Abstract
Sixty fungal cultures were isolated from agricultural soil, industrial soil, forest canopy soil having decomposed leaf litter and compost samples collected from different regions of India. Fifteen fungal cultures were selected qualitatively for the production of xylanase and cellulases and were identified employing ITS, NS and MNS primers. The enzyme cocktail consisting of 3811 IU g−1 of xylanase and 9.9 IU g−1 of cellulase from Trichoderma longibrachiatum MDU-6 was selected quantitatively for the deinking of diverse paper wastes. The enzyme production increased two fold when produced at tray level in comparison with flasks. The enzyme cocktail was effective in the deinking of old newspaper samples with significant removal of chromophores, phenolics and hydrophobic compounds and less sugar loss. While in case of examination papers and laser printed papers, ink removal was not very significant. Moreover, the sugar loss was significantly high in case of examination papers. The deinking results were further confirmed with FTIR analysis. Deinked newspaper pulp sample shows brightness of 52 %, which was 9.6 % high than its control sample. The ERIC value for deinked newspaper pulp was found to be 655.9 ppm. Thereafter, the deinked newspaper pulp was examined under light microscope after differential staining with safranin and malachite green and also examined under scanning and transmission electron microscope, which revealed fibrillation and perforation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The paper industries are mainly dependent upon three types of raw materials; forests, agro residues and waste papers. The biggest problem faced by this industry is the scarcity of conventional raw materials, i.e., forests. Therefore, the pre-eminent alternative is to increase the use of waste papers, which will reduce the dependence on conventional raw materials and also drop off the process cost and energy consumption (IARPMA estimates) [1]. In India, paperboard accounts for nearly 47.3 % of the total market size, followed by writing and printing paper (29.6 %), newsprint (19.5 %) and speciality paper (3.6 %) [CRISIL Research]. A typical writing and printing paper contains a mixture of carbohydrate polymers, i.e., cellulose and hemicelluloses, whereas the newspaper has additional modified lignin [2, 3]. Cellulose is the most abundant, unbranched biopolymer of glucose units that forms both amorphous and highly recalcitrant crystalline region through stable glycosidic bonds that are the target of cellulases (EC 3.2.1.4) [4]. Cellulases can be divided into three major groups of enzymes: endoglucanase (EC 3.2.1.4), exoglucanase (EC 3.2.1.91) also known as cellobiohydrolase and β-glucosidase (EC 3.2.1.21) that acts synergistically with different binding sites. Endoglucanase or CMCase acts randomly to break internal molecular bonds to increase the number of free reducing groups [5]. Exoglucanase or filter paper cellulase (FPase) degrades the polymeric chain from either the reducing or non-reducing ends to produce cellobiose, which is further hydrolysed by β-glucosidase into glucose monomers [6]. The cellulase activity and accessibility depends on the crystallinity of the celluloses and its association with hemicelluloses. Xylan is the dominating component of hemicelluloses and it is a heteropolysaccharide with a linear backbone chain of 1, 4-linked β-d-xylopyranose unit having various side-chains [7]. The structural heterogeneity of xylan necessitates the action of a number of hydrolytic enzymes, among which, endo-1, 4-β-xylanase (EC 3.2.1.8) plays an important role in xylan depolymerization [8].
Among various industrial enzymes; cellulases and xylanases are gaining enormous attention for their application in the recycling of various paper pulp samples and therefore are “green catalysts” [3, 9]. Cellulases and xylanases cause peeling off effect with the fiber and release the entrapped ink particles [1]. Kantelinen et al. [10] suggested that the hydrolysis of the xylan on the surface also liberate entrapped lignin and therefore increase the pulp brightness [11]. Due to the increasing demand of cost-competitive industrial enzymes, exploitation of local resources for the isolation of better fungal strains is the need of recent time [12]. Most commercial enzymes are of fungal origin, mainly because fungi secrete these enzymes extracellularly and their yield is also significantly high in comparison with yeasts and bacteria under submerged fermentation (SmF) and solid state fermentation (SSF) conditions [13, 14]. PCR amplification using various universal primers targeted to different housekeeping genes shows promising results in identifying a broad range of fungi up to the species level [15–18]. The sequence variation of internal transcribing spacer (ITS) regions has led to their use as a most significant tool in phylogenetic studies [16]. Elongation factor 1 (EF-1) primers target the tef-1α region and it is a well known primer for the identification of green spored filamentous fungi [17]. The primers NS1 through NS8 amplify nearly the entire nuclear small rDNA (18S) region in almost all fungi [18]. The objective of our current study was (a) to screen and identify different xylanase and cellulase producing filamentous fungi using various taxonomic markers, (b) production of xylanase and cellulase using wheat bran under SSF condition and (c) application of crude enzyme cocktail from T. longibrachiatum MDU-6 for the deinking of old examination papers, newspapers and laser printed paper wastes.
