Summary
Background
An increased frequency of Proteus mirabilis isolates resistant to expanded-spectrum cephalosporins was observed recently in a long-term care facility in Zagreb (Godan). The aim of this study was the molecular characterization of resistance mechanisms to new cephalosporins in P. mirabilis isolates from this nursing home.
Methods
Thirty-eight isolates collected from 2013–2015 showing reduced susceptibility to ceftazidime were investigated. Antibiotic susceptibilities were determined by broth microdilution method. Inhibitor-based tests were performed to detect extended-spectrum (ESBLs) and AmpC β-lactamases. AmpC β-lactamases were characterized by polymerase chain reaction (PCR) followed by sequencing of bla ampC genes. Quinolone resistance determinants (qnr genes) were characterized by PCR. Genotyping of the isolates was performed by repetitive element sequence (rep)-PCR and pulsed-field gel electrophoresis (PFGE).
Results
Presence of an AmpC β-lactamase was confirmed in all isolates by combined-disk test with phenylboronic acid. All isolates were resistant to amoxicillin alone and combined with clavulanate, cefotaxime, ceftriaxone, cefoxitin, and ciprofloxacin; but susceptible to cefepime, imipenem, and meropenem. PCR followed by sequencing using primers targeting bla ampc genes revealed CMY-16 β-lactamase in all but one strain. Bla cmy-16 was carried by a non-conjugative plasmid which did not belong to any known plasmid-based replicon typing (PBRT) group. Rep-PCR identified one large clone consisting of 15 isolates, three pairs or related isolates, one triplet, and four singletons. PFGE confirmed the clonality of the isolates.
Conclusions
This is the first report of multidrug resistant P. mirabilis in a nursing home in Croatia. Cephalosporin resistance was due to plasmid-mediated AmpC β-lactamase CMY-16.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The rapid emergence of antibiotic resistance among Gram-negative bacteria is a serious threat to the management of infectious diseases. β-lactam antibiotics are the most frequently used antimicrobials for empirical therapy [1]. Production of β-lactamases is one of the strategies adopted by bacteria to develop resistance to the β-lactam class of antibiotics [1, 2]. The development of highly stable expanded-spectrum cephalosporins at the beginning of the 1980s was quickly followed by the emergence of extended-spectrum β-lactamases (ESBL) in Klebsiella pneumoniae and other Enterobacteriaceae [2]. These enzymes are usually plasmid-mediated and most frequently derived from parental TEM-1, TEM-2, and SHV-1 β-lactamases by point mutations that alter the configuration of the active site to expand their spectrum of activity [3]. AmpC enzymes hydrolyze first-, second-, and third-generation cephalosporins and cephamycins, but spare cefepime and carbapenems. Unlike ESBLs they are not inhibited by clavulanic acid, sulbactam, or tazobactam [4]. Proteus mirabilis is an emerging cause of nosocomial infections, particularly of wounds and the urinary tract. The various types of P. mirabilis infections are difficult to treat because of acquisition of various resistance mechanisms, such as ESBLs or AmpC β-lactamases [5, 6]. Recently, an increased frequency of multidrug-resistant P. mirabilis isolates was observed in a long-term care facility in Zagreb (Godan). The role of P. mirabilis as an important multidrug-resistant pathogen in long-term care facilities has not been investigated yet. The previous reports on ESBLs in P. mirabilis in Croatia showed the clonal spread of TEM-52 β-lactamase-producing P. mirabilis isolates in the University Hospital Center Split [7, 8]. Spread of multidrug-resistant P. mirabilis from hospitals to nursing homes was observed recently. This prompted us to conduct the molecular characterization of antibiotic resistance in P. mirabilis isolates from a nursing home in Zagreb.
