Abstract
The arabinogalactan protein (AGP) family is one of the most complex protein families and is ubiquitous in the plant kingdom. Moreover, it has been demonstrated to play various roles during plant reproduction. A typical AGP contains a hydroxyproline-rich core protein with high heterogeneity and varying numbers of polysaccharide side chains. However, the functions of the polysaccharide components (i.e. AG sugar chains) remain largely unknown due to the general difficulties associated with studying sugar chains in glycobiology. In recent years, methodological breakthroughs have resulted in substantial progress in AGP research. Here, we summarise the multiple roles of AGPs during plant gametophyte development and male–female communication, with a focus on recent advances. In addition, we discuss the analytical tools used in AGP research, and the biosynthesis and function of AG sugar chains. A comprehensive understanding of the AGP family will help clarify the mechanisms precisely controlling reproductive processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since their isolation from various plant tissues more than half a century ago, arabinogalactans (AGs) have been proposed to be linked to a large group of hydroxyproline-containing polypeptides known as AG proteins (AGPs) (Fincher et al. 1974, 1983). The typical structure of an AGP consists of a highly variable core protein enriched in hydroxyproline residues and various carbohydrate side chains that are often anchored to the cell membrane according to glycosylphosphatidylinositol (GPI) lipid anchor signals (Seifert and Roberts 2007; Nguema-Ona et al. 2013; Pereira et al. 2015). GPI-anchored AGPs can be released into the cell wall. Since many AGPs are highly glycosylated, more than 90% of their total molecular mass comes from glycan moieties consisting of (1 → 3)-β-galactan and (1 → 6)-β-linked galactan chains, which are thought to be important for the functional diversity of AGPs (Knoch et al. 2014).
Compared with many other protein families, only a handful of AGPs have been functionally characterised, mainly due to the complexity of their AG sugar chains and the heterogeneity of their core proteins. With the help of tools such as anti-AG chain-based immunomicroscopy, β-Yariv reagents, degradation enzymes that target specific parts of AG chains, chemical synthesis of specific structures of AG chains, and bioinformatics methods, the nature of AGPs is becoming increasingly clear. AGPs have crucial roles in multiple biological processes, including cell division, cellular communication, programmed cell death, embryogenesis, postembryonic pattern regulation, secondary wall deposition, organ abscission, plant–microbe interactions, plant growth, and reproductive processes (reviewed in Majewska-Sawka and Nothnagel 2000; Seifert and Roberts 2007; Ellis et al. 2010; Nguema-Ona et al. 2012, 2013; Pereira et al. 2015, 2016a). In reproduction processes, AGPs and their sugar chains are involved in gametophyte development and male–female interactions (Fig. 1).
Arabinogalactan proteins (AGPs) involved in plant reproduction. AGPs and their functions are indicated (blue circles: male gametophyte development and function; orange circle: female gametophyte development; red circles: male–female communication). See the text for details of each function. AMOR is an AG sugar chain, for which the protein backbone is unknown
In this review, we summarise recent progresses in our understanding of the multiple functions of AGPs in plant reproduction, with a special focus on the analytical tools used to study AGPs, as well as the biosynthesis and functions of AG sugar chains.
Classification of AGPs
According to the structure of their core proteins, the AGP family can be generally categorised into two types: classical AGPs and non-classical AGPs (Showalter 2001). Classical AGPs often share a signal peptide in the N-terminal and always combine with a GPI anchor in the C-terminal; only the central domain consists of a high percentage of proline, alanine, serine, and threonine (PAST) residues (Schultz et al. 2000; Showalter 2001). Those with short mature protein backbones, usually with residues of around 10–13 amino acids, are separately designated as AG peptides (Schultz et al. 2000). Distinct from these two classical AGPs, a third type of classical AGP is the lysine-rich AGP subfamily, which is characterised by a short lysine-rich region at the C-terminal (Sun et al. 2005).
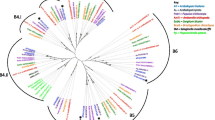
Distinct from the classical AGPs, non-classical chimeric AGPs with different domains in their core proteins can be classified into three main subfamilies: fasciclin-like AGPs (FLAs), phytocyanin-like AGPs (PAGs), and xylogen-like AGPs (XYLPs) (Ma et al. 2017). FLAs contain at least one fasciclin-like domain, a secretion signal in the N-terminal, one or two AGP regions, and often a GPI anchor in the C-terminal (Johnson et al. 2003). Members of the PAG subfamily share a similar structure with phytocyanins, except for the presence of amino acids that bind copper in their plastocyanin-like domains (Mashiguchi et al. 2009; Cao et al. 2015). The XYLP subfamily members, originally isolated from differentiating xylem cells of in vitro Zinnia elegans cultures, contain AGP and non-specific lipid transfer protein domains and act as functional extracellular proteins in various plant tissues (Motose et al. 2004; Kobayashi et al. 2011). In addition, there are many other chimeric AGPs that cannot be grouped into these three subfamilies, which are designated as “other chimeric AGPs” (Ma et al. 2017). Apart from the classical and chimeric AGPs, those containing sequence characteristics of both AGPs and extensins (EXTs) are separately referred to as hybrid AGP/EXTs (HAEs) (Showalter et al. 2010). A recent study characterised 151 AGPs from the genome database of Arabidopsis thaliana, including 42 classical AGPs, 105 chimeric AGPs, and 4 HAEs (Ma et al. 2017; Fig. 2).