Materials and methods
Materials
3, 5-dinitrosalicylic acid (DNS), guaiacol and beechwood xylan were purchased from Sigma Chemicals (St. Louis, USA). All other dyes and chemicals were of reagent grade and wheat bran was bought from local vendors. Old examination papers, newspapers and laser printed papers were obtained locally.
Fungal isolation
The fungal cultures were isolated from agricultural soil, industrial soil, forest canopy soil having decomposed leaf litter and compost samples collected from different regions of India, especially from a latitude and longitude of 30.30 N and 74.60 E, respectively. The pH of the soil samples ranges from 6.0 to 8.0 with moisture availability of 10–15 %. The soil samples were collected with the help of disinfected spatula and stored in sterilized polythene bags. The fungal cultures were isolated by serial dilution method using 0.85 % normal saline on potato dextrose agar (PDA) medium. The purified fungal cultures were maintained on PDA slants at 4 °C and also stored as glycerol stocks at −80 °C.
Qualitative screening for lignocellulolytic enzymes
Xylanase screening was done on xylan-congo-red agar plates which contains 5.0 g L−1 peptone, 5.0 g L−1 yeast extract, 1.0 g L−1 KH2PO4, 0.1 g L−1 MgSO4·7H20, 2.5 g L−1 beechwood xylan, 0.5 g L−1 congo-red dye and 20 g L−1 agar. Cellulase screening was performed by replacing xylan with carboxymethyl cellulose (CMC) and without adding congo-red dye in the medium. 500 mM guaiacol prepared in citrate–phosphate buffer pH 5.4 along with PDA media was used for laccase screening.
Six wells (8 mm dia each) were prepared in each plate, one at the center and five at periphery. The culture supernatant (200 µL) produced under solid state fermentation (SSF) conditions from five filamentous fungi were loaded in the wells at periphery. Similar screening method was performed with the supernatant from all the purified fungal cultures. The supernatant from Ganoderma lucidum MDU-7 was used as control and loaded in the central well of each plate. After loading, both the xylan and CMC plates were incubated at 30 °C for 24 h. Xylan-congo-red plates were observed for zone of hydrolysis, while CMC agar plates were stained for 10–20 min with 0.5 % congo-red dye and destained with 1 M NaCl. The guaiacol plates were observed for the appearance of red coloration within few seconds after loading the enzyme sample [14].
Fungal identification
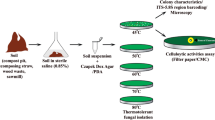
The selected fungal cultures were identified with the help of several housekeeping genes (ITS, EF-1, NS) used as taxonomic markers (Fig. 1). The genomic DNA of the fungal cultures was extracted from 72 h old growing mycelia by cell disruption method using liquid nitrogen [16]. Purification of genomic DNA was done using phenol, chloroform and isoamyl alcohol, precipitated with isopropanol and finally washed with 70 % ethanol. The primers used for PCR and DNA sequencing were specific for internal transcribed spacer (ITS) region [16], translation elongation factor 1α gene (EF-1) [17, 19] and nuclear small rRNA gene (NS) with their annealing temperatures 54, 59 and 58 °C, respectively [15, 18]. The genomic DNA of PCR negative fungal cultures were pre-denatured in the boiling water bath for 5 min with immediate cooling on ice, prior to amplification with NS primers. A pair of new degenerate primers (MNS) were also designed after multiple sequence alignment of the 18S regions of various filamentous fungi and used for fungal identification with 58 °C of annealing temperature.
All the PCR products were visualized on 1 % (w/v) agarose gel, stained with ethidium bromide. Thereafter, the amplicons were eluted from the agarose gel and sequenced commercially. Comparisons of sequences with those in the databases were made with BLASTN at NCBI (http://blast.ncbi.nlm.nih.gov). The nucleotide sequences were submitted to GenBank. The phylogenetic tree was constructed using molecular evolutionary genetics analysis (MEGA) software version 5.2 with bootstrap values calculated from 500 replicate runs.
Production of xylanases and cellulases
The qualitatively selected and identified fungal cultures were further screened quantitatively for the production of xylanase and cellulase under SSF conditions. Fresh fungal spores were used as inocula as described earlier [1]. 10 g wheat bran (washed, dried and milled to 0.5–1.0 mm particle size) was moistened with tap water in 1:3 ratio of solid substrate-to-liquid, the flasks were sterilized at 121 °C for 20 min and uniformly inoculated with fungal spores. The bran was incubated for 240 h at 30 °C under static conditions. The samples were extracted at a regular interval of 24 h with 0.05 M citrate buffer (pH 6.0) and vortexed (125 rev min−1, 1 h, and 30 °C). The extracted samples were centrifuged at 9168×g for 10 min at 4 °C. The cell free culture supernatant thus obtained was used for estimating xylanase and cellulase activity.
