solution was added with GO solution and diluted with buffer 2 to 1 mL. After homogeneous mixing, the fluorescence of the mixture was measured at room temperature. In control experiments, the measurement process was all the same with the above except the addition of Hg2+. Unless otherwise noted, each fluorescence measurement was repeated three times, and the standard deviation was plotted as the error bar.
To evaluate effects of potential experimental conditions on Hg2+ detection, other metal ions, including Fe3+, Zn2+, Cu2+, Ni2+, Co2+, Cd2+ at a concentration of 500 nM, were added respectively (in the presence and absence of 50 nM Hg2+) and the measurement was performed respectively under the same conditions . To investigate the specificity of the assay, the fluorescence response of 50 nM Hg2+ was respectively compared with that of other potentially interfering metal ions (in the absence of 50 nM Hg2+) at high concentrations (500 nM). To evaluate the selectivity of the assay, the fluorescence intensity of 50 nM Hg2+ was respectively compared with that of other potentially interfering metal ions (in the presence of 50 nM Hg2+) at high concentrations (500 nM).
Real sample analysis
To investigate the practical application of the sensing platform, the water sample from Huangjiahu Lake in Wuhan was filtered through a 0.22 μm membrane. The lake water was spiked with a stock solution of Hg2+. Next, the high concentration Tris buffer was added to the mixture. The fluorescence measurement of Hg2+ was then performed in the same manner.
Results and discussion
Bioassay strategy
The inspiration of our sensing strategy for the detection of Hg2+ is based on the following factors: (1) Exo III digests dsDNA with blunt or recessed 3-terminus, but it has limited activity to ssDNA or dsDNA with protruding 3-terminus [20, 22], (2) GO absorbs the FAM-labeled ssDNA probe and quench the fluorescence of probe, while the affinity between very short ssDNA and GO is negligible [26]. (3) Hg2+ mediates the T-rich ssDNA to self-form the well folded dsDNA [12]. (4) the well folded dsDNA with T-Hg2+-T base pairs can be digested by Exo III with the same efficiency as normal dsDNA [27]. The schematic description of the sensing strategy is illustrated in Fig. 1. In the absence of Hg2+, the FAM-labeled ssDNA probe is presented as a random coil structure and resistant to Exo III digestion. So the probe is adsorbed on the surface of GO and the fluorescence is quenched. On the contrary, in the presence of Hg2+, the fluorescence dye-labeled probe forms into a hairpin structure or a mismatch dsDNA between different probes via the Hg2+-mediated coordination of T-Hg2+-T base pairs. The hairpin structure is consisted of a 4-nt loop (CCCC) and a 10-bp stem with a recessed 3-terminus. The Exo III catalyzes the stepwise removal of mononucleotides from the 3-hydroxyl ends of the hairpin structure and mismatch dsDNA, resulting in the release of the target Hg2+ and FAM-labeled mononucleotide. The FAM-labeled mononucleotide does not adsorb on the surface of GO so that the fluorescence intensity of the solution has obvious increase with the progress of the triggered activity of Exo III. Moreover, the released Hg2+ then mediates the new probe, forming the well folded dsDNA and initiating a new cycle of digestion. Through such a catalytic cycle, small amounts of Hg2+ can exponentially trigger the cleavage of FAM-labeled DNA probe by being recycled hundreds of times. So this Exo III-assisted signal amplification technique can offer an ultrahigh sensitivity for the detection of extremely low concentrations of Hg2+.
Assay feasibility
In order to verify the feasibility of the proposed sensing system for Hg2+ assay, the fluorescence signals of Hg2+ measurements in the presence of Exo III as well as a series of control experiments were depicted in Fig. 2. As shown in Fig. 2c, the very weak fluorescence showed that the great majority of the FAM-labeled probe was adsorbed on the surface of GO just in the presence of Hg2+. Upon addition of Exo III, the fluorescence intensity was dramatically increased (shown in Fig. 2a), revealing that Exo III triggered the enzyme digestion reaction and formed the FAM-labeled mononucleotides which were not adsorbed on the surface of GO. Nevertheless, a non-negligible signal was observed just in the presence of Exo III (shown in Fig. 2b), revealing that Exo III non-specifically digests the probe and thus increase the background signal of the sensing platform. These results obviously indicate that the Exo III-assisted cleavage cycle triggered by Hg2+ has been successfully achieved. The quantitative assay of Hg2+ was realized by using the relative fluorescence change: Fr = [(F1–F0)/F0] × 100 %. F1 and F0 are the fluorescence intensities at 517 nm in the presence and absence of Hg2+, respectively.
