Abstract
Genome-wide association studies (GWASs) have successfully identified a large amount of single-nucleotide polymorphisms associated with many complex phenotypes in diverse populations. However, a comprehensive understanding of the genetic correlation of associated loci of phenotypes across populations remains lacking and the extent to which associations discovered in one population can be generalized to other populations or can be utilized for trans-ethnic genetic prediction is also unclear. By leveraging summary statistics, we proposed MAGIC to evaluate the trans-ethnic marginal genetic correlation (rm) of per-allele effect sizes for associated SNPs (P < 5E-8) under the framework of measurement error models. We confirmed the methodological advantage of MAGIC over general approaches through simulations and demonstrated its utility by analyzing 34 GWAS summary statistics of phenotypes from the East Asian (Nmax = 254,373) and European (Nmax = 1,220,901) populations. Among these phenotypes, rm was estimated to range from 0.584 (se = 0.140) for breast cancer to 0.949 (se = 0.035) for age of menarche, with an average of 0.835 (se = 0.045). We also uncovered that the trans-ethnic genetic prediction accuracy for phenotypes in the target population would substantially become low when using associated SNPs identified in non-target populations, indicating that associations discovered in the one population cannot be simply generalized to another population and that the accuracy of trans-ethnic phenotype prediction is generally dissatisfactory. Overall, our study provides in-depth insight into trans-ethnic genetic correlation and prediction for complex phenotypes across diverse populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the past decade, genome-wide association studies (GWASs) have successfully identified a large amount of single nucleotide polymorphisms (SNPs) associated with many complex diseases and traits, revolutionizing our understanding of genetic foundation underlying those phenotypes (Altshuler et al. 2008; Boyle et al. 2017; Chanock et al. 2007; Gallagher and Chen-Plotkin 2018; Klein et al. 2010; McMahon et al. 2019; Tam et al. 2019; Visscher et al. 2017). However, according to the NHGRI-EBI GWAS catalog (https://www.ebi.ac.uk/gwas/) (McMahon et al. 2019), about 65.0% (= 4428/6808, up to 2021-03-25) of current GWASs in the discovery stage focus almost completely on individuals of European (EUR) descent (Bustamante et al. 2011; Rosenberg et al. 2010; Teo et al. 2010). Given the well-known genetic differentiation of complex phenotypes among various ancestries (Guo et al. 2018; MacKinnon et al. 2000; Race and Group 2005; Robinson et al. 2015; Vuckovic et al. 2020), the lack of representation of other populations in GWAS prompts an urgent demand to undertake relevant studies with non-EUR samples for validating whether association signals discovered in the EUR population can be translatable to other ethnic groups (e.g., East Asian [EAS] and African [AFR]). It has been demonstrated that a lot of disease- or trait-associated SNPs can be replicated across worldwide ethnicities (De Candia et al. 2013; Ikeda et al. 2018; Spracklen et al. 2020; Wojcik et al. 2019), providing in-depth sights into shared pathophysiology for biological mechanism, disease etiology, drug repositioning and therapeutic intervention (Gurdasani et al. 2019; Okada et al. 2014).
The knowledge regarding common genetic components of phenotypes across populations has important implications for integrating study cohorts from various ethnicities to conduct trans-ethnic meta GWASs (Li and Keating 2014; Morris 2011; Wojcik et al. 2019; Zaitlen et al. 2010), designing trans-ethnic genetic risk prediction (Coram et al. 2017), and mapping causal genes in minority populations (Coram et al. 2015; Spracklen et al. 2017). It also holds the key to benefit more ethnic groups from current medical genomics researches (Bustamante et al. 2011; Rosenberg et al. 2010; Teo et al. 2010). At the same time, quantifying genetic similarity of diseases and traits has attracted considerable research attention over the past few years (van Rheenen et al. 2019). The concept of genetic correlation has been proposed to measure such similarity within a pair of phenotypes based on GWAS datasets (Brown et al. 2016; Galinsky et al. 2019; Lee et al. 2012; van Rheenen et al. 2019). Generally, the genetic correlation is defined as the correlation between SNP effect sizes of phenotypes to measure the extent to which the same genetic variants have the same or similar influence on phenotypic variation.
Theoretically, estimating genetic correlation between phenotypes requires individual-level genotype and phenotype measurements on the same set of individuals [e.g., bivariate linear mixed model (Lee et al. 2012)], which however is not possible to observe phenotypes of interest for individuals across populations. This difficulty is effectively circumvented using GWAS summary statistics generated from diverse populations (Brown et al. 2016; Bulik-Sullivan et al. 2015a, b; Galinsky et al. 2019; Shi et al. 2017; van Rheenen et al. 2019), which enables to assess the trans-ethnic genetic similarity for a wide range of phenotypes with an unprecedented manner. For example, the genetic correlation is estimated to be 0.76 (standard error [se] = 0.04) for Crohn’s disease and 0.79 (se = 0.04) for ulcerative colitis (Liu et al. 2015), 0.46 (se = 0.06) for rheumatoid arthritis (Brown et al. 2016), 0.68 (p = 9.17E−4) for bipolar disorder (Ikeda et al. 2018), 0.33 (95% confidence intervals [CIs] 0.27–0.39) for major depressive disorder (Bigdeli et al. 2017), and 0.39 (se = 0.15) for attention-deficit hyperactivity disorder between the EUR and EAS populations; and 0.75 (se = 0.04) for height and 0.68 (se = 0.06) for body mass index (BMI) (Guo et al. 2021) between the EUR and AFR populations. In general, these estimated genetic correlations are significantly larger than zero but less than one (Brown et al. 2016; Veturi et al. 2019).
The genome-wide genetic correlation described above characterizes the average similarity of phenotypes with all available SNPs, while the genetic correlation calculated for genes or linkage disequilibrium (LD) blocks can dramatically vary across individual genetic regions (Brown et al. 2016; Paré et al. 2018; Shi et al. 2016, 2017; van Rheenen et al. 2019; Veturi et al. 2019). Under the context of genetic heterogeneity, it is sometimes more meaningful to understand the extent of genetic similarity for only associated SNPs from both scientific and practical perspectives. Scientifically, in addition to having their own value, causal SNPs are of particular importance for biologists and geneticists in revealing genetic foundation of complex phenotypes. Practically, associated genetic variants have been employed to serve as instrumental variables of exposures in Mendelian randomization (MR) studies to investigate causal relationships between phenotypes (Davey Smith et al. 2017; Davies et al. 2018; Ding et al. 2017; Disney-Hogg et al. 2018; Greenland 2000; Qi et al. 2018; Robinson et al. 2016; Voight et al. 2012; Yu et al. 2020a, b; Zeng et al. 2019; Zeng and Zhou 2019; Zhu et al. 2016, 2018), and are applied to evaluate phenotypic differentiation of phenotypes due to natural selection (Guo et al. 2018). In addition, associated SNPs are also utilized to characterize and compare genetic architecture of phenotypes across diverse ethnicities (Ishigaki et al. 2020; Okada et al. 2014; Spracklen et al. 2020; Wojcik et al. 2019) and to implement trans-ethnic genetic risk prediction for many phenotypes (Gurdasani et al. 2019; Lewis and Vassos 2017; Márquez-Luna et al. 2017; Martin et al. 2017; Martin et al. 2019).