Materials and methods
Bacteria
Thirty-eight consecutive non-duplicate P. mirabilis isolates with reduced susceptibility to ceftazidime (zone diameter ≤22 mm) were isolated from urine samples during a period from April 29, 2013 until January 21, 2015 from a nursing home Godan in Zagreb, Croatia. The isolates were identified by conventional biochemical tests using standard recommended techniques.
Susceptibility testing
The susceptibility testing to amoxicillin alone and combined with clavulanate; and piperacillin alone and combined with tazobactam, cefazoline, cefuroxime, ceftazidime, cefotaxime, ceftriaxone, cefepime, cefoxitin, imipenem, meropenem, gentamicin, and ciprofloxacin was performed by a twofold microdilution technique according to Clinical Laboratory Standard Institution (CLSI) standard procedures [9].
Disk diffusion test was performed for all antibiotics which are routinely tested in our laboratory for diagnostic purposes (amoxycillin alone and combined with clavulanic acid; piperacillin alone and combined with tazobactam, cephalexin, cefuroxime, ceftazidime, cefotaxime, ceftriaxone, cefepime, cefoxitin, gentamicin, netilmicin, amikacin, ciprofloxacin, norfloxacin, sulphametoxazole/trimethoprim, and nitrofurantoin) prior to microdilution.
Escherichia coli ATCC 25922 and K. pneumoniae ATCC 700603 were used as quality control isolates.
Phenotypic characterization of β-lactamases
A double-disk synergy test (DDST) using the combination of amoxycillin/clavulanate with cefotaxime, ceftriaxone, ceftazidime, and aztreonam [10], as well as a combined disk test using disks of ceftazidime, cefotaxime, ceftriaxone, and cefepime with and without clavulanate (10 µg/ml) according to CLSI were performed to detect ESBLs [9]. Deformation of the inhibition zone around cephalosporin disks towards the central disk with amoxicillin/clavulanate in DDST or augmentation of inhibition zone around cephalosporin disks for at least 5 mm in the presence of clavulanic acid compared to control disks without clavulanic acid in combined disk test indicated production of ESBL. E. coli ATCC 25922 was used as a negative and K. pneumoniae ATCC 700603 as a positive control.
Presumptive test for AmpC β-lactamases is considered positive if the inhibition zone for cefoxitin was ≤18 mm [9]. AmpC β-lactamases were phenotypically detected by combined disk test using disks of ceftazidime, cefotaxime, and ceftriaxone with and without 3-amino-phenyboronic acid (PBA). AmpC production was indicated by an increase in zone size of 5 mm or more around cephalosporin disks containing PBA compared to control disks containing only cephalosporins [11].
Conjugation
P. mirabilis isolates were investigated for the transferability of their resistance determinants. Conjugation experiments were set up employing plasmid-free and sodium azide-resistant E. coli A15 R- recipient strain [12]. Transconjugants were selected on the combined plates containing ceftazidime (1 mg/L) and sodium azide (100 mg/L). The frequency of conjugation was expressed relative to the number of donor cells.
Characterization of β-lactamases
The presence of bla TEM, bla SHV, bla CTX-M, bla PER-1, and bla ampC genes was investigated by polymerase chain reaction (PCR) using primers and conditions as described previously [13–17]. In order to amplify the whole coding sequence, additional primers were used for amplification of bla CMY genes, as described previously [18]. Template DNA was extracted by the boiling method. PCR mix (50 µl) contained 25 µl of master mix (Roche, Medical Intertrade, Zagreb, Croatia), 20 µl of ultrapure water, 1 µl of each primer (10 pmol), and 3 µl of template DNA. Lysates from reference strains producing TEM-1, TEM-2, SHV-1, SHV-2, SHV-4, SHV-5, CTX-M-15, PER-1, CMY-4, MIR-1, DHA-1, FOX-1, and MOX-1 were used as positive controls for PCR. Nucleotide sequences were determined directly on PCR products on both strands by the Microgene (Macrogene, Seoul, South Korea, sDNA sequencing service) DNA sequencing service. CMY and TEM amplicons were sequenced. Sequences were analyzed using BLAST program (National Center for Biotechnology Information, NCBI). Designation of bla genes based on identified mutations was done according to the Bush, Jacoby, and Medeiros scheme. The presence of ISEcp1 and IS26 in the region upstream of bla CMY genes was investigated by combining IS26 and ISEcp1 forward primers with reverse primers for bla CMY [19].