Enzyme assays
Xylanase and cellulase activity was determined by using 0.6 % beechwood xylan and 2 % and carboxymethyl cellulose, as substrate, respectively. The reaction mixture also contains 500 µl of appropriately diluted enzyme and incubated at 50 °C for 10 min. The reactions were then terminated using DNS reagent [20]. Absorbance of the resulting color was measured at 540 nm. One unit of enzyme activity (IU) was defined as the amount of enzyme required to release one µmole of reducing sugar per ml per min under the standard assay conditions.
Scale up for enzymes production
The selected fungal culture was cultivated in enamel coated metallic trays of size (24 × 19 × 5 cm3) containing 100 g of wheat bran. The trays were sterilized at 121 °C for 20 min and inoculated with fungal spore suspension. After incubation for 4 days, the fermented bran was extracted with 0.05 M citrate buffer (pH 6.0) and the supernatant was used for the deinking of different paper wastes.
Deinking of various paper pulp samples
Three types of paper samples, i.e., old examination papers, newspapers and laser printed papers were individually shredded and pulped by soaking in tap water at 5 % (w/v) consistency with 0.5 % (v/v) Tween 80, for 2 h under shaking condition (100 rpm) at room temperature. The soaked paper pieces were washed thoroughly with running tap water and macerated manually to obtain soft cottony pulp. Thereafter, the dried pulp samples were used for the deinking with enzyme cocktail extracted from T. longibrachiatum MDU-6. The deinking experiments were performed in 250 mL Erlenmeyer flasks at 5 % (w/v) consistency with 0.05 M citrate buffer (pH 6.0). All the three different pulp samples were treated with 7500 IU g−1 of xylanase and 20 IU g−1 of cellulase with 0.5 % (v/v) Tween 80 and incubated at 40 °C and 150 rpm for 3 h [1]. The control reaction contained all the components excluding enzymes. Further, the resulting filtrate was used to estimate total reducing sugars, chromophores, hydrophobic compounds and phenolics released during deinking process. The control and decolorized pulp samples were washed thoroughly with tap water and dried for further analysis.
Analytical methods
The relation between amount of reducing sugar (λ 540 nm) and ink released with the help of fungal enzyme from three different paper pulp samples was studied. The deinking of all the three pulp samples was measured by taking absorbance of released color at 550 nm (λ max). The amount of phenolics and hydrophobic compounds released were estimated at 237 and 465 nm, respectively [1].
Fourier transformed infrared spectroscopy
The surface functional groups of the deinked pulp samples and control samples (prepared without enzymes) were investigated for surface measurement by fourier transformed infrared (FTIR) spectrometry using the attenuated total reflectance (ATR) measuring cell. FTIR spectra were recorded with a resolution of 4 cm−1 over the wave number range of 3400–600 cm−1, using 24 scans per sample (Bruker Alpha ATR). The spectra were analyzed using the FTIR database [21].
Analysis of pulp properties
Deinked newspaper pulp sample was used to make hand sheets as described in TAPPI test T220 sp-01 procedure with a grammage of 60 g/m2. The hand sheets were characterized by measuring the brightness (IS: 1060-2) and effective residual ink concentration (ERIC) according to the standard procedures.
Pulp staining
Enzyme treated pulp and control samples were stained with appropriate dye or the combination of dyes, such as 1 % safranin, which stains lignin [22] and a large number of other dyes that stains cellulose fibers (1 % methylene blue, 1 % malachite green, 0.1 % congo-red, 0.05 % RBB and 0.1 % rhodamine B) [23]. The pulp sample was placed onto glass slide and stained with a few drops of a particular dye. Sample was then heated at 60 °C in an oven until dry and then rinsed carefully with distilled water to remove excess stain and examined under light microscope. Then, a combination of two dyes were prepared in the ratio of 1:1 (safranin with methylene blue and safranin with malachite green) and used for pulp staining. Finally, safranin with malachite green prepared in the ratio of 1:1 was selected for the differential staining of enzyme treated and control pulp sample. The differential staining procedure was used to examine the specific morphological changes due to the deinking effect of fungal enzymes.
