Abstract
Common, but weakly penetrant, functional polymorphisms probably account for most of the genetic risk for breast cancer in the general population. Current polygenic risk models assume that component genes act independently. To test for potential gene–gene interactions, single nucleotide polymorphisms in ten genes with known or predicted roles in breast carcinogenesis were examined in a case-control study of 631 Caucasian women diagnosed with breast cancer under the age of 53 years and 1,504 controls under the age of 53 years. Association of breast cancer risk with individual genes and with two- and three-gene combinations was analyzed. Sixty-nine oligogenotypes from 37 distinct two- and three-gene combinations met stringent criteria for significance. Significant odds ratios (ORs) covered a 12-fold range: 0.5–5.9. Of the observed ORs, 17% differed significantly from the ORs predicted by a model of independent gene action, suggesting epistasis, i.e., that these genes interact to affect breast cancer risk in a manner not predictable from single gene effects. Exploration of the biological basis for these oligogenic interactions might reveal etiologic or therapeutic insights into breast cancer and other cancers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Relatively few breast cancers can be attributed to familial cancer predisposition syndromes that involve mutations in single, highly penetrant genes, such as BRCA1/2 or TP53 (Li-Fraumeni syndrome) (Evans et al. 2002; Miki et al. 1994; Sidransky et al. 1992; Wooster et al. 1995). These highly penetrant mutations are rare and together account only for about 5% of all breast cancers (Serova et al. 1997). They appear to have little role in sporadic breast cancers where moderate or no family history is present (Futreal et al. 1994). Consequently, risk assessment based on testing for these familial predisposition genes is uninformative for the majority of women (Malone et al. 1998; Newman et al. 1998). Although a number of twin studies clearly point to the significant role of genetics in the etiology of sporadic breast cancer (Lichtenstein et al. 2000; Peto and Mack 2000), the genetic contribution to the majority of breast cancers remains to be elucidated.
Many common polymorphisms have been examined for their association with breast cancer (reviewed in de Jong et al. 2002; Dunning et al. 1999). Generally, candidate polymorphisms are chosen for study based on probable roles in the biochemical or physiological pathways involved in breast carcinogenesis. Typically, the breast cancer risk conferred by these single polymorphisms is small to moderate, with odds ratios (ORs) that reach significance ranging from 1.3 to 2.5. Perhaps these results are not surprising because breast cancer is certainly a complex disease whose genetic risk is likely to be determined by multiple interactions among weakly penetrant polymorphisms (Antoniou et al. 2001; Pharoah et al. 2002). A few studies of limited numbers of candidate genes and often small sample sets have suggested that additive effects among common polymorphisms have a significant role in breast cancer risk (Comings et al. 2003; Feigelson and Henderson 2000; Feigelson et al. 2001; Fu et al. 2003; Huang et al. 1999). Before developing estimates of cancer risk based on multiple genes, it is important to evaluate empirically potential interactions between genes.
Here, we report the examination of ten common genetic polymorphisms in a single associative study of 631 breast cancer cases diagnosed before the age of 53 years and 1,504 cancer-free controls enrolled before age 53. The focus on women under 53 years of age arises from the generalization that genetic determinants are more obvious in young compared with older women; furthermore, earlier identification of women at high risk may change their age of first surveillance. We have studied candidate single nucleotide polymorphisms (SNPs) in ten genes likely to be involved in the physiological pathways that influence development of breast carcinoma. These include genes involved in cell cycle control (PHB, HER2, CCND1), carcinogen metabolism (SULT1A1, NQO1, GSTP1), DNA repair (ERCC2), and steroid hormone metabolism (VDR, CYP17, COMT). Whereas many more SNPs in the literature may meet these criteria, the ten SNPs chosen for this study were selected as being the most likely to influence breast cancer risk.
Most of the selected SNPs (HER2, SULT1A1, NQO1, GSTP1, ERCC2, COMT) lead to amino acid substitutions in proteins and either are known or are predicted to alter enzymatic or metabolic activity (see references in Table 1). The other SNPs are in promoter regions that regulate transcriptional activity (CYP17) or in transcribed non-coding regions that influence biological function, such as RNA splicing (CCND1), or regulatory RNA activity (PHB). For example, the promoter polymorphism in CYP17 has been associated with altered transcriptional efficiency leading to changes in circulating hormone levels (Feigelson et al. 1998; Haiman et al. 1999). The CCND1 variant A allele produces a truncated transcript, through alternative splicing, that has a longer than normal half life leading to the accumulation of cellular cyclin D1 (Betticher et al. 1995; Sawa et al. 1998). The PHB SNP is located in the 3′ untranslated region (UTR) and inactivates a novel regulatory RNA that functions as a tumor suppressor (Manjeshwar et al. 2003). The intronic variant in the VDR gene is the only polymorphism examined whose direct influence on physiology is not apparent. It is part of a VDR haplotype associated with alternative VDR signaling and activity as related to osteoporosis and breast cancer susceptibility (Bell et al. 2001; Curran et al. 1999; Ferrari et al. 1995; Friedrich et al. 2003; Hou et al. 2002). Thus, this ApaI polymorphism is used as a marker for the more extensive VDR haplotype.
The association of breast cancer risk with these SNPs singly and in combinations of two and three genes was evaluated. We also investigated whether there was evidence for epistasis, i.e., whether the breast cancer risk associated with the combination of SNPs differed from that predicted by the risks associated with the individual SNPs. The demonstration of epistasis is important for building an improved predictive model for breast cancer risk.
Materials and methods
Study participants
The current subjects were drawn from a study of 6,151 women, 1,716 with breast cancer and 4,435 controls. The participants included here were 2,135 Caucasian women representing 631 cases of breast cancer diagnosed before 53 years of age and 1,504 cancer-free controls interviewed before 53 years of age. All were enrolled in the Oklahoma area from 1996 to 2003. The threshold at age 53 years was chosen for this analysis because it was the average age of menopause in our sample, and because it was the approximate age at which allele frequencies were observed to differ between young and older women for several SNPs. Cases were defined as women with a self-reported diagnosis of breast cancer and were identified primarily from mammography centers and the offices of oncologists in the Oklahoma area. Controls were women who had never been diagnosed with any cancer and were identified primarily from mammography facilities in the Oklahoma area, from Komen Race for the Cure fund-raising events, and from several local medical clinics. To our knowledge, none were related. Informed consent was obtained from all study participants. Participants were assigned anonymous ID codes that were the sole identification used for questionnaires and biological samples. They completed a questionnaire concerning personal medical/health history and family history of cancer. The Institutional Review Boards of the Oklahoma Medical Research Foundation and the University of Oklahoma Health Sciences Center approved all grant-funded study protocols; InterGenetics study protocols were approved by the Research Consultants Review Committee (Austin, TX, USA).
