Abstract
It is well-known that obesity is a complex multifactorial and heterogeneous condition with an important genetic component. Recently, major advances in obesity research emerged concerning the molecular mechanisms contributing to the obese condition. This review outlines several studies and data concerning the genetics and other important factors in the susceptibility risk to develop obesity. Based in the genetic etiology three main categories of obesity are considered: monogenic, syndromic, and common obesity. For the monogenic forms of obesity, the gene causing the phenotype is clearly identified, whereas for the common obesity the loci architecture underlying the phenotype is still being characterized. Given that, in this review we focus mainly in this obesity form, reviewing loci found until now by genome-wide association studies related with the susceptibility risk to develop obesity. Moreover, we also detail the obesity-related loci identified in children and in different ethnic groups, trying to highlight the complexity of the genetics underlying the common obese phenotype. Importantly, we also focus in the evolutionary hypotheses that have been proposed trying to explain how natural selection favored the spread of genes that increase the risk for an obese phenotype and how this predisposition to obesity evolved. Other factors are important in the obesity condition, and thus, we also discuss the epigenetic mechanisms involved in the susceptibility and development of obesity. Covering all these topics we expect to provide a complete and recent perspective about the underlying mechanisms involved in the development and origin of obesity. Only with a full understanding of the factors and mechanisms contributing to obesity, it will be possible to provide and allow the development of new therapeutic approaches to this condition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Obesity is becoming one of the world´s greatest health public challenges, contributing to the increased risk of many chronic diseases, such as hypertension, type 2 diabetes, cardiovascular diseases and other co-morbidities (Swinburn et al. 2011). In this review we address this important heatlh problem in two main aspects: (1) providing an overview about the genetic causes of obesity, and (2) reviewing hypotheses trying to explain how natural selection acted in genes that increase the susceptibility to develop obesity. It is widely accepted that our lifestyle contribute to an increase in obesity, however, it is also clear that an important genetic component is behind the risk of developing an obese phenotype. The genetic component of obesity will be reviewed in this work from monogenic to common obesity forms, referring recent data about new loci associated with obesity. Moreover, aspects like the obesity genetics of adults versus children, or the obesity genetic differences across ethnicities will be also addressed in this review. From another point of view an important question is: how has natural selection favored the spread of genes that increase the risk for an obese phenotype and how has this predisposition to obesity evolved? Thus, the second important aspect of this review is to present some of the evolutionary hypotheses trying to explain the development and prevalence of obesity nowadays. With this review we expect to contribute to a better understanding of underlying causes of obesity, and to raise important questions about future research directions.
It is believed that differences in obesity and others traits could arise primarily as a consequence of genetic factors, as is revealed by the high heritability for body mass index (BMI) (Feinleib et al. 1977; Stunkard et al. 1986a, b; Silventoinen et al. 2010). A trait can reflect the activity of a single-gene (Mendelian or monogenic) or more than one gene (polygenic), both cases being influenced by environmental factors. The polygenic multi-factorial condition reflects the additive contribution of many genes conferring different degrees of susceptibility. Accordingly, we may understand a polygenic trait as the combined action of several genes producing a “continuously varying” phenotype. With the advent of the Human Genome Project (1990–2003), millions of DNA sequence variants were discovered in the human genome. This large and diverse database of polymorphism markers provided a novel opportunity to study the human genetic basis of several complex diseases through population approaches. In the study design of population approaches, a significant amount of individuals must be screened for a large number of polymorphisms. If a mutation increases susceptibility to a specific disease of interest, it is more common among individuals affected by this condition than among non-affected individuals.
The increase in prevalence of overweight and obesity has been described as an important problem worldwide. Epidemiological studies estimated that prevalence of overweight/obesity increased of ~41 % between 1980 and 2013 (Ng et al. 2014). This increase represents only three decades marked by lifestyle changed. Today, the majority of individuals engage in sedentary behavior and have easy access of food. If, along our evolutionary origin a genetic predisposition to obesity evolved it is obviously probable that our lifestyle changes play an important role to the risk of obesity. When human morphology is considered, there are profound individual differences, such as body size, hair color/form, eyes color/form, etc. These human variations were due, in part, to evolutionary forces, genetic drift, environmental conditions, among others. However, in all societies and subpopulations, despite different prevalence rates, there are both obese and non-obese individuals. Here, we describe four hypotheses based on the evolutionary origins of obesity. From the “thrifty genotype” to the newest and promising hypothesis suggesting that a positive selection could be occurred during the “out of Africa” migration between adaptations of metabolic rate with cold temperatures.
Obesity
Human obesity is a global public health concern and results from an excessive accumulation of body fat that can adversely affect health (Haslam and James 2005). The global rise of obesity has serious effects, contributing for a significant number of diseases including type 2 diabetes mellitus, cardiovascular diseases, metabolic syndrome, and some cancers (Haslam and James 2005; Swinburn et al. 2011). Beyond co-morbidities, obesity has a great social impact and substantial direct and indirect costs in healthcare services (Swinburn et al. 2011). Excessive fat accumulation results from a persistent positive energy balance, that is, the amount of energy consumed exceeds the amount of energy expended (Silventoinen et al. 2010). So, a simple definition of obesity could be: a consequence of an imbalance between energy intake and energy expenditure (Sandholt et al. 2012). The energy balance represents a conglomerate of traits, each one influenced by numerous variables such as behavior, diet, environment, social structures, metabolic factors and genetics (Mathes et al. 2011). The result of this complex interaction among all of these variables contributes to individual differences in the development of obesity.
Epidemiological studies indicate that adiposity, as reflected by BMI, has increased worldwide over the past decades (Finucane et al. 2011). Moreover, obesity is more common in some countries than in others, though precise cross-country comparisons can be difficult because not all samples are representative of the relevant populations. Nonetheless, available data suggest that the increase in the prevalence of obesity began to emerge during the 1980s and ever since more countries have joined the global obesity pandemic (Finucane et al. 2011; Ng et al. 2014). Between 1980 and 2008, the global change per decade for age-standardized mean BMI was increased ~0.4 and ~ 0.5 kg/m2 in men and women, respectively (Finucane et al. 2011). By 2013, the global estimated prevalence of overweight (BMI > 25 kg/m2) and obesity (BMI > 30 kg/m2) in men and women was 36.9 and 38.0 %, respectively, comparing with 28.8 and 29.8 %, respectively, in 1980. In children and adolescents, the prevalence of overweight and obesity has increased from 1980 to 2013 in developed countries (23.8 and 22.6 %, in boys and girls, respectively), but also in developing countries with an increase around 5 % of both boys and girls. In some countries like Kiribati, Federated States of Micronesia, Libya, Qatar, Samoa among others the estimated prevalence of obesity in adults exceeded 50 % of the population (Ng et al. 2014). In 1989, worldwide estimates for the prevalence of overweight and obesity among adults (>20 years) was around 857 million individuals, comparing to the 2.1 billion in 2013 (Ng et al. 2014). These values represent an increase of ~41 % in 33 years.
In modern societies, despite obesity awareness campaigns, and efforts to decrease the energy intake and increase the energy expenditure, obesity prevalence is still increasing. However, we do not yet understand why not everyone in our societies becomes obese. Obesity has a multi-factorial etiology, involving various non-genetic and genetic factors (Haslam and James 2005; Xia and Grant 2013). Probably, most cases of obesity results of a cluster towards the middle of this spectrum, which can be best described as the outcome of an adverse obesogenic environment, working on a susceptibility genotype. Effectively, the genetic susceptibility can potentially be mediated through defects in several different homeostatic mechanisms. Certainly, the exposure to an obesogenic and other environmental factors is the main cause of the increase in the prevalence of high BMI over the last 30 years (Haslam and James 2005; Finucane et al. 2011; Swinburn et al. 2011).
Interestingly, in the early 2000s a considerable number of studies emerged with a controversy surrounding the idea of the “obesity paradox”. According to it, some individuals with overweight or obesity can be considered healthy regarding to their metabolic and cardio-respiratory fitness. Concretely, this paradox suggests that individuals with a high BMI have a better prognosis than individuals with normal BMI concerning the risk association with cardio-vascular diseases and many other chronic diseases (Goyal et al. 2014). For example, Romero-Corral et al. (2006) showed that overweight individuals had a significantly lower risk of all-cause mortality, and a trend towards decreased cardiovascular mortality, comparing to individuals with normal BMI. Some hypotheses were proposed trying to explain this obesity paradox. For example, this could be a result of a possible selection bias and inability to control non-measurable confounding factors (Goel et al. 2014). In fact, BMI cannot distinguish between an elevated body weight due to high levels of lean versus fat body mass. An excess of body fat is more frequently associated with metabolic abnormalities than a high level of lean body mass (Bastien et al. 2014). The genetically inherited leptin or other adipokines deficiencies were also investigated and could have an important role in obesity paradox, in which increased levels of leptin could be cardioprotective (Blüher and Mantzoros 2015). However, until now the underlying mechanisms for obesity paradox remain unclear, and the idea controversial.
A more detailed discussion about this topic is beyond the scope of the current article, but could be found in detail in other studies (Blüher 2014; Goel et al. 2014; Lavie et al. 2015).
Genetic obesity
The increase in the obesity prevalence around the world has been broadly attributed to the change in environment, that is more obesogenic, against an evolutionary background, which could be maladaptive in this new obesogenic context. On the other hand, specific features of the energy balance mechanisms can effectively protect against obesity, possibly explaining why one-third or more of the population remains lean (O’Rahilly and Farooqi 2008). The obesity phenotype only emerges if food consumption exceeds the energy expenditure on a lasting basis, resulting in a prolonged positive energy balance. However, there are many risk factors that predict the development of obesity and generally all involve the interaction of biological and social factors. Numerous studies are consistent with the hypothesis that the personal genetic profile could be a cause for individual differences in the predisposition to weight gain. It is, therefore, interesting that most of the genes involved in the susceptibility of obesity are also related to food intake and regulation of energy balance (O’Rahilly and Farooqi 2008). Based on genetic and phenotypic characteristics, three types of obesity forms can be considered: monogenic syndromic obesity, monogenic non-syndromic obesity and polygenic (common) obesity.
Evidence for a genetic component to obesity
Over the last 30 years, the increase in the prevalence of obesity could be attributed primarily to environmental changes, or to high-calorie food intake together with the sedentary lifestyle of modern societies (Xia and Grant 2013). The fact that the prevalence of obesity in many countries has increased threefold over the last three decades seems difficult to conjugate with the notion that genetics are the primary cause of obesity as revealed by twin and adoption studies (Feinleib et al. 1977; Stunkard et al. 1986a, b; Silventoinen et al. 2010). Nevertheless, it is now believed that environmental factors can influence the genetic background contributing to the increase in obesity prevalence. Morevover, epigenetic mechanisms, in which environmental factors cause changes in the expression of genes, could also help explaining the observed increase in obesity prevalence.
Heritability represents the proportion of phenotypic variation among individuals due to genetic contribution. Hence, it is not surprising that one important risk factor for childhood and adolescent obesity is parental obesity. Whitaker et al. (1997) found that when both parents are obese there is an increase of more than double of the risk for childhood obesity. However, most of the studies found a small to medium effect of parental obesity as risk factor for childhood obesity (Danielzik et al. 2002). Other studies have found a stronger effect for maternal obesity compared to paternal obesity, which may reflect pre- and postnatal environmental factors (Magnusson and Rasmussen 2002). Moreover, maternal weight gain in pregnancy has been positively associated with BMI of the children into adulthood (Mamun et al. 2009). Several studies found that environmental conditions experienced in utero are an important factor in programming obesity (in the epigenetics section it will be discussed in more detail).
Twin studies have been used to model the genetic component of a given trait, due to the fact that monozygotic (MZ) twins are genetically identical, while non-identical dizygotic (DZ) twins share only 50 % of their genetic material (Xia and Grant 2013). In 1977, Feinleib et al. studied the correlations for weight in 250 MZ and 264 DZ male veteran twin pairs, and established for the first time that familial aggregation for obesity results mainly from genetic influence. In 1986, Stunkard et al. (1986a) confirmed these results in a 25-year follow-up study using more than 4000 MZ and DZ twin pairs. High heritability values for BMI were observed for the same subjects at 20 years (h 2 = 0.77) and at 45 years (h 2 = 0.84). The heritability of fat mass among MZ twins has been reported to range from 70 to 90 %, while in DZ twins it is 35–45 %. Adoption studies have strengthened the evidence of a strong genetic influence on human body weight. Body corpulence of adopted children correlates more strongly with BMI of their biologic parents versus the BMI of their adoptive parents (Stunkard et al. 1986b). Recently, Silventoinen et al. (2010), conduct a review of studies in twins and adopted children, suggesting that genetic factors could have a much stronger effect than environmental factors on the BMI trends in children up to the age of 18 years.
Another genetic component for obesity is highlighted through the different prevalence between racial groups. For example, it was found that obesity prevalence in Caucasian and Asian populations is of about 35 % or less compared to 50 % or more found among Pima Indians living in New Mexico (Knowler et al. 1990). Several studies support the concept that genes play a key role in the obesity etiology. However, the search for underlying genotypes that cause of obesity has been challenging due to the complex interactions involved in the regulation of adiposity. Indeed, many of individual genotypes (especially those obtained with a lower odds ratio) that have been associated with elevated body mass have not been replicated in a reliable fashion. Moreover, environmental factors and cultural diversity also account for the different obesity prevalence found across ethnicities. Studies have shown that genetic population substructure, economic disadvantages, psychosocial stress, or access to medical care could have an important impact in obesity development and prevalence. The cultural context could also influence obesity, by defining for example the type and quality of food intake. Also, in some cultural contexts the obesity phenotype could represent a signal of wealth and high social status. Another important factor is the genetic architecture, which is different across population (population substructure) and might be differentiated in ethnic clusters. This could presuppose that disease-causing alleles are more likely to be present in some groups or even be specific to other groups. This point could be considered as the population genetic predisposition to develop the disease. For example, it is well-demonstrated that in the US black-white population disparities in the risk of developing cardiovascular disease exist (Kuzawa and Sweet 2009). Also, social factors such as economics disadvantage or psychosocial stress between groups could have a real impact in causes of ill health. For example, disparities originated by the limited access of quality medical care between different ethnics groups living in the same population might influence health, causing different disease rates. The role of maternal nutrition and stress suffered during pregnancy, mostly due to social disparities or cultural differences could also influence biological processes and responses across the life cycle that will be discussed later (epigenetic section). All these factors affect the intrauterine environment reflecting differences in birth weight. However, nowadays is still evident that genetic factors play a considerable role in obesity. Three distinct forms of obesity could be found: monogenic, syndromic and polygenic obesity.
Mendelian forms of obesity
Monogenic forms of obesity result from an alteration of a single gene and are rare, affecting about 5 % of the population (Farooqi and O’Rahilly 2005; González-Jiménez et al. 2012). There are more than 200 types of human obesity that are associated with homozygous forms of a single gene mutation (Bell et al. 2005; Rankinen et al. 2006). Two forms of Mendelian inheritance of obesity could be found: syndromic and non-syndromic. Most of these monogenic forms of obesity are characterized by an early-onset of the disease and an extreme phenotype (Farooqi and O’Rahilly 2005). In the search for homologous mutations in mice, several human forms of obesity have been identified (Huszar et al. 1997). Thus, murine models appear useful to understand the molecular pathogenesis of human obesity (Lutz and Woods 2012). Family studies based on individuals with extreme obesity, also proved to be very successful in the detection of obesity-related mutations (Hinney et al. 2010). Below is presented a brief and general review about the two syndromic forms of obesity.
Non-syndromic form of obesity
Over the past 15–20 years, several gene mutations have been shown to cause autosomal recessive and dominant forms of obesity. More than 200 single-gene mutations have been found to cause human obesity (Mutch and Clément 2006). Interestingly, all these mutations can be found in only ten genes (Rankinen et al. 2006). However, these mutations are rare and lead to extreme obesity with an early-onset obesity and other endocrine disorders (González-Jiménez et al. 2012). There are eight well-known genes mutations in monogenic non-syndromic form of obesity explaining up to 10 % of cases with early-onset extreme obesity, affecting LEP, LEPR, POMC, PCSK1, MC4R, BDNF, NTRK2 and SIM1 (Table 1) (Farooqi and O’Rahilly 2005; Ranadive and Vaisse 2008; González-Jiménez et al. 2012). All these genes code for proteins with a central role in the leptin–melanocortin signaling pathway present in the hypothalamus, and therefore, affect regulation of food intake and energy expenditure (González-Jiménez et al. 2012). This pathway is activated when LEP is secreted by the adipose tissue, binds to its receptor, localized in the surface neurons in the arcuate nucleus of the hypothalamus (Dubern and Clément 2012). The signal that regulates satiety and energy homeostasis is then propagated through the POMC/cocain and amphetamine-related transcript (CART) and melanocortin system (González-Jiménez et al. 2012). While POMC/CART neurons synthesize anorexigenic peptide alpha-melanocyte-stimulating hormone (α-MSH), a distinct group of neurons synthesizes the orexigenic peptide neuropeptide Y (NPY) and agouti-related protein (AGRP), which act as inhibitors of MC3 and MC4 receptors (Harrold and Williams 2006). The derived peptide nature of POMC depends of the endoproteolytic-type enzyme present, specific in brain region. In the anterior pituitary, the PCSK1 enzyme produces adrenocorticotropic hormone (ACTH) and β-lipotropin (β-LPH), while the combined presence of PCSK1 and PCSK2 in the hypothalamus control the production of α-, β-, γ- MSH and β-endorphins (González-Jiménez et al. 2012).
The protein encoded by the MC4R gene, is a membrane-bound receptor and a member of the melanocortin receptor family (Hinney et al. 2013). The protein interacts with adrenocorticotropic and MSH hormones and is mediated by G proteins. The MC4R gene is composed by a single exon, and is located in the chromosome 18q21.3, encoding for the 332-amino acid seven-transmembrane G-protein-linked receptor, critically involved in regulating energy balance (Gantz et al. 1993). It is expressed mainly in the central nervous system, including in the hypothalamus, contributing to food intake and energy expenditure regulation (Gantz et al. 1993; Mountjoy et al. 1994). In 1998, two independent groups reported a mutation in the MC4R gene, which result in a non-functional receptor causing severe early-onset obesity (Vaisse et al. 1998; Yeo et al. 1998). In morbidly obese individuals, deficiency in the MC4R gene activity represents the most common cause (1–6 %) for the obese phenotype (Yeo et al. 1998; Farooqi et al. 2003; Beckers et al. 2006). More than 150 variants of this gene have been described, usually classified into five classes depending of their molecular effects (Hinney et al. 2013).
The LEP gene (chromosome 7q31.2) encodes a protein that is secreted by white adipocytes, which plays a central role in body weight regulation (Dubern and Clément 2012). This protein, acts as part of a signaling pathway that can inhibit food intake and/or regulate energy expenditure to maintain constancy of adipose mass. In 1997, in a screening for serum level concentrations in severely obese subjects, two children of the same family were found with undetectable levels of leptin (Montague et al. 1997). Subsequently, research revealed that leptin deficiency is inherited and produces extreme early onset obesity (Rau et al. 1999). This deficiency can be caused by a frameshift mutation (del G133), which produces a truncated protein that is not secreted (Rau et al. 1999) or a missense mutation Arg105Trp, which is associated with low levels of circulating leptin (Strobel et al. 1998).
The protein encoded by the LEPR gene (chromosome 1p31.3) belongs to the gp130 family of cytokine receptors, which stimulate gene transcription via activation of cytosolic STAT proteins, predominantly in the hypothalamic neurons (Bates and Myers 2003). This protein is a receptor for leptin and is involved in regulation of fat metabolism. A splice site mutation in the exon 16 is associated with leptin receptor deficiency, producing extreme obesity (Clément and Ferré 2003).
Syndromic form of obesity
Syndromic forms refer to obesity cases that occur in a distinct set of associated clinical phenotypes, such as mental retardation or organ-specific developmental abnormalities (Ichihara and Yamada 2008). There are more than 30 Mendelian disorders that result in obesity (Mutch and Clément 2006). Research is beginning to determine the genetic basis of some of these syndromes, thus elucidating the pathogenesis of the chronic positive energy balance. The genetic basis of these disorders is extremely heterogeneous. Table 1 presents the most common forms of early-onset syndromic obesity for which the genetic basis is, at least, partially understood, including WAGR (Wilm's tumor, aniridia, genitourinary anomalies and mental retardation), Prader-Willi, Bardet-Bield, Altröm and Cohen syndromes.
WAGR syndrome is a rare genetic disorder characterized by a deletion at chromosome 11p13 in a region containing the Wilm’s tumor 1 (WT1) and paired box 6 (PAX6) genes (Farooqi and O’Rahilly 2005). A specific type of WAGR has been associated with a deletion in the brain-derived neurotrophic factor (BDNF) gene, which results in an obese phenotype.
