Abstract
Due to environmental concern, the research to date has tended to focus on how textile dye removal can be carried out in a greener manner. Therefore, this study aims to evaluate the decolorization and biotransformation pathway of Mordant Orange-1 (MO-1) by Cylindrocephalum aurelium RY06 (C. aurelium RY06). Decolorization study was conducted in a batch experiment including the investigation of the effects of physio-chemical parameters. Enzymatic activity of C. aurelium RY06 during the decolorization was also investigated. Moreover, transformation and biodegradation of MO-1 by C. aurelium RY06 were observed using the gas chromatography–mass spectrometry. Manganese peroxidase, lignin peroxidase, laccase, 1,2-dioxygenase, and 2,3-dioxygenase enzymes were detected during the decolorization. In general, the present work concluded that the MO-1 was successfully degraded by C. aurelium RY06 and transformed to be maleic acid and to be isophtalic acid.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
As it is well known, a wide variety of colors has been extensively used for textile dyeing and paper painting [1]. Azo dyes are the most commonly used in textile industry. During the dyeing process, approximately 10–15% of the used dyes can be released into the environment, probably causing serious environmental and health problems [2, 3]. The presence of dye in the water environment can affect photosynthetic activity because it can cut light access as well as causes a shortage of oxygen thus decreasing the survival rate of flora and fauna [4]. In addition, most of dyes released from industries are greatly toxic and mutagenic agent [5]. Therefore, it is a major environmental challenge to remove dyes from industrial effluents since they are difficult to treat by conventional methods [6].
Although decolorization of dyes can be carried out using physio-chemical methods such as coagulation/adsorption, precipitation or chemical degradation, they have shortcomings such as high energy consumption, high cost, and hazardous by-product production [7,8,9,10]. Other conventional aerobic treatment processes were usually not efficient for decolorization of the effluents contaminated with azo dyes because of their strong electron-withdrawing group, which protects them against attack by the oxygenases [11]. Hence, decolorization by exploring existing microorganisms is becoming attractive because it is cost-effective and environmentally friendly, and has the potential to produce less sludge compared to chemical and physical treatments [12]. In addition, current development trends have been focused on how it can be carried out in a greener and cheaper manner as well as sustainability. Aligning with the aforementioned issues, it is evidenced in scientific literature that the dye decolorization can be carried out by microorganisms such as bacteria, yeast, algae, actinomycetes, and fungi. The capability of microorganisms to decolorize and mineralize through breakdown of intramolecular bond of organic compounds has been explored. In addition, the use of microorganisms can provide a new contribution for development of different bioremediation technologies. Among these, bacteria and fungi are capable to degrade and decolorize azo dyes and polycyclic aromatic hydrocarbons. Fungi provide many advantages and some characteristics of fungi make them better potential as a degrader compared to bacteria. In addition, they have also the ability to resist unfavorable environments [13].
Recently, there has been renewed interest in exploring fungi to decolorize and biodegrade dyes. For instance, the decolorization of Reactive Blue 268 by Aspergillus fumigatus can be obtained up to 65% removal [14]. The decolorization and biodegradation of triphenylmethane dyes, which are cotton blue and crystal violet by Coriolopsis sp. (1c3) were carried out and found significant decolorization activities up to 85% [15]. Alternatively, a newly isolated fungus, Achaetomium strumarium can provide up to 99% decolorization of acid red 88 dye [16]. Biodecolorization of brilliant green carpet industry dye using three fungi, which are Pleurotus florida (PF), Pleurotus eryngii (PE), and Pleurotus sajor-caju (PS), have also been evaluated and confirmed the highest decolorization potentialities recorded in the ordering as 99 > 91 > 83% by PF > PE > PS, respectively [17]. Using another fungus, biodegradation scarlet RR dye present in the textile industry effluent by Peyronellaea prosopidis can also be carried out and found to reduce 68, 88 and 91% of biological oxygen demand, chemical oxygen demand, and color intensity of the wastewater [18]. Moreover, dye-containing wastewaters can be treated by Trametes villosa SCS-10 [19]. Their study exhibited that up to 90% of color removal, and 80% of COD and TOC reduction can be achieved.
For the mechanism, there are two major processes for decolorization of dyes using fungi, which are biodegradation and biosorption. The biodegradation is specifically dedicated to living cells because they can produce the lignin-modifying enzymes, laccase, manganese peroxidase, and lignin peroxidase. The role of laccase, manganese peroxidase, and lignin peroxidase for decolorization of dyes may be different for each fungus. For instance, lignin peroxidase secreted by Phanerochaete chrysosporium was responsible for the decolorization of azo dyes [20]. A similar finding was also found when the fungus was employed to decolorize azo, triphenyl methane, heterocyclic and polymeric dyes [21]. A different finding was reported when the wood rotting fungus was used and their study exhibited that the manganese peroxidase played an important role in the decolorization of cotton bleaching effluent [22]. Alternatively, laccase released from Tinea versicolor and Cyathus bulleri was observed to be responsible as an enzyme degrader in decolorizing anthraquinone, azo, and indigo dyes [23, 24]. Another mechanism to remove dyes is biosorption that is basically devoted for dead cells. In this mechanism, the involvement of physio-chemical interactions such as adsorption, deposition, and ion exchange is responsible for the removal. However, the use of dead cells is commonly associated with their lower removal capacity compared to living cells as validated by investigation of Congo red decolorization by Phanerochaete chrysosporium, performing 90% and 70% removal for the living and dead cells, respectively.