Scanning electron microscopy (SEM) and transmission electron microscopy (TEM)
The deinked newspaper pulp sample was gradually dehydrated with acetone gradient (30, 50, 70, 80, 90 and 95 %) and finally suspended in 100 % acetone. The samples were kept for 15 min each; up to 95 % acetone, thereafter treated for 30 min with absolute alcohol. Then the sample was air dried, coated with a thin film of gold and examined under scanning electron microscope (JEOL, Model JSM-6100) at 10 kV in order to observe the surface morphology. Electron micro-graphs were taken at desired magnifications. Further, for TEM analysis the treated newspaper pulp samples were sonicated and the resulting fiber was casted directly on copper grids. Images were taken with Hitachi 7500 transmission electron microscope at 100 kV and different magnifications.
Results
Isolation and screening
Initially, sixty fungal cultures were isolated from agricultural soil, industrial soil, forest canopy soil having decomposed leaf litter and compost samples. All these cultures differ in their mycelial growth, spore color and extent of sporulation on PDA plates, when incubated at 30 °C (Fig. 1).
Various qualitative screening methods were compared to find out an easy, fast and evident qualitative method to screen microorganisms for the production of lignocellulolytic enzymes on agar plates (data not shown). The supernatant from xylanase positive culture showed distinct and prominent zone of hydrolysis around the wells in xylan congo-red plates. A sharp increase in the color contrast of the hydrolyzed and non-hydrolyzed zone was obtained after 24 h of incubation with enzyme (Fig. 1a). The supernatant loaded in CMC agar plates when stained with congo-red dye and destained with 1 M NaCl, then cellulase positive cultures showed zone of hydrolysis around the wells (Fig. 1a). Furthermore, the methods were also validated by loading the culture supernatant from basidiomycetes fungi in the central well of each plate, which gave negative results for xylanase and cellulase and marked as control. Whereas, all the filamentous fungi were found negative for laccases, only control gave red color with guaiacol within few seconds after enzyme loading (Fig. 1a) [14].
On the basis of qualitative screening, among sixty filamentous fungi, six cultures from agricultural soil, three from industrial soil, three cultures from forest canopy soil and three from compost samples were found positive for xylanase and cellulase and all the cultures gave negative results for laccases.
Fungal identification
These fifteen fungal cultures were identified using primers specific for various taxonomic markers (conserved regions) (Table 1). Initially, five cultures were identified by the amplification of their ITS region. EF-1 primer failed completely to identify any fungi as it produced multiple bands and also generated poor sequencing results (Fig. 2c II). Thereafter, six cultures were identified using NS primers, out of which the genomic DNA of three cultures was pre-denatured before amplification. Finally, the newly designed MNS primers were used successfully for the identification of four more fungi which failed earlier with NS primers.
a, b Locations on nuclear rDNAs and tef-1α DNAs of PCR primers. The arrowheads represent the 3′ end of each primer; c agarose gel electrophoresis of PCR products of various fungal genomic DNA amplified using; (I) ITS primers, (II) EF-1 primers, (III) NS primers and (IV) NS primer with pre-denatured DNA (V) MNS primers. M. Lambda DNA/HindIII Marker and L. 1 kb DNA ladder; d phylogenetic tree of various fungal strains identified using ITS primers and; e using NS primers. GenBank accession numbers are listed after each species name
PCR results revealed amplicon size of around 560 bp with ITS primers (Fig. 2c I), approximately 1800 bp in case of NS primers (Fig. 2c III, IV) and 1500 bp with MNS primers (Fig. 2c V). The sequences of all the fifteen fungal cultures were submitted to GenBank and their accession numbers were provided (Table 1). Two phylogenetic trees were prepared, one with cultures identified using ITS primers (Fig. 2d) and the other includes cultures identified after the amplification of their nuclear small rDNA region (Fig. 2e). Bootstrap values showed similarity and differences among the sequences of fungi belong to similar or different genera or species.
Production of xylanase and cellulase
On the basis of extensive qualitative screening and identification, the selected fifteen fungal cultures were screened quantitatively for the production of xylanase and cellulase under solid state fermentation (SSF) conditions (Table 2). Trichoderma longibrachiatum MDU-6, purified from agricultural soil with xylanase and cellulase activity of 3811 and 9.9 IU g−1, respectively was selected for the deinking of diverse paper wastes.
Scale up for enzymes production
Trichoderma longibrachiatum showed significantly high xylanase and cellulase activity; 7500 IU g−1 and 20 IU g−1, respectively at tray level. It was almost double the enzyme titer produced at flask scale. This enzyme cocktail consisting of xylanase and cellullases were used effectively for the deinking of old examination papers, newspapers and laser printed papers.
