Abstract
Cells respond to extra- and intra-cellular signals by dynamically changing their gene expression patterns. After termination of the original signal, new expression patterns are maintained by epigenetic DNA and histone modifications. This represents a powerful mechanism that enables long-term phenotypic adaptation to transient signals. Adaptation of epigenetic landscapes is important for mediating cellular differentiation during development and allows adjustment to altered environmental conditions throughout life. Work over the last decade has begun to elucidate the way that extra- and intra-cellular signals lead to changes in gene expression patterns by directly modulating the function of chromatin-associated proteins. Here, we review key signaling-to-chromatin pathways that are specifically thought to target Polycomb and Trithorax group complexes, a classic example of epigenetically acting gene silencers and activators important in development, stem cell differentiation and cancer. We discuss the influence that signals triggered by kinase cascades, metabolic fluctuations and cell-cycle dynamics have on the function of these protein complexes. Further investigation into these pathways will be important for understanding the mechanisms that maintain epigenetic stability and those that promote epigenetic plasticity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cells are constantly exposed to changing environmental signals and respond by the initiation of cellular signaling cascades. These cascades often terminate in the nucleus and induce or repress gene activity. Changes to gene activity can be transient but are also often transformed into more permanent changes by epigenetic mechanisms that maintain gene expression signatures in the absence of the original signal.
Epigenetic changes are mediated by post-translation modifications of either DNA or histone residues. The most studied DNA modifications are 5-methyl cytosine and its derivatives, 5-hydroxymethylcytosine, 5-formylcytosine and 5-carboxylcytosine (e.g., Kohli and Zhang 2013). The extent to which DNA methylation occurs is species-specific, although 5-methyl cytosine modifications have been detected in many bacteria, plant, insect and mammalian cells. Some of the possible histone modifications include acetylation, methylation, phosphorylation, sumoylation, ubiquitylation, ribosylation and glycosylation of histones H1, H2A, H2B, H3, H4 and their respective variants. These modifications can alter the ability of DNA to associate tightly with the histone octamer, thereby altering the accessibility of DNA to regulatory factors, or can serve as a molecular platform for the recruitment of other protein complexes that further modify the chromatin state.
Central to the historic notion of epigenetics, as introduced by Conrad Waddington, is the concept that gene expression patterns are stable. According to this idea, epigenetic modifications are stably inherited through cell divisions and serve as a cellular memory of past decisions. Within this framework, DNA and histone modifications maintain activated or silenced states of target genes and are crucial to the maintenance of cell differentiation and lineage restrictions throughout embryonic and adult life.
Work over the past decade, however, has highlighted the plasticity associated with chromatin modifications and epigenetically controlled switches at target genes. Cellular signal transduction in response to environmental changes, metabolic processes, cell-cycle dynamics, or disease conditions appear to alter cellular phenotypes by regulating DNA and histone modifications. The associated epigenetic changes represent a powerful mechanism for translating transient environmental signals into enduring modifications to cellular phenotype and function.
Many of these signal-mediated changes arise through the activation of classic DNA-sequence-specific transcription factors that act as signaling effectors and integrators. They can recruit, cooperate with, or compete with epigenetically acting protein complexes for the initiation of changes in the activation states of target genes (see Beisel and Paro 2011). Various studies, however, point to more direct effects that signaling transduction cascades, energy metabolism and the cell cycle can have on the function of epigenetically acting proteins and protein complexes. Post-translational modifications of chromatin-associated proteins by signaling cascades and the dependency of chromatin-modifying enzymes on metabolically derived co-factors are among the most exciting examples of recent studies that directly link cellular processes to the epigenetic state. Although the way by which these mechanisms communicate the cellular state to chromatin is currently poorly understood, these signaling-to-chromatin pathways are likely to contribute to epigenetic processes during cell differentiation, nuclear reprogramming, tissue regeneration and disease. Our current understanding, or lack thereof, about the extent to which signaling-induced changes to chromatin are inherited by daughter cells makes the use of the term “epigenetic” in its original sense sometimes problematic. However, to reflect the fact that signaling-induced chromatin modifications are potentially heritable, we will often use the term “epigenetic” interchangeably with “chromatin-modifying process”.
In this review, we will highlight several recent studies that (1) investigate functional changes in response to post-translational modifications of epigenetically acting proteins, (2) associate metabolic processes with altered enzymatic activity of histone-modifying enzymes and (3) point to cell-cycle dynamics as an important modulator of epigenetic landscapes. We will focus on studies that describe changes to gene-regulatory mechanisms rather than those affecting higher-order genome structure. We will pay particular attention to signal-induced functional changes in members of the Drosophila Polycomb group (PcG) and Trithorax group (TrxG) family, a classic example of epigenetically acting silencer and activator complexes that target genes important for development, stem cell differentiation and cancer. We will specifically highlight observations suggesting that cellular stress initiates signaling pathways directly regulating PcG and TrxG function, as they have important implications for our understanding of epigenetic adaptation to processes such as wounding healing, disease states, or aging. Principles derived from the studies discussed below probably apply to other chromtin-modifying pathways. Additionally, we will refer the reader not only to the relevant original studies but also to some excellent and extensive reviews published on these subjects.
Function of PcG and TrxG proteins
PcG and TrxG proteins are highly conserved throughout evolution (Sawarkar and Paro 2010) and function as antagonistically acting protein complexes that regulate the expression of diverse sets of genes (see Schuettengruber et al. 2009; Schwartz et al. 2010; Tolhuis et al. 2006). The first target genes to be identified were those encoding homeotic transcription factors that belong to the Hox gene family and that are necessary for determining cellular identity along the anterior-posterior body axis of invertebrates and vertebrates (Zink and Paro 1989). More recent work in Drosophila has highlighted the role of PcG and TrxG in regulating target genes involved in cell-cycle control (Iovino et al. 2013; Jacobs et al. 1999; Oktaba et al. 2008) and tissue growth (Classen et al. 2009; Martinez et al. 2009). Similarly, in mammalian cells, PcG/TrxG proteins function as important tumor suppressors and oncogenes by regulating genes controlling cell proliferation (reviewed in Mills 2010).
PcG and TrxG proteins associate to form several multi-protein complexes that primarily implement (“write”) and recognize (“read”) histone methylation, ubiquitylation and acetylation marks (Fig. 1). At least five PcG complexes both “write” and “read” inactivating histone marks: Polycomb repressive complex 1 (PRC1; Franke et al. 1992), Polycomb repressive complex 2 (PRC2; Ng et al. 2000), dRing-associated factor complex (dRAF; Lagarou et al. 2008), Polycomb repressive deubiquitinase complex (PR-DUB; Scheuermann et al. 2010) and Pho-repressive complex (Pho-RC; Klymenko et al. 2006; for reviews, see Lanzuolo and Orlando 2012; Simon and Kingston 2013). Most prominently, the conserved SET-domain protein Enhancer of Zeste (E(z)) of PRC2 catalyzes or “writes”the trimethylation of the histone 3 lysine 27 residue (H3K27me3). The chromo-domain protein Polycomb (Pc) of PRC1 “reads” and binds H3K27me3, which is thought to implement gene silencing by compacting chromatin (Francis et al. 2004), by interfering with transcription initiation (Breiling et al. 2001) and by the recruitment of additional histone-modifying complexes (Lagarou et al. 2008; Wang et al. 2004; for a review, see Sparmann and van Lohuizen 2006). In mammalian cells, the E(z) homolog EZH2 recruits DNA methyltransferases, integrating a multiple gene-silencing mechanism at target genes (Vire et al. 2006).
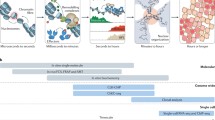
Drosophila Polycomb and Trithorax group complexes: molecular composition and function. In Drosophila, Polycomb repressive complex 1 (PRC1), Polycomb repressive complex 2 (PRC2) and the dRing-associated factor complex (dRAF) maintain silenced states by H3K27 methylation, H2AK118 ubiquitinylation and H3K36 demethylation (ME methyl group). Erasers, such as Lsd1, Lid and Rpd3, facilitate conversion from active to silenced states by H3K4 demethylation and H3K27 deacetylation (AC acetyl group). PcG-mediated silenced states are counteracted by the H3K4 methylation activities of the Trithorax-complex belonging to the COMPASS (Complex of Proteins Associated with Set 1) family. The dSet1-complex acts as general H3K4 methyltransferase important for gene activation. Utx of the Trr-complex catalyzes H3K27 demethylation, whereas association between CBP and Ash1 catalyzes H3K27 acetylation and H3K36 methylation. All three activities molecularly counteract the enzymatic activities of the PcG-associated PRC2 and dRAF complexes on H3K27 and H3K36, thereby promoting conversion from silenced to active states. Phosphorylation of H3S28 residues interferes with Polycomb function at H3K27 residues. A gradient of transcriptional activation is symbolized by increasingly open chromatin configurations from left to right, as facilitated by ATP-dependent SWI-SNF remodelers (black arrows targeting of histone residues by epigenetically acting proteins, red arrows removal of specific modifications from histone tails)
Conversely, Drosophila TrxG and mammalian myeloid/lymphoid or mixed-lineage leukemia (MLL)-protein-containing complexes work antagonistically to PcGs as activators of the same target genes. They do so by either catalyzing the implementation of activating epigenetic marks, such as the trimethylation of histone 3 lysine 4 (H3K4me3) by the SET-domain-containing Trx protein, the dimethylation of histone 3 lysine 36 (H3K36me2) by the SET-domain-containing Ash1 protein (Dorighi and Tamkun 2013), or by working as ATP-dependent chromosome remodelers (for reviews, see Ruthenburg et al. 2007; Schuettengruber et al. 2011).
H3K4me3 has been recognized as a general mark of actively transcribed genes. Three major H3K4 methylation complexes have been purified from Drosophila (for a review, see Shilatifard 2012). One of the Drosophila complexes contains the SET protein Trx protein at its catalytic core, whereas the SET-domain proteins Trithorax-related (Trr) and dSet1 are at the center of the other two. The complex containing Trx is thought to mediate H3K4 trimethylation at PcG target genes, whereas the Trr complex catalyzes H3K4 monomethylation at enhancer regions (Herz et al. 2012) and the dSet1 complex acts as a general H3K4 trimethyltransferase at actively transcribed genes (Ardehali et al. 2011; Hallson et al. 2012).
Several PcG and TrxG proteins function as “eraser” proteins that prime histones for the implementation of new marks. Many of these “eraser” proteins are also found to be in complex with histone code “writers”, facilitating immediate transition from active to inactive chromatin states or conversely from inactive to active chromatin states (see Lagarou et al. 2008; Rudolph et al. 2007).
PcG and TrxG erasers work primarily as demetylases, deubiquitinases, or deacetylases. For example, demethylation of the TrxG-mediated H3K4me3/H3K4me2 mark at active promoters is accomplished by the Drosophila KDM5a homolog little imaginal discs (lid; Eissenberg et al. 2007; Lee et al. 2007; Secombe et al. 2007), whereas dLsd1, the homolog of human KDM1a, catalyzes the removal of H3K4me2/H3K4me1 histone marks (Rudolph et al. 2007). Furthermore, the Drosophila demethylase dKdm2 associates with the PcG-dRAF complex to promote demethylation of Ash1-mediated H3K36me2 marks, allowing the dRAF complex immediately to implement the inactivation of H2A ubiquitylation. H3K36me2 demethylation in over-actively transcribed gene regions is necessary for efficient H2A ubiquitylation and for the silencing of PcG-target gene transcription (Lagarou et al. 2008). The Drosophila Trithorax-group protein Utx (mouse KDM6a homolog) is a histone demethylase that catalyzes the removal of the PcG-mediated H3K27me3 repressive mark and is found to co-localize with elongating RNA polymerase II (Smith et al. 2008). Removal of H3K27me3 may allow the Trx-associated protein dCBP/p300 to acetylate H3K27, preventing the reversion of these sites to H3K27me3 states, thus antagonizing gene silencing (Pasini et al. 2010; Schwartz et al. 2010; Tie et al. 2009, 2012). Rpd3, or dHDAC1, is a histone deacetylase capable of removing H3K27ac marks facilitating reversal to a silenced state (Reynolds et al. 2012).
