Abstract
Rat quiescin/sulphydryl oxidase (rQSOX) introduces disulphide bridges into peptides and proteins with the reduction of molecular oxygen to hydrogen peroxide. Its occurrence has been previously highlighted in a wide range of organs by reverse transcription-polymerase chain reaction (RT-PCR) and Northern blot analyses, methods that have provided information concerning its expression in whole organs but that do not reveal the cell types expressing this enzyme. In this report, in addition to RT-PCR and Western blot experiments, the cell-specific localization of rQSOX has been investigated in a wide range of male and female adult rat tissues by using in situ hybridization and immunohistochemistry. Labelling was detected in most organs and systems including the immune, endocrine and reproductive systems, the respiratory, digestive and urinary tracts and the skin. No labelling was observed in the heart, blood vessel endothelium, liver or smooth and skeletal muscles. rQSOX expression was mainly localized in epithelial cells specialized in secretion, strengthening the hypothesis that QSOX enzymes play an important role in the mechanism of secretion, notably in the folding of secreted proteins. The intracellular patterns of immunolabelling indicate that the protein usually follows the secretory pathway, which is in accordance with its secreted nature and its presumed involvement in the elaboration of the extracellular matrix. In seminiferous tubules, where a high level of expression was noticed, QSOX might play an important physiological role in sperm function and serve as a marker for the diagnosis of male infertility.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sulphydryl oxidases are enzymes that catalyze the formation of disulphide bonds in peptides and proteins with the reduction of molecular oxygen to hydrogen peroxide (Chang and Morton 1975; Ostrowski et al. 1979). Two families of flavin adenine dinucleotide (FAD)-linked sulphydryl oxidases have been described so far: the QSOX (quiescin sulphydryl oxidase) family (Hoober et al. 1999; Thorpe et al. 2002) and the ERV/ALR (essential for respiration and vegetative growth/augmenter of liver regeneration) family (Lee et al. 2000; Lisowsky et al. 2001). QSOX members share strong cDNA sequence identities and common amino-acidic sequence features (Hoober et al. 1999; Thorpe et al. 2002). They display an N-terminal thioredoxin PDI (protein disulphide isomerase)-like domain and a C-terminal ERV1-like domain, which contains the FAD-binding site (Coppock et al. 1998). The QSOX family includes the human quiescin Q6 from lung fibroblasts (Coppock et al. 1993) and sulphydryl oxidases from chicken egg white (Hoober et al. 1999), rat seminal vesicles (Ostrowski and Kistler 1980; Benayoun et al. 2001), guinea-pig endometrial cells (Musard et al. 2001) and mouse epidermis (Matsuba et al. 2002). Quiescin/sulphydryl oxidase (QSOX) proteins have been implicated in essential cellular functions such as protein folding (Thorpe et al. 2002), elaboration of the extracellular matrix, regulation of the redox state and control of the cell cycle (Coppock et al. 1993, 2000; Musard et al. 2001). Recently, in human, a paralogue gene to QSOX (originally named Qscn6 and localized on locus 1q24; Thorpe et al. 2002) has been identified on locus 9q34 in neuroblastoma cells and has been named SOXN or Qscn6LI (Wittke et al. 2003). The SOXN protein is highly homologous to members of the QSOX family and has been linked to apoptosis signalling (Wittke et al. 2003).
Rat QSOX protein (rQSOX) was first identified in the reproductive tract (Chang and Morton 1975) and later sequenced from seminal vesicles (Benayoun et al. 2001). It is a 64-kDa glycoprotein and contains 570 amino acids, including a signal sequence of 32 amino acids. High levels of rQSOX expression have been detected by reverse transcription-polymerase chain reaction (RT-PCR) and Northern blot in seminal vesicles and epididymis, with lower levels in a broad range of other peripheral organs (Benayoun et al. 2001; Mairet-Coello et al. 2002). Rat QSOX is also widely distributed throughout the brain. Two transcripts of 2.8 kb and 3.6 kb, arising from alternative splicing, have been found in the brain, whereas only the 2.8-kb mRNA form has been detected in peripheral organs (Mairet-Coello et al. 2002). The mapping of the protein in adult rat brain by immunohistochemistry has shown that it is specifically expressed by neurons and differentially distributed throughout the central nervous system (Mairet-Coello et al. 2004). In particular, high rQSOX levels have been noted in neuron populations known to secrete disulphide-bond-containing peptides such as the magnocellular neurons of the hypothalamus, which produce oxytocin and vasopressin. It is also expressed in several cell populations of the adenohypophysis but mainly in a subset of gonadotrophs (Tury et al. 2004). In the present paper, we have extended our histological observations concerning the cellular localization of rQSOX to peripheral organs by immunohistochemistry (IHC) and in situ hybridization (ISH). The analysis of the distribution of QSOX enzymes in normal tissues represents an essential step towards the elucidation of the physiological substrates and functions of these highly conserved and widely distributed proteins.
Materials and methods
Animals and tissue preparation
Sprague–Dawley rats (IFFA Credo, L'Arbresle, France) were housed in a temperature-controlled room under a natural light-dark cycle and fed with standard laboratory chow and water ad libitum. Animal manipulations and experimental procedures were performed in accordance with the recommendations of our institution and under the supervision of authorized investigators.
For RT-PCR investigations, one male and one female adult rat were killed by decapitation. Peripheral organs were removed, rapidly frozen over liquid nitrogen and stored at −80°C until RNA extraction. Another adult male rat was used for protein extractions and Western blot experiments.
For ISH and IHC investigations, two male and two female adult rats were deeply anaesthetized with 7% chloral hydrate solution (1 ml/200 g body weight) and then perfused transcardially with 400 ml 0.9% NaCl followed by 400 ml fixative, viz. ice-cold 1% paraformaldehyde (PFA) solution in 0.1 M phosphate buffer (PB), pH 7.2. Organs were removed, postfixed in the same fixative for 2 h at 4°C, immersed overnight in a 15% cryoprotective sucrose solution in 0.1 M PB at 4°C, embedded in a commercial medium (Cryomatrix, Shandon, Pittsburgh, USA), deep-frozen over liquid nitrogen and serially cut into 10-μm-thick sections with a cryostat-microtome. Sections were mounted on gelatinized slides and stored at −45°C until treatment.