Azo dyes have a complex structure and synthetic origin; as a result, this makes that it is quite difficult to be decolorized. Mordant Orange-1 (MO-1) is an azo dye characterized by the presence of at least one azo bond (–N = N–) bearing aromatic rings and has high photolytic stability and resistance towards major oxidizing agents to avoid degradation of dyes [25]. Since removal capability of fungi depends highly on the type of fungi, dyes, medium, and environmental condition, the biodegradation products formed after decolorization by fungi is also quite specific. Aligning the aforementioned research question, there is a need for research on degradation product validation formed by new potential fungus to degrade MO-1. The metabolic pathways utilized may differ and be interesting to be investigated. Hence, the aim of this study was to evaluate the capability of a newly isolated C. aurelium RY06 to decolorize and biodegrade MO-1. Also, the effect of various culture conditions such as agitation speeds, pH, carbon sources, nitrogen sources, and surfactant was completely investigated. As a new knowledge and novelty, enzymatic activity and metabolic pathway for the decolorization of MO-1 by C. aurelium RY06 were also evaluated. Although the capability of fungi to degrade some textile dyes has been reported elsewhere, novelty and contribution of this study are on the proposed new mechanism and transformation, which may be different compared to previous studies since degradation mechanism and transformation of dyes depend highly on the types of dye and fungi as well as employed procedures. Hence, findings from this study are useful for the contribution to current development of bioremediation technologies.
Materials and methods
Dye and chemicals
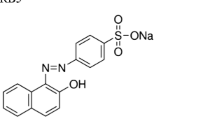
MO-1 was purchased from the Sigma-Aldrich, the United States of America. The properties of MO-1 are listed in Table 1. In addition, fructose, glucose, galactose, ammonium nitrate, ammonium sulfate, yeast extract, and Tween 80 were obtained from the Merck, Germany.
Identification of the proposed fungus
In this study, potential fungal strain was isolated from a tropical rain forest in the surrounding of the Universiti Teknologi Malaysia. Identification of the proposed fungus was carried out on the Petri dishes containing the potato dextrose agar and malt extract agar. In this study, polymerase chain reaction analysis was employed for identification purpose. The preparation product was sent to 1st BASE Laboratory Sdn. Bhd for sequencing. The DNA sequence was then read and edited using BioEdit and compared with other 18S rRNA gene sequences obtained from the National Center for Biotechnology Information (NCBI) GenBank database. It was confirmed that the identified fungus can be categorized as C. aurelium RY06.
Microorganism and culture condition
Cylindrocephalum aurelium RY06 having a size of 3 mm in diameter and about 1 g of weight was first cultured in a conical flask (250 mL) with liquid medium on an orbital shaker at a speed of 120 rpm. The liquid medium (100 mL in a volume) contains yeast extract (2 g) and glucose (2 g). The dishes with 20 mL agar medium containing glucose (20 g/L), malt extract agar (20 g/L), chloramphenicol (0.04 g/L), and ammonium nitrate (10 g/L) were inoculated with the mycelia (3 mm in diameter and 1 g of weight) cut from the body of mycelia. All medium and experimental equipments were sterilized using autoclave at a temperature of 120 °C for 15 min.
Decolorization experiment
Decolorization of MO-1 was performed using a batch study carried out in 100-mL Erlenmeyer flasks. The flasks were prepared in triplicates and contained 20 ml nutrient media with dyestuff. The agar plates and flasks were used under sterile condition. Inoculation process was carried out using a 3-mm active plug cut from the pure fungal culture grown on agar plates. The liquid medium cultures for decolorization were incubated for 30 days in dark room condition. The effect of initial pH was evaluated by performing ranging pH from 3 to 6. The pH was adjusted using 0.1 M hydrochloric acid (HCl) or 0.1 M sodium hydroxide (NaOH). Next, the effect of agitation was conducted at different speeds of 0, 50, 75, and 100 rpm. The effects of various carbon and nitrogen sources were added at a final concentration of 20 g/L. All experimental parameters can be seen in Table 2. Samples (triplicate flasks) were taken periodically and centrifuged at 4000g for 20 min at 15 °C, and the clear supernatant obtained was used to determine the percentage of decolorization by measuring absorbance in a UV–Vis Spectrometer DR 5000 at a wavelength of 385 nm. The percentage of decolorization was estimated using the following equation:
where DP is the decolorization percentage (%), C0 the initial concentration of the dye (mg/L) and C is the final concentration of the dye after decolorization (mg/L).
