Abstract
In human society, the facial surface is visible and recognizable based on the facial shape variation which represents a set of highly polygenic and correlated complex traits. Understanding the genetic basis underlying facial shape traits has important implications in population genetics, developmental biology, and forensic science. A number of single nucleotide polymorphisms (SNPs) are associated with human facial shape variation, mostly in European populations. To bridge the gap between European and Asian populations in term of the genetic basis of facial shape variation, we examined the effect of these SNPs in a European–Asian admixed Eurasian population which included a total of 612 individuals. The coordinates of 17 facial landmarks were derived from high resolution 3dMD facial images, and 136 Euclidean distances between all pairs of landmarks were quantitatively derived. DNA samples were genotyped using the Illumina Infinium Global Screening Array and imputed using the 1000 Genomes reference panel. Genetic association between 125 previously reported facial shape-associated SNPs and 136 facial shape phenotypes was tested using linear regression. As a result, a total of eight SNPs from different loci demonstrated significant association with one or more facial shape traits after adjusting for multiple testing (significance threshold p < 1.28 × 10−3), together explaining up to 6.47% of sex-, age-, and BMI-adjusted facial phenotype variance. These included EDAR rs3827760, LYPLAL1 rs5781117, PRDM16 rs4648379, PAX3 rs7559271, DKK1 rs1194708, TNFSF12 rs80067372, CACNA2D3 rs56063440, and SUPT3H rs227833. Notably, the EDAR rs3827760 and LYPLAL1 rs5781117 SNPs displayed significant association with eight and seven facial phenotypes, respectively (2.39 × 10−5 < p < 1.28 × 10−3). The majority of these SNPs showed a distinct allele frequency between European and East Asian reference panels from the 1000 Genomes Project. These results showed the details of above eight genes influence facial shape variation in a Eurasian population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Facial morphology represents the most recognizable feature in humans with a strong genetic component. Several family based studies have estimated the heritability of certain facial shape features up to 0.73 (Alkhudhairi and Alkofide 2010), but an understanding of the genetic basis of normal variation in human facial morphology remains limited.
To date, ten genome-wide association studies (GWASs) have been performed to examine the associations between DNA variants and normal facial variation. These studies reported a total of 125 SNPs at 103 distinct genomic loci with genome-wide significant association to a number of different facial features (Adhikari et al. 2016; Cha et al. 2018; Claes et al. 2018; Cole et al. 2017; Crouch et al. 2018; Lee et al. 2017a; Liu et al. 2012; Paternoster et al. 2012; Pickrell et al. 2016b; Shaffer et al. 2016). These GWASs used a variety of phenotyping approaches, ranging from questionnaires on anthropological features to the analysis of 2D images and/or 3D head MRI or facial surface data. With the exception of a few, most of the identified loci are non-overlapping between the independent GWASs. These findings are largely consistent with a highly polygenic model and suggest a high degree of population heterogeneity underlying human facial variation. Because most of the previous GWASs of facial variation were conducted on European populations, whether these findings are generalizable in Asian populations remains unclear. Here, we investigated the potential effects of the 125 facial variation-associated SNPs on facial morphology in a European–Asian admixed population.
Materials and methods
Samples
This study was approved by the Ethics Committee of the Institute of Forensic Science of China, and all individuals provided written informed consent. The participants were all volunteers. The consent was discussed in their native language and the signature was in their native language. We sampled a total of 612 unrelated Eurasian individuals living in Tumxuk City in Xinjiang Uyghur Autonomous Region, China. All individuals met the following conditions: (1) their parents and grandparents were both of Uyghur origin; (2) they had not received hormone therapy; (3) they had no thyroid disease, pituitary disease, or tumors; and (4) they had no medical conditions affecting growth and development, such as dwarfism, gigantism, and acromegaly. The 3D facial surface data were ascertained using an Artec Spider scanner in combination with Artec Studio Professional v10 software, and all volunteers were requested to maintain the same sitting position and neutral expression.
Phenotyping
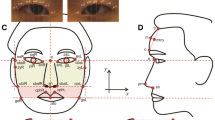
The x–y–z coordinates of 17 facial landmarks were derived from the 3D face images based on an automated pipeline developed in-house by fine-tuning a previously detailed protocol (Guo et al. 2013). The method starts with preliminary nose tip localization and pose normalization, followed by localization of the six most salient landmarks using principal component analysis (PCA) and heuristic localization of 10 additional landmarks. Trained experts reviewed all landmarks from the automated pipeline by comparing the landmark positions with example images pre-landmarked according to the definition of the landmarks (Table S1) using the FaceAnalysis software (Guo et al. 2013). Obviously inaccurately positioned landmarks were corrected using the 3dMD patient software (www.3dmd.com). After generalized procrustes analysis (GPA), a total of 136 Euclidean distances between all pairs of the 17 landmarks were quantitatively derived. Outliers with values greater than three standard deviations were removed. Z-transformed phenotypes were used in the subsequent analyses.
DNA genotyping, quality control, and imputation
Venous whole blood samples were collected in EDTA-Vacutainer tubes and stored at − 20 °C until processed. DNA samples were genotyped on an Illumina Infinium Global Screening Array 650 K. SNPs with minor-allele frequency < 1%, call-rate < 97%, Hardy–Weinberg p values < 0.0001, and samples missing > 3% of genotypes were excluded. One sample with excess of heterozygosity (F < 0.084) was excluded. Two samples were identified as second-degree relatives in identity by descent (IBD) estimation, and one was removed. Genotype imputation was performed to capture information on unobserved SNPs and sporadically missing genotypes among the genotyped SNPs, using all haplotypes from the 1000 Genomes Project Phase 3 reference panel (Genomes Project et al. 2012). Pre-phasing was performed in SHAPEIT2 (Delaneau et al. 2013), and imputation was performed using IMPUTE2 (Howie et al. 2011; Howie et al. 2009). Imputed SNPs with INFO scores < 0.8 were excluded. The imputed dataset contained genotypes for 5,289,934 SNPs. We ascertained a list of 125 SNPs that have been associated with facial morphology in previous facial morphology GWASs (Adhikari et al. 2016; Cha et al. 2018; Claes et al. 2018; Cole et al. 2016; Crouch et al. 2018; Lee et al. 2017b; Liu et al. 2012; Paternoster et al. 2012; Pickrell et al. 2016a; Shaffer et al. 2016). Out of the 125 SNPs, 10 were genotyped, 47 were imputed and passed quality control, and 68 were excluded by quality control.