Classification and expression of AGPs of Arabidopsis thaliana. Publicly available expression data of 130 of 151 AGP genes are shown as heat maps using Genevestigator (Hruz et al. 2008), and according to the classification of Ma et al. (2017). See the main text for details of AGP classification. The expression levels in the male reproductive organs/tissues are highlighted in blue, while those in female tissues are highlighted in orange
AGPs are the jack of all trades of plant reproduction
AGP genes show differential expression in various plant tissues (Fig. 2). Most AGPs are expressed in plant reproductive tissues, and some classical and non-classical AGPs show predominantly high expression levels. For example, AGP6/11/22/23/24/40 and BCP1, which are classical AGPs, ENODL6/7 and BCB from the PAG subfamily, FLA3/14 from the FLA subfamily, members of XYLP, and other chimeric AGPs are highly expressed in male reproductive tissues, including the stamen and pollen. Meanwhile, FLA1/8/10/16, many PAG members such as ENODL1/11/12/13/14/15, the XYLP subfamily members XYP11, AT5G09370, and AT1G73560, and several other chimeric AGPs accumulate in the female parts of the pistil, carpel, stigma, ovary, and ovule (Fig. 2). The expression levels of several classical AGP members have been confirmed by AGP-specific promoters fused with fluorescent proteins or β-glucuronidase, and with fluorescence in situ hybridisation (Pereira et al. 2014). These expression patterns of AGPs are consistent with their roles in plant reproduction.
AGPs in gametophyte development
Land plants have a complex life history that encompasses haploid gametophytes and diploid sporophytes, both of which are strictly regulated by numerous molecular networks. In flowering plants, the male gametophyte (i.e. pollen grain) develops during two phases, microsporogenesis and microgametogenesis, in the stamen. Meanwhile, the female gametophyte (i.e. embryo sac) originating from the ovule experiences two similar phases, known as megasporogenesis and megagametogenesis (reviewed in Borg et al. 2009; Yang et al. 2010). In the following sections, we examine the various roles of AGP members in the development of male and female gametophytes in flowering plants.
In addition to studies using monoclonal antibodies and β-Yariv reagents, AGP mutants provide insights into the function of each AGP gene. AGP18, a gene encoding a lysine-rich classical AGP in A. thaliana, is indispensable for the development of female gametophytes (Acosta-Garcia and Vielle-Calzada 2004; Demesa-Arevalo and Vielle-Calzada 2013). For example, downregulation of AGP18 was associated with decreased fertility and aborted ovules, whereby functional megaspores could not undergo normal haploid mitosis and thus failed to initiate female gametogenesis (Acosta-Garcia and Vielle-Calzada 2004). Overexpression of AGP18 also reduced fertility due to abnormal maintenance of viable megaspores, indicating its function in megaspore selection (Demesa-Arevalo and Vielle-Calzada 2013).
AGPs are not only involved in female reproductive tissues, but many have been demonstrated to be essential for the differentiation of male parts. AGP6 and AGP11, two phylogenetically similar AGP genes specifically expressed in male reproductive tissues, are required for stamen and pollen grain development (Pereira et al. 2006; Coimbra et al. 2008; Levitin et al. 2008; Coimbra et al. 2009). In double mutants of agp6 agp11, a large proportion of pollen grains collapsed, and there was a failure to release pollen, leading to reduced fertility (Levitin et al. 2008; Coimbra et al. 2009). A mutation in the AGP11 homolog BcMF8 in Brassica campestris also exhibited abnormal pollen shape and pollen tube growth, suggesting the functional conservation of this AGP branch within the family Brassicaceae (Lin et al. 2014).
Apart from the classical AGPs, FLA-like chimeric AGPs are also recruited during gametophyte development. FLA3, which encodes an FLA in pollen grains and tubes, is involved in microspore development by affecting cellulose deposition (Li et al. 2010). After downregulating FLA3 expression, male fertility was reduced, and many wrinkled and shrunken pollen grains were observed. In addition, half of the pollen grains showed defects in the pollen intine layer and were aborted during transition to the bicellular stage, indicating that FLA3 has a role in intine layer formation (Li et al. 2010). In the monocot rice, MICROSPORE AND TAPETUM REGULATOR1 (MTR1) encodes an FLA specifically expressed in male reproductive cells, and its mutant showed a complete male sterile phenotype, indicating that MTR1 is indispensable to pollen tapetum and microspore development in rice (Tan et al. 2012).
AGPs in male–female interactions
Flowering plants (angiosperms) have evolved a set of mechanisms to guide pollen tubes precisely to enter the embryo sac and ultimately accomplish double fertilisation. It has been demonstrated that AGPs are widely involved in the interactions between the male and female parts during this process.
Pioneering studies were performed using Nicotiana species more than two decades ago. The AGP protein, transmitting-tract-specific (TTS) protein from Nicotiana tabacum, stimulates pollen tubes to grow along the stylar transmitting tissue towards the ovary (Cheung et al. 1995; Wu et al. 1995). In the transmitting tissue, the extent of glycosylation of TTS proteins gradually increases from the stigma to the ovarian end, which might provide positional cues for the pollen tube (Wu et al. 1995). With suppression of TTS expression, the pollen tube growth rate is significantly reduced, which is thought to be conserved at least among the Nicotiana genus (Cheung et al. 1995; Wu et al. 2000). Another two HAEs from tobacco, class III pistil EXT-like proteins (PELPIII) and 120 kDa glycoprotein (120 K), have been shown to be involved in self-incompatibility (SI) and interspecific incompatibility (II), respectively (Hancock et al. 2005; Eberle et al. 2013). PELPIII suppression leads to an increase in interspecific pollen tube growth, suggesting its role in the specific inhibition of pollen tubes in II (Eberle et al. 2013); meanwhile, downregulation of 120 K results in failure of S-specific pollen rejection, favouring the idea that HAE-type AGPs play different roles in male–female interactions (Lind et al. 1994; Hancock et al. 2005). AGPs are also likely to have a role in acquisition of stigma receptivity in the apple flower (Losada and Herrero 2012).
