Abstract
The genetic variation and spatial distribution of the ectomycorrhizal fungus Tricholoma terreum was studied in an artificial Larix principis-rupprechtii stand in China. The 33 sporophores studied belonged to distinct genotypes, based on analysis of random amplified polymorphic DNA (RAPD) markers. The genets of T. terreum were small and no larger than 0.5 m. Two major phenetic groups, i.e. eight individuals in group 1 and 25 in group 2, were identified by principal component analysis and the unweighted pair group method with arithmetic means of simple matching coefficients, respectively. The genetic diversity as determined by the Shannon-Weaver diversity index was 4.52 for these 33 sporophores. Furthermore, there was a slightly lower diversity index in group 1 (4.28) than in group 2 (4.46). It is suggested that sexual spore propagation is very important in the life history of T. terreum in this larch forest.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ectomycorrhizas (ECM), a mutual association between plants and fungi, play a significant role in forest ecosystems (Smith and Read 1997). In the world, several thousand fungi form ECM with tree hosts in natural and managed forest habitats (Molina et al. 1992). Each fungal species exists as a population of many genetic individuals, namely genets. Genets differ from each other in their ability to colonize diverse genotypes of host plants, to utilize mineral and organic soil nutrients, and in their adaptation to abiotic factors, such as soil pH and drought (Gherbi et al. 1999; Fiore-Donno and Martin 2001; Jany et al. 2002). Research on the temporal and spatial distributions of populations of ECM fungi, along with study of the origin and maintenance of their genetic variation, is crucial when trying to understand how populations of these symbiotic fungi evolve and disappear in the succession of forest ecosystems (Dahlberg and Stenlid 1995; Egger 1995; Dahlberg 2001; Bruns et al. 2002; Weber et al. 2002).
Molecular techniques, e.g. random amplified polymorphic DNA (RAPD) marker analysis (Williams et al. 1990), have been successfully used in the detection of genetic diversity and discrimination of fungi (Pei et al. 1997; Morris et al. 2000; Calvo-Bado et al. 2001). Especially in ECM fungi, they are very useful in differentiating genotypes of closely related individuals in population studies. Jacobson et al. (1993) pointed out that the isolates of the ECM fungus Suillus granulatus (L. ex Fr.) Kuntze could display polymorphisms based on RAPD markers, although these isolates were shown to be identical by somatic incompatibility testing. The spatial distribution and temporal persistence of several ECM fungi have been studied using the RAPD method (e.g. de la Bastide et al. 1994; Anderson et al. 1998; Selosse et al. 1998; Ito and Yanagi 1999; Jany et al. 2002). A population of Laccaria amethystina Cooke showed numerous small, short-lifespan genets in a mixed forest of Switzerland based on the analysis of RAPD markers (Fiore-Donno and Martin 2001).
Tricholoma terreum (Schaeff.:Fr.) Kummer collected in the present study, with the same morphological characteristics as the T. terreum from Europe (Phillips 1981), is an ECM symbiont mainly associated with species of pine, spruce, and beech (Mao 2000). In the survey of ECM fungi in the Dongling Mountain, Beijing Forest Ecosystem Research Station of the Chinese Academy of Sciences, T. terreum was the dominant species of ECM fungi in the artificial Larix principis-rupprechtii Mayr. stand. To our knowledge, the genetic diversity of T. terreum has not been studied.
The objective of the present study was to determine the level of genetic variation among a T. terreum population collected from the artificial L. principis-rupprechtii stand using RAPD markers, and to investigate the spatial distribution of distinct T. terreum genets.
Materials and methods
Field site and sampling procedure
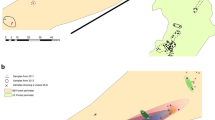
This study was conducted at the artificial Larix principis-rupprechtii stand in Dongling Mountain, Beijing Forest Ecosystem Research Station of the Chinese Academy of Sciences, 117 km west of Beijing, China (39°58′N, 115°26′E). These trees were about 30 years old and located at an altitude of 1,211 m above sea level. Because these trees stood at an even distance of ca. 1.5 m apart, they were not marked in Fig. 1. The mean annual temperature is 2~7°C and the mean annual precipitation is about 500 mm. Because it was unusually dry at the sampling site in summer (June–August 2001) and only one representative sporophore was collected within a circle of 0.5 m diameter (Bonello et al. 1998), a total of only 33 sporophores of Tricholoma terreum were collected from July to August 2001. The locations of each sporophore were mapped in Fig. 1. Each stem of the fruit bodies was cut, and each pileus was packaged with aluminium foil, labelled, and stored inside a freezer (−80 °C) until DNA extraction.