Therefore, measuring genetic correlation among associated SNPs can offer additional insightful mechanisms shared between complex phenotypes. However, previous genetic correlations (e.g., popcorn (Brown et al. 2016)) cannot be directly adopted because they are estimated with genome-wide genetic variants, most of which are not causally related to phenotypes and thus may attenuate the genetic correlation of associated SNPs. Intuitively, the genetic correlation of associated SNPs might be much more pronounced compared to its genome-wide counterpart because the effect sizes of associated SNPs are generally more consistent in direction and magnitude across populations (Spracklen et al. 2020). Moreover, the local genetic correlation approach developed for phenotypes in the same ancestry (e.g., ρ-HESS (Shi et al. 2017)) also does not work for the trans-ethnic case because of the diversity of genetic architecture (e.g., distinct LD structures (Pritchard and Przeworski 2001; Wall and Pritchard 2003)). Because of focusing only on significant loci in our study, we thus define the corresponding genetic correlation as marginal genetic correlation (rm) to distinguish it from the global definition (Brown et al. 2016) or the local definition (Shi et al. 2017) of genetic correlation.
Conceptually, the trans-ethnic marginal genetic correlation of phenotypes can be evaluated based on per-allele SNP effect sizes by leveraging publicly available GWAS summary statistics (e.g., using the classical Pearson’s correlation). However, one main challenge is that the effect sizes of SNPs are estimated with uncertainty, which needs to be considered. For example, the two-sample-based MR studies often ignores estimation errors of instrumental variables (i.e., index SNPs) for simplicity following the assumption of no measurement error (NOME) (Bowden et al. 2016); whereas such assumption might be unrealistic and can lead to bias in causal effect estimate (Bowden et al. 2016; Corbin et al. 2016; Spiller et al. 2019; van't Hof et al. 2017). To validate the NOME assumption, it is routine to perform additional sensitivity analyses adjusted by the simulation and extrapolation (SIMEX) algorithm, which is a very creative and simulation-based general method to correct for measurement error and aims to estimate the effect size in the absence of measurement error (Carroll et al. 1996; Cook and Stefanski 1994; Guolo 2008; Lockwood and McCaffrey 2017; Stefanski and Cook 1995). As will be demonstrated later, we show that such uncertainty also cannot be simply ignored when estimating trans-ethnic marginal genetic correlation.
To quantify the trans-ethnic marginal genetic similarity for a set of uncorrelated associated SNPs of phenotypes while taking the uncertainty of estimated effect sizes into account, in the present work we proposed a new approach called MAGIC (measurement-error adjusted marginal genetic correlation) under the measurement error framework (Buonaccorsi 2010; Charles 2005). Through simulations, we demonstrated that the commonly used Pearson’s correlation often underestimated the genetic correlation, whereas MAGIC effectively addressed the issue of correlation attenuation arising if failing to correct such uncertainty. We illustrated the utility of MAGIC by analyzing 34 GWAS summary statistics of phenotypes publicly accessible from the EAS (Nmax = 254,373) and EUR (Nmax = 1,220,901) populations. We showed that extensive trans-ethnic genetic overlaps existed across these phenotypes. More importantly, we discovered that the estimated marginal genetic correlation among associated SNPs (P < 5E-8) were less than one although they were generally stronger than the overall genetic correlation obtained with genome-wide SNPs using popcorn. These findings reveal that: (i) the genetic heterogeneity also simultaneously exists among associated loci across populations; therefore, associations discovered in the one population cannot be simply generalized to another population; and (ii) the accuracy of trans-ethnic phenotype prediction is generally dissatisfactory.
Materials and methods
MAGIC evaluating trans-ethnic marginal genetic correlation of associated SNPs
Here we offer a brief overview of MAGIC, with technical details demonstrated in the Supplementary File. In brief, MAGIC takes summary statistics (e.g., marginal effect sizes and standard errors) of independent SNPs as input to assess the genetic similarity of a pair of phenotypes. Suppose we have yielded a set of uncorrelated associated SNPs for the phenotype of focus and let the estimated marginal per-allele SNP effect size be \(\hat{\alpha }\). Because of the finite sample size, \(\hat{\alpha }\) trends to be intrinsic to estimation error compared to the true effect size α. To account for this uncertainty, we model \(\hat{\alpha }\) within the measurement error framework using a signal–noise model (Buonaccorsi 2010): \(\hat{\alpha }\) = α + ε, where ε is the measurement error reflecting estimation uncertainty. Based on the model shown above, MAGIC aims to assess the trans-ethnic marginal correlation, \(r_{m} \; = \;{\text{Cor}}(\alpha_{{1}} ,\;\alpha_{{2}} )\), of genetic effects of SNP for the same trait between the first population (e.g., EAS) and the second population (e.g., EUR). It needs to emphasize that the Pearson’s correlation, \(\kappa \; = \;{\text{Cor}}(\hat{\alpha }_{{1}} ,\;\hat{\alpha }_{{2}} )\), calculated without the modification of estimation error, is often biased towards zero, resulting in |κ| ≤|rm|. This bias is also well known as correlation attenuation (Charles 2005). The sampling error and confidence intervals of rm in MAGIC are estimated with the Bootstrap approach (Efron and Tibshirani 1994). MAGIC is implemented in the R statistical environment, freely available at https://github.com/biostatpzeng/MAGIC.
Simulation studies assessing the performance of MAGIC
We now evaluated the performance of MAGIC on the estimation of trans-ethnic marginal genetic correlation with simulation studies. Considering independent genetic variants can be easily obtained via LD clumping, we thus primarily focused on uncorrected SNPs and would perform another simulation to evaluate how the LD clumping can affect the performance of MAGIC later. To generate genotypes of uncorrelated SNPs, we relied on 968 genetic variants that were independently related to body mass index (BMI) under various scenarios. In brief, we set S (the number of associated SNPs) to be 100, 200 or 300, N1 (the sample size of the phenotype in the first population) to be 1 × 104, 5 × 104 or 10 × 104 but always fixed N2 (the sample size of the phenotype in the second population) to be 10 × 104. The true value of rm varied from 0.2, 0.4, 0.6 to 0.8 or from − 0.8, − 0.6, − 0.4 to − 0.2. Details of the simulation studies are described in the Supplementary File.