Investigation of enzymatic activity
Investigation of enzymatic activity was carried out spectroscopically using UV–Vis spectrophotometer. It is noted that one unit of activity is defined as the amount of enzyme that oxidized 1 μmol of substrate per minute and it was expressed in units per volume (U/L). For this purpose, investigation of laccase activity was carried out via using syringaldazine in 100 mM sodium acetate buffer. An activity of manganese peroxidase was examined using 50 mM malonate buffer and dimethoxyphenol in 20 mM MnSO4. In addition, lignin peroxidase activity was determined using veratryl alcohol. Moreover, 1,2-dioxygenase and 2,3-dioxygenase activities were observed via catechol as a substrate. It is noted that the experiments were performed in triplicates.
Characterizations
For the Fourier-transform infrared spectroscopy (FTIR) characterization, samples were examined using the FTIR spectrometer (PerkinElmer Frontier-GPOB model) equipped with the PerkinElmer Spectrum software was employed for this characterization. It used OptKBr (7800–400 cm−1) as beam splitter and MIR TGS (15,000–370 cm−1) as detector. Samples in a pellet form were prepared by hydraulic pressing (Specac model) at 10 tons in pressure. FTIR characterization was inspected using a spectrum wavelength in the range of 650–4000 cm−1 at a resolution of 4 cm−1 and accumulations of ten scans. In addition, changes in fungal morphologies were also characterized by means of field-emission scanning electron microscopy (FESEM ZEISS Supra 35VP). For this investigation, the fungal samples were collected before and after decolorization.
Identification of metabolites
The biodegradation products of MO-1 were inspected by means of gas chromatography–mass spectrometry (GC–MS). For the GC–MS analysis, extract cultures were first prepared by condensing using rotary evaporators and passed through a column. The GC–MS system equipped with DB-1 capillary column was employed for metabolite inspections. It was operated at the splitless injector temperature of 260 °C with an oven condition of 80 °C for 1 min, increased up to 160 °C with a rate of 18 °C min−1, raised up to 320 °C with a rate of 20 °C min−1, and then held at 320 °C for 15 min. This examination was operated in full scan of m/z 50–500. It was then followed by examining the retention time of the suspected compound with MO-1. The mass spectrometry of the sample was compared with the corresponding authentic standards and the mass spectra library of Wiley 275L.
Results and discussion
Effect of agitation speed
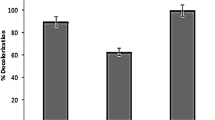
The mechanism of biodegradation of dyes can be linked to several processes, which are: (1) catalyzation of the dye by extracellular enzyme and (2) adsorption process through the biomass with or without subsequent degradation inside the cells aided by intracellular enzymes. Effects of agitation speeds on MO-1 biodegradation by C. aurelium RY06 are presented in Fig. 1a. It can be observed that the agitation treatment can enhance the percentage of the decolorization compared to that of without agitation treatment. When the decolorization was carried out at a speed of 50 rpm, the removal percentage can be slightly enhanced from 41% (0 rpm) to 44%. In addition, due to increase in the speed of up to 70 rpm, the decolorization percentage was found to be 65%. At the highest speed used in this work (100 rpm), the decolorization percentage can be enhanced by up to 85%.
In general, increase in the agitation speed can increase the decolorization percentage. Maximum removal of MO-1 can be achieved when the agitation speed of 100 rpm was applied. Mechanical treatment such as agitation implemented in this study can enhance the distribution of nutrient such as carbon sources and nitrogen sources as well as transfer of oxygen in the liquid medium. As a result, it can promote cell growth and the enhancement of biodegradation process as also observed in the previous work [26]. Findings of this study were also in line with those results obtained by another study that employed Funalia Trogii to decolorize Astrazon Red FBL, which found the optimum decolorization can be obtained at agitation speeds of 100–150 rpm [5].
Effect of pH
Effects of pH on the decolorization percentage are shown in Fig. 1b. It can be observed that pH significantly affected the removal of MO-1. Increase in pH ranging from 3 to 6 can decrease the decolorization percentage ranging from 86 to 45%. The highest removal rate of MO-1 was found at pH 3 and the lowest degradation removal occurred at pH 6. It was possible that some essential nutrients in the growth media were suitable at pH 3. The optimum pH for decolorization of dyes depends highly on the type of fungi, dyes, medium, and environmental condition. It was reported in scientific literature that the optimum pH to degrade several dyes such as Reactive orange 16 [27], Remazol Black B [28], Poly R478, blue dextran, crystal violet, cresol red, bromophenol blue [29], and Remazol Brilliant Blue R [30] using fungi generally appeared in the range of 2–4. It was probably that pH values in this range also generally enrich and maintain the culture microorganisms well for optimal decolorization [31].