Detection of quinolone resistance determinants
Plasmid borne quinolone resistance genes qnrA, qnrB and qnrS were determined by PCR as described previously [20].
Characterization of plasmids
Plasmids were extracted with the Macherey Nagel mini kit (Hilden, Germany) according to manufacturer’s recommendations. Plasmids extractions were subjected to PCR-based replicon typing (PBRT) according to Carattoli et al. [21], and to PCR with primers specific for TEM and CMY β-lactamases to determine the location of bla genes.
Genotyping of isolates
Twenty-eight isolates were subjected to molecular typing by rep-PCR as described previously [22] DNA was isolated by Ultra-Clean microbial DNA isolation kit (Mo Bio Laboratories, Carlsbad, CA, USA), as recommended by the manufacturer. The DNA concentration was measured and set between 25 and 30 ng/L. Subsequently, the DNA was amplified using the Bacterial fingerprinting kit (Bacterial barcodes, bioMerieux, Athens, GA, USA), according to the manufacturer’s instructions. PCR was performed using the following parameters: initial denaturation (94 °C) for 2 min; and then 35 cycles of 30 s of denaturation (94 °C), 30 s of annealing (60 °C), and 90 s of extension (70 °C); followed by 3 min of final extension (70 °C); and ending at 4 °C. The amplification products were separated with the Agilent B2100 bioanalyzer. Five microliters of DNA standard markers (used for normalization of sample runs) and 1 µl of the DNA product were used. All data were entered in the DiversiLab software system. Cutoff value of 97 % was used to define a clone.
Pulsed-field genotyping of SfiI-digested genomic DNA was performed on 30 isolates with a CHEF-DRIII system (Bio-Rad); the images were processed using the Gel-Compar software, and a dendrogram was computed after band intensity correlation using global alignment with 1.5 % optimization and tolerance and unweighted pair-group method using arithmetical averages (UPGMA) clustering. The strains were considered to be clonally related if they showed more than 80 % similarity of their PFGE patterns [23, 24].
Results
Patients
All patients were residents of the Godan long-term care facility. Since the Godan nursing home is located close to the University Hospital Center Zagreb where the urine samples were processed, we found in the hospital internet system that 23 of 38 patients were previously hospitalized in the University Hospital Center in the intensive care unit, pulmonary unit, abdominal surgery, hematology, ophthalmology, gastroenterology, cardiology, and oncology departments. Three patients were only examined in the emergency room in order to change the urinary catheter or to obtain the blood transfusion but did not stay in the hospital. They all had severe underlying diseases such as coronary artery disease; myocardial infarction; adenocarcinoma ventriculi; pancreatic cancer; chronic lymphocytic or myelocytic leukemia; prostatic adenocarcinoma; respiratory insufficiency; kidney failure; megaloblastic and hypochromic anemia; Morbus Alzheimer; pulmonary embolia; and diabetes mellitus. Two patients suffered from bronchopneumonia and were treated with azithromycine and ceftriaxone. All patients had urinary tract infection with > 105 CFU/ml of P. mirabilis and white blood cells in the urinary sediment. Ten patients had additional E. coli ESBL, three K. pneumoniae ESBL, and eight E. faecalis. The majority of patients received cefuroxime or ciprofloxacin for the treatment of urinary infections prior to isolation of P. mirabilis.