Statistical analyses
Linear regressions were iteratively conducted to test genetic association between the facial shape-associated SNPs and the facial phenotypes under an additive genetic model, while adjusting for sex, age, BMI, and the first three genomic principal components from the –pca function in PLINK V1.9 (Purcell et al. 2007). We conducted a genomic PCA analysis to detect the presence of potential population substructures, using three population samples, i.e., 612 Uyghurs (UYG) from the current study, 504 East Asians (EAS) and 503 Central Europeans (EUR) from the 1000 Genomes Project (Genomes Project et al. 2012), and an overlapping set of 5,085,557 SNPs. The relative contribution was derived for the top 20 PCs. An unsupervised K-means clustering analysis was used to cluster the three population samples into three clusters based on the top-contributing genomic PCs (Hartigan and Wong 1979). We used the distance matrix that was derived from the phenotypes correlation matrix to perform hierarchical clustering analysis with the –dist and –hclust function in R V3.3.2 and obtained four phenotype clusters.
To adjust for the multiple testing of multiple phenotypes, we conducted a Bonferroni correction to the effective number of independent variables, which was estimated using the Matrix Spectral Decomposition (matSpD) method (Li and Ji 2005). The fraction of trait variance explained by the SNPs was estimated using multiple regressions, where the face residuals were considered as the phenotype, i.e.: the effects of sex, age, and BMI were regressed out prior to the analysis. The distribution of allele frequencies in the 2504 subjects of the 1000 Genomes Project was visualized using Mapviewer software version 7.
Results
This study included 590 males and 22 females, ranging from 16 to 59 years of age (mean age was 34.9 years, Table S2), of admixed European-Asian ancestry. We focused on 17 anatomical landmarks (Fig. 1), and the 136 Euclidian distances (Figure S1, Table S2) between all of these landmarks. Age had a significant effect on 80.1% of all 136 face phenotypes (1.06 × 10−17 < p < 0.05, Table S3), and sex had a significant effect on 83.8% of the face phenotypes (4.88 × 10−15 < p < 0.05, Table S3). The effect of BMI was significant on 80.1% of the face phenotypes (1.38 × 10−114 < p < 0.05, Table S3) and most significantly associated with ObiR-ObiL, which is equal to the width of face, as expected.
We selected a total of 125 SNPs at 103 distinct loci that associated with facial features in previous GWASs (Table S4) (Adhikari et al. 2016; Cha et al. 2018; Claes et al. 2018; Cole et al. 2016; Crouch et al. 2018; Lee et al. 2017b; Liu et al. 2012; Paternoster et al. 2012; Pickrell et al. 2016a; Shaffer et al. 2016) and tested their association with 136 facial phenotypes in 612 individuals. We derived 20 PCs from a genomic principal component analysis using the combined dataset including 503 EUR, 504 EAS, and 612 Eurasian individuals. The 1st PC alone accounted for the majority (59.74%) of the total genomic variance explained by all 20 PCs (Figure S2A). K-means clustering of the top 2 PCs clearly differentiated the three populations into separate clusters (Figure S2B). No indications of population sub-structures were detected within the Uyghur individuals. The significance threshold was derived as p < 1.28 × 10−3 using Bonferroni correction, and the effective number of independent variables was estimated as 39 using the matSpD method. The association testing identified eight SNPs displaying significant association with facial phenotypes after adjusting for multiple testing (Table 1). Of these eight SNPs, three were genotyped and five were imputed (Table S4). These included EDAR rs3827760 (min p = 2.39 × 10−5), LYPLAL1 rs5781117 (min p = 1.43 × 10−4), PRDM16 rs4648379 (p = 8.55 × 10−4), PAX3 rs7559271 (p = 7.88 × 10−4), DKK1 rs1194708 (p = 1.77 × 10−3), TNFSF12 rs80067372 (p = 5.90 × 10−4), CACNA2D3 rs56063440 (p = 5.29 × 10−4), and SUPT3H rs227833 (p = 9.89 × 10−4). All eight SNPs together explained up to 6.47% of the sex-, age-, BMI-, and first three genetic PC-adjusted facial phenotype variance (top explained phenotypes: Entocanthion-Otobasion Inferius, Table S5, Fig. 2a). Sex-stratified analysis did not reveal any sex-specific association (Table S6), and more significant association was observed in males than in females, likely explained by the larger sample size of males.
