Abstract
The c-Jun N-terminal kinases (JNKs) are stress-activated serine-threonine kinases that have recently been linked to various neurological disorders. We previously described a patient with intellectual disability (ID) and seizures (Patient 1), carrying a de novo chromosome translocation affecting the CNS-expressed MAPK10/JNK3 gene. Here, we describe a second ID patient (Patient 2) with a similar translocation that likewise truncates MAPK10/JNK3, highlighting a role for JNK3 in human brain development. We have pinpointed the breakpoint in Patient 2, which is just distal to that in Patient 1. In both patients, the rearrangement resulted in a predicted protein interrupted towards the C-terminal end of the kinase domain. We demonstrate that these truncated proteins, although capable of weak interaction with various known JNK scaffolds, are not capable of phosphorylating the classical JNK target c-Jun in vitro, which suggests that the patient phenotype potentially arises from partial loss of JNK3 function. We next investigated JNK3-binding partners to further explore potential disease mechanisms. We identified PSD-95, SAP102 and SHANK3 as novel interaction partners for JNK3, and we demonstrate that JNK3 and PSD-95 exhibit partially overlapping expression at synaptic sites in cultured hippocampal neurons. Moreover, JNK3, like JNK1, is capable of phosphorylating PSD-95 in vitro, whereas disease-associated mutant JNK3 proteins do not. We conclude that reduced JNK3 activity has potentially deleterious effects on neuronal function via altered regulation of a set of post-synaptic proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The c-Jun N-terminal kinases, or JNKs (also sometimes referred to as stress-activated kinases, or SAPKs), belong to the mitogen-activated protein kinase (MAPK) family of signalling molecules and are the terminal kinases within one branch of the MAP kinase cascade. There are three highly conserved JNK genes in mammals (formally referred to as MAPK8, MAPK9 and MAPK10), and a total of 10 isoforms, with overlapping functions, have been described. JNK 1 and 2 isoforms are widely expressed and essential for early development and differentiation. Unlike JNKs 1 and 2, JNK3 exhibits restricted expression, primarily in the central nervous system (Martin et al. 1996). While early studies centered on the role of active JNKs in apoptosis and differentiation and in the cellular stress response in non-neural systems, more recent work implicates JNK signalling in neuronal functions. For example, numerous studies demonstrate that a hyper-phosphorylation of JNK is associated with Alzheimer’s and other neurodegenerative diseases and provide the basis for research on the use of JNK inhibitors as potential therapeutic agents [for review see (Mehan et al. 2011)]. However, it is important to acknowledge that there is a high basal activation of JNKs in neuronal cultures (Bjorkblom et al. 2005; Xu et al. 1997), suggesting that they execute important functions in neurons also under physiological, non-stress conditions. In line with this, JNK signalling aberrations have recently been observed as part of the pathophysiology of early-onset cognitive disorders. For example, the IL1RAPL1 gene has been implicated in monogenic forms of intellectual disability and autism in several unrelated patients, and recent characterisation of the IL1RAPL1 knockout mouse illuminated disease-associated alterations in the JNK signalling cascade (Pavlowsky et al. 2010a, b). These studies, paralleled by our identification of a de novo MAPK10/JNK3 (OMIM 602897) truncation mutation in a patient with a severe neurodevelopmental disorder (Shoichet et al. 2006), and the fact that this gene is among those affected by a rearrangement in another patient with a complex physical and cognitive disorder (Baptista et al. 2008), make clear that JNKs—in particular the CNS-expressed MAPK10/JNK3—are candidates for mutation identification in patients with early-onset neurological disorders. Here, we describe the clinical and genetic characterisation of an unrelated patient with a cognitive phenotype harbouring a de novo chromosome rearrangement affecting the MAPK10/JNK3 gene. We have investigated the nature of the mutation and associated disease phenotype with respect to the previously characterised patient, and we have further investigated JNK3 function. Importantly, we show that PSD-95 is a novel JNK3 substrate and interaction partner, and we demonstrate that other disease-associated synaptic scaffold proteins, namely SAP102 and SHANK3, are also capable of interacting with JNK3. In summary, this study provides critical evidence supporting the idea that defective JNK signalling is detrimental for brain function, and it highlights potential disease mechanisms.
Materials and methods
Patient description and genetic characterisation
The patient (Patient 2) was born after an uneventful pregnancy to healthy and non-consanguineous parents. Psychomotor development was delayed, and upon examination at the age of 13 years, he had mild intellectual disability. Body measurements were normal, and he had neither neurological disorders nor dysmorphic features. Brain MRI scan was normal. Routine chromosome analysis revealed a de novo balanced translocation 46,X,t(Y;4)(q12;q21.3), and whole genome array CGH (resolution 100 kb) detected no pathogenic gains or losses. For breakpoint mapping, we did array painting essentially as described in previous studies (Kalscheuer et al. 2007). Briefly, EBV-transformed patient lymphoblastoid cell lines were cultured in RPMI 1640 medium supplemented with 10 % fetal calf serum, 2 mM l-glutamine and antibiotics. Cells in log phase were treated for 16 h with colcemid (0.05 mg/mL final concentration), and metaphase chromosomes were then isolated and flow-sorted as described previously (Arkesteijn et al. 1999).
Approximately 6,000 flow-sorted chromosomes were used directly for amplification with the GenomiPhi V2 DNA Amplification Kit (GE Healthcare) according to the manufacturer’s recommendations. One microgram of amplified DNA from the derivative chromosome was labelled via Agilent’s Genomic DNA Enzymatic Labeling Kit Plus (Agilent). To each labelling reaction, 100 ng of genomic control DNA was added to ensure proper placement of the grid for subsequent image analysis. At first, these probes were hybridised to a SurePrint G3 Human CGH 400 K whole genome oligonucleotide array. Based on the results of this first hybridisation, eArray (Agilent) was used to design a custom array for high-resolution breakpoint mapping, consisting of 4,414 oligonucleotides evenly spaced across the genomic interval chr4: 86,999,606–87,024,331 (GRCh37/HG19). All hybridisations were done according to the manufacturer’s recommendations for array CGH experiments (Agilent). Further analysis and visualisation of array painting data were done using the array CGH software package CGHPRO (Chen et al. 2005).