Another epigenetic modification, called O-linked N-acetylglucosamine (O-GlcNAc), has recently been associated with PcG and TrxG function. O-GlcNAc addition to proteins is catalyzed by O-linked N-acetylglucosamine transferase (OGT) and is erased by O-GlcNAc glycoside hydrolase (OGA). Strikingly, the Drosophila OGT gene, called super sex combs (sxc), was first identified as a homeotic gene that genetically interacts with members of the PcG family (Cheng et al. 1994; Ingham 1984). These observations suggest that OGlcNAcylation and, possibly, the dynamic reversal mediated by OGT/OGA contribute to PcG-mediated gene silencing.
The molecular mechanisms described above highlight the potential that PcG/TrxG complexes possess for dynamically converting chromatin between active and inactive states. Although we are starting to understand the molecular mechanisms underlying such dynamic conversion processes, the reason and the way that these processes are initiated by cellular signals are much less clear. Increasing studies indicate that cellular signaling cascades can directly act on PcG/TrxG complexes to modify their function (Fig. 2).
PcG/TrxG activity might depend on various metabolic signals. Left Cellular production of the methyl donor S-adenosylmethionine (SAM) for DNA and histone methyltransferase reactions depend on co-factor availability for the 1-carbon-metabolism pathway. The byproduct of SAM methyl-group donation is S-adenosyl homocysteine (SAH), a potent inhibitor of methyltransferases. Middle Cellular production of flavin adenine dinucleotide (FAD) and α-ketoglutarate, co-factors for histone (and DNA) demethylases, is dependent on the tricarboxylic acid (TCA) cycle and electron-transport-chain kinetics. Furthermore, TCA cycle intermediates can inhibit the production of α-ketoglutarate. Since α-ketoglutarate-dependent demethylases also require oxygen to function, oxygen levels and hypoxia might affect their catalytic activity. Right The hexosamine biosynthesis pathway utilizes glucose to generate the O-GlcNAc donor uridine diphospho-N-acetylglucosamine (UDP-O-GlcNAc). O-GlcNAcylation of PcG and histone proteins is mediated by the enzyme O-GlcNAc transferase (OGT). O-GlcNAcase (OGA) catalyzes the removal of O-GlcNAc from proteins. OGT and OGA catalytic cycles make O-GlcNAc modifications dynamic and reversible
Regulation of PcG and TrxG function by kinase signaling cascades
Modulation of epigenetic mechanisms by signaling through kinase networks has recently received much attention. Several studies suggest that kinases can function as direct chromatin-interacting proteins (for excellent reviews, see Chow and Davis 2006; Klein et al. 2013; Vermeulen et al. 2009; Yang et al. 2013). For example, genome-wide mapping of extracellular-signal-regulated kinase 2 (ERK2) binding in human embryonic stem cells (hESC) reveals that ERK2 binds near genes important for proliferation and pluripotency. Curiously, a negative correlation between ERK2-occupied genes and PcG-binding has been observed, suggesting that extracellular-signaling-stimulated cell proliferation and PcG-mediated repression of cellular differentiation work together to maintain the identity of hESCs (Goke et al. 2013). Several recent studies expand on this notion of chromatin-associated kinases and suggest that changes in PcG and TrxG function arise either through the direct phosphorylation of PcG/TrxG proteins or through the phosphorylation of histone tails in response to various cellular signals. The way that these processes contribute to epigenetic plasticity is an emerging area of study, adding yet another level of complexity to an already intricate chromatin network.
Direct phosphorylation of PcG and TrxG proteins
Phosphorylation is utilized by signaling cascades to transmit cellular messages in response to a number of diverse stimuli. Kinases mediate the covalent addition of phosphate groups to specific serine, threonine and tyrosine residues, whereas phosphatases counteract the activity of kinases by specifically removing phosphate groups from proteins. This reversible cycle makes phosphorylation a versatile, yet also a dynamic, post-translational modification.
Several studies suggest that direct phosphorylation of PcG and TrxG proteins regulates protein function in a context-specific manner (for reviews, see Muller and Verrijzer 2009; Niessen et al. 2009) and mediates acute and long-term epigenetic modifications that are intimately coupled to cellular states. One of the first indications that phosphorylation regulates PcG/TrxG protein functions in Drosophila was the observation that PRC2 complex formation can be inhibited by the phosphorylation of Extra sex combs (Esc), a binding partner or E(z) (Ng et al. 2000; Tie et al. 2005).
More recently, cell-cycle-regulated phosphorylation of PRC2 complex member EZH2 by Cyclin-dependent kinase 1 (CDK1) and Cyclin-dependent kinase 2 (CDK2) in mammals has been shown to cause global reduction in H3K27me3 levels, increased EZH2 degradation and interference with EZH2-binding to the non-coding RNAs HOTAIR and Xist (Chen et al. 2010; Kaneko et al. 2010; Wei et al. 2011; Wu and Zhang 2011). EZH2 can also be phosphorylated by Akt, a downstream kinase in the cell growth and nutrient-responsive phosphoinositide 3-kinase (PI3K) signaling pathway, resulting in reduced histone methyltransferase activity of EZH2 and decreased cellular H3K27me3 levels (Cha et al. 2005). CDK1/2 and Akt-mediated phosphorylation of EZH2 results in increased expression of normally silenced PcG target genes. These studies link dynamic physiological processes, such as cell cycle and nutrient sensing, to dynamic EZH2 function (for a review, see Caretti et al. 2011).
Similar to EZH2, the mammalian Posterior sex combs (Psc) homolog Bmi-1 was found to be phosphorylated by Akt1. However, in contrast to EZH2, phosphorylation of Bmi-1 enhances ubiquitylation activity of a Bmi-1/Ring1B complex toward H2A, resulting in tumor suppressor gene silencing (Nacerddine et al. 2012). Phosphorylation of another mammalian Psc homolog, Mel-18, similarly increases the ubiquitylation activity of a Ring1B/Mel-18 complex toward H2A-K119, which is important for PcG target gene repression (Cao et al. 2005; Elderkin et al. 2007; Endoh et al. 2012).
In contrast to Akt-mediated signaling, mitogen-activated protein kinase (MAPK) signaling has been shown to target the phosphorylation of CBX7, a homolog of the Drosophila Polycomb protein and to increase CBX7-repressive activity at target genes (Wu et al. 2013). Although some of the upstream kinases are less-well characterized, the phosphorylation of the chromo-domain of mouse CBX2, another CBX family member also known as M33, alters its H3K27me3-binding specificity. Phosphorylated CBX2 exhibits increased binding preference toward H3K27me3- over H3K9me3-modified histone tails (Hatano et al. 2010). CBX2 has previously been demonstrated also to be phosphorylated in liver cells undergoing regenerative growth after partial hepatectomy (Noguchi et al. 2002), implying that phosphorylation in the context of tissue stress or damage modulates CBX2 function.
Tissue damage induces the activation of a number of stress-responsive signaling cascades, including the three main branches of the MAPK pathways: ERK, p38 and c-Jun NH2-terminal kinase (JNK). In flies, activation of JNK is critically required for stress responses, wound healing and tissue regeneration (Bosch et al. 2005; Ramet et al. 2002) and has been revealed to downregulate PcG function (Klebes et al. 2005; Lee et al. 2005; Owusu-Ansah and Banerjee 2009). Downregulation of PcG function has also been observed in a mouse wounding model (Shaw and Martin 2009). Transcriptional downregulation of PcG members appears to explain part of the observed functional changes in flies and mouse (Lee et al. 2005; Shaw and Martin 2009). One is tempted to speculate, however, that the direct phosphorylation of epigenetically acting proteins downstream of JNK contributes to the ability of cells to mount an efficient stress response. The phosphorylation both of PcG/TrxG protein complexes and of histones might contribute to dynamic epigenetic switches at genes necessary for tissue repair and regenerative proliferation in response to damage or cellular stress.
Incidentally, cellular stress induced by reactive oxygen species (ROS) treatment or serum starvation leads to the MAPK-mediated phosphorylation of Bmi-1 and results in the expulsion of Bmi-1 from chromatin (Nakamura et al. 2012; Prickaerts et al. 2012; Voncken et al. 2005). Another study suggests that the interplay between Akt-dependent and p38 stress-dependent phosphorylation regulates the stability of Bmi-1 (Kim et al. 2011). RNF2, the mammalian homolog of the Drosophila PRC1 protein dRING, is phosphorylated via signaling through the ERK and p38 pathways. This interferes with its function, as unphosphorylated RNF2 more efficiently silences the tumor suppressor gene p15INK4b (Rao et al. 2009). Furthermore, interaction between MK2 (mitogen-activated protein kinase-activated protein kinase 2) and the human Polyhomeotic homolog, hPH2, has been found to be essential for hematopoietic stem cell maintenance (Schwermann et al. 2009) suggesting that the MK2-mediated phosphorylation of the PRC1 component hPH2 is required for PcG-silencing of differentiation genes.
MAPK-mediated phosphorylation also plays a role in targeting TrxG complexes to chromatin during differentiation. The H3K4me3-methyltransferase-containing Ash2L-complex and the chromatin-remodeling the SWI-SNF complex are recruited to muscle-specific gene loci by the activation of the p38 MAPK pathway during myoblast differentiation (Rampalli et al. 2007; Simone et al. 2004). Phosphorylation of the SWI-SNF complex member BAF60c by p38α facilitates the formation of a larger complex between muscle determination factor MyoD and the SWI-SNF complex, causing chromatin remodeling and the activation of a number of MyoD target genes (Forcales et al. 2012). In contrast, p38α also facilitates the phosphorylation of EZH2 in satellite stem cells during muscle regeneration, thereby preventing proliferation and the expression of the Pax7 target gene (Palacios et al. 2010).
The biological significance of individual phosphorylation events in response to cellular signaling is highly context-specific and causes activation, inactivation, or changes in protein function. However, the direct phosphorylation of chromatin-bound proteins is not the only mechanism that has evolved to regulate chromatin structure and gene expression in response to extra- and intra-cellular signals.
Phosphorylation of H3S10 and H3S28 histone residues
In a landmark paper, the genome-wide profiles of JNK revealed that this kinase directly binds to a large set of active promoters and is required for the differentiation of mouse neuronal stem cells; JNK binding at promoters negatively correlates with the presence of H3K27me3 and positively correlates with the activation of target gene expression (Tiwari et al. 2012). The functional significance and potential mechanistic link of this binding are not clear and remain to be elucidated. The authors identified serine 10 on histone 3 as the main phosphorylation substrate of JNK, which is enriched at active JNK-bound promoters during neuronal stem cell differentiation.
Many other kinases have been described to phosphorylate H3S10. In mammals, Aurora B, AKT, MSK1/2, PIM1, IKK, RSK2 and, as described above, JNK have all been shown to phosphorylate H3S10 in response to a number of diverse cellular cues (Banerjee and Chakravarti 2011; Dyson et al. 2005; Sassone-Corsi et al. 1999; Soloaga et al. 2003; Tiwari et al. 2012; Zippo et al. 2007). During mitosis, this mark coincides with highly condensed, transcriptionally inactive chromosomes; however, H3S10 phosphorylation in response to differentiation signals results in the activation of a number of specific target genes (Sassone-Corsi et al. 1999; Tiwari et al. 2012; Vicent et al. 2006; Zippo et al. 2007). Some of these effects might be mediated by interference with binding of Heterochromatin-Protein 1 (HP-1) to H3K9me3 marks, thereby disrupting the establishment of a repressive heterochromatin-like state. Accordingly, the loss of H3S10 kinase Jil-1 in Drosophila results in increased H3K9 methylation along chromosome arms and increased spreading of heterochromatin (Zhang et al. 2006).
