Abstract
Developmental dyslexia, or reading disability, is a multigenic complex disease for which at least five loci, i.e. DYX1–3 and DYX5–6, have been clearly identified from the human genome. To date, DYX1C1 is the only dyslexia candidate gene cloned. We have previously reported linkage to 2p11 and 7q32 in 11 Finnish pedigrees. Here, we report the fine mapping of the approximately 40-cM linked region from chromosome 2 as we increased marker density to one per 1.8 cM. Linkage was supported with the highest NPL score of 3.0 (P=0.001) for marker D2S2216. Association analysis using the six pedigrees showing linkage pointed to marker D2S286/rs3220265 (P value <0.001) in the near vicinity of D2S2216. We went on to further characterise this ~15-cM candidate region (D2S2110-D2S2181) by adding six SNPs covering ~670 kb centred at D2S286/rs3220265. A haplotype pattern could no longer be observed in this region, which was therefore excluded from the candidate area. This also excluded the TACR1 (tachykinin receptor 1) gene, located at marker D2S286. The dyslexia candidate region on 2p11 is, therefore, now limited to the chromosomal area D2S2116-D2S2181, which is ~12 Mbp of human sequence and is at a distinct location from the previously reported DYX3 locus, raising the possibility of two distinct loci on chromosome 2p.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dyslexia, or specific or developmental reading disability, is defined by an unexpected failure in learning to read, write or spell in spite of normal senses, normal intelligence, and adequate opportunity and motivation. Dyslexia is a multifactorial, or complex phenotype, the genetic basis of which has been established in a number of twin- and family-based studies (for review, see Fisher and DeFries 2002; Grigorenko 2001). In addition to its complex etiology, dyslexia displays a wide spectrum of phenotypes, which could also reflect incomplete penetrance, and/or the effects of influencing environmental factors. To date, at least five loci have been identified, i.e. DYX1 on 15q21 (OMIM 127700 at http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=OMIM) with the newly identified DYX1C1 gene (Taipale et al. 2003), DYX2 on 6p21.3 (OMIM 600202), DYX3 on 2p16-p15 (OMIM 604254), DYX5 on 3p12-q13 (OMIM 606896) and DYX6 on 18p11.2 (OMIM 606616) (Cardon et al. 1994; Fagerheim et al. 1999; Fisher et al. 2002; Nopola-Hemmi et al. 2001; Petryshen et al. 2001; Schulte-Korne et al. 1998). Recently, genome-wide scans performed in English- (Fisher et al. 2002) and Finnish-speaking (Kaminen et al. 2003) populations not only suggested two new reading disability loci, i.e. 18p (DYX6) and 7q, but also strengthened the evidence for a dyslexia locus on 2p, located about 20–30 cM centromeric to DYX3 (Kaminen et al. 2003). Here, we report the fine mapping of this locus using the same set of Finnish families. Twenty-one microsatellite markers spread along an approximately 40-cM interval (D2S391-D2S2181) and six additional SNP markers located over ~670 kb of sequence around D2S286/rs3220265 were genotyped and the candidate region was refined to an approximately 10.5-cM interval between D2S2116 and D2S2181.
The TACR1 (tachykinin receptor 1) gene, encoding the G-protein-coupled receptor for tachykinin substance P/neurokinin 1 and involved in the modulation of neuronal activity, inflammation and mood (OMIM 162323, LocusID 6869 at http://www.ncbi.nlm.nih.gov/LocusLink) (De Felipe et al. 1998; Derocq et al. 1996; Kramer et al. 1998), was tested for mutation as it represented an excellent positional candidate gene (Gerard et al. 1991; Hopkins et al. 1991). This gene was excluded as a dyslexia gene on 2p11, as no coding mutations were detected in affected individuals. This exclusion was fully confirmed by a lack of haplotype conservation using six SNPs located within the gene and its surroundings.
