Abstract
Background
Several types of cancers, including hepatocellular carcinoma (HCC), show resistance to hypoxia and nutrient starvation. Autophagy is a means of providing macromolecules for energy generation under such stressed-conditions. The aim of this study was to clarify the role of autophagy in HCC development under hypoxic conditions.
Methods
The expression of microtubule-associated protein 1 light chain 3 (LC3), which is a key gene involved in autophagosome formation, was evaluated in human HCC using immunohistochemistry and western blot. The relationship between LC3 and hypoxia-induced factor 1α (HIF1α) expression was examined using real-time PCR. In addition, human HCC cell line Huh7 was treated with pharmacological autophagy-inhibitor and inactive mutant of Atg4B (Atg4BC74A) under hypoxic condition to evaluate the effects of hypoxia-induced autophagy on cell survival, intracellular ATP, and mitochondrial β-oxidation.
Results
LC3 was significantly highly expressed in HCC as compared with noncancerous tissues. LC3 expression, correlated with HIF1α expression, was also significantly correlated with tumor size, and only in the context of large tumors, was an independent predictor of HCC recurrence after surgery. In addition, Huh7 treated with autophagy-inhibitor under hypoxia had lower viability, with low levels of intracellular ATP due to impaired mitochondrial β-oxidation.
Conclusions
Autophagy in HCC works to promote HIF1α-mediated proliferation through the maintenance of intracellular ATP, depending on the activation of mitochondrial β-oxidation. These findings demonstrated the feasibility of anti-autophagic treatment as a potential curative therapy for HCC, and improved understanding of the factors determining adaptive metabolic responses to hypoxic conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is common and increasing in incidence worldwide [1–4]. HCC grows to a relatively large size, sometimes over 10 cm in diameter even when necrosis was observed [5] and it can easily reoccur after therapy [6]. Proliferating cancer cells in tumors growing to a large size require a good supply of nutrients and oxygen. Angiogenesis around tumors is one way of increasing blood flow to provide the required oxygen and energy to the growing tumor [7]. However, recent studies have revealed that nutrition levels, oxygen and glucose are frequently reduced in locally advanced tumors despite tumor vessels having been established [8, 9]. This suggests that the microvasculature around the tumor is structurally and functionally abnormal and not capable of supplying the blood flow needed for cancer cell growth. Furthermore, some aggressive malignant tumors, such as poorly differentiated HCC and pancreatic cancers, are clinically hypovascular [10]. Under these conditions, cancer cells are likely to encounter limited nutrients and oxygen. However, they can exhibit resistance to nutrient deprivation and continue to grow. The mechanisms by which cancer cells obtain energy sources when their external nutrient supply is limited remain is unclear.
Autophagy is a homeostatic mechanism that regulates the turnover of long-lived or damaged proteins and organelles, buffers intracellular constituents and supplies amino acids taken from degradation products of the autolysosome [11]. For the first step, the isolation membrane, a lipid bilayer structure, is developed and sequesters cytoplasmic materials such as organelles, to form autophagosomes. During this step, microtubule-associated protein 1 light chain 3 (LC3), one of the mammalian homologues of yeast autophagy-related gene (Atg) 8, is processed and activated by a ubiquitination-like reaction regulated by Atg7 and Atg3 [12]. First, LC3 proform is cleaved into a soluble form known as LC3-I, which is further modified into a membrane-bound form, LC3-II, and this is followed by recruitment into the autophagosomes. Thus, LC3 is a specific marker of autophagosome formation. Autophagosomes engulf organelles and then fuse with lysosomes to become mature autolysosomes. Accordingly, sequestrated materials are digested into amino acids in the autolysosomes by the lysosomal enzymes [13, 14].
To study the role of autophagy in HCC under hypoxia-induced metabolic stress, we examined LC3 expression as the main marker of autophagosomes, and the other corresponding autophagic genes, Atg5 and Beclin-1, in human tissue samples and HCC cell lines under hypoxic conditions. Our results suggest that high expression of autophagy has the potential to cause malignant tumors to grow in size under hypoxic condition and also promotes poor survival, which can be independently predicted by the autophagic gene LC3 in HCC.
Materials and methods
Human tissue samples
Samples from 102 patients who had undergone liver resection for HCC without preoperative treatment at the Department of Surgery and Science at the Kyushu University Hospital between January 1986 and December 2002 were analyzed using immunohistochemistry [15]. Samples from another 131 patients between January 2004 and March 2009 were analyzed by real-time polymerase chain reaction (PCR). There were no significant differences between the characteristics of HCC patients using immunohistochemistry analysis or real-time PCR analysis (Table S1). Details in Doc. S1.
Reagents and plasmid
3-Methyladenine (3MA) and an inactive mutant of Atg4B (Atg4BC74A) were prepared as described previously [15, 16]. Details in Doc. S1.
