Abstract
Localized aggressive periodontitis (LAP; previously known as localized juvenile periodontitis) is one of the rapidly progressive periodontal diseases. Certain forms of familial LAP show a simple Mendelian pattern of transmission. However, no gene mutation has been identified to be responsible for the LAP phenotype. As an initial step to identify a gene mutation associated with LAP, we have performed genetic linkage analysis with four multigenerational families exhibiting the LAP phenotype. Affected individuals in the families were identified based on clinical and laboratory criteria in an attempt to define a homogeneous phenotype, since the clinical presentation of LAP may represent a manifestation of a heterogeneous group of diseases. The LAP phenotype is linked to a DNA marker, D1S492, with LOD score 3.48, θ=0.00. The haplotype analysis of the chromosome interval associated with D1S492 indicates that a LAP locus is located between D1S196 and D1S533 on chromosome 1, covering about 26 million DNA basepairs. We have also examined the DNA sequence of prostaglandin-endoperoxide synthase 2 (PTGS2 or cyclooxygenase 2, COX2) since prostaglandin 2 (PGE2), the product of COX2, is upregulated in LAP patients and COX2 is located between D1S196 and D1S533. No mutation in COX2 was identified in the patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Localized aggressive periodontitis (LAP) is estimated to affect 0.1%–3% of adolescent children depending on the specific form of disease and the population studied (Loe and Brown 1991; Papapanou 1999). The aggressive periodontal diseases are clinically heterogeneous and include forms of disease clinically indistinguishable from other forms of periodontitis, and several forms of disease with a uniquely localized clinical presentation with significant destruction at a very early age (Caton et al. 1989; Albandar et al. 1997). These conditions show remarkable familial aggregation (Schenkein and Van Dyke 1994; Novak and Novak 1996). Reports of familial aggregation of LAP have been interpreted to support a major gene locus, with autosomal recessive and X-linked inheritance (Saxen and Nevanlinna 1984; Long et al. 1987). Whereas autosomal recessive forms of aggressive periodontitis appear in northern Europe, results of segregation analysis conducted on US populations are most consistent with autosomal dominant inheritance, although other forms of inheritance cannot be excluded (Boughman et al. 1986; Hart et al. 1993; Marazita et al. 1994).
Although there is heterogeneity within the aggressive periodontitis classification with regard to neutrophil abnormalities, it is clear that there is homogeneity within certain populations. For instance, in US-based studies, approximately 60%–70% of LAP patients exhibit a decreased neutrophil chemotaxis trait (Van Dyke et al. 1985, 1997; Daniel et al. 1993). This is far lower in northern Europe and Finland (Saxen 1980; Kinane et al. 1989). The finding is best illustrated in African-American study populations. Co-segregation of the neutrophil trait with clinical disease has been reported in certain African-American families (Van Dyke et al. 1983). In these families, the neutrophil abnormality relating to reduced chemotaxis has been well characterized by a number of laboratories. This includes decreased chemotaxis to a number of chemotactic factors associated with reduced receptor expression on the neutrophil surface for those chemotactic factors (Van Dyke et al. 1981), reduced calcium transport in LAP neutrophils (Agarwal et al. 1989), increased diacylglycerols levels in stimulated LAP neutrophils, and decreased protein kinase C activity (Kurihara et al. 1993). The finding that these observations are not universal in the LAP populations reported in different geographic locations, or even within the same geographic population, strongly supports the hypothesis that there are several distinct forms of disease with the same clinical presentation. This aspect of the genetics of LAP has not been previously explored systematically.