Materials and methods
Subjects and genomic DNA preparation
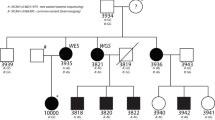
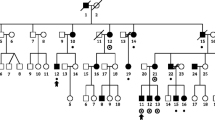
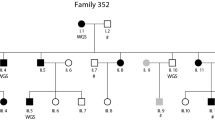
Eleven three-generation families (97 individuals), with at least two affected individuals/family (see pedigree description in Kaminen et al. 2003), were studied. Eighty-eight subjects were available for genotyping: 37 affected, 38 healthy and 13 not tested for dyslexia (phenotype unknown). Phenotypes were ascertained as described in Nopola-Hemmi et al. (2001). One affected from each of the pedigrees displaying positive linkage to 2p11 (nos. 1, 4, 9, 10 and 11 in Kaminen et al. 2003) and two affected unrelated individuals originating from Central Finland were screened for mutation in the TACR1 gene. Two unaffected individuals from those families were used as controls. For all individuals, genomic DNA was obtained from blood lymphocytes using a standard non-enzymatic extraction method (Lahiri and Nurnberger 1991). The study has been approved by the local ethical committees both in Finland and in Sweden.
Genotyping
Microsatellite markers
Eighteen microsatellite markers were successfully used for genotyping as described previously (Kaminen et al. 2003). Briefly, each marker was PCR-amplified from 10 ng of genomic DNA in 5-μl reactions, which were thereafter pooled (9–13 markers/pool) and electrophoresed on a MegaBACE 1000 instrument (Amersham Biosciences). Alleles were visualised using the Genetic Profiler v1.5 software (Amersham Biosciences).
SNPs markers
SNP markers rs1487371, rs754978, rs1477157, rs2016839 and rs718507 were chosen from the SNP consortium (TSC) database for having reported minor allele frequency of at least 30% in Caucasian populations (http://www.snp.cshl.org). SNP rs6715729 was chosen for its location in the coding part of the first exon of the TACR1 gene. All SNPs (http://www.ncbi.nlm.nih.gov/SNP/index.html, http://www.snp.cshl.org) were genotyped in all 88 DNA samples using matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry (Sequenom, San Diego, Calif.) (Jurinke et al. 2002). PCR assays and associated extension reactions were designed using the SpectroDESIGNER software (Sequenom) and primers were obtained from Metabion (Planegg-Martinsried, Germany) (sequences available on request). All amplification reactions were run in a total volume of 5 μl with 2.5 ng of genomic DNA, 1 pmol of each amplification primer, 0.2 mM of each dNTP, 2.5 mM MgCl2 and 0.2 U of HotStarTaq DNA polymerase (Qiagen). Reactions were heated at 95 °C for 15 min, subjected to 45 cycles of amplification (20 s at 94 °C, 30 s at 60 °C, 30 s at 72 °C) before a final extension of 10 min at 72 °C. Extension reactions were conducted in a total volume of 2 μl, using 5 pmol of allele-specific extension primer and the Mass EXTEND Reagents Kit, before being cleaned using SpectroCLEANER (Sequenom) on a MULTIMEK 96 robot (Beckman Coulter, Fullerton, Calif.). Clean products were loaded onto a 384-elements chip with a nanolitre pipetting system (SpectroCHIP, SpectroJet, Sequenom), analysed by a MassARRAY mass spectrometer (Bruker Daltonik, Bremen, Germany) and peaks were identified using the SpectroTYPER RT 2.0 software (Sequenom).
Two researchers independently confirmed all genotypes (microsatellite and SNP markers). Data were checked for Mendelian errors using PedCheck (O’Connell and Weeks 1998) and for Hardy-Weinberg equilibrium to identify genotyping errors.
Nomenclature
Gene symbols used in this article follow the recommendations of the HUGO Gene Nomenclature Committee (Povey et al. 2001).
Statistical methods
Linkage analysis
Non-parametric multipoint linkage analysis was performed with Genehunter 2.1 software (Kruglyak et al. 1996) for the 11 pedigrees. The significance of the NPL score was evaluated by simulation, simulating marker genotypes according to the null hypothesis distribution, in 100 replications, and calculating NPL scores for each of the replicates. The empirical significance of the observed NPL score distribution is given as the proportion of simulated scores that are higher than the observed NPL. Data for two markers (D2S2180 and D2S2333) were excluded from the linkage analysis. Their inclusion yielded multiple double-recombinant interpretations and inflation of the genetic map suggesting errors in allele calling and/or marker instability.
