Abstract
The tumor necrosis factor alpha-inducible protein 3 (TNFAIP3) gene polymorphisms have recently been reported to be associated with the susceptibility to several immune-related diseases. This study was performed to evaluate the potential association of TNFAIP3 polymorphisms with Behcet’s disease (BD) in a Chinese Han population. Five single-nucleotide polymorphisms (SNPs), rs10499194, rs610604, rs7753873, rs5029928, and rs9494885 of TNFAIP3 were genotyped in 722 BD patients and 1,415 healthy controls using a PCR-restriction fragment length polymorphism assay. Allele and genotype frequencies were compared between patients and controls using the χ 2 test. The results showed a significantly increased prevalence of the rs9494885 TC genotype and C allele in BD patients compared with controls (Bonferroni corrected p (p c) = 1.83 × 10−10, odds ratio (OR) [95 % CI] 2.03 [1.65–2.49]; p c = 8.35 × 10−10, OR [95 % CI] 1.81 [1.51–2.18], respectively).The frequency of the TT genotype and T allele of rs9494885 was markedly lower in BD patients than that in controls (p c = 1.23 × 10−10, OR [95 % CI] 0.50 [0.40–0.61]; p c = 8.35 × 10−10, OR [95 % CI] 0.55 [0.46–0.66], respectively). For rs10499194, a higher frequency of the CC genotype (p c = 0.015, OR [95 % CI] 1.96 [1.30–2.97]) and C allele (p c = 0.005, OR [95 % CI] 1.92 [1.28–2.90]), and a lower frequency of the TC genotype (p c = 0.015, OR [95 % CI] 0.51 [0.34–0.77]) and T allele (p c = 0.005, OR [95 % CI] 0.52 [0.35–2.97]) were found in BD patients. Concerning rs7753873, a higher frequency of the AC genotype (p c = 0.015, OR [95 % CI] 1.49 [1.17–1.91]) and C allele (p c = 0.025, OR [95 % CI] 1.39 [1.11–1.76]), and a lower frequency of the AA genotype (p c = 0.03, OR [95 % CI] 0.68 [0.53–0.87]) and A allele (p c = 0.025, OR [95 % CI] 0.72 [0.57–0.91]) were observed in BD patients. This study identified one strong risk SNP rs9494885 and two weak risk SNPs rs10499194 and rs7753873 of TNFAIP3 in Chinese Han BD patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Behçet’s disease (BD) is an idiopathic, multisystem, recurrent chronic inflammatory disease, clinically characterized as recurrent uveitis, recurrent oral ulceration, genital ulceration, and erythema nodosum. It is quite common along the ancient ‘Silk Road’ countries extending from China to the Mediterranean area (Keino and Okada 2007; Yang et al. 2008). Although the precise etiology of BD remains unknown, extensive studies suggest that the environment and genetic factors are all involved in its pathogenesis. It has been shown that polymorphisms of several genes such as human leukocyte antigen-B51 (HLA-B51) (Cohen et al. 2002), intercellular adhesion molecule-1(ICAM-1) (Verity et al. 2000), small ubiquitin-like modifier4 (SUMO4) (Hou et al. 2008), interleukin-23 receptor (IL-23R)-IL12RB2 (Jiang et al. 2010; Mizuki et al. 2010; Remmers et al. 2010), IL10 (Mizuki et al. 2010; Remmers et al. 2010), monocyte chemoattractant protein-1 (MCP)-1 (Hou et al. 2010), signal transducers and activators of transcription 4(STAT4) (Hou et al. 2012c; Hu et al. 2010), CCR1/CCR3 (Hou et al. 2012b), UBAC2 (Hou et al. 2012a), and Fc receptor-like 3 gene (FCRL3) (Li et al. 2008) are established or suggested to be associated with susceptibility to BD. However, the identified risk genes such as HLA-B51 only account for approximately 20 % of the genetic-risk effect in siblings of affected individuals (Gul et al. 2001). Therefore, there are many non-HLA risk genes which need to be investigated.