RNA extraction
Total RNA was extracted from the various tissues by using the RNA NOW extraction kit (Ozyme, Saint Quentin en Yvelines, France) as previously described (Mairet-Coello et al. 2002). RNA concentrations were determined by absorbance at 260 nm.
RT-PCR technique
Denatured total RNA (1 μg) was reverse-transcribed into cDNA as previously described (Tury et al. 2004). PCR amplifications were carried out in a PTC-200 thermocycler (MJ-Research, Fontenay Sous Bois, France) with 1 μl cDNA and the FastStart Taq DNA polymerase kit (Roche, Meylan, France) in a 25-μl final reaction mixture containing 2.5 μl 10× PCR buffer, 3 mM MgCl2, 0.4 mM mixture of the four deoxyribonucleotides (Roche) and 1 U Taq DNA polymerase. PCR was performed to amplify a 797-bp fragment of rQSOX cDNA (base 17 to base 813 of the 2.8-kb rQSOX mRNA, GenBank accession no. NM_053431; base (−3) to base 793 of the 3.6 kb rQSOX mRNA, GenBank accession no. AY623665) with the rQSOX sense primer 5′-ACTTGAGCGAGGTGGACAGTCAAG-3′ and the rQSOX antisense primer 5′-AGCACAGGCACTCGGGAA-3′ (Eurogentec, Seraing, Belgium; Benayoun et al. 2001; Mairet-Coello et al. 2002). A 509-bp fragment of cyclophilin cDNA (CYC, GenBank accession no. M19533) was used as an internal standard and was amplified with specific CYC primers (sense primer: 5′-CGCCGCTTGCTGCAGACATGG-3′and reverse primer: 5′-GAGTTGTCCACAGTCGGAGATGG-3′, Eurogentec). After one step of PCR enzyme activation at 96°C for 5 min, reactions were performed in a PTC-200 thermocycler for 30 cycles of 30 s at 96°C, 30 s at 60°C and 90 s at 72°C, followed by one final extension step at 72°C for 10 min. Aliquots of 20 μl of each PCR product were separated by electrophoresis on a 1% agarose gel containing ethidium bromide and were analysed by using a Gel Doc 2000 (Bio-Rad, Marnes-la-Coquette, France) and Bio-Rad Quantity One 4.2.3 software.
Cloning of rQSOX cDNA
The 797-bp rQSOX cDNA obtained by RT-PCR from brain (Mairet-Coello et al. 2002), seminal vesicles, pancreas and small intestine were purified from agarose gel by using the QIAquick gel extraction kit (QIAGEN, Courtaboeuf, France) according to the manufacturer's instructions. The purified fragments were then cloned into the pGEM-T easy vector (Promega, Charbonnières, France) as recommended by the manufacturer. The four cDNA inserts were sequenced on an automated Applied Biosystem Model 373A sequencing system (Courtaboeuf, France).
QSOX riboprobe synthesis
The plasmid containing the 797-bp rQSOX insert obtained from brain was linearized with NcoI restriction enzyme (Roche). After phenol/chloroform purification, the linearized plasmid was used for in vitro transcription with the DIG (digoxigenin) RNA labelling kit (Roche). Transcription was performed for 2 h at 37°C according to the manufacturer's instructions. As a control, an unlabelled riboprobe was also obtained by replacing the DIG-labelled NTP mixture with a mix of 10 mM each unlabelled NTP. Plasmid containing the template cDNA was then digested with 20 U RNase-free DNase I for 15 min at 37°C and the reaction was stopped by adding 2 μl 0.2 M EDTA pH 8. The riboprobe was subsequently purified through a precipitation step by addition of 2.5 μl 4 M lithium chloride and 75 μl 100% ethanol (2 h at −20°C) followed by centrifugation for 15 min a 4°C. After a washing with 100 μl 70% ethanol during centrifugation for 15 min at 4°C, the pellet was resuspended in 50 μl diethyl-pyrocarbonate-treated water (DEPC, Sigma, Saint Quentin Fallavier, France). Riboprobe concentration was determined by absorbance at 260 nm. The length and integrity of the riboprobe were monitored by gel electrophoresis.
ISH protocol
The technique was adapted from the simplified ISH protocol developed by Braissant and Wahli (1998) with DIG-labelled riboprobes to detect abundant and rare mRNA on tissue sections.
All steps prior to and during hybridization were conducted under RNase-free conditions. After a post-fixation in 4% PFA prepared in DEPC-treated phosphate-buffered saline (PBS), pH 7.5, sections were incubated twice for 15 min in PBS containing 0.1% active DEPC and equilibrated for 15 min in DEPC-treated 5× standard saline citrate buffer (SSC). Sections were then prehybridized for 2 h at 58°C in the hybridization solution (50% formamide, 50% DEPC-treated 5× SSC, 200 μg/ml salmon sperm DNA; 150 μl on each slide). The probe was added to the hybridization solution (5.3 ng probe per microliter hybridization solution) and denatured for 5 min at 80°C prior use. The hybridization reaction was carried out at 58°C for 40 h with 75 μl of this probe solution (corresponding to 400 ng riboprobe) on each slide. To avoid evaporation during the prehybridization and hybridization steps, slides were placed in a box saturated with 5× SSC and sections were covered with laboratory film (Parafilm) and sealed to the slide with rubbercement glue (Royal Talens). After incubation, sections were washed for 30 min in 2× SSC at room temperature, incubated 30 min at 37°C in RNase A solution (20 μg/ml) in 2× SSC, washed twice in 2× SSC for 5 min at room temperature, for 1 h in 2× SSC at 58°C and for 1 h in 0.1× SSC at 58°C. After equilibration for 5 min at room temperature in buffer 1 (100 mM TRIS-HCl, 150 mM NaCl, pH 7.5), the sections were incubated for 2 h at room temperature with alkaline-phosphatase-conjugated DIG antibody (Roche) diluted 1/2500 in buffer 1 containing 0.5% blocking reagent (Boehringer Mannheim-Roche). They were then washed twice for 15 min in buffer 1 and equilibrated for 5 min in buffer 2 (100 mM TRIS-HCl, 100 mM NaCl, 50 mM MgCl2, pH 9.5). Hybridization was revealed, at room temperature, with 4.5 μl nitro-blue tetrazolium chloride and 3.5 μl 5-bromo-4-chloro-3-indolylphosphate p-toluidine salt (Roche) diluted in 1 ml buffer 2. The enzymatic reaction was conducted for up to one night. Staining was stopped in 10 mM TRIS and 1 mM EDTA, pH 8, for 10 min. The precipitated TRIS was removed in a bath of distilled water. Negative controls included (1) the omission of the antisense rQSOX riboprobe, (2) the omission of the DIG antibody and (3) hybridization with a mix of labelled and unlabelled rQSOX antisense riboprobes (ratio 1:10).