The genetic effects on facial morphology in 612 Eurasian individuals. a Face map depicting the percentage of facial phenotype variance (R2) explained by eight facial shape associated SNPs: including EDAR rs3827760, LYPLAL1 rs5781117, PRDM16 rs4648379, PAX3 rs7559271, DKK1 rs1194708, TNFSF12 rs80067372, CACNA2D3 rs56063440, and SUPT3H rs227833. b Face map denoting the significance (− log10P) level for the associations between EDAR rs3827760 and facial phenotypes, as well as the direction of the genetic effect. c Face map denoting the significance (− log10P) level for the associations between LYPLAL1 rs5781117 and facial phenotypes, as well as the direction of the genetic effect
The strongest association signal was observed for EDAR rs3827760, which showed significant association with eight facial phenotypes (Fig. 2b). The derived G allele demonstrated significant length-increasing effects on eight facial phenotypes belonging to two distinct clusters, including the eye-otobasion distances (ExR-ObiR, EnR-ObiR, EnL-ObiL, ExL-ObiL, and N-ObiR, 2.39 × 10−5 < p < 9.54 × 10−4) and the distances between nosewing and center of mouth (AlR-Sto, AlL-Sto, and AlR-Li, 2.05 × 10−4 < p < 1.24 × 10−3, Figure S3). This allele was highly polymorphic in Eurasians (fUYG = 0.35) and East Asians (fEAS = 0.87), but nearly non-polymorphic in Europeans (fEUR = 0.01) and Africans (fAFR = 0.01) (Figure S4A). Rs3827760 is known as an East-Asian specific variant and has been repeatedly reported to be subjective under positive selection in East Asians (Grossman et al. 2013; Sabeti et al. 2007). Rs3827760 alone explained 2.86% of sex-, age-, BMI-, and the first three genetic PC-adjusted ExL-ObiL variance. The second significant signal belonged to LYPLAL1 rs5781117. Its ancestral T allele displayed significant length-decreasing effects on seven facial distances between otobasion and other landmarks (AlR-ObiL, AlL-ObiR, EnL-ObiL, N-ObiL, AlL-ObiL, EnR-ObiR, and ExL-ObiR, 1.43 × 10−4 < p < 1.13 × 10−3). These facial phenotypes belong to the one facial phenotype cluster that (Figure S3) was also characterized by the distances between the otobasion inferius and other facial landmarks (1.43 × 10−4 < p < 1.13 × 10−3, Fig. 2c). This allele is polymorphic in Africans (fAFR = 0.52) and Europeans (fEUR = 0.34) and minor in East Asians (fEAS = 0.18) (Figure S4B). Rs59156997 explained 2.36% of sex-, age-, BMI-, and the first three genetic PC-adjusted AlR-ObiL variance. The other six SNPs were significantly associated with only one facial phenotype (Table 1, Table S7). The effect alleles of these six SNPs also showed substantial frequency differences between European and East Asian populations, as illustrated using samples from the 1000 Genomes Project (Figure S4). DKK1 rs1194708 especially demonstrated a reversed allele frequency distribution between East Asians and Europeans.
Discussion
In an admixed Eurasian population, we identified eight SNPs (EDAR rs3827760, LYPLAL1 rs5781117, PRDM16 rs4648379, PAX3 rs7559271, DKK1 rs1194708, TNFSF12 rs80067372, CACNA2D3 rs56063440, and SUPT3H rs227833) that were significantly associated with facial features. Together, they explained a considerable proportion of facial variation. EDAR and LYPLAL1 gene variants demonstrated large effects on facial morphology in the Eurasian population, and these effects are likely further pronounced in other East Asian populations. These findings bridged the gap between European and Asian populations in terms of the genetic basis of facial shape variation.
All of the eight face associated SNPs showed significant allele frequency differences between different continental groups and four of them (rs4648379, rs3827760, rs7559271 and rs1194708) showed an inversed allele frequency between Europeans and East Asians, emphasizing population heterogeneity as a key feature underlying the genetic architecture of human facial variation. Recent population genetic studies on human nose morphology have demonstrated that climate changes have significantly contributed to the evolution of the human face (Wroe et al. 2018; Zaidi et al. 2017). The observation of the large allele frequency differences in our study is in line with the previous findings and supports the hypothesis that climatic adaptation and natural selection have shaped the human face during the history of evolution.
The most significant finding was EDAR rs3827760. EDAR encodes a cell-surface receptor important for the development of ectodermal tissues, including skin. rs3827760 is a missense variant (V370A) that affects protein activity (Bryk et al. 2008; Mou et al. 2008), and the derived G allele is associated with several ectodermal-derived traits such as chin protrusion (Adhikari et al. 2016), increased hair straightness (Tan et al. 2013) and thickness (Fujimoto et al. 2008a; Fujimoto et al. 2008b), teeth single and double incisors shoveling (Kimura et al. 2009; Park et al. 2012), increased earlobe attachment, decreased earlobe size, decreased ear protrusion, and decreased ear helix rolling (Adhikari et al. 2015; Shaffer et al. 2017). Previous population genetics studies repeatedly suggested that EDAR has undergone strong positive selection in East Asia populations (Adhikari et al. 2016; Grossman et al. 2010; Kamberov et al. 2013; Sabeti et al. 2007). In our Eurasian sample, the EDAR rs3827760 G allele was significantly associated with increases in eight facial landmark distances and showed a pronounced effect on eye-otobasion distances. This finding is consistent with an Asian-specific and pleiotropic effect of rs3827760. A previous facial shape GWAS in Latin Americans reports that rs3827760 explains 1.32% of chin protrusion variance (Adhikari et al. 2016). In this study of Eurasians, we did not quantify chin protrusion, but rs3827760 explained a considerably larger proportion (2.86%) of the phenotypic variance for a different facial phenotype, i.e., the eye-otobasion distance. Because the G allele is nearly absent (~ 0.01) in Europeans and Africans (~ 0.01), highly frequent in our Eurasian study population (~ 0.35), and abundant in East Asians (~ 0.87), we expect the effect of EDAR on facial variation is even more pronounced in East Asian populations.