Candidate polymorphisms
SNPs known or predicted to alter functional activity in ten candidate genes with a role in major pathways (cell cycle, carcinogen metabolism, DNA repair, steroid hormone metabolism, and signaling) that influence cancer development have been examined (Table 1). The exception is the polymorphism of unknown function located in intron 8 of VDR that is a haplotype marker previously associated with altered activity. The rest of these SNPs have been directly associated with enzymatic and/or physiological alterations and, thus, are not likely to be simply markers in linkage disequilibrium with the causative polymorphisms. All of these SNPs have been associated with breast cancer risk in at least one published study. For this reason, in this initial investigation of candidate genes with significant functional consequences, we have examined only one SNP in each gene as opposed to performing more extensive haplotype analyses of many SNPs in these candidate genes.
Genotyping assays
Genomic DNA was isolated from either buccal cells collected in Scope mouthwash or from venous blood by using the Gentra PureGene DNA extraction kit (Gentra, Minneapolis, MN, USA). The polymerase chain reactions (PCRs) were performed in either an Eppendorf Mastercycler or Perkin Elmer 9600 thermal cycler using either HotStarTaq DNA polymerase (QIAGEN, Valencia, CA, USA) or Ex-Taq DNA polymerase (PanVera, Madison, WI, USA). Annealing and extension temperatures were optimized for each primer set. The genotypes were determined by Luminex-based microbead methods (Iannone et al. 2000; Taylor et al. 2001) and/or PCR-restriction fragment length polymorphism (PCR-RFLP) assays. The primer sequences and specific genotyping conditions are available from the authors upon request. Genotypes were determined blinded to the case-control status of the participant.
When this project was begun, all genotyping was performed by RFLP analysis. Two investigators independently determined all genotype calls. Discrepancies were arbitrated by repeating the assays. More recently, genotyping was performed by ASPE by using the Luminex technology platform and scored on strict information criteria (available from the authors upon request). For quality assurance, 5%–10% of individuals were genotyped twice or more by one or both methods. Both genotyping methods were highly reproducible and concordant. Excluding the individuals who were only genotyped for PHB, 97% of controls and 95% of cases were successfully genotyped for at least nine of the ten polymorphisms examined.
Prior to the association analyses, compliance with Hardy–Weinberg (HW) frequency expectations was determined in the controls. Deviation from HW expectations could indicate technological errors. Chi-squared goodness-of-fit tests were used to determine the significance of deviations from HW expectation frequencies, calculated from allele frequencies estimated by gene-counting methods.
Statistical analyses
In this report, an “oligogenic combination” refers to a combination of two or more genes, such as PHB:CYP17:COMT. An “oligogenotype” refers to the set of specific SNP genotypes in the oligogenic combination, e.g., PHB C/C:CYP17 C/T:COMT G/A.
The principal statistic used in the analysis was the OR calculated for those carrying the oligogenotype (or genotype if only one gene is involved), namely the exposed at-risk group, versus those not carrying the oligogenotype, that is, the unexposed baseline group. This ratio approximated the OR for the risk of the oligogenotype compared with the population average risk. Dominance effects were also considered by examining combined genotypes such as PHB T/* that includes both T/T and T/C genotypes.
The association analyses proceeded in three steps. First, the calculated OR for an oligogenotype was compared with a null distribution of ORs. This null distribution was generated by randomizing the case-control status of the individuals in the study and by calculating the OR for this oligogenotype in each of the 10,000 randomized samples. An empirical estimate for the P-value for the observed OR was given by the proportion of ORs in the null distribution that were equal to or more extreme (further from 1.0) than the observed OR. For most of the oligogenotypes, this empirical P-value did not differ appreciably from the theoretical P-value calculated assuming large number theory. However, differences did occur for some of the less common oligogenotypes.
Second, resampling was performed to give an empirical estimate of the 95% confidence interval (95% CI) and a more reliable estimate of the OR for the oligogenotype, particularly for the less common oligogenotypes. The study sample was resampled 10,000 times with each resampling composed of 80% of the controls and 80% of the cases selected at random. The 95% CI for the OR of the oligogenotype was then defined as the 5th and 95th centile points of the distribution of ORs calculated from these resamplings. The mean of this distribution of ORs provided a more stable estimate of the true OR for an oligogenotype, especially for the rarer oligogenotypes for which a single estimate could be influenced markedly by a small random fluctuation. All ORs and 95% CIs reported in the text and in the tables are these resampled values.
Third, the frequencies of the oligogenotypes in the cases were tested for significant deviations from HW expectations. Deviation from HW expectations was further evidence of association with breast cancer risk. Oligogenotypes were selected as having significant influence on variation in breast cancer risk on the basis of: (criterion 1) an empirical P-value <0.0001; or (criterion 2) a P-value <0.05 for the test of deviation from HW equilibrium in the case population and an empirical P-value <0.002 (combined P=0.05×0.002=0.0001). For those oligogenic combinations for which at least one oligogenotype met the significance criteria, an overall significance for association of the oligogenic combination with breast cancer was calculated by using logistic regression and GLIM statistical software (Numerical Algorithms Group, Downers Grove, IL, USA). Estimates of minimum detectable ORs with 80% statistical power were calculated by using NCSS-PASS 2000 software (NCSS Statistical Software, Kaysville, UT, USA).
Analysis of epistasis between SNPs also used a resampling approach. The study sample was resampled 10,000 times with each resampling composed of 80% of the controls and 80% of the cases selected at random. For each resampling, the expected OR under the null hypothesis of no epistasis was calculated based on the product of the genotype frequencies for the component SNPs in the oligogenic combination. If there were no epistasis, one would expect that the risks estimated from the product of the frequencies of the individual SNP genotypes would approximate the observed risk of an oligogenotype composed of those SNP genotypes. An empirical estimate for the P-value for the observed interaction is given by the proportion of ORs in this null (no epistasis) distribution that are equal to or more extreme (further from 1.0) than the observed OR.
Results
Characteristics of study population
Selected demographic and risk factor data for the 631 cases and 1,504 controls are presented in Table 2. Other than family history of breast cancer in the controls, the proportions with established breast cancer risk factors were similar to those reported in the literature for both cancer cases and controls (Bernstein et al. 2002; Davis et al. 2002; Wrensch et al. 2003). Our population-based controls had a higher prevalence of history of breast cancer in a first-degree relative (28%) than the 3%–20% range in published reports (Bernstein et al. 2002; Colditz et al. 1993; Davis et al. 2002; Hall et al. 1993; Johnson et al. 1995; Slattery and Kerber 1993; Wrensch et al. 2003). This might be expected given that almost 18% of the controls were recruited from the Komen Race for the Cure and were probably motivated to participate in the present study because they had an affected relative and were concerned about their own breast cancer risk; indeed, 34% of the controls recruited from the Komen Race for the Cure had a history of breast cancer in a first-degree relative. However, even for those controls recruited from women visiting mammography clinics for routine checkups, the prevalence was higher (27%) than that reported in other studies. This high percentage of controls with a family history might be expected to attenuate the magnitude of the associations with breast cancer risk, leading to a more conservative analysis. On the other hand, there were no significant differences (P-values ranging from 0.09 to 0.92) in SNP allele frequencies between controls with a history of breast cancer in a first-degree relative and those without.