The geographical distribution of the 33 Tricholoma terreum sporophores collected at the Larix principis-rupprechtii forest, which were divided into two groups based on the analysis of the unweighted pair group method with arithmetic means (UPGMA) and the principal component analysis (PCA). The sporophores clustered into group 1 are indicated by ○. The sporophores clustered into group 2 are indicated by ●
DNA extraction
Genomic DNA was extracted using the modified CTAB protocol (Guo et al. 2000). About 600 mg of freezed pileus tissues was cut and transferred into a 1.5-ml Eppendorf microcentrifuge tube with 700 μl of preheated (60°C) 2×CTAB extraction buffer [2% (w/v) CTAB, 100 mM TRIS-HCl, 1.4 M NaCl, 20 mM EDTA, pH 8.0] and ca. 0.2 g sterilized sand (white quartz; Sigma). The pileus was ground using a glass pestle for 5–10 min and then incubated at 65°C for 1 h with occasional gentle swirling. Seven hundred microlitres of phenol:chloroform:isoamyl alcohol (25:24:1) was added to each tube and mixed thoroughly to form an emulsion. The mixture was spun at 13,000 r.p.m. for 15 min at room temperature in a microcentrifuge, and the upper aqueous phase was removed and transferred to a new 1.5-ml tube. The aqueous phase containing DNA was re-extracted with chloroform:isoamyl alcohol (24:1) until no interface was visible. Fifty microlitres of 5 M KOAc was added into the aqueous phase followed by 400 μl isopropanol and inverted gently to mix. The genomic DNA was precipitated at 9,000 r.p.m. for 2 min in a microcentrifuge. The DNA pellet was washed with 70% ethanol twice and air-dried. The DNA pellet was then resuspended in 100 µl TE buffer (10 mM TRIS-HCl, 1 mM EDTA). The final concentration of total DNA was ca. 150 ng/μl.
RAPD assay
Eleven 10-oligonucleotide primers (Shanghai Sangon Biologic Engineering Technology and Service, China) were used in this study, and these primers were S282 (5′-CATCGCCGCA-3′), S381 (5′-GGCATGACCT-3′), S382 (5′-TGGGCGTCCA-3′), S385 (5′-ACGCAGGCAC-3′), S388 (5′-AGCAGGTGGA-3′), S390 (5′-TGGGAGATGG-3′), S392 (5′-GGGCGGTACT-3′), S393 (5′-ACCGCCTGCT-3′), S396 (5′-AGGTTGCAGG-3′), S1097 (5′-CCGCAAGTGT-3′), and S1158 (5′-GGGAGCGCTT-3′).
DNA amplification was performed basically following the method of Williams et al. (1990). The DNA fragment was amplified in an automated thermal cycler (PTC-100; MJ Research, Watertown, Mass.). Amplification was performed in a 25 µl reaction volume which contained PCR buffer [10 mM KCl, 10 mM (NH4)2SO4, 20 mM TRIS-HCl, pH 8.8, 0.1% Triton X-100], 2.5 mM MgCl2, 200 μM of each deoxyribonucleotide triphosphate, 0.4 μM of each primer, ca. 30 ng template DNA, and 1 unit of Taq DNA polymerase (Promega, Madison, Wis.). A negative control using water instead of template DNA was included in the amplification process and all of the mixture was overlaid with a drop of mineral oil. The thermal cycling program was as follows: 3 min initial denaturation at 94°C, followed by 45 cycles of 30 s denaturation at 94°C, 1 min primer annealing at 35°C, 2 min extension at 72°C, and a final 10 min extension at 72°C. Five microlitres of PCR products from each PCR reaction was examined by electrophoresis at 70 V for 5 h in a 1.8% (W/V) agarose gel (Promega) in 0.5×TBE buffer. Gels were stained with ethidium bromide (0.5 μg/ml) and photographed under UV light.