The results of this study were consistent with those obtained from previous studies addressing decolorization of synthetic dye by fungus at different pH ranges. For instance, the biosorption of Reactive orange 16 dye by Rhizopus arrhizus was optimum at pH 2 and can significantly decrease at higher pH (above 4) [27]. The dye uptake capacity of Rhizopus arrhizus to remove the reactive dye Remazol Black B was also optimum in the range pH around 2 with a sharp drop off at higher values (above 3) [28]. The use of several white fungi, which are Trametes cingulata, Trametes versicolor, Trametes pocas, DSPM95, Datronia concentrica, and Pycnoporus sanguineus, to degrade dyes (Poly R478, blue dextran, crystal violet, cresol red, and bromophenol blue) was optimum at pH 3–4.5 [29]. Moreover, the optimum pH for anthraquinonic dye Remazol Brilliant Blue R decolorization using Lentinus crinitus and Psilocybe castanella ranged from 2.5 to 3.5 [30]. The surface electrical charge of fungal biomass and the ionic forms of dye pollutant can be controlled by the solution pH value. The solution pH can alter both adsorbate chemistry and fungal biomass-binding sites. For instance, the optimum removal of acidic dye can be obtained at acidic condition because there was an increased in the protonation of weak base group (biomass) that bound and degraded the anionic group of acidic dye [32].
Effect of carbon sources
In this study, fructose, galactose, xylose, and glucose were added to the culture to validate the effects of carbon sources on the decolorization of MO-1 by C. aurelium RY06 as presented in Fig. 1c. The addition of glucose and fructose in the liquid medium can provide the decolorization percentages of 76 and 71%, respectively. In addition, the use of galactose and xylose as carbon sources can stimulate decolorization percentages of 65 and 35%, respectively. Findings from this study suggested that the use of glucose and fructose was the best option in terms of the removal percentage. However, it is obvious that a slightly higher removal percentage can be obtained when the glucose was used.
These results were in line with previous study that observed that an increase in incubation time and an addition of carbon sources gave a high enzymatic production and activity [33]. This could be attributed to the fact that the decolorization was mainly due to the extracellular enzyme activity [34]. By the presence of carbon sources in the culture, it was found that the decolorization process during the dye decolorization mechanism by the presently proposed fungi was highly correlated with the growth of fungi. Previous study revealed that the decolorization of MO-1 using Trichoderma harzianum with the addition of glucose can enhance the dye decolorization percentage [35].
Effect of nitrogen sources
Effects of nitrogen sources such as ammonium nitrate, ammonium sulfate, and yeast extract were investigated for decolorization of MO-1 by C. aurelium RY06 as shown in Fig. 1d. The addition of yeast extract, ammonium nitrate, and ammonium tartrate performed the decolorization percentages of 49, 71, and 64%, respectively. It was obvious that the addition of ammonium nitrate showed the highest decolorization percentages compared to others. It is noted that nitrogen is a basic requirement for growth of all fungi. Fungi can utilize a variety of nitrogen sources, including inorganic nitrate, ammonium salts, and some organic compounds especially amino acids.
The results indicated that the nitrogen sources affected the decolorization of the current dye. The different percentages reflected capacities of the filtrates to remove dyes with diverse chemical structures. This might be due to the differences in electron distribution, charge density or steric factors [37]. Previous study reported that the nitrogen sources can influence the produced amount and type of enzymes [38]. Therefore, it was speculated that the use of ammonium nitrate as a carbon source model can stimulate a higher enzymatic production and activities compared to other carbon source models. Carbon and nitrogen sources were used to enhance the enzyme production since they contained essential minerals and nutrients to be used as energy for growth [36].
Effect of surfactant
Surfactant can also be considered to affect the decolorization. For this purpose, its effect on the decolorization was examined by applying Tween 80 at different volumes, which are 0.1, 0.5, 1.0, and 1.5 mL. The use of this surfactant was due to their internal property to enhance the decolorization performance compared to another surfactant type such as Tween 20 [39]. Figure 1e shows the effect of the surfactant on the decolorization. The present work found that the percentages of decolorization of MO-1 at 0.1, 0.5, 1.0, and 1.5 mL were 27, 49, 85, and 49%, respectively.
In general, this study exhibited that the presence of surfactant might affect the growth of fungi during chemical reaction. Furthermore, these results confirmed that the addition of the surfactant at 1 mL can achieve the highest removal percentages of decolorization compared to others. The presence of surfactant can increase the solubility of dye in water [40]. As a result, the dye can be easily degraded. This was in agreement with those results obtained from the previous study that employed Rhizopus arrhizus to decolorize Remazol Blue dye [41]. Their study found that the presence of surfactant for the dye decolorization can enhance the decolorization percentage up to 77%. Conversely, the decolorization percentage without the use of surfactant was only 21%.