TNFAIP3 encodes the ubiquitin-modifying enzyme A20, a key regulator of the NF-κB signaling pathway of tumor necrosis factor alpha (TNFα), toll-like receptor (TLR), interleukin 1 receptor (IL1R), and nucleotide-binding oligomerization domain containing 2 (NOD2) (Boone et al. 2004; Hitotsumatsu et al. 2008; Jaattela et al. 1996; Lee et al. 2000). Several studies have suggested a role for TNFAIP3 polymorphisms in the susceptibility to complex genetic autoimmune disorders, including rheumatoid arthritis (RA) (Plenge et al. 2007; Thomson et al. 2007), systemic sclerosis (SSc) (Dieude et al. 2010),systemic lupus erythematosus (SLE) (Adrianto et al. 2011; Graham et al. 2008; Musone et al. 2008), psoriasis (Musone et al. 2011; Nair et al. 2009), Sjögren’s syndrome (SS) (Musone et al. 2011), Crohn’s disease (Musone et al. 2011; WTCCC 2007), celiac disease (Dubois et al. 2010; Trynka et al. 2009; Zhernakova et al. 2011), diabetes (Fung et al. 2009), and ulcerative colitis (Wang et al. 2010a). These findings suggest that TNFAIP3 may be a common risk gene for a number of immune-related disorders. Moreover, these genetic findings described in human patients linking TNFAIP3 with autoimmune inflammatory pathology such as RA, SLE, psoriasis, and CD were experimentally confirmed in mice (Chu et al. 2011; Hammer et al. 2011; Kool et al. 2011; Matmati et al. 2011; Tavares et al. 2010; Vereecke et al. 2010). Based on this evidence, we postulated that TNFAIP3 might also be a risk gene for BD and therefore examined whether the TNFAIP3 gene was associated with the susceptibility to BD in a Chinese Han population. In this study we identified a strong association between rs9494885 of this gene with BD and two weak associations were observed with rs7753873 and rs10499194.
Methods
Study population
Seven hundred and twenty-two BD patients who all belonged to the Chinese Han population were included in this study. The control population consisted of 1,415 unrelated healthy individuals who were selected from the same geographical regions as the BD patients. There were no differences in age, sex, and ethnicity between patients and controls. The blood samples were obtained from the First Affiliated Hospital, Chongqing Medical University (Chongqing, China) or the Uveitis Study Center of the Sun Yat-sen University (Guangzhou, China). The diagnosis of BD was based on the criteria of the International Study Group (1990). The clinical characteristics of the BD patients were assessed at the time of diagnosis and are summarized in Table 1. All study participants gave written informed consent and the local ethical committee of both hospitals approved the study.
SNP selection and genotyping
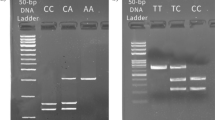
We studied 5 SNPs rs10499194, rs610604, rs7753873, rs5029928 and rs9494885 in the TNFAIP3 region on 6q23, which were demonstrated earlier by other groups to be associated with certain immune-related diseases. SNPs rs10499194, rs7753873, and rs9494885 are located in the intergenic region of TNFAIP3. SNPs rs610604, rs5029928 are located in an intron of TNFAIP3. Genomic DNA was isolated from blood leukocytes using the commercial kit Qiagen DNA Blood Mini kit (Qiagen, Valencia, CA). The extracted DNA was stored at −20 °C until use. Amplification of the target DNA was performed by PCR. The primers used in this study are shown in Table 2. These five SNPs were genotyped by restriction fragment length polymorphism analysis. The amplification was performed using initial denaturation at 95 °C for 5 min, 95 °C for 30 s, 58–62 °C for 30 s, 72 °C for 30 s, and 72 °C for 5 min followed by 37 cycles. The PCR products were incubated with restriction enzymes (Table 2) for at least 4 h. Digestion products were visualized on a 4.0 % agarose gel and stained with Goodview (SBS Genetech, Beijing, China). Direct sequencing was performed by the Invitrogen Biotechnology Company (Guangzhou, Guangdong province, China) using randomly selected subjects (20 % of all samples) to validate the method used in this study.
Real-time PCR
Peripheral blood mononuclear cells (PBMCs) were prepared from peripheral blood by Ficoll-Hypaque density-gradient centrifugation. Total RNA was extracted from PBMCs using TRIzol (Invitrogen), followed by reverse transcription using a transcriptase kit (Applied Biosysterms). The relative expression level of mRNA was normalized to that of the internal control β-actin by using the 2−ΔΔCt cycle threshold method.
Statistical analysis
Distribution of genotypes and alleles between patients and normal controls was analyzed using SPSS version 17.0 (SPSS, Inc., Chicago, IL). The Chi square test was used to compare allele and genotype distributions. In case a genotype had a frequency less than 10, the Fisher’s exact test was applied. Bonferroni correction was applied for multiple testing. We also performed a power analysis using Quanto software to verify the association between BD and the tested SNPs. The result showed a power value of 0.84 for SNP rs10499194, 0.26 for SNP rs610604, 0.05 for SNP rs5029928, 0.48 for SNP rs7753873, and 0.96 for SNP rs9494885 using a prevalence of Behcet’s disease in our population of 0.01 % (Wang et al. 2010b) (Supplementary Table 1).