Constructs
The open reading frame (ORF) of human JNK3a2 (NM_138982, 1,395 bp) was amplified from human cDNA using the primers JNK3a2-Bgl-fw (ATT AGA TCT GCC ACC ATG AGC CTC CAT TTC TTA TAC) and JNK3a2-Xma-rv (A TAC CCG GGC TGC AAC AAC CCA GGG GTC). The mutant JNK3 variants (reflecting those present in Patients 1 and 2) were amplified from human cDNA using the forward primer JNK3a2-Bgl-fw and for Patient 1 (NM_138982, exon 1 to exon 9 inclusive; cDNA coding nucleotides 1–801) the reverse primer JNK3a2-800-Xma-rv (ATA CCC GGG TCC CTT CCT GGA AAG AG); for Patient 2 (NM_138982, exon 1 to exon 8 inclusive; cDNA coding nucleotides 1–730) the same forward primer was used with the reverse primer JNK3a2-730-Xma-rv (ATA CCC GGG CGT TCT CCT TGT AGC CCA TC). PCR products were cloned with BglII and XmaI into the expression vector pEGFP-C1 (Clontech). The inactive kinase mutant pEGFP-C1-JNK3-221APF was generated by site-directed mutagenesis using the primers JNK3-221APF223-HindIIIf (TPY221APF mutation: CAC AAG CTT CAT GAT GGC TCC ATT TGT GGT GAC ACG TTA T) and JNK3a2-Xma-rv. The PCR product was cloned with HindIII and XmaI into the vector pEGFP-C1-JNK3. The pEGFP-C1-JNK1 clone was generated by amplification of the human JNK1 coding sequence NM139049 (from Addgene clone # 13751) using the primers JNK1a2-BglII-for CTC AGA TCT ATG AGC AGA AGC AAG CGT G and JNK1a2-SalI-rev CCG TCG ACT CAT CTA CAG CAG CCC AGA G, and cloned into pEGFP-C1 (Clontech). PSD-95 coding sequence (based on NM_019621; 2,175 bp) was amplified from a rat cDNA clone and subcloned into pCMV-Tag2A (Stratagene) using EcoRI and SalI restriction sites. The N-terminal GST-PSD-95 fusion construct was subsequently generated from this clone using an N-terminal FLAG primer FLAG-Sal-for TTG TCG ACA AAT GGA TTA CAA GGA TGA C and C-terminal primer DLG4-PDZ1-3-NotI-rev AAG CGG CCG CTC ATC GAC TAT ACT CTT CTG G. This fragment, encoding an N-terminal FLAG tag and the first 399 amino acids of PSD-95, was cloned into the pGEX-6P-1 vector with an N-terminal GST tag (GE Healthcare). SAP102 coding sequence (NM_021120) was amplified from a human cDNA library and subcloned into pCMV-Tag2A (Stratagene) using EcoRI and SalI restriction sites. The expression construct pBud-beta-Arrestin2-V5, harbouring the human beta-Arrestin2 transcript (NM_004313), was cloned using KpnI and BglII into the pBudCE4.1 mammalian expression vector (Invitrogen). The Shank3-HA, c-Jun-HA, and pCMV5-FLAG-JIP1b constructs were generous gifts from Morgan Sheng, Anna Maria Musti, and Alan Whitmarsh, respectively.
Cell culture and transfection experiments
COS-7 cells were maintained in DMEM (Lonza) supplemented with 10 % FBS (Sigma), 2 mM l-glutamine and penicillin/streptomycin at 37 °C with 5 % CO2. Transient transfections were done using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s recommendations.
Kinase assays
Kinases were either commercially available (BioMol) or overexpressed in COS-7 cells, harvested in lysis buffer [20 mM Tris, pH 7.5, 150 mM NaCl, 1 % Triton X and Mini Complete Inhibitors without EDTA (Roche)], cleared by centrifugation at 20,000×g and immunoprecipitated with 2 μg GFP antibody (anti-GFP, Roche) as described in detail in the next section. Comparative kinase reactions with wild-type and mutant JNK3 immunoprecipitates were performed by adding 25 μl kinase reaction buffer (20 mM Tris pH 7.5, 10 mM MgCl2, 200 μM ATP, 0.1 % beta-mercaptoethanol, and approximately 2 μg GST-c-Jun (MBL/Biozol) or GST-PSD-95 as a substrate) to the beads of the immunoprecipitated kinase variants for 30 min at 30 °C. For c-Jun kinase assays, we added 1 mM EDTA and phosphatase inhibitors (HALT, Thermo Scientific) to the lysis buffer. c-Jun phosphorylation was analysed by standard SDS-PAGE followed by western blotting with a phospho-c-Jun antibody. PSD-95 phosphorylation was assessed using the PhosTAG system (Wako), which facilitates phosphorylation-dependent retarded protein migration through a polyacrylamide gel, and subsequent western blot with the FLAG antibody. PhosTAG gel electrophoresis (6 % polyacrylamide supplemented with 75 μM PhosTAG) was established using an optimised protocol with Zn2+ in a Bis–Tris-buffered neutral-pH gel system as described previously (Kinoshita and Kinoshita-Kikuta 2011).