Rs5781117 is close to the LYPLAL1 (Lysophospholipase Like 1) gene, which is a protein coding gene. Gene ontology (GO) (Gene Ontology 2015) annotations related to this gene include hydrolase activity and lysophospholipase activity. The ancestral T allele of rs5781117 has been previously associated with an increase in nose size (Pickrell et al. 2016a). Gene variants in this region are also associated with the waist-hip ratio (Heid et al. 2010), obesity (Lv et al. 2017; Nettleton et al. 2015), and adiposity and fat distribution in different populations (Hotta et al. 2013; Lindgren et al. 2009; Liu et al. 2014; Wang et al. 2016). This may suggested that LYPLAL1 slightly affects facial phenotypes by impacting fat distribution. Although we did not ascertain the nose size ordinal phenotype in the current study, the LYPLAL1 rs5781117 SNP was significantly associated with a good number of facial phenotypes, with a pronounced effect on distances between the otobasion inferius and several other facial landmarks (including two nose landmarks) and explained a considerable proportion of the phenotypic variance (up to 2.36% for AlR-ObiL). rs59156997 is highly polymorphic in all continental groups, suggesting a rather universal effect on a variety of facial traits.
The other six SNPs (PRDM16 rs4648379, PAX3 rs7559271, DKK1 rs1194708, TNFSF12 rs80067372, CACNA2D3 rs56063440, and SUPT3H rs227833) were each only significantly associated with one facial trait. Two previous GWASs report that the ancestral A allele of PAX3 rs7559271 has a significant effect on a decreased nasion to mid-endocanthion point distance (Adhikari et al. 2016; Paternoster et al. 2012) in European and Latin American populations. The other two variants including CACNA2D3 rs56063440 and SUPT3H rs227833 are associated with the nose (nose size and nose area) (Claes et al. 2018; Pickrell et al. 2016a). The PRDM16 variant rs4648379 is reported to be associated with a decreased pronasale to left alare distance (Liu et al. 2012). Both the DKK1 variant rs1194708 and TNFSF12 variant rs80067372 are associated with chin dimples (Pickrell et al. 2016a). In our study of Eurasians, although the genetic association survived multiple testing correction, the associated traits did not exactly match the previous GWAS findings. Here, the effect of PAX3 rs7559271 was on the distance between the left entocanthion and right alare, the effect of PRDM16 rs4648379 was on the distance between the subnasale to right cheilion, the effect of DKK1 rs1194708 was on the width of the nosewing, the effect of TNFSF12 rs80067372 was on the subnasale to right ectocanthion, the effect of CACNA2D3 rs56063440 was on the right otobasion inferius to left otobasion inferius, and the effect of SUPT3H rs227833 was on the right entocanthion to left otobasion inferius. This may be explained by genetic effects on multiple facial traits, and further validations of these effects in East Asian populations are warranted. In addition, we note that the small sample size of females is a limit of the current study. Excluding these female samples showed little effect on the detected associations and did not change our conclusions. Although the sex-stratified analysis did not reveal any sex-specific association, and previous GWASs did not report any sex-specific effects of the highlighted SNPs, the effects of these SNPs in Eurasian females warrant further investigations in future studies.
A clustering analysis of the 136 facial phenotypes resulted in four clusters. These clusters followed certain anthropological patterns. The 1st two clusters of the facial phenotypes were in line with the horizontal and vertical facial variations, respectively. The 3rd cluster mainly contained the facial phenotypes involving the otobasion landmark. The 4th cluster mainly involve the phenotypes explaining the variation in the lower part of the face. It is reasonable to speculate that phenotypes in the same cluster may share more or stronger genetic factors than those in different clusters, and genetic factors involved in early stages of facial development may affect more facial phenotype across different clusters. For example, the missense variant rs3827760 of EDAR, which showed significant association with multiple facial phenotypes belonging to two different phenotype clusters, plays an important role in the early embryonic ectoderm development of mice (Kamberov et al. 2013).
Change history
30 August 2021
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1007/s00439-021-02352-6
References
Adhikari K, Reales G, Smith AJ, Konka E, Palmen J, Quinto-Sanchez M, Acuna-Alonzo V, Jaramillo C, Arias W, Fuentes M, Pizarro M, Barquera Lozano R, Macin Perez G, Gomez-Valdes J, Villamil-Ramirez H, Hunemeier T, Ramallo V, Silva de Cerqueira CC, Hurtado M, Villegas V, Granja V, Gallo C, Poletti G, Schuler-Faccini L, Salzano FM, Bortolini MC, Canizales-Quinteros S, Rothhammer F, Bedoya G, Calderon R, Rosique J, Cheeseman M, Bhutta MF, Humphries SE, Gonzalez-Jose R, Headon D, Balding D, Ruiz-Linares A (2015) A genome-wide association study identifies multiple loci for variation in human ear morphology. Nat Commun 6:7500. https://doi.org/10.1038/ncomms8500