Results
There were no differences in age and sex between patients and controls. The results showed that the five SNPS rs10499194, rs610604, rs7753873. and rs5029928 rs9494885of TNFAIP3 were in Hardy–Weinberg equilibrium in the cases and controls (P > 0.01). The distribution of both genotype frequencies and allele frequencies of the five tested TNFAIP3 polymorphisms are shown in Table 3. The call rate for the examined five SNPs is 100 %. There were no statistically significant differences in the proportions of missing genotype data between cases and controls (P > 0.05). The results showed that there were significant differences between BD patients and controls concerning the frequencies of rs9494885, rs10499194, and rs7753873. A significantly increased prevalence of the rs9494885 TC genotype and C allele was found in BD patients compared with controls (p c = 1.83 × 10−10, OR 2.02, 95 % CI 1.65–2.49; p c = 8.35 × 10−10, OR 1.81, 95 % CI 1.51–2.18, respectively). The frequencies of the TT genotype and T allele were significantly lower in BD patients than that in controls (p c = 1.23 × 10−10, OR 0.50, 95 % CI 0.40–0.61; p c = 8.35 × 10−10, OR 0.55, 95 % CI 0.46–0.66, respectively). The frequency of the rs10499194 CC genotype (p c = 0.015, OR 1.96, 95 % CI 1.30–2.97) and C allele (p c = 0.005, OR 1.924,95 % CI 1.28–2.90) were higher in BD patients than that in controls. The frequency of the rs10499194 TC genotype (p c = 0.015, OR 0.51, 95 % CI 0.34–0.77) and T allele (p c = 0.005, OR 0.52, 95 % CI 0.35–0.78) were lower in BD patients than that in controls. For rs7753873, a higher frequency of the AC genotype (p c = 0.015, OR 1.49, 95 % CI 1.17–1.91) and C allele (p c = 0.025, OR 1.39, 95 % CI 1.11–1.76), and a lower frequency of the AA genotype (p c = 0.03, OR 0.68, 95 % CI 0.53–0.87) and A allele (p c = 0.025, OR 0.72, 95 % CI 0.57–0.91) were found in BD patients compared with controls. There was no association of BD with rs610604 or with rs5029928. The haplotype block was constructed using haploview software 4.0. The results showed that the five examined SNPs in this study did not exibit strong linkage disequilibrium to each other (r 2 < 0.8). There results suggest that the association with SNPs rs10499194 and rs7753873 may be independent from SNP rs9494885.
We further investigated whether the TNFAIP3 SNPs studied were associated with certain clinical findings of BD, such as genital ulceration, hypopyon, skin lesions, and arthritis. The analysis failed to find any association of these parameters with the tested TNFAIP3 SNPs.
The results stated above showed a strong association of rs9494885 in TNFAIP3 with BD.A further study was performed on a set of PBMCs from 16 individuals (with 8 TT and 8 TC) derived from healthy individuals of known SNP (rs9494885) status to examine whether this SNP influences the expression. TNFAIP3 mRNA levels were measured by the SYBGREEN assay (Takara, Dalian, China) with available RNA samples. The results showed that there was no difference in TNFAIP3 expression between individuals with the TT or TC genotype (Fig. 1).
Discussion
This study identified a novel association between SNP rs9494885 in TNFAIP3 and BD in a Chinese Han population. The results suggest that the rs9494885 TT genotype and T allele of TNFAIP3 could provide strong protection against BD. The rs9494885 TC genotype and C allele were high-risk factors for this disease. Our study also showed that SNP rs10499194 and rs7753873 were weakly associated with susceptibility to BD. The CC genotype and C allele of rs10499194, and the AC genotype and C allele of rs7753873 were the risk factors for BD whereby the AA genotype and A allele of rs7753873, and the TC genotype and T allele of rs10499194 provided protection.
In this study we chose TNFAIP3 as a candidate gene to investigate its relationship with BD principally based on its involvement in several signaling pathways (Boone et al. 2004; Hitotsumatsu et al. 2008; Jaattela et al. 1996; Lee et al. 2000) and its association with various autoimmune diseases (Adrianto et al. 2011; Dieude et al. 2010; Graham et al. 2008; Musone et al. 2011; Nair et al. 2009; Plenge et al. 2007; Thomson et al. 2007; Wang et al. 2010a). The fact that TNF-α, an important cytokine involved in BD pathogenesis, triggers TNFAIP3 gene expression (Arida et al. 2011; Song et al. 1996), also stimulated us to investigate the relationship between this gene and BD. As to the tested SNPs, we chose these five SNPs predominantly based on the results previously reported on one or more autoimmune diseases including psoriatic arthritis (Bowes et al. 2011), psoriasis (Nair et al. 2009; Tejasvi et al. 2012), RA (Plenge et al. 2007; Shimane et al. 2010), and SLE (Adrianto et al. 2011). Although other variants in this gene have also been shown to be associated with autoimmune disease (Bowes et al. 2010; Plenge et al. 2007), they were not included in our study due to the absence of polymorphisms in the Chinese Han population.