Immunohistochemistry and immunofluorescence
Immunohistochemical staining and immunofluorescence analysis was performed as previously described [17–20]. Immunoreactivity of cytoplasmic staining in the cancerous region was independently divided into two groups, positive and negative, by two liver pathologists. Positive staining was classified if even a small area of tissue was stained. Details in Doc. S1.
Protein extraction and western blot analysis
Protein extraction and western blot analysis were performed as previously described [21]. Details in Doc. S1.
Real-time PCR
Extraction of total RNA and real-time PCR was performed as previously described [22]. Primers used for real-time PCR are shown in Table S2 and details described in Doc. S1.
Electron microscopy
Analysis of electron microscopy was performed as previously described [20, 23, 24]. For quantification of autophagosome using electron micrographs, high-powered micrographs (×8000–10000) of 10 single cells from multiple distinct low-powered fields were obtained from each specimen [23]. Details in Doc. S1.
Cell culture under hypoxic conditions
For hypoxia treatments, human HCC cell lines were incubated in a humidified hypoxic workstation (MCO-5M, Sanyo, Osaka, Japan) with final oxygen concentrations of 0.1 % O2 using a Clark-type polarographic electrode (Animas, Frazer, PA, USA). To defect autophagosome formation, cultured cells were treated with autophagy-inhibitor using two methods, 3MA and Atg4BC74A, and pre-incubated for 24 h followed by incubation under hypoxia. Details in Doc. S1.
Quantification of intracellular ATP
Total cellular ATP concentration was quantitated using an ATP Detection Reagent kit (Toyo-ink, Tokyo, Japan) as previously described [25]. Details in Doc. S1.
Cytofluorimetric analysis of ∆Ψ m and mitochondrial structure
Cytofluorimetric analysis of ∆Ψ m and alteration of mitochondrial structure during hypoxia was performed as previously described [26, 27]. Details in Doc. S1.
Measurement of β-hydroxybutyrate
β-hydroxybutyrate concentration was spectrophotometrically assayed as previously described [28]. Details in Doc. S1.
Statistical analysis
All statistical analyses were performed using JMP statistical software version 7.01 (SAS Institute Inc., Cary, NC, USA). All experiments were independently performed three times in triplicate. All variables are expressed as the mean ± standard deviation (SD). Details in Doc. S1.
Results
Upregulation of LC3 expression and autophagy activation in HCC
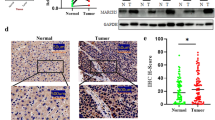
In the cancer cells of 102 HCC resected tissues, LC3 immunoreactivity was observed in the cytoplasm in 49 of 102 specimens (Fig. 1a). LC3 immunoreactivity was not observed in noncancerous lesions in any specimens. Western blot analysis using a limited set of tissue samples consisting of four matched noncancerous and HCC tissues revealed that expression of LC3-II, a well-known marker of activated autophagy, was detected in the cancerous tissue of case 3 and 4, in which LC3 immunoreactivity was immunohistochemically observed in the cancer cells (Fig. 1b). p62 regulates ubiquitin-positive protein aggregates caused by autophagic deficiency. Considering the results that the expression of p62 were higher in Case 1 and 2 than in Case 3 and 4 both at the cancerous and noncancerous tissues, it also indicated that the autophagic activity were impaired even in the noncancerous tissue and highly impaired in cancerous lesions in the Case 1 and 2.
Upregulation of LC3 protein in HCC human tissue samples (a, b) and LC3 and HIF1α expression associated with HCC tumor size (c–f). a LC3 immunoreactivity was observed in the cytoplasm of cancer cells, and hepatocytes in non-cancerous tissues were negative for LC3. b Western blot analysis of LC3 expression in the four matched HCC tissues and noncancerous tissues. Expression of LC3-II was detected in cancerous tissue in case 3 and 4, but no or only a faint signal was detected in non-cancerous tissue all of the cases. The expression of p62 was higher in case 1 and 2 than in case 3 and 4. c–f LC3 and HIF1α expression were evaluated by real-time PCR analysis of human HCC samples on the three cohorts: c, d the total patient population (n = 131); e, f patients with tumor sizes of ≥3 cm (n = 65); and g, h patients with tumor sizes of <3 cm (n = 66). They were also divided into two groups consisting of those with high and low expression of HIF1α with a cutoff value with a median of 2.85. LC3 (c) and HIF1α (d) expression was significantly correlated with tumor size (LC3, P = 0.0131; HIF1α, P = 0.0089). e, f In the group with a tumor size of ≥3 cm, LC3 expression in the high HIF1α expression group was significantly greater than that in low HIF1α expression group (P = 0.0097) and LC3 expression in the positive recurrence group was significantly higher than in the negative recurrence group (P = 0.0088). g, h In the group with a tumor size of <3 cm, there was no significant difference in LC3 expression between the high and low HIF1α expression groups, and between positive and negative recurrence groups. HCC, hepatocellular carcinoma; HIF1α, hypoxia-induced factor 1α; LC3, microtubule-associated protein 1 light chain 3
Relationship between LC3 expression and clinicopathological factors
The positive LC3 expression group had significantly higher serum tumor markers, AFP (P = 0.0467) and DCP (P = 0.0454), tumor size (P = 0.0006) and positive rates of portal vein invasion (P = 0.0001) than the negative LC3 expression group (Table 1). Unexpectedly, there were no correlation between the LC3 expression and viral infection. Some patients with HCV or HBV underwent the each pharmacological therapy such as interferon or nucleotide analog, therefore, the activities of hepatitis were difficult to be assessed accurately. That might be the reason why the autophagic activity was necessarily correlated with the presence of the hepatitis clinically.