Prader-Willi syndrome (PWS) can have several etiologies, characterized by central obesity, neonatal hypotonia, hyperphagia, hypothalamic hypogonadism and mild mental retardation, with such abnormalities as short stature and peculiar facial features (Farooqi and O’Rahilly 2005). Most of the cases were associated with loss of expression from paternal deletions of the 15q11.2-q12 chromosomal region (González-Jiménez et al. 2012).
Bardet-Biedl syndrome (BBS) is characterized by early-onset obesity, which is associated with progressive cone-rod dystrophy, morphological finger abnormalities, dyslexia, learning disabilities, and progressive renal disease (Farooqi and O’Rahilly 2005). BBS has extensive genetic heterogeneity with at least 14 loci, (often called BBS gene) and several mutations identified within these loci (González-Jiménez et al. 2012).
Alström (ALMS) and Cohen syndromes are associated with childhood mild truncal obesity and small stature (Farooqi and O’Rahilly 2005; González-Jiménez et al. 2012). Both of them are autosomal recessive and genetically homogenous. ALMS is caused by a balanced translocation of chromosome 2p13 that disrupts ALMS1 gene or by a small number of mutations in this gene. Cohen syndrome results from mutations in the COH1 gene, located at chromosome 8q22, which encodes a transmembrane protein of unknown function (Farooqi and O’Rahilly 2005).
Finally, we can also find ciliary dysfunction, collectively termed “ciliopathies”. These comprise a group of several disorders associated with genetic mutations encoding defective proteins, affecting normal function or formation of cilia (Bergmann 2012). Ciliary dysfunction can manifest as a set of heterogeneous features including retinal degeneration, renal disease, cerebral anomalies, congenital fibrocystic diseases of the liver and pancreas, diabetes, obesity and skeletal dysplasias (Waters and Beales 2011). Due to the heterogeneous phenotype, ciliopathies have been associated with mutations in more than 40 genes, including the genes involved in BBS and ALMS syndromes.
Polygenic or common obesity
In most modern societies, the environment favors promote weight gain rather than loss due to food abundance and lack of physical activity. The increase of common obesity in both adults and children resulted in a major increase in common obesity worldwide. However, the genetic and molecular mechanisms involved in body weight regulation are complex. The genetic profile of polygenic obesity results from the effects of several altered genes (Rankinen et al. 2006). In theory, the genetic basis of polygenic obesity implies that the specific set of variants relevant for obesity vary considerably from one obese person to the next (Hinney et al. 2010). For this reason, the study of common obesity is far more complex. However, the advent of new techniques facilitated this study by allowing the analysis of several loci at the same time.
Genetic approach for common obesity
The study of common obesity is based in the analysis of gene variation in genomic DNA (single nucleotide polymorphism, or repetition of bases of polyCAs or microsatellites) situated within or near candidate genes. In contrast with monogenic obesity, in polygenic obesity each polymorphism leads to a variant that confers susceptibility, requiring additionally the presence of other variants and an obesogenic environment to determine the obese phenotype (Razquin et al. 2011). There are some approaches used in the detection and analysis of a candidate gene in body weight regulation: linkage studies, candidate gene association studies and GWA studies. Their objective is to determine whether an association between a genetic variation and an obesity-related trait do exist. Until now, GWAS had identified more than 52 loci associated with obesity-related traits.
Recently, with the advent of automated DNA sequencing instruments, involving advances in engineering, chemistry, molecular biology, and software, a number of new opportunities have emerged (Mardis 2013). Currently, molecular diagnosis based on Sanger’s sequencing is restricted to only a few genes as this technology is expensive, time consuming, and labor intensive. The advent of next-generation sequencing (NGS) technology provides a new method for molecular diagnosis, allowing the sequencing of whole genomes or exomes, or several genes at the same time (Marian 2012). NGS promises to change the landscape of genetic testing with innovative cost-efficient methods for sensitive obesity multi-gene screening.
Only a few studies have used NGS technology to study obesity. Saeed et al. (2014) analysed 26 susceptible genes for obesity in a sample of 39 Pakistani children with early-onset obesity. They found two new LEPR mutations at the homozygous state: a splice site mutation in exon 15 (c.2396-1 G>T), and a nonsense mutation in exon 10 (c.1675 G>A). Sällman et al. (2013) amplified the entire region of FTO gene (412 kilo base pairs), from 524 severely obese and 527 lean Swedish children. They detected 705 single nucleotide polymorphisms, from which 19 were novel obesity-associated polymorphisms within the first intron of the FTO gene. An interesting finding was the fact that 10 of them have a stronger association with obesity (p < 0.007) when comparing with the commonly studied rs9939609 polymorphism (p < 0.012). This study concluded that within the entire region of the FTO gene the first intron was the only one associated with obesity. Bonnefond et al. (2014) searched for mutations with NGS in 40 patients, with a monogenic form of diabetes (n = 19) or obesity (n = 21), in which the causing mutation was already known. The study found the same mutations described as the phenotype cause, except for one variant (mean of 98.6 %). On the other hand novel mutations were found in 3 patients with a putative deleterious effect.
The NGS approach could be used as an efficient tool with highly sensitive screening for mutations in genes associated with obesity or other diseases. Further, sequencing the human genome can now be accomplished in the data-generation phase within 2 weeks at a cost of approximately, US $5000 (Mardis 2013). However, the price for genome sequencing continues to decrease; in 2014 Illumina announced that would produce a new system called HiSeq X Ten that can deliver full coverage of human genomes for less than US $1000. However, until now the majority of loci associated with obesity susceptibility were found by GWAS. For this reason, we describe the loci found by this technique in more detail and in a chronological order.
Common loci associated with obesity-susceptibility discovered through GWAS
The GWAS approach is the most commonly methodology used, allowing geneticists to scan numerous polymorphisms (~0.1–5 million of polymorphisms) across the entire genome using powerful statistical methods to identify loci associated with a particular phenotype. Since the start of the GWAS era in 2005, there have been five waves of GWAS’ discoveries for BMI. The first loci identified through GWAS was the fat mass and obesity-associated (FTO) gene, and until now more than 50 genetic loci have been identified as being associated with at least one obesity-related trait (Loss 2012; Sandholt et al. 2012; Xia and Grant 2013) (Fig. 1).
First discoveries by GWAS: the FTO gene
The first locus associated with obesity was the insulin-induced gene 2 (INSIG2) (Herbert et al. 2006). However, replication studies demonstrated very inconsistent results. So, the first locus unequivocally associated with obesity by a GWA study was the FTO gene (Frayling et al. 2007). Initially, Frayling et al. (2007) conducted a GWA study to test the correlation between polymorphisms across the entire human genome and type II diabetes (T2D). They found that the rs9939609 polymorphism, located in the first intron of the FTO gene was strongly associated with T2D and increased BMI. However, after adjustment for BMI, the apparent association of the polymorphism with T2D was not maintained. The effect size of FTO polymorphism on BMI is modest, with homozygous individuals for the risk allele (in this case the A allele) weighing on average 3 kg more than those homozygous for the protective allele (in this case the T), with the difference representing approximately, 0.36 kg/m2 (Xia and Grant 2013). These findings have been independently replicated and have consistently confirmed the association of rs9939609 polymorphism with the etiology of common obesity in several populations: European (Rodríguez-López et al. 2010; Albuquerque et al. 2013a; Zavattari et al. 2011; González et al. 2012), Asian (Chang et al. 2008; Hotta et al. 2008; Fang et al. 2010; Mačeková et al. 2012) and African (Grant et al. 2008; Song et al. 2008; Deliard et al. 2013), both in children and adults. Two following studies reported other polymorphisms in the intronic FTO region also consistently associated with severe early-onset childhood and adult obesity (rs1421085 and rs17817449) (Dina et al. 2007), and have extended the association to other obesity-related traits including body weight and waist-to-hip circumference ratio (WHR) (rs9930506) (Scuteri et al. 2007). The FTO polymorphisms were also associated with abdominal obesity, waist circumference and waist-to-hip ratio (WHR) (Heard-Costa et al. 2009; Lindgren et al. 2009), and also with body-fat percentage (Kilpeläinen et al. 2011a). Although these posterior reports replicate well the initial findings, the FTO polymorphisms explain only 1–3 % of the variance in BMI (Frayling et al. 2007; Scuteri et al. 2007).
To date, over 500 studies have been performed concerning the association of FTO polymorphisms with obesity in several populations worldwide, and more than 60 polymorphisms in this gene were significantly associated with obesity (Jacobsson et al. 2012). All these polymorphisms were found within a 47 kb linkage disequilibrium (LD) block encompassing parts of the first two introns as well as exon 2 of the FTO gene (Fawcett and Barroso 2010). This is a region where the sequence is strongly conserved across species, were polymorphisms are highly correlated (LD r 2 > 0.80 in CEU of the HapMap) in Caucasian populations (Jacobsson et al. 2012).
The functional mechanism underlying FTO role in obesity remains unknown, as well as the pathway underlying that role. The FTO is a very large gene with 9 exons spanning more than 400 kilobase (kb) in the chromosome 16q12.2 (Loos and Yeo 2014). It was originally identified in 1999 in the Fused toes (Ft) homologue mutants, in a deletion of 1.6 megabase (Mb) on chromosome 8 (Peters et al. 2002). Homozygosity of Ft mutants is embryonically lethal. To investigate the biological function of FTO gene, two mouse models were used. Homozygous FTO −/− mice introduced by Fischer et al. (2009) show postnatal growth retardation, significant reduction in fat and lean body mass compared to the wild-type animals (Church et al. 2009). In other mice model, Church et al. (2010) observed a lean phenotype in mice carrying a missense mutation in exon 6 of FTO (FTOI367F mice). These results seem to indicate that FTO could play a role in food intake control, energy expenditure and homeostasis.
The predicted human protein consists of 505 amino acids, characterized as a 2-oxoglutarate-dependent enzyme that is localized in the cell nucleus, belonging to the (2OG) oxygenases AlkB family of proteins (Gerken et al. 2007). The AlkB is a DNA repair enzyme, which catalyzes Fe(II)- and 2OG-dependent demethylation of damaged DNA substrates (van den Born et al. 2008). Recently, Jia et al. (2011) indicated that FTO also demethylates N6-methyladenosine (m6A) residue in nuclear RNA. FTO variation appears to lead to an increase in energy intake (Speakman et al. 2008) by modifying hypothalamic control of appetite (Jacobsson et al. 2012). The crystal structure of FTO has recently been published and reveals the basis for its substrate specificity (Han et al. 2010). Moreover, it was found that the FTO gene is also a transcriptional coactivator (Wu et al. 2010) and a possible regulator of telomere length (Dlouha et al. 2012). A recent study found that BMI-associated FTO variants interact with the promoter region of iroquois homeobox 3 (IRX3) gene in the human, mouse and zebrafish genomes (Smemo et al. 2014). The study also found that in Irx3-deficient mice, there is a reduction in body weight of 25–30 %. This study suggests that IRX3 gene is a functional long-range target of obesity-associated polymorphisms within FTO. On the other hand, the FTO deficiency remains still poorly understood in the context of obesity development and confirm the complexity of the genetics underlying common obesity.
Five waves of GWAS
Following the discovery of the FTO locus, investigators enhanced GWA studies by increasing the sample size improving statistical power to uncover additional obesity-susceptibility loci (Table 2). Subsequently, a large-scale international consortium, called the Genetic Investigation of Anthropometric Traits (GIANT) emerged. The association data of 16,876 Caucasians from seven GWAS for BMI were combined in a meta-analysis (Loos et al. 2008). This study confirmed the strong association of obesity with polymorphisms in the FTO gene, and identified one new locus near the MC4R gene which mutations are known to be the common cause of extreme childhood obesity (Farooqi and O’Rahilly 2005). The MC4R was the second gene significantly associated with common obesity (Chambers et al. 2008; Loos et al. 2008). The rs17782313 polymorphism near the MC4R gene was associated with obesity among both adults and children (Loos et al. 2008). Another polymorphism (rs12970134) near the MC4R gene also appears to increase the risk of obesity among Europeans (Thorleifsson et al. 2009). Several polymorphisms near the MC4R gene have subsequently been found and replicated in various European populations, as well as in Asians (Xi et al. 2012), African-American (Xi et al. 2012), both in children and adolescents (Grant et al. 2008; Deliard et al. 2013).
In the third wave of discoveries, a meta-analysis was performed using 15 GWAS for BMI in Caucasians (n > 32,000) and replicated in another 14 studies for a second-stage sample of 59,082 individuals (Willer et al. 2009). They confirmed the association of the FTO and MC4R genes, and found six new genes positively associated with obesity: MTCH2, GNPDA2, KCTD15, SH2B1, NEGR1 and TMEM18. At the same time, a GWAS of 31,392 individuals, predominantly from Iceland population, found seven new genetic loci near or in: BDNF, SEC16B, ETV5 and FAIM2, as well as FTO and MC4R genes associated with BMI (Thorleifsson et al. 2009). Four of the seven newly identified loci were common with the results from Willer et al. (2009).
In 2010, the fourth wave, the GIANT consortium expanded its GWAS stage to comprise 249,796 individuals of European origin, and reveal 18 new loci associated with BMI near or in: PRKD1, SLC39A8, GPRC5B, MAP2K5, QPCTL, RBJ, LRRN6C, FLJ35779, CADM2, TMEM160, FANCL, LRP1B, TNNI3 K, MTIF3, TFAP2B, ZNF608, NRXN3, RPL27A, PTBP2 and NUDT3 (Speliotes et al. 2010). By 2011, GWAS had identified 32 genetic loci unequivocally associated with BMI.
The most recent and fifth wave expanded the GIANT meta-analysis, to comprise 263,407 individuals of European ancestry (Berndt et al. 2013). Besides confirming all 32 BMI-associated loci previously identified by the fourth wave, they found seven new loci, ZZZ3, RPTOR, ADCY9, GNAT2, MRPS33P4, HS6ST3 and HNF4G, explaining an additional 0.09 % of the variability in BMI (Berndt et al. 2013).
To date, more than 35 loci have been found associated with the increase of BMI (explaining ~1–3 % of the variance in BMI), while other loci correlate with abdominal obesity, establishing 13 loci associated with it, assessed by the WHR (Heid et al. 2010). Other loci, such as the Lactase gene (LCT) have been associated with BMI and abdominal obesity, but more studies are required to confirm associations (Kettunen et al. 2010; Corella et al. 2011; Almon et al. 2012; Albuquerque et al. 2013b). A study identified two new loci with body fat percentage: IRS1 and the other near SPRY2 (Kilpeläinen et al. 2011b). There is a gap between the explained variance of BMI due to known common polymorphisms (1–4 %), and the estimated heritability (40–70 %). One of the main problems pointed out in GWAS is the failure to detect loci that are associated with traits whose effect sizes are too small to reach genome-wide statistical significance (false negative rate). To circumvent this “missing heritability” the genome-wide complex trait analysis (GCTA) method appears to show a multitude of low penetrance common polymorphisms, each with causal effects but too small to allow detection by GWA studies. Using this approach, Yang et al. (2011) estimated in 17 % the BMI variation due to common genetic variants, and a recent analysis of twin studies revealed that additive effects of multiple common polymorphisms could explain 37 % of BMI (Llewellyn et al. 2013). Most GWAS were performed in samples of Caucasians adults, and only a few in children. However, it seems important to determine the genetic predisposition in children, as obesity tends to develop from childhood into adult life.
Testing adult-discovered loci in children
Childhood obesity is a major health problem in developing countries throughout the world. Most of obesity susceptible genes were found in studies with adults, which prompted an effort to replicate findings in studies with children (Zhao et al. 2011; Albuquerque et al. 2014; Deliard et al. 2013). Knowledge of the genetic risk factors of obesity in children could be used as a first step to develop possible prevention measures. The FTO locus remains the most replicated gene and the strongest gene associated with obesity susceptibility, both in adults and children (Albuquerque et al. 2013a; Deliard et al. 2013; León-Mimila et al. 2013). Results from longitudinal studies suggested a possible age-related change in the association between the FTO rs9939609 polymorphism and higher BMI. Sovio et al. (2011) studied subjects of European ancestry aged from early infancy to 13 years. In that sample, individuals’ carriers of the minor A-allele of this polymorphism showed lower BMI in infancy and higher BMI later in childhood. In another study, Hallman et al. (2012) analysed a sample of non-Hispanic white children and adolescents (8–18 years). It was found a significant age-by-genotype interaction predicting that in individuals with AA genotype the BMI would be ~0.7 kg/m2 higher at age 8, and ~1.6 kg/m2 higher at age 17, comparing to those with AT or TT genotypes. The results reported in these studies might help to reveal mechanisms regulating body mass in humans during a critical period of development. Genes TMEM18 and GNPDA2 were also associated with obesity susceptibility, with a similar effect of the FTO gene (Zhao and Grant 2011). The remaining loci with evidence for association were INSIG2, MC4R, NEGR1, BDNF and KCTD15 (Zhao and Grant 2011; Mitchell et al. 2013).
In the GIANT meta-analysis of adult BMI in a pediatric European American sample, Zhao et al. (2011) examined 32 genetic loci in 1097 obese cases and 2760 lean controls, aged between 2 and 18 years old. They found evidence of associations with nine of these loci, namely at FTO, TMEM18, NRXN3, MC4R, SEC16B, GNPDA2, TNNI3 K, QPCTL, and BDNF. Overall, 28 of the 32 loci showed directionally consistent effects to that of the adult BMI meta-analysis.
Another similar report by the early growth genetics (EGG) consortium investigated the effect of established adult BMI with two recently associated loci with childhood obesity (HOXB5 and OLFM4 genes) (Bradfield et al. 2012) in a Greek adolescents cohort (Ntalla et al. 2013). The genetic risk score of the 34 (GRS-34) variants was calculated and found that variants at the FTO, TMEM18, FAIM2, RBJ, ZNF608 and QPCTL loci produced nominal evidence for association with BMI and/or obesity risk. Overall, 27 out 34 variants showed consistent effects with those reported by large-scale meta-analyses adult BMI.
These results showed clearly that these obesity-conferring variants operate early in life, suggesting that individual preventative lifestyle intervention in childhood could be important to obesity development.
GWAS-related investigations in other ethnicities
There are remarkable disparities in the prevalence of obesity between ethnic groups. To date most of GWAS published reports have been performed in populations of European origin. Only one study identified, at the first discovery stage, a locus near MC4R gene associated with waist circumference and insulin resistance in a cohort of South Asian population (Chambers et al. 2008). This could be partly due to the fact that some susceptible loci only affect a specific ethnic group, while others might affect any ethnic group. Indeed, the human genetic architecture differs across ethnicities, which is well-illustrated by differences in linkage disequilibrium (LD), whereas haplotype blocks vary only somewhat among human populations (Slatkin 2008).
As a case in point, FTO locus also have consistently correlated with BMI and risk of obesity in populations of African (Grant et al. 2008; Song et al. 2008; Hennig et al. 2009; Deliard et al. 2013), Asian (Chang et al. 2008; Hotta et al. 2008; Fang et al. 2010; Mačeková et al. 2012) and Pacific-Islander (Ohashi et al. 2007) ancestry. Despite the fact that effect sizes were similar to those observed in white European populations, the risk allele frequency varies substantially: around 45 % in white Europeans, ~25 % in Asian, and range of ~7–18 % in African origin (Hassanein et al. 2010). In the case of FTO gene, Peters et al. (2013) genotyped 3756 polymorphisms across a 646 kb region, encompassing the large FTO gene (16q12.2) and the flanking gene RPGRIP1L in 20,488 African Americans. Authors reported the rs56137030 polymorphism as the most significantly associated with BMI. Interestingly, they found that in individuals of Europeans ancestry, this polymorphism represents a cluster of 103 polymorphisms (r 2 > 0.50), whereas in African Americans this cluster includes only 29 polymorphisms (at r 2 > 0.50).
Two recent independent meta-analysis studies were performed in both East Asian and African populations (Wen et al. 2012; Monda et al. 2013). Wen et al. (2012) performed a meta-analysis using 27,715 individuals, followed by in silico and de novo replication studies in a further 37,691 and 17,642 individuals of East Asian origins, respectively. Seven previously identified loci were detected (FTO, SEC16B, MC4R, GIPR-QPCTL, ADCY3-RBJ, BDNF and MAP2K5) and three new loci were uncovered, near or in CDKAL1, PCSK1 and GP2 genes. Data also implicated three loci, GNPDA2, TFAP2B (previously identified) and PAX6, which all reached the genome-wide significance threshold. A recent meta-analysis was conducted to examine the association of >3.2 million polymorphisms with BMI in 39,144 adults of African ancestry (Monda et al. 2013). It identified one new locus at 5q33 (GALNT10, rs7708584 polymorphism) and another at 7p15, when data from the GIANT consortium was included (MIR148A-NFE2L3, rs10261878 polymorphism). They also found evidence of an association at 6q16 (KLHL32, rs974417 polymorphism) in African-ancestry sample. Overall, 32 of the 36 previously established BMI variants showed consistent effect in this GWAS. The 36 known BMI loci explain in average 1.30 % of the variance in BMI of African ancestry compared with 1.67 and 1.25 % in European and Asian ancestry populations, respectively (Monda et al. 2013). More recently, Tan et al. (2014) replicated six confirmed obesity genes (FTO, CTNNBL1, ADRB2, LEPR, PPARG and UCP2 genes) in eight different population samples from different ancestries (five Caucasian, one Chinese, one African-American and one Hispanic population). The main goal of this study was to explore whether the same genes contribute differentially to obesity susceptibility in populations of different ancestries. Regarding the FTO gene they found 35 polymorphisms significantly associated with obesity in Caucasian populations. However, none of them showed evidence of associations with obesity in another ethnic group.