There were no significant differences between cases and controls with respect to family history of breast or other cancers. Earlier age at menarche and later age at the first birth showed the expected trends but were only marginally associated with increased risk. A personal history of benign breast disease (BBD) showed a strong association with increased risk (OR=1.9; 31% of breast cancer cases having a history of BBD compared with 19% of controls). Remaining nulliparous by 40 years of age was not associated with increased risk, contrary to expectations. Age at interview for the controls ranged from 18 to 52 years; age of diagnosis for the cases ranged from 15 to 52 years.
DNA samples were available for all 2,135 participants. However, not all women were successfully genotyped for all ten loci. Thirty-nine cases and 257 controls were typed for PHB only. For the remaining 592 cases and 1,247 controls, on average, 566 (96%) cases (range 553–580) and 1,211 (97%) controls (range 1,181–1,236) were genotyped for nine of the SNPs examined in this study. For the SNP in the 3′ UTR of PHB, 621 cases and 1,499 controls were genotyped. For the oligogenotypes involving two or three SNPs, the average number of complete oligogenotypes available for cases was 539 (range 522–574) and for controls was 1,168 (range 1,094–1,233). For all ten SNPs, 479 cases and 1,057 controls were genotyped. For the controls, none of the frequencies of individual SNPs showed significant deviation from HW expectations.
Oligogenic combinations associated with breast cancer risk
Altogether, 16,126 oligogenotypes from the 165 possible two- and three-gene oligogenic combinations were evaluated. After elimination of the 1,499 oligogenotypes with four or fewer individuals in either the cases or controls, 14,627 oligogenotypes were tested for significant association with breast cancer. There were 69 oligogenotypes, representing 37 distinct oligogenic combinations, that met our significance criteria (Table 3 Footnote 1). Thirty-four of these had at least one oligogenotype that met criteria 1, the more stringent of the criteria. The remaining three only had oligogenotypes that met criteria 2, which included consideration of deviation from HW equilibrium. Using a significance level of P≤0.0001, only 1.5 significant results (16,126−1,499=14,627×0.0001≈1.5) were expected by chance alone. Thus, despite testing multiple hypotheses, we observed far more oligogenotypes significantly associated with breast cancer risk than were expected by mere chance.
Approximately 3/4 of the oligogenic combinations in Table 3 (and in the larger dataset) showed an OR>1. This might encourage one to conclude that the majority of combinations lead to increased rather than reduced breast cancer risk. However, consideration of the significant oligogenotypes comprising each combination showed ORs that stratified risk over a broad range. For example, the oligogenic combination GSTP1:SULT1A1:ERCC2 had six oligogenotypes that met our significance criteria, with ORs ranging from 0.7 to 2.6, indicating reduced to increased risk. The full range of ORs across all possible oligogenotypes for this oligogenic combination, including those that did not reach our criteria for statistical significance, was 0.6–3.6 (Table 5). Clearly, focusing only upon the oligogenotypes in Table 3 ignores much of the power of an oligogenic combination to stratify risk. To illustrate this point further, Table 3 shows that the oligogenic combination PHB:CYP17:COMT carries only a moderate (but significant) risk (OR=1.6). In contrast, Table 4, which shows all oligogenotypes for this oligogenic combination, demonstrates a range of ORs from 0.2 to 8.9. To allow comparison with the ORs calculated in traditional single-gene analyses, Table 4 also shows the ORs (OR B ) calculated by using a designated no-risk (OR=1.0) baseline genotype, typically selected as the most common genotype in the control population (in this case PHB C/C:CYP17 C/T:COMT G/A). For BRCA1/2, the range of OR B is 9.9–33.0 (Antoniou et al. 2003), and so the maximum OR B of 10.8 in Table 4 suggests that oligogenic combinations involving as few as three genes can generate risks in the range reported for mutations in BRCA1/2. The full range of ORs observed for each of the oligogenic combinations listed in Table 3 is shown in Table 5. In addition, P-values for association of the oligogenic combination, across all oligogenotypes, with breast cancer are presented in Table 5.
Epistasis between SNPs
Table 3 compares the observed OR with the OR expected if each of the individual SNP genotypes confers breast cancer risk independently. The ratio between the observed and expected OR is given in the last column of Table 3 and demonstrates the level of epistasis. The empirical P-values calculated to measure the significance are also shown in Table 3. Of the 26 combinations presented in Table 3, 16 show evidence of epistasis at a P≤0.0001 level of significance. Overall, 27 (39%) of the 69 significant oligogenotypes exhibit evidence of epistasis at a P≤0.0001 level of significance, and 39 (57%) reveal evidence of epistasis at a P≤0.01 level of significance. For each of the significant oligogenic combinations, the number of oligogenotypes with empirical P-values ≤0.0001 is shown in Table 5. On average, 17% of oligogenotypes encompassed by the three-gene combinations shown in Table 5 exhibit significant evidence of epistasis. For comparison, 14% of all 14,627 oligogenotypes show significant evidence for epistasis (P≤0.0001). In total, these results indicate that risks associated with oligogenotypes cannot always be predicted from the risks associated with their component SNP genotypes.
Risk associated with single genes
To provide a comparison with other published studies, the ORs and associated empirical P-values for each gene considered alone are shown in Table 6 Footnote 2. None met our stringent significance criteria (P≤0.0001). However, PHB (T/*, OR=1.4, P=0.0003), ERCC2 (C/C, OR=1.6, P=0.0009), and COMT (G/*, OR=1.4, P=0.0041) meet the significance criteria used in most published studies. No others meet even the standard (P<0.05) level of significance. Table 6 includes the OR (OR B ) calculated when the most common homozygous genotype is selected as the baseline no-risk genotype (OR=1.0). Some of the results reported here cannot be directly compared with those in the literature because of differences in the characteristics of the study populations. Thus, our earlier published result for PHB (T/*, OR=4.8, P=0.003; Jupe et al. 2001) is not comparable because that result was based on subjects aged 50 years or younger and reporting a first-degree relative with breast cancer, unlike the OR reported here.
Discussion
The three main conclusions of this study are that oligogenic combinations are strongly and significantly associated with wide variation in breast cancer risk, that there is epistasis between genes in many of these oligogenotypes, and that these oligogenic combinations can stratify risk over a broader range than their component single genes considered independently.