Association of LC3 with HIF1α expression and HCC recurrence according to HCC tumor size
LC3 (Fig. 1c, P = 0.0131) and HIF1α (Fig. 1d, P = 0.0089) expression was correlated with tumor size. In the group with a tumor size of ≥3 cm (Fig. 1e), LC3 expression in the high HIF1α expression group was greater than in the low HIF1α expression group (P = 0.0097). In any case, LC3 expression in the positive recurrence group was higher than in the negative recurrence group (Fig. 1f, P = 0.0088), whereas, in the group with a tumor size of <3 cm, there was no difference in LC3 expression between the high- and low-HIF1α expression groups (Fig. 1g, h). These indicated that only in the case of larger tumors, in which hypoxia and poor nutritional conditions were indicated, the autophagic gene LC3 was highly expressed and that autophagy enhanced tumor growth and promoted HCC malignancy.
Association of high expression of LC3 with tumor size and poor HCC prognosis
For the entire patient group (n = 131), the disease-free survival rates of the positive LC3 group (49.0 % at 3 years and 34.7 % at 5 years) was lower than for the negative LC3 group (60.4 % at 3 years and 46.8 % at 5 years) (P = 0.0056) (Fig. 2a). In patients with a tumor size of ≥3 cm (n = 65), disease-free survival in the positive LC3 group (48.4 % at 3 years and 32.3 % at 5 years) was lower than in the negative LC3 group (82.6 % at 3 years and 59.8 % at 5 years) (P = 0.0054) (Fig. 2b). In contrast, in patients with a tumor size of <3 cm (n = 66), there was no difference (Fig. 2c).
Association between LC3 expression and patients’ prognosis depending on tumor size of HCC. The disease-free survival after surgery were compared between the positive and negative LC3 expression groups by immunohistochemical analysis. The analysis was performed on three cohorts: a the total patient group (n = 102); b patients with tumor sizes of ≥3 cm (n = 54); and c patients with tumor sizes of <3 cm (n = 48). a For the entire patient group, the disease-free survival rates of the positive LC3 group (49.0 % at 3 years and 34.7 % at 5 years) was significantly lower than that of the negative LC3 group (60.4 % at 3 years and 46.8 % at 5 years) (P = 0.0056). b In the patients with a tumor size of ≥3 cm, disease-free survival in the positive LC3 group (48.4 % at 3 years and 32.3 % at 5 years) was significantly lower than in the negative LC3 group (82.6 % at 3 years and 59.8 % at 5 years) (P = 0.0054). c In patients with a tumor size of <3 cm there was no significant difference between the two groups. HCC hepatocellular carcinoma, LC3 microtubule-associated protein 1 light chain 3
Prognostic value of LC3 expression in HCC patients with a tumor size of ≥3 cm
The prognostic factors were evaluated in patients with a tumor size of ≥3 cm using univariate analysis (Table S3), which showed that three parameters namely, ICG R15 (P = 0.0001), multiple tumors (P = 0.0416) and portal vein invasion (P = 0.0280) were predictors of HCC recurrence. Positive LC3 expression was also a predictor of tumor recurrence (P = 0.0103). Furthermore, multivariate analysis was conducted with four of the variables (positive LC3 expression, ICG R15 ≥ 14.5 % as median values, multiple tumors and portal vein invasion), and there was no obvious correlation between them (Table 2). Positive LC3 expression was still the independent variable for predicting poor disease free survival (P = 0.0065).
Upregulation of LC3 expression in HCC cell lines under hypoxic conditions
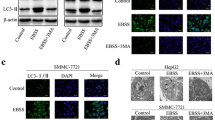
Hypoxia markedly increased the number of autophagic vacuoles in Huh7 cells, which appeared as dot-like signals by immunofluorescence analysis (Fig. 3a, b) and electron microscopy (Fig. 3c, d). Expression of LC3-II increased in a time dependent manner (Fig. 3e). These initial observations indicated that HCC cell lines had the potential power to express autophagic activity in response to hypoxic conditions.