Isolates
Detection of ESBLs and susceptibility testing
The isolates were resistant to amoxicillin alone and combined with clavulanic acid, piperacillin, cefuroxime, cefoxitin, gentamicin, and ciprofloxacin, but susceptible to cefepime, imipenem, and meropenem with MICs of imipenem being slightly higher than those of meropenem according to microdilution test (Table 1). There were variable susceptibility/resistance patterns to ceftazidime, cefotaxime, ceftriaxone and to combination of piperacillin with tazobactam as shown in Table 1. Meropenem was the most potent antibiotic with Minimum Inhibitory Concentration (MIC)90 of 0.06 mg/L. In disk diffusion tests, all isolates were resistant to sulfametoxazole/trimethoprim (cotrimoxazole) and norfloxacin. The phenotype of resistance, including resistance or reduced susceptibility to expanded-spectrum cephalosporins (ceftazidime, cefotaxime, ceftriaxone), cefoxitin, and amoxicillin/clavulante, but preserved susceptibility to cefepime was consistent with production of plasmid-mediated AmpC β-lactamase which was confirmed by an inhibitor-based test. An augmentation of the inhibition zones around cephalosporin disks of at least 5 mm was seen with PBA but not with clavulanic acid. All isolates tested phenotypically positive for AmpC but negative for ESBLs.
Conjugation
The isolates did not transfer ceftazidime resistance to an E. coli recipient strain.
Characterization of β-lactamases
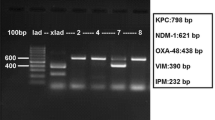
All 38 P. mirabilis strains yielded an amplicon of 1432 bp with primers specific for CMY-β-lactamase genes. Sequencing of amplicons revealed the bla cmy16 β-lactamase allele in all strains except strain 12, which was found to produce CMY-112. The isolates were positive for bla TEM-1, but negative for bla SHV, bla CTX-M, and bla PER-1 genes. ISEcp1 was identified 110 bp upstream of the bla CMY-16 starting codon.
Characterization of plasmids
Plasmid encoding CMY-16 did not belong to any known PBRT. The plasmid extractions were positive for bla TEM and bla CMY genes.
Detection of quinolone resistance determinants
Plasmid borne quinolone resistance genes-qnrA, qnrB, and qnrS were not found.
Genotyping of the isolates
Rep-PCR of 28 isolates identified one large clone consisting of 15 isolates (8, 2, 19, 14, 7, 4, 3, 20, 15, 22, 24, 26, 12, 16, 5); however, a certain degree of diversification was observed within the clone, with seven subclusters containing two or three identical isolates as shown in Fig. 1. The first one with strains 2, 8, and 19; the second one with strains 7 and 14; the third with strains 3 and 4; the fourth one with strains 22, 24 and 26; and the fifth one with strains 5 and 16.
Three pairs of related isolates (13 and 9, 23 and 25, and 38 and 35) and one triplet (28, 30, 31) were identified (Fig. 1). Four isolates were singletons: 6, 17, 18 and 36.
PFGE identified one large clone with 19 isolates out of 30 (isolates 15, 8, 20, 19, 16, 13, 11, 10, 9, 7, 4, 12, 14, 38, 36, 25, 34, 28, 30), one small cluster with three strains (1, 23, 29), two pairs (37, 33 and 27 and 31), and and four singletons (6, 18, 32, 35), as shown in Fig. 2.
Rep-PCR showed a better discriminatory effect because it identified a subcluster among the large clone and this could explain small discrepancies between the two genotyping methods.