Adhikari K, Fuentes-Guajardo M, Quinto-Sanchez M, Mendoza-Revilla J, Camilo Chacon-Duque J, Acuna-Alonzo V, Jaramillo C, Arias W, Lozano RB, Perez GM, Gomez-Valdes J, Villamil-Ramirez H, Hunemeier T, Ramallo V, Silva de Cerqueira CC, Hurtado M, Villegas V, Granja V, Gallo C, Poletti G, Schuler-Faccini L, Salzano FM, Bortolini MC, Canizales-Quinteros S, Cheeseman M, Rosique J, Bedoya G, Rothhammer F, Headon D, Gonzalez-Jose R, Balding D, Ruiz-Linares A (2016) A genome-wide association scan implicates DCHS2, RUNX2, GLI3, PAX1 and EDAR in human facial variation. Nat Commun 7:11616. https://doi.org/10.1038/ncomms11616
Alkhudhairi TD, Alkofide EA (2010) Cephalometric craniofacial features in Saudi parents and their offspring. Angle Orthod 80:1010–1017. https://doi.org/10.2319/050410-66.1
Bryk J, Hardouin E, Pugach I, Hughes D, Strotmann R, Stoneking M, Myles S (2008) Positive selection in East Asians for an EDAR allele that enhances NF-kappaB activation. PLoS One 3:e2209. https://doi.org/10.1371/journal.pone.0002209
Cha S, Lim JE, Park AY, Do JH, Lee SW, Shin C, Cho NH, Kang JO, Nam JM, Kim JS, Woo KM, Lee SH, Kim JY, Oh B (2018) Identification of five novel genetic loci related to facial morphology by genome-wide association studies. BMC Genomics 19:481. https://doi.org/10.1186/s12864-018-4865-9
Claes P, Roosenboom J, White JD, Swigut T, Sero D, Li J, Lee MK, Zaidi A, Mattern BC, Liebowitz C, Pearson L, Gonzalez T, Leslie EJ, Carlson JC, Orlova E, Suetens P, Vandermeulen D, Feingold E, Marazita ML, Shaffer JR, Wysocka J, Shriver MD, Weinberg SM (2018) Genome-wide mapping of global-to-local genetic effects on human facial shape. Nat Genet 50:414–423. https://doi.org/10.1038/s41588-018-0057-4
Cole JB, Manyama M, Kimwaga E, Mathayo J, Larson JR, Liberton DK, Lukowiak K, Ferrara TM, Riccardi SL, Li M, Mio W, Prochazkova M, Williams T, Li H, Jones KL, Klein OD, Santorico SA, Hallgrimsson B, Spritz RA (2016) Genomewide association study of African children identifies association of SCHIP1 and PDE8A with facial size and shape. PLoS Genet 12:e1006174. https://doi.org/10.1371/journal.pgen.1006174
Cole JB, Manyama M, Larson JR, Liberton DK, Ferrara TM, Riccardi SL, Li M, Mio W, Klein OD, Santorico SA, Hallgrimsson B, Spritz RA (2017) Human facial shape and size heritability and genetic correlations. Genetics 205:967–978. https://doi.org/10.1534/genetics.116.193185
Crouch DJM, Winney B, Koppen WP, Christmas WJ, Hutnik K, Day T, Meena D, Boumertit A, Hysi P, Nessa A, Spector TD, Kittler J, Bodmer WF (2018) Genetics of the human face: identification of large-effect single gene variants. Proc Natl Acad Sci USA 115:E676–E685. https://doi.org/10.1073/pnas.1708207114
Delaneau O, Zagury JF, Marchini J (2013) Improved whole-chromosome phasing for disease and population genetic studies. Nat Methods 10:5–6. https://doi.org/10.1038/nmeth.2307
Fujimoto A, Kimura R, Ohashi J, Omi K, Yuliwulandari R, Batubara L, Mustofa MS, Samakkarn U, Settheetham-Ishida W, Ishida T, Morishita Y, Furusawa T, Nakazawa M, Ohtsuka R, Tokunaga K (2008a) A scan for genetic determinants of human hair morphology: EDAR is associated with Asian hair thickness. Hum Mol Genet 17:835–843. https://doi.org/10.1093/hmg/ddm355
Fujimoto A, Ohashi J, Nishida N, Miyagawa T, Morishita Y, Tsunoda T, Kimura R, Tokunaga K (2008b) A replication study confirmed the EDAR gene to be a major contributor to population differentiation regarding head hair thickness in Asia. Hum Genet 124:179–185. https://doi.org/10.1007/s00439-008-0537-1
Gene Ontology C (2015) Gene ontology consortium: going forward. Nucleic Acids Res 43:D1049–D1056. https://doi.org/10.1093/nar/gku1179
Genomes Project C, Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, Kang HM, Marth GT, McVean GA (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491:56–65. https://doi.org/10.1038/nature11632
Grossman SR, Shlyakhter I, Karlsson EK, Byrne EH, Morales S, Frieden G, Hostetter E, Angelino E, Garber M, Zuk O, Lander ES, Schaffner SF, Sabeti PC (2010) A composite of multiple signals distinguishes causal variants in regions of positive selection. Science 327:883–886. https://doi.org/10.1126/science.1183863
Grossman SR, Andersen KG, Shlyakhter I, Tabrizi S, Winnicki S, Yen A, Park DJ, Griesemer D, Karlsson EK, Wong SH, Cabili M, Adegbola RA, Bamezai RN, Hill AV, Vannberg FO, Rinn JL, Genomes P, Lander ES, Schaffner SF, Sabeti PC (2013) Identifying recent adaptations in large-scale genomic data. Cell 152:703–713. https://doi.org/10.1016/j.cell.2013.01.035
Guo J, Mei X, Tang K (2013) Automatic landmark annotation and dense correspondence registration for 3D human facial images. BMC Bioinformatics 14:232. https://doi.org/10.1186/1471-2105-14-232