Coimmunoprecipitation studies
Kinase variants were overexpressed in COS-7 cells together with their candidate interaction partner as described above. 24 h post-transfection cells were harvested in immunoprecipitation (IP) buffer [50 mM Tris pH 7.5, 100 mM NaCl, 0.1 % NP40 and Mini Complete Inhibitors (Roche)] and lysed with 10–15 strokes using a 30G needle. Lysates were cleared by two centrifugations for 10 min at 20,000×g. For IP, lysates were incubated for 3 h at 4 °C with 2 μg of the respective antibody, and transfected proteins were pulled down with Protein-G agarose for 1 h. Beads were washed 3 times for 5 min at 4 °C. Samples were analysed by SDS-PAGE and western blot. For brain lysate IPs, adult C57Bl6 mice were killed by cervical dislocation; whole brains were removed and placed directly in ice-cold TN buffer [50 mM Tris, pH 7.5, 100 mM NaCl, and Mini Complete Inhibitors without EDTA (Roche)] and homogenised. Triton-X was added to the lysate (final concentration of 1 %) and incubated on ice for 15 min. TN buffer was added to generate a final Triton-X concentration of 0.5 %. Cell debris was removed by high-speed centrifugation (20,000×g), and lysates pre-cleared for 1.5 h with Protein-G agarose (Roche). 2 μg of JNK3 mouse monoclonal antibody (anti-MAPK10, Acris, AM06230SU-N) was used for overnight immunoprecipitation. Following subsequent centrifugation, 20 μL of Protein G agarose was used for pull-down (1 h), and beads were washed 3 × 10 min at 4 °C. Samples were analysed by SDS-PAGE and western blot.
Primary neuronal cultures
For primary rat hippocampal neuronal cultures, newborn (P0) Wistar Unilever rats were used. Following decapitation, hippocampi from 5 to 10 pups were isolated and collected in ice-cold DMEM (Lonza). Tissue was partially digested in Trypsin/EDTA (Lonza) at 37 °C for 5 min. After stopping the reaction with 10 % FBS (Biochrom) in DMEM and subsequent washing in DMEM to remove traces of trypsin, the hippocampal tissue was suspended in neuron culture medium (Neurobasal A supplemented with B27 and 0.5 mM glutamine) and further dissociated mechanically. Neurons were plated onto glass coverslips coated with a mixture of poly-d-lysine (Sigma) and Laminin (Sigma) at a density of ~105 cells/cm2. Cell debris was removed after healthy neurons adhered to the cover slips (~1 h post-plating), and neurons were maintained for 2–5 weeks with minimal perturbation at 37 °C with 5 % CO2 in neuron culture medium. All animals used in this study were treated according to the German regulations on the use of animals for research purposes and killed under the permit T0280/10.
Immunocytochemistry
After 2–5 weeks in culture, differentiated neurons were fixed in 4 % PFA in phosphate-buffered saline (PBS) for 10 min and immunofluorescence was performed as described previously (Kunde et al. 2011) according to standard IF protocols. Briefly, cells were permeabilised in 0.2 % Triton-X in PBS for 5 min and blocked with 4 % bovine serum albumin (BSA) in PBS for 1 h at room temperature. They were then incubated overnight with the primary antibodies in the same solution at 4 °C, washed with PBS and subsequently incubated with secondary antibodies in blocking solution. After final washes in PBS, coverslips were dipped in water and mounted with Fluoromount-G (Southern Biotech). Images were acquired with a Leica laser-scanning confocal microscope (Leica TCS-SP5 II) using a 63× objective.
GST pull-down assays and protein purification
GST fusion proteins were purified essentially according to the manufacturer’s recommended protocol (GE Healthcare). Briefly, BL21 Escherichia coli harbouring the GST fusion constructs were cultured O/N and recombinant protein expression was induced with 0.1 mM IPTG. Cells were suspended in Tris-buffered saline (TBS) and lysed by sonication plus 1 % Triton-X. Lysates were cleared by centrifugation at 20,000×g and GST fusion proteins were purified using glutathione agarose (Pierce) and eluted with 50 mM glutathione in 200 mM Tris (pH 8). They were desalted using a Zeba Spin Desalting Column (40 K MWCO Thermo Scientific).
Antibodies
Antibodies used in this study include anti-phospho-c-Jun (CST, rabbit, WB 1:2,000), anti-c-Jun (CST, rabbit, WB 1:2,000), anti-JNK3 (Upstate/Millipore 04-893, WB 1:5,000; IF 1:500), anti-JNK3 (Acris, mouse, AM06230SU-N, IP 2 μg), anti-PSD-95 (NeuroMab, #75-028, mouse, WB 1:2,000; IF 1:500), anti-GFP (Roche, mouse, IP 2 μg), anti-GFP (Abcam, goat, WB 1:5,000), anti-FLAG (Sigma, rabbit, WB 1:5,000), anti-V5 (Millipore, rabbit, WB 1:5,000), anti-HA (Covance, mouse, WB 1:5,000), anti-GST (GE Healthcare, goat, WB 1:5,000), anti-mouse-HRP (Dianova, WB 1:5,000), anti-goat-HRP (SantaCruz, WB 1:5,000) and anti-rabbit-HRP (Dianova, WB 1:5,000). All primary and secondary antibodies were diluted in 5 % milk/PBST for WB, apart from anti-phospho-c-Jun, which was diluted in 4 % BSA/TBST, as well as the following anti-rabbit-HRP. Unspecific mouse IgGs, as required (SantaCruz, 2 μg), were used for negative control coimmunoprecipitation studies.
Results
Related de novo MAPK10/JNK3 truncation mutations in patients with neurodevelopmental disturbances
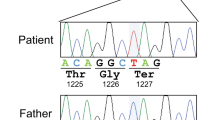
We previously reported on a patient with severe cognitive and developmental delay carrying a balanced translocation affecting the c-Jun N-terminal kinase JNK3 (Shoichet et al. 2006). On the derivative chromosome 4, the breakpoint interrupts the MAPK10/JNK3 gene between exons 9 and 10, and the predicted truncated JNK3, presumably expressed predominantly in neural tissue, harbours 267 of 464 amino acids (Shoichet et al. 2006). Here, we describe a second patient carrying a similar chromosome rearrangement involving the same region of chromosome 4 and presenting with intellectual disability (ID). Routine karyotyping revealed a de novo balanced Y;4 translocation (depicted in Fig. 1a). Using a whole-genome oligonucleotide 400 k array, the chromosome breakpoint was localised within chr4: 87,019,606–87,022,331 (depicted schematically in Fig. 1b). Fine-mapping employing a custom array covering this region with high density (Fig. 1c) further narrowed down the breakpoint to chr4: 87,019,870–87,019,980 (GRCh37/HG19), just distal to exon 9 (coding exon 7), i.e. the breakpoint in this patient falls between exons 8 and 9 (coding exons 6 and 7) of MAPK10/JNK3, which is on the reverse strand of chromosome 4, and thereby interrupts the coding sequence just after nucleotide 730. The two truncation mutations affect essentially the same region of the predicted protein, i.e. they both lie towards the C-terminal end of the conserved kinase domain (see comparative schematic diagram, Fig. 2a); however, the predicted truncated proteins differ by 24 amino acids. These genetic data strongly support the idea that JNK signalling is of essential importance for normal brain function.