Application to 34 complex phenotypes
We applied MAGIC to 34 various phenotypes which were publicly available from EAS and EUR GWAS consortia. These phenotypes included diseases (e.g., breast cancer, type 2 diabetes, and prostate cancer), social behaviors (e.g., smoking), white blood cell traits (e.g., neutrophil and monocyte count), blood lipids (e.g., triglyceride), and anthropometric traits (e.g., BMI and height). More information about these phenotypes is summarized in Table 1 and Tables S1–S2. For each analyzed phenotype, we reserved common SNPs (MAF > 1%) that were shared in both populations and then performed stringent quality control: (i) excluded non-biallelic SNPs and those with strand-ambiguous alleles; (ii) excluded SNPs having no rs labels; (iii) removed duplicated SNPs; and (iv) removed SNPs whose alleles were inconsistent between EAS and EUR populations. For each trait, we also aligned the effect allele of SNP between the two populations before the analysis.
Selecting independent SNPs associated with analyzed phenotypes
Note that, MAGIC requires independent associated genetic variants to calculate rm. Here, we considered two important applications of MAGIC. First, we wished to assess the marginal genetic correlation among potential population-common associated SNPs for every trait to quantify the genetic similarity of effect sizes. To avoid the so-called winner’s curse if choosing associated SNPs only from one single population (Kraft 2008; Zollner and Pritchard 2007), we first conducted an inverse-variance weighted meta-analysis for each shared SNP in the two populations (Willer et al. 2010). Then, based on the meta-analysis result, we selected uncorrelated index SNPs by applying the clumping procedure of PLINK (version v1.90b3.38) (Purcell et al. 2007). We set the LD threshold and the physical distance for clumping to be 0.01 or 10 Mb, respectively, with genotypes of 504 EAS and 503 EUR (a total of 1007) individuals in the 1000 Genomes Project as the reference panel for LD calculation (The 1000 Genomes Project Consortium 2015). Notably, when LD clumping, we selected index SNPs with the significance level of 5E−8 which is the widely used genome-wide significance threshold in GWAS. Here, it needs to emphasize that the applicability of MAGIC does not fully rely on the LD clumping; rather than, it primarily depends on how to define the significance and independence of SNPs that would be analyzed. Therefore, the LD clumping is only employed to obtain significant and independent SNPs analyzed by MAGIC.
Second, the genetic score prediction calculated with associated SNPs has become a useful tool for identifying individuals with high risk of complex diseases or classifying individuals into various sub-groups (De La Vega and Bustamante 2018; Dudbridge 2013; Khera et al. 2018; Krapohl et al. 2018; Lewis and Vassos 2017; Martin et al. 2019; Power et al. 2015; Zhang et al. 2018). It is of great interest to understand how accurate the trans-ethnic prediction of complex phenotypes is when using associated SNPs detected in the non-target population. The genetic correlation quantifies the maximal boundary of trans-ethnic prediction power (Brown et al. 2016; Martin et al. 2017); that is, \(R_{\max } = r_{m} \sqrt {h^{2} }\), with Rmax being the phenotypic correlation of the trans-ethnic prediction, and h2 being the heritability of the phenotype in the target population. Therefore, to examine the prediction performance using EUR-specific associated SNPs in the EAS population or vice versa, we obtained population-specific associated genetic variants for each phenotype using the similar clumping procedure described above at the same genome-wide significance level of 5E-8, with genotypes of 504 EAS or 503 EUR individuals of the 1000 Genomes Project as the reference panel (Table 1). In addition, we separately estimated h2 of the phenotype in each population using the LD score regression (LDSC) (Bulik-Sullivan et al. 2015a), with genotypes of the corresponding EAS or EUR individuals of the 1000 Genomes Project as the reference panel for LD computation.
Briefly, in the first application of MAGIC, we calculated rm with population-common associated SNPs to assess effect similarity of the same phenotypes across populations. In the second application, we calculated rm with population-specific associated SNPs to evaluate the accuracy of the trans-ethnic phenotypic prediction.
Results
Results of simulation studies
First and most importantly, in terms of the results of simulations, it is seen that MAGIC effectively handles the issue of measurement error (i.e., estimation error) that is inherent in estimated effect sizes of SNPs and generates unbiased estimates for rm (Fig. 1). In contrast, because of the incapability of accounting for such measurement error, the Pearson’s method generally underestimates rm, confirming the phenomenon of correlation attenuation (Charles 2005). Second, as anticipated, the bias in the Pearson’s correlation relies on sample size and becomes less pronounced with the increases of sample size (see below for results from another simulation). Third, the bias of the Pearson’s correlation becomes more evident as the number of associated SNPs increases, partly due to the cumulative impact of more uncertainties not explained. For example, when the true rm = 0.8 and N1 = N2 = 1 × 105, the average bias in the Pearson’s correlation is − 0.017 when S = 100, − 0.044 when S = 300 and − 0.072 when S = 500. Fourth, the two strategies used to estimate r0 (i.e., the correlation for null SNPs; see the Supplementary File for details) lead to very similar estimates of rm, implying that r0 has limited influence on \(\hat{r}_{m}\) because r0 approaches zero in our simulation settings as well as in many practical situations. These patterns are also consistently observed when rm is set to be negative (Fig. S2). We also demonstrate via simulations that MAGIC can generate approximately unbiased estimates for rm when the correlation among SNPs was simulated and the LD clumping was applied (Fig. S3).
Comparison of estimated genetic correlation rm in simulated datasets with the true value varying from 0.2, 0.4, 0.6 to 0.8 and the number of associated SNPs being set to 100, 300 or 500. In each panel, the sample size is set to 1 × 104, 5 × 104 or 10 × 104 in one population; the sample size in another population is set to 10 × 104, and PVE is set to 5% in both populations. Here, κ denotes the estimated Pearson’s correlation, rm1 denotes the genetic correlation estimated with r0 computed using null SNPs, while rm2 denotes the genetic correlation estimated with r0 = 0. (a-d) The number of associated SNPs is set to 100; (e-h) The number of associated SNPs is set to 300; (i-l) The number of associated SNPs is set to 500.