Deinking of various paper pulp samples
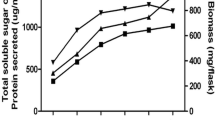
The enzyme cocktail extracted from T. longibrachiatum MDU-6 was successfully deinked the newspaper pulp samples apart from the fact that it contains modified lignin along with cellulose and hemicelluloses (Fig. 3a). Hydrolysis of xylan helps in the removal of lignin. In case of laser printed papers, non-impact ink (toners) was also removed to some extent with very less sugar loss (Fig. 3a). On the contrary, the removal of ball pen ink was very less but as there was no hindrance for fungal enzymes, it resulted in the liberation of higher amount of sugars (Fig. 3a). The deinking of all the paper samples was performed only for 3 h to prevent the additional sugar loss as optimized earlier [1]. These findings confirm the successful deinking of newspaper samples with the xylanase and cellulase extracted from T. longibrachiatum MDU-6.
a The extent of sugar, phenolics, hydrophobic compounds and chromophores released from examination papers, newspapers and laser printed papers treated with the extracellular enzymes of T. longibrachiatum MDU-6, b FTIR spectra of examination paper pulp, c newspaper pulp and d laser printed paper pulp showing the comparison between control and enzyme (extracted from T. longibrachiatum MDU-6) treated pulp samples
FTIR spectral analysis
The characteristic peak between 3600 and 3200 cm−1 is assigned to the concentration of hydroxyl groups. The area of this peak has similar intensity for both, enzyme treated and control sample of laser printed paper fibers, while the peak is slightly less intense in case of enzyme treated examination papers in comparison with its respective control. The difference between the intensities is high in case of newspapers fibers. The peak is less intense at 2897 cm−1, assigned to CH asymmetrical stretching vibration in CH3, CH2 and CH in enzyme treated newspaper and laser printed pulp samples, which indicate the degradation of aliphatic side chains [1].
The peak intensity was found to be low at 1730 and 1245 cm−1, which attribute to vibrational group C=O and C–O, respectively, present in the structure of lignin and hemicelluloses. The peak at 1634 cm−1 suggested the release of free carbonyl groups (C=O) from the aromatic ring. This peak is less intense in examination and laser printed papers and completely absent in newspaper fibers. The peak at 1434 cm−1 can be assigned to CH2OH group in ink and is less intense in case of enzyme treated pulp samples mainly with laser printed fibers. The peak at 1162 cm−1 in enzyme treated pulp indicated degradation of syringyl groups. The lower intensities of the peaks in all three enzyme treated samples at 1033 and 1060 cm−1 match the vibration of deformation of C–O–C bonds in the cellulose and hemicelluloses (Fig. 3b–d). These results confirm the maximum release of phenolics, hydrophobic compounds and chromophores from newspaper fibers, with the help of hydrolytic enzymes of T. longibrachiatum MDU-6. The sugar removal was maximum with examination papers due to the absence of inhibitory effect of lignin and pigments, which are present in newspapers and laser-printing papers.
Pulp properties
The hand sheets of control and deinked newspaper pulp samples were prepared with a grammage of 60 g/m2 for the analysis of brightness and ERIC. Deinked newspaper pulp sample shows brightness of 52 % which was 9.6 % high than its respective control sample. The ERIC value for deinked newspaper pulp was found to be 655.9 ppm. Enzyme treatment reduced the ERIC value in comparison with its control.
Pulp staining
Safranin shows affinity with lignin present in the newspaper pulp samples and therefore imparts red coloration to the pulp (Fig. 4a I). Methylene blue and malachite green were selected among various other dyes such as congo-red, RBB and rhodamine B, due to their higher affinity for celluloses (Fig. 4a II, III). Thereafter, pulp samples were compared between safranin:methylene blue and safranin:malachite green differential staining, which was prepared in the ratio of 1:1 (Fig. 4a IV, V). The results were prominent with the differential staining of pulp samples with safranin and malachite green (1:1), which revealed red color on the outer wall and green coloration on the inner side of the pulp fiber due to the binding of lignin and celluloses with safranin and malachite green, respectively (Fig. 4a V).
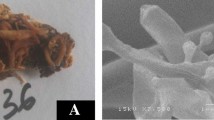
Staining of pulp samples with a (I) safranin, (II) methylene blue, (III), malachite green, (IV), safranin and methylene blue in the ratio of 1:1 and (V), safranin and malachite green in the ratio of 1:1, b differential staining of (I) control prepared without enzyme and (II) deinked newspaper pulp treated with enzyme from T. longibrachiatum MDU-6. Scanning electron micrographs (c-I, II) and transmission electron micrographs (d-I, II) of enzyme treated newspaper pulp at different magnifications, arrow indicate fibrillation and perforation on the fiber surface
Therefore, newspapers pulp samples treated with enzymes from T. longibrachiatum MDU-6 showed significant fibrillation and perforation on the fiber surface when compared with control sample (Fig. 4b). The control sample appears bright red in comparison with the enzyme treated newspaper samples due to the removal of lignin as shown in Fig. 4b.