Upregulation of LC3 protein in HCC cell lines under hypoxic conditions. a, b Huh7 cells exposed to 2 h hypoxic conditions had markedly increased numbers of autophagic vacuoles, which appeared as dot-like signals by immunofluorescence analysis using antibody against LC3. c, d Extensive autophagosome formation (arrowed) in Huh7 cells exposed to 2 h hypoxic conditions was monitored using electron microscopy. e The expression of LC3-II increased in a time-dependent manner when evaluated by western blot analysis using the proteins extracted from Huh7 cells under hypoxic conditions for the indicated times. The expression of the other autophagic genes, Beclin-1 and Atg5, also increased in a time-dependent manner. Atg Autophagy-related genes, HCC hepatocellular carcinoma, LC3 microtubule-associated protein 1 light chain 3
Suppression of autophagy and proliferation of HCC cells under hypoxic conditions
LC3 expression was suppressed using two methods namely, pharmacological inhibition by means of 3MA and transfection of Atg4BC74A, which involved type III PI3K inhibition and hampering of the conversion of LC3-I to LC3-II, respectively (Fig. S1). Subsequently, the down-regulation of autophagy under hypoxia inhibited the proliferation of HCC cells (Fig. 4a). In addition, the growth of Huh7 cells transfected with GFP-LC3 was higher than the cells receiving no treatment under hypoxic condition (Fig. 4a). These findings indicated that hypoxia-induced autophagy in HCC cells works to promote cell proliferation through preventing accumulation of damaged protein and organelles.
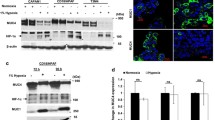
Impaired proliferation of HCC cell lines treated with autophagy-inhibitor and deterioration of intracellular ATP maintenance through impairment of mitochondrial β-oxidation due to suppressed autophagy activity. Huh7 cells receiving no treatment or treated with autophagy-inhibitor using two methods, pharmacological inhibitor as 3MA and transfection of Atg4BC74A, were incubated under hypoxic conditions of 0.1 % O2 final concentration for 2 h. a Growth of both Huh7 cells treated using the two methods of autophagy-inhibition were significantly lower than that of the cells receiving no treatment under hypoxic conditions. In addition, the growth of Huh7 cells transfected with GFP-LC3 was higher than the cells receiving no treatment under hypoxic condition. b The intracellular ATP in the Huh7 cells treated with autophagy-inhibitor under hypoxic conditions was present at significantly lower levels than in cells receiving no treatment. c The proportion of Huh7 cells with low levels of mitochondrial membrane permeability treated with autophagy-inhibitor was higher than that of non-treated cells by by FACScan analysis using JC-1 antibody. d The gene expression related to mitochondrial β-oxidation in Huh7 cells were examined by real-time PCR. The levels of the enzymes MCAD and CPT, L-FABP, and FATP in Huh7 cells treated with autophagy-inhibitor were significantly lower than that in non-treated cells. e The intracellular β-hydroxybutyrate levels in Huh7 cells treated with autophagy-inhibitor under hypoxic conditions were significantly lower than in cells receiving no treatment. ACC Acetyl-CoA carboxylase, ATP adenosine 5′-triphosphate, CPT carnitine palmitoyltransferase, FAS fatty acid synthase, FAT fatty acid translocase, FATP fatty acid transport protein, L-FABP fatty acid binding protein, LC3 microtubule-associated protein 1 light chain 3, MCAD medium-chain acyl-CoA dehydrogenase, PCR polymerase chain reaction, 3MA 3-methyladenine
Suppression of autophagy activity and mitochondrial β-oxidation
The levels of intracellular ATP in autophagy-inhibited HCC cells were lower than in non-treated cells (Fig. 4b). The proportion of HCC cells with low mitochondria membrane permeability treated with autophagy-inhibitor was higher (Fig. 4c). Gene expression levels of MCAD and CPT, L-FABP, and FATP enzymes in autophagy-inhibited cells, which were indicated to have a rate-controlling effect on β-oxidation, the transportation of free fatty acids to mitochondria, and the transport of free fatty acids into hepatocytes, respectively, were significantly lower than those in non-treated cells (Fig. 4d). Consequently, β-hydroxybutyrate levels, final ketone body product, in autophagy-inhibited cells were decreased more than in non-treated cells (Fig. 4e). These findings indicated that HCC cells exposed to hypoxia had the potential power to maintain intracellular ATP through the activation of mitochondrial β-oxidation, which may have been due to the prompt removal of the damaged mitochondria by activated autophagy as a selective degradation system.