Association studies across different populations can help us to define more precisely which loci or variants could play a role in the obesity etiology, and help to understand the genetic and environmental factors contributing to obesity. As we can see, polymorphisms in the FTO gene are associated with obesity in several ethnicities. However, the allele frequencies of the BMI-associated FTO polymorphisms vary substantially across the different ethnicities. The highest prevalence of minor risk allele is observed in Europeans and the smaller frequency in Asian and African populations. The LD cluster of FTO polymorphisms was clearly demonstrated by Loos and Yeo (2014), in which Europeans present the larger cluster, followed by Asian populations that do not overlap with the African cluster (probably without association with obesity-related traits). The discovery of new loci in replication studies at established loci found in other populations reflect differences in allele frequency and effect size. Further studies will be needed to test the biological function at the associated loci and take into account difference observed regarding LD.
Obesity risk-allele scores
As noted, several GWAS have identified a large number of obesity susceptibility loci. Nevertheless, the major part of these studies only identified single genetic loci associated with obesity. It has indeed been demonstrated that combining information from all these obesity loci into genetic risk-allele scores (GRS) could be a convenient way to summarize risk-associated variations across the genome (Horne et al. 2005) and more useful when individual genetic effects are moderate (Belsky et al. 2013). The simplest way to calculate a GRS is by summing the number of accumulated risk alleles associated with the disease. Using this approach, Zhu et al. (2014) analysed 28 BMI-associated polymorphisms in a sample of Han Chinese and found 26 nominally associated with BMI. To assess the combined effect of all polymorphisms studied with BMI, they create a GRS which was associated with increased risk of obesity (OR 1.06; CI 95 % 1.03–1.10), and each additional BMI-increasing allele in the GRS was associated with 0.11 kg/m2 higher BMI (p = 1.54 × 10−7). Willer et al. (2009) found effect sizes between 0.06 and 0.33 kg/m2 per allele in BMI changes and that account for 0.40 % of the variance of BMI analysing six loci together (TMEM18, KCTD15, GNPDA2, SH2B1, MTCH2 and NEGR1). When they included the FTO and MC4R genes in the combined effect the variance increases to 0.84 %. Similar results have also been found in other studies trying to explain the variance of BMI. Combining 12 polymorphisms in a sample of 20,431 of European descent, the GRS obtained by Li et al. (2010) explained 0.9 % of BMI variation. Apart the nominal association between 15 polymorphisms located in or near the INSIG2, FTO, MC4R, TMEM18, GNPDA2, NEGR1, BDNF, KCTD15, and 1q25 genes with BMI, Zhao et al. (2009) explained 1.12 % of the total variation for BMI z-score in a sample of children of European ancestry. In other sample of European descent González et al. (2013) create a GRS including six polymorphisms located in the FTO, TFAP2B, SEC16B, ETV5 and SH2B1 genes and found that individuals carrying ≥7 risk-alleles had 3.1 (OR 3.11; CI 95 % 1.58–6.61) times increase in the odds of developing the obese phenotype. Individually, each risk allele conferred an estimated increased risk of 1.69 (OR 1.69; CI 95 % 1.46–1.97) times to develop obesity.
The use of a combined genetic score is considered as a better tool to determine the susceptibility of a common trait, than using each genetic locus alone. This is particularly, more evident when the allele score consists either of many common polymorphisms with small effects, or of rare polymorphisms (Belsky et al. 2013). Generally, when several polymorphisms are combined, the estimated genetic score may explain a considerable proportion of variation in the risk factor, even if none of the polymorphisms individually does. This is partially due to the fact that the “signal” obtained from a GRS is more robust to imperfect linkage than each polymorphism individually (Belsky et al. 2013). In complex diseases it is likely that the effects of different genetic loci-related to obesity operate in an interactive fashion. Future research should investigate this possibility using classification or regression tree analyses, which are well-suited to detecting complex non-linear interactions. The identification of the complex interplay among all genes in the genome-wide context is essential to unravel the molecular mechanisms in the obesity etiology. However, as previously demonstrated there are differences between populations regarding to allele frequencies. Belsky et al. (2013) developed a GRS for obesity using results obtained in 16 previously published GWAS in European descent samples. Analysing 32 locus they found a significantly predictor of BMI and obesity among Europeans. However, the predictive effects for this GRS did not replicate among African Americans due particularly to the differences in risk-allele distributions. In less than 10 years, we assisted to the discovery of several genes associated with obesity-related traits. Despite the discoveries, all these genes only explain a small percentage of obesity susceptibility. Of course, several genes remain to be found and certainly in the next year’s novel candidate loci will appear. However, and more recently, a new field called epigenetic emerged as a new potential factor influencing the obese phenotype and helping to find differences in obesity risk based in the environment that surrounds us.
Epigenetics
Epigenetic regulation of gene expression emerged in the last few years as a potential factor that might explain individual differences in obesity risk (Campión et al. 2009). Epigenetics can be defined as heritable changes that are mitotically stable (and potentially meiotically) and affect gene function but do not involve changes in the DNA sequence (Bird 2002). At the molecular level, epigenetic markers include genomic DNA methylation, changes in chromatic organization by histone modifications, the non-coding micro RNAs (miRNA), genomic imprinting, non-covalent mechanisms, and other nuclear proteins that are critical for epigenetic gene regulation (Kim et al. 2009). Currently, there is a growing interest in the study of the relations between genetic variation, epigenetic variation, and disease simultaneously.
Emerging studies have characterized the potential mechanisms by which epigenetic factors could increase the risk for obesity (Table 3). Moreover, unlike DNA genotypes, epigenetic markers can change during lifetime, and have a heterogeneous distribution in tissues. DNA methylation is the most well-known epigenetic marker, which has been proposed as a new generation of biomarkers. It is a biologic process that consists of the addition of a methyl group at the carbon-5 position of cytosine, in the context of the CpG dinucleotides, and usually associated with gene silencing in the promoter regions (Costello and Plass 2001; Bird 2002). The universal methyl donor is DNA methyltransferases (Dnmts) that maintain the cellular DNA methylation patterns (Campión et al. 2009). Despite the high number of DNA methylation candidate genes and some epigenome-wide association studies (EWAS), most of the associations have not yet been replicated in other samples to further confirm and establish whether those loci are reliably associated with obesity.
Using a genome-wide approach, obesity has been related to changes in DNA methylation status in peripheral blood leukocytes of lean and obese adolescents for two genes. In the ubiquitin-associated and SH3 domain-containing protein A (UBASH3A) gene, a CpG site showed higher methylation levels in obese cases, and one CpG site in the promoter region of Tripartite motif-containing 3 (TRIM3) gene, showed lower methylation levels in the obese cases (Wang et al. 2010). In a recent work, Godfrey et al. (2011) measured the methylation status of 68 CpGs 5′ from five candidate genes in umbilical cord tissue DNA from healthy neonates, and found that methylation higher levels within promoter region of retinoid X receptor-a (RXRA) gene, measured at birth, was strongly correlated with greater adiposity in later childhood (Godfrey et al. 2011). A positive correlation between maternal BMI and promoter methylation in peroxisome proliferator-activated receptor-gamma co-activator 1alpha (PPARGC1A), a gene encoding a transcriptional coactivator of the peroxisome proliferator-activated receptor (PPAR) α and У, playing an essential role in energy homeostasis, was observed when analysing promoter genomic DNA from umbilical cord newborns (Gemma et al. 2009).
The obesity risk allele of FTO has been associated with higher methylation of sites within the first intron of the FTO gene, suggesting an interaction between genetic and epigenetic factors (Gemma et al. 2009). Moreover, Almén et al. (2012) determined the methylation profile on a genome-wide scale by sampling DNA from peripheral whole blood in female preadolescents. The sample included obese and a normal weight groups, both of which contains homozygous carriers of both the FTO normal and risk alleles (rs9939609). They analysed how the risk allele for rs9939609 polymorphism affects the methylation status of sites related to other genes (KARS, TERF2IP, DEXI, MSI1, STON1 and BCAS3), showing that the FTO gene may influence the methylation level of other genes (Almén et al. 2012).
A study examined the MC4R gene, which is associated with common and morbid obesity and encodes for a protein that is a membrane-bound receptor and member of the melanocortin receptor family controlling food intake and energy expenditure. Mouse genomic DNA of brain tissue was examined to determine the methylation status of the MC4R exon. Results indicated that methylation of the CpGs was decreased in response to high-fat diet (Widiker et al. 2010). A study examining whether a high-energy diet may affect promoter methylation of LEP gene, encoding an adipokine involved in body weight and food intake regulation, showed in DNA isolated from retroperitoneal adipocytes in rats that leptin methylation pattern can be influenced by diet-induced obesity (Milagro et al. 2009). Zhao et al. (2013) demonstrated that promoter hypermethylation in the serotonin transporter gene (SLC6A4) was associated with an increase in BMI, body weight and waist circumference. Xu et al. (2013) studied 470,000 CpG sites from 48 obese and lean youth African-American (14–20 years-old); they found a differential variability in CpG sites which was more variable in obese than lean subjects, constituting an important feature of obesity related with methylation changes. In another recent EWA study, Rönn et al. (2013) analyzed 476,753 CpG sites to evaluate the possible alteration of DNA methylation patterns after a 6-month exercise intervention. A global DNA methylation changes were found in 17,975 individual CpG sites altering the levels of DNA methylation in response to physical activity (Rönn et al. 2013).
Thus, most of these DNA methylation sites need to be confirmed as being associated with obesity, taking into account the tissue sampled, obesity history, and eating behaviors. However, the high number of new studies concerning obesity epigenetics will undoubtedly permit the confirmation some of these associations, thereby establishing an epigenetic basis for human obesity. Interestingly, one recent work of genome-wide analysis revealed that carriers of the FTO risk allele (rs9939609) had a significant differential methylation level in 6 loci (KARS, TERF2IP, DEXI, MSI1, STON1 and BCAS3) compared to non-carriers controls (Widiker et al. 2010). This work could elucidate the mechanisms underlying the association of obesity with genetic variants, possibly due to epigenetic factors.
Studies based on pre-conceptual, in utero, and postnatal developmental environment showed an impact on long-term risk for adult-onset obesity by a set point of adaptive changes. It could be understood as a “critical period” where environmental conditions experienced in utero may have a life-long effect on the propensity to develop the obese phenotype. The Agouti mouse viable yellow (A vy) model is one of the best examples on how early environmental exposures interact with epigenetic gene regulation influencing the phenotype (Wolff et al. 1998). Briefly, the murine agouti gene influences DNA methylation in early development, affecting coat color, which correlates with adult body weight. Varying the mother´s diet tends to produce offspring with a wide variation in individual coat color and obese phenotype as epigenetic modifications of agouti gene were established in early development (Waterland and Jirtle 2003). These variations in phenotypes are caused by DNA methylation patterns, which were acquired during early embryonic development and passed through the female germline that results in stable intergenerational transmission (Khosla et al. 2001).
In a recent report, Relton et al. (2012) evidenced that DNA methylation patterns in 9 of 24 (37.5 %) genes at birth, show association with at least one index of body composition (BMI, fat mass, lean mass, height) at age of 9 years. This observation suggests that variation in DNA methylation patterns at birth in multiple target genes may influence body size in childhood. Moreover, maternal diet can alter later child's adiposity, accompanied by epigenetic changes in genes controlling the energy homeostasis. Parental pre-conceptional environmental exposures could also have an effect in the health status of the offspring in later life. In two recent studies regarding parental obesity conducted by Soubry et al. (2013a, b) it has been observed an association between DNA methylation profiles at human imprinted genes, such as MEST, PEG3, and NNAT, in children born from obese parents, when compared with children born from non-obese parents. Changes related to maternal obesity were also detected at loci PLAGL1, MEG3 and H19 (Soubry et al. 2013a, b). Hypomethylation at the IGF2 gene was associated with paternal obesity (Soubry et al. 2013a). These results points to a pre-conceptional influence of parental life-style or over-nutrition on the reprogramming of imprint marks during gametogenesis and early development (Soubry et al. 2013a, b). The trans-generational effects of parental obesity can influence the offspring’s future health status. These reports evidenced that peri-natal events are important in defining the epigenetic marks that will persist until the adult age. In the future, it might be used as early prognostic markers to identify those individuals with more risk to develop obesity. However, the knowledge of mechanisms by which maternal nutritional environment induces such changes remains largely unknown.
microRNAs
Another type of epigenetic mechanisms is microRNAs (miRNAs). The gene expression in humans is precisely controlled in cellular, temporal, and condition specific manner. Because miRNAs have been shown to be important in gene regulation, it is not surprising that they have been implicated in the development of obesity (Williams and Mitchell 2012). Therefore, the understanding of the regulatory mechanisms of gene expression can shed some light on the underlying mechanisms causing obesity. miRNAs are endogenous short single-stranded non-protein-coding RNAs with about 21/25 nucleotides in length which are involved in post-transcriptional regulation of gene expression by partially complementary binding to the 3′ untranslated region (3′ UTR) of target mRNAs (Ambros 2004; Bartel 2004).
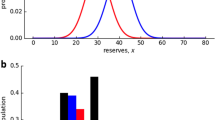
Several miRNAs expression patterns have been profiled during adipocyte differentiation (Kajimoto et al. 2006; Xie et al. 2009; Ortega et al. 2010), others have been linked to adipocyte phenotype, and other obesity parameters (Kajimoto et al. 2006; Takanabe et al. 2008; Klöting et al. 2009; Xie et al. 2009; Ortega et al. 2010, 2013; Heneghan et al. 2011; Keller et al. 2011) (Fig. 2). For example, miR-21 was strongly expressed in human adipose tissue and positively correlated with BMI (Keller et al. 2011). These studies revealed that miRNAs may represent biomarkers for obesity, and could also be implicated in the molecular mechanisms leading to this disease. However, further studies are needed to elucidate the effect of miRNAs and other epigenetic mechanisms in the etiology of obesity.
Continuous advances in research show promising results about the implication of epigenetics mechanisms in the etiology of obesity. Epigenetics has shown that our genes are not the only factor to determine our phenotype and that our behaviors can alter the expression of our genotypes. However, additional research is needed, particularly with regard to which cell types should be explored in EWAS.
Evolutionary explanations for obesity
The evidence for a genetic component of obesity has been well-established in recent years (Bell et al. 2005). The question about the evolution of obesity is: how has natural selection favored the spread of genes that increase risk for an obese phenotype and how has this predisposition to obesity evolved? Answers to these questions should advance the understanding of the etiology of obesity. However, our knowledge about the evolution of body-weight regulation mechanisms in humans remains incomplete. Nevertheless, four different types of evolutionary perspective have been proposed in an attempt to address these questions (Speakman 2013; Sellayah et al. 2014).
The first hypothesis, called “thrifty gene”, is that the modern genetic predisposition to obesity was adaptive in the past, when storing large amounts of fat could have been selectively advantageous (Neel 1962). It explain the prevalence of obesity and diabetes in modern societies due to a change in lifestyle from that of Paleolithic hunters–gatherers to a subsistence based on agriculture, a pattern characterized by more sedentary occupations. The basis of this hypothesis states that during evolution of the modern humans, genes that promoted efficient fat accumulation would be extremely advantageous for primitive humans, because they allow their holders to survive famine periods (Speakman 2006). In modern societies, where food supply is always available, such genes are disadvantageous and the result is the widespread obesity (Speakman 2006, 2013).
Studies were conducted to try and identify genes under positive selection that have a role on obesity. A study lead by Myles et al. (2011), suggested that the high frequency of the risk allele of the Gly482Ser variant in the PPARGC1A gene in Polynesians populations remains a thrifty allele in the Pacific populations. Another variant, the PC-1 Gln121, was also considered as a possible thrifty gene supported by studies in African and other groups (Rey et al. 2012). A recent study provides evidence for a positive selection of TRIB2 gene, which influences visceral fat accumulation in East Asians (Nakayama et al. 2013). In addition to these few examples showing a possibly positive selection in our evolutionary history with metabolic traits, other loci have been extensively studied, one of them being the LCT gene, at ~7000 years bp, which is considered a prototypic example of selective advantage leading to rapid human evolution compatible with the agricultural innovations (Bersaglieri et al. 2004). In European populations, the −13,910 C>T (LCT) polymorphism has been associated with the persistence of the lactase enzyme in adulthood: individuals carrying the CC genotype possess insufficient enzyme activity in intestinal cells and are classified as lactase non-persistence (i.e., show lactose intolerance), which is considered the ancestral condition in humans, whereas individuals carrying at least one T allele are considered lactase persistent (Enattah et al. 2002). Nevertheless, this adaptive hypothesis reveals some problems: if accumulating extra adipose tissue was advantageous in the past populations, many people with these thrifty genotypes in modern society do not develop the obese phenotype, despite the environmental change favoring fat storage. On the other hand, population genetic models predict that thrifty genes would not have sufficient advantage or even time to spread in the human population (Speakman 2004).
A second explanation, for the evolutionary selection favoring obesity is that most mutations in the obesity susceptibility genes are neutral and have been drifting over evolutionary time (Speakman 2008, 2013). The neutral theory of molecular evolution, postulates that most evolutionary changes at the molecular level is not caused by natural selection, but by genetic drift (Kimura 1983). According to this theory, the majority of genetic variation observed within and between species is selectively neutral, i.e. does not affect the fitness of individuals, in contrast to the theory of natural selection for which most of the genetic variation observed in populations affect the fitness of individuals and thus is subject to selection (Nielsen 2005). This new “drifty genes” hypothesis is a non-adaptive scenario providing an explanation for why some individuals get obese while others remain obesity resistant (Speakman 2008, 2013).
The maladaptive viewpoint is another hypothesis that suggests that obesity is not adaptive and may never even have existed in human evolution history, except in some individuals with unusual genetic modifications such as the monogenic forms of obesity (Speakman 2013). Nevertheless, genes that actually predispose to obesity could be favored as a maladaptive by-product of positive selection on some other advantageous trait. One example of this maladaptive interpretation is a work suggesting that obesity could result from individual differences in brown adipose tissue (BAT) (Sellayah et al. 2014). This recent point of view emerged highlighting the differences in genetic susceptibility to develop an obese phenotype based on BAT. Sellayah et al. (2014) proposed the thermogenesis hypothesis, in which climatic selection pressures in the evolutionary history could exert a strong influence on genes. Effectively, climate changes played a key role in our evolution, representing the principal engine of evolutionary change (climate is an important factor in determining anatomical differences among different geographical populations). In fact, we can see around the world several differences among warm-adapted and cold-adapted species.
Regarding human populations there is a significant variation in their body form, by an adaptation to different climates, suggesting a link between body shape and climate, probably related to thermoregulation. In warm climates, individuals have a large surface area relative to body mass (e.g. slim, long trunk) that facilitate heat loss, whereas in cold climates individuals have a small surface area relative to body mass (e.g. bulky, short trunk) allowing heat retention. In most eutherian mammals, BAT is an essential factor in thermogenesis helping to maintain their body temperature regardless of the ambient temperatures. The uncoupling protein 1 (UCP1) gene was found with a key role to maintain body temperature in cold climates, which is highly expressed in BAT (Cannon and Nedergaard 2004). Some polymorphisms were found in the UCP1 gene associated with body fat accumulation, body weight gain and BMI in response to a high-fat diet (Jia et al. 2010). Feldmann et al. (2009) demonstrated in mice exempt from thermal stress that UCP1 ablation induced obesity, even in mice fed with control diet. They conclude that ambient temperatures have an important role in UCP1 in mediating diet-induced adrenergic thermogenesis. They suggest that UCP1 activity could be determinative for obesity in mice, and possibly in humans. Epidemiological studies found different rates of obesity (including other metabolic disorders) across certain ethnic groups. These studies demonstrated that in United States there are ethnically differences; with blacks, Hispanics, and people of Native American ancestry being more prone to develop an obese phenotype than European Caucasians and people of East Asian ancestry (Chinese, Japanese, and Koreans). When we look to these ethnic population distributions it could be observed that people living in warm climates have a higher obesity prevalence compared to people living in cold temperatures. In evolution history, genes with an essential role to survival, especially in newborn and young children, were positively selected.