Enzymatic activities
Data shown in Fig. 2 presented the enzymatic activities during decolorization. Lignin peroxidase (LiP), laccase, and manganese peroxidase (MnP) were detected to be present in the liquid culture. In addition, 1,2-dioxygenase (1,2-DO) and 2,3-dioxygenase (2,3-DO) were also detected during decolorization in different activities. For instance, the presence of manganese peroxidase, lignin peroxidase, laccase, 1,2-dioxygenase, and 2,3-dioxygenase in the medium was 38, 37, 122, 10, and 6 U/L, respectively. Findings from this study exhibited that the highest enzyme activity was provided by laccase, indicating the main role of extracellular enzyme in the decolorization of MO-1 using the presently employed fungus. It is well established that the degradation of dyes can be mediated by several enzymes such as laccase, manganese peroxidase, lignin peroxidase, and some extents by dioxygenase, depending on type of fungi.
The capability of these enzymes to degrade various textile dyes has been well documented in literature [42,43,44,45]. Several studies have mentioned that the capability of these enzymes in decolorization of dyes depended highly on microorganism. For instance, laccase secreted by Pycnoporus sanguineus acted as an important enzyme to degrade azo and triphenylmethane dyes [44]. In addition, degradation of the anthraquinonic dye Remazol Brilliant Blue R can be facilitated using laccase produced by the white-rot fungus Trametes pubescens [43]. Moreover, laccases produced by Trametes hirsuta, Sclerotium rolfsii, Trametes villosa, Trametes modesta, and Trametes pubescens were found to have capability for decolorizarion of several synthetic dyes [42, 43, 45]. Following this logic, this work proposed that the white-rot fungus C. aurelium RY06 can degrade MO-1 via the enzymatic mechanism.
Surface morphology and FTIR characteristics
It was well established that synthetic dyes provided a harmful effect on living organisms such as bacteria and fungi. For this respect, Fig. 3 shows the morphological structure of C. aurelium RY06 before and after decolorization of MO-1. It was obvious from the figure that C. aurelium RY06 performed a perforated surface structure and rough surface in nature (see Fig. 3a). Several pores can also be observed on C. aurelium RY06 before decolorization of MO-1. A significant change in terms of their structure and roughness of C. aurelium RY06 can be observed after decolorization of MO-1. It can be seen from Fig. 3b that their surface morphology transformed to be smoother and unperforated surface structure. This was a result from the deposition of MO-1 on the surface of C. aurelium RY06. In addition, MO-1 can also fill the pores of C. aurelium RY06.
To investigate biomolecule transformations of MO-1 before and after decolorization, FTIR spectra analysis was employed as depicted in Fig. 4. It can be observed that the original MO-1 (see Fig. 4a) showed several prominent peaks around 900–1000 cm−1 that can be correlated to the C–O stretching. As a comparison, FTIR spectra at these regions for MO-1 after decolorization (see Fig. 4b) became more intense compared to before decolorization. In addition, a small peak at 1346 cm−1 that can be attributed to stretching vibration of C–N for MO-1 before decolorization transformed to be more intense after decolorization. In addition, a new sharp peak at 1707 cm−1 due to strong C=O can be observed for MO-1 after decolorization, which cannot be found in the control dye. This study also found that the peaks at 2973 and 3707 cm−1 attributed to C–H, and O–H stretching, respectively, exhibited more intensify compared to the control dye. By considering the different characteristics of the FTIR spectra of MO-1 before and after decolorization, it can be speculated that the dye can be degraded by C. aurelium RY06.
Proposed pathway for decolorization mechanisms
Table 3 presents mass spectra analysis of the metabolic product produced from MO-1 by C. aurelium RY06 complete with their correspondence UV–Vis spectra. In addition, Fig. 5 shows mass spectra of each metabolite of MO-1 by C. aurelium RY06. GC–MS chromatograms of MO-1 biodegradation exhibited the metabolite I with the molecular ion (M+-15) at m/z 245 and the substantial fragment ions at m/z 147 and 73. These characteristics can be associated with the sequential losses of OSi(CH3)3 and COOSi (CH3)3, respectively, which can be categorized as maleic acid (see Fig. 5a). In addition, the metabolite II showed major spectrum at m/z 310 and the significant fragment ions at m/z 295, 221, and 73 that can be correlated to the sequential losses of CH3 (CH3)3Si, and OSi(CH3)3, respectively. These characteristics can be linked to isopthalic acid as validated to the authentic compound (see Fig. 5b).