Similarly, the phosphorylation of serine 28 on histone 3 (H3S28P) next to the Polycomb-targeted H3K27 residue is thought to interfere with Polycomb-binding and the silencing function at target genes. H3S28P is facilitated by the activation of ERK and mitogen- and stress-activated kinases (MSK) pathways in response to hormonal stimuli and to UVB-induced DNA damage (Vicent et al. 2006; Zhong et al. 2001). Additionally, stress and mitogenic and retinoic acid signaling activates MSK1 and MSK2 and promotes H3S28 phosphorylation (Dyson et al. 2005; Gehani et al. 2010; Soloaga et al. 2003). MK3 kinase activation causes H3S28 phosphorylation and subsequent dissociation of PRC1 from target genes (Prickaerts et al. 2012). In all studies, signaling-induced H3S28P is thought to allow the re-activation of PcG-silenced target genes (Gehani et al. 2010; Prickaerts et al. 2012).
MSK phosphorylation of H3S28 is additionally thought to promote a switch from H3K27me3 to H3K27ac modifications, further antagonizing PcG gene silencing (Lau and Cheung 2011). In contrast, both H3S10P and H3S28P recruit the TrxG-associated ATP-dependent chromatin remodeler SWI-SNF to immediate early genes in response to Ras-MAPK signaling, which facilitates nucleosomal remodeling of target genes and transcriptional activation (Drobic et al. 2010). H3S28P might also govern the extent and rate of Polycomb dissociation from chromatin during mitosis, thereby potentially influencing the stability of epigenetic inheritance during cell division (Fonseca et al. 2012).
Although we have only focused our discussion on two specific histone phosphorylation sites, namely H3S10P and H3S28P, a number of other histone phosphorylation sites have an impact on chromatin structure (for reviews, see Baek 2011; Banerjee and Chakravarti 2011; Rossetto et al. 2012). As described above, the phosphorylation of PcG/TrxG components and histone residues represents a class of highly dynamic post-translational modifications that are able to communicate extra- and intra-cellular signals directly to chromatin. Signaling through the three branches of the MAPK pathway (ERK, p38, JNK) is an attractive candidate for further studies, as these cascades are primary signaling pathways communicating the need for cellular adaptation in response to changing environments and cellular stressors. Nevertheless, the transient interactions between kinases, phosphatases and their targets make it difficult to identify specific enzyme-substrate pairs. In addition, cross talk between kinase cascades makes deciphering the phosphorylation code all the more complicated. This might be the reason that, to our knowledge, only a few studies have identified chromatin-targeted phosphatases and epigenetically acting substrates in the context of gene regulation (see Koshibu et al. 2009, 2011; Rudenko et al. 2004; Zhang et al. 2013).
Regulation of PcG and TrxG function by metabolic processes
In cells, energy-rich nutrients are converted into ATP. Carbohydrates are metabolized to produce ATP by the metabolic pathways of glycolysis and the tricarboxylic acid (TCA) cycle coupled to oxidative phosphorylation. Cells alternatively utilize fatty acid oxidation and amino acid deamination/transamination to generate metabolic substrates for the TCA cycle. In addition to ATP, these pathways produce metabolites at various steps, most importantly acetyl coenzyme A (acetyl-CoA), nicotinamide adenine dinucleotides (NAD+/NADH), flavin adenine dinucleotides (FAD/FADH2), α-ketoglutarate (α-KG) and S-adenosylmethionine (SAM). Together, these metabolites and ATP are used as co-factors by histone and DNA-modifying enzymes to drive chromatin modifications and to change target gene expression, coupling cellular nutrient and energy states to the modulation of epigenetic signatures (for reviews, see Gut and Verdin 2013; Hitchler and Domann 2009; Ladurner 2006; Liu and Ward 2010; Smith and Denu 2009).
Emerging evidence suggests that histone acetyltransferases (HAT), like Drosophila dCBP/p300 and histone deacetylases (HDAC), like dRpd3, act as energy sensors that are activated by high and low cellular energy levels, respectively. HATs employ acetyl-CoA, whereas the SIRTUIN HDACs utilize NAD+ as co-factor substrates. Fluctuations in levels of these metabolites according to cellular energy status is thought to be translated into altered chromatin structure and gene transcription (for a review, see Kaelin and McKnight 2013). Since these mechanisms have recently been extensively reviewed, we will focus here on metabolic mechanisms that affect PcG and TrxG epigenetic function through the regulation of histone methylation dynamics, a paradigm for PcG/TrxG function.
S-adenosylmethionine-dependent histone and DNA methylation
The SET-domain containing methyltransferases, such as Drosophila E(z) and its mammalian homologs EZH1 and EZH2, plus the mammalian DNMT family of methyltransferases, use S-adenosylmethionine (SAM) as a methyl group donor to transfer methyl groups to H3K27 residues and cytosine bases in DNA, respectively (Smith and Denu 2009). SAM is a product of 1-carbon (1C) metabolism that utilizes zinc, methionine and vitamin B family members, such as folate, choline, vitamin B6 and B12, in enzymatic reactions. SAM is synthesized from methionine by the S-adenosylmethionine synthetase enzyme and serves as the chief physiological methyl group donor in cellular methyltransferase reactions.
The byproduct of SAM methyl-group donation is S-adenosyl homocysteine (SAH), a potent inhibitor of methyltransferases. Together, SAM and SAH can be seen as rate-limiting substrates and products of methyltransferase reactions. Decreases in cellular SAM concentration attributable to reduced intracellular energy levels might cause decreases in SAM:SAH ratios. This effect, in turn, might reduce cellular methylation potential and link energy homeostasis to the methylation of DNA and histone proteins (Hitchler and Domann 2009). The way that dietary or exercise-dependent changes in cellular energy levels alter SAM levels remains unclear.
However, the dietary intake of relevant methyl donor precursors, such as methionine, folate and choline, has been strongly linked to changes in cellular methylation potential. Currently, the most widely investigated are the consequences that these co-factors impose on DNA methylation (for a review, see Anderson et al. 2012).
Studies in mice and sheep have indicated that, during pregnancy, the nutrient availability of metabolites required for SAM synthesis can induce epigenetically mediated phenotypes (Sinclair et al. 2007; Waterland et al. 2006). A striking example of the way in which diet can affect the epigenome is the ability to alter coat color in offspring litters by supplemention of the diet of pregnant female mice with substrates needed for the production of SAM. Female agouti viable yellow mice, which have a yellow fur color, were fed a diet highly supplemented with methionine, vitamin B and zinc. They gave birth to offspring with heavily mottled, agouti fur color, as a result of increased levels of methylation at the agouti gene (Cooney et al. 2002; Dolinoy et al. 2007; Waterland and Jirtle 2003). This finding demonstrates that maternal diet regulates the establishment of gene expression patterns in utero and that environmental cues, such as dietary co-factors, can modulate the activity of chromatin-modifying enzymes in vivo.
In general, diets deficient for methyl donors are thought to cause DNA hypomethylation, correlating with a change in SAM and SAH ratios. DNA hypomethylation causes the reactivation of a number of oncogenes and genomic instability, resulting in the increased occurrence of tumors in mice (see Ghoshal and Farber 1984; Liu and Ward 2010; Mikol et al. 1983; Toyota and Suzuki 2010; Wainfan and Poirier 1992). Studies have generated conflicting results concerning the effect of dietary supplements of co-factors necessary for SAM biosynthesis on cancer risk in humans. Folate intake and high levels of B6 have been associated with a lower risk of developing colon, breast (Kato et al. 1999; Prinz-Langenohl et al. 2001), cervical (Butterworth et al. 1982) and lung (Hartman et al. 2001; Heimburger et al. 1988) cancer (for a review, see Liu and Ward 2010). Other trials have failed to demonstrate a reduction in cancer occurrence by increasing the dietary intake of such methyl donor precursors (for example Andreeva et al. 2012; Cole et al. 2007; Zhang et al. 2003). Nevertheless, hypomorphic mutations in folate- and methionine-metabolizing enzymes have been associated with increased cancer risk in humans (Cheng et al. 2012; Lightfoot et al. 2008). These mutations are thought to decrease the cellular production of the methyl-donor SAM, thereby inhibiting SAM-dependent transmethylations of DNA.
Whether histone methyltransferase activity and histone methylation levels are also affected by changes in SAM metabolism is less clear. A recent study suggested that folate and methionine deficiency leads to a reduced methylation of TrxG-targeted H3K4 residues in yeast and human cells causing changes to gene transcription (Sadhu et al. 2013). Furthermore, feeding female mice a zinc-deficient diet before ovulation dramatically disrupts the epigenetic signatures of oocytes and disturbs preimplantation development. The study revealed that such dietary deficiency causes decreased TrxG-mediated H3K4 trimethylation, decreased global DNA methylation and a strong increase in the transcription of repetitive DNA elements normally silenced by DNA methylation. Interestingly, dietary supplementation with a methyl donor (SAM) restores H3K4 trimethylation and improves the viability of oocytes (Tian and Diaz 2013). Similarly, feeding rats a methyl-deficient diet induces a global decrease in histone H3K9me3 and H4K20me3 marks and promotes carcinogenesis, although the mechanism behind these processes is less clear (Pogribny et al. 2007).
Recently, the nuclear accumulation of active S-adenosylmethionine synthetase has been demonstrated to correlate with increased PcG-mediated H3K27 trimethylation, indicating the necessity for nuclear S-adenosylmethionine synthetase to provide sufficient SAM for H3K27 methylation reactions (Reytor et al. 2009). In Drosophila, S-adenosylmethionine synthetase is highly expressed and essential for viability. Curiously, mutations in the S-adenosylmethionine synthetase gene affect position-effect variegation, a Drosophila model for studying the effect of proteins on the establishment of heterochromatin by assaying random silencing of reporter constructs located along the heterochromatin-euchromatin border. Furthermore, S-adenosylmethionine synthetase mutations also genetically interact with mutant alleles of Polycomb and enhance Polycomb-derived homeotic phenotypes (Larsson et al. 1996).
Even though these studies are only beginning to shape our understanding of the metabolically driven dynamics of histone and DNA methylation, such observations suggest that physiological SAM levels might, under certain circumstances, affect the active histone methylation potential of PcG/TrxG methyltransferases and of mammalian DNA methyltransferases.
FAD- and Fe(II)/α-ketoglutarate-dependent histone demethylases
Implementation of methylation marks is important for establishing epigenetic signatures; however, mediation of epigenetic transitions requires the activity of demethylases. Histone demethylases are generally grouped into two major classes: FAD-dependent demethylases and Fe(II)/α-ketoglutarate-dependent demethylases. They are distinguished by the nature of their catalytic domains. LSD1-domain demethylases, such as dLsd1 (mammalian KDM1A), are FAD-containing enzymes that remove histone mono- and dimethylation. Jumonji-domain demethylases, such as dLid (mammalian KDM5A) and dUtx (mammalian KDM6), use oxygen, Fe(II) and α-ketoglutarate as co-factors and catalyze demethylation of all three histone lysine methylation states (mono, di, tri; Hou and Yu 2010; Teperino et al. 2010; Walport et al. 2012).