Sellayah et al. (2014) proposed in the thermogenesis hypothesis that migration to colder climates could result in a more efficient BAT and UCP1 gene function. This fact, would endow the capacity of higher energy expenditure and energy-burning capacity, providing a higher metabolic rate, which then could reduce body fat. At the opposite side, Africans and South Asians, whose ancestors had no need to evolve efficient BAT and UCP1 function due to the warm climate, show an increased propensity for obesity when subjected to sedentary and hypercaloric lifestyle.
Regarding the thrifty and drifty genotype hypotheses that attempt to explain how human obesity evolved, at least one fact remains unclear. Obesity emerged in industrialized countries, which then exported their sedentary and hypercaloric lifestyle. One of the principal drawbacks in these two hypotheses is that it cannot explain the clear evidence for ethnic differences in the susceptibility to develop obesity. On the opposite side, the thermogenesis point of view it will be interesting; as during the out of Africa migration, genes involved in the BAT thermogenic function could be positively selected to a better cold adaptation, although it has also some drawbacks. So, further investigation in the UCP1 gene and other in genes involved in metabolic regulation could help unravel the causes of obesity susceptibility, and explain the differences among populations and why not all people in the same environment seem to have the same predisposition for obesity. However, it should be mentioned that these hypotheses are not mutually exclusive and it is possible that all have some valid arguments explaining the evolutionary origin of obesity. Thus, understanding human evolution could help us to understand modern human behavior and traits.
Prevention and treatment based on genotyping
Nutrigenetics
Nutrition is one of the lifestyle factors contributing to the development and progression of obesity. An appropriate intake of energy and nutrients has been commonly accepted to prevent weight gain. Furthermore, epigenetics studies have demonstrated that several nutrients and bioactive food could play a role in the complex machinery involving the interaction between genome and the epigenome, which regulate gene expression (McKay and Mathers 2011). The ingestion of these nutrients introduces some bioactive components that have signal molecules that carry information from the external environment (Milner 2004). Many dietary components can modulate epigenetic phenomena by inhibiting enzymes such as DNA methyltransferases and histone deacetylases (McKay and Mathers 2011), with the most well-known vitamin B-12 and folate providing methyl groups for DNA methylation reaction (Brunaud et al. 2003; Choi et al. 2004). New research has attempted to understand the variability in metabolic responses to diet and food components, which could affect health. Nutrigenetics recognizes the effect of genetic variation on nutrient requirements, while nutrigenomics study the interaction between nutrients and genes (Steemburgo et al. 2009). These areas aim to develop diagnostic tools that can “read” genetic susceptible loci to offer a personalized diet, taking into account the individual needs. Interactions among genetic loci and diet were found for obesity in interleukin-6 (IL-6), with daily food intake, peroxisome proliferator-activated receptor gamma 2 (PPAR-gama2) and FTO with fat intake (Steemburgo et al. 2009). The Mediterranean diet is known to be rich in folates, which is crucial for the DNA methylation status. Ortega-Azorín et al. (2012) found a significant gene-diet interaction of the FTO rs9939609 and MC4R rs17782313 polymorphisms with type 2 diabetes depending on diet, in which the Mediterranean diet counteracts the genetic predisposition. A cross-sectional study found that individuals carrying both AA risk allele of the rs9939609 polymorphism were positively associated with a high intake of total fat (>34 % energy) and low-fiber consumption (<16 g/day), independently of BMI (Steemburgo et al. 2013). It has also been reported in a recent study that obesity susceptibility genes (FAIM2, FLJ35779, FTO, LRRN6C, RBJ, and SEC16B) were found to interact with dietary carbohydrates (sugar-sweetened beverages) to increase BMI when one or more servings are consumed per day (Qi et al. 2012). Other two genes, β-adrenergic receptor 2 (ADRB2) and MC4R were also suggested being related with carbohydrate intake (Steemburgo et al. 2009).
During pregnancy and early postnatal life, an individual can be programmed for nutritional thrift to adapt and survive in an environment scarce in resources. In 2008, Heijmans et al. (2008) studied the degree of methylation at five DNA sites in the insulin-like growth factor 2 (IGF2) gene on the population exposed to the Dutch famine of 1944–1945. Prenatal exposure to the Dutch famine was associated with the risk of obesity. Kucharski et al. (2008), provided evidence that epigenetic information could be differentially altered by the nutritional input in honeybee (Apis mellifera). Moreover, they found that epigenetic modifications could provoke profound changes in developmental fates with implications in reproductive and behavioral status. When bee larvae are fed royal jelly, it turns off the expression of DNA Dnmt3 and other genes are expressed, turning some of them into a queen, whereas bee larvae that are not fed royal jelly, Dnmt3 remains active and the larval development produces the worker variety of bees.
In the past few years, a new window of research opened concerning the possible influence of the diversity of human gut microbiota (microorganisms that populate the adult intestines) in obesity (Million et al. 2013). Some studies found that the gut microbiota of nonobese individuals is more diverse than that of obese individuals (Chen et al. 2014). Furthermore, several studies found an increase or decrease of different phyla between diet modification (e.g. reduced-carbohydrate, fiber, high-fat, etc.) and weight gain/loss (Million et al. 2013; Chen et al. 2014). However, the mechanisms underlying gut microbiota affecting obesity in humans remain largely unknown. Thus, new studies and discoveries about how the gut microbiota affects the host metabolism could provide a more comprehensive understanding on how it affects obesity.
A dietary intervention could be helpful in prevention as a potential instrument that can complement dietary advice. However, there are some limitations concerning nutrigenetics applications, such as, the lack of studies analysing the evidence of common polymorphisms, polymorphisms differ on ethnic background, and the high cost of the genetic analyses. More generally, compliance with nutrient-based recommendations, such as reducing intake of fat and sugar, has been very poor.
Physical activity-genotype interactions in obesity
Physical activity is another important component involved in the heterogeneous set of factors influencing obesity. Regular exercise is one of the most promising behavioral candidates for preventing and reducing weight gain, with other health and psychological benefits (Richardson et al. 2014). The most extensively studied example of a gene interaction with physical activity in obesity is the FTO locus; evidencing that physical activity attenuates the association of FTO variants with obesity-related traits (Andreasen et al. 2008; Rampersaud et al. 2008; Lee et al. 2010; Ruiz et al. 2010; Xie et al. 2009; Richardson et al. 2014). A meta-analysis by Kilpeläinen et al. (2011a) observed that the estimated effect of the A risk allele of rs9939609 increased the odds ratio of obesity by 1.23-fold/allele, but this effect is attenuated by 27 % in physical active adults (p interaction = 0.001). Similarly, the meta-analysis conducted by Ahmad et al. (2013) showed a statistical significant GRS and physical activity interaction effect in obesity (p interaction = 0.015). In this analysis of 111,421 adults of European ancestry, data support the interaction effect between physical activity and a genetic risk score (combining 12 polymorphisms) in obesity disposition (p interaction = 0.015). So, higher levels of physical activity may attenuate the influence of obesity susceptibility polymorphisms on BMI during adolescence.
However, several studies have provided evidence that the propensity to be physically active has also a strong genetic component in both animals and humans (Herring et al. 2014). In humans, physical activity has been shown to aggregate in families; more active parents have more active children relative to inactive parents (Moore et al. 1991). The physical activity heritability ranges from 9 % in Mexican-American families, to almost 80 % in European twins (Herring et al. 2014). Common polymorphisms in the MC4R gene were also found associated with self-reported physical inactivity in French-Canadian families and Mexican-Americans (Loos et al. 2005; Cai et al. 2006). Another variant, the Gln223Arg polymorphism located in LEPR gene, was found to be associated with lower 24 h energy expenditure and physical activity levels in individual homozygotes for the Arg223 allele compared to Gln homozygotes in Pima Indians population (Stefan et al. 2002). Summarizing, it appears that some variation in our DNA could contribute to the variation in the physical activity level. Thus, new studies and the identification of new loci implicated in this interaction could better enlighten and help to understand the causes contributing to the development of obesity.
Drug-genotype interaction
The use of drugs as a treatment option for obesity could be indicated for individuals with a BMI > 30 kg/m2 with existing co-morbidities such as diabetes, dyslipidemia or hypertension (O'Connor and Swick 2013). In the last decade, with the discovery that some drugs were affected by hereditarily variation, the concept of “pharmacogenetics” emerged (Cascorbi et al. 2013). This new field focuses on the study of polymorphisms within one or more candidate genes for associations with pharmacologic phenotypes. So, common polymorphisms may alter the response to pharmacotherapy affecting drug metabolism, drug transport or drug targets (Cascorbi et al. 2013; O'Connor and Swick 2013). Relating to obesity, at least 35 loci were validated as being associated with BMI and the advent of GWAS and next generation sequencing will likely lead to the identification of additional genetic biomarkers. Until now, only three obesity-related drugs were approved for continuous use in the United States of America (USA): orlistat (Xenical®, Alli®), lorcaserin HCL (Belviq®), and phentermine and topiramate extended release (Qsymia™) (O'Connor and Swick 2013). Orlistat is a drug that alters metabolism by inhibiting the gastro-intestinal absorption of triglycerides (Ravussin and Bouchard 2000). Lorcaserin HCL and Phentermine are drugs that act centrally as an appetite suppressant (Cosentino et al. 2013; O'Connor and Swick 2013). At the end of 2014, the US Food and Drug Administration (FDA) approved a new drug the Saxenda® (Novo Nordisk), an agonist of a glucagon-like peptide 1 (GLP-1).
In the future, it may be possible to determine which sub-populations will respond optimally to particular doses of drugs, allowing more effective personalized pharmacologic intervention. To achieve this end, it would be ideal if pharmacogenetic studies could identify differences in drug response and tolerability, and investigate gene regulation, epigenetic modifications, and DNA–protein interactions that could explain individual differences in responses to drugs beyond genetic variation. Ultimately, it will also be necessary for clinical trials to evaluate pharmacologic interventions that are guided by genetic tests.
Surgical intervention
For patients with morbid obesity (BMI ≥ 40 kg/m2) and overweight (BMI ≥ 35 kg/m2), suffering also of obesity-related comorbidities, which failed diet, exercise, and drug therapy, a surgical intervention could be the only option for the resolution of obesity problem. This approach could be a definitive way in many situations to reduce loss weight. However, some patients present a significant weight gain after chirurgical intervention. There are several guidelines and procedures that surgeons/gastroenterologists need to follow (Elrazek et al. 2014). In this section, we will not detail about surgical intervention strategies, which have been reviewed elsewhere (Vu et al. 2013), but about a possible relation between some genetic variants and the success to maintain weight loss after surgical intervention.
Throughout this review, we discuss the importance that genetics factors play in the etiology of obesity, and for that genetics should also be a factor taking into account in patients undergoing surgery. Effectively, there is a high degree of inter-subject variability for surgical outcomes (Sevilla and Hubal 2014). Generally, patients submitted to a surgical intervention have a durable weight loss (Hatoum et al. 2013). However, despite its effectiveness not all people lose the same amount of weight or obtain the same clinical benefits after the intervention. Some studies emerged associating specific polymorphisms with the response to bariatric surgery. Still et al. (2011) used a summative allele risk score to found the presence of an association between several polymorphisms (including the FTO, and MC4R genes) with postoperative weight loss. De Luis et al. (2014) investigated the role of rs6923761 polymorphism at GLP-1R gene on outcomes after biliopancreatic diversion. They found that homozygous individuals for the rs6923761 G allele showed higher weight loss 12 and 18 months after bariatric surgery than individuals A allele carriers. In another study, Hatoum et al. (2013) found that a 15q26.1 locus was significantly associated with weight loss after Roux-en-Y gastric bypass surgery. Using the same surgery intervention, in 2013, a GWAS identified 17 polymorphisms whose frequencies were significantly different between two phenotypic extremes of weight loss at 2 years after surgery (Rinella et al. 2013).
There are some evidences for the use of genomics to identify response to surgical procedures (Hatoum et al. 2011, 2013). Thus, the identification of genetic contributors could be useful to select those individuals who will obtain a better benefit from a weight-loss surgery. However, these results need to be interpreted with some caution due to the few number of replication studies.
Final remarks
Obesity is a complex phenotype resulting from the interaction of several internal and external factors. Although most scientists and clinicians now acknowledge that genes contribute to obesity, nowadays relatively little is known regarding the specific loci involved and the mechanism by which they lead to the expression of obesity. Also, it is very intriguing how natural selection favored the spread of genes that increase the risk for an obese phenotype and how this predisposition to obesity evolved. Trying to answer these questions several different evolutionary hypotheses have been proposed, however, none of them have consensus and for now our knowledge about the evolution of obesity underlying mechanisms remains incomplete. In the monogenic forms of obesity the causing gene is clearly identified, however, for common obesity the question is far more complex. In the last years with an improvement in technology, the number of loci associated with common obesity increased. Nevertheless, some problems were found in the replication of several studies. Moreover, differences between adults and children and across ethnicities also account for a more complex scenario in the genetic causes for common obesity. Of all currently identified loci, the FTO locus has the largest effect on obesity-susceptibility. Even though, the FTO locus explains only 0.31 % of the inter-individual variation in BMI. Thus, we might speculate that obesity-susceptibility is more likely to be caused by low-frequency variants that have not yet been identified. New genotyping approaches focusing on low frequency and rare variants, studying populations of diverse ancestry will become even more important as such variants are more likely to be ancestry-specific. Moreover, like for many other complex human traits, environmental factors play a major role in the etiology of obesity. In this aspect, recent advances in nutrigenetics or nutrigenomics, relating the diet with the susceptibility for several health conditions, could also account for the obese phenotype, and should be a very important topic of research in the future. Moreover, evidences suggest that interactions between genetic and environmental factors may contribute to epigenetic changes, which have recently emerged as important players in the obesity phenotype. Molecular mechanisms like microRNAs and gene expression are also being implicated in obesity, and together with other epigenetic factors constitute an important area of research in obesity. Altogether, these factors and their possible interactions contribute to the complexity of causes underlying the development of obesity. Although remarkable advances in our understanding of the factors that give rise to obesity have occurred, further research on the etiology, genetics and other factors affecting of obesity is still needed and probably will be a hot topic in obesity research in the next years. In fact, the understanding of the mechanisms accounting for obesity is an important and essential step to develop new therapeutic approaches.