Discussion
In the present study, the inalterable role of autophagy in human HCC exposed to hypoxic conditions as tumors grew in size was demonstrated. The LC3 expression was correlated with tumor size, and only in large tumors, was correlated with the expression of HIF1α, hypoxia and the under nutrition marker. Additionally, high expression of LC3 was shown to be an independent predictor of HCC recurrence. Further, analysis of HCC cell lines using autophagy-inhibitor revealed that the hypoxia-induced autophagy in HCC cells worked to promote cell proliferation through maintenance of intracellular ATP. This depended on the activation of mitochondrial β-oxidation with the prompt removal of damaged mitochondria due to activated autophagy. This is the first report demonstrating the mechanism involved in maintaining the intracellular energy sources by means of activated autophagy, as HCC tumors developed in large size under hypoxic stress. Recently some reports demonstrated that the emerging role of autophagy for promoting cell viability in HCC progressions during ischemia-hypoxia condition only in the rodent models [29, 30]. We also demonstrated these results even in human samples clinically and emphasized that ‘only in the large tumors’ in which the impact of hypoxia and poor conditions were involved, the autophagic activity was highly expressed and its enhancement for the tumor growth and promotion of HCC malignancy, therefore, the newly targeted therapy for autophagy pathway of these adaptive metabolic responses is desired to becomes major challenges to overcome the large sized HCC tumors.
The role of autophagy in cell fate decision remains controversial. Autophagy is claimed to be an indispensable physiological reaction that sustains cell viability under nutrient-starved conditions [31]. Regarding cancer cells, it has been reported that autophagy was highly expressed in many of these cells and that this high expression was a strong factor related to tumor progression [32]. However, autophagy has recently attracted attention in connection with programmed or autophagic cell death. Colell et al. [33] demonstrated that cell death resulting from progressive cellular consumption can be attributed to unrestrained autophagy, which has led to the belief that autophagy is a nonapoptotic form of programmed cell death [34, 35]. However, the role of autophagy, an alternative caspase-independent cell death program, and its underlying molecular mechanism, is still controversial in cancer, especially in tumor progression. About the mTOR pathway, mTOR inhibitors such as RAD001, also an autophagy inducer, does not promote the proliferation of HCC cells and results in cell death [36, 37]. Weiner et al. [38], demonstrated that autophagy is observed in established cancers, but its inhibition during early carcinogenesis actually promotes tumor progression, suggesting that an autophagic switch promotes the transition of a tumor into a state of so-called autophagy addiction in order to maintain viability in hypoxic, nutrient-limited microenvironments. They also discussed that the autophagic function depends on the extent to which cells are capable of enhancing basal levels of autophagy. In fact, the hyperdynamic state of autophagy might be the crisis of cellular life because of its consumption of the intracellular even healthy organelle and proteins. Considering the reports of mTOR inhibitors resulted in HCC cell death, the basal levels of autophagy in these setting of HCC cells and whether its inhibitor might excessively induce the autophagic activity or not should be carefully assessed because mTOR is a strong key component in a series of pathways involved in tumor growth and development. We revealed the protective role played by autophagy that involves proliferation of HCC cells due to activation of mitochondrial β-oxidation. Furthermore, we investigated the clinical significance of this finding regarding the link between autophagy and tumor progression. Three HIF1α-dependent molecular mechanisms have been reported by which cells adapt their energy metabolism under hypoxic conditions: inhibition of mitochondrial biogenesis by repression of c-Myc activity [39]; inhibition of acetyl-CoA synthesis by activation of PDK1 [40], and COX4 subunit switching [41]. We demonstrated that protection of mitochondria by autophagy is a fourth component of the HIF1α-mediated metabolic adaptation required to prevent cell death and damaged mitochondria, due to increased ROS levels in HCC cells under low nutrient conditions.
Ding et al. [42] demonstrated that only in an apoptosis compromised background, the expression of the autophagic gene, Beclin 1, and their corresponding autophagic activities were suppressed in HCC. They indicated that the loss of a survival pathway, autophagy, enhanced tumor growth by promoting genome damage and instability in an apoptosis-deficient background, a Bcl-xL positive background. However, their data depended on the assessment of the expression of Beclin-1 and there were multiple molecular machineries involved in the formation of the autophagosome downstream of Beclin-1. Therefore, it was not established whether autophagy was truly activated [39]. Recently, some reports demonstrated Beclin-1 independent autophagy, which acted as a caspase-independent cell death mechanism [43–45]. Additionally, Beclin-1 is the multifunctional gene involved in apoptosis [46, 47]. Thus, Beclin-1 plays a key role in autophagy, however, Beclin-1 should be carefully analyzed for the assessment of activated autophagy. The standard method for assessing autophagic activity is the demonstration of autophagic vesicles using electron microscopy, [48] and another method is the LC3-based assay. When autophagy occurs, conversion of a fraction of the cytosolic form of LC3-I to the autophagic membrane form of LC3-II can be detected using western blots of LC3 proteins. This change in intracellular localization of LC3 protein is considered to correlate with autophagic activity [49], and LC3 expression detected using immunohistochemistry represents the steady-state LC3 level including both LC3-I and LC3-II. We examined activated autophagy promptly with LC3-II using western blotting and confirmed the high expression of autophagosomes using an electron microscope.