Scanning electron microscopy and transmission electron microscopy
SEM and TEM data of enzyme treated newspaper pulp confirms fibrillation and irregular surface morphology, indicating delignification and slight hydrolysis on the fiber surface (Fig. 4c, d). Enzyme also excavates fiber surface and effectively develops cracks and perforations, therefore confirms the peeling off effect, which facilitates ink detachment from the surface due to the removal of small-fibrils after enzymatic treatment. The resolution of TEM provides advantages for the study of fiber ultra-structure.
Discussion
The cell free culture supernatant was used for the qualitative screening of lignocellulolytic enzymes because with fungal mycelial growth, it became very difficult to observe the clear zone of hydrolysis or the appearance of distinct coloration after the degradation of a particular substrate [24]. It is because the fungi grow with the radial spreading of their mycelia and there is a huge difference between the mycelial growth and sporulation among various filamentous fungi [25]. The enzymatic hydrolysis of xylan with congo-red media resulted in the appearance of sharply discernible clear zone due to reddish brown complex formation of dye with xylan (polysaccharide) but not with glucose (monosaccharide) [26]. The flooding of hydrolysed CMC media with congo-red dye followed by washing with 1 M NaCl provides an efficient and highly sensitive screening test for cellulolytic fungi [23, 26]. The zone of hydrolysis was not very clear with CMC-congo-red media as in case with xylanase because the cellulose is less soluble and does not mix properly with the congo-red dye. These outcomes contradict with the earlier reports that showed better results with gram’s iodine [25] and resembles with the reports of Mouelhi and co-worker [27]. In case of the screening of laccase enzyme, guaiacol produced significant results [14]. These qualitative screening results are easy to interpret and usually unambiguous, although a control loading of laccase was preferred in the central well with the supernatant from basidiomycetes fungi [26]. These methods worked well with both bacteria as well as fungi; hence, it can be employed for the screening of hydrolases and phenol oxidases produced from a large number of microorganisms.
Due to vast genomic diversity, a single marker study cannot fix the phylogeny of a diverse group of filamentous fungi. Therefore, various taxonomic markers including ITS primers [28], EF-1 primers [17] and NS primers [18] were used. The ITS primers make use of the conserved region of 18S, 5.8S and 28S rRNA genes to amplify the non-coding regions between them [18]. The sequence variations of ITS region has led to their use as a significant tool in phylogenetic studies and successfully identified few fungal cultures [16]. Thereafter, EF-1 primer was used that target the tef-1α region and it is well known primer for the identification of green spored filamentous fungi but it failed completely in identifying any fungal culture [19]. Finally, NS1 and NS8 primers were used that amplify the entire 18S gene in almost all fungi. NS primer gave good detection results but failed at certain points because the difference between the Tm and GC content of NS1 and NS8 primers were very high [28]. Therefore, the sequence information from various filamentous fungi and their phylogenetic relationships provides the basis for developing a modified NS (MNS) primer specific for the 18S rDNA region for the identification of other fungal cultures. Comparatively slow rate of molecular evolution makes the 18S rDNA sequence, a good taxonomic tool for finding consensus conserved regions suitable for fungal identification [29]. Those DNA regions can be selected for primer designing which shows similarity between 70 and 100 % among various fungal groups [29]. The PCR conditions were optimized for obtaining good yield of an amplified product of various sizes produced using all these primers [18].