Protein extraction and Western blot
Proteins were extracted from various tissues according to the method recommended by the supplier of the TriReagent kit (Euromedex, Souffelweyersheim, France). The rQSOX antiserum raised against the purified QSOX protein from rat seminal vesicles was prepared and checked at INSERM U618 (Protéases et Vectorisation Pulmonaires), Tours (Benayoun et al. 2001). Samples containing 25 μg total proteins extracted from rat tissues were separated on 12% SDS-polyacrylamide gels without reducing agent in the sample buffer and transferred to a nitrocellulose membrane. The membrane was incubated with the rQSOX antiserum diluted 1:600, followed by incubation with goat peroxidase conjugated anti-rabbit IgG (DAKO, Trappes, France). Signals were visualized by using the RPN 2106 chemiluminescence detection system (Amersham Pharmacia Biotech, Orsay, France).
IHC protocol
After being rinsed in 0.1 M PBS containing 0.3% Triton X-100 (PBS-T), cryostat sections were submitted to an indirect immunofluorescence protocol. They were incubated overnight with the rQSOX antiserum diluted 1:500 in PBS-T containing 10% lactoproteins, 1% bovine serum albumin and 0.01% sodium azide. Labelling was revealed by incubation for 1 h in secondary goat anti-rabbit IgG conjugated to Alexa Fluor 488 (Molecular Probes, Interchim, Montluçon, France) diluted 1:400 in the same solution as the primary antibody. A control step was performed for each tissue by incubating the rQSOX antiserum for 4 h with the antigen solution (0.03 nmol rQSOX protein purified from seminal vesicle fluid per microliter of non-diluted antiserum) before the immunolabelling procedure.
A double immunofluorescence protocol was conducted for the simultaneous detection of rQSOX and insulin (DAKO), glucagon (G2654; Sigma), somatostatin (MAB354; Chemicon International, Temecula, California, USA) and mitochondrial cytochrome oxidase subunit I (Molecular probes; see Table 1). Sections were first incubated with the rQSOX antiserum and revealed as described above with goat anti-rabbit IgG conjugated to Alexa Fluor 488 or CY3-conjugated donkey anti-rabbit IgG (Jackon Immunoresearch Laboratories, Interchim). They were then incubated with the second primary antibody, which was revealed with the appropriate secondary antibody conjugated to CY3 or to Alexa Fluor 488. rQSOX and phenylethanolamine N-methyl transferase (PNMT), the final enzyme in the synthetic pathway for adrenalin, were also detected on consecutive sections of adrenal glands. The antiserum against PNMT was kindly provided by Dr. Orsini (Laboratoire de Neurobiologie, CNRS, Marseille, France); its preparation and specificity have been reported by Kitahama et al. (1985).
Photomicrograph production
Sections were observed under a fluorescence microscope (Olympus BX51). Digital image acquisitions were carried out with a DP50 Olympus camera and AnalySIS 3.1 software (Soft Imaging System) and imported into Adobe Photoshop software for subsequent treatment, minimally altering the captured images.
Results
RT-PCR and Western blot analyses
The expected 797-bp rQSOX cDNA fragment was detected by RT-PCR in all analysed organs. The rQSOX mRNA level varied from one tissue to another (Fig. 1). The highest level of expression of rQSOX was found in seminal vesicles and testis. It was strongly expressed in lung, salivary glands, stomach, small intestine, adrenal glands, kidney, uterus and ovary, with lower expression levels being noted in heart, spleen, large intestine, liver, skeletal muscle and eye. Sequencing the 797-bp cDNA fragment isolated from seminal vesicles, pancreas and small intestine confirmed that it corresponded to the 5′ common part of the two 2.8-kb and 3.6-kb rQSOX mRNAs (Mairet-Coello et al. 2002; bases 17 to 813, GenBank accession no. NM_053431 for the 2.8-kb rQSOX mRNA and no. AY623665 for the 3.6-kb rQSOX mRNA).
RT-PCR detection of QSOX and CYC in rat peripheral organs. RT-PCR were performed on 1 μg total RNA extracted from each sample by using rQSOX and CYC primers. After Agarose gel electrophoresis and ethidium bromide staining, 797-bp (rQSOX) and a 509-bp (CYC) bands were detected in all the analysed tissues (lane 1 1-kb Plus DNA Ladder (Invitrogen), AG adrenal glands, E eye, H heart, K kidney, Li liver, LI long intestine, Lu lung, O ovary, SI small intestine, SG salivary glands, SM skeletal muscle, Sp spleen, St stomach, SV seminal vesicles, T testis, U uterus)
A single 64-kDa band was detected by Western blot with the rQSOX antiserum in homogenates from seminal vesicles, testis, epididymis, lung and pancreas (Fig. 2). The most intense signal was observed in seminal vesicles. No signal was revealed in homogenates from heart, skeletal muscle or liver.