Many recent studies have revealed the role of AGPs in male–female interactions in A. thaliana. A group of AGPs from the PAG subfamily, ENODL11/12/13/14/15, show high expression levels in the funiculus and ovules, which redundantly control pollen tube reception (Hou et al. 2016). Single, double, and triple mutants of these ENODL genes did not exhibit any obvious phenotype; however, in the ovules of quintuple mutants, wild-type pollen tubes failed to release sperm cells, indicating their function in coordinating male–female communication and facilitating double fertilisation (Hou et al. 2016). JAGGER encodes AGP4, which is specifically expressed in the transmitting tract, stigma, and integuments before fertilisation, and is responsible for the polytubey block (Pereira et al. 2016b). In a null mutant of JAGGER, the persistent synergid cell survived, which is normally disorganised after fertilisation, leading to partial failure of polytubey block (Pereira et al. 2016b).
AMOR has been identified as a bioactive AG sugar chain derived from the ovules of Torenia fournieri, which makes the pollen tube competent for the ovular attraction signal (Mizukami et al. 2016). Parallels can be drawn between the animal capacitation and plant pollen tube activation by pistil tissue, and AMOR was first identified as a responsible pistil molecule (Mizukami et al. 2016; Sankaranarayanan and Higashiyama 2018). AMOR activates the LURE (attractant peptides) signalling cascade in the pollen tube. The protein backbone of AMOR remains unknown. Interestingly, the terminal disaccharide structure, the β isomer of methyl-glucuronosyl galactose (4-Me-GlcA-β-1,6-Gal), has been shown to be both necessary and sufficient for AMOR activity (Mizukami et al. 2016). The strict structural activity relationship of AMOR (Jiao et al. 2017) might imply the existence of a receptor in the pollen tube that can recognise its disaccharide structure.
Tools used to identify the roles of AGPs and their sugar chains
Recent progress regarding our understanding of AGPs and their sugar chains has relied both on classical methods and on the development of new methods. In this section, we summarise the tools used to study AGPs and their sugar chains.
Adding to the tools summarised below, molecular genetics (mutant analysis) and glycan structure analysis have been powerful in AGP research. As shown in the study of ENODLs (Hou et al. 2016), some AGPs are likely to act redundantly. Highly efficient CRISPR/Cas9-mediated gene knockout in Arabidopsis (e.g. Tsutsui and Higashiyama 2017) contributes to explore the large AGP family. For details of glycan structure analysis, for example, Gane et al. (1995) reported structure analysis of AG sugar chains from stigmas and styles of Nicotiana alata by nuclear magnetic resonance (NMR). Tryfona et al. (2012) reported structure analysis of AG sugar chains from leaves of Arabidopsis by enzyme degradation, mass spectrometry, and carbohydrate gel electrophoresis.
Monoclonal antibodies
The localisation of AG sugar chains in plant tissues can be visualised using monoclonal antibodies that can detect different glycosidic AGP epitopes (Pennell and Roberts 1990; Penell et al. 1991; Yates et al. 1996; Willats et al. 1998; Qin and Zhao 2006; Moller et al. 2008). More than two decades ago, the alteration in AGP levels detected by MAC207 antibody was first proposed as a developmental switch occurring during primordia initiation in stamen and carpel (Pennell and Roberts 1990). Many more monoclonal antibodies have since been used to characterise the distribution of AG sugar chains and other carbohydrates. For example, JIM4, JIM8, and JIM13 recognise an AG epitope similar to MAC207 (Penell et al. 1991; Yates et al. 1996), LM6, and LM13 detect the location of arabinans and the pectin structure, and LM14 binds to the epitope of type II AG that may occur on AGPs (Willats et al. 1998; Moller et al. 2008). The immunolocation detected by these antibodies has been of great value for elucidating the functions of AGPs and their sugar chains, even in recent studies (Corral-Martinez et al. 2016; Da et al. 2017; Olmos et al. 2017; Suzuki et al. 2017). The reproductive organs of flowering plants show divergence in the distribution of specific sugar chains (e.g. Coimbra et al. 2007), implying that specific AG sugar chains might be involved in local cellular development and cell-to-cell communication. Epitopes of monoclonal antibodies have been examined with enzyme-linked immunosorbent assay (ELISA) using chemically defined sugar chains, and a microarray with various types of printed, synthesised sugar chains was developed recently to rapidly show the specificity of each monoclonal antibody (Ruprecht et al. 2017).
β-Yariv reagents
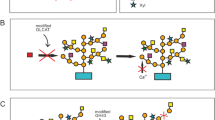
Yariv reagents are a group of synthetic phenylglycosides that were initially used for the purification of sugar-binding proteins, which were later demonstrated to selectively bind to AGPs depending on the glycosyl residue (Yariv et al. 1967; Kitazawa et al. 2013). For example, β-Yariv reagents bind to the β-1,3-galactan main chain (Fig. 3), a structure conserved among AGPs (Kitazawa et al. 2013). These reagents not only visualise the distribution of AGPs, but also perturb their biological functions, which has led to the wide application of β-Yariv reagents to study the functions of AGPs during plant development and growth.