Discussion
Previous studies found TEM-52 and PER-1 ESBLs to be dominant resistance determinants to expanded-spectrum cephalosporins in P. mirabilis [6]. This study demonstrated predominance of plasmid-mediated AmpC β-lactamase CMY-16 among tested isolates. AmpC β-lactamases detection is not routinely carried out in many microbiology laboratories. This could be attributed to lack of awareness, or lack of resources and facilities to conduct β-lactamase identification. Currently available tests for detection of plasmid-mediated AmpC β-lactamases are inconvenient, subjective, and lack sensitivity and specificity [4, 11]. AmpC β-lactamases are inhibited by PBA and cloxacillin. There are several inhibitor-based tests for identification of AmpC β-lactamases, including the disk test and the E-test [25]. The production of CMY β-lactamase was associated with resistance or reduced susceptibility to third-generation cephalosporins and the combination of amoxicillin with clavulanic acid. The isolates showed variable levels of susceptibility/resistance to piperacillin/tazobactam, which would lead to the conclusion that this combination is less affected by production of AmpC β-lactamase compared to amoxicillin/clavulanate. This could be attributed to better intrinsic activity of piperacillin against P. mirabilis compared to amoxicillin. The susceptibility to cefepime and carbapenems was maintained, with meropenem having slightly lower MICs. Ceftazidime resistance was not transferred by conjugation to an E. coli recipient isolate, indicating that CMY genes were encoded on non-transferable plasmids. P. mirabilis lacks the ampC gene and, thus, AmpC β-lactamases are always plasmid mediated in this species, although some studies found incorporation of the bla CMY gene in the chromosome [26]. In our study, the plasmid extract did not belong to any known PBRT but yielded amplicons with primers specific for TEM and CMY β-lactamases. However, it is not possible to exclude the possibility of chromosomal contamination of plasmid extract and chromosomal location of the bla ampC gene. Fifteen of the isolates were found to be clonally related, although three pairs, one triplet, and four singleton isolates were observed. This finding points to clonal dissemination of related isolates within the nursing home, probably due to the contaminated urinary catheters, but horizontal spread of the bla CMY gene also occurred, most likely mediated by the ISEcp1 insertion sequence upstream of the gene. All the patients had severe underlying diseases and were previously hospitalized in one of the large hospital centers in Zagreb (University Hospital Center Zagreb, Sisters of Mercy University Hospital, and University Hospital Merkur) and there is a possibility that they were colonized with multiresistant P. mirabilis during their stay in the hospital, thus raising the possibility of multiple independent introduction routes of AmpC-positive P. mirabilis into the long-term care facility. The first plasmid-mediated AmpC β-lactamase reported in Croatia was DHA-1, identified in E. coli in 2003 [27]. Recent studies found plasmid-mediated AmpC β-lactamases of the CMY family among hospital P. mirabilis isolates from Split [28] and among E. coli isolates from companion animals in Croatia [29]. Moreover, CMY-4 was identified as an additional β-lactamase in Enterobacteriacea producing VIM or NDM metallo-β-lactamases [30]. In the present study, we found an alarming number of AmpC-producing P. mirabilis in a nursing home in Zagreb. CMY β-lactamases originate from chromosomal AmpC β-lactamases of Citrobacter freundii [26]. The acquired bla CMY genes have escaped from the chromosome of C. freundii following mobilization mediated by ISEcp1, IS26, or ISCR1. CMY-1, CMY-12, and CMY-16 were found to be the most prevalent variants of plasmid-mediated AmpC β-lactamases in Europe [26]. In addition, mobile insertion sequences such as IS26 and/or ISEcp1, which can be found upstream of bla AmpC genes, can facilitate their mobilization. Similar genetic context with ISEcp1 preceding bla CMY-16 was previously reported [31]. Simultaneous production of ESBLs and AmpC β-lactamases was also reported in P. mirabilis in recent studies [5]. CMY-16 was previously reported in P. mirabilis from a long-term care facility in Italy [32]. In the latter study, TEM-92, which is an ESBL, and plasmid-mediated AmpC β-lactamase CMY-16 were found. Similar to our study, CMY-16-producing organisms were clonally related, unlike those possessing ESBL [32]. The production of additional TEM-1 β-lactamase could increase the level of resistance to amoxycillin combined with clavulanate.