Evaluate the influence of sample size on marginal genetic correlation
As shown above, the standard error of estimated SNP effect sizes has non-negligible impact on the estimation of rm and because the standard error directly depending on sample size, we further performed another simulation to assess how the sample size influences the estimation of rm. Again, we took the 968 BMI-associated SNPs as example. Of note, for these SNPs we had \(\hat{\kappa }\) = 0.814 (se = 0.012) and \(\hat{r}_{m}\) = 0.891 (se = 0.013) (Fig. 2a). To accomplish our aim, we increased the sample size in each population by dividing the standard errors of effect sizes in summary statistics by \(a_{1}^{1/2}\) or \(a_{2}^{1/2}\) with a1 or a2 varying from 1 to 10, approximately representing the sample size increases a1 or a2 times in either population.
a Relationship of effect sizes of the 968 BMI-associated SNPs in the EAS and EUR populations; b Relationship between the genetic correlation, the sample size in the EAS population (a1) and the sample size in the EUR population (a2); c Relationship between the bias of the Pearson’s correlation, the sample size in the EAS population (a1) and the sample size in the EUR population (a2). In panel b, the red dot in the top left side indicates \(\hat{r}_{m}\) of BMI estimated with current sample sizes, while the blue dot in the bottom right indicates \(\hat{r}_{m}\) of BMI estimated with sample sizes increased to ten times. In panel c, the red and blue dots indicate the corresponding biases.
The estimated genetic correlation \(\hat{r}_{m}\) with various combinations of a1 and a2 is shown in Fig. 2b and the bias between \(\hat{r}_{m}\) and \(\hat{\kappa }\) (here we assume \(\hat{\kappa }\) is always fixed to be 0.814) is displayed in Fig. 2c. In particular, when a1 = a2 = 10, equivalently to increasing the sample size up to 1,582,840 in the EAS population and 7,135,090 in the EUR population, both of which are far larger than the sample size of currently available largest GWAS of BMI, we have \(\hat{r}_{m}\) = 0.821 and the bias = 0.007, confirming that \(\hat{r}_{m}\) would approach \(\hat{\kappa }\) as the increase of sample size because the standard error is asymptotically close to zero. In summary, these findings clearly demonstrate how the sample sizes of the two populations would influence the estimate of rm and further display the importance of adjusting for the uncertainty in estimated effect sizes when leveraging GWASs summary statistics to calculate marginal genetic correlation.
Evaluate the robustness of MAGIC with phenotypes in the same population
To evaluate the robustness and serving as a control, we further applied MAGIC to T2D summary statistics from two different European cohorts, with one available from the Wellcome Trust Case Control Consortium (WTCCC) (1924 cases and 2938 controls) (The Wellcome Trust Case Control Consortium 2007) and another obtained from the Genetic Epidemiology Research on Adult Health and Aging (GERA) cohort (7638 cases and 54,675 controls) (Banda et al. 2015). We also applied MAGIC to two sets of summary statistics of seven phenotypes from various East Asian cohorts (Table S3), including BMI (N = 141,431), height (N = 141,431), HDL (N = 69,414), LDL (N = 69,414), TC (N = 69,414), TG (N = 69,414), and HbA1c (N = 9244). To avoid the bias induced by sample overlap, for all pairs of phenotypes we had examined that no individuals were simultaneously analyzed in the two cohorts under consideration. For each pair of phenotypes, we first performed the similar quality control as described before; then we conducted the inverse-variance weighted meta-analysis for each SNP and selected uncorrelated index SNPs using the similar PLINK clumping procedure as done before. Based on these independent associated SNPs (P < 5E−8), we employed MAGIC to evaluate the genetic correlation for each phenotype within the same population. Theoretically, it is expected that the marginal genetic correlation of these phenotypes calculated within the same population (EUR or EAS) would be close to one or at least substantially high.
The estimates of marginal genetic correlation for all the eight phenotypes are shown in Table S3, which confirm our expectation that the genetic effect sizes of associated SNP for a given phenotype in the same population is highly correlated. For instance, we have a large correlation for LDL, TC and TG (\(\hat{r}_{m}\) = ~ 0.98) between Asian Genetic Epidemiology Network (AGEN) and BioBank Japan Project (BBJ) cohorts, and for height (\(\hat{r}_{m}\) = ~ 0.97) between Chinese Millionome Database (CMDB) and BBJ cohorts. The genetic correlation of T2D between WTCCC and GERA cohorts is ~ 0.94. These estimated genetic correlations are slightly smaller than one, which might be primarily due to the between-cohort heterogeneity of collected individuals.
Trans-ethnic genetic similarity of complex phenotypes
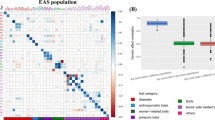
Among all the 34 analyzed phenotypes, the effective sample size varies from 12,788 for BRC to 254,373 for T2D in the EAS population and from 43,290 for RA to 1,220,901 for smoking initiation in the EUR population (Table 1 and Tables S1–S2). To evaluate the genetic similarity of SNP effect sizes between the two populations, we first obtained a set of independent population-common associated genetic variants for each trait, with the number of SNPs ranging from 53 for HbA1c to 968 for BMI (i.e., S12 in Table 1). The scatter plots of effect sizes for these phenotypes display clearly positive correlation between the two populations (Fig. S4). With these SNPs, the marginal genetic correlation is estimated using MAGIC. For comparison, we also evaluated the Pearson’s correlation (κ) and the global genetic correlation (ρg), with the former calculated with these uncorrelated associated SNPs and the latter calculated with all available genome-wide SNPs.
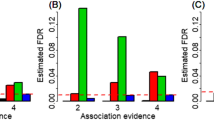
The main results of MAGIC are shown in Fig. 3. The estimate \(\hat{r}_{m}\) ranges from 0.584 (se = 0.140) for breast cancer to 0.949 (se = 0.035) for age of menarche, with an average of 0.835 (se = 0.045) (Fig. 3a), partly reflecting the genetic ethnic discrepancy across phenotypes. We observe that on average the genetic correlation for continuous phenotypes is higher compared to that for binary phenotypes, with an average of 0.844 (se = 0.043) for the former vs. 0.804 (se = 0.062) for the latter. As mentioned before, due to the correction of measurement error, \(\hat{r}_{m}\) is generally larger than the Pearson’s correlation \(\hat{\kappa }\) (Fig. 3b) and is also much greater than the average genetic correlation across the whole genome (Fig. 3c), in line with previous observations that local genetic correlation may be substantial while the overall genetic correlation is weak (Shi et al. 2017). Moreover, on average, \(\hat{r}_{m}\) increases about 6.8% compared to \(\hat{\kappa }\), with the maximal improvement of 25.8% for breast cancer from \(\hat{\kappa }\) = 0.465 (se = 0.106) to \(\hat{r}_{m}\) = 0.584 (se = 0.140), followed by pulse pressure (16.2%), diastolic blood pressure (16.1%) and age of menarche (15.1%). In addition, it is seen that \(\hat{r}_{m}\) is also generally larger than its global counterpart \(\hat{\rho }_{g}\) for most of these analyzed phenotypes (Fig. 3d), suggesting that associated SNPs often exhibit higher similarity between diverse populations compared with these general genetic variants.
a Estimated marginal genetic correlation and its 95% confidence intervals for each analyzed trait; b Comparison between the marginal genetic correlation and the Pearson’s correlation; c Comparison of the marginal genetic correlation among associated SNPs and the average gene-based genetic correlation across the whole genome; d Comparison of the marginal genetic correlation among associated SNPs and the global genetic correlation across the whole genome.