The cellular production of FAD and α-ketoglutarate, co-factors utilized by histone demethylases, is highly dependent on cellular metabolism. FAD is produced in the mitochondria from riboflavin (vitamin B2) and is an important co-factor for several oxidation and reduction reactions necessary for generating ATP. Under aerobic conditions, glucose or fatty acid and amino acid catabolites are metabolized through the TCA cycle, which converts FAD into the high-energy electron carrier FADH2. FADH2 donates its electrons to the electron transport chain in the inner mitochondrial membrane, providing energy for the generation of ATP and in the process reverting back to FAD. Fluctuations in the levels of FAD and FADH2 might therefore be linked to cellular energy and oxygen availability, influencing the activity of histone demethylases.
LSD1 is one of the few nuclear proteins that utilize FAD. Recent evidence suggests that FAD-biosynthesis affects LSD1 activity, which, in turn, feeds back onto metabolic gene activity. For example, FAD-potentiated LSD1-catalyzed demethylation removes H3K4me1/me2 marks at energy-expenditure genes in mouse adipocytes. By limiting H3K4me1/me2 levels, a marker of actively transcribed genes, LSD1 might cause the repression of energy-expenditure genes in precisely those cells that store excess energy as triglycerides (Hino et al. 2012). Therefore, FAD availability guides a direct feedback loop that couples cellular nutrients to metabolic processes via LSD1.
One important source of α-ketoglutarate is the TCA cycle in which it is a key intermediate produced from isocitrate and is subsequently converted to succinyl CoA. These reactions are highly regulated and depend on the presence of essential co-factors, such as ATP, FAD and NAD+ and on levels of TCA cycle intermediates that control enzymatic activities by negative feedback loops. For example, succinyl-CoA, FADH2, NADH and high intracellular ATP levels have all been shown to inhibit α-ketoglutarate synthesis.
Other TCA cycle intermediates, such as fumarate and succinate, are oncometabolites accumulating in tumors when mutations arise in the TCA cycle enzymes isocitrate dehydrogenase, succinate dehydrogenase and fumarate hydratase (for reviews, see Adam et al. 2013; Thompson 2009). When present in elevated concentrations, fumarate and succinate are capable of inhibiting several α-ketoglutarate-dependent demethylases, including the JmjC-class of histone demethylases (Cervera et al. 2009; Chowdhury et al. 2011; Lu et al. 2012; Smith et al. 2007). However, to what extent physiological changes in α-ketoglutarate, succinate and fumarate levels modulate epigenetic signatures remains to be determined.
A further suggestion is that α-ketoglutarate-dependent demethylases are sensitive to environmental oxygen levels; this would largely depend on their (often unknown) Km values for oxygen (Mimura et al. 2011). Interestingly, hypoxic conditions appear to induce the inhibition of JARID and JmjD-family demethylases, resulting in increased H3K4me3, H3K9me3 and H3K36me3 histone methylation levels in tissue culture cells (Tausendschon et al. 2011; Zhou et al. 2010). These observations suggest that, at least some demethylases are sensitive to oxygen levels, potentially altering their catalytic behavior under hypoxic conditions known to occur at wound sites or in tumor environments.
α-Ketoglutarate-dependent demethylases might also be sensitive to ROS. ROS are often generated in response to cellular stress conditions, such as tissue damage or during tumor growth (see Gauron et al. 2013; Ziech et al. 2011). Similarly, carcinogenic metals, such as arsenite, trigger ROS production and oxidative stress, rendering histone demethylases susceptible to a decrease in catalytic activity (Chervona et al. 2012). Accordingly, demethylase activity can be enhanced in the presence of the ROS scavenger ascorbic acid (Blaschke et al. 2013; Yin et al. 2013).
As outlined above, histone methyltransferase and demethylase activity is dependent on metabolic co-enzymes, such as SAM, FAD and α-ketoglutarate, whose availabilities are governed by intracellular energy levels. Conceivably, histone methylation cycles can therefore synchronize dynamic transcriptional changes with metabolic information. However, the evidence in support of this concept in the context of histone methylation is still circumstantial. In the future, we need to improve our understanding of the way that dynamic physiological changes affect levels of the described metabolic co-factors in vivo. We also need to determine the way that fluctuations in co-factor and metabolite concentrations generally found in mitochondria or cytoplasm alter the function of nuclear-localized enzymes and to address the contradiction that intracellular energy levels positively regulate co-factor synthesis for both methyltransferases and demethylases.
O-GlcNAc modifications
The addition of O-GlcNAc to serine or threonine residues is a post-translational modification that regulates the stability, activity, or subcellular localization of many cellular proteins. One way by which O-GlcNAc mediates its effects on proteins is by competing with kinases for phosphorylation sites at specific serine or threonine residues, influencing the efficiency of signal transduction through cellular kinase pathways.
The enzyme uridine diphosphate N-acetylglucosaminepolypeptidyl transferase, known as OGT, catalyzes the transfer of O-GlcNAc sugar molecules from the donor uridine diphospho-N-acetylglucosamine (UDP-O-GlcNAc) onto target proteins. A glycoside hydrolase known as OGA catalyzes the removal of covalently linked O-GlcNAc from proteins. The antagonistic relationship between OGT and OGA makes O-GlcNAc modifications highly dynamic and reversible.
UDP-O-GlcNAc is the end product of the hexosamine biosynthesis pathway, which assimilates glucose (Vocadlo 2012). Therefore, UDP-O-GlcNAc levels depend on the availability of glucose and UDP-O-GlcNAcylation metabolism constitutes a powerful mechanism by which cellular processes are integrated with the nutritional status of the cell. For these reasons, O-GlcNAcylation kinetics are linked to insulin signaling and diabetic conditions (Issad et al. 2010) and to cancer (Fardini et al. 2013), among other human diseases.
Several studies co-localize O-GlcNAc modifications with transcription factors and with chromatin. Recent findings suggest that histones H2A, H2B, H3 and H4 are OGlcNAc modified and that epigenetic OGlcNAcylation is associated with transcriptional activation, gene silencing, mitotic chromatin regulation and the modulation of other histone modifications (for a review, see Hanover et al. 2012).
ChIP approaches have shown that OGlcNAcylation localizes to DNA-sequences also targeted by PcG complexes in fly tissues. Whether OGlcNAcylation localizes to these regions by OGlcNAc-modification of Polyhomeotic, a core member of PRC1, or by other modified substrates, such as histones, is not clear (Gambetta et al. 2009; Sinclair et al. 2009). Biochemical studies suggest that OGT itself can be recruited to chromatin and associates with distinct chromatin-regulating complexes (Chikanishi et al. 2010; Love et al. 2010; Yang et al. 2002). Intriguingly, the SET domain of the mammalian trithoraxrelated MLL5 protein becomes O-GlcNAcylated to enhance MLL5 H3K4-methyltransferase activity at target gene promoters (Fujiki et al. 2009). The interaction of OGlcNAcylation with both members of PcG and TrxG complexes suggests that this post-translational modification plays an important yet dynamic role in maintaining the fine balance between the activating and repressive activities of the PcG and TrxG complexes. Although the transcriptional consequences of PcG/TrxG-associated OGlcNAcylation remain to be fully elucidated, the glucose-dependent kinetics of the hexosamine biosynthesis pathway probably affect this novel epigenetic mark and imply that OGT modifications are under nutrient-sensitive control.
The studies discussed in the previous sections suggest that cellular nutrient sensing has a larger impact on epigenetic dynamics than has previously been imagined. The availability of vitamins and co-factors might play a huge role in the cellular ability to generate methyl-donors such as SAM, on the enzymatic activity of demethylases, such as LSD1 or UTX and on the substrate availability for glycosylating enzymes such as OGT. Many open questions remain about the way that cellular metabolism can directly effect chromatin structure and gene expression. However, initial evidence hints at an intricate feedback network between the cellular nutrient environment and post-translational modifications to either DNA or histones.
Regulation of PcG and TrxG function by cell-cycle dynamics
The function of PcG and TrxG genes as tumor suppressors and oncogenes and as regulators of the cell cycle has received much attention over the last decade. We now know that many cell-cycle genes are under the transcriptional or post-translational control of PcG and TrxG activity (see Iovino et al. 2013; Jacobs et al. 1999; Mohd-Sarip et al. 2012; Oktaba et al. 2008; Sen et al. 2011). More recently, several studies have addressed the opposite relationship: the way that cell-cycle dynamics can modulate PcG and TrxG function. These reports are particularly exciting, as they suggest that the processing of cellular signals during the various cell-cycle phases may impact PcG/TrxG target gene activity in distinct ways.
Cell divisions and corresponding cell-cycle dynamics are essential features of cell physiology. Progression through S-phase causes the disruption of the nucleosome architecture as the replication machinery passes through and epigenetic marks need to be re-established on both the newly synthesized DNA daughter strands. Mitotic chromosome condensation during M-phase hinders the access of protein complexes to DNA sequences and contributes to the exclusion of most regulatory proteins from chromatin during mitosis. Within this framework, the various cell-cycle dynamics observed during developmental proliferation, stem cell division, tumor growth and cellular senescence all impose specific kinetics on epigenetic processes. The integration of chromatin-modifying processes with these kinetics not only creates challenges to epigenetic stability but also offers opportunities for epigenetic plasticity in the context of cellular signaling environments.
PcG and TrxG complexes during DNA replication
Recent studies have addressed the molecular means by which epigenetic signatures can be maintained in the context of cell division (also reviewed in Probst et al. 2009; Simon and Kingston 2013). DNA replication needs to be coupled to the segregation of DNA and histone modifications. Cytosine-methylation marks are divided during S-phase in a semi-conservative manner, a mechanism inherent to semi-conservative DNA replication. Segregation of histone modifications and associated chromatin complexes could in principle involve the semi-conservative separation of these features between the two newly synthesized DNA strands. Alternatively, the copying of epigenetic signatures from one newly synthesized daughter strand inheriting all modifications onto the other newly synthesized, naked strand might be a possibility.
A series of recent studies in Drosophila suggests that both PRC1 and PRC2 components remain associated with replicating DNA and are found on both newly synthesized daughter strands immediately after the passage of the replication machinery (Francis et al. 2009; Lanzuolo et al. 2011, 2012; Petruk et al. 2012). As tested for PRC1, the association with replicating DNA is independent of other eukaryotic co-factors (Lengsfeld et al. 2012). The protein Psc, a subunit of PRC1, is sufficient to maintain PRC1 association with replicating DNA. Psc-Psc interactions between neighboring PRC1 complexes have been proposed to be able to bridge the complexes on parent and daughter strands through replication to support stable template association as the replication machinery passes (Lo et al. 2012).
Strikingly, a recent study suggests that the binding of PcG proteins to target sequences, in Drosophila and mammalian cells, appears to increase before replication, when compared with levels of bound PcG during the G1/S transition or after replication (Lanzuolo et al. 2011). These observations give rise to the hypothesis that PcG binding increases locally before replication to provide a sufficient number of protein complexes that can be segregated to both daughter strands and, therefore, ensures the re-establishment of PcG silencing after replication.
Investigation of inheritance patterns of PcG/TrxG-associated epigenetic histone modifications has provided conflicting results. Similar to the increase in PcG binding described above, an increase in H3K27me3 and H3K4me3 has been observed to occur before DNA replication (Lanzuolo et al. 2011). Other studies confirm the association of PcG and TrxG with replicating DNA but find no evidence for the presence of H3K27me3- and H3K4me3-modified histones at DNA during replication. Instead, methylated H3 histones are replaced by unmodified histones on nascent DNA daughter strands (Petruk et al. 2012). These unmodified histones might be immediately modified by the enzymatic activities of PRC2, as the complex remains bound to replicating DNA (Hansen et al. 2008; Petruk et al. 2012).