We have performed genetic linkage analysis of candidate regions with four LAP multigenerational families. With regard to the complicated etiology of LAP, we selected the families based on clinical examination and laboratory tests to minimize the negative influence of non-allelic heterogeneity. The results from the linkage analysis showed that the DNA marker, D1S492, on chromosome 1 had a LOD score 3.48 with θ at 0.00. Haplotype analysis of the chromosome interval associated with D1S492 was performed, and the result from the analysis indicated that the LAP locus resided between D1S196 and D1S533. The DNA sequence of prostaglandin-endoperoxide synthase 2 (PTGS2; formerly known as cyclooxygenase 2, COX2) in LAP patients was also examined since prostaglandin E2 (PGE2), the product of COX2, is upregulated in LAP patients, and COX2 is located between D1S196 and D1S533. No mutation in COX2 was found in the patients.
Materials and methods
Recruitment of LAP families for genetic linkage analysis
Although there is no direct genetic evidence that LAP is a heterogeneous disorder, there is published evidence of heterogeneity in clinical presentation. We, therefore, collected LAP families with caution to minimize the potential negative influence of a heterogeneous etiology of the disorder on the genetic linkage analysis. All of the families were recruited from Boston University Medical and affiliated health centers, in accordance with the human subject protocol approved by Boston University Medical Center Institutional Review Board.
Clinical examination
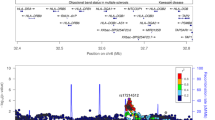
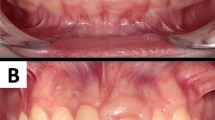
At the beginning of each clinical visit, subjects were asked to fill out a questionnaire to obtain detailed medical and dental histories. A routine dental examination was performed including periodontal, dental, and radiological examinations. Subjects were diagnosed with LAP according to the clinical characteristics of the disease as defined by the American Academy of Periodontology, including attachment and bone loss limited to first molars and incisors. The bone pattern was characteristic of LAP as indicated in Fig. 1. Individuals with active infectious diseases such as hepatitis, AIDS, and tuberculosis or those being treated with any medication known to affect periodontal status were excluded from the genetic linkage study. Pregnant women were also excluded.
The cemento-enamel junction is defined as the joining point of the enamel of the tooth crown and the cementum that covers the radicular surface of the tooth. The distance between the tip of the alveolar crest and the enamel is a clinical and radiographic measure by which the levels of supporting alveolar bone are assessed. This distance, shown in solid bars, is 1–2 mm in healthy individuals without any periodontal disease. An increased distance suggests bone loss. This figure shows the average alveolar bone levels of each tooth in the LAP patients. As a characteristic of LAP, the bone destruction is localized around the incisor (numbers 7–10, 23–26) and first molar (numbers 3, 14, 19, 30) teeth. A significant amount of bone loss is also observed around second molar teeth (numbers 2, 15, 18, 31). On the other hand, no bone loss is observed around canines (numbers 6, 11, 22, 27) and premolar teeth (numbers 4, 5, 12, 13, 20, 21, 28, 29). We used this information for the estimation of bone loss in our clinical examination
Laboratory evaluation of neutrophil function
One of the phenotypic characteristics of most LAP subjects of African-American origin is abnormal in vitro neutrophil function manifest as reduced chemotaxis and elevated superoxide production consistent with cell priming. To characterize our subjects further, chemotaxis and superoxide generation by peripheral blood neutrophils was analyzed in all subjects.
After obtaining informed consent, 30 ml peripheral venous blood was collected. A normal individual in each family served as a control for each laboratory analysis together with an unrelated healthy subject. The laboratory tests included the examination of levels of superoxide generation and chemotaxis to formyl-L-methionyl-L-leucyl-L-phenylalanine (fMLP) by neutrophils. LAP patients in each family demonstrated increased levels of superoxide generation and decreased chemotaxis. In addition, the identification of Actinobacillus actinomycetemcomitans, the causative microorganism of LAP, by DNA-DNA hybridization checkerboard analysis was performed. If subjects who had been clinically diagnosed as having LAP did not meet the laboratory criteria (decrease chemotaxis, elevated O2 −) and microbiological characteristics (A.a. infection), the families were not included in the genetic linkage study.