Note that, among these selected SNPs, some may be related to both phenotypes and some may be associated with the trait in a population-specific manner. One might conjecture that the genetic correlation would become higher if more population-common associated SNPs are incorporated in MAGIC. To examine this, we calculated the proportion of independent SNPs which were simultaneously associated with the phenotypes of focus in both EAS and EUR populations at the significance level for clumping. However, we observe little evidence supporting the existence of the relationship between this proportion and the estimated \(\hat{r}_{m}\) (Fig. S5).
In addition, it is discovered that the popcorn method designed to estimate ρg using genome-wide SNPs can suffer from the issue of numerical instability when using it to estimate genetic correlation with only a set of selected SNPs. For instance, it failed to convergence for approximately 48.5% of genes and 31.0% of the estimated genetic correlations were outside of the boundary of 1 and − 1 across all the 34 analyzed phenotypes when applying popcorn to evaluate gene-based similarity. The similar numerical problem also emerged when applying popcorn to assess genetic correlation of associated SNPs selected for these phenotypes. For example, popcorn failed to convergence for 12 of 34 phenotypes and the estimates of genetic correlation for another 16 of 34 phenotypes were outside of the boundary of 1 and -1.
Trans-ethnic genetic prediction accuracy for complex phenotypes
To assess the trans-ethnic genetic prediction accuracy of complex phenotypes, we yielded another set of independent population-specific associated SNPs for each trait (i.e., S1 and S2 in Table 1). Based on these SNPs, we calculated rm1 (i.e., marginal genetic correlation based on pairs of EAS-specific associated SNPs) and rm2 (i.e., marginal genetic correlation based on pairs of EUR-specific associated SNPs); but we estimated h2 using genome-wide SNPs. For comparison, we denoted rm calculated above as rm12. There are several interesting findings needing to be highlighted. First, both rm1 and rm2 are generally smaller than rm12 (Fig. 4). For instance, on average an approximately 9.8% decrease in genetic correlation is observed when using EAS-specific associated genetic variants, and a larger reduction (~ 14.6%) occurs when applying EUR-specific associated SNPs; suggesting that the effect sizes of the population-specific associated SNPs are more variable compared to those of the population-common associated SNPs.
Estimated genetic correlation based on different sets of independent associated SNPs for each trait. The grey box represents genetic correlation calculated with EAS-specific associated SNPs (rm1); the blue box represents genetic correlation calculated with EUR-specific associated SNPs (rm2); and the pink box represents genetic correlation calculated with population-common SNPs (rm12)
Second, besides the difference in magnitude, compared with rm12, rm1 and rm2 also vary dramatically across phenotypes. For instance, rm2 is only 0.076 for age at menarche but is 0.820 for BMI and has a variance of 0.028 across all phenotypes when using EUR-specific associated SNPs (the variance is 0.028 for rm1 and 0.022 for rm12). Third, the trans-ethnic genetic prediction accuracy becomes low compared to that assessed within individuals of the same descent. For example, Rmax would reduce approximately 38.5% from 0.19 to 0.12 if predicting T2D in the EAS population using associated SNPs identified in the EUR population (Table S3), and the other seven phenotypes would on average suffer from a reduction of 20.6% in prediction accuracy if predicting them in the EUR population using associated SNPs discovered in the EAS population; demonstrating less satisfactory performance in trans-ethnic prediction for complex phenotypes.
Discussion
Previous work revealed that there existed substantial overall genetic overlap in the same phenotypes in diverse populations (Brown et al. 2016). However, as shown in the population with the same ancestry, the global genetic correlation ρg sometimes might offer a biased knowledge about the local correlation for certain genetic regions of interest (Shi et al. 2017). In the present study, we have proposed a novel approach called MAGIC to evaluate the marginal genetic correlation rm of complex phenotypes in diverse populations (van Rheenen et al. 2019). Conceptually, rm is supplementary to ρg and may be more attractive because one is often more interesting in associated SNPs compared to genome-wide genetic variants in many practical applications (Guo et al. 2018; Gurdasani et al. 2019; Ishigaki et al. 2020; Lewis and Vassos 2017; Márquez-Luna et al. 2017; Martin et al. 2017; Martin et al. 2019; Okada et al. 2014; Spracklen et al. 2020; Wojcik et al. 2019; Yu et al. 2020a, b; Zeng et al. 2019). Both simulations and real-life data analyses have demonstrated that the estimation error of marginal effect size of associated SNPs has non-ignorable influence on the estimation of rm and the adjustment of such error is necessary.
In the analysis of 34 phenotypes, consistent with previous finding (Brown et al. 2016), it was shown that the estimated genetic correlation was generally larger than zero but less than one, implying potential genetic diversity in complex phenotypes across populations. Indeed, using the Cochran's Q test we found that on average 33.2% (ranging from 16.5% for breast cancer to 64.2% for HbA1c) of these population-common associated SNPs showed genetic effect heterogeneity between the EAS and EUR populations at the significance level of 0.05; this proportion decreased to 11.3% after taking multiple comparisons into account and adjusting with the Bonferroni’s method. We also observed some of associated SNPs had obviously distinct genetic effects on the same phenotypes between the two populations (Fig. S6a), implying the population-specific effects for these genetic variants. Moreover, it was identified that 70.6% (24 out of 34) of analyzed phenotypes exhibited ethnic difference in MAF at the significance level of 0.05, decreasing to 50.0% (17 out of 34) if adjusting the issue of multiple comparisons (Fig. S6b). It is noted, consistent with the results reported recently (Ishigaki et al. 2020), that some phenotypes (e.g., BMI, diastolic blood pressure, systolic blood pressure, and pulse pressure) generally had smaller MAF in the EUR population compared to that in the EAS population, indicating the global genetic differentiation of complex phenotypes (Guo et al. 2018; Schoech et al. 2019; Zeng et al. 2018). The possible reasons for such genetic differentiation may include natural selection (Guo et al. 2018) and gene-environment interaction (Wang et al. 2019).