Numerous factors have been reported to influence the results of studies on the association of gene polymorphisms with complex diseases. Efforts were made to decrease the influence of confounding factors on the results. BD patients were strictly selected according to the criteria of the International Study Group. Unrelated healthy individuals acting as controls were selected from the same geographical regions as the BD patients. Furthermore, we selected age- and sex-matched normal individuals as controls. Finally, 20 % of the samples were randomly chosen and investigated by direct sequencing for the purpose of validating the method used in this study and the results showed no difference.
We found a significantly increased prevalence of the rs9494885 TC genotype and a lower frequency of the TT genotype in BD patients. This result is consistent with that reported in SLE in European-ancestry and Korean populations (Adrianto et al. 2011). We also found a higher frequency of the CC genotype and a lower frequency TC in rs10499194, in BD patients compared with controls. This result is consistent with that reported in RA and SLE in Japanese patients (Shimane et al. 2010). In addition, we observed a higher frequency of the AC genotype and a lower frequency of the AA genotype in rs7753873.This result is consistent with that reported in SLE in European-ancestry and Korean populations (Adrianto et al. 2011). These results seem to suggest that BD has, to a certain extent, similarity in genetic background with SLE and RA. The latter are considered autoimmune diseases, whereas BD is generally seen as an auto-inflammatory disease. We failed to find any association between rs610604 and rs5029928 with BD though both SNPs have been reported to be associated with a number of autoimmune diseases (Table 4).
Previous genome-wide association studies have identified multiple risk genes for Behcet’s disease including IL23R-IL12RB2, IL10, UBAC2, and STAT4 (Fei et al. 2009; Hou et al. 2012c; Mizuki et al. 2010; Remmers et al. 2010). However, these studies did not find the association of TNFAIP3 with Behcet’s disease. These results suggest that Behcet’s disease displays genetic heterogeneity in different ethnic populations.
The above findings suggest that rs9494885 may be an important susceptibility factor for autoimmune disease. Functional studies of the TNFAIP3 polymorphism will be an important advance to explain how polymorphisms of this gene affect the susceptibility to autoimmune disease. As yet little is known concerning its effect on the function or expression of this gene. We therefore evaluated the association of the rs9494885 genotypes and TNFAIP3 expression. The results showed no differences in TNFAIP3 expression between the individuals with the TT and those with the TC genotype of rs9494885. This study suggests that the diseases associated SNP rs9494885 may be involved in this disease through an unknown mechanisms rather than directly regulating TNFAIP3 transcriptional regulation. In addition to mechanisms other than transcription, rs94948855 may be in linkage disequilibrium (LD) with the causal variant. Although other sites such as rs6920220, rs6933404, and rs6927172 in TNFAIP3 have also been shown to be associated with rheumatoid arthritis (Elsby et al. 2010) and showed a functional role in disease, we did not study the association of these SNPs with Behcet’s disease due to nonpolymorphism in Chinese population based on the database of HapMap.
It is worth mentioning that there are some limitations in the present study. BD is a systemic disease involving various organs and the patients enrolled in our study originated from an ophthalmology department and may, therefore, represent a subpopulation of this disease. The susceptible SNPs identified in this study are, therefore, associated only with uveitis in BD and more studies are needed to confirm the present results using BD patients from other medical departments.
As far as we are aware this is the first study to report an association between TNFAIP3 SNPs rs10499194, rs7753873, and rs9494885 with BD in Chinese Han patients.