Fine-mapping of the chromosome 4 translocation breakpoint by array painting. a Ideogram depicting translocation chromosomes from the ID patient described (referred to throughout the text as Patient 2) with karyotype 46,X,t(Y;4)(q12;q21.3). b Graphical representation of array painting results with the 400 k SurePrint G3 Human CGH Microarray (Agilent) following hybridisation of DNA from flow-sorted derivative chromosome 4 from Patient 2. Data is depicted with the software CGHPRO, in which each DNA probe is plotted as a dark red spot to the right of the ideogram of chromosome 4 according to its genomic position and its ratio of Cy3 to Cy5 signal intensities (see “Materials and methods” section for details). The abrupt ratio shift from left to right indicates the translocation breakpoint. c Depiction of results following high-resolution analysis of the same translocation chromosome employing a custom eArray (Agilent) with an average oligo spacing of less than 100 bp, also viewed using CGHPRO. To the right is the expanded view of oligos mapping to the breakpoint region
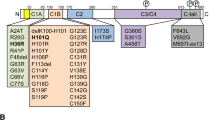
a Schematic diagram indicating the intron–exon structure of the MAPK10/JNK3 coding sequence with respect to the conserved serine-threonine kinase domain. The locations of the intronic breakpoints (BP) found in Patients 1 and 2 (P1 and P2) are indicated; transcripts are labelled as wild-type (wt) or truncated (tr). b In vitro kinase assay in which the abilities of wild-type (WT) and mutant JNK3 proteins to phosphorylate the classical JNK substrate c-Jun are compared. Following overexpression and pull-down of JNK variants and controls from COS-7 cell lysates, phosphorylated c-Jun is observed by western blot using an antibody specific for phosphorylated c-Jun (anti-ph-c-Jun). Presence of non-phosphorylated c-Jun substrate is validated using a c-Jun antibody on the same lysates (middle panel), and comparable pull-down of kinase variants is validated by WB with an anti-GFP antibody. WT refers to the wild-type EGFP-JNK3 construct, APF to the EGFP-JNK3 phospho-site mutant, and P1 and P2 to the patient-associated variants. Negative controls include pull-down following overexpression of EGFP alone (EGFP), and beads alone (beads), in which the kinase reaction was performed in the absence of cell lysate
Disease-associated JNK3 mutant proteins are not active kinases
Given the fact that both patients described here are heterozygous mutation carriers with one wild-type allele and a truncated JNK3 transcript that is potentially subject to nonsense-mediated decay, it is reasonable to propose that partial loss of JNK3 function might be the cause of the observed cognitive defects. To determine if loss-of-function can be expected, we first tested the mutant JNK3, which is potentially expressed as a truncated protein in the patient CNS, in classical kinases assays. We made use of both disease-associated recombinant JNK3 proteins (herewith referred to as JNK3 truncations P1 and P2), which, respectively, harbour the N-terminal 267 and 243 amino acids of human JNK3 (see schematic diagram in Fig. 2a). Following overexpression of EGFP-tagged JNK3 constructs in COS-7 cells and subsequent pull-down of the overexpressed JNK3 variants, we assessed the catalytic activity in an in vitro kinase reaction involving phosphorylation of the classical JNK substrate c-Jun. We also performed this assay using an EGFP-tagged recombinant JNK3 kinase-defective mutant that we generated, which is based on previously described JNK1 kinase-dead mutants with targeted point mutations at critical phosphorylation sites (Derijard et al. 1994; Matsuura et al. 2002). As expected, when compared with the wild-type JNK3, this mutant was inactive in our kinase assay (see Fig. 2b). The disease-associated JNK3 truncation mutants were likewise unable to phosphorylate c-Jun to detectable levels (Fig. 2b), suggesting that any transcripts expressed from the patients’ defective alleles would encode mutant JNK3 proteins that are indeed catalytically inactive. Clearly, reduced JNK3 kinase activity can be expected in the patients, unless there is significant compensatory catalytic action by the normal protein encoded by the intact gene.
The JNK3 mutant proteins interact with a subset of known JNK-associated scaffold proteins
Based on our genetic data, together with our in vitro data validating that truncated JNK3 proteins indeed lack classical kinase activity, we conclude that both patients very likely suffer from reduced total JNK3 enzymatic activity in the CNS. In line with published expression data for JNK3, we were unable to detect truncated JNK3 proteins in the patient lymphoblastoid cell lines with the antibodies currently available to us; it is, therefore, possible that one or both mutant transcripts are subject to nonsense-mediated decay. However, given that JNK3 is predominantly expressed in the CNS (Martin et al. 1996) and that comparative analysis of patient lymphoblastoid cell lines and mouse brain lysates suggests that lymphoblastoid cell lines do not harbour the same prominent JNK3 isoforms present in brain (data not shown), we cannot use lymphoblastoid cell lines to reliably determine if truncated JNK3 proteins are expressed in patient neurons. Indeed the expression of subtly different truncated JNK3 transcripts could potentially explain the observed phenotypic differences between the two patients we describe. Using coimmunoprecipitation studies, we have therefore compared the ability of wild-type and mutant recombinant JNK3 proteins to interact with known JNK-binding partners. In line with previous studies in which the conserved docking (CD) site is absent or mutated, we observe dramatically reduced interaction between both truncated JNK3 mutants and c-Jun (Fig. 3a). Using similar techniques, we investigated the ability of truncated proteins to bind cytoplasmic JNK3 interaction partners. Compared to the wild-type JNK3, truncated JNK3 proteins are capable of weak binding to JIP1 (Fig. 3b) and to β-Arrestin 2 (Fig. 3c). Importantly, we do not observe obvious differences between the two mutant proteins in these binding assays, suggesting that phenotypic differences between the two patients do not likely result from distinct interactions of the mutant proteins with known binding partners.