Isolation of neutrophils
Neutrophils were separated by discontinuous one-step Ficoll-Hypaque density gradient centrifugation (Kalmar et al. 1988). After centrifugation at 900 g for 20 min at room temperature, the neutrophils were isolated and washed in 4 ml phosphate-buffered saline (PBS). The samples were centrifuged at 300 g for 10 min at room temperature, the supernatants were removed, and the contaminating erythrocytes were lysed with ice-cold NH4Cl buffer for 10 min. The samples containing neutrophils were washed with PBS and centrifuged at 300 g for 10 min at room temperature. The purified cells were counted by using a hemocytometer, and the concentration was adjusted depending on the assay.
Superoxide production
Superoxide production (oxidative burst) of neutrophils was analyzed by using a superoxide-dismutase-inhibitable cytochrome C reduction assay (Tanigawa et al. 1993). The neutrophils were suspended in PBS at a concentration of 6×106/ml in a well of a 96-well microplate. Each sample was run in triplicate with 5×105 neutrophils in each well.
Superoxide production was measured both at rest and after stimulation with fMLP (10−6 M). The plate was then incubated at 37°C for 5 min and read in the microplate reader at a wavelength of 550 nm. The amount of superoxide produced was expressed as nanomoles O2 − per 5×105 neutrophils per minute.
Chemotaxis
Cells were suspended in Gey’s buffer (GBSS, Life Technologies, Gaithersburg, Md.) supplemented with 2% bovine serum albumin at a concentration of 2.5×106 cells/ml. The cells suspension was placed in the upper compartment of a modified Boyden chamber separated by a 5-μm pore-size micropore filter (Nucleopore, Pleasanton, Calif.), whereas the lower compartment was loaded with the buffer solution containing fMLP (10 nM) or buffer alone. Cell migratory response was evaluated by counting the number of neutrophils that accumulated on the distal surface of the filter after a 2-h incubation in a 37°C, 5% CO2 incubator. Three representative high-power microscopic fields (×400) were counted for each of the triplicate filters. Neutrophil chemotactic migration in response to fMLP was compared with random migration in the absence of chemotactic agent (Van Dyke et al. 1980).
Checkerboard analysis
Subgingival plaque samples were collected from the teeth and placed into sterile Eppendorf tubes containing 150 μl TE buffer (TE buffer = 10 mM TRIS HCl pH 7.4, 1 mM EDTA), and 100 μl 0.5 M NaOH was added. The samples were then blotted on nylon membranes. The membranes were hybridized with DNA probes including the following species: Fusobacterium nucleatum ss.vincentii (364), Campylobacter concisus (484), Campylobacter rectus (371), Bacteriodes forsythus (338), Porphyromonas gingivalis (381), Prevotella intermedia (25611), Prevotella nigrescens (33563), Capnocytophaga sputigena (33562), Streptococcus oralis (SS11), Actinomyces naeslundii (43146), Treponema denticola (Td), Campylobacter curva (9584), Eikenella corrodens (23834), Actinomyces israeli (12102), and Actinobacillus actinomycetemcomitans (Y4). The hybridization was performed on a Miniblotter 45 apparatus (Immunetics, Cambridge, Mass.). The probes and hybridization buffer were placed in individual lanes of the Miniblotter. The apparatus was placed in a sealed plastic bag and incubated overnight at 42°C. After hybridization, the filters were washed and incubated with phosphatase-conjugated antibody against digoxigenin. Lumiphos 530 solution was applied to the filters, and the chemiluminescence signals were detected with X-ray film. The signals were compared with standards on the membrane (0: no signal detected; 1: <105; 2: ~105; 3: >105 and <106; 4: ~106; 5: >106 cells).
Genetic linkage analysis
In practice, 20 individuals from a large family or several small families are required to perform a linkage analysis to obtain a LOD score greater than 3. Although more than 20 individuals from the families were collected, a simulation study with the SLINK program was performed to estimate the maximum expected LOD score (ELOD; Ott 1989; Weeks et al. 1990).