Hartigan JA, Wong MA (1979) Algorithm AS 136: a K-means clustering algorithm. J R Stat Soc 28:100–108
Heid IM, Jackson AU, Randall JC, Winkler TW, Qi L, Steinthorsdottir V, Thorleifsson G, Zillikens MC, Speliotes EK, Magi R, Workalemahu T, White CC, Bouatia-Naji N, Harris TB, Berndt SI, Ingelsson E, Willer CJ, Weedon MN, Luan J, Vedantam S, Esko T, Kilpelainen TO, Kutalik Z, Li S, Monda KL, Dixon AL, Holmes CC, Kaplan LM, Liang L, Min JL, Moffatt MF, Molony C, Nicholson G, Schadt EE, Zondervan KT, Feitosa MF, Ferreira T, Lango Allen H, Weyant RJ, Wheeler E, Wood AR, Magic Estrada K, Goddard ME, Lettre G, Mangino M, Nyholt DR, Purcell S, Smith AV, Visscher PM, Yang J, McCarroll SA, Nemesh J, Voight BF, Absher D, Amin N, Aspelund T, Coin L, Glazer NL, Hayward C, Heard-Costa NL, Hottenga JJ, Johansson A, Johnson T, Kaakinen M, Kapur K, Ketkar S, Knowles JW, Kraft P, Kraja AT, Lamina C, Leitzmann MF, McKnight B, Morris AP, Ong KK, Perry JR, Peters MJ, Polasek O, Prokopenko I, Rayner NW, Ripatti S, Rivadeneira F, Robertson NR, Sanna S, Sovio U, Surakka I, Teumer A, van Wingerden S, Vitart V, Zhao JH, Cavalcanti-Proenca C, Chines PS, Fisher E, Kulzer JR, Lecoeur C, Narisu N, Sandholt C, Scott LJ, Silander K, Stark K et al (2010) Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nat Genet 42:949–960. https://doi.org/10.1038/ng.685
Hotta K, Kitamoto A, Kitamoto T, Mizusawa S, Teranishi H, So R, Matsuo T, Nakata Y, Hyogo H, Ochi H, Nakamura T, Kamohara S, Miyatake N, Kotani K, Itoh N, Mineo I, Wada J, Yoneda M, Nakajima A, Funahashi T, Miyazaki S, Tokunaga K, Masuzaki H, Ueno T, Chayama K, Hamaguchi K, Yamada K, Hanafusa T, Oikawa S, Sakata T, Tanaka K, Matsuzawa Y, Nakao K, Sekine A (2013) Replication study of 15 recently published Loci for body fat distribution in the Japanese population. J Atheroscler Thromb 20:336–350
Howie BN, Donnelly P, Marchini J (2009) A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 5:e1000529. https://doi.org/10.1371/journal.pgen.1000529
Howie B, Marchini J, Stephens M (2011) Genotype imputation with thousands of genomes. G3 (Bethesda) 1:457–470. https://doi.org/10.1534/g3.111.001198
Kamberov YG, Wang S, Tan J, Gerbault P, Wark A, Tan L, Yang Y, Li S, Tang K, Chen H, Powell A, Itan Y, Fuller D, Lohmueller J, Mao J, Schachar A, Paymer M, Hostetter E, Byrne E, Burnett M, McMahon AP, Thomas MG, Lieberman DE, Jin L, Tabin CJ, Morgan BA, Sabeti PC (2013) Modeling recent human evolution in mice by expression of a selected EDAR variant. Cell 152:691–702. https://doi.org/10.1016/j.cell.2013.01.016
Kimura R, Yamaguchi T, Takeda M, Kondo O, Toma T, Haneji K, Hanihara T, Matsukusa H, Kawamura S, Maki K, Osawa M, Ishida H, Oota H (2009) A common variation in EDAR is a genetic determinant of shovel-shaped incisors. Am J Hum Genet 85:528–535. https://doi.org/10.1016/j.ajhg.2009.09.006
Lee MK, Shaffer JR, Leslie EJ, Orlova E, Carlson JC, Feingold E, Marazita ML, Weinberg SM (2017a) Genome-wide association study of facial morphology reveals novel associations with FREM1 and PARK2. Plos One. https://doi.org/10.1371/journal.pone.0176566(ARTN e0176566)
Lee MK, Shaffer JR, Leslie EJ, Orlova E, Carlson JC, Feingold E, Marazita ML, Weinberg SM (2017b) Genome-wide association study of facial morphology reveals novel associations with FREM1 and PARK2. PLoS One 12:e0176566. https://doi.org/10.1371/journal.pone.0176566
Li J, Ji L (2005) Adjusting multiple testing in multilocus analyses using the eigenvalues of a correlation matrix. Heredity (Edinb) 95:221–227. https://doi.org/10.1038/sj.hdy.6800717
Lindgren CM, Heid IM, Randall JC, Lamina C, Steinthorsdottir V, Qi L, Speliotes EK, Thorleifsson G, Willer CJ, Herrera BM, Jackson AU, Lim N, Scheet P, Soranzo N, Amin N, Aulchenko YS, Chambers JC, Drong A, Luan J, Lyon HN, Rivadeneira F, Sanna S, Timpson NJ, Zillikens MC, Zhao JH, Almgren P, Bandinelli S, Bennett AJ, Bergman RN, Bonnycastle LL, Bumpstead SJ, Chanock SJ, Cherkas L, Chines P, Coin L, Cooper C, Crawford G, Doering A, Dominiczak A, Doney AS, Ebrahim S, Elliott P, Erdos MR, Estrada K, Ferrucci L, Fischer G, Forouhi NG, Gieger C, Grallert H, Groves CJ, Grundy S, Guiducci C, Hadley D, Hamsten A, Havulinna AS, Hofman A, Holle R, Holloway JW, Illig T, Isomaa B, Jacobs LC, Jameson K, Jousilahti P, Karpe F, Kuusisto J, Laitinen J, Lathrop GM, Lawlor DA, Mangino M, McArdle WL, Meitinger T, Morken MA, Morris AP, Munroe P, Narisu N, Nordstrom A, Nordstrom P, Oostra BA, Palmer CN, Payne F, Peden JF, Prokopenko I, Renstrom F, Ruokonen A, Salomaa V, Sandhu MS, Scott LJ, Scuteri A, Silander K, Song K, Yuan X, Stringham HM, Swift AJ, Tuomi T, Uda M, Vollenweider P, Waeber G, Wallace C, Walters GB, Weedon MN et al (2009) Genome-wide association scan meta-analysis identifies three Loci influencing adiposity and fat distribution. PLoS Genet 5:e1000508. https://doi.org/10.1371/journal.pgen.1000508