AG sugar chain biosynthesis. A typical AG sugar chain structure (e.g. Tryfona et al. 2012) is shown with glycosylation enzymes. See the main text for details of the enzymes. The indications of KNS4/UPEX1, GALT29A, GALT31A, RAY1, and FUT4, and 6 for some residues are omitted. GALT31A has been suggested to work cooperatively with GALT29A by forming a protein complex (Dilokpimol et al. 2014)
In mature ovary cryosections of T. fournieri, the β-1,3-galactan of AG is abundant in the ovule and cell layer of the placenta surface, and AG accumulation could only be detected by β-Yariv reagents rather than α-Yariv reagents, indicating a high specificity of this chemical compound (Mizukami et al. 2016). In Z. elegans, the function of xylogen, an AGP required for xylem differentiation, was impaired in the presence of β-Yariv but not α-Yariv reagents (Motose et al. 2004). In N. tabacum, the size of prefertilised ovules significantly decreased with increasing concentrations of β-Yariv, but not α-Yariv reagents, in the culture medium (Qin and Zhao 2006). Overall, studies based on β-Yariv reagents have improved our knowledge of the functions of AGPs.
Degradation enzymes of AG sugar chains
Many glycoside hydrogenases of microbial and plant origin have been identified (reviewed in Knoch et al. 2014). These enzymes are important for the metabolism of AG sugar chains, including AG chain turnover. In addition, these enzymes are useful tools for studying the function of AG sugar chains. To examine the function of the sugar moiety of AGPs, acid treatment to remove the sugar moiety and conventional glycosidase treatment, non-specific to AG sugar chains, have been used. However, it is debated as to whether acid treatment can damage protein backbones and whether glycosidase treatment, with a wide spectrum of activity, can affect non-AGP glycoproteins in the fraction. Using degradation enzymes targeted to specific structures of AG sugar chains, the function of AG sugar chains can be examined in a more reliable manner. Moreover, it is possible to characterise the critical structures of AG sugar chains with respect to their function. For example, AMOR in T. fournieri maintained its activity even when the β-1,6-galactan side chain (Fig. 3) was digested by endo-β-1,6-galactanase and α-L-arabinofuranosidase, but lost its activity when treated with β-glucuronidase, which removed β-glucuronosyl and 4-O-methyl-glucuronosyl residues at the terminal (for details of these enzyme treatment, see also Kotake et al. 2004; Konishi et al. 2008; Takata et al. 2010). This led to identification of the terminal disaccharide structure, 4-Me-GlcA-β-1,6-Gal, as the structure responsible for AMOR activity (Mizukami et al. 2016).
Degradation enzymes are also useful for glycan structure analysis to release oligosaccharides from AG sugar chains (e.g. Tryfona et al. 2012) and to quantify sugars in AG fractions (e.g. Mizukami et al. 2016).
Chemical synthesis of AG sugar chains
Chemical synthesis of specific components of AG sugar chains is a powerful method to obtain large amounts of pure molecule. Chemically synthesised AG sugar chains have been used to elucidate epitopes of monoclonal antibodies (e.g. ELISA and the aforementioned microarray). For example, chemical synthesis was sufficiently powerful to show that 4-Me-GlcA-β-1,6-Gal has specific AMOR activity (Mizukami et al. 2016; Jiao et al. 2017), whereas the α isomer of the disaccharide showed less activity (approx. 1/100), and a disaccharide without the methyl residue showed no AMOR activity. Chemically synthesised disaccharide AMOR is now commercially available from a Japanese company (Tokyo Chemical Industry Co Ltd). Chemical synthesis must be sufficiently powerful to search for other structures responsible for the bioactivity of AG sugar chains, although chemical synthesis of long and branched AG sugar chains, like those of native AGPs, is difficult.
Omics approaches to identify AGPs
The omics era has yielded substantial increases in big data in biology. Moreover, bioinformatics approaches have proven able to efficiently mine AGP sequences from published genomic and transcriptomic databases (Schultz et al. 2002; Showalter et al. 2010; Johnson et al. 2017a, b; Ma et al. 2017). Developed using a Perl script, the BIO OHIO software program, which is based on the proportion of PAST in each protein and specific amino acid motifs associated with known hydroxyproline-rich glycoproteins (HRGPs), has successfully identified 85 AGPs, moderately glycosylated EXTs, 18 lightly glycosylated proline-rich proteins, and other HRGPs from the A. thaliana genome (Schultz et al. 2002; Showalter et al. 2010). Based on the original version, an improved version of BIO OHIO (ver. 2.0) was released to screen for HRGPs from the complete genome of Populus trichocarpa (Showalter et al. 2016).
Based on BIO OHIO ver. 2.0, a small Python script, “Finding-AGP,” has successfully identified AGP gene family members from various lineages of the plant kingdom (Ma et al. 2017). The Finding-AGP program can not only discover AGPs with large proportions of PAST, but can also identify chimeric AGPs with smaller PAST percentages, covering more than half of all AGPs (Ma et al. 2017). However, more accurate, manual curation of the output results is still required, which would be time-consuming when dealing with large datasets. More recently, an automated motif and amino acid bias (MAAB) pipeline was introduced to classify HRGP sequences and has been applied to the 1000 Plants project, which includes sequencing data for over 1,000 plant species (Johnson et al. 2017a, b).