Association analysis
Genetic associations were analysed by the haplotype pattern mining (HPM) method (Toivonen et al. 2000). HPM is a data-mining-based approach to genetic association analysis, in which frequent haplotype patterns associated with a trait are sought. HPM analyses case-control data using non-transmitted parental chromosomes as controls (i.e., pseudo-controls). Independent family trios were extracted and haplotyped from the pedigrees using an algorithm which picks up all possible non-overlapping trios, even if they are only partly haplotyped and/or genotyped. The association analysis was thus carried out in a case-control set consisting of 43 disease-associated chromosomes and 29 control chromosomes and repeated in a subset of data consisting of only those trios from pedigrees showing a positive NPL score in linkage analysis (pedigrees 1, 4, 5, 9, 10 and 11 in Kaminen et al. 2003). This data subset comprised 28 disease-associated and 20 control chromosomes. HPM was carried out for the whole chromosome, first including all 45 microsatellite markers alone and then adding to these, the six SNPs surrounding marker D2S286/rs3220265. The following parameters were used: maximum length of the pattern, eight markers; maximum number of gaps, one; minimum χ2 for a pattern, 2. To compensate for differing marker densities and marker information along the studied chromosomal area, a total of 10,000 permutations in the same fashion as described previously (Toivonen et al. 2000), were run to obtain empirical P values.
Expression analysis
Exons 1–5 of the TACR1 gene were separately PCR-amplified from 20 ng of genomic DNA from a normal individual using the following conditions: 1.5 mM MgCl2, 1 μM of each primer (sequences available upon request), 0.2 mM of each dNTP and 0.04 U/μl of AmpliTaq DNA polymerase (Applied Biosystems). Reactions were heated at 94 °C for 5 min, subjected to 35 cycles of amplification (45 s at 94 °C, 45 s at 42–62 °C and 1 min at 72 °C) before a final extension of 10 min at 72 °C. Each exon was then re-amplified (same conditions as above, 25 cycles) separately in a 10-μl reaction volume containing 1–2 μl of the first PCR reaction, 0.2 mM of each dNTP (dATP, dTTP and dGTP), 1.5 mM MgCl2, 1 μM of each primer, 0.04 U/μl of AmpliTaq DNA polymerase (Applied Biosystems) and 50 μCi P32-dCTP. The radioactively labelled products were then pooled, cleaned from unincorporated P32-dCTP and used as probe on a human multiple tissues Northern blot containing 2 μg of poly(A) RNA from brain, placenta, skeletal muscle, heart, kidney, pancreas, liver, lung, spleen and thymus (cat. no. 3140-1, Ambion). Pre-hybridisation, hybridisation and washing of the membrane were done according to the manufacturer’s recommendations. Exposure to Hyperfilm MP (Amersham Biosciences) was done for two weeks.
Mutation screening
Exons 1–5 of the human TACR1 gene were PCR-amplified (products ranging from 382 to 622 bp) as described above in the primary PCR-amplification step. Products were cleaned from unincorporated primers and dNTPs using the GFX PCR DNA purification kit (no. 27-9602-01, Amersham Biosciences) and further sequenced using the DYEnamic ET Dye terminator kit (no. US81090, Amersham Biosciences), following the manufacturer’s instructions. Sequencing products were injected for 40 s at 3 kW and electrophoresed 100 min at 9 kW using a MegaBACE long read matrix (no. US79676, Amersham Biosciences) on a MegaBACE 500 (Amersham Biosciences). Each exon was sequenced from both directions using the same primers as in the PCR. Sequences were visualised and analysed using the Sequence Analyser v3.0 software (Amersham Biosciences). FASTA output of each sequencing result was then compared, using Blast 2 sequences (http://www.ncbi.nlm.nih.gov/blast/bl2seq/bl2.html), to corresponding genomic sequences (AC_007400 and AC_007681) (http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=Nucleotide). Sequences were also inspected visually to detect heterozygous polymorphisms.