References
Ahmad S, Rukh G, Varga TV, Ali A, Kurbasic A, Shungin D, Ericson U, Koivula RW, Chu AY, Rose LM, Ganna A, Qi Q, Stančáková A, Sandholt CH, Elks CE, Curhan G, Jensen MK, Tamimi RM, Allin KH, Jørgensen T, Brage S, Langenberg C, Aadahl M, Grarup N, Linneberg A, Paré G, InterAct Consortium, DIRECT Consortium, Magnusson PK, Pedersen NL, Boehnke M, Hamsten A, Mohlke KL, Pasquale LT, Pedersen O, Scott RA, Ridker PM, Ingelsson E, Laakso M, Hansen T, Qi L, Wareham NJ, Chasman DI, Hallmans G, Hu FB, Renström F, Orho-Melander M, Franks PW (2013) Gene × physical activity interactions in obesity: combined analysis of 111,421 individuals of European ancestry. PLoS Genet 9(7):e1003607
Albuquerque D, Nóbrega C, Manco L (2013a) Association of FTO polymorphisms with obesity and obesity-related outcomes in Portuguese children. PLoS One 8(1):e54370
Albuquerque D, Nóbrega C, Manco L (2013b) The lactase persistence-13910C>T polymorphism shows indication of association with abdominal obesity among Portuguese children. Acta Paediatr 102(4):e153–e157
Albuquerque D, Nóbrega C, Rodríguez-López R, Manco L (2014) Association study of common polymorphisms in MSRA, TFAP2B, MC4R, NRXN3, PPARGC1A, TMEM18, SEC16B, HOXB5, and OLFM4 genes with obesity-related traits among Portuguese children. J Hum Genet 59:307–313
Almén MS, Jacobsson JA, Moschonis G, Benedict C, Chrousos GP, Fredriksson R, Schiöth HB (2012) Genome wide analysis reveals association of a FTO gene variant with epigenetic changes. Genomics 99(3):132–137
Almon R, Alvarez-Leon EE, Serra-Majem L (2012) Association of the European lactase persistence variant (LCT-13910 C>T polymorphism) with obesity in the Canary Islands. PLoS ONE 7:e43978
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355
Andreasen CH, Stender-Petersen KL, Mogensen MS, Torekov SS, Wegner L, Andersen G, Nielsen AL, Albrechtsen A, Borch-Johnsen K, Rasmussen SS, Clausen JO, Sandbaek A, Lauritzen T, Hansen L, Jørgensen T, Pedersen O, Hansen T (2008) Low physical activity accentuates the effect of the FTO rs9939609 polymorphism on body fat accumulation. Diabetes 57(1):95–101
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bastien M, Poirier P, Lemieux I, Després JP (2014) Overview of epidemiology and contribution of obesity to cardiovascular disease. Prog Cardiovasc Dis 56(4):369–381
Bates SH, Myers MG (2003) The role of leptin receptor signaling in feeding and neuroendocrine function. Trends Endocrinol Metab 14:447–452
Beckers S, Mertens I, Peeters A, Van Gaal L, Van Hul W (2006) Screening for melanocortin-4 receptor mutations in a cohort of Belgian morbidly obese adults and children. Int J Obes (Lond) 30:221–225
Bell AC, Kremer PJ, Magarey AM, Swinburn BA (2005) Contribution of ‘noncore’ foods and beverages to the energy intake and weight status of Australian children. Eur J Clin Nutr 59:639–645
Belsky DW, Moffitt TE, Sugden K, Williams B, Houts R, McCarthy J, Caspi A (2013) Development and evaluation of a genetic risk score for obesity. Biodemogr Soc Biol 59(1):85–100
Bergmann C (2012) Ciliopathies. Eur J Pediatr 171(9):1285–1300
Berndt SI, Gustafsson S, Mägi R, Ganna A, Wheeler E, Feitosa MF, Justice AE, Monda KL, Croteau-Chonka DC, Day FR, Esko T, Fall T, Ferreira T, Gentilini D, Jackson AU, Luan J, Randall JC, Vedantam S, Willer CJ, Winkler TW, Wood AR, Workalemahu T, Hu YJ, Lee SH, Liang L, Lin DY, Min JL, Neale BM, Thorleifsson G, Yang J, Albrecht E, Amin N, Bragg-Gresham JL, Cadby G, den Heijer M, Eklund N, Fischer K, Goel A, Hottenga JJ, Huffman JE, Jarick I, Johansson Å, Johnson T, Kanoni S, Kleber ME, König IR, Kristiansson K, Kutalik Z, Lamina C, Lecoeur C, Li G, Mangino M, McArdle WL, Medina-Gomez C, Müller-Nurasyid M, Ngwa JS, Nolte IM, Paternoster L, Pechlivanis S, Perola M, Peters MJ, Preuss M, Rose LM, Shi J, Shungin D, Smith AV, Strawbridge RJ, Surakka I, Teumer A, Trip MD, Tyrer J, Van Vliet-Ostaptchouk JV, Vandenput L, Waite LL, Zhao JH, Absher D, Asselbergs FW, Atalay M, Attwood AP, Balmforth AJ, Basart H, Beilby J, Bonnycastle LL, Brambilla P, Bruinenberg M, Campbell H, Chasman DI, Chines PS, Collins FS, Connell JM, Cookson WO, de Faire U, de Vegt F, Dei M, Dimitriou M, Edkins S, Estrada K, Evans DM, Farrall M, Ferrario MM, Ferrières J, Franke L, Frau F, Gejman PV, Grallert H, Grönberg H, Gudnason V, Hall AS, Hall P, Hartikainen AL, Hayward C, Heard-Costa NL, Heath AC, Hebebrand J, Homuth G, Hu FB, Hunt SE, Hyppönen E, Iribarren C, Jacobs KB, Jansson JO, Jula A, Kähönen M, Kathiresan S, Kee F, Khaw KT, Kivimäki M, Koenig W, Kraja AT, Kumari M, Kuulasmaa K, Kuusisto J, Laitinen JH, Lakka TA, Langenberg C, Launer LJ, Lind L, Lindström J, Liu J, Liuzzi A, Lokki ML, Lorentzon M, Madden PA, Magnusson PK, Manunta P, Marek D, März W, Mateo Leach I, McKnight B, Medland SE, Mihailov E, Milani L, Montgomery GW, Mooser V, Mühleisen TW, Munroe PB, Musk AW, Narisu N, Navis G, Nicholson G, Nohr EA, Ong KK, Oostra BA, Palmer CN, Palotie A, Peden JF, Pedersen N, Peters A, Polasek O, Pouta A, Pramstaller PP, Prokopenko I, Pütter C, Radhakrishnan A, Raitakari O, Rendon A, Rivadeneira F, Rudan I, Saaristo TE, Sambrook JG, Sanders AR, Sanna S, Saramies J, Schipf S, Schreiber S, Schunkert H, Shin SY, Signorini S, Sinisalo J, Skrobek B, Soranzo N, Stančáková A, Stark K, Stephens JC, Stirrups K, Stolk RP, Stumvoll M, Swift AJ, Theodoraki EV, Thorand B, Tregouet DA, Tremoli E, Van der Klauw MM, van Meurs JB, Vermeulen SH, Viikari J, Virtamo J, Vitart V, Waeber G, Wang Z, Widén E, Wild SH, Willemsen G, Winkelmann BR, Witteman JC, Wolffenbuttel BH, Wong A, Wright AF, Zillikens MC, Amouyel P, Boehm BO, Boerwinkle E, Boomsma DI, Caulfield MJ, Chanock SJ, Cupples LA, Cusi D, Dedoussis GV, Erdmann J, Eriksson JG, Franks PW, Froguel P, Gieger C, Gyllensten U, Hamsten A, Harris TB, Hengstenberg C, Hicks AA, Hingorani A, Hinney A, Hofman A, Hovingh KG, Hveem K, Illig T, Jarvelin MR, Jöckel KH, Keinanen-Kiukaanniemi SM, Kiemeney LA, Kuh D, Laakso M, Lehtimäki T, Levinson DF, Martin NG, Metspalu A, Morris AD, Nieminen MS, Njølstad I, Ohlsson C, Oldehinkel AJ, Ouwehand WH, Palmer LJ, Penninx B, Power C, Province MA, Psaty BM, Qi L, Rauramaa R, Ridker PM, Ripatti S, Salomaa V, Samani NJ, Snieder H, Sørensen TI, Spector TD, Stefansson K, Tönjes A, Tuomilehto J, Uitterlinden AG, Uusitupa M, van der Harst P, Vollenweider P, Wallaschofski H, Wareham NJ, Watkins H, Wichmann HE, Wilson JF, Abecasis GR, Assimes TL, Barroso I, Boehnke M, Borecki IB, Deloukas P, Fox CS, Frayling T, Groop LC, Haritunian T, Heid IM, Hunter D, Kaplan RC, Karpe F, Moffatt MF, Mohlke KL, O’Connell JR, Pawitan Y, Schadt EE, Schlessinger D, Steinthorsdottir V, Strachan DP, Thorsteinsdottir U, van Duijn CM, Visscher PM, Di Blasio AM, Hirschhorn JN, Lindgren CM, Morris AP, Meyre D, Scherag A, McCarthy MI, Speliotes EK, North KE, Loos RJ, Ingelsson E (2013) Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nat Genet 45(5):501–512
Bersaglieri T, Sabeti PC, Patterson N, Vanderploeg T, Schaffner SF, Drake JA, Rhodes M, Reich DE, Hirschhorn JN (2004) Genetic signatures of strong recent positive selection at the lactase gene. Am J Hum Genet 74(6):1111–1120
Bird A (2002) DNA methylation patterns and epigenetic memory. Genes Dev 16(1):6–21
Blüher M (2014) Mechanisms in endocrinology: are metabolically healthy obese individuals really healthy? Eur J Endocrinol 171(6):R209–R219
Blüher M, Mantzoros CS (2015) From leptin to other adipokines in health and disease: facts and expectations at the beginning of the 21st century. Metabolism 64(1):131–145
Bonnefond A, Philippe J, Durand E, Muller J, Saeed S, Arslan M, Martínez R, De Graeve F, Dhennin V, Rabearivelo I, Polak M, Cavé H, Castaño L, Vaxillaire M, Mandel JL, Sand O, Froguel P (2014) Highly sensitive diagnosis of 43 monogenic forms of diabetes or obesity through one-step PCR-based enrichment in combination with next-generation sequencing. Diabetes Care 37(2):460–467
Bradfield JP, Taal HR, Timpson NJ, Scherag A, Lecoeur C, Warrington NM, Hypponen E, Holst C, Valcarcel B, Thiering E, Salem RM, Schumacher FR, Cousminer DL, Sleiman PM, Zhao J, Berkowitz RI, Vimaleswaran KS, Jarick I, Pennell CE, Evans DM, St Pourcain B, Berry DJ, Mook-Kanamori DO, Hofman A, Rivadeneira F, Uitterlinden AG, van Duijn CM, van der Valk RJ, de Jongste JC, Postma DS, Boomsma DI, Gauderman WJ, Hassanein MT, Lindgren CM, Mägi R, Boreham CA, Neville CE, Moreno LA, Elliott P, Pouta A, Hartikainen AL, Li M, Raitakari O, Lehtimäki T, Eriksson JG, Palotie A, Dallongeville J, Das S, Deloukas P, McMahon G, Ring SM, Kemp JP, Buxton JL, Blakemore AI, Bustamante M, Guxens M, Hirschhorn JN, Gillman MW, Kreiner-Møller E, Bisgaard H, Gilliland FD, Heinrich J, Wheeler E, Barroso I, O’Rahilly S, Meirhaeghe A, Sørensen TI, Power C, Palmer LJ, Hinney A, Widen E, Farooqi IS, McCarthy MI, Froguel P, Meyre D, Hebebrand J, Jarvelin MR, Jaddoe VW, Smith GD, Hakonarson H, Grant SF, Early Growth Genetics Consortium (2012) A genome-wide association meta-analysis identifies new childhood obesity loci. Nat Genet 44:526–531
Brunaud L, Alberto JM, Ayav A, Gérard P, Namour F, Antunes L, Braun M, Bronowicki JP, Bresler L, Guéant JL (2003) Vitamin B12 is a strong determinant of low methionine synthase activity and DNA hypomethylation in gastrectomized rats. Digestion 68:133–140
Cai G, Cole SA, Butte N, Bacino C, Diego V, Tan K, Göring HH, O’Rahilly S, Farooqi IS, Comuzzie AG (2006) A quantitative trait locus on chromosome 18q for physical activity and dietary intake in Hispanic children. Obesity (Silver Spring) 14:1596–1604
Campión F, Milagro F, Martínez J (2009) Etiology and pathophysiology: individuality and epigenetics in obesity. Obes Rev 10:383–392
Cannon B, Nedergaard J (2004) Brown adipose tissue: function and physiological significance. Physiol Rev 84:277–359
Cascorbi I, Bruhn O, Werk AN (2013) Challenges in pharmacogenetics. Eur J Clin Pharmacol 69(Suppl 1):S17–S23
Chambers JC, Elliott P, Zabaneh D, Zhang W, Li Y, Froguel P, Balding D, Scott J, Kooner JS (2008) Common genetic variation near MC4R is associated with waist circumference and insulin resistance. Nat Genet 40:716–718
Chang YC, Liu PH, Lee WJ, Chang TJ, Jiang YD, Li HY, Kuo SS, Lee KC, Chuang LM (2008) Common variation in the fat mass and obesity-associated (FTO) gene confers risk of obesity and modulates BMI in the Chinese population. Diabetes 57:2245–2252
Chen J, He X, Huang J (2014) Diet effects in gut microbiome and obesity. J Food Sci 79(4):R442–R451
Choi SW, Friso S, Ghandour H, Bagley PJ, Selhub J, Mason JB (2004) Vitamin B-12 deficiency induces anomalies of base substitution and methylation in the DNA of rat colonic epithelium. J Nutr 134:750–755
Church C, Lee S, Bagg EA, McTaggart JS, Deacon R, Gerken T, Lee A, Moir L, Mecinović J, Quwailid MM, Schofield CJ, Ashcroft FM, Cox RD (2009) A mouse model for the metabolic effects of the human fat mass and obesity associated FTO gene. PLoS Genet 5:e1000599
Church C, Moir L, McMurray F, Girard C, Banks GT, Teboul L, Wells S, Brüning JC, Nolan PM, Ashcroft FM, Cox RD (2010) Overexpression of Fto leads to increased food intake and results in obesity. Nat Genet 42:1086–1092
Clément K, Ferré P (2003) Genetics and the pathophysiology of obesity. Pediatr Res 53:721–725
Corella D, Arregui M, Coltell O, Portolés O, Guillem-Sáiz P, Carrasco P, Sorlí JV, Ortega-Azorín C, González JI, Ordovás JM (2011) Association of the LCT-13910C > T polymorphism with obesity and its modulation by dairy products in a Mediterranean population. Obesity (Silver Spring) 19:1707–1714
Cosentino G, Conrad AO, Uwaifo GI (2013) Phentermine and topiramate for the management of obesity: a review. Drug Des Devel Ther 7:267–278
Costello JF, Plass C (2001) Methylation matters. J Med Genet 38:285–303
Danielzik S, Langnäse K, Mast M, Spethmann C, Müller MJ (2002) Impact of parental BMI on the manifestation of overweight 5–7 year old children. Eur J Nutr 41(3):132–138
de Luis DA, Pacheco D, Aller R, Izaola O (2014) Role of the rs6923761 gene variant in glucagon-like peptide 1 receptor gene on cardiovascular risk factors and weight loss after Biliopancreatic diversion surgery. Ann Nutr Metab 65(4):259–263
Deliard S, Panossian S, Mentch FD, Kim CE, Hou C, Frackelton EC, Bradfield JP, Glessner JT, Zhang H, Wang K, Sleiman PM, Chiavacci RM, Berkowitz RI, Hakonarson H, Zhao J, Grant SF (2013) The missense variation landscape of FTO, MC4R, and TMEM18 in obese children of African ancestry. Obesity (Silver Spring) 21(1):159–163
Dina C, Meyre D, Gallina S, Durand E, Körner A, Jacobson P, Carlsson LM, Kiess W, Vatin V, Lecoeur C, Delplanque J, Vaillant E, Pattou F, Ruiz J, Weill J, Levy-Marchal C, Horber F, Potoczna N, Hercberg S, Le Stunff C, Bougnères P, Kovacs P, Marre M, Balkau B, Cauchi S, Chèvre JC, Froguel P (2007) Variation in FTO contributes to childhood obesity and severe adult obesity. Nat Genet 39(6):724–726
Dlouha D, Pitha J, Lanska V, Hubacek JA (2012) Association between FTO 1st intron tagging variant and telomere length in middle aged females. 3PMFs study. Clin Chim Acta 413(15–16):1222–1225
Dubern B, Clément K (2012) Leptin and leptin receptor-related monogenic obesity. Biochimie 94:2111–2115
Elrazek AE, Elbanna AE, Bilasy SE (2014) Medical management of patients after bariatric surgery: principles and guidelines. World J Gastrointest Surg 6(11):220–228
Enattah NS, Sahi T, Savilahti E, Terwilliger JD, Peltonen L, Jãverlã I (2002) Identification of a variant associated with adult-type hypolactasia. Nat Genet 30:233–237
Fang H, Li Y, Du S, Hu X, Zhang Q, Liu A, Ma G (2010) Variant rs9939609 in the FTO gene is associated with body mass index among Chinese children. BMC Med Genet 11:e136
Farooqi S, O’Rahilly S (2005) Monogenic obesity in humans. Annu Rev Med 56:443–458
Farooqi IS, Keogh JM, Yeo GS, Lank EJ, Cheetham T, O’Rahilly S (2003) Clinical spectrum of obesity and mutations in the melanocortin 4 receptor gene. N Engl J Med 348:1085–1095
Fawcett KA, Barroso I (2010) The genetics of obesity: FTO leads the way. Trends Genet 26(6):266–274
Feinleib M, Garrison RJ, Fabsitz R, Christian JC, Hrubec Z, Borhani NO, Kannel WB, Rosenman R, Schwartz JT, Wagner JO (1977) The NHLBI twin study of cardiovascular disease risk factors: methodology and summary of results. Am J Epidemiol 106(4):284–285
Feldmann HM, Golozoubova V, Cannon B, Nedergaard J (2009) UCP1 ablation induces obesity and abolishes diet-induced thermogenesis in mice exempt from thermal stress by living at hermoneutrality. Cell Metab 9:203–209
Finucane MM, Stevens GA, Cowan MJ, Danaei G, Lin JK, Paciorek CJ, Singh GM, Gutierrez HR, Lu Y, Bahalim AN, Farzadfar F, Riley LM, Ezzati M (2011) National, regional, and global trends in body-mass index since 1980: systematic analysis of health examination surveys and epidemiological studies with 960 country-years and 9.1 million participants. Lancet 377(9765):557–567
Fischer J, Koch L, Emmerling C, Vierkotten J, Peters T, Brüning JC, Rüther U (2009) Inactivation of the Fto gene protects from obesity. Nature 458:894–898
Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM, Perry JR, Elliott KS, Lango H, Rayner NW, Shields B, Harries LW, Barrett JC, Ellard S, Groves CJ, Knight B, Patch AM, Ness AR, Ebrahim S, Lawlor DA, Ring SM, Ben-Shlomo Y, Jarvelin MR, Sovio U, Bennett AJ, Melzer D, Ferrucci L, Loos RJ, Barroso I, Wareham NJ, Karpe F, Owen KR, Cardon LR, Walker M, Hitman GA, Palmer CN, Doney AS, Morris AD, Smith GD, Hattersley AT, McCarthy MI (2007) A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316(5826):889–894
Gantz I, Miwa H, Konda Y, Shimoto Y, Tashiro T, Watson SJ, DelValle J, Yamada T (1993) Molecular cloning, expression, and gene localization of a fourth melanocortin receptor. J Biol Chem 268:15174–15179
Gemma C, Sookoian S, Alvariñas J, García SI, Quintana L, Kanevsky D, González CD, Pirola CJ (2009) Maternal pregestational BMI is associated with methylation of the PPARGC1A promoter in newborns. Obesity (Silver Spring) 17:1032–1039
Gerken T, Girard CA, Tung YC, Webby CJ, Saudek V, Hewitson KS, Yeo GS, McDonough MA, Cunliffe S, McNeill LA, Galvanovskis J, Rorsman P, Robins P, Prieur X, Coll AP, Ma M, Jovanovic Z, Farooqi IS, Sedgwick B, Barroso I, Lindahl T, Ponting CP, Ashcroft FM, O’Rahilly S, Schofield CJ (2007) The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318:1469–1472
Godfrey KM, Sheppard A, Gluckman PD, Lillycrop KA, Burdge GC, McLean C, Rodford J, Slater-Jefferies JL, Garratt E, Crozier SR, Emerald BS, Gale CR, Inskip HM, Cooper C, Hanson MA (2011) Epigenetic gene promoter methylation at birth is associated with child’s later adiposity. Diabetes 60:1528–1534
Goel K, Lopez-Jimenez F, De Schutter A, Coutinho T, Lavie CJ (2014) Obesity paradox in different populations: evidence and controversies. Future Cardiol 10(1):81–91
González JR, González-Carpio M, Hernández-Sáez R, Serrano Vargas V, Torres Hidalgo G, Rubio-Rodrigo M, García-Nogales A, Núñez Estévez M, Luengo Pérez LM, Rodríguez-López R (2012) FTO risk haplotype among early onset and severe obesity cases in a population of western Spain. Obesity (Silver Spring) 20(4):909–915