A directed genomic screening approach was used to perform the genetic linkage analysis. In this approach, we did not randomly choose chromosome regions or evenly select regions across the genome for the analysis. Instead, the regions chosen were based on a chromosome interval containing functional candidate gene or genes for LAP. Fourteen autosomal chromosome intervals were selected (Table 1).
All of the DNA markers used were commercially available (Applied Biosystems, California). Genomic DNA was isolated from 3 ml peripheral blood from individuals by using a Puregene DNA Isolation Kit from Gentra Systems (Minneapolis, Minn.). For the polymerase chain reaction (PCR), 15 μl 1×PCR buffer containing 75 ng DNA, 5 μM each primer, and 200 nM dNTPs, and 1 U Taq polymerase were used. The PCR conditions were 95°C for 12 min, followed by 10 cycles of 94°C for 15 s, 55°C for 15 s, and 72°C for 30 s, followed by 20 cycles of 89°C for 15 s, 55°C for 15 s, and 72°C for 30 s. Finally, the PCR was extended at 72°C for 10 min. The PCR products were loaded on an ABI 3100 Genetic Analyzer to examine the size of the PCR products.
A direct linkage analysis method was used to identify a candidate chromosome interval for further testing. In this method, a DNA marker with the least recombinant fraction among all affected individuals was identified. A chromosomal region around this DNA marker was considered a region of interest for further haplotype analysis. If a DNA marker showed 30% recombination, a chromosome region 5 cM on each side of the marker would not be considered the region of interest for further linkage testing; DNA markers with 10-cM intervals across the genome were planned, if needed.
Once a chromosome region of interest was identified, haplotype analysis of this region was performed in all families. Two-point LOD scores were calculated by using the MLINK option of the LINKAGE package (version 5.10) and FASTLINK (version 4.0P) under the assumption of 0.001 disease-gene frequency and 95% penetrance.
Examination of DNA sequence of prostaglandin-endoperoxide synthase 2 (cyclooxygenase 2, COX2)
The DNA sequence of COX2 from LAP patients and normal individuals was directly examined. PCR fragments were generated, and primers for PCR were designed according to the published DNA sequence of COX2 (Table 2).
For the PCR, 50 μl 1×PCR reaction buffer containing 200 ng genomic DNA, 1 μM of each primer, and 200 nM dNTPs, and 5 U Taq polymerase were used. The PCR conditions were 95°C for 3 min, followed by 35 cycles of 95°C for 40 s, 60°C for 45 s, and 72°C for 60 s. Finally, the PCR was extended at 72°C for 10 min. The PCR products were purified by gel extraction kit (Qiagen). The purified PCR products were then sequenced on an ABI 3100 Genetic Analyzer.
Results
Recruitment of LAP families for genetic linkage analysis
Eight African-American families with LAP phenotype were recruited for the genetic linkage analysis.
Clinical examination and laboratory evaluation of neutrophil function
Superoxide production
Affected individuals exhibited significantly higher superoxide generation than non-affected relatives and normal controls (Fig. 2).
Chemotaxis
Neutrophil chemotactic migration in response to fMLP was dramatically reduced in LAP. Random migration was equivalent between groups (Fig. 3).
Chemotactic migration of polymorphonuclear neutrophils (PMNs) from LAP and age-, race-, and gender-matched healthy controls (n=6). Chemotaxis was measured by subtracting random migration of neutrophils from the migration of neutrophils toward a chemotactic agent (fMLP, 10 nM). LAP neutrophils show a statistically significant reduction in chemotaxis compared with neutrophils from healthy donors (*P<0.05, ANOVA)
Checkerboard analysis
LAP affected individuals harbored significantly more A. actinomycetemcomitans (Fig. 4).
Based on clinical and laboratory evaluations, four families (A–D) were suitable for genetic linkage analysis. Two families did not meet our laboratory criteria, and the other two families were too small to be informative for genetic linkage analysis.