Following these evidences, C. aurelium RY06 can degrade MO-1 via two major routes, which are: (1) maleic acid and (2) isopthalic acid. For a comprehensive overview, a proposed pathway of MO-1 biodegradation by C. aurelium RY06 is shown in Fig. 6. Basically, dyes can be cleaved symmetrically or asymmetrically depending on the structure of substrate and involved enzymes [46]. For this study, MO-1 was asymmetrically cleaved by C. aurelium RY06 into 4-nitroaniline and 5-amino-2-hydroxybenzoic acid as observed in Fig. 6. In addition, it was noted from Fig. 6 that the transformation mechanism of MO-1 into the maleic acid was initiated by the asymmetric cleavage before deamination and hydroxylation, denitration, and carboxylation. For the metabolite II, the processes were deamination, carboxylation and dihydroxylation, and decarboxylation. Deamination, decarboxylation, and carboxylation and dihydroxylation are likely catalyzed by laccase [47]. Deamination and hydroxylation mechanism was possible with the help of laccase and dioxygenase enzymes. Moreover, the denitration mechanism can be associated with the presence of LiP enzyme [18].
It was well known that the transformation and biodegradation of pollutants depended highly on the types of microorganisms and pollutants. For instance, several metabolites, which are chrysenequinone, 1-hydroxy-2-naphthoic acid, phthalic acid, salicylic acid, protocatechuic acid, gentisic acid, and catechol can be detected when chrysene degraded using Polyporus sp. S133 [48]. Another study also observed that benzo[a]pyrene can be degraded by Armillaria sp. F022 to be benzo[a]pyrene-1,6-quinone, 1-hydroxy-2-benzoic acid, and benzoic acid [49]. The degradation of acid red 88 dye by Achaetomium strumarium can be identified by metabolites, namely sodium naphthalene-1-sulfonate and naphthalen-2-ol as well as finally degraded to be 1,4-dihydronaphthalene [16]. Direct blue-14 dye by Bacillus fermus can be degraded by metabolites, which are 3,5-diamino-4-hydroxynaphthalene-2,7-disulfonic acid, 3,3′-dimethylbiphenyl-4,4′-diamine, 4-hydroxynaphthalene-2,7-disulfonic acid, 3,3′-dimethylbiphenyl (2Z)-3-(4-hydroxyphenyl)prop-2-enoic acid, and (2Z)-3-(4-methylphenyl)prop-2-enoic acid [50].
Conclusions
This work was carried out to investigate the decolorization and biotransformation of MO-1 by C. aurelium RY06. The maximum decolorization percentage up to 86% can be obtained when glucose, ammonium nitrate, pH 3, agitation speed of 100 rpm, and 1 mL Tween 80 were employed. Although the maximum removal cannot achieve 100%, novelty and contribution of this study are on the proposed new mechanism and transformation, which are useful as basic knowledge to improve the method. The presence of manganese peroxidase, lignin peroxidase, laccase, 1,2-dioxygenase, and 2,3-dioxygenase can be detected with the highest concentration as laccase (122 U/L). The degradation of MO-1 using C. aurelium RY06 can be achieved by two different routes, which are to be maleic acid and to be isophtalic acid. This work was useful as a framework to employ C. aurelium RY06 for decolorization of dyes for foreseeable wastewater purification application.
References
Pearce C, Lloyd J, Guthrie J (2003) The removal of colour from textile wastewater using whole bacterial cells: a review. Dyes Pigm 58:179–196
Chen K-C, Wu J-Y, Liou D-J, Hwang S-CJ (2003) Decolorization of the textile dyes by newly isolated bacterial strains. J Biotechnol 101:57–68
Selvam K, Swaminathan K, Chae K-S (2003) Microbial decolorization of azo dyes and dye industry effluent by Fomes lividus. World J Microbiol Biotechnol 19:591–593
Rigas F, Dritsa V (2006) Decolourisation of a polymeric dye by selected fungal strains in liquid cultures. Enzyme Microb Technol 39:120–124
Zouari-Mechichi H, Mechichi T, Dhouib A, Sayadi S, Martinez AT, Martinez MJ (2006) Laccase purification and characterization from Trametes trogii isolated in Tunisia: decolorization of textile dyes by the purified enzyme. Enzyme Microb Technol 39:141–148
Gomes C, Piccin J, Gutterres M (2016) Optimizing adsorption parameters in tannery-dye-containing effluent treatment with leather shaving waste. Process Saf Environ Prot 99:98–106