Disruption of epigenetic signatures during replication has not been linked to changes in PcG/TrxG target gene transcription. One is tempted to speculate that the need to re-establish epigenetic signatures after DNA replication provides a unique opportunity to alter epigenetic states by cellular signaling pathways during S-phase. S-phase dynamics and cell division have been associated with phenotypic plasticity in various cellular contexts (Fisher and Mechali 2003; Skora and Spradling 2010; Sustar et al. 2011; Sustar and Schubiger 2005). However, the regular histone turnover during the cell cycle in Drosophila is suggested to be much faster than the cell cycle itself, especially at actively transcribed genes and Polycomb-targeted regulatory DNA elements (Deal et al. 2010). Similar observations regarding fast turnover kinetics with respect to cell-cycle length have been made for the chromatin-association of PcG and TrxG proteins (Ficz et al. 2005). This argues against a simple scenario in which, during S-phase, disruption of nucleosome architecture challenges the maintenance of epigenetic memory, providing a crucial opportunity for altering chromatin signatures in the context of various cellular signals. Instead, fast turnover rates of histones and chromatin-associated complexes, especially at transcribed genes, provide additional opportunities for epigenetic reprogramming throughout G1 and G2 phases of the cell cycle. However, more studies are needed to define what role molecular mechanisms of DNA replication play in guaranteeing epigenetic stability or in promoting chromatin plasticity.
PcG and TrxG complexes during mitosis
During mitosis, dramatic chromatin architecture changes occur in the context of chromosome condensation and segregation. Many regulatory proteins are excluded from chromatin during mitosis; however, PcG-repressive and TrxG-activated states at target genes must be preserved.
Several studies in Drosophila suggest that the majority of PcG and TrxG proteins tested dissociate from mitotic chromosomes, although some do indeed remain bound. Older studies in Drosophila revealed that the core PRC1 components Polycomb and Polyhomeotic largely dissociate during mitosis, whereas Psc remains bound, albeit to a lesser extent than in interphase (Buchenau et al. 1998; Dietzel et al. 1999). The partial association of Psc with mitotic chromosomes has recently been confirmed by biochemical fractionation and ChIP approaches. These studies verified that a large fraction of Psc remains associated with chromosomes but only at a fraction of the target sites normally occupied by Psc during interphase (Follmer et al. 2012). Recent imaging studies extended the observations of pronounced mitotic dissociation to the PRC2 component E(z) and the PcG recruitment factor Pleiohomeotic (Fonseca et al. 2012; Steffen et al. 2013). In contrast, the TrxG member Ash1 remains associated with chromatin throughout mitosis (Steffen et al. 2013). This is reminiscent of a phenomenon in mammalian cells termed bookmarking, a mechanism whereby some regulatory proteins that remain bound to chromatin during mitosis mark previously active loci for fast post-mitotic re-activation (Zhao et al. 2011). Additional studies suggest that several PcG and TrxG components in mammalian cells also dissociate from chromosomes during mitosis (Aoto et al. 2008; Miyagishima et al. 2003; Voncken et al. 1999).
Although individual results vary, the studies described above highlight a pronounced role for chromatin-associated complexes in maintaining epigenetic memory during DNA replication by remaining associated with replicating DNA and re-establishing an appropriate set of modifications at newly incorporated histones. In contrast, established histone marks might guide the re-association of dissociated protein complexes to de-condensing chromatin after mitosis. We currently do not understand the nature of signals that guide chromatin association and dissociation. Possibly, cell-cycle-related phosphorylation by cyclin-dependent kinases (CDK) plays a role. As discussed above, human EZH2 is phosphorylated by CDK1 and CDK2 kinases and interferes with the EZH2 silencing function at target genes (Chen et al. 2010; Kaneko et al. 2010; Wei et al. 2011; Wu and Zhang 2011). The fly MSK homolog Jil-1 has been suggested to control H3S28 phosphorylation governing the extent and rate of Polycomb dissociation from chromatin during mitosis (Fonseca et al. 2012). However, a detailed dissection of this pathway, which is potentially able to integrate cellular signals with cell-cycle dynamics, is still lacking. Therefore, future studies need to investigate the dynamics of PcG/TrxG chromatin dissociation during mitosis and to examine the regulation of the re-association of chromatin factors in daughter cells by cellular signaling pathways guiding differentiation and proliferation. These mechanisms are particularly important for understanding the way that daughter cells acquire different fates, such as during the asymmetric divisions of embryonic and adult stem cells.
Concluding remarks and future perspectives
The way cells respond to environmental cues by changing gene expression patterns is important for cell survival and adaptation. These changes might be short-lived and mediated by acute activation of DNA-sequence-specific transcription factors. However, the observations described in the studies above suggest that direct signaling-induced functional changes of epigenetically acting protein complexes contribute to the initiation and maintenance of altered gene expression patterns. We have outlined, in this review, the manner in which the gene activating and silencing complexes of the PcG and TrxG protein families regulate chromatin dynamics in response to signaling through kinase cascades, in response to changes in cellular metabolic state, or in response to cell-cycle dynamics. Although these pathways are probably important mediators of PcG/TrxG function in differentiation decisions during development, the acute nature and the wiring of some of these pathways under cellular stress responses suggests that PcG/TrxG function is highly dynamic and can be altered in response to environmental cues throughout the lifetime of a cell.
Future studies will face a multitude of challenges as they aim to elucidate the exact regulation and function of these signaling pathways to chromatin. How is the phosphorylation of PcG/TrxG proteins integrated to guide their activity at target genes? This is completely unclear at present. Additionally, the role that nutrition plays in driving the metabolic remodeling of epigenetic landscapes and the way that remodeling, in turn, promotes adaptation to energy availability are insufficiently understood. The maintenance of epigenetic stability during cell division is an area of intense study; however, few have addressed the opportunity that this process provides to promote epigenetic plasticity.
One of the biggest challenges will be to understand the way in which the specificity in gene regulation arises from global signaling events. Complex kinase signaling networks, energetic and metabolic fluctuations and large-scale remodeling processes during cell division can globally alter chromatin-modifying proteins within a cell. However, phenotypic adaptation to these cellular signals might only be mediated by the activation or silencing of specific target genes. The means of achieving specificity have been highlighted in this review: kinases, such as JNK and ERK, or metabolically driven enzymes, such as OGT, are recruited to chromatin and localize to specific gene-regulatory regions, thereby focusing potentially global effects to the local regulation of substrate activity and target gene transcription.
The plasticity that is increasingly associated with epigenetic processes further raises the question regarding at which time scales such signaling-induced changes are translated into changes of gene expression patterns. The cellular signals that are epigenetically integrated as acute short-term responses and the signals that cause long-term, sometimes inheritable, changes to gene expression patterns are unknown. The studies described here suggest that the dietary nutrition of female mice during pregnancy causes changes to the expression of fur color genes in their offspring. These new expression patterns are initiated by local DNA methylation and maintained throughout the lifetime of the animal. In contrast, acute modulation of PcG/TrxG activity via the direct phosphorylation of the respective protein complexes probably causes functional changes that revert upon signal termination. Short-term signals might also be integrated to establish long-term epigenetic changes at specific target genes.
We are only beginning to understand the molecular tools that are available to cells and that can implement epigenetic shifts in chromatin state. Much remains to be learned about why these shifts occur, how they are initiated and for how long these changes are implemented to allow cells to attain acute or long-term changes in cellular phenotypes.
References
Adam J, Yang M, Soga T, Pollard PJ (2013) Rare insights into cancer biology. Oncogene. doi:10.1038/onc.2013.222
Anderson OS, Sant KE, Dolinoy DC (2012) Nutrition and epigenetics: an interplay of dietary methyl donors, one-carbon metabolism and DNA methylation. J Nutr Biochem 23:853–859
Andreeva VA, Touvier M, Kesse-Guyot E, Julia C, Galan P, Hercberg S (2012) B vitamin and/or ω-3 fatty acid supplementation and cancer: ancillary findings from the supplementation with folate, vitamins B6 and B12, and/or omega-3 fatty acids (SU.FOL.OM3) randomized trial.Arch Intern Med 172:540–547
Aoto T, Saitoh N, Sakamoto Y, Watanabe S, Nakao M (2008) Polycomb group protein-associated chromatin is reproduced in post-mitotic G1 phase and is required for S phase progression. J Biol Chem 283:18905–18915
Ardehali MB, Mei A, Zobeck KL, Caron M, Lis JT, Kusch T (2011) Drosophila Set1 is the major histone H3 lysine 4 trimethyltransferase with role in transcription. EMBO J 30:2817–2828
Baek SH (2011) When signaling kinases meet histones and histone modifiers in the nucleus. Mol Cell 42:274–284
Banerjee T, Chakravarti D (2011) A peek into the complex realm of histone phosphorylation. Mol Cell Biol 31:4858–4873
Beisel C, Paro R (2011) Silencing chromatin: comparing modes and mechanisms. Nat Rev Genet 12:123–135
Blaschke K, Ebata KT, Karimi MM, Zepeda-Martinez JA, Goyal P, Mahapatra S, Tam A, Laird DJ, Hirst M, Rao A et al (2013) Vitamin C induces Tet-dependent DNA demethylation and a blastocyst-like state in ES cells. Nature 500:222–226