Genetic linkage analysis
There were more than 20 individuals in families A, B, C, and D, as required for linkage analysis. To verify further that a LOD score greater than 3 from those families was possible, a simulation study with the SLINK program was performed to obtain a maximum expected LOD scores (ELOD) of 3.75 with θ at 0.00 based on 1,000 replicates.
The initial result from our direct linkage analysis showed that one specific DNA marker, D1S238, was shared by all affected individuals but not by normal individuals in all four families. Another DNA marker, D1S413 (10 cM telomeric side of D1S238), was also shared by all of the affected individuals with the exception of one recombination in an affected individual. The chromosome region around D1S238 was chosen as the candidate region to contain LAP. Therefore, haplotype analysis was performed on this chromosome region (Fig. 5).
Haplotype analysis of four families (arrows probands). The results from the haplotype analysis shows that several recombinant events occurred within the families (in C.II-3 and D.I-1), which positioned a LAP locus between D1S196 and D1S533. The length of this chromosome interval is about 26 million DNA basepairs
The result from the haplotype analysis showed that a number of recombinant events had occurred among several individuals. The recombinant events in C.II-3 and in D.I-1 positioned the LAP locus between D1S196 and D1S533. The physical distance of this chromosome interval is 26,316,987 bp. To confirm the linkage result from the haplotype analysis, two-point LOD scores for each marker were calculated (Table 3) by using the MLINK option of the LINKAGE package (version 5.10) and FASTLINK (version 4.0P) under the assumption of 0.001 disease-gene frequency and 95% penetrance. The pedigrees were not sensitive to estimates of marker allele frequencies.
Examination of DNA sequence of prostaglandin-endoperoxide synthase 2 (cyclooxygenase 2, COX 2)
The entire coding region and a 1.7-kb 5’-upstream and 2.7-kb 3’-untranslated region of COX2 were sequenced. No DNA mutation of COX2 in LAP patients was identified.
Discussion
In this paper, a LAP locus was assigned to chromosome 1q25 in four African-American families. The families were chosen based on clinical periodontal presentation and specific host response and microbiologic parameters. Although there is no direct genetic evidence that LAP is a heterogeneous disorder, previous clinical studies have indicated a simple Mendelian pattern of transmission in certain numbers of LAP families with the wide-range of clinical phenotype. This suggests that LAP can be caused by either different mutations in different genes (non-allelic heterogeneity) or different mutations in the same gene (allelic heterogeneity). To minimize the negative influence of non-allelic heterogeneity on our genetic linkage analysis, we performed laboratory tests to screen LAP families before genetic linkage analysis.
Although it is evident that LAP is a rare genetic disorder, few genetic linkage analyses have been performed to identify a locus associated with the disease. In 1986, Boughman et al. conducted a study involving a large five-generation kindred family including more than 70 individuals. This family had two distinct dental disorders, type III dentinogenesis imperfecta (DGI-III) and aggressive periodontitis (juvenile periodontitis, JP). The pedigree showed that the two disorders did not completely co-segregate. Subsequently, genetic linkage analysis showed that JP and DGI-III mapped to two different loci on the same region of chromosome 4 within less than 7 cM. However, two main questions remained to be addressed in this linkage study. The two dental disorder loci showed a higher recombinant frequency (5/15=0.33) than expected (less than 0.07) if the loci were located within less than 7 cM on an autosomal chromosome, unless they were in a chromosomal interval with a very high number of “recombinant hot spots”. Another issue was that the genetic linkage analysis was carried out with a protein polymorphic marker. Protein polymorphic markers for genetic linkage analysis are often not as sensitive and specific as DNA markers. Therefore, it would be more informative if the linkage data could be confirmed by using DNA polymorphic markers. Another genetic linkage study was performed with affected sib-pair (ASP) analysis and included 29 African-American families and 15 Caucasian families (Wang et al. 1996); 385 DNA polymorphic markers covering the entire human genome were used for the ASP analysis. Two chromosome loci, viz., a locus around HLA region on chromosome 6 and a locus on chromosome 9, were suggested to be the loci of interest for aggressive periodontitis. One of the advantages of the ASP approach is that genetic linkage analysis can be performed, without knowledge of the mode of transmission of a genetic disorder, by evaluation of the distribution pattern of a specific marker among ASPs. A marker presenting in ASPs with a higher frequency than expected indicates the possibility of linkage between the marker and the disease phenotype. However, it has to be assumed that this marker is identical by descent (IBD), since a marker IBD is not distinguishable from a marker identical by state in ASP analysis. In addition, non-allelic heterogeneity in human genetic disorders reduces dramatically the power of ASP analysis. Obviously, the loci of interest for LAP on chromosomes 4, 6, and 9 need to be confirmed by further genetic linkage analyses.