Results
Genotyping
Eighteen microsatellite markers (Fig. 1, http://www.ncbi.nlm.nih.gov/mapview) were successfully PCR-amplified and informative enough to proceed with the genotyping of 88 individuals from 11 families (37 affected, 38 healthy and 13 of unknown phenotype). The success rate for those markers was 83.9% and the average information content of 0.93 (range 0.91–0.97). Within the region of fine mapping (~40 cM), we achieved a marker density of one per 1.8 cM. Results were merged with the previous genome-wide scan data (Kaminen et al. 2003), giving a total of 45 informative markers (Fig. 1). The six SNP markers were genotyped with an average success rate of 97.4%.
Multipoint non-parametric linkage (NPL) analysis on chromosome 2 in eleven dyslexic Finnish pedigrees. a NPL results with 45 informative markers. The NPL curve was obtained by Genehunter 2.1 software (Kruglyak et al. 1996) using 45 microsatellite markers. Chromosome 2 centromere is indicated by Cen and a black filled circle, while 2pter and 2qter denote the telomeres. Markers are depicted on the x-axis according to the DeCode genetic map (Kong et al. 2002) and http://www.ncbi.nlm.nih.gov/mapview. A thickened portion of the x-axis denotes the ~40-cM region of the chromosome subjected to fine mapping by adding 18 new microsatellite markers (bold characters). Vertical dotted lines highlight the emplacement of the peak observed for marker D2S2352 (NPL=0.9, P=0.1) and the highest peak at marker D2S2216 (NPL=3.0, P<0.002), as well as the telomeric border of our candidate region at marker D2S2110 (NPL=2.1, P<0.02). An arrow points to marker D2S286/rs3220265, which showed the highest HPM score when the six pedigrees with positive linkage results were chosen for association analysis using HPM. b Enlarged ~40-cM interval between D2S391 (2p21) and D2S2181 (2p11.1). DYX3 denotes the dyslexia susceptibility locus on 2p16-p15 (OMIM 604254) summarizing the candidate regions (white rectangles) described in two independent linkage studies (Fagerheim et al. 1999; Petryshen et al. 2002). OTX1, SEMA4F and TACR1 (black vertical boxes) are the three genes from the 2q dyslexia candidate region which, to date have been tested for mutation analysis (Francks et al. 2002 and present study). Three small black squares denote some of the 2p markers which displayed significant results in quantitative-trait loci (QTL)-based genome-wide scans in US and UK English-speaking dyslexic individuals (Fisher et al. 2002). Markers in italics have not been used in our study but in previously published linkage studies. A two-gray-toned rectangle denotes the dyslexia candidate region described in this study. In light gray is the ~2.7-Mbp chromosomal area which could be excluded using six additional SNP markers, finally narrowing the candidate region to the D2S2116-D2S2181 interval (dark gray)
Confirmation of linkage to 2p11
NPL analysis was performed on the whole data set using the above-described set of 45 markers, except D2S2180 and D2S2333. These markers were excluded because a 4.5- to 24-times inflation of expected recombination was observed in their vicinity (data not shown), strongly suggesting errors in allele calling or mutation events in the marker sequences. A graphical representation of the NPL scores obtained along the 250-cM long chromosome 2 is presented on Fig. 1a. This analysis revealed the highest NPL score (3.0; P=0.001) for marker D2S2216 with NPL scores >2 and nominal P values <0.01 for all markers spanning from D2S2110 (2p12) to D2S2181 (2p11.1). This ~15-cM candidate region (D2S2110-S2181) is depicted in Fig. 1b. Pedigree-wise NPL scores were positive in only six of the pedigrees (nos. 1, 4, 5, 9, 10 and 11 in Kaminen et al. 2003) and their respective contributions to the highest score at marker D2S2216 are: 3.0 (P=0.03), 1.4 (P=0.25), 1.12 (P=0.25), 2.1 (P=0.12), 0.65 (P=0.18) and 2.1 (P=0.12).