Biosynthesis of AG sugar chains
Post-translational modification of AG sugar chains to the protein backbone occurs in the secretory pathway of cells. O-glycosylation is the predominant form of AGP glycosylation. Hydroxylated prolines (Hyps) in repetitive dipeptide motifs (e.g. alanine–proline, serine–proline, threonine–proline, and valine–proline) are first glycosylated with a galactose to form Gal-β-1,4-Hyp. This initiation is critical for the successful elongation of AG sugar chains. The enzymes in this initiation, hydroxyproline O-galactosyltransferases (HPGTs), and their genes have been identified (Basu et al. 2013; Ogawa-Ohnishi and Matsubayashi 2015). HPGT1, 2, 3 of A. thaliana are three potent HPGTs, in which a triple loss-of-function mutant resulted in considerable reductions in β-Yariv-precipitated AGPs (Ogawa-Ohnishi and Matsubayashi 2015; Fig. 3). In this triple mutant, many plant growth and development processes were defective, including plant reproduction, suggesting that AG sugar chains are involved in various plant growth and development processes.
Other glycosyltransferases required for the step-wise elongation of AG chains have been identified, including KAONASHI4 (KNS4)/UNEVEN PATTERN OF EXINE1 (UPEX1) as a b-(1,3)-galactosyltransferase for main chains (Suzuki et al. 2017), GALT29A and GALT31A as β-(1,6)-galactosyltransferases for side chains (Dilokpimol et al. 2014), FUT4 and FUT6 as α-1,2-fucosyltransferases (Wu et al. 2010), and RAY1 as an arabinofuranosyltransferase (Gille et al. 2013) (Fig. 3; reviewed in Knoch et al. 2014). A loss-of-function mutant of AtGALT31A showed defects in embryogenesis (Geshi et al. 2013). Related to AMOR in T. fournieri, β-glucuronosyltransferases that add a glucuronic acid to galactan, AtGlcAT14A, B, C have been identified in A. thaliana (Knoch et al. 2013; Dilokpimol and Geshi 2014). Detailed analysis of mutants defective in AtGlcAT14 genes is of interest to examine how these genes and β-glucuronosyl residues of AG chains are involved in plant reproduction in A. thaliana. The identification of AMOR and differential localisation of specific structures of AG sugar chains suggest that the functions of AGPs and their sugar chains are locally controlled by these AG biosynthesis enzymes.
Conclusions
From gametogenesis to fertilisation, AGPs are recruited into crosstalk between male and female parts, and between sporophyte and gametophyte generations. AGPs possibly act as plant wall components, calcium capacitors, and nutritional or signalling molecules (Tan et al. 2013; Lamport et al. 2018). However, it remains unknown how these AGPs and their sugar chains function during plant reproduction processes, which is of great academic interest.
In future research, elucidation of the mechanisms of AGPs, and especially their interactors, will require a combination of multidisciplinary tools, as partly summarised here. Decoding the precise structure, and dissecting the functions, of AG sugar chains linked to the core proteins will be challenging. However, achieving an integrated understanding of the roles of AGPs and their sugar chains will help unveil the mysteries of plant reproductive processes and could ultimately lead to agricultural advances.
References
Acosta-Garcia G, Vielle-Calzada JP (2004) A classical arabinogalactan protein is essential for the initiation of female gametogenesis in Arabidopsis. Plant Cell 16:2614–2628
Basu D, Liang Y, Liu X, Himmeldirk K, Faik A, Kieliszewski M, Held M, Showalter AM (2013) Functional identification of a hydroxyproline-O-galactosyltransferase specific for arabinogalactan protein biosynthesis in Arabidopsis. J Biol Chem 288:10132–10143
Borg M, Brownfield L, Twell D (2009) Male gametophyte development: a molecular perspective. J Exp Bot 60:1465–1478
Cao J, Li X, Lv Y, Ding L (2015) Comparative analysis of the phytocyanin gene family in 10 plant species: a focus on Zea mays. Front Plant Sci 6:515
Cheung AY, Wang H, Wu HM (1995) A floral transmitting tissue-specific glycoprotein attracts pollen tubes and stimulates their growth. Cell 82:383–393
Coimbra S, Almeida J, Junqueira V, Costa ML, Pereira LG (2007) Arabinogalactan proteins as molecular markers in Arabidopsis thaliana sexual reproduction. J Exp Bot 58:4027–4035
Coimbra S, Jones B, Pereira LG (2008) Arabinogalactan proteins (AGPs) related to pollen tube guidance into the embryo sac in Arabidopsis. Plant Signal Behav 3:455–456
Coimbra S, Costa M, Jones B, Mendes MA, Pereira LG (2009) Pollen grain development is compromised in Arabidopsis Agp6 Agp11 null mutants. J Exp Bot 60:3133–3142
Corral-Martinez P, Garcia-Fortea E, Bernard S, Driouich A, Seguí-Simarro JM (2016) Ultrastructural immunolocalization of arabinogalactan protein, pectin and hemicellulose epitopes through anther development in Brassica napus. Plant Cell Physiol 57:2161–2174