Our linkage study positioned a LAP locus between D1S196 and D1S533. The data is the first report of a chromosome interval that contains a LAP locus. The fact that all families in the genetic linkage study have different haplotypes suggests that the LAP phenotype in the families is probably attributable to multiple mutations in a same gene. Based on the chromosome interval that contains the LAP locus, it should be possible to identify the DNA polymorphic markers through genome databases. DNA markers will ultimately be useful for clinical genetic consultations and the diagnosis of certain forms of LAP.
Initially, COX2 was considered a primary candidate gene for LAP, because previous studies had shown that prostaglandin E2, the product of COX2, was over-expressed in LAP patient neutrophils (Shapira et al. 1996). Thus, the chromosome region containing COX2 was selected as a candidate locus for genetic linkage analysis. Once this chromosome region was linked to the LAP phenotype, DNA sequences of COX2 in LAP patients including a 1.7-kb 5’-upstream region (assumed to be the promotor region), the entire coding region, and a 2.7-kb 3’-untranslated region were examined. No mutation was detected in COX2 in LAP patients. Although COX2 cannot be completely ruled out as the primary defect of LAP, simple DNA alterations in COX2, such as a point mutation, insertion, or small deletion, do not appear to be responsible for the disease.
References
Agarwal S, Reynolds MA, Duckett LD, Suzuki JB (1989) Altered free cytosilic calcium changes and neutrophil chemotaxis in patients with juvenile periodontitis. J Periodont Res 24:149–154
Albandar JM, Brown LJ, Genco RJ, Loe H (1997) Clinical classification of periodontitis in adolescents and young adults. J Periodontol 68:545–555
Boughman JA, Halloran SL, Roulston D, Schwartz S, Suzuki JB, Weitkamp LR, Wenk RE, Wooten R, Cohen MM (1986) An autosomal-dominant form of juvenile periodontitis: its localization to chromosome 4 and linkage to dentinogenesis imperfecta and Gc. J Craniofac Genet Dev Biol:341–350
Caton J, Bouwsma O, Polson A, Espeland M (1989) Effects of personal oral hygiene and subgingival scaling on bleeding interdental gingiva. J Periodontol 60:84–90
Daniel MA, McDonald G, Offenbacher S, Van Dyke TE (1993) Defective chemotaxis and calcium response in localized juvenile periodontitis neutrophils. J Periodontol 64:617–621
Hart TC, Marazita ML, McCanna KM, Schenkein HA, Diehl SR (1993) Reevaluation of the chromosome 4q candidate region for early onset periodontitis. Hum Genet 91:416–422
Kalmar JR, Arnold RR, Warbington ML, Gardner MK (1988) Superior leukocyte separation with a discontinuous one-step Ficoll-Hypaque gradient for the isolation of human neutrophils. J Immunol Methods 110:275–281
Kinane DF, Cullen CF, Johnston FA, Evans CW (1989) Neutrophil chemotactic behaviour in patients with early-onset forms of periodontitis (I). Leading front analysis in Boyden chambers. J Clin Periodontol 16:242–246
Kurihara H, Murayama Y, Warbington M, Champagne CME, Van Dyke TE (1993) Depressed protein kinase C (PKC) activity of neutrophils in localized juvenile periodontitis. Infect Immun 61:3137–3142