González JR, Estévez MN, Giralt PS, Cáceres A, Pérez LM, González-Carpio M, Ballester F, Sunyer J, Rodríguez-López R (2013) Genetic risk profiles for a childhood with severely overweight. Pediatr Obes. doi:10.1111/j.2047-6310.2013.00166.x. [Epub ahead of print]
González-Jiménez E, Aguilar Cordero MJ, Padilla López CA, García García I (2012) Monogenic human obesity: role of the leptin-melanocortin system in the regulation of food intake and body weight in humans. An Sist Sanit Navar 35(2):285–293
Goyal A, Nimmakayala KR, Zonszein J (2014) Is there a paradox in obesity? Cardiol Rev 22(4):163–170
Grant SF, Li M, Bradfield JP, Kim CE, Annaiah K, Santa E, Glessner JT, Casalunovo T, Frackelton EC, Otieno FG, Shaner JL, Smith RM, Imielinski M, Eckert AW, Chiavacci RM, Berkowitz RI, Hakonarson H (2008) Association analysis of the FTO gene with obesity in children of Caucasian and African ancestry reveals a common tagging SNP. PLoS ONE 3:e1746
Hallman DM, Friedel VC, Eissa MA, Boerwinkle E, Huber JC Jr, Harrist RB, Srinivasan SR, Chen W, Dai S, Labarthe DR, Berenson GS (2012) The association of variants in the FTO gene with longitudinal body mass index profiles in non-Hispanic white children and adolescents. Int J Obes (Lond). 36(1):61–68
Han Z, Niu T, Chang J, Lei X, Zhao M, Wang Q, Cheng W, Wang J, Feng Y, Chai J (2010) Crystal structure of the FTO protein reveals basis for its substrate specificity. Nature 464:1205–1209
Harrold JA, Williams G (2006) Melanocortin-4 receptors, beta-MSH and leptin: key elements in the satiety pathway. Peptides 27:365–371
Haslam DW, James WP (2005) Obesity. Lancet 366:1197–1209
Hassanein MT, Lyon HN, Nguyen TT, Akylbekova EL, Waters K, Lettre G, Tayo B, Forrester T, Sarpong DF, Stram DO, Butler JL, Wilks R, Liu J, Le Marchand L, Kolonel LN, Zhu X, Henderson B, Cooper R, McKenzie C, Taylor HA Jr, Haiman CA, Hirschhorn JN (2010) Fine mapping of the association with obesity at the FTO locus in African derived populations. Hum Mol Genet 19:2907–2916
Hatoum IJ, Greenawalt DM, Cotsapas C, Reitman ML, Daly MJ, Kaplan LM (2011) Heritability of the weight loss response to gastric bypass surgery. J Clin Endocrinol Metab 96:E1630–E1633
Hatoum IJ, Greenawalt DM, Cotsapas C, Daly MJ, Reitman ML, Kaplan LM (2013) Weight loss after gastric bypass is associated with a variant at 15q26.1. Am J Hum Genet 92(5):827–834
Heard-Costa NL, Zillikens MC, Monda KL, Johansson A, Harris TB, Fu M, Haritunians T, Feitosa MF, Aspelund T, Eiriksdottir G, Garcia M, Launer LJ, Smith AV, Mitchell BD, McArdle PF, Shuldiner AR, Bielinski SJ, Boerwinkle E, Brancati F, Demerath EW, Pankow JS, Arnold AM, Chen YD, Glazer NL, McKnight B, Psaty BM, Rotter JI, Amin N, Campbell H, Gyllensten U, Pattaro C, Pramstaller PP, Rudan I, Struchalin M, Vitart V, Gao X, Kraja A, Province MA, Zhang Q, Atwood LD, Dupuis J, Hirschhorn JN, Jaquish CE, O’Donnell CJ, Vasan RS, White CC, Aulchenko YS, Estrada K, Hofman A, Rivadeneira F, Uitterlinden AG, Witteman JC, Oostra BA, Kaplan RC, Gudnason V, O’Connell JR, Borecki IB, van Duijn CM, Cupples LA, Fox CS, North KE (2009) NRXN3 is a novel locus for waist circumference: a genome-wide association study from the CHARGE Consortium. PLoS Genet 5:e1000539
Heid IM, Jackson AU, Randall JC, Winkler TW, Qi L, Steinthorsdottir V, Thorleifsson G, Zillikens MC, Speliotes EK, Mägi R, Workalemahu T, White CC, Bouatia-Naji N, Harris TB, Berndt SI, Ingelsson E, Willer CJ, Weedon MN, Luan J, Vedantam S, Esko T, Kilpeläinen TO, Kutalik Z, Li S, Monda KL, Dixon AL, Holmes CC, Kaplan LM, Liang L, Min JL, Moffatt MF, Molony C, Nicholson G, Schadt EE, Zondervan KT, Feitosa MF, Ferreira T, Lango Allen H, Weyant RJ, Wheeler E, Wood AR, MAGIC, Estrada K, Goddard ME, Lettre G, Mangino M, Nyholt DR, Purcell S, Smith AV, Visscher PM, Yang J, McCarroll SA, Nemesh J, Voight BF, Absher D, Amin N, Aspelund T, Coin L, Glazer NL, Hayward C, Heard-Costa NL, Hottenga JJ, Johansson A, Johnson T, Kaakinen M, Kapur K, Ketkar S, Knowles JW, Kraft P, Kraja AT, Lamina C, Leitzmann MF, McKnight B, Morris AP, Ong KK, Perry JR, Peters MJ, Polasek O, Prokopenko I, Rayner NW, Ripatti S, Rivadeneira F, Robertson NR, Sanna S, Sovio U, Surakka I, Teumer A, van Wingerden S, Vitart V, Zhao JH, Cavalcanti-Proença C, Chines PS, Fisher E, Kulzer JR, Lecoeur C, Narisu N, Sandholt C, Scott LJ, Silander K, Stark K, Tammesoo ML, Teslovich TM, Timpson NJ, Watanabe RM, Welch R, Chasman DI, Cooper MN, Jansson JO, Kettunen J, Lawrence RW, Pellikka N, Perola M, Vandenput L, Alavere H, Almgren P, Atwood LD, Bennett AJ, Biffar R, Bonnycastle LL, Bornstein SR, Buchanan TA, Campbell H, Day IN, Dei M, Dörr M, Elliott P, Erdos MR, Eriksson JG, Freimer NB, Fu M, Gaget S, Geus EJ, Gjesing AP, Grallert H, Grässler J, Groves CJ, Guiducci C, Hartikainen AL, Hassanali N, Havulinna AS, Herzig KH, Hicks AA, Hui J, Igl W, Jousilahti P, Jula A, Kajantie E, Kinnunen L, Kolcic I, Koskinen S, Kovacs P, Kroemer HK, Krzelj V, Kuusisto J, Kvaloy K, Laitinen J, Lantieri O, Lathrop GM, Lokki ML, Luben RN, Ludwig B, McArdle WL, McCarthy A, Morken MA, Nelis M, Neville MJ, Paré G, Parker AN, Peden JF, Pichler I, Pietiläinen KH, Platou CG, Pouta A, Ridderstråle M, Samani NJ, Saramies J, Sinisalo J, Smit JH, Strawbridge RJ, Stringham HM, Swift AJ, Teder-Laving M, Thomson B, Usala G, van Meurs JB, van Ommen GJ, Vatin V, Volpato CB, Wallaschofski H, Walters GB, Widen E, Wild SH, Willemsen G, Witte DR, Zgaga L, Zitting P, Beilby JP, James AL, Kähönen M, Lehtimäki T, Nieminen MS, Ohlsson C, Palmer LJ, Raitakari O, Ridker PM, Stumvoll M, Tönjes A, Viikari J, Balkau B, Ben-Shlomo Y, Bergman RN, Boeing H, Smith GD, Ebrahim S, Froguel P, Hansen T, Hengstenberg C, Hveem K, Isomaa B, Jørgensen T, Karpe F, Khaw KT, Laakso M, Lawlor DA, Marre M, Meitinger T, Metspalu A, Midthjell K, Pedersen O, Salomaa V, Schwarz PE, Tuomi T, Tuomilehto J, Valle TT, Wareham NJ, Arnold AM, Beckmann JS, Bergmann S, Boerwinkle E, Boomsma DI, Caulfield MJ, Collins FS, Eiriksdottir G, Gudnason V, Gyllensten U, Hamsten A, Hattersley AT, Hofman A, Hu FB, Illig T, Iribarren C, Jarvelin MR, Kao WH, Kaprio J, Launer LJ, Munroe PB, Oostra B, Penninx BW, Pramstaller PP, Psaty BM, Quertermous T, Rissanen A, Rudan I, Shuldiner AR, Soranzo N, Spector TD, Syvanen AC, Uda M, Uitterlinden A, Völzke H, Vollenweider P, Wilson JF, Witteman JC, Wright AF, Abecasis GR, Boehnke M, Borecki IB, Deloukas P, Frayling TM, Groop LC, Haritunians T, Hunter DJ, Kaplan RC, North KE, O’Connell JR, Peltonen L, Schlessinger D, Strachan DP, Hirschhorn JN, Assimes TL, Wichmann HE, Thorsteinsdottir U, van Duijn CM, Stefansson K, Cupples LA, Loos RJ, Barroso I, McCarthy MI, Fox CS, Mohlke KL, Lindgren CM (2010) Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nat Genet 42(11):949–960
Heijmans BT, Tobi EW, Stein AD, Putter H, Blauw GJ, Susser ES, Slagboom PE, Lumey LH (2008) Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci 105(44):17046–17049
Heneghan HM, Miller N, McAnena OJ, O’Brien T, Kerin MJ (2011) Differential miRNA expression in omental adipose tissue and in the circulation of obese patients identifies novel metabolic biomarkers. J Clin Endocrinol Metab 96:e846–e850
Hennig BJ, Fulford AJ, Sirugo G, Rayco-Solon P, Hattersley AT, Frayling TM, Prentice AM (2009) FTO gene variation and measures of body mass in an African population. BMC Med Genet 10:21
Herbert A, Gerry NP, McQueen MB, Heid IM, Pfeufer A, Illig T, Wichmann HE, Meitinger T, Hunter D, Hu FB, Colditz G, Hinney A, Hebebrand J, Koberwitz K, Zhu X, Cooper R, Ardlie K, Lyon H, Hirschhorn JN, Laird NM, Lenburg ME, Lange C, Christman MF (2006) A common genetic variant is associated with adult and childhood obesity. Science 312:279–283
Herring MP, Sailors MH, Bray MS (2014) Genetic factors in exercise adoption, adherence and obesity. Obes Rev 15(1):29–39
Hinney A, Vogel CI, Hebebrand J (2010) From monogenic to polygenic obesity: recent advances. Eur Child Adolesc Psychiatry 19(3):297–310
Hinney A, Volckmar AL, Knoll N (2013) Melanocortin-4 receptor in energy homeostasis and obesity pathogenesis. Prog Mol Biol Transl Sci 114:147–191
Horne BD, Anderson JL, Carlquist JF, Muhlestein JB, Renlund DG, Bair TL, Pearson RR, Camp NJ (2005) Generating genetic risk scores from intermediate phenotypes for use in association studies of clinically significant endpoints. Ann Hum Genet 69(Pt 2):176–186
Hotta K, Nakata Y, Matsuo T, Kamohara S, Kotani K, Komatsu R, Itoh N, Mineo I, Wada J, Masuzaki H, Yoneda M, Nakajima A, Miyazaki S, Tokunaga K, Kawamoto M, Funahashi T, Hamaguchi K, Yamada K, Hanafusa T, Oikawa S, Yoshimatsu H, Nakao K, Sakata T, Matsuzawa Y, Tanaka K, Kamatani N, Nakamura Y (2008) Variations in the FTO gene are associated with severe obesity in the Japanese. J Hum Genet 53(6):546–553
Huszar D, Lynch CA, Fairchild-Huntress V, Dunmore JH, Fang Q, Berkemeier LR, Gu W, Kesterson RA, Boston BA, Cone RD, Smith FJ, Campfield LA, Burn P, Lee F (1997) Targeted disruption of the melanocortin-4 receptor results in obesity in mice. Cell 88:131–141
Ichihara S, Yamada Y (2008) Genetic factors for human obesity. Cell Mol Life Sci 65(7–8):1086–1098
Jacobsson J, Schiöth H, Fredriksson R (2012) The impact of intronic single nucleotide polymorphisms and ethnic diversity for studies on the obesity gene FTO. Obes Rev 13(12):1096–1109
Jia JJ, Tian YB, Cao ZH, Tao LL, Zhang X, Gao SZ, Ge CR, Lin QY, Jois M (2010) The polymorphisms of UCP1 genes associated with fat metabolism, obesity and diabetes. Mol Biol Rep 37:1513–1522
Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, Yi C, Lindahl T, Pan T, Yang YG, He C (2011) N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 7:885–887
Kajimoto K, Naraba H, Iwai N (2006) MicroRNA and 3T3-L1 pre-adipocyte differentiation. RNA 12:1626–1632
Karolina DS, Tavintharan S, Armugam A, Sepramaniam S, Pek SLT, Wong MTK, Lim SC, Sum CF, Jeyaseelan K (2012) Circulating miRNA profiles in patients with metabolic syndrome. J Clin Endocrinol Metab 97:e2271–e2276
Keller P, Gburcik V, Petrovic N, Gallagher IJ, Nedergaard J, Cannon B, Timmons JA (2011) Gene-chip studies of adipogenesis-regulated microRNAs in mouse primary adipocytes and human obesity. BMC Endocr Disord 11:7
Kettunen J, Silander K, Saarela O, Amin N, Müller M, Timpson N, Surakka I, Ripatti S, Laitinen J, Hartikainen AL, Pouta A, Lahermo P, Anttila V, Männistö S, Jula A, Virtamo J, Salomaa V, Lehtimäki T, Raitakari O, Gieger C, Wichmann EH, Van Duijn CM, Smith GD, McCarthy MI, Järvelin MR, Perola M, Peltonen L (2010) European lactase persistence genotype shows evidence of association with increase in body mass index. Hum Mol Genet 19:1129–1136
Khosla S, Dean W, Brown D, Reik W, Feil R (2001) Culture of preimplantation mouse embryos affects fetal development and the expression of imprinted genes. Biol Reprod 64:918–926
Kilpeläinen TO, Qi L, Brage S, Sharp SJ, Sonestedt E, Demerath E, Ahmad T, Mora S, Kaakinen M, Sandholt CH, Holzapfel C, Autenrieth CS, Hyppönen E, Cauchi S, He M, Kutalik Z, Kumari M, Stančáková A, Meidtner K, Balkau B, Tan JT, Mangino M, Timpson NJ, Song Y, Zillikens MC, Jablonski KA, Garcia ME, Johansson S, Bragg-Gresham JL, Wu Y, van Vliet-Ostaptchouk JV, Onland-Moret NC, Zimmermann E, Rivera NV, Tanaka T, Stringham HM, Silbernagel G, Kanoni S, Feitosa MF, Snitker S, Ruiz JR, Metter J, Larrad MT, Atalay M, Hakanen M, Amin N, Cavalcanti-Proença C, Grøntved A, Hallmans G, Jansson JO, Kuusisto J, Kähönen M, Lutsey PL, Nolan JJ, Palla L, Pedersen O, Pérusse L, Renström F, Scott RA, Shungin D, Sovio U, Tammelin TH, Rönnemaa T, Lakka TA, Uusitupa M, Rios MS, Ferrucci L, Bouchard C, Meirhaeghe A, Fu M, Walker M, Borecki IB, Dedoussis GV, Fritsche A, Ohlsson C, Boehnke M, Bandinelli S, van Duijn CM, Ebrahim S, Lawlor DA, Gudnason V, Harris TB, Sørensen TI, Mohlke KL, Hofman A, Uitterlinden AG, Tuomilehto J, Lehtimäki T, Raitakari O, Isomaa B, Njølstad PR, Florez JC, Liu S, Ness A, Spector TD, Tai ES, Froguel P, Boeing H, Laakso M, Marmot M, Bergmann S, Power C, Khaw KT, Chasman D, Ridker P, Hansen T, Monda KL, Illig T, Järvelin MR, Wareham NJ, Hu FB, Groop LC, Orho-Melander M, Ekelund U, Franks PW, Loos RJ (2011a) Physical activity attenuates the influence of FTO variants on obesity risk: a meta-analysis of 218,166 adults and 19,268 children. PLoS Med 8(11):e1001116
Kilpeläinen TO, Zillikens MC, Stančákova A, Finucane FM, Ried JS, Langenberg C, Zhang W, Beckmann JS, Luan J, Vandenput L, Styrkarsdottir U, Zhou Y, Smith AV, Zhao JH, Amin N, Vedantam S, Shin SY, Haritunians T, Fu M, Feitosa MF, Kumari M, Halldorsson BV, Tikkanen E, Mangino M, Hayward C, Song C, Arnold AM, Aulchenko YS, Oostra BA, Campbell H, Cupples LA, Davis KE, Döring A, Eiriksdottir G, Estrada K, Fernández-Real JM, Garcia M, Gieger C, Glazer NL, Guiducci C, Hofman A, Humphries SE, Isomaa B, Jacobs LC, Jula A, Karasik D, Karlsson MK, Khaw KT, Kim LJ, Kivimäki M, Klopp N, Kühnel B, Kuusisto J, Liu Y, Ljunggren O, Lorentzon M, Luben RN, McKnight B, Mellström D, Mitchell BD, Mooser V, Moreno JM, Männistö S, O’Connell JR, Pascoe L, Peltonen L, Peral B, Perola M, Psaty BM, Salomaa V, Savage DB, Semple RK, Skaric-Juric T, Sigurdsson G, Song KS, Spector TD, Syvänen AC, Talmud PJ, Thorleifsson G, Thorsteinsdottir U, Uitterlinden AG, van Duijn CM, Vidal-Puig A, Wild SH, Wright AF, Clegg DJ, Schadt E, Wilson JF, Rudan I, Ripatti S, Borecki IB, Shuldiner AR, Ingelsson E, Jansson JO, Kaplan RC, Gudnason V, Harris TB, Groop L, Kiel DP, Rivadeneira F, Walker M, Barroso I, Vollenweider P, Waeber G, Chambers JC, Kooner JS, Soranzo N, Hirschhorn JN, Stefansson K, Wichmann HE, Ohlsson C, O’Rahilly S, Wareham NJ, Speliotes EK, Fox CS, Laakso M, Loos RJ (2011b) Genetic variation near IRS1 associates with reduced adiposity and an impaired metabolic profile. Nat Genet 43:753–760
Kim JK, Samaranayake M, Pradhan S (2009) Epigenetic mechanisms in mammals. Cell Mol Life Sci 66(4):596–612
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Klöting N, Berthold S, Kovacs P, Schön MR, Fasshauer M, Ruschke K, Stumvoll M, Blüher M (2009) MicroRNA expression in human omental and subcutaneous adipose tissue. PLoS ONE 4:e4699
Knowler WC, Pettitt DJ, Saad MF, Bennett PH (1990) Diabetes mellitus in the Pima Indians: incidence, risk factors and pathogenesis. Diabetes Metab Rev 6(1):1–27
Kucharski R, Maleszka J, Foret S, Maleszka R (2008) Nutritional control of reproductive status in honeybees via DNA methylation. Science 319:1827–1830
Kuzawa C, Sweet E (2009) Epigenetics and the embodiment of race: developmental origins of US racial disparities in cardiovascular health. Am J Hum Biol 21:2–15
Lavie CJ, De Schutter A, Milani RV (2015) Healthy obese versus unhealthy lean: the obesity paradox. Nat Rev Endocrinol 11(1):55–62
Lee IM, Djoussé L, Sesso HD, Wang L, Buring JE (2010) Physical activity and weight gain prevention. JAMA 303(12):1173–1179
León-Mimila P, Villamil-Ramírez H, Villalobos-Comparán M, Villarreal-Molina T, Romero-Hidalgo S, López-Contreras B, Gutiérrez-Vidal R, Vega-Badillo J, Jacobo-Albavera L, Posadas-Romeros C, Canizalez-Román A, Río-Navarro BD, Campos-Pérez F, Acuña-Alonzo V, Aguilar-Salinas C, Canizales-Quinteros S (2013) Contribution of common genetic variants to obesity and obesity-related traits in Mexican children and adults. PLoS ONE 8(8):e70640
Li S, Zhao JH, Luan J, Luben RN, Rodwell SA, Khaw KT, Ong KK, Wareham NJ, Loos RJ (2010) Cumulative effects and predictive value of common obesity-susceptibility variants identified by genome-wide association studies. Am J Clin Nutr 91(1):184–190
Lindgren CM, Heid IM, Randall JC, Lamina C, Steinthorsdottir V, Qi L, Speliotes EK, Thorleifsson G, Willer CJ, Herrera BM, Jackson AU, Lim N, Scheet P, Soranzo N, Amin N, Aulchenko YS, Chambers JC, Drong A, Luan J, Lyon HN, Rivadeneira F, Sanna S, Timpson NJ, Zillikens MC, Zhao JH, Almgren P, Bandinelli S, Bennett AJ, Bergman RN, Bonnycastle LL, Bumpstead SJ, Chanock SJ, Cherkas L, Chines P, Coin L, Cooper C, Crawford G, Doering A, Dominiczak A, Doney AS, Ebrahim S, Elliott P, Erdos MR, Estrada K, Ferrucci L, Fischer G, Forouhi NG, Gieger C, Grallert H, Groves CJ, Grundy S, Guiducci C, Hadley D, Hamsten A, Havulinna AS, Hofman A, Holle R, Holloway JW, Illig T, Isomaa B, Jacobs LC, Jameson K, Jousilahti P, Karpe F, Kuusisto J, Laitinen J, Lathrop GM, Lawlor DA, Mangino M, McArdle WL, Meitinger T, Morken MA, Morris AP, Munroe P, Narisu N, Nordström A, Nordström P, Oostra BA, Palmer CN, Payne F, Peden JF, Prokopenko I, Renström F, Ruokonen A, Salomaa V, Sandhu MS, Scott LJ, Scuteri A, Silander K, Song K, Yuan X, Stringham HM, Swift AJ, Tuomi T, Uda M, Vollenweider P, Waeber G, Wallace C, Walters GB, Weedon MN, Wellcome Trust Case Control Consortium, Witteman JC, Zhang C, Zhang W, Caulfield MJ, Collins FS, Davey Smith G, Day IN, Franks PW, Hattersley AT, Hu FB, Jarvelin MR, Kong A, Kooner JS, Laakso M, Lakatta E, Mooser V, Morris AD, Peltonen L, Samani NJ, Spector TD, Strachan DP, Tanaka T, Tuomilehto J, Uitterlinden AG, van Duijn CM, Wareham NJ, Watkins H, Procardis Consortia, Waterworth DM, Boehnke M, Deloukas P, Groop L, Hunter DJ, Thorsteinsdottir U, Schlessinger D, Wichmann HE, Frayling TM, Abecasis GR, Hirschhorn JN, Loos RJ, Stefansson K, Mohlke KL, Barroso I, McCarthy MI, Giant Consortium (2009) Genome-wide association scan meta-analysis identifies three loci influencing adiposity and fat distribution. PLoS Genet 5:e1000508