Coimmunoprecipitation of EGFP-JNK3 variants with HA-c-Jun (a), FLAG-JIP1b (b) or β-Arrestin2-V5 (c), respectively. Tagged constructs were coexpressed in COS-7 cells and proteins were precipitated from cell lysates with either GFP antibody or mouse IgGs (negative control). In the left panel, the input is observed by western blot of lysates. In the right panels, western blot reflect the presence of proteins in the immunoprecipitates. Transfected proteins are indicated on the left, and their presence in each lysate by the + and − signals above the lanes. Antibodies used for western blots are indicated to the right; blots shown are representative examples of experiments repeated at least 2–4 times
JNK3 is capable of interacting with several synaptic scaffold proteins and endogenous JNK3 colocalises with PSD-95 in primary hippocampal neurons
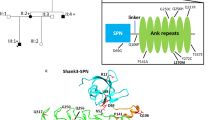
We next searched for novel JNK3 substrates that serve as potential molecular links between JNK3 activity and cognitive function using a candidate gene approach. To identify novel neuronal substrates, we generated a set of candidate interaction partners considering the following criteria: (1) documented expression in neurons, (2) the presence of a predicted JNK docking site (Whisenant et al. 2010), (3) identification of a conserved proline-directed putative phosphorylation site (Edbauer et al. 2009; Jaffe et al. 2004), and (4) association with neurodevelopmental disorders. Candidates were tested for their ability to interact with JNK3 in immunoprecipitation assays. With this strategy, we identified PSD-95, SAP102, and SHANK3 (Fig. 4a–c) as novel JNK3-interaction partners. PSD-95 and SAP102 are members of the MAGUK family of membrane-associated guanylate kinase-like scaffold molecules with an established role in receptor trafficking, and SHANK3 is a synaptic scaffold protein that resides somewhat deeper within the post-synaptic density (PSD) of glutamatergic neurons. Both SAP102 and SHANK3 have been implicated in neurodevelopmental disorders in patients (Tarpey et al. 2004; Durand et al. 2007). Further experiments centered on PSD-95, which is arguably the major scaffolding protein at the PSD and was recently identified as a phosphorylation target of JNK1 (Kim et al. 2007). Following pull-down of endogenous JNK3 from adult brain lysates, PSD-95 is indeed present in the immunoprecipitates (Fig. 4d), and in subsequent analysis of endogenous JNK3 expression in primary neurons, we observed expression throughout the neurites that partially overlapped with sites of enriched PSD-95 expression (Fig. 4e). We, thus, conclude that JNK3 might participate in regulation of PSD-95 at the synapse.
Novel interaction partners of JNK3. Coimmunoprecipitation following coexpression of EGFP-JNK3 with either FLAG-PSD-95 (a), FLAG-SAP102 (b) or HA-Shank3 (c) in COS-7 cells. Following pull-down with the GFP antibody (GFP-IP) or mouse IgGs (negative control, mIgG-IP), proteins are observed by western blot of the immunoprecipitates. Expression of proteins in the lysates (input) was also confirmed by western blot using anti-GFP (goat) and either anti-FLAG (rabbit) or anti-HA (mouse). d Coimmunoprecipitation of endogenous PSD-95 and JNK3 in brain lysates. Following pull-down of JNK3 from adult mouse whole brain lysates using the JNK3-specific mouse monoclonal antibody (Acris, AM06230SU-N), immunoprecipitates were tested for the presence of both PSD-95 and JNK3 (JNK3-specific antibody, rabbit, Upstate/Millipore # 04-893) by western blot. Lysate controls are on the left (input); mouse IgGs were used as a negative control for the immunoprecipitation (mIgG-IP). e Images of primary hippocampal rat neurons following immunostaining with primary antibodies that recognise endogenous PSD-95 and JNK3 (JNK3-specific, Upstate/Millipore # 04-893) and corresponding secondary antibodies (anti-mouse Alexa-Fluor 594, red, for PSD-95 and anti-rabbit Alexa Fluor 488, green, for JNK3, both from Invitrogen). On the left is a representative image depicting the merged channels from a single Z-stack plane at low magnification, with the region of interest highlighted and amplified in the neighbouring panel; regions of overlapping PSD-95 and JNK3 expression are observable in yellow (arrows). Panels on the right reflect single channel images from the same Z-stack plane. Magnification indicated with integrated scale bars
PSD-95 is a novel JNK3 substrate
It has been shown that JNK1 is capable of phosphorylating PSD-95 at serine 295 (Kim et al. 2007). We validated this result (data not shown), and we next asked whether JNK3—in addition to JNK1—might also be responsible for phosphorylation of PSD-95. To explore this question, we used an in vitro kinase assay similar to that used for testing JNK phosphorylation of c-Jun. Unlike in the case of c-Jun, however, there is no commercially available GST-fusion protein harbouring the JNK target site; we therefore generated a comparable construct harbouring the known JNK site within the N-terminal half of PSD-95 (Fig. 5a). Following purification of GST-PSD-95 using glutathione sepharose beads, we used commercially available JNK3 and JNK1 (Biomol) for in vitro kinase reactions. Unlike that for phospho-c-Jun, the phospho-PSD-95 antibody available for purchase is not specific (data not shown); we, therefore, analysed our kinase reaction using the PhosTAG system (Wako, Japan), in which phosphorylated proteins are observably shifted relative to non-phosphorylated forms in polyacrylamide gel electrophoresis. Following kinase reaction with either JNK1 or JNK3, we observed phosphorylated forms of PSD-95 that we do not observe in phosphatase-treated control assays (data not shown), or kinase reactions that lack ATP, indicating that both purified JNK1 and JNK3 are indeed capable of phosphorylating the N-terminal region of PSD-95 (see Fig. 5b), and suggesting that JNK3 might also be important for regulation of PSD-95 phosphorylation at the PSD.
