Abstract
ATP-citrate lyase (ACL, EC4.1.3.8) catalyzes citrate to oxaloacetate and acetyl-CoA in the cell cytosol, and has important roles in normal plant growth and in the biosynthesis of some secondary metabolites. We identified three ACL genes, CitACLα1, CitACLα2, and CitACLβ1, in the citrus genome database. Both CitACLα1 and CitACLα2 encode putative ACL α subunits with 82.5 % amino acid identity, whereas CitACLβ1 encodes a putative ACL β subunit. Gene structure analysis showed that CitACLα1 and CitACLα2 had 12 exons and 11 introns, and CitACLβ1 had 16 exons and 15 introns. CitACLα1 and CitACLβ1 were predominantly expressed in flower, and CitACLα2 was predominantly expressed in stem and fibrous roots. As fruits ripen, the transcript levels of CitACLα1, CitACLβ1, and/or CitACLα2 in cultivars ‘Niuher’ and ‘Owari’ increased, accompanied by significant decreases in citrate content, while their transcript levels decreased significantly in ‘Egan No. 1’ and ‘Iyokan’, although citrate content also decreased. In ‘HB pummelo’, in which acid content increased as fruit ripened, and in acid-free pummelo, transcript levels of CitACLα2, CitACLβ1, and/or CitACLα1 increased. Moreover, mild drought stress and ABA treatment significantly increased citrate contents in fruits. Transcript levels of the three genes were significantly reduced by mild drought stress, and the transcript level of only CitACLβ1 was significantly reduced by ABA treatment. Taken together, these data indicate that the effects of ACL on citrate use during fruit ripening depends on the cultivar, and the reduction in ACL gene expression may be attributed to citrate increases under mild drought stress or ABA treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
ATP-citrate lyase (ACL, EC4.1.3.8) catalyzes the conversion of citrate to oxaloacetate and acetyl-CoA in the cell cytosol (Rangasamy and Ratledge 2000). This reaction acts as a switch to control the shifts among carbohydrates, organic acids, fatty acids, isoprenoids, flavonoids, and other compounds (Chypre et al. 2012; Fatland et al. 2002). Citrate is the main acid in citrus fruits. Controlling the acid content in fruit is extremely important for improving fruit quality in terms of sourness as well as other parameters, such as color and aroma, because citrate affects these traits via ACL catalysis (Crifò et al. 2012; Fatland et al. 2002; Lo Piero et al. 2014). Although ACL genes and polypeptides have been detected in citrus fruit (Cercós et al. 2006; Katz et al. 2007), a genome-wide identification is lacking, although three citrus genomes have been published (Xu et al. 2012; www.phytozome.net). Understanding the ACL family in the citrus genome is important for their functional identification.
In most citrus cultivars, citric acid begins to accumulate during the second phase of fruit development and continues for a few weeks, peaking when the fruit is about half of its final volume and then declining gradually as the fruit matures (Baldwin 1993). Citric acid content in ripening fruits differs among cultivars. For example, the titratable acid levels in sour lemon, sweet orange, and acidless orange are about 4–5 %, 0.8–1.2 %, and less than 0.1 %, respectively (Albertini et al. 2006). A model for regulating the accumulation and decline of citric acid in citrus fruits was presented by Sadka et al. (2000) and complemented by Cercós et al. (2006). Citric acid is synthesized in the mitochondria as part of the tricarboxylic acid cycle via the condensation of acetyl-CoA with oxaloacetate, catalyzed by citrate synthase (Popova and Pinheiro de Carvalho 1998). Some citric acid is then transported and stored in the vacuole because of the partial block of mitochondrial aconitase activity (Sadka et al. 2000). As fruit matures, citric acid in most cultivars is then removed to the cytosol and degraded by cytosolic aconitase to biosynthesize some amino acids through the γ-aminobutyrate (GABA) pathway (Cercós et al. 2006; Degu et al. 2011; Sadka et al. 2000).