References
(1990) Criteria for diagnosis of Behcet’s disease. International Study Group for Behcet’s Disease. Lancet 335:1078–10association study identifies susceptible locus80
Adrianto I, Wen F, Templeton A, Wiley G, King JB, Lessard CJ, Bates JS, Hu Y, Kelly JA, Kaufman KM, Guthridge JM, Alarcon-Riquelme ME, Anaya JM, Bae SC, Bang SY, Boackle SA, Brown EE, Petri MA, Gallant C, Ramsey-Goldman R, Reveille JD, Vila LM, Criswell LA, Edberg JC, Freedman BI, Gregersen PK, Gilkeson GS, Jacob CO, James JA, Kamen DL, Kimberly RP, Martin J, Merrill JT, Niewold TB, Park SY, Pons-Estel BA, Scofield RH, Stevens AM, Tsao BP, Vyse TJ, Langefeld CD, Harley JB, Moser KL, Webb CF, Humphrey MB, Montgomery CG, Gaffney PM (2011) Association of a functional variant downstream of TNFAIP3 with systemic lupus erythematosus. Nat Genet 43:253–258
Arida A, Fragiadaki K, Giavri E, Sfikakis PP (2011) Anti-TNF agents for Behcet’s disease: analysis of published data on 369 patients. Semin Arthritis Rheum 41:61–70
Boone DL, Turer EE, Lee EG, Ahmad RC, Wheeler MT, Tsui C, Hurley P, Chien M, Chai S, Hitotsumatsu O, McNally E, Pickart C, Ma A (2004) The ubiquitin-modifying enzyme A20 is required for termination of Toll-like receptor responses. Nat Immunol 5:1052–1060
Bowes J, Lawrence R, Eyre S, Panoutsopoulou K, Orozco G, Elliott KS, Ke X, Morris AP, Thomson W, Worthington J, Barton A, Zeggini E (2010) Rare variation at the TNFAIP3 locus and susceptibility to rheumatoid arthritis. Hum Genet 128:627–633
Bowes J, Orozco G, Flynn E, Ho P, Brier R, Marzo-Ortega H, Coates L, McManus R, Ryan AW, Kane D, Korendowych E, McHugh N, FitzGerald O, Packham J, Morgan AW, Bruce IN, Barton A (2011) Confirmation of TNIP1 and IL23A as susceptibility loci for psoriatic arthritis. Ann Rheum Dis 70:1641–1644
Chu Y, Vahl JC, Kumar D, Heger K, Bertossi A, Wojtowicz E, Soberon V, Schenten D, Mack B, Reutelshofer M, Beyaert R, Amann K, van Loo G, Schmidt-Supprian M (2011) B cells lacking the tumor suppressor TNFAIP3/A20 display impaired differentiation and hyperactivation and cause inflammation and autoimmunity in aged mice. Blood 117:2227–2236
Cohen R, Metzger S, Nahir M, Chajek-Shaul T (2002) Association of the MIC-A gene and HLA-B51 with Behcet’s disease in Arabs and non-Ashkenazi Jews in Israel. Ann Rheum Dis 61:157–160
Dieude P, Guedj M, Wipff J, Ruiz B, Riemekasten G, Matucci-Cerinic M, Melchers I, Hachulla E, Airo P, Diot E, Hunzelmann N, Cabane J, Mouthon L, Cracowski JL, Riccieri V, Distler J, Meyer O, Kahan A, Boileau C, Allanore Y (2010) Association of the TNFAIP3 rs5029939 variant with systemic sclerosis in the European Caucasian population. Ann Rheum Dis 69:1958–1964
Dubois PC, Trynka G, Franke L, Hunt KA, Romanos J, Curtotti A, Zhernakova A, Heap GA, Adany R, Aromaa A, Bardella MT, van den Berg LH, Bockett NA, de la Concha EG, Dema B, Fehrmann RS, Fernandez-Arquero M, Fiatal S, Grandone E, Green PM, Groen HJ, Gwilliam R, Houwen RH, Hunt SE, Kaukinen K, Kelleher D, Korponay-Szabo I, Kurppa K, MacMathuna P, Maki M, Mazzilli MC, McCann OT, Mearin ML, Mein CA, Mirza MM, Mistry V, Mora B, Morley KI, Mulder CJ, Murray JA, Nunez C, Oosterom E, Ophoff RA, Polanco I, Peltonen L, Platteel M, Rybak A, Salomaa V, Schweizer JJ, Sperandeo MP, Tack GJ, Turner G, Veldink JH, Verbeek WH, Weersma RK, Wolters VM, Urcelay E, Cukrowska B, Greco L, Neuhausen SL, McManus R, Barisani D, Deloukas P, Barrett JC, Saavalainen P, Wijmenga C, van Heel DA (2010) Multiple common variants for celiac disease influencing immune gene expression. Nat Genet 42:295–302
Elsby LM, Orozco G, Denton J, Worthington J, Ray DW, Donn RP (2010) Functional evaluation of TNFAIP3 (A20) in rheumatoid arthritis. Clin Exp Rheumatol 28:708–714