Regarding the energy supply system in surviving cancer cells, several cancer cell lines including pancreatic cancer-derived and colorectal cancer-derived cell lines, are resistant to nutrient-deprived culture conditions [50]. These cells might use some alternative metabolic process to obtain energy for survival. However, untransformed human fibroblasts were completely abolished under the same conditions [51]. These indicated that this phenomenon of starvation-resistance may contribute to survival of cancer cells in nutrient-deficient microenvironments. As an alternative energy source, mammalian cells can use amino acids [52]. If the starvation-resistant cancer cells can raise amino acids from the inside of cells, these amino acids become potential energy sources. Autophagy is an evolutionarily conserved process involving lysosomal degradation of cytoplasmic and cellular organelles [11, 26]. Autophagy constitutes a stress adaptation that avoids tumor death and has roles in protecting cells against the shortage of nutrients [26, 53]. Furthermore, autophagy is a catabolic process by which cells supply amino acids from self-digested organelles, as an alternative energy source for survival [11, 13, 31]. This function of autophagy seems ideal in fostering the survival of cancer cells in an unfavorable starved microenvironment. We demonstrated that autophagy plays an essential role by maintaining an optimal balance between the competing demands of energy and proliferation under hypoxic conditions. Whereas consideration of cellular energetics favors oxidative metabolism as the most efficient means of producing adequate levels of ATP to maintain cell survival, mitochondrial respiration is also associated with increased amino acid consumption due to autophagy. It was reasonable that the two aspects of HIF1α mediated-autophagic function, namely the protection of mitochondria and the supply amino acids from inside the cell contributed to cell survival in response to hypoxia.
For the recurrence of HCC after surgery, high expression of LC3 is the independent predictive factor in the context of larger sized tumors. Other predictive factors were liver dysfunction defined as ICG R15 < 14.5 %, multiple tumors and portal vein invasion. As HCC tumors grow, it becomes difficult to secure an exogenous energy supply, in particular for the central nest of tumors. However, in small HCC tumors with adequate oxygen supply and enriched nutrient conditions, cells did not struggle to obtain energy from an exogenous nutrient supply. For this reason, in patients with tumors of <3 cm, there was no correlation between LC3 expression, and HIF1α expression and tumor size. However, in patients with tumors of ≥3 cm, LC3 expression was significantly associated with HIF1α expression. To curatively treat HCC, an effective approach may be not only to inhibit the autophagy machinery, but also the HIF1α machinery molecule to prevent energy supply in conditions in which exogenous nutrient supply is extremely limited.
We have provided experimental evidence supporting the conclusion that HIF1α-mediated autophagy is involved in mitochondrial metabolism and is essential to understanding the mechanisms and consequences of the maintenance of intracellular energy in cancer cells under hypoxic conditions. This process requires the HIF1α-dependent induction of autophagy machinery as demonstrated by means of LC3-deficient HCC cell analysis. Understanding the factors determining which of these adaptive metabolic responses to conditions of hypoxia and under nutrition in HCC cells are utilized in vitro and in vivo, and whether these adaptations are successful in preventing cell death, remains a challenge.
Abbreviations
- AFP:
-
Alpha-fetoprotein
- Atg:
-
Autophagy-related genes
- ATP:
-
Adenosine 5′-triphosphate
- DCP:
-
Des-gamma-carboxy prothrombin
- HCC:
-
Hepatocellular carcinoma
- HIF1α:
-
Hypoxia-induced factor 1α
- ICG R15:
-
Indocyanine green retention test at 15 min
- LC3:
-
Microtubule-associated protein 1 light chain 3
- PBS:
-
Phosphate-buffered saline
- PCR:
-
Polymerase chain reaction
- PI3K:
-
Phosphatidylinositol 3-kinase
- ROS:
-
Reactive oxygen species
- SD:
-
Standard deviation
- 3MA:
-
3-Methyladenine
References
Shirabe K, Toshima T, Taketomi A, et al. Hepatic aflatoxin B1-DNA adducts and TP53 mutations in patients with hepatocellular carcinoma despite low exposure to aflatoxin B1 in southern Japan. Liver Int. 2011;31:1366–72.
Shirabe K, Itoh S, Yoshizumi T, et al. The predictors of microvascular invasion in candidates for liver transplantation with hepatocellular carcinoma-with special reference to the serum levels of des-gamma-carboxy prothrombin. J Surg Oncol. 2007;95:235–40.
Shirabe K, Kajiyama K, Harimoto N, Tsujita E, Wakiyama S, Maehara Y. Risk factors for massive bleeding during major hepatectomy. World J Surg. 2010;34:1555–62.
Bruix J, Llovet JM. Prognostic prediction and treatment strategy in hepatocellular carcinoma. Hepatology. 2002;35:519–24.
Yamashita Y, Taketomi A, Shirabe K, et al. Outcomes of hepatic resection for huge hepatocellular carcinoma (≥10 cm in diameter). J Surg Oncol. 2011;104:292–8.