Optimization of assay conditions
The concentration of Exo III and GO, enzymatic reaction temperature and time were investigated systematically in order to obtain the optimal experimental conditions and attain a high signal-to-noise ratio for the detection of Hg2+ by fixing the 30 nM probe (The details can be found in Electronic Supplementary Material and Figure S1). 1 U mL−1 Exo III, 50 μg mL−1 GO, 37 °C enzymatic reaction temperature, 5 min enzymatic reaction time were chosen as the optimum condition.
Sensitivity of the assay towards Hg2+ detection
Under the optimal experiment conditions, the sensitivity of the bioassay was evaluated. In the presence of various Hg2+ concentrations, the fluorescence emission spectra of the bioassay were shown in Fig. 3. Clearly, the fluorescence signal was dynamically enhanced with increased Hg2+ concentrations from 0 to 350 nM. However, when the Hg2+ ions concentration was higher than 0.5 μM, the fluorescence signal reached a plateau and decreased with further increase of the Hg2+ concentrations (The results were not shown). This may be ascribed to the fact that the very high Hg2+ ions concentrations diminish the activity of Exo III. The inset depicted the relationship between Fr and the Hg2+ concentrations. As shown in the inset of Fig. 3, the value of Fr was proportional to the Hg2+ concentrations in the range of 3 nM to 350 nM. The linear regression equation was Fr = 1.4124C+ 130.4188 with a correlation coefficient of 0.9924, and the detection limit was determined to be 0.1 nM (three times the standard deviation of the blank solution. C is the concentration of Hg2+). The detection limit of this assay is far below the maximum allowable level of Hg2+ in drinking water (10 nM) set by the U.S. EPA. The sensitivity of this assay is also higher than that reported in most of the previous reported methods (as shown in Table 1). These results indicate that the designed bioassay might possess a great application for the detection of Hg2+ in environmental fields.
Selectivity and specificity
In order to evaluate the selectivity and specificity of the bioassay system for possessing great application, various metal ions were added to test the change of the fluorescence intensity. Firstly, we investigated whether these metal ions may markedly change the fluorescence intensity of the bioassay. A variety of environmentally relevant metal ions such as Fe3+, Zn2+, Cu2+, Ni2+, Co2+, Cd2+ were respectively added in the absence of Hg2+. Compared with the blank solution, the fluorescence intensity just had slight change after adding these metal ions (shown in Fig. 4). This indicates that these potentially interfering metal ions do not mediate the same formation of dsDNA as Hg2+ does. In this work, we added 50 μM potentially interfering metal ions in the presence of 50 nM Hg2+. It was found that there was no strongly significant effects upon addition of Fe3+, Zn2+ and Cu2+ ions (the decrease of fluorescence intensity was all less than 20 %), but addition of Cd2+ and Ni+ ions led to a large decrease in the fluorescence intensity (The results were not shown). Therefore, 500 nM of these potentially interfering metal ions were respectively added in the presence of 50 nM Hg2+ in the work. As demonstrated in Fig. 4, the bioassay can detect Hg2+ in the presence of other potentially interfering metal ions at high concentrations. All the above observations show that this bioassay for Hg2+ detection has high selectivity and specificity against the interferences of other metal ions, which are mainly attributed to the specific T-Hg2+-T base pairing [30]. Nevertheless, in view of the strong interference of high concentrations of metal ions in real samples (for instance, Cd2+ and Ni+ ions), the potential solution to reduce the interference of high concentrations of these metal ions in real samples may be that citric acid is added to the analyzed samples, resulting in the chelating ligand between citric acid and heavy metal ions, as described in the previously reported work [16].
Real samples assay
In order to evaluate the performance of this bioassay for the detection of Hg2+ in natural samples, the lake water samples spiked with three different concentrations of Hg2+ were applied. According to the linear regression equation in the range from 3 to 350 nM Hg2+, the ‘Found’ Hg2+ concentrations in the lake water samples were obtained. All the measurements were performed three times, and the results were summarized in Table 2. We observed that the recoveries were in the range of 94.6–106.0 %, and the average RSD was less than 5.0 %. The results clearly indicate that the bioassay could realize the quantification analysis of Hg2+ in real samples.