Western blot analysis of QSOX in protein extracts from rat peripheral tissues; 25 μg total protein was separated on 12% SDS-polyacrylamide gel under non-reducing conditions. The nitrocellulose membrane was probed with the rQSOX antiserum and bands were visualized by chemiluminescence. A single band of about 64 kDa was detected in homogenates from the following tissues: E epididymis, Lu lung, P prostate, SV seminal vesicles, T testis. No signal was revealed in extracts from the following: H heart, Li liver, SM skeletal muscle. Lane 1 MagicMark protein molecular weight markers (Invitrogen)
ISH and IHC localization of rQSOX in organs
ISH and IHC controls
No signal was observed on tissue sections when the riboprobe or the DIG antiserum was omitted. Moreover, no or only slight labelling was obtained on tissue sections hybridized with a mix of labelled antisense riboprobe added with an excess of the same unlabelled riboprobe, as shown in seminal vesicles (Fig. 4e,f).
For IHC, preadsorption of the antiserum with rat QSOX protein purified from seminal vesicle fluid before its application on tissue sections completely abolished immunolabelling (Fig. 4g,h).
General observations
ISH and IHC results are summarized in Table 2. The distribution of rQSOX mRNA observed by ISH correlated well with that of the protein observed by IHC. Rat QSOX expression was detected in most systems and tracts, including the immune, endocrine and reproductive systems, the respiratory, digestive and urinary tracts, the skin and the retina. However, neither ISH nor IHC labelling was observed in liver or smooth and skeletal muscles. Heart and blood vessel endothelia were also devoid of rQSOX labelling. A more detailed description of the localization of rQSOX in the various systems and tracts is provided below.
Immune system
In the thymus, ISH staining and reticular cytoplasmic immunolabelling were seen in granulocytes localized around the blood vessels but no labelling was observed in lymphoid cells. In spleen, labelling was also found in granulocytes localized around sinuses of the red pulp (Fig. 3a,b) whereas lymphoid follicles of the white pulp were not or only slightly labelled.
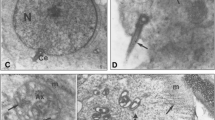
ISH and IHC localization of rQSOX expression in rat peripheral tissue sections. rQSOX ISH staining (a) and IHC labelling (b) of granulocytes cells (arrows) located around sinuses of the red pulp of the spleen. Note the reticular aspect of the immunolabelling in these cells (inset in b). c Immunolabelling of bronchiolar epithelial cells (arrows and inset) in lung. d Intense immunoreactivity bordering the apical side of acinar cells (arrows) of salivary glands. In the oesophagus epithelium, rQSOX mRNA (e) and protein (f) were exclusively localized in the granular layer (f arrows and inset). g In stomach, the most superficial cells of the epithelium (arrows) and neck cells (arrowheads and inset) were labelled for rQSOX. No labelling was observed in gastric glandular cells. In small intestine, ISH staining (h) and intense granular immunolabelling (i) filled the whole cytoplasm (inset in i) of goblet cells (arrows), whereas no labelling was found in enterocytes (arrowheads). j ISH staining in acinar (arrows) and islet of Langherans (arrowheads) cells of the pancreas. k The highest rQSOX immunolabelling was localized at the periphery of the islets of Langherans (arrows) and exhibited a granular aspect filling the whole cytoplasm (inset). Double immunodetection of rQSOX (green) and insulin (red) showed that rQSOX was not or only slightly expressed in β cells. l Double immunofluorescence for rQSOX (green) and glucagon (red) revealed that all the α cells were positive for rQSOX (yellow, arrows). ISH staining (m) and IHC labelling (n) of the most superficial cells of the urinary bladder transitional epithelium (n arrows and inset). o In the epiphysis (Ep), numerous pinealocytes were intensely labelled for rQSOX. Note also in this section the labelling of neurons of the inferior colliculus (IC), a nucleus of the brainstem. In adrenal glands (cx cortex), ISH staining (p) and diffuse IHC labelling (q, see also inset) was exclusively localized in the chromaffin cells of the medulla (m). Immunodetection of QSOX (q) and PNMT (r) on two consecutive sections indicated that rQSOX was the most expressed in the adrenergic cell population of the adrenal medulla (m). Bars 100 μm
Respiratory tract
In trachea, the majority of goblet and ciliated epithelial cells of the epithelium and seromucous glandular cells of the submucosa were strongly immunolabelled for rQSOX. In the various divisions of the lung, from the bronchi to the terminal bronchioles, ciliated epithelial, goblet and Clara cells displayed intense immunolabelling (Fig. 3c). No labelled cells were seen in the most distal subdivisions, i.e. alveolar ducts, alveolar sacs and alveoli.
Digestive tract
Rat QSOX was expressed along the digestive tract from salivary glands to colon. Broadly speaking, it was mainly detected in epithelium and glands, whereas no or only a few labelled cells were observed in the lamina propria, muscularis mucosa, submucosa or smooth muscle. In salivary glands, intense rQSOX immunoreactivity was observed next to the lumen of the acini (Fig. 3d). Weak labelling was also sometimes observed in epithelial cells of excretory ducts. In the oesophagus, epithelial cells of the granular layer were strongly labelled for rQSOX both by ISH and IHC (Fig. 3e,f) but no positive cells were observed in any other layers of the epithelium. In the stomach, the most superficial cells of the epithelium lay close to the gastric cavity. The mucous neck cells localized in the back of the gastric crypts were labelled for rQSOX (Fig. 3g), whereas gastric glandular cells were unlabelled. In the small intestine, i.e. duodenum, jejunum and ileum, ISH staining and a granular and diffuse cytoplasmic IHC labelling were found in the majority of goblet cells localized in villi and glandular epithelium but absorptive cells were virtually devoid of labelling (Fig. 3h,i). Brunner's gland cells of the duodenum submucosa were intensely labelled, whereas glandular cells of the jejunum were barely labelled. In the ileum, chromaffin Paneth cells localized in the back of folds were also rQSOX-positive. rQSOX labelling was also seen in mastocytes and polynuclear cells localized in the lamina propria of the mucosa and in some endocrine glandular cells. Surprisingly, a few labelled goblet cells were noted in the large intestine. In the pancreas, ISH staining was found in both the exocrine and endocrine portions, the most intense staining being observed in acinar cells. Granular immunolabelling was also seen in the two portions of the pancreas but the strongest labelling was observed in specific cells located at the periphery of islets of Langerhans (Fig. 3j,k). Double IHC showed that they corresponded to the subset of glucagon-secreting α cells (Fig. 3l). No or very weak granular rQSOX labelling was seen in the insulin-secreting β (Fig. 3k) and somatostatin-secreting δ (not shown) cells.