Llewellyn CH, Trzaskowski M, Plomin R, Wardle J (2013) Finding the missing heritability in pediatric obesity: the contribution of genome-wide complex trait analysis. Int J Obes (Lond) 37(11):1506–1509
Loos RJ, Lindgren CM, Li S, Wheeler E, Zhao JH, Prokopenko I, Inouye M, Freathy RM, Attwood AP, Beckmann JS, Berndt SI, Prostate, Lung, Colorectal, and Ovarian (PLCO) Cancer Screening Trial, Jacobs KB, Chanock SJ, Hayes RB, Bergmann S, Bennett AJ, Bingham SA, Bochud M, Brown M, Cauchi S, Connell JM, Cooper C, Smith GD, Day I, Dina C, De S, Dermitzakis ET, Doney AS, Elliott KS, Elliott P, Evans DM, Sadaf Farooqi I, Froguel P, Ghori J, Groves CJ, Gwilliam R, Hadley D, Hall AS, Hattersley AT, Hebebrand J, KORA, Heid IM, Lamina C, Gieger C, Illig T, Meitinger T, Wichmann HE, Herrera B, Hinney A, Hunt SE, Jarvelin MR, Johnson T, Jolley JD, Karpe F, Keniry A, Khaw KT, Luben RN, Mangino M, Marchini J, McArdle WL, McGinnis R, Meyre D, Munroe PB, Morris AD, Ness AR, Neville MJ, Nica AC, Ong KK, O’Rahilly S, Owen KR, Palmer CN, Papadakis K, Potter S, Pouta A, Qi L, Nurses’ Health Study, Randall JC, Rayner NW, Ring SM, Sandhu MS, Scherag A, Sims MA, Song K, Soranzo N, Speliotes EK, Diabetes Genetics Initiative, Syddall HE, Teichmann SA, Timpson NJ, Tobias JH, Uda M, SardiNIA Study, Vogel CI, Wallace C, Waterworth DM, Weedon MN, Welcome Trust Case Control Consortium, Willer CJ, FUSION, Wraight, Yuan X, Zeggini E, Hirschhorn JN, Strachan DP, Ouwehand WH, Caulfield MJ, Samani NJ, Frayling TM, Vollenweider P, Waeber G, Mooser V, Deloukas P, McCarthy MI, Wareham NJ, Barroso I, Jacobs KB, Chanock SJ, Hayes RB, Lamina C, Gieger C, Illig T, Meitinger T, Wichmann HE, Kraft P, Hankinson SE, Hunter DJ, Hu FB, Lyon HN, Voight BF, Ridderstrale M, Groop L, Scheet P, Sanna S, Abecasis GR, Albai G, Nagaraja R, Schlessinger D, Jackson AU, Tuomilehto J, Collins FS, Boehnke M, Mohlke KL (2008) Common variants near MC4R are associated with fat mass, weight and risk of obesity. Nat Genet 40(6):768–775
Loos RJ, Yeo GSH (2014) The bigger picture of FTO—the first GWAS-identified obesity gene. Nat Rev Endocrinol 10(1):51–61
Loos RJ, Rankinen T, Tremblay A, Perusse L, Chagnon Y, Bouchard C (2005) Melanocortin-4 receptor gene and physical activity in the Quebec Family Study. Int J Obes (Lond) 29:420–428
Loss RJ (2012) Genetic determinants of common obesity and their value in prediction. Best Pract Res Clin Endocrinol Metab 26:211–226
Lutz T, Woods S (2012) Overview of animal models of obesity. Curr Protoc Pharmacol 5:61
Mačeková S, Bernasovský I, Gabriková D, Bôžiková A, Bernasovská J, Boroňová I, Behulová R, Svíčková P, Petrejčíková E, Soták M, Sovičová A, Carnogurská J (2012) Association of the FTO rs9939609 polymorphism with obesity in Roma/Gypsy population. Am J Phys Anthropol 147(1):30–34
Magnusson PK, Rasmussen F (2002) Familial resemblance of body mass index and familial risk of high and low body mass index. A study of young men in Sweden. Int J Obes Relat Metab Disord 26:1225–1231
Mamun AA, O´Callaghan M, Callaway L, Najman J, Lawlor DA DA (2009) Associations of gestational weight gain with offspring body mass index and blood pressure at 21 years of age: evidence from a birth cohort study. Circulation 119:1720–1727
Mardis ER (2013) Next-generation sequencing platforms. Annu Rev Anal Chem 6:287–303
Marian AJ (2012) Molecular genetic studies of complex phenotypes. Transl Res 159(2):64–79
Mathes WF, Aylor DL, Miller DR, Churchill GA, Chesler EJ, de Villena FP, Threadgill DW, Pomp D (2011) Architecture of energy balance traits in emerging lines of the collaborative cross. Am J Physiol Endocrinol Metab 300(6):e1124–e1134
McKay J, Mathers J (2011) Diet induced epigenetic changes and their implications for health. Acta Physiol 202:103–118
Milagro FI, Campión J, Garcia-Diaz DF, Goyenechea E, Paternain L, Martinez JÁ (2009) High fat diet-induced obesity modifies the methylation pattern of leptin promoter in rats. J Physiol Biochem 65:1–9
Million M, Lagier JC, Yahav D, Paul M (2013) Gut bacterial microbiota and obesity. Clin Microbiol Infect 19(4):305–313
Milner JA (2004) Molecular targets for bioactive food components. J Nutr 134(9):2492S–2498S
Mitchell JA, Hakonarson H, Rebbeck TR, Grant SF (2013) Obesity-susceptibility loci and the tails of the pediatric BMI distribution. Obesity 21:1256–1260
Monda KL, Chen GK, Taylor KC, Palmer C, Edwards TL, Lange LA, Ng MC, Adeyemo AA, Allison MA, Bielak LF, Chen G, Graff M, Irvin MR, Rhie SK, Li G, Liu Y, Liu Y, Lu Y, Nalls MA, Sun YV, Wojczynski MK, Yanek LR, Aldrich MC, Ademola A, Amos CI, Bandera EV, Bock CH, Britton A, Broeckel U, Cai Q, Caporaso NE, Carlson CS, Carpten J, Casey G, Chen WM, Chen F, Chen YD, Chiang CW, Coetzee GA, Demerath E, Deming-Halverson SL, Driver RW, Dubbert P, Feitosa MF, Feng Y, Freedman BI, Gillanders EM, Gottesman O, Guo X, Haritunians T, Harris T, Harris CC, Hennis AJ, Hernandez DG, McNeill LH, Howard TD, Howard BV, Howard VJ, Johnson KC, Kang SJ, Keating BJ, Kolb S, Kuller LH, Kutlar A, Langefeld CD, Lettre G, Lohman K, Lotay V, Lyon H, Manson JE, Maixner W, Meng YA, Monroe KR, Morhason-Bello I, Murphy AB, Mychaleckyj JC, Nadukuru R, Nathanson KL, Nayak U, N’diaye A, Nemesure B, Wu SY, Leske MC, Neslund-Dudas C, Neuhouser M, Nyante S, Ochs-Balcom H, Ogunniyi A, Ogundiran TO, Ojengbede O, Olopade OI, Palmer JR, Ruiz-Narvaez EA, Palmer ND, Press MF, Rampersaud E, Rasmussen-Torvik LJ, Rodriguez-Gil JL, Salako B, Schadt EE, Schwartz AG, Shriner DA, Siscovick D, Smith SB, Wassertheil-Smoller S, Speliotes EK, Spitz MR, Sucheston L, Taylor H, Tayo BO, Tucker MA, Van Den Berg DJ, Edwards DR, Wang Z, Wiencke JK, Winkler TW, Witte JS, Wrensch M, Wu X, Yang JJ, Levin AM, Young TR, Zakai NA, Cushman M, Zanetti KA, Zhao JH, Zhao W, Zheng Y, Zhou J, Ziegler RG, Zmuda JM, Fernandes JK, Gilkeson GS, Kamen DL, Hunt KJ, Spruill IJ, Ambrosone CB, Ambs S, Arnett DK, Atwood L, Becker DM, Berndt SI, Bernstein L, Blot WJ, Borecki IB, Bottinger EP, Bowden DW, Burke G, Chanock SJ, Cooper RS, Ding J, Duggan D, Evans MK, Fox C, Garvey WT, Bradfield JP, Hakonarson H, Grant SF, Hsing A, Chu L, Hu JJ, Huo D, Ingles SA, John EM, Jordan JM, Kabagambe EK, Kardia SL, Kittles RA, Goodman PJ, Klein EA, Kolonel LN, Le Marchand L, Liu S, McKnight B, Millikan RC, Mosley TH, Padhukasahasram B, Williams LK, Patel SR, Peters U, Pettaway CA, Peyser PA, Psaty BM, Redline S, Rotimi CN, Rybicki BA, Sale MM, Schreiner PJ, Signorello LB, Singleton AB, Stanford JL, Strom SS, Thun MJ, Vitolins M, Zheng W, Moore JH, Williams SM, Ketkar S, Zhu X, Zonderman AB, NABEC Consortium; UKBEC Consortium; BioBank Japan Project; AGEN Consortium, Kooperberg C, Papanicolaou GJ, Henderson BE, Reiner AP, Hirschhorn JN, Loos RJ, North KE, Haiman CA (2013) A meta-analysis identifies new loci associated with body mass index in individuals of African ancestry. Nat Genet 45(6):690–696
Montague CT, Farooqi IS, Whitehead JP, Soos MA, Rau H, Wareham NJ, Sewter CP, Digby JE, Mohammed SN, Hurst JA, Cheetham CH, Earley AR, Barnett AH, Prins JB, O’Rahilly S (1997) Congenital leptin deficiency is associated with severe early-onset obesity in humans. Nature 387:903–908
Moore LL, Lombardi DA, White MJ, Campbell JL, Oliveria SA, Ellison RC (1991) Influence of parents’ physical activity levels on activity levels of young children. J Pediatr 118:215–219
Mountjoy KG, Mortrud MT, Low MJ, Simerly RB, Cone RD (1994) Localization of the melanocortin-4 receptor (MC4-R) in neuroendocrine and autonomic control circuits in the brain. Mol Endocrinol 8:1298–1308
Mutch DM, Clément K (2006) Unravelling the genetics of human obesity. PLoS Genet 2:e188
Myles S, Lea RA, Ohashi J, Chambers GK, Weiss JG, Hardouin E, Engelken J, Macartney-Coxson DP, Eccles DA, Naka I, Kimura R, Inaoka T, Matsumura Y, Stoneking M (2011) Testing the thrifty gene hypothesis: the Gly482Ser variant in PPARGC1A is associated with BMI in tongans. BMC Med Genet 12:10
Nakayama K, Ogawa A, Miyashita H, Tabara Y, Igase M, Kohara K, Miki T, Kagawa Y, Yanagisawa Y, Katashima M, Onda T, Okada K, Fukushima S, Iwamoto S (2013) Positive natural selection of TRIB2, a novel gene that influences visceral fat accumulation East Asia. Hum Genet 132(2):201–217
Neel JV (1962) Diabetes mellitus: a “thrifty” genotype rendered detrimental by “progress”? Bull WHO 77(8):694–703
Ng M, Fleming T, Robinson M, Thomson B, Graetz N, Margono C, Mullany EC, Biryukov S, Abbafati C, Abera SF, Abraham JP, Abu-Rmeileh NM, Achoki T, AlBuhairan FS, Alemu ZA, Alfonso R, Ali MK, Ali R, Guzman NA, Ammar W, Anwari P, Banerjee A, Barquera S, Basu S, Bennett DA, Bhutta Z, Blore J, Cabral N, Nonato IC, Chang JC, Chowdhury R, Courville KJ, Criqui MH, Cundiff DK, Dabhadkar KC, Dandona L, Davis A, Dayama A, Dharmaratne SD, Ding EL, Durrani AM, Esteghamati A, Farzadfar F, Fay DF, Feigin VL, Flaxman A, Forouzanfar MH, Goto A, Green MA, Gupta R, Hafezi-Nejad N, Hankey GJ, Harewood HC, Havmoeller R, Hay S, Hernandez L, Husseini A, Idrisov BT, Ikeda N, Islami F, Jahangir E, Jassal SK, Jee SH, Jeffreys M, Jonas JB, Kabagambe EK, Khalifa SE, Kengne AP, Khader YS, Khang YH, Kim D, Kimokoti RW, Kinge JM, Kokubo Y, Kosen S, Kwan G, Lai T, Leinsalu M, Li Y, Liang X, Liu S, Logroscino G, Lotufo PA, Lu Y, Ma J, Mainoo NK, Mensah GA, Merriman TR, Mokdad AH, Moschandreas J, Naghavi M, Naheed A, Nand D, Narayan KM, Nelson EL, Neuhouser ML, Nisar MI, Ohkubo T, Oti SO, Pedroza A, Prabhakaran D, Roy N, Sampson U, Seo H, Sepanlou SG, Shibuya K, Shiri R, Shiue I, Singh GM, Singh JA, Skirbekk V, Stapelberg NJ, Sturua L, Sykes BL, Tobias M, Tran BX, Trasande L, Toyoshima H, van de Vijver S, Vasankari TJ, Veerman JL, Velasquez-Melendez G, Vlassov VV, Vollset SE, Vos T, Wang C, Wang X, Weiderpass E, Werdecker A, Wright JL, Yang YC, Yatsuya H, Yoon J, Yoon SJ, Zhao Y, Zhou M, Zhu S, Lopez AD, Murray CJ, Gakidou E (2014) Global, regional, and national prevalence of overweight and obesity in children and adults during 1980–2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet 384(9945):766–781
Nielsen R (2005) Molecular signatures of natural selection. Annu Rev Genet 39:197–218
Ntalla I, Panoutsopoulou K, Vlachou P, Southam L, William Rayner N, Zeggini E, Dedoussis GV (2013) Replication of established common genetic variants for adult BMI and childhood obesity in Greek adolescents: The TEENAGE Study. Ann Hum Genet 77(3):268–274
O'Connor A, Swick A (2013) Interface between pharmacotherapy and genes in human obesity. Hum Hered 75:116–126
Ohashi J, Naka I, Kimura R, Natsuhara K, Yamauchi T, Furusawa T, Nakazawa M, Ataka Y, Patarapotikul J, Nuchnoi P, Tokunaga K, Ishida T, Inaoka T, Matsumura Y, Ohtsuka R (2007) FTO polymorphisms in oceanic populations. J Hum Genet 52:1031–1035
O’Rahilly S, Farooqi SI (2008) Human obesity: a heritable neurobehavioral disorder that is highly sensitive to environmental conditions. Diabetes 57(11):2905–2910
Ortega FJ, Moreno-Navarrete JM, Pardo G, Sabater M, Hummel M, Ferrer A, Rodriguez-Hermosa JI, Ruiz B, Ricart W, Peral B, Fernández-Real JM (2010) MiRNA expression profile of human subcutaneous adipose and during adipocyte differentiation. PLoS ONE 5:e9022
Ortega FJ, Mercader JM, Catalán V, Moreno-Navarrete JM, Pueyo N, Sabater M, Gómez-Ambrosi J, Anglada R, Fernández-Formoso JA, Ricart W, Frühbeck G, Fernández-Real JM (2013) Targeting the circulating MicroRNA signature of obesity. Clin Chem 59(5):781–792
Ortega-Azorín C, Sorlí JV, Asensio EM, Coltell O, Martínez-González MÁ, Salas-Salvadó J, Covas MI, Arós F, Lapetra J, Serra-Majem L, Gómez-Gracia E, Fiol M, Sáez-Tormo G, Pintó X, Muñoz MA, Ros E, Ordovás JM, Estruch R, Corella D (2012) Associations of the FTO rs9939609 and the MC4R rs17782313 polymorphisms with type 2 diabetes are modulated by diet, being higher when adherence to the Mediterranean diet pattern is low. Cardiovasc Diabetol 11:137
Peters T, Ausmeier K, Dildrop R, Rüther U (2002) The mouse fused toes (Ft) mutation is the result of a 1.6-Mb deletion including the entire Iroquois B gene cluster. Mamm Genome 13(4):186–188
Peters U, North KE, Sethupathy P, Buyske S, Haessler J, Jiao S, Fesinmeyer MD, Jackson RD, Kuller LH, Rajkovic A, Lim U, Cheng I, Schumacher F, Wilkens L, Li R, Monda K, Ehret G, Nguyen KD, Cooper R, Lewis CE, Leppert M, Irvin MR, Gu CC, Houston D, Buzkova P, Ritchie M, Matise TC, Le Marchand L, Hindorff LA, Crawford DC, Haiman CA, Kooperberg C (2013) A systematic mapping approach of 16q12.2/FTO and BMI in more than 20,000 African Americans narrows in on the underlying functional variation: results from the Population Architecture using Genomics and Epidemiology (PAGE) study. PLoS Genet 9(1):e1003171
Qi Q, Chu AY, Kang JH, Jensen MK, Curhan GC, Pasquale LR, Ridker PM, Hunter DJ, Willett WC, Rimm EB, Chasman DI, Hu FB, Qi L (2012) Sugar-sweetened beverages and genetic risk of obesity. N Eng J Med 367:1387–1396
Rampersaud E, Mitchell BD, Pollin TI, Fu M, Shen H, O'Connell JR, Ducharme JL, Hines S, Sack P, Naglieri R, Shuldiner AR, Snitker S (2008) Physical activity and the association of common FTO gene variants with body mass index and obesity. Arch Intern Med 168(16):1791–1797
Ranadive SA, Vaisse C (2008) Lessons from extreme human obesity: monogenic disorders. Endocrinol Metab Clin North Am 37(3):733–751
Rankinen T, Zuberi A, Chagnon YC, Weisnagel SJ, Argyropoulos G, Walts B, Pérusse L, Bouchard C (2006) The human obesity gene map: the 2005 update. Obesity (Silver Spring) 14:529–644
Rau H, Reaves BJ, O’Rahilly S, Whitehead JP (1999) Truncated human leptin (delta133) associated with extreme obesity undergoes proteasomal degradation after defective intracellular transport. Endocrinology 140:1718–1723
Ravussin E, Bouchard C (2000) Human genomics and obesity: finding appropriate drug targets. Eur J Pharmacol 410(2–3):131–145
Razquin C, Marti A, Martinez JA (2011) Evidences on three relevant obesogenes: MC4R, FTO and PPARγ. Approaches for personalized nutrition. Mol Nutr Food Res 55:136–149
Relton CL, Groom A, St Pourcain B, Sayers AE, Swan DC, Embleton ND, Pearce MS, Ring SM, Northstone K, Tobias JH, Trakalo J, Ness AR, Shaheen SO, Davey Smith G (2012) DNA methylation patterns in cord blood DNA and body size in childhood. PLoS ONE 7:e31821
Rey D, Fernandez-Honrado M, Areces C, Algora M, Abd-El-Fatah-Khalil S, Enriquez-de-Salamanca M, Coca C, Arribas I, Arnaiz-Villena A (2012) Amerindians show no association of PC-1 gene Gln121 allele and obesity: a thrifty gene population genetics. Mol Biol Rep 39(7):7687–7693
Richardson AS, North KE, Graff M, Young KM, Mohlke KL, Lange LA, Lange EM, Harris KM, Gordon-Larsen P (2014) Moderate to vigorous physical activity interactions with genetic variants and body mass index in a large US ethnically diverse cohort. Pediatric Obesity 9(2):e35–e46
Rinella E, Still C, Shao Y, Wood GC, Chu X, Salerno B, Gerhard GS, Ostrer H (2013) Genome-wide association of single-nucleotide polymorphisms with weight loss outcomes after roux-en-Y gastric bypass surgery. J Clin Endocrinol Metab 98(6):E1131–E1136
Rodríguez-López R, González-Carpio M, Serrano MV, Torres G, García de Cáceres MT, Herrera T, Román A, Rubio M, Méndez P, Hernández-Sáez R, Núñez M, Luengo LM (2010) Association of FTO gene polymorphisms and morbid obesity in the population of extremadura. Endocrinol Nutr 57(5):203–209
Romero-Corral A, Montori VM, Somers VK, Korinek J, Thomas RJ, Allison TG, Mookadam F, Lopez-Jimenez F (2006) Association of bodyweight with total mortality and with cardiovascular events in coronary artery disease: a systematic review of cohort studies. Lancet 368(9536):666–678
Rönn T, Volkov P, Davegårdh C, Dayeh T, Hall E, Olsson AH, Nilsson E, Tornberg A, Dekker Nitert M, Eriksson KF, Jones HA, Groop L, Ling C (2013) A six months exercise intervention influences the genome-wide DNA methylation pattern in human adipose tissue. PLoS Genet 9(6):e1003572
Ruiz JR, Labayen I, Ortega FB, Legry V, Moreno LA, Dallongeville J, Martínez-Gómez D, Bokor S, Manios Y, Ciarapica D, Gottrand F, De Henauw S, Molnár D, Sjöström M, Meirhaeghe A; HELENA Study Group (2010) Attenuation of the effect of the FTO rs9939609 polymorphism on total and central body fat by physical activity in adolescents: the HELENA study. Arch Pediatr Adolesc Med 164(4):328–333
Saeed S, Bonnefond A, Manzoor J, Philippe J, Durand E, Arshad M, Sand O, Butt TA, Falchi M, Arslan M, Froguel P (2014) Novel LEPR mutations in obese Pakistani children identified by PCR-based enrichment and next generation sequencing. Obesity (Silver Spring) 22(4):1112–1117
Sällman Almén M, Rask-Andersen M, Jacobsson JA, Ameur A, Kalnina I, Moschonis G, Juhlin S, Bringeland N, Hedberg LA, Ignatovica V, Chrousos GP, Manios Y, Klovins J, Marcus C, Gyllensten U, Fredriksson R, Schiöth HB (2013) Determination of the obesity-associated gene variants within the entire FTO gene by ultra-deep targeted sequencing in obese and lean children. Int J Obes (Lond) 37(3):424–431
Sandholt CH, Hansen T, Pedersen O (2012) Beyond the fourth wave of genome-wide obesity association studies. Nutr Diabetes 2:e37
Scuteri A, Sanna S, Chen WM, Uda M, Albai G, Strait J, Najjar S, Nagaraja R, Orrú M, Usala G, Dei M, Lai S, Maschio A, Busonero F, Mulas A, Ehret GB, Fink AA, Weder AB, Cooper RS, Galan P, Chakravarti A, Schlessinger D, Cao A, Lakatta E, Abecasis GR (2007) Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genet 3(7):1200–1210