Moreover, we observed that on average the genetic correlation of continuous phenotypes was higher compared with that for binary diseases, which may indicate larger genetic heterogeneity in diseases among diverse populations due to the difference in disease definition, diagnose and classification. We also offered an intuitively reasonable conclusion that rm should be in general greater than ρg since the former only quantifies the genetic similarity of causal genetic variants, which often have higher effect consistence in direction and magnitude (Spracklen et al. 2020). This observation also suggests that associated SNPs, compared to general genetic variants, contribute disproportionately to genetic sharing between phenotypes in diverse populations. Because the genetic correlation quantifies the maximal boundary of trans-ethnic prediction power; our results further indicate that the genetic risk score prediction in the target population (i.e., EAS) using associated SNPs identified non-target populations (EUR) should be interpreted with caution (Brown et al. 2016; Martin et al. 2017). Together, our study provides a definite answer for the previous question that the associations identified in the EUR population cannot be simply applied to other non-EUR populations, highlighting the necessary of performing GWASs with various ancestries.
As shown in Fig. S6, SNPs often exhibit difference in allele frequency across populations. To explain the influence of such allele variances, we can also assess local genetic impact correlation Cor(α1σ1, α2σ2)-based normalized effect sizes of associated SNPs relying on the similar principle of rm, where σ = 2f (1 − f) with f being MAF in the EUR or EAS population. While we found that the local genetic impact correlation was almost identical to rm (results not shown), in line with the prior finding that the overall trans-ethnic genetic impact correlation was also very consistent with the overall trans-ethnic genetic effect correlation (Brown et al. 2016), indicating that the genetic correlation between each pair of phenotypes may be primarily derived by effect sizes of SNPs rather than allele frequencies. However, it is not known whether this conclusion still holds for rare genetic variants because only common SNPs with MAF > 1% were analyzed in our study.
Compared with previous approaches, MAGIC possesses several appealing advantages. First, compared to the traditional Pearson’s correlation which often underestimates rm, MAGIC takes the uncertainty of summary statistics into account and thus corrects the so-called issue of correlation attenuation due to estimation error (Buonaccorsi 2010; Charles 2005). Second, MAGIC does not require the assignment of effect distribution for SNPs and thus has wider applicability. While previous genetic correlation methods often assume a special distribution (e.g., the normal distribution in the infinitesimal model supposed by popcorn (Brown et al. 2016)) for effect sizes of SNPs, which may be not realistic in practice. Third, due to only a small set of independent SNPs involved, MAGIC does not need reference panels to calculate LD matrix and is thus much computationally efficient. Fourth, MAGIC models the genetic correlation directly using effect sizes of SNPs; it can be applied for not only continuous phenotypes, but also binary phenotypes. Previous genetic correlation methods often suppose a liability threshold model when the phenotypes are binary (Brown et al. 2016; Shi et al. 2017). In summary, MAGIC represents a conceptually general and computationally fast approach for the evaluation of genetic correlation.
Finally, some limitations of the present study should be described. First, one of the main limitations is that multiple phenotypes in the EAS GWAS had a relatively smaller sample size compared to the EUR GWAS (92 K vs. 290 K on average across all analyzed phenotypes), meaning that many trait-associated SNPs with weak effect sizes cannot be successfully detected in the EAS population due to low statistical power, which would consequently undermine the estimation of the local genetic correlation rm. Second, because most of current GWASs only analyzed common SNPs; therefore, we cannot explore the genetic correlation of phenotypes for rare genetic variants (e.g., MAF < 1%). It can be reasonably speculated that the genetic correlation among rare SNPs may be much weaker than that among common SNPs because trait-associated rare genetic variants are generally population-specific (Bomba et al. 2017; Visscher et al. 2017). Third, because of the lack of relevant datasets, we cannot conduct a two-step MAGIC analysis so that we can select SNPs in one dataset but estimate the marginal genetic correlation in another dataset. Instead, we had to perform the two steps using the same set of GWAS summary statistics for each trait. Doing this might lead to potentially upward bias; however, we found that the resulting bias was minimal (Supplementary File).
Conclusion
Overall, based on a compressive evaluation of summary statistics of 34 various phenotypes, our study provides new insight into genetic foundation for pairs of phenotypes among diverse populations and provides better understanding of trans-ethnic genetic correlation and prediction for complex phenotypes.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information file.
References
Altshuler D, Daly M, Lander E (2008) Genetic mapping in human disease. Science 322:881–888
Banda Y, Kvale MN, Hoffmann TJ et al (2015) Characterizing race/ethnicity and genetic ancestry for 100,000 subjects in the Genetic Epidemiology Research on Adult Health and Aging (GERA) cohort. Genetics 200:1285–1295
Bigdeli TB, Ripke S, Peterson RE et al (2017) Genetic effects influencing risk for major depressive disorder in China and Europe. Transl Psychiatry 7:e1074–e1074
Bomba L, Walter K, Soranzo N (2017) The impact of rare and low-frequency genetic variants in common disease. Genome Biol 18:77
Bowden J, Del Greco MF, Minelli C et al (2016) Assessing the suitability of summary data for two-sample Mendelian randomization analyses using MR-Egger regression: the role of the I 2 statistic. Int J Epidemiol 45:1961–1974
Boyle EA, Li YI, Pritchard JK (2017) An expanded view of complex traits: from polygenic to omnigenic. Cell 169:1177–1186
Brown BC, Ye CJ, Price AL et al (2016) Transethnic genetic-correlation estimates from summary statistics. Am J Hum Genet 99:76–88
Bulik-Sullivan B, Finucane HK, Anttila V et al (2015a) An atlas of genetic correlations across human diseases and traits. Nat Genet 47:1236–1241
Bulik-Sullivan BK, Loh P-R, Finucane HK et al (2015b) LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 47:291–295
Buonaccorsi JP (2010) Measurement error: models, methods, and applications. Chapman and Hall/CRC, New York
Bustamante CD, Burchard EG, De la Vega FM (2011) Genomics for the world. Nature 475:163–165