Association analyses by microsatellite markers
HPM analysis was performed for the whole chromosome using either (1) trios from the whole set of individuals (11 pedigrees) or (2) trios from the six pedigrees which had shown a positive NPL score in some part of chromosome (see linkage analysis results). At the first stage of analysis, only the microsatellite markers (45 markers) were used. Selecting independent trios from the whole data (43 affected and 29 control chromosomes) gave only modest scores, with non-significant empirical P values. However, the associations were much stronger in the six families showing linkage (28 affected and 20 control chromosomes). Significant scores with empirical P values ≤0.01 were obtained for three neighbouring markers: D2S2110, D2S286 and D2S2116. The highest score was for marker D2S286/rs3220265 (P value <0.001), for which allele 3 (93 bp) (Fig. 1b, 2) was part of a pattern common to ten affected and one control chromosomes.
Nine-marker haplotypes of the eleven chromosomes displaying the associated pattern containing allele 3 of D2S286/rs3220265. All chromosomes containing the associated pattern with allele 3 (93 bp) for marker D2S286/rs3220265 (arrow) are horizontally depicted by their nine-marker haplotypes spanning the interval D2S2110-D2S2116. One affected chromosome from a pedigree that did not show positive linkage to our candidate region (pedigree 6 in Kaminen et al. 2003) also contained this common pattern and is included. All haplotypes except the last one are from affected individuals. First and last haplotypes were found twice but are only represented once here. Markers D2S2110 and D2S21116 are located ~2.6–2.7 cM apart from D2S286/rs3220265 (DeCode genetic map) and their relative physical distances (denoted by horizontal bars) are according to their positions in the dbSNP (build116) (http://www.ncbi.nlm.nih.gov/SNP/index.html). All SNP markers are located on the same sequence contig, NT_022184 (http://www.ncbi.nlm.nih.gov/mapview, build 33) and the physical distance (in kb) from each marker to its adjacent markers is indicated by small horizontal bars. In italics are two genotypes which could not be obtained but were inferred from parental chromosomes
Refined candidate region
In order to verify the positive association obtained for marker D2S286/rs3220265, we added two, respectively four, SNPs covering 226.4 kb and 441.3 kb from D2S286/rs3220265, in the directions of neighbouring markers D2S2110 and D2S2116 (Fig. 2). HPM analysis was re-run with the integrated map of 45 microsatellite markers and the new six SNPs on the two data sets described above (all families and linked-only families). Only modest scores with non-significant empirical P values were obtained in both analyses that did not confirm the association from fine mapping using microsatellite markers only. All 11 chromosomes that first displayed the associated pattern containing allele 3 of D2S286/rs3220265, are depicted in Fig. 2 with their nine markers haplotype (six SNPs and three microsatellites). Three haplotype combinations (1-3-2, 1-3-1 and 2-3-1) could be observed with D2S286/rs3220265 and its two immediate SNP markers (rs754978 and rs6715729) located only 70–85 kb from D2S286. A total of five haplotypes could be detected when one more SNP was added on each side (five markers haplotype). As no single conserved haplotype could be observed even at small distance from D2S286/rs3220265, it is therefore unlikely that the sequence surrounding this marker represents the exact location of a dyslexia gene on chromosome 2p11. Thus, the dyslexia candidate region on 2p11 could be narrowed to the chromosomal area D2S2116-D2S2181 or ~12 Mbp on the human sequence map (~10.5 cM on the DeCode genetic map).
Expression and mutation analyses of the human TACR1 gene
Linkage and association results from our fine mapping pointed to a candidate region in which marker D2S286/rs3220265 was our primary “hot-spot” for the location of a dyslexia gene on 2p11. The TACR1 gene encompasses marker D2S286 and from previous studies is known to have a role in the CNS. We first studied its pattern of expression on Northern blot in a panel of human tissues and could confirm its expression in human brain. Two main transcripts, one ~2.1–2.2 kb and the other nearly 6 kb, could be detected in all ten tissues, but at variable levels of expression (Fig. 3). The smallest transcript is 2 kb in liver and ~1.8 kb in pancreas, while in placenta both a 1.8-kb and 2.1- to 2.2-kb mRNA can be found. Two tissues show high levels of expression of both small and large transcripts, i.e. brain and thymus. The size of the smallest transcript is in accordance with the 2.05-kb previously cloned cDNA and with gene prediction based on the human genome sequence (Gerard et al. 1991; Hopkins et al. 1991, contig NT_022184, GenBank build 33 at http://www.ncbi.nlm.nih.gov/mapview). However, the nearly 6-kb long messenger seen in all ten tissues tested has, to our knowledge, not been reported and probably represents a new splice variant containing at least one of the five exons we used to detect the TACR1 gene expression pattern. In conclusion, the human TACR1 gene is ubiquitously expressed and two transcripts of high levels of expression can be detected in total brain.