Sellayah D, Cagampang FR, Cox RD (2014) On the evolutionary origins of obesity: a new hypothesis. Endocrinology 155(5):1573–1588
Sevilla S, Hubal MJ (2014) Genetic modifiers of obesity and bariatric surgery outcomes. Semin Pediatr Surg 23(1):43–48
Silventoinen K, Rokholm B, Kaprio J, Sørensen TI (2010) The genetic and environmental influences on childhood obesity: a systematic review of twin and adoption studies. Int J Obes 34(1):29–40
Slatkin M (2008) Linkage disequilibrium-understanding the evolutionary past and mapping the medical future. Nat Rev 9(6):477–485
Smemo S, Tena JJ, Kim KH (2014) Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature 507(7492):371–375
Song Y, You NC, Hsu YH, Howard BV, Langer RD, Manson JE, Nathan L, Niu T, Tinker F, Liu LS (2008) FTO polymorphisms are associated with obesity but not diabetes risk in postmenopausal women. Obesity (Silver Spring) 16(11):2472–2480
Soubry A, Schildkraut JM, Murtha A, Wang F, Huang Z, Bernal A, Kurtzberg J, Jirtle RL, Murphy SK, Hoyo C (2013a) Paternal obesity is associated with IGF2 hypomethylation in newborns: results from a Newborn Epigenetics Study (NEST) cohort. BMC Med 11:29
Soubry A, Murphy SK, Wang F, Huang Z, Vidal AC, Fuemmeler BF, Kurtzberg J, Murtha A, Jirtle RL, Schildkraut JM, Hoyo C (2013b) Newborns of obese parents have altered DNA methylation patterns at imprinted genes. Int J Obes (Lond). doi:10.1038/ijo.2013.193 [Epub ahead of print]
Sovio U, Mook-Kanamori DO, Warrington NM, Lawrence R, Briollais L, Palmer CN, Cecil J, Sandling JK, Syvänen AC, Kaakinen M, Beilin LJ, Millwood IY, Bennett AJ, Laitinen J, Pouta A, Molitor J, Davey Smith G, Ben-Shlomo Y, Jaddoe VW, Palmer LJ, Pennell CE, Cole TJ, McCarthy MI, Järvelin MR, Timpson NJ, Early Growth Genetics Consortium (2011) Association between common variation at the FTO locus and changes in body mass index from infancy to late childhood: the complex nature of genetic association through growth and development. PLoS Genet 7(2):e1001307
Speakman JR (2004) Obesity: the integrated roles of environment and genetics. J Nutrition 134:2090S–2105S
Speakman JR (2006) Thrifty genes for obesity and the metabolic syndrome-time to call off the search? Diab Vasc Dis Res 3(1):7–11
Speakman JR (2008) Thrifty genes for obesity, an attractive but flawed idea, and an alternative perspective: the ‘drifty gene’ hypothesis. Int J Obes (Lond) 32(11):1611–1617
Speakman JR (2013) Evolutionary perspectives on the obesity epidemic: adaptive, maladaptive, and neutral viewpoints. Annu Rev Nutr 33:289–317
Speakman JR, Rance KA, Johnstone AM (2008) Polymorphisms of the FTO gene are associated with variation in energy intake, but not energy expenditure. Obesity (Silver Spring) 16:1961–1965
Speliotes EK, Willer CJ, Berndt SI, Monda KL, Thorleifsson G, Jackson AU, Lango Allen H, Lindgren CM, Luan J, Mägi R, Randall JC, Vedantam S, Winkler TW, Qi L, Workalemahu T, Heid IM, Steinthorsdottir V, Stringham HM, Weedon MN, Wheeler E, Wood AR, Ferreira T, Weyant RJ, Segrè AV, Estrada K, Liang L, Nemesh J, Park JH, Gustafsson S, Kilpeläinen TO, Yang J, Bouatia-Naji N, Esko T, Feitosa MF, Kutalik Z, Mangino M, Raychaudhuri S, Scherag A, Smith AV, Welch R, Zhao JH, Aben KK, Absher DM, Amin N, Dixon AL, Fisher E, Glazer NL, Goddard ME, Heard-Costa NL, Hoesel V, Hottenga JJ, Johansson A, Johnson T, Ketkar S, Lamina C, Li S, Moffatt MF, Myers RH, Narisu N, Perry JR, Peters MJ, Preuss M, Ripatti S, Rivadeneira F, Sandholt C, Scott LJ, Timpson NJ, Tyrer JP, van Wingerden S, Watanabe RM, White CC, Wiklund F, Barlassina C, Chasman DI, Cooper MN, Jansson JO, Lawrence RW, Pellikka N, Prokopenko I, Shi J, Thiering E, Alavere H, Alibrandi MT, Almgren P, Arnold AM, Aspelund T, Atwood LD, Balkau B, Balmforth AJ, Bennett AJ, Ben-Shlomo Y, Bergman RN, Bergmann S, Biebermann H, Blakemore AI, Boes T, Bonnycastle LL, Bornstein SR, Brown MJ, Buchanan TA, Busonero F, Campbell H, Cappuccio FP, Cavalcanti-Proença C, Chen YD, Chen CM, Chines PS, Clarke R, Coin L, Connell J, Day IN, den Heijer M, Duan J, Ebrahim S, Elliott P, Elosua R, Eiriksdottir G, Erdos MR, Eriksson JG, Facheris MF, Felix SB, Fischer-Posovszky P, Folsom AR, Friedrich N, Freimer NB, Fu M, Gaget S, Gejman PV, Geus EJ, Gieger C, Gjesing AP, Goel A, Goyette P, Grallert H, Grässler J, Greenawalt DM, Groves CJ, Gudnason V, Guiducci C, Hartikainen AL, Hassanali N, Hall AS, Havulinna AS, Hayward C, Heath AC, Hengstenberg C, Hicks AA, Hinney A, Hofman A, Homuth G, Hui J, Igl W, Iribarren C, Isomaa B, Jacobs KB, Jarick I, Jewell E, John U, Jørgensen T, Jousilahti P, Jula A, Kaakinen M, Kajantie E, Kaplan LM, Kathiresan S, Kettunen J, Kinnunen L, Knowles JW, Kolcic I, König IR, Koskinen S, Kovacs P, Kuusisto J, Kraft P, Kvaløy K, Laitinen J, Lantieri O, Lanzani C, Launer LJ, Lecoeur C, Lehtimäki T, Lettre G, Liu J, Lokki ML, Lorentzon M, Luben RN, Ludwig B, MAGIC, Manunta P, Marek D, Marre M, Martin NG, McArdle WL, McCarthy A, McKnight B, Meitinger T, Melander O, Meyre D, Midthjell K, Montgomery GW, Morken MA, Morris AP, Mulic R, Ngwa JS, Nelis M, Neville MJ, Nyholt DR, O’Donnell CJ, O’Rahilly S, Ong KK, Oostra B, Paré G, Parker AN, Perola M, Pichler I, Pietiläinen KH, Platou CG, Polasek O, Pouta A, Rafelt S, Raitakari O, Rayner NW, Ridderstråle M, Rief W, Ruokonen A, Robertson NR, Rzehak P, Salomaa V, Sanders AR, Sandhu MS, Sanna S, Saramies J, Savolainen MJ, Scherag S, Schipf S, Schreiber S, Schunkert H, Silander K, Sinisalo J, Siscovick DS, Smit JH, Soranzo N, Sovio U, Stephens J, Surakka I, Swift AJ, Tammesoo ML, Tardif JC, Teder-Laving M, Teslovich TM, Thompson JR, Thomson B, Tönjes A, Tuomi T, van Meurs JB, van Ommen GJ, Vatin V, Viikari J, Visvikis-Siest S, Vitart V, Vogel CI, Voight BF, Waite LL, Wallaschofski H, Walters GB, Widen E, Wiegand S, Wild SH, Willemsen G, Witte DR, Witteman JC, Xu J, Zhang Q, Zgaga L, Ziegler A, Zitting P, Beilby JP, Farooqi IS, Hebebrand J, Huikuri HV, James AL, Kähönen M, Levinson DF, Macciardi F, Nieminen MS, Ohlsson C, Palmer LJ, Ridker PM, Stumvoll M, Beckmann JS, Boeing H, Boerwinkle E, Boomsma DI, Caulfield MJ, Chanock SJ, Collins FS, Cupples LA, Smith GD, Erdmann J, Froguel P, Grönberg H, Gyllensten U, Hall P, Hansen T, Harris TB, Hattersley AT, Hayes RB, Heinrich J, Hu FB, Hveem K, Illig T, Jarvelin MR, Kaprio J, Karpe F, Khaw KT, Kiemeney LA, Krude H, Laakso M, Lawlor DA, Metspalu A, Munroe PB, Ouwehand WH, Pedersen O, Penninx BW, Peters A, Pramstaller PP, Quertermous T, Reinehr T, Rissanen A, Rudan I, Samani NJ, Schwarz PE, Shuldiner AR, Spector TD, Tuomilehto J, Uda M, Uitterlinden A, Valle TT, Wabitsch M, Waeber G, Wareham NJ, Watkins H, Procardis Consortium, Wilson JF, Wright AF, Zillikens MC, Chatterjee N, McCarroll SA, Purcell S, Schadt EE, Visscher PM, Assimes TL, Borecki IB, Deloukas P, Fox CS, Groop LC, Haritunians T, Hunter DJ, Kaplan RC, Mohlke KL, O’Connell JR, Peltonen L, Schlessinger D, Strachan DP, van Duijn CM, Wichmann HE, Frayling TM, Thorsteinsdottir U, Abecasis GR, Barroso I, Boehnke M, Stefansson K, North KE, McCarthy MI, Hirschhorn JN, Ingelsson E, Loos RJ (2010) Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat Genet 42(11):937–948
Steemburgo T, Azevedo MJ, Martínez JÁ (2009) Gene-nutrient interaction and its association with obesity and diabetes mellitus. Arq Bras Endocrinol Metabol 53(5):497–508
Steemburgo T, Azevedo MJ, Gross JL, Milagro FI, Campión J, Martínez JÁ (2013) The rs9939609 polymorphism in the FTO gene is associated with fat and fiber intakes in patients with type 2 diabetes. J Nutrigenet Nutrigenomics 6(2):97–106
Stefan N, Vozarova B, Del Parigi A, Ossowski V, Thompson DB, Hanson RL, Ravussin E, Tataranni PA (2002) The Gln223Arg polymorphism of the leptin receptor in Pima Indians: influence on energy expenditure, physical activity and lipid metabolism. Int J Obes Relat Metab Disord 26:1629–1632
Still CD, Wood GC, Chu X, Erdman R, Manney CH, Benotti PN, Petrick AT, Strodel WE, Mirshahi UL, Mirshahi T, Carey DJ, Gerhard GS (2011) High allelic burden of four obesity SNPs is associated with poorer weight loss outcomes following gastric bypass surgery. Obesity (Silver Spring) 19(8):1676–1683
Strobel A, Issad T, Camoin L, Ozata M, Strosberg AD (1998) A leptin missense mutation associated with hypogonadism and morbid obesity. Nat Genet 18:213–215
Stunkard AJ, Foch TT, Hrubec Z (1986a) A twin study of human obesity. JAMA 256(1):51–54
Stunkard AJ, Sorensen TI, Hanis C, Teasdale TW, Chakraborty R, Schull WJ, Schulsinger F (1986b) An adoption study of human obesity. N Engl J Med 314(4):193–198
Swinburn BA, Sacks G, Hall KD, McPherson K, Finegood DT, Moodie ML, Gortmaker SL (2011) The global obesity pandemic: shaped by global drivers and local environments. Lancet 378(9793):804–814
Takanabe R, Ono K, Abe Y, Takaya T, Horie T, Wada H, Kita T, Satoh N, Shimatsu A, Hasegawa K (2008) Up-regulated expression of microRNA-143 in association with obesity in adipose tissue of mice fed high-fat diet. Biochem Biophys Res Commun 376:728–732
Tan LJ, Zhu H, He H, Wu KH, Li J, Chen XD, Zhang JG, Shen H, Tian Q, Krousel-Wood M, Papasian CJ, Bouchard C, Pérusse L, Deng HW (2014) Replication of 6 obesity genes in a meta-analysis of genome-wide association studies from diverse ancestries. PLoS ONE 9(5):e96149
Thorleifsson G, Walters GB, Gudbjartsson DF, Steinthorsdottir V, Sulem P, Helgadottir A, Styrkarsdottir U, Gretarsdottir S, Thorlacius S, Jonsdottir I, Jonsdottir T, Olafsdottir EJ, Olafsdottir GH, Jonsson T, Jonsson F, Borch-Johnsen K, Hansen T, Andersen G, Jorgensen T, Lauritzen T, Aben KK, Verbeek AL, Roeleveld N, Kampman E, Yanek LR, Becker LC, Tryggvadottir L, Rafnar T, Becker DM, Gulcher J, Kiemeney LA, Pedersen O, Kong A, Thorsteinsdottir U, Stefansson K (2009) Genome-wide association yields new sequence variants at seven loci that associate with measures of obesity. Nat Genet 42(11):937–948
Vaisse C, Clément K, Guy-Grand B, Froguel P (1998) A frameshift mutation in human MC4R is associated with a dominant form of obesity. Nat Genet 20:113–114
van den Born E, Omelchenko MV, Bekkelund A, Leihne V, Koonin EV, Dolja VV, Falnes PØ (2008) Viral AlkB proteins repair RNA damage by oxidative demethylation. Nucl Acids Res 36:5451–5461
Vu L, Switzer NJ, De Gara C, Karmali S (2013) Surgical interventions for obesity and metabolic disease. Best Pract Res Clin Endocrinol Metab 27(2):239–246
Wang X, Zhu H, Snieder H, Su S, Munn D, Harshfield G, Maria BL, Dong Y, Treiber F, Gutin B, Shi H (2010) Obesity related methylation changes in DNA of peripheral blood leukocytes. BMC Med 8:87
Waterland RA, Jirtle RL (2003) Transposable elements: targets for early nutritional effects on epigenetic gene regulation. Mol Cell Biol 23:5293–5300
Waters AM, Beales PL (2011) Ciliopathies: an expanding disease spectrum. Pediatr Nephrol 26:1039–1056
Wen W, Cho YS, Zheng W, Dorajoo R, Kato N, Qi L, Chen CH, Delahanty RJ, Okada Y, Tabara Y, Gu D, Zhu D, Haiman CA, Mo Z, Gao YT, Saw SM, Go MJ, Takeuchi F, Chang LC, Kokubo Y, Liang J, Hao M, Le Marchand L, Zhang Y, Hu Y, Wong TY, Long J, Han BG, Kubo M, Yamamoto K, Su MH, Miki T, Henderson BE, Song H, Tan A, He J, Ng DP, Cai Q, Tsunoda T, Tsai FJ, Iwai N, Chen GK, Shi J, Xu J, Sim X, Xiang YB, Maeda S, Ong RT, Li C, Nakamura Y, Aung T, Kamatani N, Liu JJ, Lu W, Yokota M, Seielstad M, Fann CS, Genetic Investigation of ANthropometric Traits (GIANT) Consortium, Wu JY, Lee JY, Hu FB, Tanaka T, Tai ES, Shu XO (2012) Meta-analysis identifies common variants associated with body mass index in East Asians. Nat Genet 44(3):307–311
Whitaker RC, Wright JA, Pepe MS, Seidel KD, Dietz WH (1997) Predicting obesity in young adulthood from childhood and parental obesity. N Engl J Med 337(13):869–873
Widiker S, Karst S, Wagener A, Brockmann GA (2010) High-fat diet leads to a decreased methylation of the Mc4r gene in the obese BFMI and the lean B6 mouse lines. J Appl Genet 51:193–197
Willer CJ, Speliotes EK, Loos RJ, Li S, Lindgren CM, Heid IM, Berndt SI, Elliott AL, Jackson AU, Lamina C, Lettre G, Lim N, Lyon HN, McCarroll SA, Papadakis K, Qi L, Randall JC, Roccasecca RM, Sanna S, Scheet P, Weedon MN, Wheeler E, Zhao JH, Jacobs LC, Prokopenko I, Soranzo N, Tanaka T, Timpson NJ, Almgren P, Bennett A, Bergman RN, Bingham SA, Bonnycastle LL, Brown M, Burtt NP, Chines P, Coin L, Collins FS, Connell JM, Cooper C, Smith GD, Dennison EM, Deodhar P, Elliott P, Erdos MR, Estrada K, Evans DM, Gianniny L, Gieger C, Gillson CJ, Guiducci C, Hackett R, Hadley D, Hall AS, Havulinna AS, Hebebrand J, Hofman A, Isomaa B, Jacobs KB, Johnson T, Jousilahti P, Jovanovic Z, Khaw KT, Kraft P, Kuokkanen M, Kuusisto J, Laitinen J, Lakatta EG, Luan J, Luben RN, Mangino M, McArdle WL, Meitinger T, Mulas A, Munroe PB, Narisu N, Ness AR, Northstone K, O’Rahilly S, Purmann C, Rees MG, Ridderstråle M, Ring SM, Rivadeneira F, Ruokonen A, Sandhu MS, Saramies J, Scott LJ, Scuteri A, Silander K, Sims MA, Song K, Stephens J, Stevens S, Stringham HM, Tung YC, Valle TT, Van Duijn CM, Vimaleswaran KS, Vollenweider P, Waeber G, Wallace C, Watanabe RM, Waterworth DM, Watkins N, Wellcome Trust Case Control Consortium, Witteman JC, Zeggini E, Zhai G, Zillikens MC, Altshuler D, Caulfield MJ, Chanock SJ, Farooqi IS, Ferrucci L, Guralnik JM, Hattersley AT, Hu FB, Jarvelin MR, Laakso M, Mooser V, Ong KK, Ouwehand WH, Salomaa V, Samani NJ, Spector TD, Tuomi T, Tuomilehto J, Uda M, Uitterlinden AG, Wareham NJ, Deloukas P, Frayling TM, Groop LC, Hayes RB, Hunter DJ, Mohlke KL, Peltonen L, Schlessinger D, Strachan DP, Wichmann HE, McCarthy MI, Boehnke M, Barroso I, Abecasis GR, Hirschhorn JN, Genetic Investigation of ANthropometric Traits Consortium (2009) Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat Genet 41:25–34
Williams MD, Mitchell GM (2012) MicroRNAs in insulin resistance and obesity. Exp Diabetes Res 2012:484696
Wolff GL, Kodell RL, Moore SR, Cooney CA (1998) Maternal epigenetics and methyl supplements affect agouti gene expression in Avy/a mice. FASEB J 12:949–957
Wu Q, Saunders RA, Szkudlarek-Mikho M, Serna Ide L, Chin KV (2010) The obesity-associated Fto gene is a transcriptional coactivator. Biochem Biophys Res Commun 401(3):390–395
Xi B, Chandak GR, Shen Y, Wang Q, Zhou D (2012) Association between common polymorphism near the MC4R gene and obesity risk: a systematic review and meta-analysis. PLoS ONE 7(9):e45731
Xia Q, Grant SF (2013) The genetics of human obesity. Ann N Y Acad Sci 1281:178–190
Xie H, Lim B, Lodish HF (2009) MicroRNAs induced during adipogenesis that accelerate fat cell development are downregulated in obesity. Diabetes 58:1050–1057
Xu X, Su S, Barnes VA, De Miguel C, Pollock J, Ownby D, Shi H, Zhu H, Snieder H, Wang X (2013) A genome-wide methylation study on obesity: differential variability and differential methylation. Epigenetics 8(5):522–533
Yang J, Manolio TA, Pasquale LR, Boerwinkle E, Caporaso N, Cunningham JM, de Andrade M, Feenstra B, Feingold E, Hayes MG, Hill WG, Landi MT, Alonso A, Lettre G, Lin P, Ling H, Lowe W, Mathias RA, Melbye M, Pugh E, Cornelis MC, Weir BS, Goddard ME, Visscher PM (2011) Genome partitioning of genetic variation for complex traits using common SNPs. Nat Genet 43(6):519–525
Yeo GS, Farooqi IS, Aminian S, Halsall DJ, Stanhope RG, O’Rahilly S (1998) A frameshift mutation in MC4R associated with dominantly inherited human obesity. Nat Genet 20:111–112
Zavattari P, Loche A, Pilia S, Ibba A, Moi L, Guzzetti C, Casini MR, Loche S (2011) rs9939609 in the FTO gene is associated with obesity but not with several biochemical parameters in Sardinian obese children. Ann Hum Genet 75:648–654
Zhao J, Grant SF (2011) Genetics of childhood obesity. J Obes 2011:e845148
Zhao J, Bradfield JP, Li M, Wang K, Zhang H, Kim CE, Annaiah K, Glessner JT, Thomas K, Garris M, Frackelton EC, Otieno FG, Shaner JL, Smith RM, Chiavacci RM, Berkowitz RI, Hakonarson H, Grant SF (2009) The role of obesity-associated loci identified in genome-wide association studies in the determination of pediatric BMI. Obesity (Silver Spring) 17(12):2254–2257
Zhao J, Bradfield JP, Zhang H, Sleiman PM, Kim CE, Glessner JT, Deliard S, Thomas KA, Frackelton EC, Li M, Chiavacci RM, Berkowitz RI, Hakonarson H, Grant SF (2011) Role of BMI-associated loci identified in GWAS meta-analyses in the context of common childhood obesity in European Americans. Obesity (Silver Spring) 19:2436–2439
Zhao J, Goldberg J, Vaccarino V (2013) Promoter methylation of serotonin transporter gene is associated with obesity measures: a monozygotic twin study. Int J Obes 37(1):140–145
Zhu J, Loos RJ, Lu L, Zong G, Gan W, Ye X, Sun L, Li H, Lin X (2014) Associations of genetic risk score with obesity and related traits and the modifying effect of physical activity in a chinese han population. PLoS ONE 9(3):e91442
Acknowledgments
David Albuquerque as a PhD grant (SFRH/BD/68774/2010) from Fundação para a Ciência e a Tecnologia (FCT).
Conflict of interest
All the authors recognize and disclose to have no conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Graw.
Rights and permissions
About this article
Cite this article
Albuquerque, D., Stice, E., Rodríguez-López, R. et al. Current review of genetics of human obesity: from molecular mechanisms to an evolutionary perspective. Mol Genet Genomics 290, 1191–1221 (2015). https://doi.org/10.1007/s00438-015-1015-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-015-1015-9