From the therapeutic point of view, it is important to distinguish between ESBLs and AmpC β-lactamases because infections caused by AmpC positive isolates can be effectively treated with cefepime and cefpirome. On the other hand, uncomplicated urinary tract infections due to ESBL-positive organisms can be treated with β-lactam/inhibitor combinations which are not recommended for AmpC producing organisms [33], although our isolates demonstrated in-vitro susceptibility to piperacillin/tazobactam. Some authorities recommend all expanded-spectrum cephalosporins to be reported as resistant if the isolate produces plasmid-mediated AmpC β-lactamase, regardless of the in-vitro susceptibility results, to avoid therapeutic failures [33]. CLSI has yet to establish a testing and reporting algorithm specifically for organisms containing AmpC β-lactamases. Identification of AmpC β-lactamases in E. coli, P. mirabilis, and Klebsiella spp. can increase the accuracy of antimicrobial testing reports for expanded-spectrum cephalosporins if the results are used to modify the interpretations of cephalosporin results [33]. Recent studies demonstrated a high rate of clinical failure among patients who were infected in the bloodstream with AmpC-producing organisms and who received cephalosporin treatment [33, 34]. There are no data on efficacy of cephalosporin therapy for urinary tract infections associated with AmpC-producing organisms. The spread of AmpC-producing P. mirabilis in Europe pose a serious laboratory and therapeutic challenge [34]. Recently, P. mirabilis has demonstrated great capacity to accumulate resistance genes such as those encoding ESBLs, plasmid-mediated AmpC β-lactamases, carbapenemases, and fluoroquinolone resistance genes.
Considering the gravity of the implication of inappropriate therapy in chronically ill and debilitated patients in long-term care facilities, looking for AmpC β-lactamases must be mandatory in all microbiological laboratories, and clinicians should be educated on the importance of ESBLs and AmpC β-lactamases and therapeutic challenges that they pose [33, 34].
References
Grover CN, Sahni BAK, Bhattacharya CS. Therapeutic challenges of ESBLs and AmpC β-lactamase producers in a tertiary care center. Med J Armed Forces. 2013;69:4–10.
Bush K. Is it important to identify extended-spectrum β-lactamase-producing isolates? Eur J Clin Microbiol Infect Dis. 1996;15:361–4.
Phillipon A, Arlet G, Lagrange H. Origin and impact of plasmid-mediated extended-spectrum β-lactamases. Eur J Clin Microbiol Infect Dis. 1994;13(Suppl 1):17–29.
Tan TJ, Yong NG, Koh TK, Hsu LY. Evaluation of screening methods to detect plasmid-mediated AmpC in Escherichia coli, Klebsiella pneumoniae and Proteus mirabilis. Antimicrob Agents Chemother. 2009;53:146–9.
Datta P, Gupta V, Arora S, Garg S, Chander J. Epidemiology of extended-spectrum β-lactamase, AmpC, and carbapenemase production in Proteus mirabilis. Jpn J Infect Dis. 2014;67:44–6.
Pagani L, Migliavacca R, Pallechi L. Emerging extended-spectrum β-lactamases in Proteus mirablis. J Clin Microbiol. 2002;40:1549–52.
Sardelić S, Bedenić B, Šijak D, Colinon C, Kalenić S. Emergence of Proteus mirabilis isolates producing TEM-52 β-lactamase in Croatia. Chemotherapy. 2010;56:208–13.
Tonkic M, Mohar B, Sisko-Kraljević K, Mesko-Meglic K, Goić-Barisić I, Novak A, Kovacić A, Punda-Polić V. High prevalence and molecular characterization of extended-spectrum β-lactamase-producing Proteus mirabilis isolates in southern Croatia. J Med Microbiol. 2010;59:1185–90.
CLSI. Clinical and Laboratory Standards Institute Performance Standards for Antimicrobial Susceptibility Testing. 21st informational supplement. Wayne, PA: CLSI; 2011, pp. M100–21.