Xylanase and cellulase production was found to vary among different species of the same genera or even among same species. T. longibrachiatum MDU-6 produced higher amount of xylanase and cellulase when compared with other fungal strains, under SSF conditions, using wheat bran as substrate. Wheat bran was used because it is cost-competitive and nutrient-rich substrate and contains proteins (14 %), carbohydrates (27 %; comprising 64 % cellulose and 36 % hemicellulose), lipids (6 %), minerals (5 %), vitamin B and high percentage of digestible nitrogen (64 %) as reported earlier [1]. Enzyme production was carried out under SSF condition because of higher biomass generation, lower catabolic repression and less proteolysis. It also provides natural environment for fungal growth and due to the absence of free flowing water bacteria are unable to grow and as a result there are no chances of bacterial contamination [14]. Therefore, SSF has gained significant interest for enzymes production during recent years using wheat bran as substrate. The crude enzyme preparation was used for deinking as it contains all the accessory enzymes for the complete hydrolysis of xylan and cellulose significantly in comparison with the purified enzyme preparation [30]. In the traditional deinking process, a large number of chemicals are used, which makes the process environmental damaging and also increase the process cost [31]. On the contrary, the use of enzymes makes the deinking process eco-friendly and significantly improves the optical properties of treated paper samples. Cellulase and xylanase, both are responsible for the mild hydrolysis on the fiber surface, in the fiber/ink inter-bonding regions, a process known as “peeling-off fibers” which facilitates ink detachment and removal of small fibrils from the surface [1, 32]. Additionally, these enzymes along with surfactants increase the relative hydrophobicity of ink particles, which facilitates their separation during washing [33]. Cellulases along with xylanases are well known enzymes for the deinking of various paper pulp samples [32, 34]. The hydrolytic enzyme preparation from A. oryzae MDU-4 [1], T. harzianum PPDDN10 NFCCI-2925 [32], Bacillus halodurans FNP 135 [31] and B. pumilus AJK [34] were found effective for the deinking of different types of papers. On the basis of deinking results, the hydrolases extracted from T. longibrachiatum MDU-6 were found highly effective for the deinking of newspaper samples. Examination papers released higher amount of sugars than newspapers and laser printed papers. The removal of phenolics, hydrophobic compounds and chromophores was maximum in case of newspapers [1, 31]. To study the morphological changes in the treated pulp samples, differential staining was performed. Safranin has affinity for lignin regardless of whether cellulose is present or not [22], while cellulose staining was compared with two different dyes; methylene blue and malachite green. Finally, safranin and malachite green were used in the ratio of 1:1 for the differential staining of treated pulp samples as reported by Yu et al. [35]. Our deinking results correlates with the earlier reports of Lee et al. [36], showing 50 % brightness of newspapers, which was deinked using commercial xylanase and cellulase extracted from Aspergillus niger. The ERIC value was in the range of 500 to 1200 ppm as reported by Vahey et al. [37]. FTIR results also corroborate with the earlier reports of enzymatic deinking [1]. Due to the presence of lignin in newspaper fibers, higher amount of phenolics, hydrophobic compounds and chromophores were released from them than laser printed papers with less sugar yield as compared to examination papers [1, 31, 34]. Enzyme treated newspaper pulp samples shows fibrillation and perforation on the fiber surface under scanning and transmission electron microscopy and therefore confirms the successful deinking of newspaper samples.
References
Chutani P, Sharma KK (2015) Biochemical evaluation of xylanases from various filamentous fungi and their application for the deinking of ozone treated newspaper pulp. Carbohydr Polym 127:54–63
Mohandass C, Raghukumar C (2005) Biological deinking of inkjet-printed paper using Vibrio alginolyticus and its enzymes. J Ind Microbiol Biotechnol 32:424–429
Kuhad RC, Mehta G, Gupta R, Sharma KK (2010) Fed batch enzymatic saccharification of newspaper cellulosics improves the sugar content in the hydrolysates and eventually the ethanol fermentation by Saccharomyces cerevisiae. Biomass Bioenergy 34:1189–1194
Berlin A (2013) No barriers to cellulose breakdown. Science 342:1454–1456
Torres CE, Negro C, Fuente E, Blanco A (2012) Enzymatic approaches in paper industry for pulp refining and biofilm control. Appl Microbiol Biotechnol 96:327–344
Lynd LR, Weimer PJ, Van Zyl WH, Pretorius IS (2002) Microbial cellulose utilization: fundamentals and biotechnology. Microbiol Mol Bio Rev 66:506–577
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Polizeli ML, Rizzatti AC, Monti R, Terenzi HF, Jorge JA, Amorim DS (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67:577–591
Torres CE, Negro C, Fuente E, Blanco A (2012) Enzymatic approaches in paper industry for pulp refining and biofilm control. Appl Microbiol Biotechnol 96:327–344
Kantelinen A, Hortling B, Sundquist J, Linko M, Viikari L (1993) Proposed mechanism of the enzymatic bleaching of kraft pulp with xylanases. Holzforschung 47:318–324