Conclusions
In summary, a fluorescence bioassay based on GO and Exo III-assisted signal amplification has been developed for the highly sensitive and selective detection of Hg2+ in the present work. Mediated by Hg2+, the FAM-labeled ssDNA forms the well-folded dsDNA and is digested by the Exo III, resulting in the release of FAM-labeled mononucleotide and Hg2+ ions. The FAM-labeled mononucleotide is not adsorbed on the surface of GO, leading to the increase of the fluorescence intensity. Furthermore, the released Hg2+ mediates a new cycle of digestion. In such a way, a small amount of Hg2+ can induce the large number of enzyme digestion reactions and form a significantly amplified fluorescence signal. This bioassay possesses high sensitivity and selectivity, and has fast response (8 min) in comparison to the published work (as shown in table 1). The practical applications in lake water also indicate that this bioassay exhibits great perspective for the detection of Hg2+ in the environmental samples.
References
Wang G, Xu G, Zhu Y, Zhang X (2014) A “turn-on” carbon nanotube-Ag nanoclusters fluorescent sensor for sensitive and selective detection of Hg2+ with cyclic amplification of exonuclease III activity. Chem Commun 50:747–750
Tan D, He Y, Xing X, Zhao Y, Tang H, Pang D (2013) Aptamer functionalized gold nanoparticles based fluorescent probe for the detection of mercury (II) ion in aqueous solution. Talanta 113:26–30
Wang R-Z, Zhou D-L, Huang H, Zhang M, Feng J-J, Wang A-J (2013) Water-soluble homo-oligonucleotide stabilized fluorescent silver nanoclusters as fluorescent probes for mercury ion. Microchim Acta 180:1287–1293
Zhang L, Li T, Li B, Li J, Wang E (2010) Carbon nanotube-DNA hybrid fluorescent sensor for sensitive and selective detection of mercury(ii) ion. Chem Commun 46:1476–1478
Huang J, Gao X, Jia J, Kim J-K, Li Z (2014) Graphene oxide-based amplified fluorescent biosensor for Hg2+ Detection through hybridization chain reactions. Anal Chem 86:3209–3215
Li M, Zhou X, Ding W, Guo S, Wu N (2013) Fluorescent aptamer-functionalized graphene oxide biosensor for label-free detection of mercury(II). Biosens Bioelectron 41:889–893
Li Y, Yan X-P, Dong L-M, Wang S-W, Jiang Y, Jiang D-Q (2005) Development of an ambient temperature post-column oxidation system for high-performance liquid chromatography on-line coupled with cold vapor atomic fluorescence spectrometry for mercury speciation in seafood. J Anal At Spectrom 20:467–472
Pourreza N, Ghanemi K (2009) Determination of mercury in water and fish samples by cold vapor atomic absorption spectrometry after solid phase extraction on agar modified with 2-mercaptobenzimidazole. J Hazard Mater 161:982–987
Zhu X, Alexandratos SD (2007) Determination of trace levels of mercury in aqueous solutions by inductively coupled plasma atomic emission spectrometry: elimination of the ‘memory effect’. Microchem J 86:37–41
Hsu K-C, Lee C-F, Tseng W-C, Chao Y-Y, Huang Y-L (2014) Selective and eco-friendly method for determination of mercury(II) ions in aqueous samples using an on-line AuNPs-PDMS composite microfluidic device/ICP-MS system. Talanta 128:408–413
Ono A, Togashi H (2004) Highly selective oligonucleotide-based sensor for Mercury(II) in aqueous solutions. Angew Chem Int Ed 43:4300–4302
Miyake Y, Togashi H, Tashiro M, Yamaguchi H, Oda S, Kudo M, Tanaka Y, Kondo Y, Sawa R, Fujimoto T, Machinami T, Ono A (2006) MercuryII-mediated formation of Thymine − HgII − Thymine base pairs in DNA Duplexes. J Am Chem Soc 128:2172–2173
Chung CH, Kim JH, Jung J, Chung BH (2013) Nuclease-resistant DNA aptamer on gold nanoparticles for the simultaneous detection of Pb2+ and Hg2+ in human serum. Biosens Bioelectron 41:827–832
Du J, Liu M, Lou X, Zhao T, Wang Z, Xue Y, Zhao J, Xu Y (2012) Highly sensitive and selective chip-based fluorescent sensor for mercuric ion: development and comparison of turn-on and turn-off systems. Anal Chem 84:8060–8066
He Y, Zhang X, Zeng K, Zhang S, Baloda M, Gurung AS, Liu G (2011) Visual detection of Hg2+ in aqueous solution using gold nanoparticles and thymine-rich hairpin DNA probes. Biosens Bioelectron 26:4464–4470