Urinary tract
Few elements of the kidney were labelled for rQSOX. Some positive cells were observed in the epithelium of renal tubules but not in glomeruli. The most superficial cells of the transitional epithelium of the urinary bladder were intensely labelled for rQSOX (Fig. 3m,n); however the immunoreaction was not continuous along the wall.
Endocrine system
Numerous cells of the anterior lobe and pars intermedia of pituitary were immunolabelled for rQSOX, as were fibres and probably pituicytes of the posterior lobe (not shown). In the epiphysis, numerous pinealocytes were also intensely labelled for rQSOX by IHC (Fig. 3o). In the adrenal glands, ISH staining and intense IHC labelling filling the whole cytoplasm were seen in catecholamine-secreting cells of the medulla (Fig. 3p,q). Consecutive sections treated with rQSOX and PNMT antisera strongly suggested that the most intense labelling was localized in adrenergic cells (Fig. 3q,r). No labelling was found in any zones of the steroid hormone-secreting cortical region.
Reproductive system
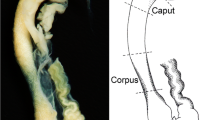
In seminiferous tubules of testis, ISH staining was found in germinal cells of all stages (Fig. 4a) but immunolabelling appeared in a stage-dependent manner (Fig. 4b). Intense crescent-shaped cytoplasmic labelling concentrated at one pole of the nucleus (Fig. 4c) was observed at various stages of spermatocytes and spermatids, whereas spermatogonia and spermatozoids were not labelled. Immunoreaction was also detected in interstitial Leydig cells but not in Sertoli cells. Double IHC with an antibody against mitochondrial cytochrome oxidase subunit I (Fig. 4d) showed that rQSOX protein was not localised in the mitochondria of these cells. Epithelial cells of seminal vesicles strongly expressed rQSOX, as shown by both ISH and IHC (Fig. 4e,g); the immunoreactivity was punctiform and concentrated at the apical side. In the epididymis, epithelial cells displayed strong reticular immunolabelling (Fig. 4i). The ovary exhibited no labelling in either the oocytes, follicle cells or corpus luteum (not shown). In the uterus, luminal and glandular epithelial cells of the endometrium were stained by ISH (Fig. 4j). IHC observations confirmed this localization, the immunolabelling accumulating at the apical side of the cells (Fig. 4k). The most superficial cells of the stratified squamous epithelium of the vagina were strongly labelled (Fig. 4l).
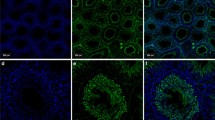
Cellular localization of rQSOX expression observed by ISH and IHC on rat peripheral tissue sections (continued). ISH (a) and immunohistochemical detection (b) of rQSOX in seminiferous tubules of testis (arrows). c In round spermatids, an intense juxtanuclear crescent-shaped labelling (green, arrows) surrounded one pole of the nucleus (coloured red by ethidium bromide). d Double immunohistochemistry for rQSOX (red) and cytochrome oxidase subunit I (green) showing that rQSOX is not localised in the mitochondria of these cells (arrows). e ISH staining of seminal vesicle epithelial cells (arrows). f No labelling was obtained on the consecutive section hybridized with a mix of labelled antisense riboprobe with an excess of the same but unlabelled riboprobe. g Immunoreactivity is concentrated at the apical side of the epithelial cells of the seminal vesicle (arrows and inset). h Preincubation of rQSOX antiserum with the antigen completely abolishes the labelling on the consecutive section. i Epithelial cells of the epididymis (arrows) displaying strong reticular immunolabelling (inset). ISH (j) and IHC (k) labelling in luminal (arrows) and glandular (arrowheads) epithelial cells of the endometrium. A higher magnification of a gland shows that the immunolabelling accumulates at the secretory apical side of the cells (inset in k). l The most superficial cells of the vagina epithelium (arrows and inset) were strongly labelled. m In the skin, labelling was observed in the granular layer of epidermis (arrows). Immunoreaction also affected sebaceous glands and hair follicles (arrowheads) dispersed in dermis. In retina, ISH (n) and IHC (o) labelling were observed in the photoreceptor, inner nuclear and ganglion layers (GCL ganglion layer, INL inner nuclear layer, IPL inner plexiform layer, OPL outer plexiform layer, Ph photoreceptor layer). Bars 100 μm (a, b, e–o), 20 μm (c, d)
Skin
rQSOX labelling was observed in the granular layer of epidermis but the other layers were unlabelled. Intense immunoreaction was also associated with the sebaceous glands and hair follicles in the dermis (Fig. 4m).
Retina
Among the various layers of the retina, three were labelled for rQSOX by ISH and IHC (Fig. 4n,o). Weak immunolabelling was localised in the photoreceptor layer mainly in the rods. An intense filiform labelling was observed through the inner nuclear layer and the majority of cells of the ganglion layer were also strongly labelled.
Discussion
In the present paper, we report rQSOX expression in a large panel of tissues belonging to various systems and tracts including the respiratory, digestive and urinary tracts, the endocrine, immune and reproductive systems and the skin. On the whole, our histological observations are in accordance with our biomolecular results and with previous studies (Coppock et al. 2000; Benayoun et al. 2001; Musard et al. 2001; Mairet-Coello et al. 2002; Thorpe et al. 2002). They confirm the occurrence of QSOX in numerous peripheral organs and emphasize its strong level of expression in secretory glands and the male reproductive system.