Da CM, Lopes AL, Amorim MI, Coimbra S (2017) Immunolocalization of AGPs and pectins in Quercus suber Gametophytic Structures. Methods Mol Biol 1669:117–137
Demesa-Arevalo E, Vielle-Calzada JP (2013) The classical arabinogalactan protein AGP18 mediates megaspore selection in Arabidopsis. Plant Cell 25:1274–1287
Dilokpimol A, Geshi N (2014) Arabidopsis thaliana glucuronosyltransferase in family GT14. Plant Signal Behav 9:e28891
Dilokpimol A, Poulsen CP, Vereb G, Kaneko S, Schulz A, Geshi N (2014) Galactosyltransferases from Arabidopsis thaliana in the biosynthesis of type II arabinogalactan: molecular interaction enhances enzyme activity. BMC Plant Biol 14:90
Eberle CA, Anderson NO, Clasen BM, Hegeman AD, Smith AG (2013) PELPIII: the class III pistil-specific extensin-like Nicotiana tabacum proteins are essential for interspecific incompatibility. Plant J 74:805–814
Ellis M, Egelund J, Schultz CJ, Bacic A (2010) Arabinogalactan-proteins: key regulators at the cell surface? Plant Physiol 153:403–419
Fincher GB, Sawyer WH, Stone BA (1974) Chemical and physical properties of an arabinogalactan-peptide from wheat endosperm. Biochem J 139:535–545
Fincher GB, Stone BA, Clarke AE (1983) Arabinogalactan-proteins: structure, biosynthesis and functions. Annu Rev Plant Physiol 34:47–70
Gane AM, Craik D, Munro SL, Howlett GJ, Clarke AE, Bacic A (1995) Structural analysis of carbohydrate moiety of arabinogalactan-proteins from stigma and styles of Nicotiana alata. Carbohydr Res 277:67–85
Geshi N, Johansen JN, Dilokpimol A, Rolland A, Belcram K, Verger S, Kotake T, Tsumuraya Y, Kaneko S, Tryfona T, Dupree P, Scheller HV, Höfte H, Mouille G (2013) A galactosyltransferase acting on arabinogalactan protein glycans is essential for embryo development in Arabidopsis. Plant J 76:128–137
Gille S, SharmaV Baidoo EEK, Keasling JD, Scheller HV, Pauly M (2013) Arabinosylation of a yariv-precipitable cell wall polymer impacts plant growth as exemplified by the Arabidopsis glycosyltransferase mutant ray1. Mol Plant 6:1369–1372
Hancock CN, Kent L, McClure BA (2005) The stylar 120 kDa glycoprotein is required for S-specific pollen rejection in Nicotiana. Plant J 43:716–723
Hou Y, Guo X, Cyprys P, Zhang Y, Bleckmann A, Cai L, Huang Q, Luo Y, Gu H, Dresselhaus T, Dong J, Qu LJ (2016) Maternal ENODLs are required for pollen tube reception in Arabidopsis. Curr Biol 26:2343–2350
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator V3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinform 2008:420747
Jiao J, Mizukami AG, Sankaranarayanan S, Yamguchi J, Itami K, Higashiyawma T (2017) Structure-activity relation of AMOR sugar molecule that activates pollen-tubes for ovular guidance. Plant Physiol 173:354–363
Johnson KL, Jones BJ, Bacic A, Schultz CJ (2003) The fasciclin-like arabinogalactan proteins of Arabidopsis. A multigene family of putative cell adhesion molecules. Plant Physiol 133:1911–1925
Johnson KL, Cassin AM, Lonsdale A, Wong GK, Soltis DE, Miles NW, Melkonian M, Melkonian B, Deyholos MK, Leebens-Mack J, Rothfels CJ, Stevenson DW, Graham SW, Wang X, Wu S, Pires JC, Edger PP, Carpenter EJ, Bacic A, Doblin MS, Schultz CJ (2017a) Insights into the evolution of hydroxyproline-rich glycoproteins from 1000 plant transcriptomes. Plant Physiol 174:904–921
Johnson KL, Cassin AM, Lonsdale A, Bacic A, Doblin MS, Schultz CJ (2017b) Pipeline to Identify hydroxyproline-rich glycoproteins. Plant Physiol 174:886–903
Kitazawa K, Tryfona T, Yoshimi Y, Hayashi Y, Kawauchi S, Antonov L, Tanaka H, Takahashi T, Kaneko S, Dupree P, Tsumuraya Y, Kotake T (2013) β-Galactosyl Yariv reagent binds to the beta-1,3-galactan of arabinogalactan proteins. Plant Physiol 161:1117–1126
Knoch E, Dilokpimol A, Tryfona T, Poulsen CP, Xiong G, Harholt J, Petersen BL, Ulvskov P, Hadi MZ, Kotake T, Tsumuraya Y, Pauly M, Dupree P, Geshi N (2013) A β-glucuronosyltransferase from Arabidopsis thaliana involved in biosynthesis of type II arabinogalactan has a role in cell elongation during seedling growth. Plant J 76:1016–1029
Knoch E, Dilokpimol A, Geshi N (2014) Arabinogalactan proteins: focus on carbohydrate active enzymes. Front Plant Sci 5:198
Kobayashi Y, Motose H, Iwamoto K, Fukuda H (2011) Expression and genome-wide analysis of the xylogen-type gene family. Plant Cell Physiol 52:1095–1106
Konishi T, Kotake T, Soraya D, Matsuoka K, Koyama T, Kaneko S, Igarashi K, Samejima M, Tsumuraya Y (2008) Properties of family 79 beta-glucuronidases that hydrolyze beta-glucuronosyl and 4-O-methyl-beta-glucuronosyl residues of arabinogalactan-protein. Carbohydr Res 343:1191–1201