Liu F, van der Lijn F, Schurmann C, Zhu G, Chakravarty MM, Hysi PG, Wollstein A, Lao O, de Bruijne M, Ikram MA, van der Lugt A, Rivadeneira F, Uitterlinden AG, Hofman A, Niessen WJ, Homuth G, de Zubicaray G, McMahon KL, Thompson PM, Daboul A, Puls R, Hegenscheid K, Bevan L, Pausova Z, Medland SE, Montgomery GW, Wright MJ, Wicking C, Boehringer S, Spector TD, Paus T, Martin NG, Biffar R, Kayser M (2012) A genome-wide association study identifies five loci influencing facial morphology in Europeans. PLoS Genet 8:e1002932. https://doi.org/10.1371/journal.pgen.1002932
Liu CT, Buchkovich ML, Winkler TW, Heid IM, African Ancestry Anthropometry Genetics C, Consortium G, Borecki IB, Fox CS, Mohlke KL, North KE, Adrienne Cupples L (2014) Multi-ethnic fine-mapping of 14 central adiposity loci. Hum Mol Genet 23:4738–4744. https://doi.org/10.1093/hmg/ddu183
Lv D, Zhou D, Zhang Y, Zhang S, Zhu YM (2017) Two obesity susceptibility loci in LYPLAL1 and ETV5 independently associated with childhood hypertension in Chinese population. Gene 627:284–289. https://doi.org/10.1016/j.gene.2017.06.030
Members BIGDC (2018) Database resources of the BIG data center in 2018. Nucleic Acids Res 46:D14–D20. https://doi.org/10.1093/nar/gkx897
Mou C, Thomason HA, Willan PM, Clowes C, Harris WE, Drew CF, Dixon J, Dixon MJ, Headon DJ (2008) Enhanced ectodysplasin-A receptor (EDAR) signaling alters multiple fiber characteristics to produce the East Asian hair form. Hum Mutat 29:1405–1411. https://doi.org/10.1002/humu.20795
Nettleton JA, Follis JL, Ngwa JS, Smith CE, Ahmad S, Tanaka T, Wojczynski MK, Voortman T, Lemaitre RN, Kristiansson K, Nuotio ML, Houston DK, Perala MM, Qi Q, Sonestedt E, Manichaikul A, Kanoni S, Ganna A, Mikkila V, North KE, Siscovick DS, Harald K, McKeown NM, Johansson I, Rissanen H, Liu Y, Lahti J, Hu FB, Bandinelli S, Rukh G, Rich S, Booij L, Dmitriou M, Ax E, Raitakari O, Mukamal K, Mannisto S, Hallmans G, Jula A, Ericson U, Jacobs DR Jr, Van Rooij FJ, Deloukas P, Sjogren P, Kahonen M, Djousse L, Perola M, Barroso I, Hofman A, Stirrups K, Viikari J, Uitterlinden AG, Kalafati IP, Franco OH, Mozaffarian D, Salomaa V, Borecki IB, Knekt P, Kritchevsky SB, Eriksson JG, Dedoussis GV, Qi L, Ferrucci L, Orho-Melander M, Zillikens MC, Ingelsson E, Lehtimaki T, Renstrom F, Cupples LA, Loos RJ, Franks PW (2015) Gene x dietary pattern interactions in obesity: analysis of up to 68 317 adults of European ancestry. Hum Mol Genet 24:4728–4738. https://doi.org/10.1093/hmg/ddv186
Park JH, Yamaguchi T, Watanabe C, Kawaguchi A, Haneji K, Takeda M, Kim YI, Tomoyasu Y, Watanabe M, Oota H, Hanihara T, Ishida H, Maki K, Park SB, Kimura R (2012) Effects of an Asian-specific nonsynonymous EDAR variant on multiple dental traits. J Hum Genet 57:508–514. https://doi.org/10.1038/jhg.2012.60
Paternoster L, Zhurov AI, Toma AM, Kemp JP, St Pourcain B, Timpson NJ, McMahon G, McArdle W, Ring SM, Smith GD, Richmond S, Evans DM (2012) Genome-wide association study of three-dimensional facial morphology identifies a variant in PAX3 associated with nasion position. Am J Hum Genet 90:478–485. https://doi.org/10.1016/j.ajhg.2011.12.021
Pickrell JK, Berisa T, Liu JZ, Segurel L, Tung JY, Hinds DA (2016a) Detection and interpretation of shared genetic influences on 42 human traits. Nat Genet 48:709–717. https://doi.org/10.1038/ng.3570
Pickrell JK, Berisa T, Liu JZ, Segurel L, Tung JY, Hinds DA (2016b) Detection and interpretation of shared genetic influences on 42 human traits (vol 48, pg 709, 2016). Nat Genet 48:1296
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. https://doi.org/10.1086/519795
Sabeti PC, Varilly P, Fry B, Lohmueller J, Hostetter E, Cotsapas C, Xie X, Byrne EH, McCarroll SA, Gaudet R, Schaffner SF, Lander ES, International HapMap C, Frazer KA, Ballinger DG, Cox DR, Hinds DA, Stuve LL, Gibbs RA, Belmont JW, Boudreau A, Hardenbol P, Leal SM, Pasternak S, Wheeler DA, Willis TD, Yu F, Yang H, Zeng C, Gao Y, Hu H, Hu W, Li C, Lin W, Liu S, Pan H, Tang X, Wang J, Wang W, Yu J, Zhang B, Zhang Q, Zhao H, Zhao H, Zhou J, Gabriel SB, Barry R, Blumenstiel B, Camargo A, Defelice M, Faggart M, Goyette M, Gupta S, Moore J, Nguyen H, Onofrio RC, Parkin M, Roy J, Stahl E, Winchester E, Ziaugra L, Altshuler D, Shen Y, Yao Z, Huang W, Chu X, He Y, Jin L, Liu Y, Shen Y, Sun W, Wang H, Wang Y, Wang Y, Xiong X, Xu L, Waye MM, Tsui SK, Xue H, Wong JT, Galver LM, Fan JB, Gunderson K, Murray SS, Oliphant AR, Chee MS, Montpetit A, Chagnon F, Ferretti V, Leboeuf M, Olivier JF, Phillips MS, Roumy S, Sallee C, Verner A, Hudson TJ, Kwok PY, Cai D, Koboldt DC, Miller RD, et al. (2007) Genome-wide detection and characterization of positive selection in human populations. Nature 449:913–918. https://doi.org/10.1038/nature06250