Recently, we have shown that the rat QSOX gene, which is located on locus 13q21, is widely expressed throughout the brain with two transcripts (2.8 kb and 3.6 kb) arising from alternative splicing (Mairet-Coello et al. 2002). The 3.6-kb rQSOX mRNA has not been revealed by Northern blot in peripheral organs (Benayoun et al. 2001; Mairet-Coello et al. 2002). The paralogue gene of rat QSOX, corresponding to the human SOXN (Wittke et al. 2003), is predicted to be localized on locus 3p13 (Genbank GeneID 296586). The cDNA transcribed from this paralogue gene (Genbank accession no. XM_231083.1) displays no similarities with the rQSOX cDNA, indicating that only the 2.8-kb rQSOX mRNA, which encodes the 64-kDa rQSOX protein, is probably detected by ISH. The deduced protein translated from the paralogue cDNA has an estimated molecular weight of 87 kDa and displays 41% of identity with the rQSOX protein from rat seminal vesicles. Although both proteins contain some regions of identity, Western blot analyses performed with our polyclonal antibody on protein extracts from rat brains (Mairet-Coello et al. 2004) and several peripheral organs have revealed a unique protein of 64 kDa that corresponds to the QSOX protein from rat seminal vesicles. This result, together with protein liquid-phase adsorption of the antiserum performed by IHC, suggests that the same protein is detected with both techniques. In addition, the mRNA distribution observed by ISH and that of the protein observed by IHC are well correlated in all tissues. One small discrepancy between RT-PCR and the other methods of analysis (Western blot, ISH and IHC) concerns the heart, liver and skeletal muscle; this is probably attributable to the differences of sensitivity between the techniques.
Distinct patterns of immunolabelling were noticed depending on the cell type, giving indications regarding the intracellular localization of the protein. The rQSOX labelling was exclusively present in the cytoplasm of the cells. In several cell types, such as goblet cells, acinar cells or epithelial cells of seminal vesicles and endometrium, the immunolabelling was granular, filling the whole cytoplasm or being restricted to the apical pole of the cells. This suggests a localization of the protein in secretory granules as previously observed in cells of the adenohypohysis (Tury et al. 2004). In the granulocytes of the thymus and spleen and in the epithelial cells of the epididymis, labelling displays a reticular appearance reminiscent of that observed in neurons (Mairet-Coello et al. 2004), strongly suggesting a localization of the enzyme in the Golgi apparatus. The perinuclear crescent-shaped labelling observed in seminiferous tubules is in accordance with a Golgi localization but could also correspond to a labelling of the acrosome. Double IHC experiments with an antibody specific to mitochondria has excluded a mitochondrial localization of rQSOX, an observation in disagreement with the initial findings of Aumüller and co-workers who have identified a sulphydryl oxidase of 65 kDa probably corresponding to QSOX, in mitochondria of rat, hamster and human germinal cells (Kumari et al. 1990; Aumüller et al. 1991). A thorough electron-microscopical study at various stages of spermatogenesis should help to clarify this discrepancy.
Proteins of the QSOX family display a signal peptide sequence and have been detected extracellularly. They are notably found in seminal vesicle secretions (Ostrowski and Kistler 1980; Benayoun et al. 2001) and tissue culture medium (Coppock et al. 2000; Musard et al. 2001). These data and our present work suggest that QSOX proteins follow the secretory pathway in many cell types and may participate in the elaboration of the extracellular matrix or intervene in extracellular redox processes as previously proposed (Coppock et al. 1998, 2000; Hoober et al. 1999; Thorpe et al. 2002). Sulphydryl oxidases of the ERV/ALR family display different intracellular localizations. Mammalian ALR protein and its orthologue yeast ERV1 are localized in the mitochondrial intermembrane space where they play a role in the assembly of cytosolic iron/sulphur proteins and cellular iron homeostasis (Lange et al. 2001). ERV2, the yeast paralogue of ERV1, is a resident protein of the endoplasmic reticulum and is involved in an oxidative folding pathway (Gerber et al. 2001).
Human QSOX paralogue SOXN (Wittke et al. 2003) and rat ALR (Hagiya et al. 1994; Klissenbauer et al. 2002) were detected in several peripheral organs and in the central nervous system by using biomolecular techniques but few histological studies have described their precise localization in tissues (Gandhi et al. 1999; Klissenbauer et al. 2002). To date, only the distribution of the bovine milk metallosulphydryl oxidase had been thoroughly reported in rat peripheral organs by using IHC (Clare et al. 1984). This enzyme is localized in the endothelial cells of the capillaries of kidney, heart and small intestine and in centroacinar cells of the pancreas but has not been found in brain and thymus. Our results show that rat QSOX is distributed in a large variety of peripheral tissues in which it is mainly localized in epithelial cells specialized in secretion. In the central nervous system, the enzyme is ubiquitously distributed and specifically expressed by neurons (Mairet-Coello et al. 2004). Thus, the metallosulphydryl oxidase and rQSOX display distinct patterns of expression, probably because they have different specific substrates and/or functions.