Carroll RJ, Küchenhoff H, Lombard F et al (1996) Asymptotics for the SIMEX estimator in nonlinear measurement error models. J Am Stat Assoc 91:242–250
Chanock S, Manolio T, Boehnke M et al (2007) Replicating genotype–phenotype associations. Nature 447:655–660
Charles E (2005) The correction for attenuation due to measurement error: clarifying concepts and creating confidence sets. Psychol Methods 10:206–226
Cook JR, Stefanski LA (1994) Simulation-extrapolation estimation in parametric measurement error models. J Am Stat Assoc 89:1314–1328
Coram MA, Candille SI, Duan Q et al (2015) Leveraging multi-ethnic evidence for mapping complex traits in minority populations: an empirical Bayes approach. Am J Hum Genet 96:740–752
Coram MA, Fang H, Candille SI et al (2017) Leveraging multi-ethnic evidence for risk assessment of quantitative traits in minority populations. Am J Hum Genet 101:218–226
Corbin LJ, Richmond RC, Wade KH et al (2016) BMI as a modifiable risk factor for type 2 diabetes: refining and understanding causal estimates using Mendelian randomization. Diabetes 65:3002–3007
Davey Smith G, Paternoster L, Relton C (2017) When will Mendelian randomization become relevant for clinical practice and public health? JAMA 317:589–591
Davies NM, Dickson M, Davey Smith G et al (2018) The causal effects of education on health outcomes in the UK Biobank. Nat Hum Behav 2:117–125
De Candia TR, Lee SH, Yang J et al (2013) Additive genetic variation in schizophrenia risk is shared by populations of African and European descent. Am J Hum Genet 93:463–470
De La Vega FM, Bustamante CD (2018) Polygenic risk scores: a biased prediction? Genome Med 10:100
Ding M, Huang T, Bergholdt HK et al (2017) Dairy consumption, systolic blood pressure, and risk of hypertension: Mendelian randomization study. BMJ 356:j1000
Disney-Hogg L, Cornish AJ, Sud A et al (2018) Impact of atopy on risk of glioma: a Mendelian randomisation study. BMC Med 16:42
Dudbridge F (2013) Power and predictive accuracy of polygenic risk scores. PLoS Genet 9:e1003348
Efron B, Tibshirani RJ (1994) An introduction to the bootstrap. CRC Press
Galinsky KJ, Reshef YA, Finucane HK et al (2019) Estimating cross-population genetic correlations of causal effect sizes. Genet Epidemiol 43:180–188
Gallagher MD, Chen-Plotkin AS (2018) The post-GWAS era: from association to function. Am J Hum Genet 102:717–730
Greenland S (2000) An introduction to instrumental variables for epidemiologists. Int J Epidemiol 29:722–729
Guo J, Wu Y, Zhu Z et al (2018) Global genetic differentiation of complex traits shaped by natural selection in humans. Nat Commun 9:1865
Guo J, Bakshi A, Wang Y et al (2021) Quantifying genetic heterogeneity between continental populations for human height and body mass index. Sci Rep 11:1–9
Guolo A (2008) Robust techniques for measurement error correction: a review. Stat Methods Med Res 17:555–580
Gurdasani D, Barroso I, Zeggini E et al (2019) Genomics of disease risk in globally diverse populations. Nat Rev Genet 20:520–535
Ikeda M, Takahashi A, Kamatani Y et al (2018) A genome-wide association study identifies two novel susceptibility loci and trans population polygenicity associated with bipolar disorder. Mol Psychiatry 23:639–647
Ishigaki K, Akiyama M, Kanai M et al (2020) Large-scale genome-wide association study in a Japanese population identifies novel susceptibility loci across different diseases. Nat Genet 52:669–679
Khera AV, Chaffin M, Aragam KG et al (2018) Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet 50:1219–1224
Klein RJ, Xu X, Mukherjee S et al (2010) Successes of genome-wide association studies. Cell 142:350–351
Kraft P (2008) Curses—winner’s and otherwise—in genetic epidemiology. Epidemiology 19:649–651
Krapohl E, Patel H, Newhouse S et al (2018) Multi-polygenic score approach to trait prediction. Mol Psychiatry 23:1368–1374
Lee SH, Yang J, Goddard ME et al (2012) Estimation of pleiotropy between complex diseases using single-nucleotide polymorphism-derived genomic relationships and restricted maximum likelihood. Bioinformatics 28:2540–2542
Lewis CM, Vassos E (2017) Prospects for using risk scores in polygenic medicine. Genome Med 9:96
Li YR, Keating BJ (2014) Trans-ethnic genome-wide association studies: advantages and challenges of mapping in diverse populations. Genome Med 6:91
Liu JZ, van Sommeren S, Huang H et al (2015) Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat Genet 47:979–986
Lockwood J, McCaffrey DF (2017) Simulation-extrapolation with latent heteroskedastic error variance. Psychometrika 82:717–736
MacKinnon DP, Krull JL, Lockwood CM (2000) Equivalence of the mediation, confounding and suppression effect. Prev Sci 1:173–181
Márquez-Luna C, Loh P-R, Consortium SATD et al (2017) Multiethnic polygenic risk scores improve risk prediction in diverse populations. Genet Epidemiol 41:811–823
Martin AR, Gignoux CR, Walters RK et al (2017) Human demographic history impacts genetic risk prediction across diverse populations. Am J Hum Genet 100:635–649
Martin AR, Kanai M, Kamatani Y et al (2019) Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet 51:584–591
McMahon A, Malangone C, Suveges D et al (2019) The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res 47:D1005–D1012
Morris AP (2011) Transethnic meta-analysis of genomewide association studies. Genet Epidemiol 35:809–822
Okada Y, Wu D, Trynka G et al (2014) Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506:376–381
Paré G, Mao S, Deng WQ (2018) A robust method to estimate regional polygenic correlation under misspecified linkage disequilibrium structure. Genet Epidemiol 42:636–647
Power RA, Steinberg S, Bjornsdottir G et al (2015) Polygenic risk scores for schizophrenia and bipolar disorder predict creativity. Nat Neurosci 18:953–955
Pritchard JK, Przeworski M (2001) Linkage disequilibrium in humans: models and data. Am J Hum Genet 69:1–14
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Qi T, Wu Y, Zeng J et al (2018) Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood. Nat Commun 9:2282–2282
Race E, Group GW (2005) The use of racial, ethnic, and ancestral categories in human genetics research. Am J Hum Genet 77:519–532
Robinson MR, Hemani G, Medina-Gomez C et al (2015) Population genetic differentiation of height and body mass index across Europe. Nat Genet 47:1357–1362
Robinson PC, Choi HK, Do R et al (2016) Insight into rheumatological cause and effect through the use of Mendelian randomization. Nat Rev Rheumatol 12:486–496
Rosenberg NA, Huang L, Jewett EM et al (2010) Genome-wide association studies in diverse populations. Nat Rev Genet 11:356–366
Schoech AP, Jordan DM, Loh P-R et al (2019) Quantification of frequency-dependent genetic architectures in 25 UK Biobank traits reveals action of negative selection. Nat Commun 10:790
Shi H, Kichaev G, Pasaniuc B (2016) Contrasting the genetic architecture of 30 complex traits from summary association data. Am J Hum Genet 99:139–153