a Comparative diagram depicting the structures of the substrates used for kinase assays shown in b, c and Fig. 2b, and control for expression of GST-PSD-95. The commercially available c-Jun substrate harbours an N-terminal GST tag fused to an N-terminal fragment of the established JNK target c-Jun. Below is the GST-PSD-95 construct that we generated for establishment of a comparable kinase assay with this novel substrate. In both cases, GST is fused to the N-terminal region of the protein harbouring the JNK target site; in the case of c-Jun, GST is followed by amino acids 1–79 of c-Jun; for the case of GST-PSD-95, GST is followed by the FLAG-tag sequence MDYKDDDK and amino acids 1–399 of rat PSD-95. Below is a western blot with a GST antibody to control for expression of the GST-PSD-95 fusion protein generated (compared to GST alone). b Kinase assay using commercial JNK1 and JNK3 proteins (Biomol) for in vitro phosphorylation of the GST-PSD-95 substrate described in a. The kinase used (100 ng) is indicated above, as is the presence or absence of ATP in the kinase reaction. Phosphorylation of the substrate is observable by the shifted GST-PSD-95 signal, which—like the non-phosphorylated form—is observable by western blot with the FLAG antibody after polyacrylamide gel electrophoresis using the PhosTAG system. c Comparative kinase assays in which the abilities of wild-type (WT) and mutant JNK3 proteins to phosphorylate the novel substrate PSD-95 are compared. Following overexpression and pull-down of EGFP-JNK3 variants and controls from COS-7 cell lysates, phosphorylated PSD-95 is observed by western blot using a FLAG antibody with the PhosTAG system. In the top panel, western blot with a GFP antibody indicates comparable pull-down efficiency of JNK3 variants. WT refers to the wild-type EGFP-JNK1/3 construct, and P1 and P2 to the patient-associated variants. Negative control is a pull-down following overexpression of EGFP alone (EGFP). In the bottom panel, the non-phosphorylated (non-shifted) form of GST-PSD-95 is detected (by western blot of the PhosTAG gel using the FLAG antibody). The middle panel is a long exposure of the same PhosTAG experiment in which the shifted (phospho-) GST-PSD-95 can be observed. Note enriched phospho-signal in lanes where the wild-type kinases are present. Blots shown are representative examples of three experiments. d Coimmunoprecipitation of EGFP-JNK3 variants with FLAG-PSD-95. Tagged constructs were coexpressed in COS-7 cells and proteins were precipitated from cell lysates with either GFP antibody or mouse IgGs (negative control). In the left panel, the input is observed by western blot of lysates. In the right panels, western blot reflects the presence of proteins in the immunoprecipitates. Transfected proteins are indicated on the left and their presence in each lysate by the + and − signals above the lanes. Antibodies used for western blots are indicated to the right; blots shown are representative examples of experiments repeated at least 2–4 times
Disease-associated mutant JNK3 variants are not capable of phosphorylating PSD-95
We then investigated whether disease-associated variants might also be able to interact with and/or phosphorylate PSD-95. Truncated versions of JNK3 are not commercially available as purified kinases; we therefore utilised activated lysates overexpressing wild-type or mutant forms of JNK3 for the subsequent set of assays. For these experiments, we again used the purified GST-PSD-95 protein we generated as a substrate. Active kinases were pulled-down from COS-7 cell lysates using the GFP antibody (as for c-Jun phosphorylation assays in Fig. 2b), and phosphorylation of GST-PSD-95 fusion protein was assessed by western blot of a PhosTAG polyacrylamide gel. While substrate is present at comparable levels in all lanes (Fig. 5c bottom panel), and pull-down of EGFP-tagged JNK3 variants occurred with similar efficiency (Fig. 5c top panel), a clear shifted PSD-95 signal was detectable in samples in which wild-type kinases were present, whereas a band of comparable intensity was not observed in samples harbouring the mutant kinases (Fig. 5c middle panel). This suggests that—as for the case of c-Jun—an intact kinase domain is required for efficient phosphorylation of PSD-95 by JNK3. We can, therefore, be confident that patients indeed carry functional JNK3 kinase on only one allele. We next tested the mutant JNK3 proteins for their ability to interact with PSD-95, as we did earlier for established JNK3-binding proteins (see Fig. 3). Following pull-down of either mutant or wild-type JNK3, we observe an interaction with PSD-95; however, similar to the case for other cytoplasmic proteins, the binding affinity of both mutant JNK3 variants is clearly reduced relative to wild-type (Fig. 5d). This is also in line with our phosphorylation data (Fig. 5c).
Discussion
In this study, we have identified an ID patient carrying a de novo chromosome rearrangement affecting MAPK10/JNK3, which provides further evidence in support of a causal relationship between JNK aberrations and defective brain function. Although one allele remains intact, the coding sequence of MAPK10/JNK3 is clearly disrupted on the other allele in both patients. The resulting predicted mutant proteins both lack a large portion of their C-termini, and we have demonstrated both a loss of kinase activity and reduced c-Jun binding capacity for the two related disease-linked recombinant mutant JNK3 proteins. We thus conclude that the classical function of JNK3 as an activator for this essential transcription factor is likely reduced by 50 % in the patients. Importantly, in light of the specific CNS expression of most JNK3 isoforms (Martin et al. 1996), we could not validate the presence of truncated protein in patient material; however, we cannot exclude dominant-interfering effects of the mutant protein in nervous tissue where JNK3 isoforms are normally expressed at high levels. Our previous studies on the mutant protein associated with Patient 1 (Shoichet et al. 2006) suggest that the truncated JNK3 may actively contribute to defective CNS function via toxic effects. Here we demonstrate that truncated JNK3 proteins are also capable of weak interactions with a set of JNK-associated scaffold proteins, suggesting another potential mode of dominant interference in cells that are haploinsufficient for JNK3.