Although the precise roles of sulphydryl oxidases have not as yet been elucidated, their cellular distribution provides clues regarding their potential substrates and physiological functions. As proposed by Thorpe et al. (2002), the most obvious substrates for QSOX enzymes are secreted proteins that possess one or several disulphide bridges. Thus, mucins, the major constituents of mucus, contain disulphide bonds (Perez-Vilar and Hill 1999) and are secreted by goblet cells, salivary and Brunner's glands. They could therefore represent good substrates for the enzyme. Seminal vesicle epithelium also secretes disulphide-bonded proteins in seminal fluid (Luo et al. 2001; Wagner and Kistler 1987). In the adenohypophysis, although most of the secreted hormones contain disulphide bridges, rQSOX is strongly expressed in a subset of gonadotrophs that synthesize luteinizing hormone and/or follicle-stimulating hormone (Tury et al. 2004). In keratinized stratified epithelia that cover the skin and rat oesophagus, rQSOX is specifically localized in the granular layer, supporting the hypothesis that it plays an important biological role in the structural integrity of the upper epidermis by catalysing the introduction of disulphide bonds into precursor proteins such as loricrin and involucrin (Hashimoto et al. 2000, 2001; Matsuba et al. 2002). Interestingly, in seminiferous tubules, rQSOX expression appears in a stage-dependent manner, no labelling being observed in the early stages of spermatogenesis or in mature spermatozoids, suggesting that rQSOX is of transient importance in the maturation of germinal cells. Redox regulation has been shown to be crucial in the physiology of normal sperm function and a dysregulation of this process has been linked to spermatogenetic abnormalities (Aitken and Baker 2004; Baker and Aitken 2004). It is noteworthy that another redox protein, the thioredoxin SPTRX-3, displays an expression pattern resembling that of rQSOX in seminiferous tubules (Jimenez et al. 2004). Thioredoxins are potential substrates of sulphydryl oxidases (Thorpe et al. 2002) and, given the possible colocalization of these proteins, they could be involved in a common function in germinal cells, such as acrosomal biogenesis or post-translational modifications of acrosomal enzymes in the Golgi apparatus (Jimenez et al. 2004). In the endocrine pancreas, the highest rQSOX expression occurs in cells secreting glucagon, which does not contain a disulphide bond, whereas the enzyme is not or only faintly expressed in cells secreting insulin or somatostatin, both of which display disulphide bonds. This observation suggests that insulin and somatostatin are not preferential substrates for rQSOX and that a specific substrate for the enzyme is present in α cells of the islets of Langherans. Stanniocalcin 2, an hormone containing 15 cystein residues allowing the formation of a disulphide-bridged homodimer (Conlon 2000), is reported to be specifically expressed in α cells of the pancreas (Moore et al. 1999) and might thus be a target for QSOX. In adrenal glands, rQSOX is exclusively localized in catecholaminergic cells, more precisely in the PNMT-expressing adrenergic cell population of the medulla, indicating that it might play a specific role in the metabolism of adrenalin.
In conclusion, through the formation and/or maintenance of disulphide bonds in secreted proteins, QSOX enzymes are probably involved in the secretory mechanism in a large variety of tissues. They could also be co-secreted with their substrates in order to continue their oxidative folding activity outside the cell. In addition, they may participate extracellularly in different processes such as the elaboration of the extracellular matrix. In testis, QSOX could fulfil an important physiological role in sperm function and has previously been proposed to serve as a marker for spermatogenic efficiency and the diagnosis of male infertility (Bergmann et al. 1992).
References
Aitken RJ, Baker MA (2004) Oxidative stress and male reproductive biology. Reprod Fertil Dev 16:581–588
Aumüller G, Bergmann M, Seitz J (1991) Immunohistochemical distribution of sulfhydryl oxidase in the human testis. Cell Tissue Res 266:23–28
Baker MA, Aitken RJ (2004) The importance of redox regulated pathways in sperm cell biology. Mol Cell Endocrinol 216:47–54
Benayoun B, Esnard-Feve A, Castella S, Courty Y, Esnard F (2001) Rat seminal vesicle FAD-dependent sulfhydryl oxidase. Biochemical characterization and molecular cloning of a member of the new sulfhydryl oxidase/quiescin Q6 gene family. J Biol Chem 276:13830–13837
Bergmann M, Aumüller G, Seitz J, Nieschlag E (1992) Sulfhydryl oxidase immunoreactivity in seminiferous tubules of infertile men. Cell Tissue Res 267:209–214
Braissant O, Wahli W (1998) A simplified in situ hybridization protocol using non-radioactively labeled probes to detect abundant and rare mRNAs on tissue sections. Biochemica 1:10–16
Chang TS, Morton B (1975) Epididymal sulfhydryl oxidase: a sperm-protective enzyme from the male reproductive tract. Biochem Biophys Res Commun 66:309–315
Clare DA, Horton HR, Stabel TJ, Swaisgood HE, Lecce JG (1984) Tissue distribution of mammalian sulfhydryl oxidase. Arch Biochem Biophys 230:138–145
Conlon JM (2000) Singular contributions of fish neuroendocrinology to mammalian regulatory peptide research. Regul Pept 93:3–12
Coppock DL, Kopman C, Scandalis S, Gilleran S (1993) Preferential gene expression in quiescent human lung fibroblasts. Cell Growth Differ 4:483–493
Coppock DL, Cina-Poppe D, Gilleran S (1998) The quiescin Q6 gene (QSCN6) is a fusion of two ancient gene families: thioredoxin and ERV1. Genomics 54:460–468
Coppock D, Kopman C, Gudas J, Cina-Poppe DA (2000) Regulation of the quiescence-induced genes: quiescin Q6, decorin, and ribosomal protein S29. Biochem Biophys Res Commun 269:604–610
Gandhi CR, Kuddus R, Subbotin VM, Prelich J, Murase N, Rao AS, Nalesnik MA, Watkins SC, DeLeo A, Trucco M, Starzl TE (1999) A fresh look at augmenter of liver regeneration in rats. Hepatology 29:1435–1445