Fei Y, Webb R, Cobb BL, Direskeneli H, Saruhan-Direskeneli G, Sawalha AH (2009) Identification of novel genetic susceptibility loci for Behcet’s disease using a genome-wide association study. Arthritis Res Ther 11:R66
Fung EY, Smyth DJ, Howson JM, Cooper JD, Walker NM, Stevens H, Wicker LS, Todd JA (2009) Analysis of 17 autoimmune disease-associated variants in type 1 diabetes identifies 6q23/TNFAIP3 as a susceptibility locus. Genes Immun 10:188–191
Graham RR, Cotsapas C, Davies L, Hackett R, Lessard CJ, Leon JM, Burtt NP, Guiducci C, Parkin M, Gates C, Plenge RM, Behrens TW, Wither JE, Rioux JD, Fortin PR, Graham DC, Wong AK, Vyse TJ, Daly MJ, Altshuler D, Moser KL, Gaffney PM (2008) Genetic variants near TNFAIP3 on 6q23 are associated with systemic lupus erythematosus. Nat Genet 40:1059–1061
Gul A, Hajeer AH, Worthington J, Barrett JH, Ollier WE, Silman AJ (2001) Evidence for linkage of the HLA-B locus in Behcet’s disease, obtained using the transmission disequilibrium test. Arthritis Rheum 44:239–240
Hammer GE, Turer EE, Taylor KE, Fang CJ, Advincula R, Oshima S, Barrera J, Huang EJ, Hou B, Malynn BA, Reizis B, DeFranco A, Criswell LA, Nakamura MC, Ma A (2011) Expression of A20 by dendritic cells preserves immune homeostasis and prevents colitis and spondyloarthritis. Nat Immunol 12:1184–1193
Hitotsumatsu O, Ahmad RC, Tavares R, Wang M, Philpott D, Turer EE, Lee BL, Shiffin N, Advincula R, Malynn BA, Werts C, Ma A (2008) The ubiquitin-editing enzyme A20 restricts nucleotide-binding oligomerization domain containing 2-triggered signals. Immunity 28:381–390
Hou S, Yang P, Du L, Zhou H, Lin X, Liu X, Kijlstra A (2008) SUMO4 gene polymorphisms in Chinese Han patients with Behcet’s disease. Clin Immunol 129:170–175
Hou S, Yang P, Du L, Jiang Z, Mao L, Shu Q, Zhou H, Kijlstra A (2010) Monocyte chemoattractant protein-1 -2518 A/G single nucleotide polymorphism in Chinese Han patients with ocular Behcet’s disease. Hum Immunol 71:79–82
Hou S, Shu Q, Jiang Z, Chen Y, Li F, Chen F, Kijlstra A, Yang P (2012a) Replication study confirms the association between UBAC2 and Behcet’s disease in two independent Chinese sets of patients and controls. Arthritis Res Ther 14:R70
Hou S, Xiao X, Li F, Jiang Z, Kijlstra A, Yang P (2012b) Two-stage association study in Chinese Han identifies two independent associations in CCR1/CCR3 locus as candidate for Behcet’s disease susceptibility. Hum Genet (Epub ahead of print)
Hou S, Yang Z, Du L, Jiang Z, Shu Q, Chen Y, Li F, Zhou Q, Ohno S, Chen R, Kijlstra A, Rosenbaum JT, Yang P (2012c) Genome-wide association study identifies susceptible locus in STAT4 for Behçet’s disease in Han Chinese. Arthritis Rheum (Epub ahead of print)
Hu K, Yang P, Jiang Z, Hou S, Du L, Li F (2010) STAT4 polymorphism in a Chinese Han population with Vogt-Koyanagi-Harada syndrome and Behcet’s disease. Hum Immunol 71:723–726
Jaattela M, Mouritzen H, Elling F, Bastholm L (1996) A20 zinc finger protein inhibits TNF and IL-1 signaling. J Immunol 156:1166–1173
Jiang Z, Yang P, Hou S, Du L, Xie L, Zhou H, Kijlstra A (2010) IL-23R gene confers susceptibility to Behcet’s disease in a Chinese Han population. Ann Rheum Dis 69:1325–1328
Keino H, Okada AA (2007) Behcet’s disease: global epidemiology of an Old Silk Road disease. Br J Ophthalmol 91:1573–1574
Kool M, van Loo G, Waelput W, De Prijck S, Muskens F, Sze M, van Praet J, Branco-Madeira F, Janssens S, Reizis B, Elewaut D, Beyaert R, Hammad H, Lambrecht BN (2011) The ubiquitin-editing protein A20 prevents dendritic cell activation, recognition of apoptotic cells, and systemic autoimmunity. Immunity 35:82–96
Lee EG, Boone DL, Chai S, Libby SL, Chien M, Lodolce JP, Ma A (2000) Failure to regulate TNF-induced NF-kappaB and cell death responses in A20-deficient mice. Science 289:2350–2354