To further elucidate potential disease mechanisms, we searched for novel interaction partners of the neuronal JNK3 using a candidate gene approach, and we identified three synaptic scaffold proteins—PSD-95, SAP102 and SHANK3—as novel JNK3 interactors in coimmunoprecipitation experiments. All three of these proteins harbour predicted JNK docking sites and putative phosphorylation sites for proline-directed MAPK family kinases. Further analysis of the links between JNK3 and PSD-95 led us to novel insights into JNK3 function in neurons. Like JNK1, wild-type JNK3 is capable of binding and also phosphorylating PSD-95, and in line with this result, endogenous JNK3 and PSD-95 have overlapping expression in primary hippocampal neurons. Given that PSD-95 is important for both synaptic maturation and synaptic transmission (El-Husseini et al. 2000), this novel function for JNK3 may be of critical relevance for understanding disease mechanism in MAPK10/JNK3 mutation carriers. Patients presumably exhibit reduced total JNK activity in the CNS, and this is perhaps especially relevant at synaptic sites where PSD-95 plays an important functional role. It is plausible that JNK3 also regulates SAP102 and SHANK3, as well as numerous other as yet unknown neuronal JNK3 substrates. Importantly, mutations in both SAP102 (Tarpey et al. 2004) and SHANK3 (Durand et al. 2007) are associated with neurodevelopmental phenotypes in patients, and links between PSD-95 and cognitive function have been observed in studies on both probands and mouse models (Feyder et al. 2010); clearly, defective regulation of these synaptic proteins could have detrimental developmental consequences.
With the identification and characterisation of an additional de novo ID-associated MAPK10/JNK3 mutation that is similar to the one we described previously, we affirm the association of MAPK10/JNK3 truncation mutations with an ID phenotype. The phenotype of the patient described here (Patient 2) differs from the previously described patient (Patient 1, see Shoichet et al. 2006), in that the seizure disorder is absent and the intellectual disability phenotype is somewhat milder. It is possible that truncated JNK3 proteins are expressed in patient nervous tissue; in this case, differences between the mutant proteins could conceivably give rise to the observed phenotypic differences. It is also plausible that environmental and/or other unknown genetic factors potentially contribute in some way. Importantly, given that JNKs are stress-activated kinases, it is conceivable that patients with aberrations in one of these proteins might be more sensitive to environmental stresses that would otherwise have no influence during development. It is also possible that there are undetected additional genetic aberrations (e.g. sub-resolution copy number variations on other chromosomes or polymorphic disease susceptibility variants in other genes) that might contribute to the more severe phenotype in Patient 1. Within this context, it is interesting that a third ID patient with a chromosome rearrangement involving the MAPK10/JNK3 gene has been described (Baptista et al. 2008). This patient has a complex rearrangement that is accompanied by a deletion affecting six genes, some of which may contribute to the complex phenotype, which includes respiratory problems and scoliosis in addition to learning deficits. One of the breakpoints directly interrupts MAPK10/JNK3 near the middle of the gene (based on Baptista et al. 2008 and personal communication with J. Baptista and J. Crolla), thereby resulting in a truncated transcript that is potentially very similar to those we describe here. Especially in light of our data, it is plausible that the cognitive aspects of the phenotype in the patient described by Baptista et al. are in part mediated by MAPK10/JNK3 alterations.
In conclusion, our study affirms that MAPK10/JNK3 mutations are associated with cognitive deficits and supports the idea that JNK signalling is essential for normal neuronal network formation and cognitive development via interactions with important synaptic proteins. Our data are in line with the idea that known disease-associated MAPK10/JNK3 mutations potentially exert their effects via both loss-of-function and dominant-interfering pathways. Further studies will aim to determine precisely how perturbations in the JNK signalling pathway influence neuronal function.
References
Arkesteijn G, Jumelet E, Hagenbeek A, Smit E, Slater R, Martens A (1999) Reverse chromosome painting for the identification of marker chromosomes and complex translocations in leukemia. Cytometry 35:117–124
Baptista J, Mercer C, Prigmore E, Gribble SM, Carter NP, Maloney V, Thomas NS, Jacobs PA, Crolla JA (2008) Breakpoint mapping and array CGH in translocations: comparison of a phenotypically normal and an abnormal cohort. Am J Hum Genet 82:927–936
Bjorkblom B, Ostman N, Hongisto V, Komarovski V, Filen JJ, Nyman TA, Kallunki T, Courtney MJ, Coffey ET (2005) Constitutively active cytoplasmic c-Jun N-terminal kinase 1 is a dominant regulator of dendritic architecture: role of microtubule-associated protein 2 as an effector. J Neurosci 25:6350–6361
Chen W, Erdogan F, Ropers HH, Lenzner S, Ullmann R (2005) CGHPRO—a comprehensive data analysis tool for array CGH. BMC Bioinformatics 6:85
Derijard B, Hibi M, Wu IH, Barrett T, Su B, Deng T, Karin M, Davis RJ (1994) JNK1: a protein kinase stimulated by UV light and Ha-Ras that binds and phosphorylates the c-Jun activation domain. Cell 76:1025–1037