Data analysis
Amplified fragments were scored 1 for presence and 0 for absence. All faint bands occurring in the sporophores were ignored. Band sizes below 300 bp and above 2,000 bp were amplified inconsistently and thus excluded from this analysis. All fragments generated by the primers were recorded and combined into one data matrix. Statistical analysis of the data was performed using the NTSYS-pc version 1.70 (Rohlf 1992). The degree of genetic relatedness or similarity was estimated using simple matching coefficients (Sneath and Sokal 1973): S=(a+d)/(a+b+c+d), where a is the number of (1, 1) matches between pairs, b and c equal the number of (0, 1) and (1, 0) mismatches respectively, and d equals the number of (0, 0) matches. Based on simple matching coefficients, a dendrogram was constructed using SAHN clustering in the unweighted pair group method with arithmetic means (UPGMA) in NTSYS-pc. MXCOMP was used to determine the degree of correlation between the original similarity matrix and the cophenetic value matrix derived from a dendrogram using COPH.
Principal component analysis (PCA) was performed using SAS PROC PRINCOMP. PCA is a multivariate procedure for examining relationships among several quantitative variables by deriving a smaller number of linear combinations (principal components) that retain as much of the information in the original data as possible (SAS Institute 1990).
Genetic diversity within the population of T. terreum was estimated using the Shannon-Weaver diversity index (H 0), which was calculated by the formula \( H_0 = - \sum\limits_{i = 1}^k {pi \times \ln pi} \), where k is the total number of RAPD fragments, and pi is the frequency of a given band (Chalmers et al. 1992).
Results
Thirty-three sporophores of Tricholoma terreum were collected in this survey. A total of 53 RAPD primers was screened for polymorphisms in the pilot study, of which 11 primers produced reproducible clear RAPD bands and were employed in the present study. The size of amplified fragments ranged from 300 bp to 2,000 bp (Fig. 2). From the 11 primers, a total of 109 bands could be observed, of which 100 (91.7%) were polymorphic.
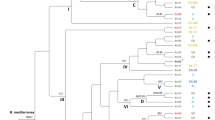
Genetic similarity coefficients between these sporophores ranged from 0.431 to 0.963. Genetic relationships among the 33 individuals were revealed by UPGMA cluster analysis based on similarity coefficients (Fig. 3). The cophenetic correlation of the dendrogram with the original data matrix was 0.89, and this value indicated that there was a high goodness of fit of a cluster analysis to the original data. These 33 fruit bodies were clustered into two major phenetic groups, i.e. eight sporophores within group 1 and 25 within group 2. The genetic similarity within group 1 (75.2%) was higher than that within group 2 (64.6%), and there was a low genetic similarity (55.1%) between these two groups.
The RAPD data were also subjected to PCA, which provided a method independent of UPGMA clustering to evaluate variation among the sporophores. The first three components explained 33.5%, 14.8%, and 4.8% of the total variation, respectively. The two-dimensional plots therefore summarized 48.3% of the total variation in the collection, and these 33 individuals formed two distinct phenetic groups (Fig. 4).
Markers S381–460, S392–400, and S392–2000 presented in six, five, and four of the eight individuals within group 1, respectively, but they were absent in all sporophores of group 2. Marker S382–500 was amplified in 23 out of 25 individuals within group 2. In addition, seven markers (S381–2000, S385–1800, S388–1600, S390–1600, S396–600, S1097–500, and S1097–1400) were present in most individuals (14~22 out of 25 individuals) within group 2, whereas they were all absent within group 1.
The genotypes of the 33 sporophores of T. terreum were all different, and the minimum distance among these samples was 0.5 m. Therefore, the genet size of T. terreum was no more than 0.5 m. Three individuals, i.e. T25, T27, and T30, were clustered in group 2, whereas they were located in the geographic area of group 1.
The genetic diversity as determined by the Shannon-Weaver diversity index was 4.52 of these 33 sporophores, furthermore, there was a slightly lower diversity index within group 1 (4.28) than within group 2 (4.46).
Discussion
Although the size ranges of RAPD markers were different in previous studies, they were all within the similar ranges from 170 to 2,770 bp. Therefore, our values for the ranges of band sizes were comparable with previous studies (Jacobson et al. 1993; de la Bastide et al. 1994; Anderson et al. 1998; Selosse et al. 1998; Fiore-Donno and Martin 2001; Jany et al. 2002).