Kadam AA, Lade HS, Lee DS, Govindwar SP (2015) Zinc chloride as a coagulant for textile dyes and treatment of generated dye sludge under the solid state fermentation: hybrid treatment strategy. Bioresour Technol 176:38–46
Nair V, Panigrahy A, Vinu R (2014) Development of novel chitosan–lignin composites for adsorption of dyes and metal ions from wastewater. Chem Eng J 254:491–502
Okesola BO, Smith DK (2013) Versatile supramolecular pH-tolerant hydrogels which demonstrate pH-dependent selective adsorption of dyes from aqueous solution. Chem Commun 49:11164–11166
Sarioglu M, Bali U, Bisgin T (2007) The removal of CI Basic Red 46 in a mixed methanogenic anaerobic culture. Dyes Pigm 74:223–229
Nigam P, Banat IM, Singh D, Marchant R (1996) Microbial process for the decolorization of textile effluent containing azo, diazo and reactive dyes. Process Biochem 31:435–442
Robinson T, McMullan G, Marchant R, Nigam P (2001) Remediation of dyes in textile effluent: a critical review on current treatment technologies with a proposed alternative. Bioresour Technol 77:247–255
Boochan M, Sudarat B, Grant A (2000) Degradation of high molecular weight polycyclic aromatic hydrocarbon by defined fungi-bacteria cocultures. Appl Environ Microbiol 66:1007–1019
Ekramul Karim M, Dhar K, Towhid Hossain M (2017) Co-metabolic decolorization of a textile reactive dye by Aspergillus fumigatus. Int J Environ Sci Technol (Tehran) 14:177–186
Munck C, Thierry E, Gräßle S, Chen SH, Ting ASY (2018) Biofilm formation of filamentous fungi Coriolopsis sp. on simple muslin cloth to enhance removal of triphenylmethane dyes. J Environ Manage 214:261–266
Bankole PO, Adekunle AA, Govindwar SP (2018) Enhanced decolorization and biodegradation of acid red 88 dye by newly isolated fungus, Achaetomium strumarium. J Env Chem Eng 6:1589–1600
Naraian R, Kumari S, Gautam RL (2018) Biodecolorization of brilliant green carpet industry dye using three distinct Pleurotus spp. Environ Sustainab 1:141–148
Bankole PO, Adekunle AA, Obidi OF, Chandanshive VV, Govindwar SP (2018) Biodegradation and detoxification of Scarlet RR dye by a newly isolated filamentous fungus, Peyronellaea prosopidis. Sustain Env Res 28:214–222
Ortiz-Monsalve S, Valente P, Poll E, Jaramillo-García V, Pegas Henriques JA, Gutterres M (2019) Biodecolourization and biodetoxification of dye-containing wastewaters from leather dyeing by the native fungal strain Trametes villosa SCS-10. Biochem Eng J 141:19–28
Pasti-Grigsby M, Paszczynski A, Goszczynski S, Crawford D, Crawford R (1992) Influence of aromatic substitution patterns on azo dye degradability by Streptomyces spp. and Phanerochaete chrysosporium. Appl Environ Microbiol 58:3605–3613
Ollikka P, Alhonmäki K, Leppänen V-M, Glumoff T, Raijola T, Suominen I (1993) Decolorization of azo, triphenyl methane, heterocyclic, and polymeric dyes by lignin peroxidase isoenzymes from Phanerochaete chrysosporium. Appl Environ Microbiol 59:4010–4016
Zhang F-M, Knapp JS, Tapley KN (1999) Decolourisation of cotton bleaching effluent with wood rotting fungus. Water Res 33:919–928
Vasdev K, Kuhad RC, Saxena RK (1995) Decolorization of triphenylmethane dyes by the bird’s nest fungus Cyathus bulleri. Curr Microbiol 30:269–272
Wong Y, Yu J (1999) Laccase-catalyzed decolorization of synthetic dyes. Water Res 33:3512–3520
Hsueh C-C, Chen B-Y, Yen C-Y (2009) Understanding effects of chemical structure on azo dye decolorization characteristics by Aeromonas hydrophila. J Hazard Mater 167:995–1001
Banat IM, Nigam P, Singh D, Marchant R (1996) Microbial decolorization of textile-dye containing effluents: a review. Bioresour Technol 58:217–227
O’Mahony T, Guibal E, Tobin JM (2002) Reactive dye biosorption by Rhizopus arrhizus biomass. Enzyme Microb Technol 31:456–463
Aksu Z, Tezer S (2000) Equilibrium and kinetic modelling of biosorption of Remazol Black B by Rhizopus arrhizus in a batch system: effect of temperature. Process Biochem 36:431–439
Tekere M, Mswaka AY, Zvauya R, Read JS (2001) Growth, dye degradation and ligninolytic activity studies on Zimbabwean white rot fungi. Enzyme Microb Technol 28:420–426
Moreira Neto SL, Matheus DR, Machado KMG (2009) Influence of pH on the growth, laccase activity and RBBR decolorization by tropical basidiomycetes. Braz Arch Biol Technol 52:1075–1082