Durand CM, Betancur C, Boeckers TM, Bockmann J, Chaste P, Fauchereau F, Nygren G, Rastam M, Gillberg IC, Anckarsater H, Sponheim E, Goubran-Botros H, Delorme R, Chabane N, Mouren-Simeoni MC, de Mas P, Bieth E, Roge B, Heron D, Burglen L, Gillberg C, Leboyer M, Bourgeron T (2007) Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet 39:25–27
Edbauer D, Cheng D, Batterton MN, Wang CF, Duong DM, Yaffe MB, Peng J, Sheng M (2009) Identification and characterization of neuronal mitogen-activated protein kinase substrates using a specific phosphomotif antibody. Mol Cell Proteomics 8:681–695
El-Husseini AE, Schnell E, Chetkovich DM, Nicoll RA, Bredt DS (2000) PSD-95 involvement in maturation of excitatory synapses. Science 290:1364–1368
Feyder M, Karlsson RM, Mathur P, Lyman M, Bock R, Momenan R, Munasinghe J, Scattoni ML, Ihne J, Camp M, Graybeal C, Strathdee D, Begg A, Alvarez VA, Kirsch P, Rietschel M, Cichon S, Walter H, Meyer-Lindenberg A, Grant SG, Holmes A (2010) Association of mouse Dlg4 (PSD-95) gene deletion and human DLG4 gene variation with phenotypes relevant to autism spectrum disorders and Williams’ syndrome. Am J Psychiatry 167:1508–1517
Jaffe H, Vinade L, Dosemeci A (2004) Identification of novel phosphorylation sites on postsynaptic density proteins. Biochem Biophys Res Commun 321:210–218
Kalscheuer VM, FitzPatrick D, Tommerup N, Bugge M, Niebuhr E, Neumann LM, Tzschach A, Shoichet SA, Menzel C, Erdogan F, Arkesteijn G, Ropers HH, Ullmann R (2007) Mutations in autism susceptibility candidate 2 (AUTS2) in patients with mental retardation. Hum Genet 121:501–509
Kim MJ, Futai K, Jo J, Hayashi Y, Cho K, Sheng M (2007) Synaptic accumulation of PSD-95 and synaptic function regulated by phosphorylation of serine-295 of PSD-95. Neuron 56:488–502
Kinoshita E, Kinoshita-Kikuta E (2011) Improved Phos-tag SDS-PAGE under neutral pH conditions for advanced protein phosphorylation profiling. Proteomics 11:319–323
Kunde SA, Musante L, Grimme A, Fischer U, Muller E, Wanker EE, Kalscheuer VM (2011) The X-chromosome-linked intellectual disability protein PQBP1 is a component of neuronal RNA granules and regulates the appearance of stress granules. Hum Mol Genet 20:4916–4931
Martin JH, Mohit AA, Miller CA (1996) Developmental expression in the mouse nervous system of the p493F12 SAP kinase. Brain Res Mol Brain Res 35:47–57
Matsuura H, Nishitoh H, Takeda K, Matsuzawa A, Amagasa T, Ito M, Yoshioka K, Ichijo H (2002) Phosphorylation-dependent scaffolding role of JSAP1/JIP3 in the ASK1-JNK signaling pathway. A new mode of regulation of the MAP kinase cascade. J Biol Chem 277:40703–40709
Mehan S, Meena H, Sharma D, Sankhla R (2011) JNK: a stress-activated protein kinase therapeutic strategies and involvement in Alzheimer’s and various neurodegenerative abnormalities. J Mol Neurosci 43:376–390
Pavlowsky A, Gianfelice A, Pallotto M, Zanchi A, Vara H, Khelfaoui M, Valnegri P, Rezai X, Bassani S, Brambilla D, Kumpost J, Blahos J, Roux MJ, Humeau Y, Chelly J, Passafaro M, Giustetto M, Billuart P, Sala C (2010a) A postsynaptic signaling pathway that may account for the cognitive defect due to IL1RAPL1 mutation. Curr Biol 20:103–115
Pavlowsky A, Zanchi A, Pallotto M, Giustetto M, Chelly J, Sala C, Billuart P (2010b) Neuronal JNK pathway activation by IL-1 is mediated through IL1RAPL1, a protein required for development of cognitive functions. Commun Integr Biol 3:245–247
Shoichet SA, Duprez L, Hagens O, Waetzig V, Menzel C, Herdegen T, Schweiger S, Dan B, Vamos E, Ropers HH, Kalscheuer VM (2006) Truncation of the CNS-expressed JNK3 in a patient with a severe developmental epileptic encephalopathy. Hum Genet 118:559–567
Tarpey P, Parnau J, Blow M, Woffendin H, Bignell G, Cox C, Cox J, Davies H, Edkins S, Holden S, Korny A, Mallya U, Moon J, O’Meara S, Parker A, Stephens P, Stevens C, Teague J, Donnelly A, Mangelsdorf M, Mulley J, Partington M, Turner G, Stevenson R, Schwartz C, Young I, Easton D, Bobrow M, Futreal PA, Stratton MR, Gecz J, Wooster R, Raymond FL (2004) Mutations in the DLG3 gene cause nonsyndromic X-linked mental retardation. Am J Hum Genet 75:318–324
Whisenant TC, Ho DT, Benz RW, Rogers JS, Kaake RM, et al (2010) Computational Prediction and Experimental Verification of New MAP Kinase Docking Sites and Substrates Including Gli Transcription Factors. PLoS Comput Biol 6(8):e1000908. doi:10.1371/journal.pcbi.1000908
Xu X, Raber J, Yang D, Su B, Mucke L (1997) Dynamic regulation of c-Jun N-terminal kinase activity in mouse brain by environmental stimuli. Proc Natl Acad Sci USA 94:12655–12660
Acknowledgments
This study was funded by the ‘Deutsche Forschungsgemeinschaft’ through the Excellence Cluster NeuroCure (EXC257), by a grant from the German Ministry of Education and Research through the MRNET and the EU FP7 project GENCODYS (Grant # 241995). We thank S. Freier, M. Fuchs, and I. Müller for excellent technical assistance, and we are grateful to the patients and their families for their willingness to participate in this study.
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kunde, SA., Rademacher, N., Tzschach, A. et al. Characterisation of de novo MAPK10/JNK3 truncation mutations associated with cognitive disorders in two unrelated patients. Hum Genet 132, 461–471 (2013). https://doi.org/10.1007/s00439-012-1260-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-012-1260-5