Our results showed that the Tricholoma terreum population was clustered into two groups by both UPGMA and PCA. The sporocarps in the same group are located close to one another and have a high genetic similarity. It is highly possible that these sporocarps in the same group may originate from closely related spores such as those from the same sporocarps (Zhou et al. 1999). Furthermore, the close location of genetically similar genets suggests that the spores from the same sporocarps do not tend to be dispersed far.
Colpaert et al. (2000) used Shannon's information measure H 0 to calculate the diversity of ECM fungus Suillus luteus (L.:Fr.) Roussel in Belgium. Their results indicated that the average diversity within the population of S. luteus was 3.169 and the mean diversity within species of S. luteus was 4.082. A similar Shannon-Weaver diversity index (4.52) was obtained in the T. terreum population, and similar diversity indices were found in group 1 (4.28) and group 2 (4.46) in the present study. It indicated that the genetic variation of T. terreum was almost the same between group 1 and group 2.
The results showed that the genet size of the T. terreum population was small (no larger than 0.5 m). Results of previous studies indicated that there were wide-ranging sizes of genets in different ECM fungal species, from a few millimetres in Collybia fusipes (Bull.:Fr.) Quel (Marcais et al. 1998) to more than 1 km in Armillaria bulbosa (J. B. Barla) G. A. Kile and R. Watling (Smith et al. 1992). Numerous small (<few millimetres) genets were found in some ECM fungi. Amanita francheti (Boud.) Fayod, Lactarius xanthogalactus Perk, and Russula cremoricolor Earle appeared to have small genets. The minimum sizes for their largest genets were 1.5, 9.3 and 1.1 m2, respectively (Redecker et al. 2001). Hebeloma cylindrosporum Romagnesi, an early colonizer, showed genet sizes of no more than 3.5 m in diameter (Gryta et al. 1997, 2000). Laccaria amethystina had numerous small genets <1 m (Gherbi et al. 1999; Fiore-Donno and Martin 2001), while a big genet size of L. bicolor was 12.5 m (Baar et al. 1994). The occurrence of numerous small genetic individuals might suggest a recent colonization by spores (Dahlberg and Stenlid 1990). This population structure was a preferred reproductive strategy of early stage, ruderal ECM species occurring under young trees or on small-scale disturbed sites (de la Bastide et al. 1994; Selosse et al. 1998, 1999).
However, relatively large genet size was found in Cortinarius rotundisporus (Clel. and Cheel) Horak and Wood. This ECM fungus was the late-stage species analysed and had a large genet of up to 30 m in diameter (Sawyer et al. 1999). Most Suillus species were reported to form few and relatively large genets due to persistence and mycelial spread. Suillus pungens Thiers and Smith was one of the largest genets in the ECM fungi, with a mycelial spread of up to 300 m2 (Bonello et al. 1998). Suillus bovinus (L.:Fr.) Roussel (Dahlberg and Stenlid 1994) and S. variegatus (S.:Fr.) Kuntze (Dahlberg 1997) appeared to form relatively large genets of 20–40 m in diameter, while a little small genet (ca. 6 m) was found in the population of S. grevillei (Klotz.) Sing. (Zhou et al. 1999; 2001). In addition, a large genet of Pisolithus tinctorius could spread vegetatively through soil over 30 m (Anderson et al. 1998). Fewer, larger genets, which were indicative of old mycelial structures that had grown from a point source over decades, were often found in mature closed forest ecosystems (Dahlberg and Stenlid 1995).
Our results suggest that proliferation by sexual spores may play a very important role in the life history of T. terreum in the artificial Larix principis-rupprechtii forest. Other factors, such as the age of the forest and the ability to form rhizomorphs for long-distance transport, may provide greater opportunity for large genet production. This feature was found in the species of the Armillaria, Cortinarius, Pisolithus, and Suillus.
Because the sporocarps of T. terreum were small and often aggregated, we only collected one representative sporocarp within a circle of 0.5 m in diameter (Bonello et al. 1998). However, as the results indicate that each sporocarp represents one genet, it is highly probable that different genotypes still exist within the circle of 0.5 m in diameter. Therefore, the minimum genet size of the ECM fungus T. terreum should be investigated in future.