Shaffer JR, Li J, Lee MK, Roosenboom J, Orlova E, Adhikari K, andMe Research T, Gallo C, Poletti G, Schuler-Faccini L, Bortolini MC, Canizales-Quinteros S, Rothhammer F, Bedoya G, Gonzalez-Jose R, Pfeffer PE, Wollenschlaeger CA, Hecht JT, Wehby GL, Moreno LM, Ding A, Jin L, Yang Y, Carlson JC, Leslie EJ, Feingold E, Marazita ML, Hinds DA, Cox TC, Wang S, Ruiz-Linares A, Weinberg SM (2017) Multiethnic GWAS reveals polygenic architecture of earlobe attachment. Am J Hum Genet 101:913–924. https://doi.org/10.1016/j.ajhg.2017.10.001
Shaffer JR, Orlova E, Lee MK, Leslie EJ, Raffensperger ZD, Heike CL, Cunningham ML, Hecht JT, Kau CH, Nidey NL, Moreno LM, Wehby GL, Murray JC, Laurie CA, Laurie CC, Cole J, Ferrara T, Santorico S, Klein O, Mio W, Feingold E, Hallgrimsson B, Spritz RA, Marazita ML, Weinberg SM (2016) Genome-wide association study reveals multiple loci influencing normal human facial morphology. PLoS Genet 12:e1006149. https://doi.org/10.1371/journal.pgen.1006149
Song S, Tian D, Li C, Tang B, Dong L, Xiao J, Bao Y, Zhao W, He H, Zhang Z (2018) Genome variation map: a data repository of genome variations in BIG data center. Nucleic Acids Res 46:D944–D949. https://doi.org/10.1093/nar/gkx986
Tan J, Yang Y, Tang K, Sabeti PC, Jin L, Wang S (2013) The adaptive variant EDARV370A is associated with straight hair in East Asians. Hum Genet 132:1187–1191. https://doi.org/10.1007/s00439-013-1324-1
Wang T, Ma X, Peng D, Zhang R, Sun X, Chen M, Yan J, Wang S, Yan D, He Z, Jiang F, Bao Y, Hu C, Jia W (2016) Effects of obesity related genetic variations on visceral and subcutaneous fat distribution in a Chinese population. Sci Rep 6:20691. https://doi.org/10.1038/srep20691
Wroe S, Parr WCH, Ledogar JA, Bourke J, Evans SP, Fiorenza L, Benazzi S, Hublin JJ, Stringer C, Kullmer O, Curry M, Rae TC, Yokley TR (2018) Computer simulations show that Neanderthal facial morphology represents adaptation to cold and high energy demands, but not heavy biting. Proc Biol Sci. https://doi.org/10.1098/rspb.2018.0085
Zaidi AA, Mattern BC, Claes P, McEvoy B, Hughes C, Shriver MD (2017) Investigating the case of human nose shape and climate adaptation. PLoS Genet 13:e1006616. https://doi.org/10.1371/journal.pgen.1006616
Acknowledgements
This project was supported by the National Key R&D Program of China (2017YFC0803501), National Natural Science Foundation of China (91651507), Fund from institute of forensic science (2018JB046), and open projects of the National Engineering Laboratory for Forensic Science (2017NELKFKT05). Biological samples were provided by the National Science and Technological Resources Platform (YCZYPT[2017]01-3 and 2017JB025). Author FL is supported by “The Thousand Talents Plan for Young Professionals”. Author CXL is supported by “The Beijing Leading Talent Program (Z18110006318006)”.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest regarding this work.
Data Availability Statement
The dataset analyzed during the current study is restricted due to participant confidentially. Restrictions apply to the availability of these data, which were used under license for the current study, and so are not entirely publicly available. However, the variation data reported in this paper have been deposited in the Genome Variation Map (Song et al. 2018) in the BIG Data Center (Members BIGDC 2018), Beijing Institute of Genomics (BIG), Chinese Academy of Sciences, under accession number GVM000031 and can be publicly accessed at http://bigd.big.ac.cn/gvm/getProjectDetail?project=GVM000031.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article has been retracted. Please see the retraction notice for more detail:https://doi.org/10.1007/s00439-021-02352-6
Electronic supplementary material
Below is the link to the electronic supplementary material.
439_2019_2023_MOESM2_ESM.tif
Supplementary Figure 2: (A) The relative proportion of variance of the first 20 PCs derived from genotyped data, including our 612 Eurasian samples and the EAS and EUR samples from the 1000 Genomes Project. (B) The PC1 and PC2 plots of genotyped data, including our 612 Eurasian samples and the EAS and EUR samples from the 1000 Genomes Project (TIFF 22719 kb)
Supplementary Figure 3: The hierarchical clustering results of 136 facial phenotypes (TIFF 13680 kb)
439_2019_2023_MOESM4_ESM.tif
Supplementary Figure 4: Worldwide allele-frequency distributions for the eight identified variants. Map displaying the geospatial distribution of the allele frequency of rs3827760 (EDAR), rs5781117 (LYPLAL1), rs4648379 (PRDM16), rs7559271 (PAX3), rs1194708 (DKK1), rs80067372 (TNFSF12), rs56063440 (CACNA2D3), and rs227833 (SUPT3H) across the world. The map is drawn based on allele frequencies of 2504 subjects obtained from 1000 Genome project datasets for each SNP. The pie denotes the sampling locations (TIFF 34735 kb)
About this article
Cite this article
Li, Y., Zhao, W., Li, D. et al. RETRACTED ARTICLE: EDAR, LYPLAL1, PRDM16, PAX3, DKK1, TNFSF12, CACNA2D3, and SUPT3H gene variants influence facial morphology in a Eurasian population. Hum Genet 138, 681–689 (2019). https://doi.org/10.1007/s00439-019-02023-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-019-02023-7