Jarlier V, Nicolas MH, Fournier G, Philippon A. Extended broad-spectrum β-lactamases conferring transferable resistance to newer β-lactam agents in Enterobacteriaceae: hospital prevalence and susceptibility patterns. Rev Infect Dis. 1998;10:867–78.
Coudron PE. Inhibitor-based methods for detection of plasmid-mediated AmpC β-lactamases in Klebsiella spp., Escherichia coli and Proteus mirabilis. J Clin Microbiol. 2005;43:4163–7.
Elwell S, Falkow LP. The characterization of R plasmids and the detection of plasmid-specified genes. In: Lorian V, editor. Antibiotics in Laboratory Medicine, 2nd edn. Baltimore: Williams and Wilkins; 1986. pp. 683–721.
Nüesch-Inderbinen MT, Haechler H, Kayser FH. Detection of genes coding for extended-spectrum SHV β-lactamases in clinical isolates by a molecular genetic method, and comparison with the E test. Eur J Clin Microbiol Infect Dis. 1996;15:398–402.
Arlet G, Brami D, Decre D, Flippo A, Gaillot O, Lagrange PH, Philippon A. Molecular characterization by PCR restriction fragment polymorphism of TEM β-lactamases. FEMS Microbiol Lett. 1995;134:203–8.
Woodford N, Ward ME, Kaufmann ME, Turton J, Fagan EJ, James D, Johnson AP, Pike R, Warner M, Cheasty T, Pearson A, Harry S, Leach JB, Loughrey A, Lowes JA, Warren RE, Livermore DM. Community and hospital spread of Escherichia coli producing CTX-M extended-spectrum β-lactamases in the UK. J Antimicrob Chemother. 2004;54:735–43.
Pagani L, Mantengoli E, Migliavacca R, Nucleo E, Pollini S, Spalla M, Daturi R, Romero E, Rossolini GM. Multifocal detection of multidrug-resistant Pseudomonas aeruginosa producing PER-1 extended-spectrum β-lactamase in Northern Italy. J Clin microbiol. 2004;42:2523–9.
Perez-Perez FJ, Hanson ND. Detection of plasmid-mediated AmpC β-lactamase genes in clinical isolates by using multiplex PCR. J Clin Microbiol. 2002;40:2153–62.
Hanson ND, Moland ES, Hossain A, Neville SA, Gosbell IB, Unusual TKS. Salmonella enterica serotype typhimurium isolate producing CMY-7, SHV-9 and OXA-30 β-lactamases. J Antimicrob Chemother. 2002;49(6):1011–4.
Saladin M, Cao VY, Lambert T. Diversity of CTX-M beta-lactamases and their promoter regions from Enterobacteriaceae isolated in three Parisian hospitals. FEMS Microbiol Lett. 2002;209:161–8.
Robischek A, Jacoby GA, Hooper DC. The worldwide emergence of plasmid-mediated quinolone resistance. Lancet Infect Dis. 2006;6:629–40.
Carattoli A, Bertini L, Villa L, Falbo V, Hopkins KL, Threfall EJ. Identification of plasmids by PCR-based replicon typing. J Microbiol Methods. 2005;63:219.
Overdevest S, Willemsen C, Elberts M, Verhulsts P, Rijnsburger J, Savelkoulnd A, Kluytmans JW. Evaluation of the DiversiLab Typing Method in a Multicenter Study. Assessing Horizontal Spread of Highly Resistant Gram-Negative Rods. J Clin Microbiol. 2011;49:3551–54.
Kaufman ME. Pulsed-Field Gel Electrophoresis. In: Woodford N, Johnsons A, editors. Molecular bacteriology. Protocols and clinical applications, 1st edn. New York: Humana Press Inc.; 1998. pp. 33–51.
Tenover F, Arbeit R, Goering RV, Mickelsen PA, Murray BE, Persing DH, Swaminthan B. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33:2233–9.