Bosch M, Serras F, Martin-Blanco E, Baguna J (2005) JNK signaling pathway required for wound healing in regenerating Drosophila wing imaginal discs. Dev Biol 280:73–86
Breiling A, Turner BM, Bianchi ME, Orlando V (2001) General transcription factors bind promoters repressed by Polycomb group proteins. Nature 412:651–655
Buchenau P, Hodgson J, Strutt H, Arndt-Jovin DJ (1998) The distribution of polycomb-group proteins during cell division and development in Drosophila embryos: impact on models for silencing. J Cell Biol 141:469–481
Butterworth CE Jr, Hatch KD, Gore H, Mueller H, Krumdieck CL (1982) Improvement in cervical dysplasia associated with folic acid therapy in users of oral contraceptives. Am J Clin Nutr 35:73–82
Cao R, Tsukada Y, Zhang Y (2005) Role of Bmi-1 and Ring1A in H2A ubiquitylation and Hox gene silencing. Mol Cell 20:845–854
Caretti G, Palacios D, Sartorelli V, Puri PL (2011) Phosphoryl-EZH-ion. Cell Stem Cell 8:262–265
Cervera AM, Bayley JP, Devilee P, McCreath KJ (2009) Inhibition of succinate dehydrogenase dysregulates histone modification in mammalian cells. Mol Cancer 8:89
Cha TL, Zhou BP, Xia W, Wu Y, Yang CC, Chen CT, Ping B, Otte AP, Hung MC (2005) Akt-mediated phosphorylation of EZH2 suppresses methylation of lysine 27 in histone H3. Science 310:306–310
Chen S, Bohrer LR, Rai AN, Pan Y, Gan L, Zhou X, Bagchi A, Simon JA, Huang H (2010) Cyclin-dependent kinases regulate epigenetic gene silencing through phosphorylation of EZH2. Nat Cell Biol 12:1108–1114
Cheng NN, Sinclair DA, Campbell RB, Brock HW (1994) Interactions of polyhomeotic with Polycomb group genes of Drosophila melanogaster. Genetics 138:1151–1162
Cheng H, Deng Z, Wang Z, Zhang W, Su J (2012) MTHFR C677T polymorphisms are associated with aberrant methylation of the IGF-2 gene in transitional cell carcinoma of the bladder. J Biomed Res 26:77–83
Chervona Y, Arita A, Costa M (2012) Carcinogenic metals and the epigenome: understanding the effect of nickel, arsenic, and chromium. Metallomics 4:619–627
Chikanishi T, Fujiki R, Hashiba W, Sekine H, Yokoyama A, Kato S (2010) Glucose-induced expression of MIP-1 genes requires O-GlcNAc transferase in monocytes. Biochem Biophys Res Commun 394:865–870
Chow CW, Davis RJ (2006) Proteins kinases: chromatin-associated enzymes? Cell 127:887–890
Chowdhury R, Yeoh KK, Tian YM, Hillringhaus L, Bagg EA, Rose NR, Leung IK, Li XS, Woon EC, Yang M et al (2011) The oncometabolite 2-hydroxyglutarate inhibits histone lysine demethylases. EMBO Rep 12:463–469
Classen AK, Bunker BD, Harvey KF, Vaccari T, Bilder D (2009) A tumor suppressor activity of Drosophila Polycomb genes mediated by JAK-STAT signaling. Nat Genet 41:1150–1155
Cole BF, Baron JA, Sandler RS, Haile RW, Ahnen DJ, Bresalier RS et al (2007) Folic acid for the prevention of colorectal adenomas: a randomized clinical trial. JAMA 297:2351-2359
Cooney CA, Dave AA, Wolff GL (2002) Maternal methyl supplements in mice affect epigenetic variation and DNA methylation of offspring. J Nutr 132:2393S–2400S
Deal RB, Henikoff JG, Henikoff S (2010) Genome-wide kinetics of nucleosome turnover determined by metabolic labeling of histones. Science 328:1161–1164
Dietzel S, Niemann H, Bruckner B, Maurange C, Paro R (1999) The nuclear distribution of Polycomb during Drosophila melanogaster development shown with a GFP fusion protein. Chromosoma 108:83–94
Dolinoy DC, Huang D, Jirtle RL (2007) Maternal nutrient supplementation counteracts bisphenol A-induced DNA hypomethylation in early development. Proc Natl Acad Sci U S A 104:13056–13061
Dorighi KM, Tamkun JW (2013) The trithorax group proteins Kismet and ASH1 promote H3K36 dimethylation to counteract Polycomb group repression in Drosophila. Development 140:4182–4192
Drobic B, Perez-Cadahia B, Yu J, Kung SK, Davie JR (2010) Promoter chromatin remodeling of immediate-early genes is mediated through H3 phosphorylation at either serine 28 or 10 by the MSK1 multi-protein complex. Nucleic Acids Res 38:3196–3208
Dyson MH, Thomson S, Inagaki M, Goto H, Arthur SJ, Nightingale K, Iborra FJ, Mahadevan LC (2005) MAP kinase-mediated phosphorylation of distinct pools of histone H3 at S10 or S28 via mitogen- and stress-activated kinase 1/2. J Cell Sci 118:2247–2259
Eissenberg JC, Lee MG, Schneider J, Ilvarsonn A, Shiekhattar R, Shilatifard A (2007) The trithorax-group gene in Drosophila little imaginal discs encodes a trimethylated histone H3 Lys4 demethylase. Nat Struct Mol Biol 14:344–346
Elderkin S, Maertens GN, Endoh M, Mallery DL, Morrice N, Koseki H, Peters G, Brockdorff N, Hiom K (2007) A phosphorylated form of Mel-18 targets the Ring1B histone H2A ubiquitin ligase to chromatin. Mol Cell 28:107–120
Endoh M, Endo TA, Endoh T, Isono K, Sharif J, Ohara O, Toyoda T, Ito T, Eskeland R, Bickmore WA et al (2012) Histone H2A mono-ubiquitination is a crucial step to mediate PRC1-dependent repression of developmental genes to maintain ES cell identity. PLoS Genet 8:e1002774
Fardini Y, Dehennaut V, Lefebvre T, Issad T (2013) O-GlcNAcylation: a new cancer hallmark? Front Endocrinol (Lausanne) 4:99
Ficz G, Heintzmann R, Arndt-Jovin DJ (2005) Polycomb group protein complexes exchange rapidly in living Drosophila. Development 132:3963–3976
Fisher D, Mechali M (2003) Vertebrate HoxB gene expression requires DNA replication. EMBO J 22:3737–3748
Follmer NE, Wani AH, Francis NJ (2012) A polycomb group protein is retained at specific sites on chromatin in mitosis. PLoS Genet 8:e1003135
Fonseca JP, Steffen PA, Muller S, Lu J, Sawicka A, Seiser C, Ringrose L (2012) In vivo Polycomb kinetics and mitotic chromatin binding distinguish stem cells from differentiated cells. Genes Dev 26:857–871
Forcales SV, Albini S, Giordani L, Malecova B, Cignolo L, Chernov A, Coutinho P, Saccone V, Consalvi S, Williams R et al (2012) Signal-dependent incorporation of MyoD-BAF60c into Brg1-based SWI/SNF chromatin-remodelling complex. EMBO J 31:301–316
Francis NJ, Kingston RE, Woodcock CL (2004) Chromatin compaction by a polycomb group protein complex. Science 306:1574–1577
Francis NJ, Follmer NE, Simon MD, Aghia G, Butler JD (2009) Polycomb proteins remain bound to chromatin and DNA during DNA replication in vitro. Cell 137:110–122
Franke A, DeCamillis M, Zink D, Cheng N, Brock HW, Paro R (1992) Polycomb and polyhomeotic are constituents of a multimeric protein complex in chromatin of Drosophila melanogaster. EMBO J 11:2941–2950
Fujiki R, Chikanishi T, Hashiba W, Ito H, Takada I, Roeder RG, Kitagawa H, Kato S (2009) GlcNAcylation of a histone methyltransferase in retinoic-acid-induced granulopoiesis. Nature 459:455–459
Gambetta MC, Oktaba K, Muller J (2009) Essential role of the glycosyltransferase sxc/Ogt in polycomb repression. Science 325:93–96
Gauron C, Rampon C, Bouzaffour M, Ipendey E, Teillon J, Volovitch M, Vriz S (2013) Sustained production of ROS triggers compensatory proliferation and is required for regeneration to proceed. Sci Rep 3:2084
Gehani SS, Agrawal-Singh S, Dietrich N, Christophersen NS, Helin K, Hansen K (2010) Polycomb group protein displacement and gene activation through MSK-dependent H3K27me3S28 phosphorylation. Mol Cell 39:886–900
Ghoshal AK, Farber E (1984) The induction of liver cancer by dietary deficiency of choline and methionine without added carcinogens. Carcinogenesis 5:1367–1370
Goke J, Chan YS, Yan J, Vingron M, Ng HH (2013) Genome-wide kinase-chromatin interactions reveal the regulatory network of ERK signaling in human embryonic stem cells. Mol Cell 50:844–855
Gut P, Verdin E (2013) The nexus of chromatin regulation and intermediary metabolism. Nature 502:489–498
Hallson G, Hollebakken RE, Li T, Syrzycka M, Kim I, Cotsworth S, Fitzpatrick KA, Sinclair DA, Honda BM (2012) dSet1 is the main H3K4 di- and tri-methyltransferase throughout Drosophila development. Genetics 190:91–100
Hanover JA, Krause MW, Love DC (2012) Bittersweet memories: linking metabolism to epigenetics through O-GlcNAcylation. Nat Rev Mol Cell Biol 13:312–321
Hansen KH, Bracken AP, Pasini D, Dietrich N, Gehani SS, Monrad A, Rappsilber J, Lerdrup M, Helin K (2008) A model for transmission of the H3K27me3 epigenetic mark. Nat Cell Biol 10:1291–1300
Hartman TJ, Woodson K, Stolzenberg-Solomon R, Virtamo J, Selhub J, Barrett MJ, Albanes D (2001) Association of the B-vitamins pyridoxal 5′-phosphate (B(6)), B(12), and folate with lung cancer risk in older men. Am J Epidemiol 153:688–694
Hatano A, Matsumoto M, Higashinakagawa T, Nakayama KI (2010) Phosphorylation of the chromodomain changes the binding specificity of Cbx2 for methylated histone H3. Biochem Biophys Res Commun 397:93–99
Heimburger DC, Alexander CB, Birch R, Butterworth CE Jr, Bailey WC, Krumdieck CL (1988) Improvement in bronchial squamous metaplasia in smokers treated with folate and vitamin B12. Report of a preliminary randomized, double-blind intervention trial. JAMA 259:1525–1530
Herz HM, Mohan M, Garruss AS, Liang K, Takahashi YH, Mickey K, Voets O, Verrijzer CP, Shilatifard A (2012) Enhancer-associated H3K4 monomethylation by Trithorax-related, the Drosophila homolog of mammalian Mll3/Mll4. Genes Dev 26:2604–2620
Hino S, Sakamoto A, Nagaoka K, Anan K, Wang Y, Mimasu S, Umehara T, Yokoyama S, Kosai K, Nakao M (2012) FAD-dependent lysine-specific demethylase-1 regulates cellular energy expenditure. Nat Commun 3:758
Hitchler MJ, Domann FE (2009) Metabolic defects provide a spark for the epigenetic switch in cancer. Free Radic Biol Med 47:115–127
Hou H, Yu H (2010) Structural insights into histone lysine demethylation. Curr Opin Struct Biol 20:739–748
Ingham PW (1984) A gene that regulates the bithorax complex differentially in larval and adult cells of Drosophila. Cell 37:815–823
Iovino N, Ciabrelli F, Cavalli G (2013) PRC2 controls Drosophila oocyte cell fate by repressing cell cycle genes. Dev Cell 26:431–439
Issad T, Masson E, Pagesy P (2010) O-GlcNAc modification, insulin signaling and diabetic complications. Diabetes Metab 36:423–435
Jacobs JJ, Kieboom K, Marino S, DePinho RA, van Lohuizen M (1999) The oncogene and Polycomb-group gene bmi-1 regulates cell proliferation and senescence through the ink4a locus. Nature 397:164–168
Kaelin WG Jr, McKnight SL (2013) Influence of metabolism on epigenetics and disease. Cell 153:56–69
Kaneko S, Li G, Son J, Xu CF, Margueron R, Neubert TA, Reinberg D (2010) Phosphorylation of the PRC2 component Ezh2 is cell cycle-regulated and up-regulates its binding to ncRNA. Genes Dev 24:2615–2620
Kato I, Dnistrian AM, Schwartz M, Toniolo P, Koenig K, Shore RE, Akhmedkhanov A, Zeleniuch-Jacquotte A, Riboli E (1999) Serum folate, homocysteine and colorectal cancer risk in women: a nested case–control study. Br J Cancer 79:1917–1922