Northern blot pattern of expression of the human TACR1 gene. Each lane contains 2 μg of poly(A+) RNA from ten normal human tissues (Ambion). Sk. Muscle skeletal muscle. Size (in kb) markers are indicated by black dots on the right side. Horizontal lanes point to the three main transcripts detected. The smallest one (~1.8 kb) is mostly seen in pancreas and placenta, the second one (~2.1–2.2 kb) is in all tissues except liver, while the third one (~6 kb) is detected ubiquitously. An asterisk denotes an intermediate size transcript only seen in liver (~2 kb)
In order to test the TACR1 gene as a candidate gene in our dyslexic subjects, we first used direct sequencing and screened for mutations all five exons containing the entire coding region of the gene, and their respective exon-intron junctions. Five of the six Finnish families positive for linkage to 2p11, i.e. pedigrees 1, 4, 9, 10 and 11 (Kaminen et al. 2003), as well as two unrelated sporadic dyslexic individuals, were tested. For exons 2–4, 40–50 bp of intronic sequences were screened on each side of the exons as well as on the 5′ border of exon 1 and the 3′ border of exon 5. Exon 1 contains the 5′UTR of the gene and there we screened 103 bp upstream of the start codon. Exon 5 contains the 3′UTR of the ~2.1- to 2.2-kb variant and 154 bp downstream of the stop codon were screened. Two already known SNPs could be detected in our set of individuals, both seen in dyslexic and non-dyslexic individuals: rs6715729, a silent polymorphism affecting the third base of Phe codon 111 in exon 1 and rs2024512, located in intron 1 (http://www.ncbi.nlm.nih.gov/SNP/index.html). We further genotyped the entire set of families for those SNPs, but no association could be found between any of them and the dyslexia phenotype (data not shown).
Discussion
Our previous genome-wide scan had shown linkage between dyslexia and chromosome 2p11, with the highest NPL score for marker D2S2216 (2.55, P=0.004). Moreover, a two-point parametric test (autosomal dominant model) had also revealed a lod score of 3.01 for marker D2S286 (Kaminen et al. 2003). By increasing marker density to one per 1.8 cM over ~40 cM within the linked region, we could confirm the linkage peak at marker D2S2216 (NPL score of 3.0, P=0.001) and detect a common allelic pattern for neighbouring markers D2S2110, D2S286 and D2S2116. This enabled us to decrease the size of the candidate region to ~15 cM, i.e. the interval D2S2110-D2S2181.
The region chosen for our fine mapping also covered the previously reported DYX3 candidate locus (OMIM 604254) on 2p16-p15 (Fagerheim et al. 1999; Petryshen et al. 2002). In our data set, we could observe a modest NPL score of 0.91 (P=0.1) for marker D2S2352 located in the DYX3 region suggesting the possibility of two dyslexia loci on chromosome 2p.
We have investigated this chromosomal area spanning about 15 cM using a candidate gene approach in combination with reconstruction of haplotypes using additional SNP markers. The initially detected association with the highest score for marker D2S286/rs3220265, made us focus on an approximately 670-kb area surrounding this marker. We first screened for mutations in an obvious candidate gene, TACR1, which contains marker D2S286 and that had been described to be of importance in functions of the CNS, such as modulation of neuronal activity, inflammation and mood. No mutations could be detected in dyslexic subjects. However, two common polymorphisms (SNPs rs6715729 and rs2024512) were detected, but no association to dyslexia could be seen with either of them separately or with their haplotypes. As a second step in characterising the surroundings of marker D2S286, we genotyped rs6715729 and five additional SNPs in all our family members, but could not confirm the strong association we had previously observed. This supports that the association to D2S286, and thereby TACR1, was unlikely to carry a founder mutation as all chromosomes that had shown to contain the associated pattern were reconstructed and no longer shared a common haplotype pattern. Since the associated pattern initially detected using HPM could not be confirmed, it probably did not represent a true identical-by-descent/founder haplotype associated with dyslexia. However, our results allowed us to exclude the approximately 2.7-Mbp chromosomal area from D2S2110 to rs718507, narrowing our fine mapping candidate region to D2S2116-D2S2181, i.e. ~12 Mbp on the human sequence map or ~10.5 cM on the DeCode genetic map.