Black JA, Moland ES, Thomson KA. AmpC disk test for identification of plasmid-mediated-AmpC β-lactamases in Enterobacteriaceae lacking chromosomal AmpC β-lactamase. J Clin Microbiol. 2005;43:3110–3.
D’Andrea MM, Literacka E, Zioga A, Giani T, Baraniak A, Fiett J, Sadowi E, Tassios T, Rossolini GM, Gniadkowski M, Miriagou V. Evolution and spread of multidrug-resistant Proteus mirabilis clone with chromosomal AmpC β-lactamase in Europe. Antimicrob Agents Chemother. 2011;55:2735–42.
Giakoupi P, Tambić-Andrašević A, Vourli S, Škrlin J, Šestan-Crnek S, Tzouvelekis LS, Vatoupoulos AC. Transferable DHA-1 cephalosporinase in Escherichia coli. Int J Antimicrob Agents. 2006;27:77–80.
Rubić Ž, Soprek S, Jelić M, Radić M, Novak A, GoićBarišić I, Tonkić M, Tambić-Andrašević A. The first detection of plasmid-mediated AmpC β-lactamase in multidrug-resistant Proteus mirabilis isolates from University Hospital Split, Croatia 24th European congress of clinical Microbiology and Infectios Diseases, Barcelona, Spain. 2014.
Bedenić B, Matanović K, Mekić S, Varda-Brkić D, Šeol-Martinac B. Coproduction of CTX-M-15 and CMY-2 in animal Escherichia coli isolate from Croatia 24th European congress of clinical Microbiology and Infectios Diseases, Barcelona, Spain. 2014.
Zujić-Atalić V, Bedenić B, Kocsis E, Mazzariol A, Sardelić S, Barišić M, Plečko V, Bošnjak Z, Mijač M, Jajić I, Vranić-Ladavac M, Cornaglia G. Diversity of carbapenemases in clinical isolates of Enterobacteriaceae in Croatia-the results of the multicenter study. Clin Microbiol Infect. 2004;20:894–903.
Literacka E, Empel J, Baraniak A, Sadowy E, Hryniewic W, Gniadkowski M. Four variants of the Citrobacter freundii AmpC type cephalosporinase including tow novel enzymes, CMY-14 and CMY-15 in a Proteus mirabilis clone widespread in Poland. Antimicrob Agents Chemother. 2004;48:4136–43.
Migliavacca R, Migliavacca A, Nucleo E, Ciaponi A, Spalla M, Luca C De, Pagani L. Molecular epidemiology of ESBL producing Proteus miraiblis isolates from a long-term care and rehabilitation facility in Italy. New Microbiol. 2007;30:362–366.
Tenover F, Emery SL, Spiegel CA. Identification of plasmid-mediated AmpC β-lactamases in Escherichia coli, Klebsiella spp and Proteus mirabilis can potentially improve reporting of cephalosporin susceptibility testing results. J Clin Microbiol. 2009;47:294–299.
Luzzaro F. Spread of multidrug-resistant Proteus mirabilis isolates producing an AmpC-type β-lactamase:epidemiology and clinical management. Int J Antimicrob Agents. 2009;33:328–333.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
B. Bedenić, N. Firis, V. Elveđi-Gašparović, M. Krilanović, K. Matanović, I. Štimac, J. Luxner, J. Vraneš, T. Meštrović, G. Zarfel and A. Grisold state that there are no conflicts of interest.
Ethical standards
This was in vitro study which did not involve human or animal subject and the permission from the Ethical Comittee was not necessary.
Rights and permissions
About this article
Cite this article
Bedenić, B., Firis, N., Elveđi-Gašparović, V. et al. Emergence of multidrug-resistant Proteus mirabilis in a long-term care facility in Croatia. Wien Klin Wochenschr 128, 404–413 (2016). https://doi.org/10.1007/s00508-016-1005-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00508-016-1005-x