Many oligogenotypes had significant and strong associations with variation in breast cancer risk, with ORs comparable with those seen for inherited mutations in BRCA1/2, even at the stringent level of significance (P≤0.0001) necessitated by our multiple hypothesis testing. We considered all possible combinations drawn two and three at a time from a set of ten SNPs in genes involved in cell cycle control, carcinogen metabolism, DNA repair, and steroid hormone metabolism and signaling. We also tested all possible oligogenotypes at these combinations with no a priori designation of a baseline no-risk oligogenotype. After eliminating from consideration those oligogenotypes with four or fewer individuals in either cases or controls, we evaluated 14,627 ORs. Of these, 69 were significant at a level of P≤0.0001. One would have expected perhaps at most two of the tested genotypes to be significant at this level of stringency just by chance, if no real associations existed. Since more than 30 times as many significant associations were observed than would be expected by random chance, we concluded that the majority of these significant oligogenotypes were true positives.
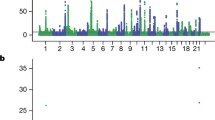
Testing multiple hypotheses can lead to high rates of false-positive findings. To avoid this problem, we chose a P-value ≤0.0001 as our criterion for accepting an association. This stringency risks the rejection of associations that might be true. The minimum ORs that can be detected with 80% power with our sample of 631 cases and 1,504 controls are shown in Fig. 1 for a range of oligogenotype frequencies. As indicated in Fig. 1, approximately 50% of the oligogenotypes had a frequency in the case sample of 0.08 or more for which an OR as low as 2.0 could be detected as significant at α=0.0001. The caveat is that the power calculations presented in Fig. 1 are based on large sample assumptions. However, Fig. 1 indicates that our study maintained reasonable statistical power despite the stringent level of significance. Thus, we have been able to show that two- and three-gene combinations can stratify breast cancer risk over a broad range.
Minimum detectable odds ratios (OR) with 80% power for 631 cases and 1,504 controls, and an α = 0.0001 level of significance for oligogenotype frequencies ranging from 0.01 to 0.50. Superimposed on this graph is a cumulative percent curve showing the percentage of oligogenotypes that have a frequency as large as or larger than indicated on the x-axis. This curve is cumulative from frequency 1.0 to 0.0. To show the way in which these curves may be interpreted, a line has been drawn upward from an oligogenotype frequency of 0.08. This intersects the minimum detectable OR curve at 2.0 and the cumulative percent curve at 50% indicating that, at a frequency of 0.08, an OR of 2.0 or larger can be detected as significant, and that 50% of the 16,126 oligogenotypes in this study have a frequency of 0.08 or larger.
Throughout, we have used empirical P-values to test oligogenotypes for significance to avoid making large-sample assumptions. However, examination of the theoretical P-values calculated based on large-sample assumptions for each oligogenotype shows that 60 of the 69 oligogenotypes listed in Table 3 are also significant at P≤0.0001, with none of the remaining nine combinations having a theoretical P-value greater than 0.0005 (data not shown). If we had chosen to use observed theoretical P-values as our measure of significance instead of empirically determined P-values, 71 combinations would have met our significance criteria (data not shown). All of the 11 (=71–60) oligogenotypes not meeting criteria by using empirical P-values identify an oligogenic combination listed in Table 5. Therefore, although theoretical and empirical P-values do not return exactly the same significant oligogenotypes, they identify the same oligogenic combinations.
The principal statistic used was the OR calculated for those carrying a particular oligogenotype, viz., the exposed at-risk group, versus those not carrying the oligogenotype, viz., the unexposed baseline group. This ratio approximated the OR for the risk of the oligogenotype compared with the population average risk. We chose these groups rather than designating a baseline no-risk oligogenotype to provide a risk estimate more applicable to genetic counseling. ORs based on a designated no-risk oligogenotype are perhaps most useful for an understanding of the biological basis for the altered risk, giving a more direct comparison between oligogenotypes. However, a difficulty arises in that the a priori choice of a baseline oligogenotype is not obvious.
The ORs reported here have not been adjusted for other risk factors such as age, age at onset of menarche, or age at birth of the first child. These factors have been omitted from the current analyses for two reasons. First, to some degree, limiting the study population to women under the age of 53 years, a largely pre-menopausal group, reduces the variation in risk attributable to age. We also note that, in Table 2, there is little difference between cases and controls in most of these other factors. Second, these factors have been omitted for computational and inferential simplicity. The specific consequences of this choice, although thought to be slight, are unpredictable. However, further analyses of larger data samples are needed to examine this.
Some of the associations in Table 3 may reflect survivorship rather than breast cancer risk. To examine this issue, we stratified the cases by those enrolled ≤24 months after diagnosis and those enrolled >24 months after diagnosis. Of the cases, 38% fell into the ≤24 months category; the lag time between diagnosis and enrollment ranged from 0 to 37 years (median 3.1 years). Analyses were repeated for each category. None of the 69 oligogenotypes showed appreciable differences in ORs between those interviewed ≤24 months and those interviewed >24 months after diagnosis (data not shown). These results suggest that the significant associations are not an artifact of survivorship.
For cases, the reference age was their age at diagnosis with breast cancer; for controls, the reference age was their age at recruitment. The present analyses included only women under 53 years of age. This stratification was chosen for its relationship to age of menopause, and because of the age-related changes in allele frequencies that we saw in preliminary tabulations by age. In the controls, the allele frequencies for several of these SNPs (VDR, CCND1, and CYP17) differed significantly between controls less than 45 years of age and those over 55 years, i.e., approximately the upper and lower age tertiles of the controls (data not shown). The reason for this is not known, although it may be attributable to competing causes of mortality, such as other cancers or heart disease, that are related to age and particularly to age of menopause. When we determined the age threshold that maximized the difference in SNP allele frequencies between young and older women, the average of these age thresholds across the SNPs was 52.3 years. Menopause is not an instantaneous event, and interventions such as surgical menopause and hormone replacement therapy confuse the issue further. The mean age of natural menopause, reported to be around 51–52 years (Bromberger et al. 1997; Gold et al. 2001; Kato et al. 1998), was 52.5 years of age in our sample. Thus, based on these values of 52.3 and 52.5 years, we chose an age stratification of under 53 years for our analysis.
To investigate the effect of our choice of age threshold, we reanalyzed our data using an age threshold of 50 years (under 51 years), the age most commonly used in the literature (Akhmedkhanov et al. 2000; Brandt et al. 2004; Breslow et al. 2001; Decensi et al. 2001; Rutter et al. 2003; Sidransky et al. 1992; Ursin et al. 2003; Wang-Gohrke et al. 2000; Weston et al. 1997). This analysis included 558 cases and 1,353 controls. The results were essentially unchanged by moving the age threshold. Of the oligogenotypes listed in Table 3, 46% also met our significance criteria in the under 51 years analysis (data not shown). Of the remainder, 64% and 81% had P≤0.0005 and 0.001, respectively, in the under 51 years analysis, and all had P≤0.01. This rise in P-value is not entirely unexpected given the decreasing sample sizes with the younger age threshold. Two oligogenotypes that did not appear in Table 3 had P≤0.0001 in the under 51 years analysis; both of these had P≤0.0002 in the under 53 years analysis. These two additional oligogenotypes did not add further oligogenic combinations to those previously seen in Table 5. We also considered an older threshold of 54 years, with similarly small differences in the results. Thus, the choice of age threshold, at least from under 51 years to under 54 years, did not affect the results appreciably.