Georis J, Giannotta F, de Buyl E, Granier B, Frere JM (2000) Purification and properties of three endo-β-1,4-xylanases produced by Streptomyces sp. strain S38 which differ in their ability to enhance the bleaching of kraft pulp. Enz Microbial Technol 26:178–186
Qaisar S, Zohra RR, Aman A, Qader SAU (2014) Enhanced production of cellulose degrading CMCase by newly isolated strain of Aspergillus versicolor. Carbohydr Polym 104:199–203
Raj KC, Chandra TS (1996) Purification and characterization of xylanase from alkali-tolerant Aspergillus fischeri Fxn1. FEMS Microbiol Lett 145:457–461
Sharma KK, Kapoor M, Kuhad RC (2005) In vivo enzymatic digestion, in vitro xylanase digestion, metabolic analogues, surfactants and polyethylene glycol ameliorate laccase production from Ganoderma sp. kk-02. Lett Appl Microbiol 41:24–31
Bakri Y, Masson M, Thonart P (2010) Isolation and identification of two new fungal strains for xylanase production. Appl Biochem Biotechnol 162:1626–1634
Henry T, Iwen PC, Hinrichs SH (2000) Identification of Aspergillus species using internal transcribed spacer regions 1 and 2. J Clin Microbiol 38:1510–1515
Roger AJ, Sandblom O, Doolittle WF, Philippe H (1999) An evaluation of elongation factor 1α as a phylogenetic marker for eukaryotes. Mol Biol Evol 16:218–233
White TJ, Bruns T, Lee S, Taylor J (1990) PCR protocols: a guide to methods and applications. Academic, New York
Chaverri P, Castlebury LA, Overton BE, Samuels GJ (2003) Hypocrea/Trichoderma: species with conidiophore elongations and green conidia. Mycologia 95:1100–1140
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Buta JG, Zardrazil F, Galletti GC (1989) FT-IR determination of lignin degradation in wheat straw by white rot fungus Stropharia rugosoannulata with different oxygen concentrations. J Agric Food Chem 37:1382–1384
Srebotnik E, Messner K (1994) A simple method that uses differential staining and light microscopy to assess the selectivity of wood delignification by white rot fungi. Appl Environ Microbiol 60:1383–1386
Teather RM, Wood PJ (1982) Use of congo red-polysaccharide interactions in enumeration and characterization of cellulolytic bacteria from the bovine rumen. Appl Environ Microbiol 43:777–780
Hankin L, Anagnostakis SL (1977) Solid media containing carboxymethyl-cellulose to detect CX cellulase activity of microorganisms. J Gen Microbiol 98:109–115
Kasana RC, Salwan R, Dhar H, Dutt S, Gulati A (2008) A rapid and easy method for the detection of microbial cellulases on agar plates using Gram’s iodine. Curr Microbiol 57:503–507
Pointing SB (1999) Qualitative methods for the determination of lignocellulolytic enzyme production by tropical fungi. Fungal Divers 2:17–33
Mouelhi FM, Moisan JK, Beauregard M (2014) A comparison of plate assay methods for detecting extracellularcellulase and xylanase activity. Enz Microbial Technol 66:16–19
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque A, Chen W (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for fungi. PNAS 109:6241–6246
Wu Z, Tsumura Y, Blomquist G, Wang XR (2003) 18S rRNA gene variation among common airborne fungi, and development of specific oligonucleotide probes for the detection of fungal isolates. Appl Environ Microbiol 69:5389–5397
Saha BC (2002) Production, purification and properties of xylanase from a newly isolated Fusarium proliferatums. Process Biochem 37:1279–1284
Virk AP, Puri M, Gupta V, Capalash N, Sharma P (2013) Combined enzymatic and physical deinking methodology for efficient eco-friendly recycling of old newsprint. PLoS ONE 8:1–8
Pathak P, Bhardwaj NK, Singh AK (2014) Production of crude cellulase and xylanase from Trichoderma harzianum PPDDN10 NFCCI-2925 and its application in photocopier waste paper recycling. Appl Biochem Biotechnol 172:3776–3797
Jeffries TW, Klungness JH, Sykes MS, Rutledge-Cropsey KR (1994) Comparison of enzyme-enhanced with conventional deinking of xerographic and laser-printed paper. Tappi J 77(4):173–179
Singh A, Yadav RD, Kaur A, Mahajan R (2012) An ecofriendly cost effective enzymatic methodology for deinking of school waste paper. Bioresour Technol 120:322–327
Yu X, Minor JL, Atalla RH (1995) Mechanism of action of Simons’ stain. Tappi J 78:175–180
Lee CK, Ibrahim D, Omar IC (2013) Enzymatic deinking of various types of waste paper: efficiency and characteristics. Process Biochem 48:299–305
Vahey DW, Zhu JY, Houtman CJ (2006) On measurements of effective residual ink concentration (ERIC) of deinked papers using Kubelka-Munk theory. Prog Pap Recycl 16:3–12
Acknowledgments
The authors are grateful to the Council of Scientific and Industrial Research (CSIR), Government of India, New Delhi [Grant No. 38(1313)/11/EMR-II] for the financial assistance to carry out this work. We are also thankful to Dr. Rajvinder Singh and Ekta Saini, Department of Forensic Sciences, M.D.U., Rohtak, for providing the facility for FTIR studies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Rights and permissions
About this article
Cite this article
Chutani, P., Sharma, K.K. Concomitant production of xylanases and cellulases from Trichoderma longibrachiatum MDU-6 selected for the deinking of paper waste. Bioprocess Biosyst Eng 39, 747–758 (2016). https://doi.org/10.1007/s00449-016-1555-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-016-1555-3