Taketomi A, Sanefuji K, Soejima Y, et al. Impact of des-gamma-carboxy prothrombin and tumor size on the recurrence of hepatocellular carcinoma after living donor liver transplantation. Transplantation. 2009;87:531–7.
Vaupel P, Thews O, Hoechkel M. Treatment resistance of solid tumors: role of hypoxia and anemia. Med Oncol. 2001;18:243–59.
Jain RK. Molecular regulation of vessel maturation. Nat Med. 2003;9:685–93.
Harris AL. Hypoxia—a key regulatory factor in tumour growth. Nat Rev Cancer. 2002;2:38–47.
Kitano M, Kudo M, Maekawa K, et al. Dynamic imaging of pancreatic diseases by contrast enhanced coded phase inversion harmonic ultrasonography. Gut. 2004;53:854–9.
Klionsky DJ, Emr SD. Autophagy as a regulated pathway of cellular degradation. Science. 2000;290:1717–21.
Tanida I, Tanida-Miyake E, Ueno T, Kominami E. The human homolog of Saccharomyces cerevisiae Apg7p is protein-activating enzyme for multiple substrates including human Apg12p, GATE-16, GABARAP, and MAP-LC3. J Biol Chem. 2001;276:1701–6.
Mizushima N, Yoshimori T, Levine B. Methods in mammalian autophagy research. Cell. 2010;140:313–26.
Fleming A, Noda T, Yoshimori T, Rubinsztein DC. Chemical modulators of autophagy as biological probes and potential therapeutics. Nat Chem Biol. 2011;7:9–17.
Liver Cancer Study Group. The general rules for the clinical and pathological study of primary liver cancer. 7th ed. Tokyo: Kanehara Publications; 2006.
Fujita N, Hayashi-Nishino M, Fukumoto H, et al. An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol Biol Cell. 2008;19:4651–9.
Aishima S, Fujita N, Mano Y, et al. Different roles of S100P overexpression in intrahepatic cholangiocarcinoma: carcinogenesis of perihilar type and aggressive behavior of peripheral type. Am J Surg Pathol. 2011;35:590–8.
Kabeya Y, Mizushima N, Yamamoto A, Oshitani-Okamoto S, Ohsumi Y, Yoshimori T. LC3, GABARAP and GATE16 localize to autophagosomal membrane depending on form-II formation. J Cell Sci. 2004;117:2805–12.
Mizushima N, Yoshimori T, Levine B. Methods in mammalian autophagy research. Cell. 2010;140:313–26.
Klionsky DJ, Abdalla FC, Abeliovich H, Abraham RT, Acevedo-Arozena A, Adeli K, et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy. 2012;8:445–544.
Anegawa G, Kawanaka H, Yoshida D, et al. Defective endothelial nitric oxide synthase signaling is mediated by rho-kinase activation in rats with secondary biliary cirrhosis. Hepatology. 2008;47:966–77.
Sadagurski M, Cheng Z, Rozzo A, et al. IRS2 increases mitochondrial dysfunction and oxidative stress in a mouse model of Huntington disease. J Clin Invest. 2011;121:4070–81.
Nanjundan M, Nakayama Y, Cheng KW, et al. Amplification of MDS1/EVI1 and EVI1, located in the 3q26.2 amplicon, is associated with favorable patient prognosis in ovarian cancer. Cancer Res. 2007;67:3074–84.
Klionsky DJ, Abeliovich H, Agostinis P, Agrawal DK, Aliev G, Askew DS, et al. Guidelines for the use and interpretation of assays for monitoring autophagy in higher eukaryotes. Autophagy. 2008;4:151–75.
Kasahara A, Ishikawa K, Yamaoka M, et al. Generation of trans-mitochondrial mice carrying homoplasmic mtDNAs with a missense mutation in a structural gene using ES cells. Hum Mol Genet. 2006;15:871–81.
Du H, Yang W, Chen L, Shen B, Peng C, Li H, et al. Emerging role of autophagy during ischemia-hypoxia and reperfusion in hepatocellular carcinoma. Int J Oncol. 2012;40:2049–57.
Chang Y, Yan W, He X, Zhang L, Li C, Huang H, et al. miR-375 inhibits autophagy and reduces viability of hepatocellular carcinoma cells under hypoxic conditions. Gastroenterology. 2012;143:177–87.
Petit PX, et al. Alterations in mitochondrial structure and function are early events of dexamethasone-induced thymocyte apoptosis. J Cell Biol. 1995;130:157–67.
Hanson GT, Aggeler R, Oglesbee D, et al. Investigating mitochondrial redox potential with redox-sensitive green fluorescent protein indicators. J Biol Chem. 2004;279:13044–53.
Aleksunes LM, Reisman SA, Yeager RL, Goedken MJ, Klaassen CD. Nuclear factor erythroid 2-related factor 2 deletion impairs glucose tolerance and exacerbates hyperglycemia in type 1 diabetic mice. J Pharmacol Exp Ther. 2010;333:140–51.