Kotake T, Kaneko S, Kubomoto A, Haque MA, Kobayashi H, Tsumuraya Y (2004) Molecular cloning and expression in Escherichia coli of a Trichoderma viride endo-β-(1 → 6)-galactanase gene. Biochem J 377:749–755
Lamport D, Tan L, Held MA, Kieliszewski MJ (2018) Pollen tube growth and guidance: occam’s razor sharpened on a molecular arabinogalactan glycoprotein Rosetta Stone. New Phytol 217:491–500
Levitin B, Richter D, Markovich I, Zik M (2008) Arabinogalactan proteins 6 and 11 are required for stamen and pollen function in Arabidopsis. Plant J 56:351–363
Li J, Yu M, Geng LL, Zhao J (2010) The fasciclin-like arabinogalactan protein gene, FLA3, is involved in microspore development of Arabidopsis. Plant J 64:482–497
Lin S, Dong H, Zhang F, Qiu L, Wang F, Cao J, Huang L (2014) BcMF8, a putative arabinogalactan protein-encoding gene, contributes to pollen wall development, aperture formation and pollen tube growth in Brassica campestris. Ann Bot 113:777–788
Lind JL, Bacic A, Clarke AE, Anderson MA (1994) A style-specific hydroxyproline-rich glycoprotein with properties of both extensins and arabinogalactan proteins. Plant J 6:491–502
Losada JM, Herrero M (2012) Arabinogalactan-protein secretion is associated with the acquisition of stigmatic receptivity in the apple flower. Ann Bot 110:573–584
Ma Y, Yan C, Li H, Wu W, Liu Y, Wang Y, Chen Q, Ma H (2017) Bioinformatics prediction and evolution analysis of arabinogalactan proteins in the plant kingdom. Front Plant Sci 8:66
Majewska-Sawka A, Nothnagel EA (2000) The multiple roles of arabinogalactan proteins in plant development. Plant Physiol 122:3–10
Mashiguchi K, Asami T, Suzuki Y (2009) Genome-wide identification, structure and expression studies, and mutant collection of 22 early nodulin-like protein genes in Arabidopsis. Biosci Biotechnol Biochem 73:2452–2459
Mizukami AG, Inatsugi R, Jiao J, Kotake T, Kuwata K, Ootani K, Okuda S, Sankaranarayanan S, Sato Y, Maruyama D, Iwai H, Garénaux E, Sato C, Kitajima K, Tsumuraya Y, Mori H, Yamaguchi J, Itami K, Sasaki N, Higashiyama T (2016) The AMOR arabinogalactan sugar chain induces pollen-tube competency to respond to ovular guidance. Curr Biol 26:1091–1097
Moller I, Marcus SE, Haeger A, Verhertbruggen Y, Verhoef R, Schols H, Ulvskov P, Mikkelsen JD, Knox JP, Willats W (2008) High-throughput screening of monoclonal antibodies against plant cell wall glycans by hierarchical clustering of their carbohydrate microarray binding profiles. Glycoconj J 25:37–48
Motose H, Sugiyama M, Fukuda H (2004) A proteoglycan mediates inductive interaction during plant vascular development. Nature 429:873–878
Nguema-Ona E, Coimbra S, Vicre-Gibouin M, Mollet JC, Driouich A (2012) Arabinogalactan proteins in root and pollen-tube cells: distribution and functional aspects. Ann Bot 110:383–404
Nguema-Ona E, Vicre-Gibouin M, Cannesan MA, Driouich A (2013) Arabinogalactan proteins in root-microbe interactions. Trends Plant Sci 18:440–449
Ogawa-Ohnishi M, Matsubayashi Y (2015) Identification of three potent hydroxyproline O-galactosyltransferases in Arabidopsis. Plant J 81:736–746
Olmos E, Garcia DLGJ, Gomez-Jimenez MC, Fernandez-Garcia N (2017) Arabinogalactan proteins are involved in salt-adaptation and vesicle trafficking in tobacco BY-2 cell cultures. Front Plant Sci 8:1092
Penell RI, Janniche L, Kjellbom P, Scofield GN, Peart JM, Roberts K (1991) Developmental regulation of a plasma membrane arabinogalactan protein epitope in Oilseed rape flowers. Plant Cell 3:1317–1326
Pennell RI, Roberts K (1990) Sexual development in the pea is presaged by altered expression of arabinogalactan protein. Nature 344:547–549
Pereira LG, Coimbra S, Oliveira H, Monteiro L, Sottomayor M (2006) Expression of arabinogalactan protein genes in pollen tubes of Arabidopsis thaliana. Planta 223:374–380
Pereira AM, Masiero S, Nobre MS, Costa ML, Solís MT, Testillano PS, Sprunck S, Coimbra S (2014) Differential expression patterns of arabinogalactan proteins in Arabidopsis thaliana reproductive tissues. J Exp Bot 65:5459–5471
Pereira AM, Pereira LG, Coimbra S (2015) Arabinogalactan proteins: rising attention from plant biologists. Plant Reprod 28:1–15
Pereira AM, Lopes AL, Coimbra S (2016a) Arabinogalactan proteins as interactors along the crosstalk between the pollen tube and the female tissues. Front Plant Sci 7:1895
Pereira AM, Nobre MS, Pinto SC, Lopes AL, Costa ML, Masiero S, Coimbra S (2016b) “Love is strong, and you’re so sweet”: JAGGER is essential for persistent synergid degeneration and polytubey block in Arabidopsis thaliana. Mol Plant 9:601–614
Qin Y, Zhao J (2006) Localization of arabinogalactan proteins in egg cells, zygotes, and two-celled proembryos and effects of beta-D-glucosyl Yariv reagent on egg cell fertilization and zygote division in Nicotiana tabacum L. J Exp Bot 57:2061–2074