References
Anderson IC, Chambers SM, Cairney JWG (1998) Use of molecular methods to estimate the size and distribution of mycelial individuals of the ectomycorrhizal basidiomycete Pisolithus tinctorius. Mycol Res 102:295–300
Baar J, Ozinga WA, Kuyper THW (1994) Spatial distribution of Laccaria bicolor genets reflected by sporocarps after removal of litter and humus layers in a Pinus sylvestris forest. Mycol Res 98:726–728
Bastide PY de la, Kropp BR, Piche Y (1994) Spatial distribution and temporal persistence of discrete genotypes of the ectomycorrhizal fungus Laccaria bicolor (Maire) Orton. New Phytol 127:547–556
Bonello P, Bruns TD, Gardes M (1998) Genetic structure of a natural population of the ectomycorrhizal fungus Suillus pungens. New Phytol 138:533–542
Bruns T, Tan J, Bidartondo M, Szaro T, Redecker D (2002) Survival of Suillus pungens and Amanita francheti ectomycorrhizal genets was rare or absent after a stand-replacing wildfire. New Phytol 155:517–523
Calvo-Bado LA, Challen MP, Thurston CF, Elliott TJ (2001) RAPD characterisation of heterogeneity in spore progenies and sexuality in the genus Agaricus. Mycol Res 105:370–376
Chalmers KJ, Waugh R, Sprent JI (1992) Detection of genetic variation between and within populations of Gliricidia sepium and G. maculata using RAPD makers. Heredity 69:465–472
Colpaert JV, Vandenkoornhuyse P, Adrlaensen K, Vangronsveld J (2000) Genetic variation and heavy metal tolerance in the ectomycorrhizal basidiomycete Suillus luteus. New Phytol 147:367–379
Dahlberg A (1997) Population ecology of Suillus variegatus in old Swedish Scots pine forests. Mycol Res 101:47–54
Dahlberg A (2001) Community ecology of ectomycorrhizal fungi: an advancing interdisciplinary field. New Phytol 150:555–562
Dahlberg A, Stenlid J (1990) Population structure and dynamics in Suillus bovinus as indicated by spatial distribution of fungal clones. New Phytol 115:487–494
Dahlberg A, Stenlid J (1994) Size, distribution and biomass of genets in populations of Suillus bovinus (L.:Fr.) Roussel revealed by somatic incompatibility. New Phytol 128:225–234
Dahlberg A, Stenlid J (1995) Spatiotemporal patterns in ectomycorrhizal populations. Can J Bot 73:S1222-S1230
Egger KN (1995) Molecular analysis of ectomycorrhizal fungal communities. Can J Bot 73:S1415-S1422
Fiore-Donno AM, Martin F (2001) Populations of ectomycorrhizal Laccaria amethystina and Xerocomus spp. show contrasting colonization patterns in a mixed forest. New Phytol 152:533–542
Gherbi H, Delaruelle C, Selosse MA, Martin F (1999) High genetic diversity in a population of the ectomycorrhizal basidiomycete Laccaria amethystina in a 150-year-old beech forest. Mol Ecol 8:2003–2013
Gryta H, Debaud JC, Effosse A, Gay G, Marmeisse R (1997) Fine-scale structure of populations of the ectomycorrhizal fungus Hebeloma cylindrosporum in coastal sand dune forest ecosystems. Mol Ecol 6:353–364
Gryta H, Debaud JC, Marmeisse R (2000) Population dynamics of the symbiotic mushroom Hebeloma cylindrosporum: mycelial persistence and inbreeding. Heredity 84:294–302
Guo LD, Hyde KD, Liew ECY (2000) Identification of endophytic fungi from Livistona chinensis (Palmae) using morphological and molecular techniques. New Phytol 147:617–630
Ito Y, Yanagi SO (1999) Discrimination of basidiomycete species and strains by random amplified polymorphic DNA (RAPD) analysis. Jpn Agric Res Q 33:149–154
Jacobson KM, Miller OK Jr, Turner BJ (1993) Randomly amplified polymorphic DNA markers are superior to somatic incompatibility tests for discriminating genotypes in natural populations of the ectomycorrhizal fungus Suillus granulatus. Proc Natl Acad Sci USA 90:9159–9163
Jany JL, Garbaye J, Martin F (2002) Cenococcum goephilum populations show a high degree of genetic diversity in beech forests. New Phytol 154:651–659