Kim J, Hwangbo J, Wong PK (2011) p38 MAPK-mediated Bmi-1 down-regulation and defective proliferation in ATM-deficient neural stem cells can be restored by Akt activation. PLoS One 6:e16615
Klebes A, Sustar A, Kechris K, Li H, Schubiger G, Kornberg TB (2005) Regulation of cellular plasticity in Drosophila imaginal disc cells by the Polycomb group, trithorax group and lama genes. Development 132:3753–3765
Klein AM, Zaganjor E, Cobb MH (2013) Chromatin-tethered MAPKs. Curr Opin Cell Biol 25:272–277
Klymenko T, Papp B, Fischle W, Kocher T, Schelder M, Fritsch C, Wild B, Wilm M, Muller J (2006) A Polycomb group protein complex with sequence-specific DNA-binding and selective methyl-lysine-binding activities. Genes Dev 20:1110–1122
Kohli RM, Zhang Y (2013) TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502:472–479
Koshibu K, Graff J, Beullens M, Heitz FD, Berchtold D, Russig H, Farinelli M, Bollen M, Mansuy IM (2009) Protein phosphatase 1 regulates the histone code for long-term memory. J Neurosci 29:13079–13089
Koshibu K, Graff J, Mansuy IM (2011) Nuclear protein phosphatase-1: an epigenetic regulator of fear memory and amygdala long-term potentiation. Neuroscience 173:30–36
Ladurner AG (2006) Rheostat control of gene expression by metabolites. Mol Cell 24:1–11
Lagarou A, Mohd-Sarip A, Moshkin YM, Chalkley GE, Bezstarosti K, Demmers JA, Verrijzer CP (2008) dKDM2 couples histone H2A ubiquitylation to histone H3 demethylation during Polycomb group silencing. Genes Dev 22:2799–2810
Lanzuolo C, Orlando V (2012) Memories from the polycomb group proteins. Annu Rev Genet 46:561–589
Lanzuolo C, Lo Sardo F, Diamantini A, Orlando V (2011) PcG complexes set the stage for epigenetic inheritance of gene silencing in early S phase before replication. PLoS Genet 7:e1002370
Lanzuolo C, Lo Sardo F, Orlando V (2012) Concerted epigenetic signatures inheritance at PcG targets through replication. Cell Cycle 11:1296–1300
Larsson J, Zhang J, Rasmuson-Lestander A (1996) Mutations in the Drosophila melanogaster gene encoding S-adenosylmethionine synthetase [corrected] suppress position-effect variegation. Genetics 143:887–896
Lau PN, Cheung P (2011) Histone code pathway involving H3 S28 phosphorylation and K27 acetylation activates transcription and antagonizes polycomb silencing. Proc Natl Acad Sci U S A 108:2801–2806
Lee N, Maurange C, Ringrose L, Paro R (2005) Suppression of Polycomb group proteins by JNK signalling induces transdetermination in Drosophila imaginal discs. Nature 438:234–237
Lee N, Zhang J, Klose RJ, Erdjument-Bromage H, Tempst P, Jones RS, Zhang Y (2007) The trithorax-group protein Lid is a histone H3 trimethyl-Lys4 demethylase. Nat Struct Mol Biol 14:341–343
Lengsfeld BM, Berry KN, Ghosh S, Takahashi M, Francis NJ (2012) A Polycomb complex remains bound through DNA replication in the absence of other eukaryotic proteins. Sci Rep 2:661
Lightfoot TJ, Barrett JH, Bishop T, Northwood EL, Smith G, Wilkie MJ, Steele RJ, Carey FA, Key TJ, Wolf R et al (2008) Methylene tetrahydrofolate reductase genotype modifies the chemopreventive effect of folate in colorectal adenoma, but not colorectal cancer. Cancer Epidemiol Biomarkers Prev 17:2421–2430
Liu JJ, Ward RL (2010) Folate and one-carbon metabolism and its impact on aberrant DNA methylation in cancer. Adv Genet 71:79–121
Lo SM, Follmer NE, Lengsfeld BM, Madamba EV, Seong S, Grau DJ, Francis NJ (2012) A bridging model for persistence of a polycomb group protein complex through DNA replication in vitro. Mol Cell 46:784–796
Love DC, Ghosh S, Mondoux MA, Fukushige T, Wang P, Wilson MA, Iser WB, Wolkow CA, Krause MW, Hanover JA (2010) Dynamic O-GlcNAc cycling at promoters of Caenorhabditis elegans genes regulating longevity, stress, and immunity. Proc Natl Acad Sci U S A 107:7413–7418
Lu C, Ward PS, Kapoor GS, Rohle D, Turcan S, Abdel-Wahab O, Edwards CR, Khanin R, Figueroa ME, Melnick A et al (2012) IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483:474–478
Martinez AM, Schuettengruber B, Sakr S, Janic A, Gonzalez C, Cavalli G (2009) Polyhomeotic has a tumor suppressor activity mediated by repression of Notch signaling. Nat Genet 41:1076–1082
Mikol YB, Hoover KL, Creasia D, Poirier LA (1983) Hepatocarcinogenesis in rats fed methyl-deficient, amino acid-defined diets. Carcinogenesis 4:1619–1629
Mills AA (2010) Throwing the cancer switch: reciprocal roles of polycomb and trithorax proteins. Nat Rev Cancer 10:669–682
Mimura I, Tanaka T, Wada Y, Kodama T, Nangaku M (2011) Pathophysiological response to hypoxia—from the molecular mechanisms of malady to drug discovery: epigenetic regulation of the hypoxic response via hypoxia-inducible factor and histone modifying enzymes. J Pharmacol Sci 115:453–458
Miyagishima H, Isono K, Fujimura Y, Iyo M, Takihara Y, Masumoto H, Vidal M, Koseki H (2003) Dissociation of mammalian Polycomb-group proteins, Ring1B and Rae28/Ph1, from the chromatin correlates with configuration changes of the chromatin in mitotic and meiotic prophase. Histochem Cell Biol 120:111–119
Mohd-Sarip A, Lagarou A, Doyen CM, van der Knaap JA, Aslan U, Bezstarosti K, Yassin Y, Brock HW, Demmers JA, Verrijzer CP (2012) Transcription-independent function of Polycomb group protein PSC in cell cycle control. Science 336:744–747
Muller J, Verrijzer P (2009) Biochemical mechanisms of gene regulation by polycomb group protein complexes. Curr Opin Genet Dev 19:150–158
Nacerddine K, Beaudry JB, Ginjala V, Westerman B, Mattiroli F, Song JY, van der Poel H, Ponz OB, Pritchard C, Cornelissen-Steijger P et al (2012) Akt-mediated phosphorylation of Bmi1 modulates its oncogenic potential, E3 ligase activity, and DNA damage repair activity in mouse prostate cancer. J Clin Invest 122:1920–1932
Nakamura S, Oshima M, Yuan J, Saraya A, Miyagi S, Konuma T, Yamazaki S, Osawa M, Nakauchi H, Koseki H et al (2012) Bmi1 confers resistance to oxidative stress on hematopoietic stem cells. PLoS One 7:e36209
Ng J, Hart CM, Morgan K, Simon JA (2000) A Drosophila ESC-E(Z) protein complex is distinct from other polycomb group complexes and contains covalently modified ESC. Mol Cell Biol 20:3069–3078
Niessen HE, Demmers JA, Voncken JW (2009) Talking to chromatin: post-translational modulation of polycomb group function. Epigenetics Chromatin 2:10
Noguchi K, Shiurba R, Higashinakagawa T (2002) Nuclear translocation of mouse Polycomb M33 protein in regenerating liver. Biochem Bioph Res Co 291:508–515
Oktaba K, Gutierrez L, Gagneur J, Girardot C, Sengupta AK, Furlong EE, Muller J (2008) Dynamic regulation by polycomb group protein complexes controls pattern formation and the cell cycle in Drosophila. Dev Cell 15:877–889
Owusu-Ansah E, Banerjee U (2009) Reactive oxygen species prime Drosophila haematopoietic progenitors for differentiation. Nature 461:537–541
Palacios D, Mozzetta C, Consalvi S, Caretti G, Saccone V, Proserpio V, Marquez VE, Valente S, Mai A, Forcales SV et al (2010) TNF/p38alpha/polycomb signaling to Pax7 locus in satellite cells links inflammation to the epigenetic control of muscle regeneration. Cell Stem Cell 7:455–469
Pasini D, Malatesta M, Jung HR, Walfridsson J, Willer A, Olsson L, Skotte J, Wutz A, Porse B, Jensen ON et al (2010) Characterization of an antagonistic switch between histone H3 lysine 27 methylation and acetylation in the transcriptional regulation of Polycomb group target genes. Nucleic Acids Res 38:4958–4969
Petruk S, Sedkov Y, Johnston DM, Hodgson JW, Black KL, Kovermann SK, Beck S, Canaani E, Brock HW, Mazo A (2012) TrxG and PcG proteins but not methylated histones remain associated with DNA through replication. Cell 150:922–933
Pogribny IP, Tryndyak VP, Muskhelishvili L, Rusyn I, Ross SA (2007) Methyl deficiency, alterations in global histone modifications, and carcinogenesis. J Nutr 137:216S–222S
Prickaerts P, Niessen HE, Mouchel-Vielh E, Dahlmans VE, van den Akker GG, Geijselaers C, Adriaens ME, Spaapen F, Takihara Y, Rapp UR et al (2012) MK3 controls Polycomb target gene expression via negative feedback on ERK. Epigenetics Chromatin 5:12
Prinz-Langenohl R, Fohr I, Pietrzik K (2001) Beneficial role for folate in the prevention of colorectal and breast cancer. Eur J Nutr 40:98–105
Probst AV, Dunleavy E, Almouzni G (2009) Epigenetic inheritance during the cell cycle. Nat Rev Mol Cell Biol 10:192–206
Ramet M, Lanot R, Zachary D, Manfruelli P (2002) JNK signaling pathway is required for efficient wound healing in Drosophila. Dev Biol 241:145–156
Rampalli S, Li L, Mak E, Ge K, Brand M, Tapscott SJ, Dilworth FJ (2007) p38 MAPK signaling regulates recruitment of Ash2L-containing methyltransferase complexes to specific genes during differentiation. Nat Struct Mol Biol 14:1150–1156
Rao PS, Satelli A, Zhang S, Srivastava SK, Srivenugopal KS, Rao US (2009) RNF2 is the target for phosphorylation by the p38 MAPK and ERK signaling pathways. Proteomics 9:2776–2787
Reynolds N, Salmon-Divon M, Dvinge H, Hynes-Allen A, Balasooriya G, Leaford D, Behrens A, Bertone P, Hendrich B (2012) NuRD-mediated deacetylation of H3K27 facilitates recruitment of Polycomb Repressive Complex 2 to direct gene repression. EMBO J 31:593–605
Reytor E, Perez-Miguelsanz J, Alvarez L, Perez-Sala D, Pajares MA (2009) Conformational signals in the C-terminal domain of methionine adenosyltransferase I/III determine its nucleocytoplasmic distribution. FASEB J 23:3347–3360
Rossetto D, Avvakumov N, Cote J (2012) Histone phosphorylation: a chromatin modification involved in diverse nuclear events. Epigenetics 7:1098–1108
Rudenko A, Bennett D, Alphey L (2004) PP1beta9C interacts with Trithorax in Drosophila wing development. Dev Dyn 231:336–341
Rudolph T, Yonezawa M, Lein S, Heidrich K, Kubicek S, Schafer C, Phalke S, Walther M, Schmidt A, Jenuwein T et al (2007) Heterochromatin formation in Drosophila is initiated through active removal of H3K4 methylation by the LSD1 homolog SU(VAR)3-3. Mol Cell 26:103–115
Ruthenburg AJ, Allis CD, Wysocka J (2007) Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol Cell 25:15–30
Sadhu MJ, Guan Q, Li F, Sales-Lee J, Iavarone AT, Hammond MC, Cande WZ, Rine J (2013) Nutritional control of epigenetic processes in yeast and human cells. Genetics 195:831–844
Sassone-Corsi P, Mizzen CA, Cheung P, Crosio C, Monaco L, Jacquot S, Hanauer A, Allis CD (1999) Requirement of Rsk-2 for epidermal growth factor-activated phosphorylation of histone H3. Science 285:886–891
Sawarkar R, Paro R (2010) Interpretation of developmental signaling at chromatin: the Polycomb perspective. Dev Cell 19:651–661