Gerber J, Muhlenhoff U, Hofhaus G, Lill R, Lisowsky T (2001) Yeast ERV2p is the first microsomal FAD-linked sulfhydryl oxidase of the Erv1p/Alrp protein family. J Biol Chem 276:23486–23491
Hagiya M, Francavilla A, Polimeno L, Ihara I, Sakai H, Seki T, Shimonishi M, Porter KA, Starzl TE(1994) Cloning and sequence analysis of the rat augmenter of liver regeneration (ALR) gene: expression of biologically active recombinant ALR and demonstration of tissue distribution. Proc Natl Acad Sci U S A 91:8142—8146 (Erratum in Proc Natl Acad Sci U S A 92:3076)
Hashimoto Y, Suga Y, Matsuba S, Mizoguchi M, Takamori K, Seitz J, Ogawa H (2000) Immunohistochemical localization of sulfhydryl oxidase correlates with disulfide crosslinking in the upper epidermis of rat skin. Arch Dermatol Res 292:570–572
Hashimoto Y, Suga Y, Matsuba S, Mizoguchi M, Takamori K, Seitz J, Ogawa H (2001) Inquiry into the role of skin sulfhydryl oxidase in epidermal disulfide bond formation: implications of the localization and regulation of skin SOx as revealed by TPA, retinoic acid, and UVB radiation. J Invest Dermatol 117:752–754
Hoober KL, Glynn NM, Burnside J, Coppock DL, Thorpe C (1999) Homology between egg white sulfhydryl oxidase and quiescin Q6 defines a new class of flavin-linked sulfhydryl oxidases. J Biol Chem 274:31759–31762
Jimenez A, Zu W, Rawe VY, Pelto-Huikko M, Flickinger CJ, Sutovsky P, Gustafsson JA, Oko R, Miranda-Vizuete A (2004) Spermatocyte/spermatid-specific thioredoxin−3, a novelGolgi apparatus-associated thioredoxin, is a specific marker of aberrant spermatogenesis. J Biol Chem 279:34971–34982
Kitahama K, Pearson J, Denoroy L, Kopp N, Ulrich J, Maeda T, Jouvet M (1985) Adrenergic neurons in human brain demonstrated by immunohistochemistry with antibodies to phenylethanolamine-N-methyltransferase (PNMT): discovery of a new group in the nucleus tractus solitarius. Neurosci Lett 53:303–308
Klissenbauer M, Winters S, Heinlein UA, Lisowsky T (2002) Accumulation of the mitochondrial form of the sulphydryl oxidase Erv1p/Alrp during the early stages of spermatogenesis. J Exp Biol 205:1979–1986
Kumari M, Aumüller G, Bergmann M, Meinhardt A, Seitz J (1990) Stage-dependent appearance of sulfhydryl oxidase during spermatogenesis in the testis of rat and hamster. An immunohistochemical study. Histochemistry 94:365–371
Lange H, Lisowsky T, Gerber J, Muhlenhoff U, Kispal G, Lill R (2001) An essential function of the mitochondrial sulfhydryl oxidase Erv1p/ALR in the maturation of cytosolic Fe/S proteins. EMBO Rep 2:715–720
Lee J, Hofhaus G, Lisowsky T (2000) Erv1p from Saccharomyces cerevisiae is a FAD-linked sulfhydryl oxidase. FEBS Lett 477:62–66
Lisowsky T, Lee JE, Polimeno L, Francavilla A, Hofhaus G (2001) Mammalian augmenter of liver regeneration protein is a sulfhydryl oxidase. Dig Liver Dis 33:173–180
Luo CW, Lin HJ, Chen YH (2001) A novel heat-labile phospholipid-binding protein, SVS VII, in mouse seminal vesicle as a sperm motility enhancer. J Biol Chem 276:6913–6921
Mairet-Coello G, Tury A, Fellmann D, Jouvenot M, Griffond B (2002) Expression of SOx-2, a member of the FAD-dependent sulfhydryl oxidase/quiescin Q6 gene family, in rat brain. Neuroreport 13:2049–2051
Mairet-Coello G, Tury A, Esnard-Feve A, Fellmann D, Risold PY, Griffond B (2004) FAD-linked sulfhydryl oxidase QSOX: topographic, cellular, and subcellular immunolocalization in adult rat central nervous system. J Comp Neurol 473:334–363
Matsuba S, Suga Y, Ishidoh K, Hashimoto Y, Takamori K, Kominami E, Wilhelm B, Seitz J, Ogawa H (2002) Sulfhydryl oxidase (SOx) from mouse epidermis: molecular cloning, nucleotide sequence, and expression of recombinant protein in the cultured cells. J Dermatol Sci 30:50–62
Moore EE, Kuestner RE, Conklin DC, Whitmore TE, Downey W, Buddle MM, Adams RL, Bell LA, Thompson DL, Wolf A, Chen L, Stamm MR, Grant FJ, Lok S, Ren H, De Jongh KS (1999) Stanniocalcin 2: characterization of the protein and its localization to human pancreatic alpha cells. Horm Metab Res 31:406–414
Musard JF, Sallot M, Dulieu P, Fraichard A, Ordener C, Remy-Martin JP, Jouvenot M, Adami P (2001) Identification and expression of a new sulfhydryl oxidase SOx−3 during the cell cycle and the estrus cycle in uterine cells. Biochem Biophys Res Commun 287:83–91
Ostrowski MC, Kistler WS (1980) Properties of a flavoprotein sulfhydryl oxidase from rat seminal vesicle secretion. Biochemistry 19:2639–2645
Ostrowski MC, Kistler WS, Williams-Ashman HG (1979) A flavoprotein responsible for the intense sulfhydryl oxidase activity of rat seminal vesicle secretion. Biochem Biophys Res Commun 87:171–176
Perez-Vilar J, Hill RL (1999) The structure and assembly of secreted mucins. J Biol Chem 274:31751–31754
Thorpe C, Hoober KL, Raje S, Glynn NM, Burnside J, Turi GK, Coppock DL (2002) Sulfhydryl oxidases: emerging catalysts of protein disulfide bond formation in eukaryotes. Arch Biochem Biophys 405:1–12
Tury A, Mairet-Coello G, Poncet F, Jacquemard C, Risold PY, Fellmann D, Griffond B (2004) QSOX sulfhydryl oxidase in rat adenohypophysis: localization and regulation by estrogens. J Endocrinol 183:353–363
Wagner CL, Kistler WS (1987) Analysis of the major large polypeptides of rat seminal vesicle secretion: SVS I, II, and III. Biol Reprod 36:501–510
Wittke I, Wiedemeyer R, Pillmann A, Savelyeva L, Westermann F, Schwab M (2003) Neuroblastoma-derived sulfhydryl oxidase, a new member of the sulfhydryl oxidase/quiescin6 family, regulates sensitization to interferon gamma-induced cell death in human neuroblastoma cells. Cancer Res 63:7742–7752
Acknowledgements
The authors thank Mrs. A. Laroche (Laboratoire d'Histologie, Besançon, France) for her expert technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
This research was supported by grants from the Ministère de l'Enseignement Supérieur et de la Recherche.
Rights and permissions
About this article
Cite this article
Tury, A., Mairet-Coello, G., Esnard-Fève, A. et al. Cell-specific localization of the sulphydryl oxidase QSOX in rat peripheral tissues. Cell Tissue Res 323, 91–103 (2006). https://doi.org/10.1007/s00441-005-0043-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-005-0043-x