Nor NM, Hadibarata T, Zubir MMFA, Lazim ZM, Adnan LA, Fulazzaky MA (2015) Mechanism of triphenylmethane Cresol Red degradation by Trichoderma harzianum M06. Bioprocess Biosyst Eng 38:2167–2175
Palmieri G, Cennamo G, Sannia G (2005) Remazol Brilliant Blue R decolourisation by the fungus Pleurotus ostreatus and its oxidative enzymatic system. Enzyme Microb Technol 36:17–24
Stajić M, Persky L, Friesem D, Hadar Y, Wasser SP, Nevo E, Vukojević J (2006) Effect of different carbon and nitrogen sources on laccase and peroxidases production by selected Pleurotus species. Enzyme Microb Technol 38:65–73
Kaal EEJ, Field JA, Joyce TW (1995) Increasing ligninolytic enzyme activities in several white-rot Basidiomycetes by nitrogen-sufficient media. Bioresour Technol 53:133–139
Hadibarata T, Syafiuddin A, Al-Dhabaan FA, Elshikh MS (2018) Biodegradation of Mordant orange-1 using newly isolated strain Trichoderma harzianum RY44 and its metabolite appraisal. Bioprocess Biosyst Eng 41:621–632
Arotupin D (2007) Effect of different carbon sources on the growth and polygalacturonase activity of Aspergillus flavus isolated from cropped soils. Res J Microbiol 2:362–368
Knapp J, Newby P, Reece L (1995) Decolorization of dyes by wood-rotting basidiomycete fungi. Enzyme Microb Technol 17:664–668
Moldes D, Lorenzo M, Sanromán MA (2004) Different proportions of laccase isoenzymes produced by submerged cultures of Trametes versicolor grown on lignocellulosic wastes. Biotechnol Lett 26:327–330
Hadibarata T, Yusoff ARM, Kristanti RA (2012) Decolorization and metabolism of anthraquionone-type dye by laccase of white-rot fungi Polyporus sp. s133. Water Air Soil Pollut 223:933–941
Lu Y, Yan L, Wang Y, Zhou S, Fu J, Zhang J (2009) Biodegradation of phenolic compounds from coking wastewater by immobilized white rot fungus Phanerochaete chrysosporium. J Hazard Mater 165:1091–1097
Dudu Gül Ü, Dönmez G (2012) Comparison the dye removal activity of systems contained surfactants and fungus. J Chilean Chem Soc 57:1170–1173
Couto SR, Osma JF, Saravia V, Gübitz G, Herrera JT (2007) Coating of immobilised laccase for stability enhancement: a novel approach. Appl Catal A 329:156–160
Osma JF, Toca-Herrera JL, Rodríguez-Couto S (2010) Transformation pathway of Remazol Brilliant Blue R by immobilised laccase. Bioresour Technol 101:8509–8514
Pointing SB, Vrijmoed L (2000) Decolorization of azo and triphenylmethane dyes by Pycnoporus sanguineus producing laccase as the sole phenoloxidase. World J Microbiol Biotechnol 16:317–318
Ryan S, Schnitzhofer W, Tzanov T, Cavaco-Paulo A, Gübitz G (2003) An acid-stable laccase from Sclerotium rolfsii with potential for wool dye decolourization. Enzyme Microb Technol 33:766–774
Dawkar VV, Jadhav UU, Jadhav SU, Govindwar SP (2008) Biodegradation of disperse textile dye Brown 3REL by newly isolated Bacillus sp. VUS. J Appl Microbiol 105:14–24
Patel VR, Bhatt NS, Bbhatt H (2013) Involvement of ligninolytic enzymes of Myceliophthora vellerea HQ871747 in decolorization and complete mineralization of Reactive Blue 220. Chem Eng J 233:98–108
Hadibarata T, Tachibana S, Itoh K (2009) Biodegradation of chrysene, an aromatic hydrocarbon by Polyporus sp. S133 in liquid medium. J Hazard Mater 164:911–917
Hadibarata T, Kristanti RA (2012) Fate and cometabolic degradation of benzo[a]pyrene by white-rot fungus Armillaria sp. F022. Bioresour Technol 107:314–318
Neetha JN, Sandesh K, Kumar KG, Chidananda B, Ujwal P (2019) Optimization of Direct Blue-14 dye degradation by Bacillus fermus (Kx898362) an alkaliphilic plant endophyte and assessment of degraded metabolite toxicity. J Hazard Mater 364:742–751
Acknowledgements
The authors thank the Universiti Teknologi Malaysia for financial supporting under the research university grant (Vote No. QJI.3000.2522.02H65). The Authors also extend their appreciation to the Deanship of Scientific Research at King Saud University for funding the work through the research group Project No RGP-1438-90.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mostafa, A.AF., Elshikh, M.S., Al-Askar, A.A. et al. Decolorization and biotransformation pathway of textile dye by Cylindrocephalum aurelium. Bioprocess Biosyst Eng 42, 1483–1494 (2019). https://doi.org/10.1007/s00449-019-02144-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-019-02144-3