The second observation from this study is that ORs associated with significant oligogenotypes frequently deviate significantly from the ORs that would be expected if their component SNP genotypes acted independently to affect risk. This deviation from independent interaction is often termed epistasis. Overall, 27 (39%) of the 69 significant oligogenotypes in Table 3 showed significant interaction at a P≤0.0001 level of significance (16 are shown in Table 3). Among all 14,627 oligogenotypes, 2,506 (17%) showed significant evidence of epistasis (P≤0.0001). These findings suggest that epistasis plays an important role in determining the breast cancer risk associated with these oligogenic combinations. However, these are statistical results, and eventually these oligogenic combinations will have to be examined to ensure that they make sense biologically.
Although the genome-wide prevalence of epistasis is not known, 17% is probably an overestimate. All of these genes and their polymorphisms have been shown to be associated with breast cancer risk in at least one other study. Because of this shared association, they might be more likely to show epistasis. On the other hand, this estimate may be an underestimate, since only multiplicative interactions are modeled. Biological interaction can yield either a multiplicative or an additive (albeit non-epistatic) statistical model (Cordell 2002), and for simplicity, the latter are not considered. However, in either case, epistasis is still clearly shown between these oligogenes.
The third observation is that oligogenic combinations significantly associated with breast cancer risk can be used to stratify risk further when all oligogenotypes within those gene combinations are considered. This is true even if the other oligogenotypes do not reach the threshold of significance. Thus, all oligogenotypes within a significant three-gene combination can be used to stratify risk over a broad range, as shown for PHB:CYP17:COMT in Table 4 in which the ORs range from 0.2 to 8.9. Risk stratification ranges for the 37 significant non-redundant oligogenic combinations are shown in Table 5. For the two-gene combination COMT:ERCC2, the range is fairly narrow varying from an OR of 0.8–1.9. Division of the largest OR by the lowest OR in this two-gene group shows that the oligogenotypes can stratify risk over a 2.5-fold range; the two-gene combination PHB:COMT returns ORs ranging from 0.3 to 1.8 representing a six-fold variation in risk. In comparison, the significant three-gene combinations return even wider variation in risk ranging from a modest four-fold for COMT:ERCC2:NQO1 to a 40-fold variation in risk for the combination PHB:COMT:SULT1A1. Thus, each oligogenic combination with at least one significant oligogenotype can stratify risk over a much broader range than that determined by considering the genes separately.
Confirmation of our results in additional large case-control studies and the addition of more SNPs to the epistatic analysis could lead to the development of a clinical genetic model to improve risk assessment for non-familial breast cancer. These combinations of low penetrance SNPs may also have relevance to breast cancers with a strongly familial basis. Highly penetrant mutations in the genes for breast cancer (BRCA1, BRCA2) do not always cause breast cancer. This lack of complete penetrance may be attributable in part to the same type of oligogenic interactions that we have described, in which breast cancer predisposition from BRCA1 and BRCA2 mutations is strongly modified by combinations of other low penetrance genes. Improved clinical risk assessment to identify those women at highest risk, regardless of family history, would assist in directing the use of mammography, sonography, and breast magnetic resonance imaging, perhaps leading to the earlier identification of breast cancer (Smith et al. 2003). As additional chemopreventatives are developed for breast cancer, a preventive care model can be envisioned that identifies high risk prior to diagnosis, that treats the patient prophylactically, and that avoids the disease directly.
Like breast cancer, most cancers are probably complex diseases with genetic components, and the approach of examining multiple genetic polymorphisms as performed here might also accurately assess predisposition for most other types of cancer. Moreoever, cardiovascular disease and diabetes probably have oligogenic influences predictable by this type of model (Yang et al. 2003). The etiology of breast cancer risk has always been alluded to as being complex, particularly for apparently “non-hereditary” breast cancer. However, our results suggest that, although the genetic etiology is complicated, many significant oligogenic associations can be identified when relatively large sample sets are analyzed.
Notes
Twenty-six of the 69 significant oligogenotypes are presented in Table 3. A complete table with all 69 significant oligogenotypes is available at http://www.intergenetics.com/intergenetics/publications.html.
Table 6 presents results only for the genes with significant association with variation in breast cancer risk. A complete table with the results for all ten genes (PHB, HER2, CCND1, COMT, SULT1A1, NQO1, GSTP1, ERCC2, VDR, CYP17, COMT) is available at http://www.intergenetics.com/intergenetics/publications.html.
References
Akhmedkhanov A, Toniolo P, Zeleniuch-Jacquotte A, Pettersson K, Huhtaniemi I (2000) Genetic variant of luteinizing hormone and risk of breast cancer in older women. Cancer Epidemiol Biomarkers Prev 9:839–842
Antoniou AC, Pharoah PD, McMullan G, Day NE, Ponder BA, Easton D (2001) Evidence for further breast cancer susceptibility genes in addition to BRCA1 and BRCA2 in a population-based study. Genet Epidemiol 21:1–18
Antoniou A, Pharoah PD, Narod S, Risch HA, Eyfjord JE, Hopper JL, Loman N, Olsson H, Johannsson O, Borg A, Pasini B, Radice P, Manoukian S, Eccles DM, Tang N, Olah E, Anton-Culver H, Warner E, Lubinski J, Gronwald J, Gorski B, Tulinius H, Thorlacius S, Eerola H, Nevanlinna H, Syrjakoski K, Kallioniemi OP, Thompson D, Evans C, Peto J, Lalloo F, Evans DG, Easton DF (2003) Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case series unselected for family history: a combined analysis of 22 studies. Am J Hum Genet 72:1117–1130