Scheuermann JC, de Ayala Alonso AG, Oktaba K, Ly-Hartig N, McGinty RK, Fraterman S, Wilm M, Muir TW, Muller J (2010) Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB. Nature 465:243–247
Schuettengruber B, Ganapathi M, Leblanc B, Portoso M, Jaschek R, Tolhuis B, van Lohuizen M, Tanay A, Cavalli G (2009) Functional anatomy of polycomb and trithorax chromatin landscapes in Drosophila embryos. PLoS Biol 7:e13
Schuettengruber B, Martinez AM, Iovino N, Cavalli G (2011) Trithorax group proteins: switching genes on and keeping them active. Nat Rev Mol Cell Biol 12:799–814
Schwartz YB, Kahn TG, Stenberg P, Ohno K, Bourgon R, Pirrotta V (2010) Alternative epigenetic chromatin states of polycomb target genes. PLoS Genet 6:e1000805
Schwermann J, Rathinam C, Schubert M, Schumacher S, Noyan F, Koseki H, Kotlyarov A, Klein C, Gaestel M (2009) MAPKAP kinase MK2 maintains self-renewal capacity of haematopoietic stem cells. EMBO J 28:1392–1406
Secombe J, Li L, Carlos L, Eisenman RN (2007) The Trithorax group protein Lid is a trimethyl histone H3K4 demethylase required for dMyc-induced cell growth. Genes Dev 21:537–551
Sen N, Satija YK, Das S (2011) PGC-1alpha, a key modulator of p53, promotes cell survival upon metabolic stress. Mol Cell 44:621–634
Shaw T, Martin P (2009) Epigenetic reprogramming during wound healing: loss of polycomb-mediated silencing may enable upregulation of repair genes. EMBO Rep 10:881–886
Shilatifard A (2012) The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu Rev Biochem 81:65–95
Simon JA, Kingston RE (2013) Occupying chromatin: polycomb mechanisms for getting to genomic targets, stopping transcriptional traffic, and staying put. Mol Cell 49:808–824
Simone C, Forcales SV, Hill DA, Imbalzano AN, Latella L, Puri PL (2004) p38 pathway targets SWI-SNF chromatin-remodeling complex to muscle-specific loci. Nat Genet 36:738–743
Sinclair KD, Allegrucci C, Singh R, Gardner DS, Sebastian S, Bispham J, Thurston A, Huntley JF, Rees WD, Maloney CA et al (2007) DNA methylation, insulin resistance, and blood pressure in offspring determined by maternal periconceptional B vitamin and methionine status. Proc Natl Acad Sci U S A 104:19351–19356
Sinclair DA, Syrzycka M, Macauley MS, Rastgardani T, Komljenovic I, Vocadlo DJ, Brock HW, Honda BM (2009) Drosophila O-GlcNAc transferase (OGT) is encoded by the Polycomb group (PcG) gene, super sex combs (sxc). Proc Natl Acad Sci U S A 106:13427–13432
Skora AD, Spradling AC (2010) Epigenetic stability increases extensively during Drosophila follicle stem cell differentiation. Proc Natl Acad Sci U S A 107:7389–7394
Smith BC, Denu JM (2009) Chemical mechanisms of histone lysine and arginine modifications. Biochim Biophys Acta 1789:45–57
Smith EH, Janknecht R, Maher LJ 3rd (2007) Succinate inhibition of alpha-ketoglutarate-dependent enzymes in a yeast model of paraganglioma. Hum Mol Genet 16:3136–3148
Smith ER, Lee MG, Winter B, Droz NM, Eissenberg JC, Shiekhattar R, Shilatifard A (2008) Drosophila UTX is a histone H3 Lys27 demethylase that colocalizes with the elongating form of RNA polymerase II. Mol Cell Biol 28:1041–1046
Soloaga A, Thomson S, Wiggin GR, Rampersaud N, Dyson MH, Hazzalin CA, Mahadevan LC, Arthur JS (2003) MSK2 and MSK1 mediate the mitogen- and stress-induced phosphorylation of histone H3 and HMG-14. EMBO J 22:2788–2797
Sparmann A, van Lohuizen M (2006) Polycomb silencers control cell fate, development and cancer. Nat Rev Cancer 6:846–856
Steffen PA, Fonseca JP, Ganger C, Dworschak E, Kockmann T, Beisel C, Ringrose L (2013) Quantitative in vivo analysis of chromatin binding of Polycomb and Trithorax group proteins reveals retention of ASH1 on mitotic chromatin. Nucleic Acids Res 41:5235–5250
Sustar A, Schubiger G (2005) A transient cell cycle shift in Drosophila imaginal disc cells precedes multipotency. Cell 120:383–393
Sustar A, Bonvin M, Schubiger M, Schubiger G (2011) Drosophila twin spot clones reveal cell division dynamics in regenerating imaginal discs. Dev Biol 356:576–587
Tausendschon M, Dehne N, Brune B (2011) Hypoxia causes epigenetic gene regulation in macrophages by attenuating Jumonji histone demethylase activity. Cytokine 53:256–262
Teperino R, Schoonjans K, Auwerx J (2010) Histone methyl transferases and demethylases; can they link metabolism and transcription? Cell Metab 12:321–327
Thompson CB (2009) Metabolic enzymes as oncogenes or tumor suppressors. N Engl J Med 360:813–815
Tian X, Diaz FJ (2013) Acute dietary zinc deficiency before conception compromises oocyte epigenetic programming and disrupts embryonic development. Dev Biol 376:51–61
Tie F, Siebold AP, Harte PJ (2005) The N-terminus of Drosophila ESC mediates its phosphorylation and dimerization. Biochem Biophys Res Commun 332:622–632
Tie F, Banerjee R, Stratton CA, Prasad-Sinha J, Stepanik V, Zlobin A, Diaz MO, Scacheri PC, Harte PJ (2009) CBP-mediated acetylation of histone H3 lysine 27 antagonizes Drosophila Polycomb silencing. Development 136:3131–3141
Tie F, Banerjee R, Conrad PA, Scacheri PC, Harte PJ (2012) Histone demethylase UTX and chromatin remodeler BRM bind directly to CBP and modulate acetylation of histone H3 lysine 27. Mol Cell Biol 32:2323–2334
Tiwari VK, Stadler MB, Wirbelauer C, Paro R, Schubeler D, Beisel C (2012) A chromatin-modifying function of JNK during stem cell differentiation. Nat Genet 44:94–100
Tolhuis B, Muijrers I, de Wit E, Teunissen H, Talhout W, van Steensel B, van Lohuizen M(2006) Genome-wide profiling of PRC1 and PRC2 Polycomb chromatin binding in Drosophila melanogaster. Nat Genet 38:694–699
Toyota M, Suzuki H (2010) Epigenetic drivers of genetic alterations. Adv Genet 70:309–323
Vermeulen L, Vanden Berghe W, Beck IM, De Bosscher K, Haegeman G (2009) The versatile role of MSKs in transcriptional regulation. Trends Biochem Sci 34:311–318
Vicent GP, Ballare C, Nacht AS, Clausell J, Subtil-Rodriguez A, Quiles I, Jordan A, Beato M (2006) Induction of progesterone target genes requires activation of Erk and Msk kinases and phosphorylation of histone H3. Mol Cell 24:367–381
Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, Morey L, Van Eynde A, Bernard D, Vanderwinden JM et al (2006) The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439:871–874
Vocadlo DJ (2012) O-GlcNAc processing enzymes: catalytic mechanisms, substrate specificity, and enzyme regulation. Curr Opin Chem Biol 16:488–497
Voncken JW, Schweizer D, Aagaard L, Sattler L, Jantsch MF, van Lohuizen M (1999) Chromatin-association of the Polycomb group protein BMI1 is cell cycle-regulated and correlates with its phosphorylation status. J Cell Sci 112:4627–4639
Voncken JW, Niessen H, Neufeld B, Rennefahrt U, Dahlmans V, Kubben N, Holzer B, Ludwig S, Rapp UR (2005) MAPKAP kinase 3pK phosphorylates and regulates chromatin association of the polycomb group protein Bmi1. J Biol Chem 280:5178–5187
Wainfan E, Poirier LA (1992) Methyl groups in carcinogenesis: effects on DNA methylation and gene expression. Cancer Res 52:2071s–2077s
Walport LJ, Hopkinson RJ, Schofield CJ (2012) Mechanisms of human histone and nucleic acid demethylases. Curr Opin Chem Biol 16:525–534
Wang H, Wang L, Erdjument-Bromage H, Vidal M, Tempst P, Jones RS, Zhang Y (2004) Role of histone H2A ubiquitination in Polycomb silencing. Nature 431:873–878
Waterland RA, Jirtle RL (2003) Transposable elements: targets for early nutritional effects on epigenetic gene regulation. Mol Cell Biol 23:5293–5300
Waterland RA, Dolinoy DC, Lin JR, Smith CA, Shi X, Tahiliani KG (2006) Maternal methyl supplements increase offspring DNA methylation at Axin Fused. Genesis 44:401–406
Wei Y, Chen YH, Li LY, Lang J, Yeh SP, Shi B, Yang CC, Yang JY, Lin CY, Lai CC et al (2011) CDK1-dependent phosphorylation of EZH2 suppresses methylation of H3K27 and promotes osteogenic differentiation of human mesenchymal stem cells. Nat Cell Biol 13:87–94
Wu SC, Zhang Y (2011) Cyclin-dependent kinase 1 (CDK1)-mediated phosphorylation of enhancer of zeste 2 (Ezh2) regulates its stability. J Biol Chem 286:28511–28519
Wu HA, Balsbaugh JL, Chandler H, Georgilis A, Zullow H, Shabanowitz J, Hunt DF, Gil J, Peters G, Bernstein E (2013) MAP kinase signaling mediates phosphorylation of Polycomb ortholog Cbx7. J Biol Chem 288:36398–36408
Yang X, Zhang F, Kudlow JE (2002) Recruitment of O-GlcNAc transferase to promoters by corepressor mSin3A: coupling protein O-GlcNAcylation to transcriptional repression. Cell 110:69–80
Yang SH, Sharrocks AD, Whitmarsh AJ (2013) MAP kinase signalling cascades and transcriptional regulation. Gene 513:1–13
Yin R, Mao SQ, Zhao B, Chong Z, Yang Y, Zhao C, Zhang D, Huang H, Gao J, Li Z et al (2013) Ascorbic acid enhances Tet-mediated 5-methylcytosine oxidation and promotes DNA demethylation in mammals. J Am Chem Soc 135:10396–10403
Zhang SM, Willett WC, Selhub J, Hunter DJ, Giovannucci EL, Holmes MD, Colditz GA, Hankinson SE (2003) Plasma folate, vitamin B6, vitamin B12, homocysteine, and risk of breast cancer. J Natl Cancer Inst 95:373-380
Zhang W, Deng H, Bao X, Lerach S, Girton J, Johansen J, Johansen KM (2006) The JIL-1 histone H3S10 kinase regulates dimethyl H3K9 modifications and heterochromatic spreading in Drosophila. Development 133:229–235
Zhang H, Wang Z, Zhang Z (2013) PP1alpha, PP1beta and Wip-1 regulate H4S47 phosphorylation and deposition of histone H3 variant H3.3. Nucleic Acids Res 41:8085–8093
Zhao R, Nakamura T, Fu Y, Lazar Z, Spector DL (2011) Gene bookmarking accelerates the kinetics of post-mitotic transcriptional re-activation. Nat Cell Biol 13:1295–1304
Zhong S, Jansen C, She QB, Goto H, Inagaki M, Bode AM, Ma WY, Dong Z (2001) Ultraviolet B-induced phosphorylation of histone H3 at serine 28 is mediated by MSK1. J Biol Chem 276:33213–33219
Zhou X, Sun H, Chen H, Zavadil J, Kluz T, Arita A, Costa M (2010) Hypoxia induces trimethylated H3 lysine 4 by inhibition of JARID1A demethylase. Cancer Res 70:4214–4221
Ziech D, Franco R, Pappa A, Panayiotidis MI (2011) Reactive oxygen species (ROS)–induced genetic and epigenetic alterations in human carcinogenesis. Mutat Res 711:167–173
Zink B, Paro R (1989) In vivo binding pattern of a trans-regulator of homoeotic genes in Drosophila melanogaster. Nature 337:468–471
Zippo A, De Robertis A, Serafini R, Oliviero S (2007) PIM1-dependent phosphorylation of histone H3 at serine 10 is required for MYC-dependent transcriptional activation and oncogenic transformation. Nat Cell Biol 9:932–944
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kolybaba, A., Classen, AK. Sensing cellular states—signaling to chromatin pathways targeting Polycomb and Trithorax group function. Cell Tissue Res 356, 477–493 (2014). https://doi.org/10.1007/s00441-014-1824-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-014-1824-x