Mao XL (ed) (2000) The macrofungi in China. Henan Science and Technology, Zhengzhou
Marcais B, Martin F, Delatour C (1998) Structure of Collybia fusipes populations in two infected oak stands. Mycol Res 102:361-367
Molina R, Massicotte H, Trappe JM (1992) Specificity phenomena in mycorrhizal symbioses: community-ecological consequences. In: Allen MF (ed) Mycorrhizal functioning and integrative plant--fungal process. Chapman and Hall, New York, pp 357–423
Morris PF, Connolly MS, St Clair DA (2000) Genetic diversity of Alternaria alternata isolated from tomato in California assessed using RAPDs. Mycol Res 104:286–292
Pei MH, Whelan MJ, Halford NG, Royle DJ (1997) Distinction between stem- and leaf-infecting forms of Melampsora rust on Salix viminalis using RAPD markers. Mycol Res 101:7–10
Phillips R (ed) (1981) Mushrooms and other fungi of Great Britain and Europe. Toppan, Hong Kong
Redecker D, Szaro TM, Bowman RJ, Bruns TD (2001) Small genets of Lactarius xanthogalactus, Russula cremoricolor and Amanita francheti in late-stage ectomycorrhizal successions. Mol Ecol 10:1025–1034
Rohlf FJ (ed) (1992) NTSYS-pc numerical taxonomy and multivariate analysis system, 1.70. Exeter Software, New York
SAS Institute (1990) SAS/STAT user's guide, version 6, 4th edn. SAS Institute, Cary, N.C.
Sawyer NA, Chambers SM, Cairney JWG (1999) Molecular investigation of genet distribution and genetic variation of Cortinarius rotundisporus in eastern Australian sclerophyll forests. New Phytol 142:561–568
Selosse MA, Jacquot D, Bouchard D, Martin F, Le Tacon F (1998) Temporal persistence and spatial distribution of an American inoculant strain of the ectomycorrhizal basidiomycete Laccaria bicolor in a French forest plantation. Mol Ecol 7:561–573
Selosse MA, Martin F, Le Tacon F (1999) Structure and dynamics of experimentally introduced and naturally occurring Laccaria sp. discrete genotypes in a Douglas fir plantation. Appl Environ Microbiol 65:2006–2014
Smith ML, Bruhn JN, Anderson JB (1992) The fungus Armillaria bulbosa is among the largest and oldest living organisms. Nature 356:428–431
Smith SE, Read DJ (eds) (1997) Mycorrhizal symbiosis, 2nd edn. Academic Press, London
Sneath PHA, Sokal RR (1973) Numerical taxonomy. Freeman, San Francisco, Calif.
Weber J, Díez J, Selosse MA, Tagu D, Le Tacon F (2002) SCAR markers to detect mycorrhizas of an American Laccaria bicolor strain inoculated in European Douglas-fir plantations. Mycorrhiza 12:19–27
Williams JGK, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acid Res 18:6531–6535
Zhou Z, Miwa M, Hogetsu T (1999) Analysis of genetic structure of a Suillus grevillei population in a Larix kaempferi stand by polymorphism of inter-simple sequence repeat (ISSR). New Phytol 144:55–63
Zhou Z, Miwa M, Hogetsu T (2001) Polymorphism of simple sequence repeats reveals gene flow within and between ectomycorrhizal Suillus grevillei populations. New Phytol 149:339–348
Acknowledgements
We are grateful to Dr R. M. Miller, U.S. Department of Energy, University of Chicago for his comment on the manuscript, and would like to thank Mr G. R. Huang and the staff of Beijing Forest Ecosystem Research Station of the Chinese Academy of Sciences for their help in sampling. This work was supported by a State Key Basic Research and Development Plan of China grant (G2000046802), National Natural Science Foundation of China grants (30070018, 30230020, 39893360), and Chinese Academy of Sciences grant (KSCX2-SW-101C).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huai, WX., Guo, LD. & He, W. Genetic diversity of an ectomycorrhizal fungus Tricholoma terreum in a Larix principis-rupprechtii stand assessed using random amplified polymorphic DNA. Mycorrhiza 13, 265–270 (2003). https://doi.org/10.1007/s00572-003-0227-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-003-0227-8