Bell NH, Morrison NA, Nguyen TV, Eisman J, Hollis BW (2001) ApaI polymorphisms of the vitamin D receptor predict bone density of the lumbar spine and not racial difference in bone density in young men. J Lab Clin Med 137:133–140
Bernstein L, Allen M, Anton-Culver H, Deapen D, Horn-Ross PL, Peel D, Pinder R, Reynolds P, Sullivan-Halley J, West D, Wright W, Ziogas A, Ross RK (2002) High breast cancer incidence rates among California teachers: results from the California Teachers Study (United States). Cancer Causes Control 13:625–635
Betticher DC, Thatcher N, Altermatt HJ, Hoban P, Ryder WD, Heighway J (1995) Alternate splicing produces a novel cyclin D1 transcript. Oncogene 11:1005–1011
Brandt B, Hermann S, Straif K, Tidow N, Buerger H, Chang-Claude J (2004) Modification of breast cancer risk in young women by a polymorphic sequence in the Egfr gene. Cancer Res 64:7–12
Breslow RA, Ballard-Barbash R, Munoz K, Graubard BI (2001) Long-term recreational physical activity and breast cancer in the National Health and Nutrition Examination Survey I epidemiologic follow-up study. Cancer Epidemiol Biomarkers Prev 10:805–808
Bromberger JT, Matthews KA, Kuller LH, Wing RR, Meilahn EN, Plantinga P (1997) Prospective study of the determinants of age at menopause. Am J Epidemiol 145:124–133
Colditz GA, Willett WC, Hunter DJ, Stampfer MJ, Manson JE, Hennekens CH, Rosner BA (1993) Family history, age, and risk of breast cancer. Prospective data from the nurses’ health study. JAMA 270:338–343
Comings DE, Gade-Andavolu R, Cone L, Muhleman D, MacMurray JP (2003) A multigene test for the risk of sporadic breast carcinoma. Cancer 97:2160–2170
Cordell HJ (2002) Epistasis: what it means, what it doesn’t mean, and statistical methods to detect it in humans. Hum Mol Genet 11:2463–2468
Curran JE, Vaughan T, Lea RA, Weinstein SR, Morrison NA, Griffiths LR (1999) Association of a vitamin D receptor polymorphism with sporadic breast cancer development. Int J Cancer 83:723–726
Davis S, Mirick DK, Stevens RG (2002) Residential magnetic fields and the risk of breast cancer. Am J Epidemiol 155:446–454
Decensi A, Johansson H, Miceli R, Mariani L, Camerini T, Cavadini E, Di Mauro MG, Barreca A, Gonzaga AG, Diani S, Sandri MT, De Palo G, Formelli F (2001) Long-term effects of fenretinide, a retinoic acid derivative, on the insulin-like growth factor system in women with early breast cancer. Cancer Epidemiol Biomarkers Prev 10:1047–1053
Dunning AM, Healey CS, Pharoah PD, Teare MD, Ponder BA, Easton DF (1999) A systematic review of genetic polymorphisms and breast cancer risk. Cancer Epidemiol Biomarkers Prev 8:843–854
Evans DG, Birch JM, Thorneycroft M, McGown G, Lalloo F, Varley JM (2002) Low rate of TP53 germline mutations in breast cancer/sarcoma families not fulfilling classical criteria for Li-Fraumeni syndrome. J Med Genet 39:941–944
Falany CN (1997) Enzymology of human cytosolic sulfotransferases. FASEB J 11:206–216
Feigelson HS, Henderson BE (2000) Future possibilities in the prevention of breast cancer: role of genetic variation in breast cancer prevention. Breast Cancer Res 2:277–282
Feigelson HS, Shames LS, Pike MC, Coetzee GA, Stanczyk FZ, Henderson BE (1998) Cytochrome P450c17alpha gene (CYP17) polymorphism is associated with serum estrogen and progesterone concentrations. Cancer Res 58:585–587
Feigelson HS, McKean-Cowdin R, Coetzee GA, Stram DO, Kolonel LN, Henderson BE (2001) Building a multigenic model of breast cancer susceptibility: CYP17 and HSD17B1 are two important candidates. Cancer Res 61:785–789
Ferrari S, Rizzoli R, Chevalley T, Slosman D, Eisman JA, Bonjour JP (1995) Vitamin-D-receptor-gene polymorphisms and change in lumbar-spine bone mineral density. Lancet 345:423–424
Friedrich M, Rafi L, Mitschele T, Tilgen W, Schmidt W, Reichrath J (2003) Analysis of the vitamin D system in cervical carcinomas, breast cancer and ovarian cancer. Recent Results Cancer Res 164:239–246
Fu YP, Yu JC, Cheng TC, Lou MA, Hsu GC, Wu CY, Chen ST, Wu HS, Wu PE, Shen CY (2003) Breast cancer risk associated with genotypic polymorphism of the nonhomologous end-joining genes: a multigenic study on cancer susceptibility. Cancer Res 63:2440–2446
Futreal PA, Liu Q, Shattuck-Eidens D, Cochran C, Harshman K, Tavtigian S, Bennett LM, Haugen-Strano A, Swensen J, et al (1994) BRCA1 mutations in primary breast and ovarian carcinomas. Science 266:120–122
Gold EB, Bromberger J, Crawford S, Samuels S, Greendale GA, Harlow SD, Skurnick J (2001) Factors associated with age at natural menopause in a multiethnic sample of midlife women. Am J Epidemiol 153:865–874
Goodman JE, Lavigne JA, Wu K, Helzlsouer KJ, Strickland PT, Selhub J, Yager JD (2001) COMT genotype, micronutrients in the folate metabolic pathway and breast cancer risk. Carcinogenesis 22:1661–1665
Gudmundsdottir K, Tryggvadottir L, Eyfjord JE (2001) GSTM1, GSTT1, and GSTP1 genotypes in relation to breast cancer risk and frequency of mutations in the p53 gene. Cancer Epidemiol Biomarkers Prev 10:1169–1173
Haiman CA, Hankinson SE, Spiegelman D, Colditz GA, Willett WC, Speizer FE, Kelsey KT, Hunter DJ (1999) The relationship between a polymorphism in CYP17 with plasma hormone levels and breast cancer. Cancer Res 59:1015–1020
Hall JA, Murphy DC, Hall BR, Hall KA (1993) Mammographic abnormalities and the detection of carcinoma of the breast. Am J Obstet Gynecol 168:1677–1680
Helzlsouer KJ, Selmin O, Huang HY, Strickland PT, Hoffman S, Alberg AJ, Watson M, Comstock GW, Bell D (1998) Association between glutathione S-transferase M1, P1, and T1 genetic polymorphisms and development of breast cancer. J Natl Cancer Inst 90:512–518
Hou MF, Tien YC, Lin GT, Chen CJ, Liu CS, Lin SY, Huang TJ (2002) Association of vitamin D receptor gene polymorphism with sporadic breast cancer in Taiwanese patients. Breast Cancer Res Treat 74:1–7