Loe H, Brown LJ (1991) Early onset periodontitis in the United States of America. J Periodontol 62:608–816
Long JC, Nance WE, Waring P, Burmeister JA, Ranney RR (1987) Early onset periodontitis: a comparison and evaluation of two proposed modes of inheritance. Genet Epidemiol 4:13–24
Marazita ML, Burmeister JA, Gunsolley JC, Koertge TE, Lake K, Schenkein HA (1994) Evidence for autosomal dominant inheritance and race-specific heterogeneity in early-onset periodontitis. J Periodontol 65:623–630
Novak MJ, Novak KF (1996) Early-onset periodontitis. Curr Opin Periodontol 3:45–58
Ott J (1989) Computer-simulation methods in human linkage analysis. Proc Natl Acad Sci USA 86:4175–4178
Papapanou PN (1999) Epidemiology of periodontal diseases: an update. J Int Acad Periodontol:110–116
Saxen L (1980) Prevalence of juvenile periodontitis in Finland. J Clin Periodontol 7:177–186
Saxen L, Nevanlinna HR (1984) Autosomal recessive inheritance of juvenile periodontitis: test of a hypothesis. Clin Genet 25:332–335
Schenkein HA, Van Dyke TE (1994) Early-onset periodontitis: systemic aspects of etiology and pathogenesis. Periodontology 2000 6:7–25
Shapira L, Soskolne WA, Van Dyke TE (1996) Prostaglandin E2 secretion, cell maturation, and CD14 expression by monocyte-derived macrophages from localized juvenile periodontitis patients. J Periodontol 67:224–228
Tanigawa T, Kotake Y, Reinke LA (1993) Spin trapping of superoxide radicals following stimulation of neutrophils with fMLP is temperature dependent. Free Radic Biol Med 4:425–433
Van Dyke TE, Horoszewicz HU, Cianciola LJ, Genco RJ (1980) Neutrophil chemotaxis dysfunction in human periodontitis. Infect Immun 27:124–132
Van Dyke TE, Levine MJ, Tabak LA, Genco RG (1981) Reduced chemotactic peptide binding in juvenile periodontitis: a model for neutrophil function. Biochem Biophys Res Commun 100:1278–1284
Van Dyke TE, Levine MJ, Tabak L, Genco RG (1983) Juvenile periodontitis as a model for neutrophil function: reduced binding of the complement chetactic fragment, C5a. J Dent Res 62:870–872
Van Dyke TE, Schweinebraten M, Cianciola MJ, Offenbacher S, Genco RG (1985) Neutrophil chemotaxis in families with localized juvenile periodontitis. J Periodont Res 20:503–514
Van Dyke TE, Schweinebraten M, Ciancola LJ, Offenbacher S, Genco RJ (1997) Neutrophil chemotaxis in families with localized juvenile periodontitis. J Periodontal Res 20:503–514
Wang S, Sun C, Gillanders E, Wang Y, Duffy D, Bock C, Freas-Lutz D, Zhang Y, Lopes N, Schenkein H, Diehl S (1996) Genome scan for susceptibility loci to the complex disorder early onset periodontitis (abstract). Am J Hum Genet 59:1386
Weeks DE, Ott J, Lathrop GM (1990) SLINK: a general simulation program for linkage analysis. Am J Hum Genet (Suppl) 47:S204
Acknowledgments
This work is supported by research funding from Harvard School of Dental Medicine (to Y.L.) and NIH grants PO1 DE 13499 and MO1-PR00533 (to T.E.V.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, Y., Xu, L., Hasturk, H. et al. Localized aggressive periodontitis is linked to human chromosome 1q25. Hum Genet 114, 291–297 (2004). https://doi.org/10.1007/s00439-003-1065-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-003-1065-7