Ruprecht C, Bartetzko MP, Senf D, Dallabernadina P, Boos I, Andersen MCF, Kotake T, Knox JP, Hahn MG, Clausen MH, Pfrengle F (2017) A synthetic glycan microarray enables epitope mapping of plant cell wall glycan-directed antibodies. Plant Physiol 175:1094–1104
Sankaranarayanan S, Higashiyama T (2018) Capacitation in plant and animal fertilization. Trends Plant Sci 23:129–139
Schultz CJ, Johnson KL, Currie G, Bacic A (2000) The classical arabinogalactan protein gene family of Arabidopsis. Plant Cell 12:1751–1768
Schultz CJ, Rumsewicz MP, Johnson KL, Jones BJ, Gaspar YM, Bacic A (2002) Using genomic resources to guide research directions: the arabinogalactan protein gene family as a test case. Plant Physiol 129:1448–1463
Seifert GJ, Roberts K (2007) The biology of arabinogalactan proteins. Annu Rev Plant Biol 58:137–161
Showalter AM (2001) Arabinogalactan-proteins: structure, expression and function. Cell Mol Life Sci 58:1399–1417
Showalter AM, Keppler B, Lichtenberg J, Gu D, Welch LR (2010) A bioinformatics approach to the identification, classification, and analysis of hydroxyproline-rich glycoproteins. Plant Physiol 153:485–513
Showalter AM, Keppler BD, Liu X, Lichtenberg J, Welch LR (2016) Bioinformatic identification and analysis of hydroxyproline-rich glycoproteins in Populus trichocarpa. BMC Plant Biol 16:229
Sun W, Xu J, Yang J, Kieliszewski MJ, Showalter AM (2005) The lysine-rich arabinogalactan-protein subfamily in Arabidopsis: gene expression, glycoprotein purification and biochemical characterization. Plant Cell Physiol 46:975–984
Suzuki T, Narciso JO, Zeng W, van de Meene A, Yasutomi M, Takemura S, Lampugnani ER, Doblin MS, Bacic A, Ishiguro S (2017) KNS4/UPEX1: a type II arabinogalactan beta-(1,3)-galactosyltransferase required for pollen exine development. Plant Physiol 173:183–205
Takata R, Tokita K, Mori S, Shimoda R, Harada N, Ichinose H, Kaneko S, Igarashi K, Samejima M, Tsumuraya Y (2010) Degradation of carbohydrate moieties of arabinogalactan-proteins by glycoside hydrolases from Neurospora crassa. Carbohydr Res 345:2516–2522
Tan H, Liang W, Hu J, Zhang D (2012) MTR1 encodes a secretory fasciclin glycoprotein required for male reproductive development in rice. Dev Cell 22:1127–1137
Tan L, Eberhard S, Pattathil S, Warder C, Glushka J, Yuan C, Hao Z, Zhu X, Avci U, Miller JS, Baldwin D, Pham C, Orlando R, Darvill A, Hahn MG, Kieliszewski MJ, Mohnen D (2013) An Arabidopsis cell wall proteoglycan consists of pectin and arabinoxylan covalently linked to an arabinogalactan protein. Plant Cell 25:270–287
Tryfona T, Liang HC, Kotake T, Tsumuraya Y, Stephens E, Dupree P (2012) Structural characterization of Arabidopsis leaf arabinogalactan polysaccharides. Plant Physiol 160:653–666
Tsutsui H, Higashiyama T (2017) pKAMA-ITACHI vectors for highly efficient CRISPR/Cas9-mediated gene knockout in Arabidopsis thaliana. Plant Cell Physiol 58:46–56
Willats WG, Marcus SE, Knox JP (1998) Generation of monoclonal antibody specific to (1-5)-alpha-L-arabinan. Carbohydr Res 308:149–152
Wu HM, Wang H, Cheung AY (1995) A pollen tube growth stimulatory glycoprotein is deglycosylated by pollen tubes and displays a glycosylation gradient in the flower. Cell 82:395–403
Wu HM, Wong E, Ogdahl J, Cheung AY (2000) A pollen tube growth-promoting arabinogalactan protein from Nicotiana alata is similar to the tobacco TTS protein. Plant J 22:165–176
Wu Y, Williams M, Bernard S, Driouich A, Showalter AM, Faik A (2010) Functional identification of two nonredundant Arabidopsis alpha(1,2)fucosyltransferases specific to arabinogalactan proteins. J Biol Chem 285:13638–13645
Yang WC, Shi DQ, Chen YH (2010) Female gametophyte development in flowering plants. Annu Rev Plant Biol 61:89–108
Yariv J, Kalb AJ, Katchalski E (1967) Isolation of an L-fucose binding protein from Lotus tetragonolobus seed. Nature 215:890–891
Yates EA, Valdor JF, Haslam SM, Morris HR, Dell A, Mackie W, Knox JP (1996) Characterization of carbohydrate structural features recognized by anti-arabinogalactan-protein monoclonal antibodies. Glycobiology 6:131–139
Acknowledgements
TH was supported by the Japan Society for the Promotion of Science (Grant-in-Aid for Scientific Research on Innovative Areas Nos. JP16H06465, JP16H06464, and JP16K21727) and by Mizutani Foundation for Glycoscience (No. 180190).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Mengxiang Sun.
A contribution to the special issue ‘Plant Reproduction Research in Asia’.
Rights and permissions
About this article
Cite this article
Su, S., Higashiyama, T. Arabinogalactan proteins and their sugar chains: functions in plant reproduction, research methods, and biosynthesis. Plant Reprod 31, 67–75 (2018). https://doi.org/10.1007/s00497-018-0329-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00497-018-0329-2