Huang CS, Chern HD, Chang KJ, Cheng CW, Hsu SM, Shen CY (1999) Breast cancer risk associated with genotype polymorphism of the estrogen-metabolizing genes CYP17, CYP1A1, and COMT: a multigenic study on cancer susceptibility. Cancer Res 59:4870–4875
Iannone MA, Taylor JD, Chen J, Li MS, Rivers P, Slentz-Kesler KA, Weiner MP (2000) Multiplexed single nucleotide polymorphism genotyping by oligonucleotide ligation and flow cytometry. Cytometry 39:131–140
Johansson AS, Stenberg G, Widersten M, Mannervik B (1998) Structure-activity relationships and thermal stability of human glutathione transferase P1–1 governed by the H-site residue 105. J Mol Biol 278:687–698
Johnson N, Lancaster T, Fuller A, Hodgson SV (1995) The prevalence of a family history of cancer in general practice. Fam Pract 12:287–289
Jong MM de, Nolte IM, Meerman GJ te, Graaf WT van der, Oosterwijk JC, Kleibeuker JH, Schaapveld M, Vries EG de (2002) Genes other than BRCA1 and BRCA2 involved in breast cancer susceptibility. J Med Genet 39:225–242
Jupe ER, Badgett AA, Neas BR, Craft MA, Mitchell DS, Resta R, Mulvihill JJ, Aston CE, Thompson LF (2001) Single nucleotide polymorphism in prohibitin 3′ untranslated region and breast-cancer susceptibility. Lancet 357:1588–1589
Kato I, Toniolo P, Akhmedkhanov A, Koenig KL, Shore R, Zeleniuch-Jacquotte A (1998) Prospective study of factors influencing the onset of natural menopause. J Clin Epidemiol 51:1271–1276
Kong S, Wei Q, Amos CI, Lynch PM, Levin B, Zong J, Frazier ML (2001) Cyclin D1 polymorphism and increased risk of colorectal cancer at young age. J Natl Cancer Inst 93:1106–1108
Lachman HM, Papolos DF, Saito T, Yu YM, Szumlanski CL, Weinshilboum RM (1996) Human catechol-O-methyltransferase pharmacogenetics: description of a functional polymorphism and its potential application to neuropsychiatric disorders. Pharmacogenetics 6:243–250
Lavigne JA, Helzlsouer KJ, Huang HY, Strickland PT, Bell DA, Selmin O, Watson MA, Hoffman S, Comstock GW, Yager JD (1997) An association between the allele coding for a low activity variant of catechol-O-methyltransferase and the risk for breast cancer. Cancer Res 57:5493–5497
Lichtenstein P, Holm NV, Verkasalo PK, Iliadou A, Kaprio J, Koskenvuo M, Pukkala E, Skytthe A, Hemminki K (2000) Environmental and heritable factors in the causation of cancer–analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med 343:78–85
Lunn RM, Helzlsouer KJ, Parshad R, Umbach DM, Harris EL, Sanford KK, Bell DA (2000) XPD polymorphisms: effects on DNA repair proficiency. Carcinogenesis 21:551–555
Malone KE, Daling JR, Thompson JD, O’Brien CA, Francisco LV, Ostrander EA (1998) BRCA1 mutations and breast cancer in the general population: analyses in women before age 35 years and in women before age 45 years with first-degree family history. JAMA 279:922–929
Manjeshwar S, Branam DE, Lerner MR, Brackett DJ, Jupe ER (2003) Tumor suppression by the prohibitin gene 3′untranslated region RNA in human breast cancer. Cancer Res 63:5251–5256
McKean-Cowdin R, Kolonel LN, Press MF, Pike MC, Henderson BE (2001) Germ-line HER-2 variant and breast cancer risk by stage of disease. Cancer Res 61:8393–8394
Miki Y, Swensen J, Shattuck-Eidens D, Futreal PA, Harshman K, Tavtigian S, Liu Q, Cochran C, Bennett LM, Ding W, et al (1994) A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266:66–71
Mitrunen K, Jourenkova N, Kataja V, Eskelinen M, Kosma VM, Benhamou S, Kang D, Vainio H, Uusitupa M, Hirvonen A (2001) Polymorphic catechol-O-methyltransferase gene and breast cancer risk. Cancer Epidemiol Biomarkers Prev 10:635–640
Morgan J, Siegel D, Ross D (1999) A potential mechanism underlying the increased susceptibility of individuals with a polymorphism in NAD(P)H;quinone oxidoreductase 1 (NQO1) to benzene toxicity. Proc Natl Acad Sci USA 96:8150–8155
Newman B, Mu H, Butler LM, Millikan RC, Moorman PG, King MC (1998) Frequency of breast cancer attributable to BRCA1 in a population-based series of American women. JAMA 279:915–921
Palli D, Russo A, Masala G, Saieva C, Guarrera S, Carturan S, Munnia A, Matullo G, Peluso M (2001) DNA adduct levels and DNA repair polymorphisms in traffic-exposed workers and a general population sample. Int J Cancer 94:121–127
Peto J, Mack TM (2000) High constant incidence in twins and other relatives of women with breast cancer. Nat Genet 26:411–414
Pharoah PD, Antoniou A, Bobrow M, Zimmern RL, Easton DF, Ponder BA (2002) Polygenic susceptibility to breast cancer and implications for prevention. Nat Genet 31:33–36
Raftogianis RB, Wood TC, Otterness DM, Van Loon JA, Weinshilboum RM (1997) Phenol sulfotransferase pharmacogenetics in humans: association of common SULT1A1 alleles with TS PST phenotype. Biochem Biophys Res Commun 239:298–304
Rutter JL, Chatterjee N, Wacholder S, Struewing J (2003) The HER2 I655V polymorphism and breast cancer risk in Ashkenazim. Epidemiology 14:694–700
Sawa H, Ohshima TA, Ukita H, Murakami H, Chiba Y, Kamada H, Hara M, Saito I (1998) Alternatively spliced forms of cyclin D1 modulate entry into the cell cycle in an inverse manner. Oncogene 16:1701–1712
Serova OM, Mazoyer S, Puget N, Dubois V, Tonin P, Shugart YY, Goldgar D, Narod SA, Lynch HT, Lenoir GM (1997) Mutations in BRCA1 and BRCA2 in breast cancer families: are there more breast cancer-susceptibility genes? Am J Hum Genet 60:486–495
Seth P, Lunetta KL, Bell DW, Gray H, Nasser SM, Rhei E, Kaelin CM, Iglehart DJ, Marks JR, Garber JE, Haber DA, Polyak K (2000) Phenol sulfotransferases: hormonal regulation, polymorphism, and age of onset of breast cancer. Cancer Res 60:6859–6863
Sidransky D, Tokino T, Helzlsouer K, Zehnbauer B, Rausch G, Shelton B, Prestigiacomo L, Vogelstein B, Davidson N (1992) Inherited p53 gene mutations in breast cancer. Cancer Res 52:2984–2986
Siegel D, McGuinness SM, Winski SL, Ross D (1999) Genotype-phenotype relationships in studies of a polymorphism in NAD(P)H:quinone oxidoreductase 1. Pharmacogenetics 9:113–121
Slattery ML, Kerber RA (1993) A comprehensive evaluation of family history and breast cancer risk. The Utah Population Database. JAMA 270:1563–1568
Smith MT, Wang Y, Kane E, Rollinson S, Wiemels JL, Roman E, Roddam P, Cartwright R, Morgan G (2001) Low NAD(P)H:quinone oxidoreductase 1 activity is associated with increased risk of acute leukemia in adults. Blood 97:1422–1426