Hu P, Jin L, Zhu C, Dong S (2011) A simple and sensitive fluorescent sensing platform for Hg2+ ions assay based on G-quenching. Talanta 85:713–717
Guo S, Du D, Tang L, Ning Y, Yao Q, Zhang G-J (2013) PNA-assembled graphene oxide for sensitive and selective detection of DNA. Analyst 138:3216–3220
Zhang Q, Kong D-M (2013) A general fluorescent sensor design strategy for “turn-on” activity detection of exonucleases and restriction endonucleases based on graphene oxide. Analyst 138:6437–6444
Qi L, Zhao Y, Yuan H, Bai K, Zhao Y, Chen F, Dong Y, Wu Y (2012) Amplified fluorescence detection of mercury(ii) ions (Hg2+) using target-induced DNAzyme cascade with catalytic and molecular beacons. Analyst 137:2799–2805
Xuan F, Luo X, Hsing IM (2013) Conformation-dependent Exonuclease III activity mediated by metal ions reshuffling on thymine-rich DNA duplexes for an ultrasensitive electrochemical method for Hg2+ detection. Anal Chem 85:4586–4593
Wolfbeis OS (2013) Editorial: probes, sensors, and labels: why is real progress slow? Angew Chem Int Ed 52:9864–9865
Xu Q, Cao A, Zhang L-F, Zhang C-Y (2012) Rapid and label-free monitoring of Exonuclease III-assisted target recycling amplification. Anal Chem 84:10845–10851
Liu S, Wang C, Zhang C, Wang Y, Tang B (2013) Label-free and ultrasensitive electrochemical detection of nucleic acids based on autocatalytic and Exonuclease III-assisted target recycling strategy. Anal Chem 85:2282–2288
Chen H, Wang J, Liang G, Zhang P, Kong J (2012) A novel exonuclease III aided amplification method for sensitive nucleic acid detection based on single walled carbon nanotube induced quenching. Chem Commun 48:269–271
Chen C, Xiang X, Liu Y, Zhou G, Ji X, He Z (2014) Dual-color determination of protein via terminal protection of small-molecule-linked DNA and the enzymolysis of exonuclease III. Biosens Bioelectron 58:205–208
Cheng G, Wang Z-G, Liu Y-L, Zhang J-L, Sun D-H, Ni J-Z (2012) A graphene-based multifunctional affinity probe for selective capture and sequential identification of different biomarkers from biosamples. Chem Commun 48:10240–10242
Xuan F, Luo X, Hsing IM (2012) Ultrasensitive solution-phase electrochemical molecular beacon-based DNA detection with signal amplification by Exonuclease III-assisted target recycling. Anal Chem 84:5216–5220
Cui X, Zhu L, Wu J, Hou Y, Wang P, Wang Z, Yang M (2015) A fluorescent biosensor based on carbon dots-labeled oligodeoxyribonucleotide and graphene oxide for mercury (II) detection. Biosens Bioelectron 63:506–512
Jiang Y, Tian J, Hu K, Zhao Y, Zhao S (2014) Sensitive aptamer-based fluorescence polarization assay for mercury(II) ions and cysteine using silver nanoparticles as a signal amplifier. Microchim Acta 181:1423–1430
Deng L, Zhou Z, Li J, Li T, Dong S (2011) Fluorescent silver nanoclusters in hybridized DNA duplexes for the turn-on detection of Hg2+ ions. Chem Commun 47:11065–11067
Guo L, Yin N, Nie D et al (2011) An ultrasensitive electrochemical sensor for the mercuric ion via controlled assembly of SWCNTs. Chem Commun 47:10665–10667
Hao Y, Guo Q, Wu H et al (2014) Amplified colorimetric detection of mercuric ions through autonomous assembly of G-quadruplex DNAzyme nanowires. Biosens Bioelectron 52:261–264
Wang G, Huang H, Zhang X et al (2012) Electrically contacted enzyme based on dual hairpin DNA structure and its application for amplified detection of Hg2+. Biosens Bioelectron 35:108–114
Xue X, Wang F, Liu X (2008) One-step, room temperature, colorimetric detection of Mercury (Hg2+) using DNA/nanoparticle conjugates. J Am Chem Soc 130:3244–3245
Acknowledgments
The authors acknowledge the support of Natural Science Foundation of China (Nos. 21275040 and 21475034), and the Natural Science Foundation of Hubei Province (No. 2013CFA061).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 756 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Tang, L., Yang, F. et al. Highly sensitive DNA-based fluorometric mercury(II) bioassay based on graphene oxide and exonuclease III-assisted signal amplification. Microchim Acta 182, 1535–1541 (2015). https://doi.org/10.1007/s00604-015-1482-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-015-1482-z