In addition to isomerization by cytosolic aconitase, citrate can be degraded by ACL to oxaloacetate and acetyl-CoA in cell cytosol (Rangasamy and Ratledge 2000). Alterations in the expression or activity of ACL in animals have shown that ACL plays pivotal roles in fetal growth and development, tumor cell growth, survival, and metabolic disorders (Chypre et al. 2012; Zaidi et al. 2012). Because of its instability, however, the study and cloning of ACL only began in 2002 in Arabidopsis (Fatland et al. 2002) and lupin (Langlade et al. 2002), respectively. While animal ACL comprises one polypeptide, plant ACL is a heteromeric enzyme composed of two distinct subunits (ACLα and ACLβ). In lupin, ACLα and ACLβ are each encoded by one gene (Langlade et al. 2002), whereas in Arabidopsis they are encoded by three and two genes, respectively (Fatland et al. 2002). Reverse genetic analysis indicated that even moderately reduced ACL activity could produce a complex bonsai phenotype with miniaturized organs, smaller cells, aberrant plastid morphology, and hyperaccumulation of starch and anthocyanin in vegetative tissue, implying that ACL is required for normal growth and development in higher plants (Fatland et al. 2005). Overexpression of ACL in Arabidopsis activated the wax, cutin, and rubber biosynthetic pathways (Xing et al. 2014). In addition, ACL activity was also suggested to be responsible for the switch between malate and citrate excretion during the development of cluster roots in lupin (Langlade et al. 2002). Recently, genes of sugarcane ACL were reported, and their expressions were induced by abscisic acid (ABA) treatment and/or drought stress (Li et al. 2012).
In citrus, the ACL gene was first discovered in a microarray analysis of gene expression during fruit development and ripening of Citrus clementina (Cercós et al. 2006). Subsequently, its polypeptide was found during a proteomics analysis of citrus fruit (Katz et al. 2007). Recent studies indicated that the ACL gene was induced by low temperature, and citrate might be catabolized by ACL for anthocyanin and flavonoid biosynthesis in blood orange under cold storage (Crifò et al. 2011, 2012; Lo Piero et al. 2014). Although the mitochondrial aconitase–GABA pathway is involved in citrate use by citrus fruits (Cercós et al. 2006; Degu et al. 2011; Katz et al. 2011; Sadka et al. 2000), whether ACL also participates in citrate use during fruit ripening should be further explored. Citrus genome sequences have been published (Xu et al. 2012), making possible the isolation of more ACL genes. The main objective of this study was to identify ACL genes from genome-wide data and evaluate their roles in citrate use by investigating their expression profiles in relation to citric acid content in the developing and ripening fruits of different varieties and in response to drought and ABA injection.
Materials and methods
Plant materials
Ripening fruits of ‘Niuher’ navel orange (Citrus sinensis cv. Niuher), ‘Owari’ Satsuma mandarin (C. unshiu cv. Owari), ‘Egan No. 1’ Ponkan (C. reticulata cv. Egan No. 1), ‘HB pummelo’ (C. grandis Osbeck cv. HB pummelo), acid-free pummelo, and ‘Iyokan’ (C. iyo Hort. ex Tanaka) from the citrus germplasm orchard of Huazhong Agricultural University (Hubei Province, China) were used in the present study. These cultivars are orange (‘Niuher’), loose mandarin (‘Owari’ and ‘Egan No. 1’), pummelo (‘HB pummelo’ and acid-free pummelo), and a hybrid (‘Iyokan’) and have different citrate contents in the ripening fruits. Fruits were sampled at two stages (T1 and T2) during ripening. T1 was around the fruit peel color break stage, while T2 was in the middle ripening stage for ‘Egan No. 1’ and ‘Iyokan’ and at harvest time for the other four cultivars. These stages were, respectively, 134 and 197 days after florescence (DAF) for ‘Niuher’, 129 and 177 DAF for ‘Owari’, 154 and 188 DAF for ‘Egan No. 1’, 128 and 183 DAF for ‘HB pummelo’ and acid-free pummelo, and 126 and 189 DAF for ‘Iyokan’. At each sampling time for each cultivar, three to five healthy fruits were randomly taken from the outer tree crown. Fruit juice sacs (JS) were isolated, mixed, ground in liquid nitrogen, and stored at −80 °C for use.