Smith RA, Saslow D, Sawyer KA, Burke W, Costanza ME, Evans WP 3rd, Foster RS Jr, Hendrick E, Eyre HJ, Sener S (2003) American Cancer Society guidelines for breast cancer screening: update 2003. CA Cancer J Clin 53:141-169
Spurdle AB, Hopper JL, Dite GS, Chen X, Cui J, McCredie MR, Giles GG, Southey MC, Venter DJ, Easton DF, Chenevix-Trench G (2000) CYP17 promoter polymorphism and breast cancer in Australian women under age forty years. J Natl Cancer Inst 92:1674–1681
Tang D, Cho S, Rundle A, Chen S, Phillips D, Zhou J, Hsu Y, Schnabel F, Estabrook A, Perera FP (2002) Polymorphisms in the DNA repair enzyme XPD are associated with increased levels of PAH-DNA adducts in a case-control study of breast cancer. Breast Cancer Res Treat 75:159–166
Taylor JD, Briley D, Nguyen Q, Long K, Iannone MA, Li MS, Ye F, Afshari A, Lai E, Wagner M, Chen J, Weiner MP (2001) Flow cytometric platform for high-throughput single nucleotide polymorphism analysis. Biotechniques 30:661–669
Thompson PA, Shields PG, Freudenheim JL, Stone A, Vena JE, Marshall JR, Graham S, Laughlin R, Nemoto T, Kadlubar FF, Ambrosone CB (1998) Genetic polymorphisms in catechol-O-methyltransferase, menopausal status, and breast cancer risk. Cancer Res 58:2107–2110
Tommasi S, Fedele V, Lacalamita R, Crapolicchio A, Perlino E, Bellizzi A, Paradiso A (2004) Molecular and functional characteristics of erbB2 in normal and cancer breast cells. Cancer Lett 209:215–222
Ursin G, Ma H, Wu AH, Bernstein L, Salane M, Parisky YR, Astrahan M, Siozon CC, Pike MC (2003) Mammographic density and breast cancer in three ethnic groups. Cancer Epidemiol Biomarkers Prev 12:332–338
Vodicka P, Kumar R, Stetina R, Sanyal S, Soucek P, Haufroid V, Dusinska M, Kuricova M, Zamecnikova M, Musak L, Buchancova J, Norppa H, Hirvonen A, Vodickova L, Naccarati A, Matousu Z, Hemminki K (2004) Genetic polymorphisms in DNA repair genes and possible links with DNA repair rates, chromosomal aberrations and single-strand breaks in DNA. Carcinogenesis 25:757–763
Wang L, Habuchi T, Takahashi T, Mitsumori K, Kamoto T, Kakehi Y, Kakinuma H, Sato K, Nakamura A, Ogawa O, Kato T (2002) Cyclin D1 gene polymorphism is associated with an increased risk of urinary bladder cancer. Carcinogenesis 23:257–264
Wang L, Habuchi T, Mitsumori K, Li Z, Kamoto T, Kinoshita H, Tsuchiya N, Sato K, Ohyama C, Nakamura A, Ogawa O, Kato T (2003) Increased risk of prostate cancer associated with AA genotype of cyclin D1 gene A870G polymorphism. Int J Cancer 103:116–120
Wang-Gohrke S, Chang-Claude J, Becher H, Kieback DG, Runnebaum IB (2000) Progesterone receptor gene polymorphism is associated with decreased risk for breast cancer by age 50. Cancer Res 60:2348–2350
Watson MA, Stewart RK, Smith GB, Massey TE, Bell DA (1998) Human glutathione S-transferase P1 polymorphisms: relationship to lung tissue enzyme activity and population frequency distribution. Carcinogenesis 19:275–280
Weston A, Pan CF, Ksieski HB, Wallenstein S, Berkowitz GS, Tartter PI, Bleiweiss IJ, Brower ST, Senie RT, Wolff MS (1997) p53 haplotype determination in breast cancer. Cancer Epidemiol Biomarkers Prev 6:105–112
Wooster R, Bignell G, Lancaster J, Swift S, Seal S, Mangion J, Collins N, Gregory S, Gumbs C, Micklem G (1995) Identification of the breast cancer susceptibility gene BRCA2. Nature 378:789–791
Wrensch M, Chew T, Farren G, Barlow J, Belli F, Clarke C, Erdmann CA, Lee M, Moghadassi M, Peskin-Mentzer R, Quesenberry CP Jr, Souders-Mason V, Spence L, Suzuki M, Gould M (2003) Risk factors for breast cancer in a population with high incidence rates. Breast Cancer Res 5:R88–R102
Xie D, Shu XO, Deng Z, Wen WQ, Creek KE, Dai Q, Gao YT, Jin F, Zheng W (2000) Population-based, case-control study of HER2 genetic polymorphism and breast cancer risk. J Natl Cancer Inst 92:412–417
Yang Q, Khoury MJ, Botto L, Friedman JM, Flanders WD (2003) Improving the prediction of complex diseases by testing for multiple disease-susceptibility genes. Am J Hum Genet 72:636-649
Ye Z, Parry JM (2002) The CYP17 MspA1 polymorphism and breast cancer risk: a meta-analysis. Mutagenesis 17:119–126
Zheng W, Xie D, Cerhan JR, Sellers TA, Wen W, Folsom AR (2001a) Sulfotransferase 1A1 polymorphism, endogenous estrogen exposure, well-done meat intake, and breast cancer risk. Cancer Epidemiol Biomarkers Prev 10:89–94
Zheng Y, Shen H, Sturgis EM, Wang LE, Eicher SA, Strom SS, Frazier ML, Spitz MR, Wei Q (2001b) Cyclin D1 polymorphism and risk for squamous cell carcinoma of the head and neck: a case-control study. Carcinogenesis 22:1195–1199
Zimniak P, Nanduri B, Pikula S, Bandorowicz-Pikula J, Singhal SS, Srivastava SK, Awasthi S, Awasthi YC (1994) Naturally occurring human glutathione S-transferase GSTP1-1 isoforms with isoleucine and valine in position 104 differ in enzymic properties. Eur J Biochem 224:893–899
Acknowledgements
This work was supported by funding from the US Army Breast Cancer Research Program (E.R.J.), the American Cancer Society (E.R.J.), Oklahoma Center for the Advancement of Science and Technology (E.R.J. and D.A.R.), the University of Oklahoma Health Sciences Center General Clinical Research Center, the National Center for Research Resources—NIH, The Presbyterian Health Foundation, The Oklahoma Life Sciences Fund, and the Swisher Family Trust. We thank A. Whelan, A. Lee, and B. Howry for technical assistance, M. Bondy for critical comments, and the many clinicians and women who participated. The Molecular Biology Resource Facility at the University of Oklahoma Health Sciences Center provided oligonucleotide synthesis services. L.F.T. holds the Putnam City Schools Chair in Cancer Research. J.J.M. holds the Kimberly V. Talley/Children’s Medical Research Institute Chair in Genetics.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aston, C.E., Ralph, D.A., Lalo, D.P. et al. Oligogenic combinations associated with breast cancer risk in women under 53 years of age. Hum Genet 116, 208–221 (2005). https://doi.org/10.1007/s00439-004-1206-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-004-1206-7