Levine B, Klionsky DJ. Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell. 2004;6:463–77.
Kuwahara Y, Oikawa T, Ochiai Y, et al. Enhancement of autophagy is a potential modality for tumors refractory to radiotherapy. Cell Death Dis. 2011;2:e177.
Colell A, Ricci JE, Tait S, et al. GAPDH and autophagy preserve survival after apoptotic cytochrome c release in the absence of caspase activation. Cell. 2007;129:983–7.
Kirkegaard K, Taylor MP, Jackson WT. Cellular autophagy: surrender, avoidance and subversion by microorganisms. Nat Rev Microbiol. 2004;2:301–14.
Kondo Y, Kanzawa T, Sawaya R, Kondo S. The role of autophagy in cancer development and response to therapy. Nat Rev Cancer. 2005;5:726–34.
Thomas HE, Mercer CA, Carnevalli LS, Park J, Andersen JB, Conner EA, et al. mTOR inhibitors synergize on regression, reversal of gene expression, and autophagy in hepatocellular carcinoma. Sci Transl Med. 2012;4:139ra84.
Altmeyer A, Josset E, Denis JM, Gueulette J, Slabbert J, Mutter D, Noël G, Bischoff P. The mTOR inhibitor RAD001 augments radiation-induced growth inhibition in a hepatocellular carcinoma cell line by increasing autophagy. Int J Oncol. 2012. doi:10.3892/ijo.2012.1583.
Weiner LM, Lotze MT. Tumor-cell death, autophagy, and immunity. N Engl J Med. 2012;366:1156–8.
Lu Z, Dono K, Gotoh K, et al. Participation of autophagy in the degeneration process of rat hepatocytes after transplantation following prolonged cold preservation. Arch Histol Cytol. 2005;68:71–80.
Degenhardt K, Mathew R, Beaudoin B, et al. Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell. 2006;10:51–64.
Pouyssegur J, Dayan F, Mazure NM. Hypoxia signalling in cancer and approaches to enforce tumour regression. Nature. 2006;441:437–43.
Ding ZB, Shi YH, Zhou J, et al. Association of autophagy defect with a malignant phenotype and poor prognosis of hepatocellular carcinoma. Cancer Res. 2008;68:9167–75.
Scarlatti F, Maffei R, Beau I, Codogno P, Ghidoni R. Role of non-canonical Beclin 1-independent autophagy in cell death induced by resveratrol in human breast cancer cells. Cell Death Differ. 2008;15:1318–29.
Chu CT, Zhu J, Dagda R. Beclin 1-independent pathway of damage-induced mitophagy and autophagic stress: implications for neurodegeneration and cell death. Autophagy. 2007;3:663–6.
Pickford F, Masliah E, Britschgi M, et al. The autophagy-related protein beclin 1 shows reduced expression in early Alzheimer disease and regulates amyloid beta accumulation in mice. J Clin Invest. 2008;118:2190–9.
Boya P, González-Polo RA, Casares N, et al. Inhibition of macroautophagy triggers apoptosis. Mol Cell Biol. 2005;25:1025–40.
Ait-Mohamed O, Battisti V, Joliot V, et al. Acetonic extract of Buxus sempervirens induces cell cycle arrest, apoptosis and autophagy in breast cancer cells. PLoS One. 2011;6:e24537.
Mizushima N, et al. Methods for monitoring autophagy. Int J Biochem Cell Biol. 2004;36:2491–502.
Kabeya Y, Mizushima N, Ueno T, et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 2000;19:5720–8.
Sato K, Tsuchihara K, Fujii S, et al. Autophagy is activated in colorectal cancer cells and contributes to the tolerance to nutrient deprivation. Cancer Res. 2007;67:9677–84.
Esumi H, Izuishi K, Kato K, et al. Hypoxia and nitric oxide treatment confer tolerance to glucose starvation in a 5′-AMP-activated protein kinase-dependent manner. J Biol Chem. 2002;277:32791–8.
Kuma A, Hatano M, Matsui M, et al. The role of autophagy during the early neonatal starvation period. Nature. 2004;432:1032–6.
Rouschop KM, van den Beucken T, Dubois L, et al. The unfolded protein response protects human tumor cells during hypoxia through regulation of the autophagy genes MAP1LC3B and ATG5. J Clin Invest. 2010;120:127–41.
Acknowledgments
We are grateful to T. Yoshimori (Osaka University) for kindly providing the inactive mutant of Atg4B (Atg4BC74A). We also thank N. Yamashita (Kyushu University) for her expert advice related to statistical analysis.
Conflict of interest
The authors have no conflicts of interest to declare and have no financial interests linked to this work.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Toshima, T., Shirabe, K., Matsumoto, Y. et al. Autophagy enhances hepatocellular carcinoma progression by activation of mitochondrial β-oxidation. J Gastroenterol 49, 907–916 (2014). https://doi.org/10.1007/s00535-013-0835-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-013-0835-9