Shi H, Mancuso N, Spendlove S et al (2017) Local genetic correlation gives insights into the shared genetic architecture of complex traits. Am J Hum Genet 101:737–751
Spiller W, Davies NM, Palmer TM (2019) Software application profile: mrrobust—a tool for performing two-sample summary Mendelian randomization analyses. Int J Epidemiol 48:684
Spracklen CN, Chen P, Kim YJ et al (2017) Association analyses of East Asian individuals and trans-ancestry analyses with European individuals reveal new loci associated with cholesterol and triglyceride levels. Hum Mol Genet 26:1770–1784
Spracklen CN, Horikoshi M, Kim YJ et al (2020) Identification of type 2 diabetes loci in 433,540 East Asian individuals. Nature 582:240–245
Stefanski LA, Cook JR (1995) Simulation-extrapolation: the measurement error jackknife. J Am Stat Assoc 90:1247–1256
Tam V, Patel N, Turcotte M et al (2019) Benefits and limitations of genome-wide association studies. Nat Rev Genet 20:467–484
Teo Y-Y, Small KS, Kwiatkowski DP (2010) Methodological challenges of genome-wide association analysis in Africa. Nat Rev Genet 11:149–160
The 1000 Genomes Project Consortium (2015) A global reference for human genetic variation. Nature 526:68–74
The Wellcome Trust Case Control Consortium (2007) Genome-wide association study of 14,000 cases of seven common diseases and 3000 shared controls. Nature 447:661–678
van Rheenen W, Peyrot WJ, Schork AJ et al (2019) Genetic correlations of polygenic disease traits: from theory to practice. Nat Rev Genet 20:567–581
van’t Hof FNG, Vaucher J, Holmes MV et al (2017) Genetic variants associated with type 2 diabetes and adiposity and risk of intracranial and abdominal aortic aneurysms. Eur J Hum Genet 25:758–762
Veturi Y, de los Campos G, Yi N et al (2019) Modeling heterogeneity in the genetic architecture of ethnically diverse groups using random effect interaction models. Genetics 211:1395–1407
Visscher PM, Wray NR, Zhang Q et al (2017) 10 years of GWAS discovery: biology, function, and translation. Am J Hum Genet 101:5–22
Voight BF, Peloso GM, Orho-Melander M et al (2012) Plasma HDL cholesterol and risk of myocardial infarction: a mendelian randomisation study. Lancet 380:572–580
Vuckovic D, Bao EL, Akbari P et al (2020) The polygenic and monogenic basis of blood traits and diseases. Cell 182:1214-1231.e11
Wall JD, Pritchard JK (2003) Haplotype blocks and linkage disequilibrium in the human genome. Nat Rev Genet 4:587–597
Wang H, Zhang F, Zeng J et al (2019) Genotype-by-environment interactions inferred from genetic effects on phenotypic variability in the UK Biobank. Sci Adv 5:eaaw3538
Willer CJ, Li Y, Abecasis GR (2010) METAL: fast and efficient meta-analysis of genome-wide association scans. Bioinformatics 26:2190–2191
Wojcik GL, Graff M, Nishimura KK et al (2019) Genetic analyses of diverse populations improves discovery for complex traits. Nature 570:514–518
Yu X, Wang T, Chen Y et al (2020a) Alcohol drinking and amyotrophic lateral sclerosis: an instrumental variable causal inference. Ann Neurol 88:195–198
Yu X, Yuan Z, Lu H et al (2020b) Relationship between birth weight and chronic kidney disease: evidence from systematics review and two-sample Mendelian randomization analysis. Hum Mol Genet 29:2261–2274
Zaitlen N, Paşaniuc B, Gur T et al (2010) Leveraging genetic variability across populations for the identification of causal variants. Am J Hum Genet 86:23–33
Zeng P, Zhou X (2019) Causal effects of blood lipids on amyotrophic lateral sclerosis: a Mendelian randomization study. Hum Mol Genet 28:688–697
Zeng J, De Vlaming R, Wu Y et al (2018) Signatures of negative selection in the genetic architecture of human complex traits. Nat Genet 50:746–753
Zeng P, Wang T, Zheng J et al (2019) Causal association of type 2 diabetes with amyotrophic lateral sclerosis: new evidence from Mendelian randomization using GWAS summary statistics. BMC Med 17:225
Zhang X, Rice M, Tworoger SS et al (2018) Addition of a polygenic risk score, mammographic density, and endogenous hormones to existing breast cancer risk prediction models: a nested case–control study. PLoS Med 15:e1002644
Zhu Z, Zhang F, Hu H et al (2016) Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet 48:481–487
Zhu Z, Zheng Z, Zhang F et al (2018) Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat Commun 9:224
Zollner S, Pritchard J (2007) Overcoming the winner’s curse: estimating penetrance parameters from case-control. Am J Hum Genet 80:605–615
Acknowledgements
We thank all the GWAS consortia for making summary statistics publicly available for us and are grateful to all the investigators and participants contributed to those studies. The data analyses in the present study were carried out with the high-performance computing cluster that was supported by the special central finance project of local universities for Xuzhou Medical University. We are especially grateful to two anonymous referees for making a lot of constructive comments that have led to substantial improvements of our manuscript.
Funding
The research of Ping Zeng was supported in part by the Youth Foundation of Humanity and Social Science funded by Ministry of Education of China (18YJC910002), the Natural Science Foundation of Jiangsu Province of China (BK20181472), the China Postdoctoral Science Foundation (2018M630607 and 2019T120465), the QingLan Research Project of Jiangsu Province for Outstanding Young Teachers, the Six-Talent Peaks Project in Jiangsu Province of China (WSN-087), the Training Project for Youth Teams of Science and Technology Innovation at Xuzhou Medical University (TD202008), the Postdoctoral Science Foundation of Xuzhou Medical University, the National Natural Science Foundation of China (81402765), and the Statistical Science Research Project from National Bureau of Statistics of China (2014LY112). The research of Shuiping Huang was supported in part by the Social Development Project of Xuzhou City (KC19017). The research of Ting Wang was supported in part by the Social Development Project of Xuzhou City (KC20062).
Author information
Authors and Affiliations
Contributions
PZ conceived the idea for the study. PZ, TW, and SH obtained and cleared the datasets; PZ, HL, TW, JZ, and SZ performed the data analyses. PZ, HL, and TW interpreted the results of the data analyses. PZ and HL wrote the manuscript with the help from other authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Consent for publication
All the authors agreed that this manuscript be submitted to the journal of Human Genetics for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lu, H., Wang, T., Zhang, J. et al. Evaluating marginal genetic correlation of associated loci for complex diseases and traits between European and East Asian populations. Hum Genet 140, 1285–1297 (2021). https://doi.org/10.1007/s00439-021-02299-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-021-02299-8