Li K, Zhao M, Hou S, Du L, Kijlstra A, Yang P (2008) Association between polymorphisms of FCRL3, a non-HLA gene, and Behcet’s disease in a Chinese population with ophthalmic manifestations. Mol Vis 14:2136–2142
Matmati M, Jacques P, Maelfait J, Verheugen E, Kool M, Sze M, Geboes L, Louagie E, Mc Guire C, Vereecke L, Chu Y, Boon L, Staelens S, Matthys P, Lambrecht BN, Schmidt-Supprian M, Pasparakis M, Elewaut D, Beyaert R, van Loo G (2011) A20 (TNFAIP3) deficiency in myeloid cells triggers erosive polyarthritis resembling rheumatoid arthritis. Nat Genet 43:908–912
Mizuki N, Meguro A, Ota M, Ohno S, Shiota T, Kawagoe T, Ito N, Kera J, Okada E, Yatsu K, Song YW, Lee EB, Kitaichi N, Namba K, Horie Y, Takeno M, Sugita S, Mochizuki M, Bahram S, Ishigatsubo Y, Inoko H (2010) Genome-wide association studies identify IL23R-IL12RB2 and IL10 as Behcet’s disease susceptibility loci. Nat Genet 42:703–706
Musone SL, Taylor KE, Lu TT, Nititham J, Ferreira RC, Ortmann W, Shifrin N, Petri MA, Kamboh MI, Manzi S, Seldin MF, Gregersen PK, Behrens TW, Ma A, Kwok PY, Criswell LA (2008) Multiple polymorphisms in the TNFAIP3 region are independently associated with systemic lupus erythematosus. Nat Genet 40:1062–1064
Musone SL, Taylor KE, Nititham J, Chu C, Poon A, Liao W, Lam ET, Ma A, Kwok PY, Criswell LA (2011) Sequencing of TNFAIP3 and association of variants with multiple autoimmune diseases. Genes Immun 12:176–182
Nair RP, Duffin KC, Helms C, Ding J, Stuart PE, Goldgar D, Gudjonsson JE, Li Y, Tejasvi T, Feng BJ, Ruether A, Schreiber S, Weichenthal M, Gladman D, Rahman P, Schrodi SJ, Prahalad S, Guthery SL, Fischer J, Liao W, Kwok PY, Menter A, Lathrop GM, Wise CA, Begovich AB, Voorhees JJ, Elder JT, Krueger GG, Bowcock AM, Abecasis GR (2009) Genome-wide scan reveals association of psoriasis with IL-23 and NF-kappaB pathways. Nat Genet 41:199–204
Plenge RM, Cotsapas C, Davies L, Price AL, de Bakker PI, Maller J, Pe’er I, Burtt NP, Blumenstiel B, DeFelice M, Parkin M, Barry R, Winslow W, Healy C, Graham RR, Neale BM, Izmailova E, Roubenoff R, Parker AN, Glass R, Karlson EW, Maher N, Hafler DA, Lee DM, Seldin MF, Remmers EF, Lee AT, Padyukov L, Alfredsson L, Coblyn J, Weinblatt ME, Gabriel SB, Purcell S, Klareskog L, Gregersen PK, Shadick NA, Daly MJ, Altshuler D (2007) Two independent alleles at 6q23 associated with risk of rheumatoid arthritis. Nat Genet 39:1477–1482
Remmers EF, Cosan F, Kirino Y, Ombrello MJ, Abaci N, Satorius C, Le JM, Yang B, Korman BD, Cakiris A, Aglar O, Emrence Z, Azakli H, Ustek D, Tugal-Tutkun I, Akman-Demir G, Chen W, Amos CI, Dizon MB, Kose AA, Azizlerli G, Erer B, Brand OJ, Kaklamani VG, Kaklamanis P, Ben-Chetrit E, Stanford M, Fortune F, Ghabra M, Ollier WE, Cho YH, Bang D, O’Shea J, Wallace GR, Gadina M, Kastner DL, Gul A (2010) Genome-wide association study identifies variants in the MHC class I, IL10, and IL23R-IL12RB2 regions associated with Behcet’s disease. Nat Genet 42:698–702
Shimane K, Kochi Y, Horita T, Ikari K, Amano H, Hirakata M, Okamoto A, Yamada R, Myouzen K, Suzuki A, Kubo M, Atsumi T, Koike T, Takasaki Y, Momohara S, Yamanaka H, Nakamura Y, Yamamoto K (2010) The association of a nonsynonymous single-nucleotide polymorphism in TNFAIP3 with systemic lupus erythematosus and rheumatoid arthritis in the Japanese population. Arthritis Rheum 62:574–579
Song HY, Rothe M, Goeddel DV (1996) The tumor necrosis factor-inducible zinc finger protein A20 interacts with TRAF1/TRAF2 and inhibits NF-kappaB activation. Proc Natl Acad Sci USA 93:6721–6725
Tavares RM, Turer EE, Liu CL, Advincula R, Scapini P, Rhee L, Barrera J, Lowell CA, Utz PJ, Malynn BA, Ma A (2010) The ubiquitin modifying enzyme A20 restricts B cell survival and prevents autoimmunity. Immunity 33:181–191
Tejasvi T, Stuart PE, Chandran V, Voorhees JJ, Gladman DD, Rahman P, Elder JT, Nair RP (2012) TNFAIP3 gene polymorphisms are associated with response to TNF blockade in psoriasis. J Invest Dermatol 132:593–600