With the exception of two original reports of the cloning of the human TACR1 cDNA (Gerard et al. 1991; Hopkins et al. 1991), no expression study of the gene in human tissues has been reported. Here, we present the relative expression levels and transcript sizes of the human TACR1 gene in a panel of ten normal tissues. Its ubiquitous expression suggests a role as a housekeeping gene and its high expression level in brain, compared with most other tissues tested, underlines its role in the CNS function. However, TACR1 is not implicated in dyslexia development.
References
Cardon LR, Smith SD, Fulker DW, Kimberling WJ, Pennington BF, DeFries JC (1994) Quantitative trait locus for reading disability on chromosome 6. Science 266:276–279
De Felipe C, Herrero JF, O’Brien JA, Palmer JA, Doyle CA, Smith AJ, Laird JM, Belmonte C, Cervero F, Hunt SP (1998) Altered nociception, analgesia and aggression in mice lacking the receptor for substance P. Nature 392:394–397
Derocq JM, Segui M, Blazy C, Emonds-Alt X, Le Fur G, Brelire JC, Casellas P (1996) Effect of substance P on cytokine production by human astrocytic cells and blood mononuclear cells: characterization of novel tachykinin receptor antagonists. FEBS Lett 399:321–325
Fagerheim T, Raeymaekers P, Tonnessen FE, Pedersen M, Tranebjaerg L, Lubs HA (1999) A new gene (DYX3) for dyslexia is located on chromosome 2. J Med Genet 36:664–669
Fisher SE, DeFries JC (2002) Developmental dyslexia: genetic dissection of a complex cognitive trait. Nat Rev Neurosci 3:767–780
Fisher SE, Francks C, Marlow AJ, MacPhie IL, Newbury DF, Cardon LR, Ishikawa-Brush Y, Richardson AJ, Talcott JB, Gayan J, Olson RK, Pennington BF, Smith SD, DeFries JC, Stein JF, Monaco AP (2002) Independent genome-wide scans identify a chromosome 18 quantitative-trait locus influencing dyslexia. Nat Genet 30:86–91
Francks C, Fisher SE, Olson RK, Pennington BF, Smith SD, DeFries JC, Monaco AP (2002) Fine mapping of the chromosome 2p12-16 dyslexia susceptibility locus: quantitative association analysis and positional candidate genes SEMA4F and OTX1. Psychiatr Genet 12:35–41
Gerard NP, Garraway LA, Eddy RL Jr, Shows TB, Iijima H, Paquet JL, Gerard C (1991) Human substance P receptor (NK-1): organization of the gene, chromosome localization, and functional expression of cDNA clones. Biochemistry 30:10640–10646
Grigorenko EL (2001) Developmental dyslexia: an update on genes, brains, and environments. J Child Psychol Psychiatry 42:91–125
Hopkins B, Powell SJ, Danks P, Briggs I, Graham A (1991) Isolation and characterisation of the human lung NK-1 receptor cDNA. Biochem Biophys Res Commun 180:1110–1117
Jurinke C, Van der Boom D, Cantor CR, Köster H (2002) Automated genotyping using DNA MassARRAY technology. Methods Mol Biol 187:179–192
Kaminen N, Hannula-Jouppi K, Kestilä M, Lahermo P, Muller K, Kaaranen M, Myllyluoma B, Voutilainen A, Lyytinen H, Nopola-Hemmi J, Kere J (2003) A genome scan for developmental dyslexia confirms linkage to chromosome 2p11 and suggests a new locus on 7q32. J Med Genet 40:340–345
Kong A, Gudbjartsson DF, Sainz J, Jonsdottir GM, Gudjonsson SA, Richardsson B, Sigurdardottir S, Barnard J, Hallbeck B, Masson G, Shlien A, Palsson ST, Frigge ML, Thorgeirsson TE, Gulcher JR, Stefansson K (2002) A high-resolution recombination map of the human genome. Nat Genet 31:241–247
Kramer MS, Cutler N, Feighner J, Shrivastava R, Carman J, Sramek JJ, Reines SA, Liu G, Snavely D, Wyatt-Knowles E, Hale JJ, Mills SG, MacCoss M, Swain CJ, Harrison T, Hill RG, Hefti F, Scolnick EM, Cascieri MA, Chicchi GG, Sadowski S, Williams AR, Hewson L, Smith D, Rupniak NM, et al (1998) Distinct mechanism for antidepressant activity by blockade of central substance P receptors. Science 281:1640–1645
Kruglyak L, Daly MJ, Reeve-Daly MP, Lander ES (1996) Parametric and nonparametric linkage analysis: a unified multipoint approach. Am J Hum Genet 58:1347–1363
Lahiri DK, Nurnberger JI (1991) A rapid non-enzymatic method for the preparation of HMW DNA from blood for RFLP studies. Nucleic Acids Res 19:5444
Nopola-Hemmi J, Myllyluoma B, Haltia T, Taipale M, Ollikainen V, Ahonen T, Voutilainen A, Kere J, Widen E (2001) A dominant gene for developmental dyslexia on chromosome 3. J Med Genet 38:658–664
O’Connell JR, Weeks DE (1998) PedCheck: a program for identification of genotype incompatibilities in linkage analysis. Am J Hum Genet 63:259–266
Petryshen TL, Kaplan BJ, Fu Liu M, de French NS, Tobias R, Hughes ML, Field LL (2001) Evidence for a susceptibility locus on chromosome 6q influencing phonological coding dyslexia. Am J Med Genet 105:507–517
Petryshen TL, Kaplan BJ, Hughes ML, Tzenova J, Field LL (2002) Supportive evidence for the DYX3 dyslexia susceptibility gene in Canadian families. J Med Genet 39:125–126
Povey S, Lovering R, Bruford E, Wright M, Lush M, Wain H (2001) The HUGO Gene Nomenclature Committee (HGNC). Hum Genet 109:678–680
Schulte-Korne G, Grimm T, Nothen MM, Muller-Myhsok B, Cichon S, Vogt IR, Propping P, Remschmidt H (1998) Evidence for linkage of spelling disability to chromosome 15. Am J Hum Genet 63:279–282
Taipale M, Kaminen N, Nopola-Hemmi J, Haltia T, Myllyluoma B, Lyytinen H, Muller K, Kaaranen M, Lindsberg PJ, Hannula-Jouppi K, Kere J (2003) A candidate gene for developmental dyslexia encodes a nuclear tetratricopeptide repeat domain protein dynamically regulated in brain. Proc Natl Acad Sci USA 100:11553–11558
Toivonen HT, Onkamo P, Vasko K, Ollikainen V, Sevon P, Mannila H, Herr M, Kere J (2000) Data mining applied to linkage disequilibrium mapping. Am J Hum Genet 67:133–145
Acknowledgements
We thank Dr. Cecilia Lindgren for her valuable comments on the manuscript and for supervising the SNP genotyping work at the Mutation Analysis Facility (MAF), Clinical Research Center, Huddinge Hospital, Sweden. This work was supported by the Sigrid Jusélius Foundation, the Academy of Finland and the Swedish Medical Research Council (“Vetenskapsrådet”).
Author information
Authors and Affiliations
Corresponding author
Additional information
H. Anthoni and P. Onkamo contributed equally to this work
Rights and permissions
About this article
Cite this article
Peyrard-Janvid, M., Anthoni, H., Onkamo, P. et al. Fine mapping of the 2p11 dyslexia locus and exclusion of TACR1 as a candidate gene. Hum Genet 114, 510–516 (2004). https://doi.org/10.1007/s00439-004-1103-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-004-1103-0