Thomson W, Barton A, Ke X, Eyre S, Hinks A, Bowes J, Donn R, Symmons D, Hider S, Bruce IN, Wilson AG, Marinou I, Morgan A, Emery P, Carter A, Steer S, Hocking L, Reid DM, Wordsworth P, Harrison P, Strachan D, Worthington J (2007) Rheumatoid arthritis association at 6q23. Nat Genet 39:1431–1433
Trynka G, Zhernakova A, Romanos J, Franke L, Hunt KA, Turner G, Bruinenberg M, Heap GA, Platteel M, Ryan AW, de Kovel C, Holmes GK, Howdle PD, Walters JR, Sanders DS, Mulder CJ, Mearin ML, Verbeek WH, Trimble V, Stevens FM, Kelleher D, Barisani D, Bardella MT, McManus R, van Heel DA, Wijmenga C (2009) Coeliac disease-associated risk variants in TNFAIP3 and REL implicate altered NF-kappaB signalling. Gut 58:1078–1083
Vereecke L, Sze M, Mc Guire C, Rogiers B, Chu Y, Schmidt-Supprian M, Pasparakis M, Beyaert R, van Loo G (2010) Enterocyte-specific A20 deficiency sensitizes to tumor necrosis factor-induced toxicity and experimental colitis. J Exp Med 207:1513–1523
Verity DH, Vaughan RW, Kondeatis E, Madanat W, Zureikat H, Fayyad F, Marr JE, Kanawati CA, Wallace GR, Stanford MR (2000) Intercellular adhesion molecule-1 gene polymorphisms in Behcet’s disease. Eur J Immunogenet 27:73–76
Wang K, Baldassano R, Zhang H, Qu HQ, Imielinski M, Kugathasan S, Annese V, Dubinsky M, Rotter JI, Russell RK, Bradfield JP, Sleiman PM, Glessner JT, Walters T, Hou C, Kim C, Frackelton EC, Garris M, Doran J, Romano C, Catassi C, Van Limbergen J, Guthery SL, Denson L, Piccoli D, Silverberg MS, Stanley CA, Monos D, Wilson DC, Griffiths A, Grant SF, Satsangi J, Polychronakos C, Hakonarson H (2010a) Comparative genetic analysis of inflammatory bowel disease and type 1 diabetes implicates multiple loci with opposite effects. Hum Mol Genet 19:2059–2067
Wang LY, Zhao DB, Gu J, Dai SM (2010b) Clinical characteristics of Behcet’s disease in China. Rheumatol Int 30:1191–1196
WTCCC (2007) Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447:661–678
Yang P, Fang W, Meng Q, Ren Y, Xing L, Kijlstra A (2008) Clinical features of chinese patients with Behcet’s disease. Ophthalmology 115(312–318):e4
Zhernakova A, Stahl EA, Trynka G, Raychaudhuri S, Festen EA, Franke L, Westra HJ, Fehrmann RS, Kurreeman FA, Thomson B, Gupta N, Romanos J, McManus R, Ryan AW, Turner G, Brouwer E, Posthumus MD, Remmers EF, Tucci F, Toes R, Grandone E, Mazzilli MC, Rybak A, Cukrowska B, Coenen MJ, Radstake TR, van Riel PL, Li Y, de Bakker PI, Gregersen PK, Worthington J, Siminovitch KA, Klareskog L, Huizinga TW, Wijmenga C, Plenge RM (2011) Meta-analysis of genome-wide association studies in celiac disease and rheumatoid arthritis identifies fourteen non-HLA shared loci. PLoS Genet 7:e1002004
Acknowledgments
The authors would like to thank all donors enrolled in the present study. This work was supported by the Natural Science Foundation Major International (RegionaJoint Research Project (30910103912), National Basic Research Program of China (973 Program) (2011CB510200), Key Project of Natural Science Foundation (81130019), National Natural Science Foundation Proj (30973242), Chongqing Key Laboratory of Ophthalmology (CSTC, 2008CA5003), Program for the Training of a Hundred Outstanding S&T Leaders of Chongqing Municipality and Fund for PAR-EU Scholars Program.
Conflict of interest
All authors do not have any conflict of interest to disclose.
Ethical standard
This study was conducted with the approval of the Ethical Committee of Chongqing Medical University.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, H., Liu, Q., Hou, S. et al. TNFAIP3 gene polymorphisms confer risk for Behcet’s disease in a Chinese Han population. Hum Genet 132, 293–300 (2013). https://doi.org/10.1007/s00439-012-1250-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-012-1250-7