A seedy orange cultivar, ‘Anliu’ orange (C. sinensis cv. Anliu), was selected for gene organ/tissue-specific expression analysis. Fully opened flowers (FL) and mature leaves (ML) were collected from ‘Anliu’ trees at the florescence stage; fruit JS were collected at 70 DAF; and fibrous roots (FR) and stems (ST) were harvested from seedlings as the height was over 10 cm. The seedlings were propagated as described by Zhou et al. (2014). All samples were frozen immediately with liquid nitrogen and stored at −80 °C.
Drought treatment and ABA injection
‘Owari’ was selected for drought and ABA treatments. Mild drought stress (MDS, no obvious phenotypic change observed in leaves and other tissues) was created using film mulch over soil during the rainy season, as previously described (Jiang et al. 2014). Healthy and uniform fruits of control and drought-treated trees were collected 15 days after film-mulch application. Ten pairs of ‘Owari’ shoots were selected for ABA treatment. Each pair grew on the same fruit-bearing shoot and was of similar size. ABA was injected 150 DAF as previously noted (Liu et al. 2014). Fruits were harvested 3 days after the last injection. Juice sacs of all samples were isolated, frozen immediately in liquid nitrogen, and stored at −80 °C until use.
Citric acid determination
Citric acid was determined by gas–liquid chromatography (Bartolozzi et al. 1997).
Gene isolation and sequence analysis
Three gene sequences (ID: Cs6g01210, Cs7g08950, and Cs9g02230) putatively encoding ACL were directly obtained by screening the sweet orange genome database at Huazhong Agricultural University (Xu et al. 2012). Two other citrus genomes, for sweet orange and clementine, from Phytozome (www.phytozome.net) were used to screen the ACL genes. Total RNA was isolated as described by Liu et al. (2006). Gene-specific primers (Table 1) were designed using Primer 3.0 (Koressaar and Remm 2007) based on the genomic sequences. The open reading frame (ORF), molecular weight, and pI were predicted using EditSeq in the Lasergene program (DNASTAR, Madison, WI, USA). Gene intron/exon structures were analyzed using the Gene Structure Display Server (GSDS, gsds.cbi.pku.edu.cn) (Guo et al. 2007). Cis-regulatory elements in the promoter region of the three ACL genes were scanned with the PLACE program (Higo et al. 1999). Amino acid sequence similarities were calculated using MegAlign within Lasergene. Alignment of multiple sequences was conducted using CLUSTALX (version 1.81) (Thompson et al. 1997).
Quantitative real-time PCR analysis
Total RNA of all samples was isolated as described before (Liu et al. 2006). Five micrograms of high-quality total RNA was treated with DNase I (Fermentas, Vilnius, Lithuania) at 37 °C for 1 h and then used for first-strand cDNA synthesis with the RevertAidTM M-MuLV Kit (Fermentas). Specific primers for citrus ACL and actin genes were designed using Primer 3.0 (Koressaar and Remm 2007) and are listed in Table 1. Additionally, before quantitative real-time PCR (qRT-PCR), the products amplified with each pair of primers in the seven cultivars were sequenced to confirm that no nucleotide differences existed among these cultivars. The qRT-PCR involved three biological replicates, each with two technical replicates. All qRT-PCR reactions were arranged in a 384-well plate. qRT-PCR was performed in a 10-µL reaction volume using the Thunderbird TM SYBR qPCR Mix (TOYOBO, Osaka, Japan) on an ABI Vii7 Real Time System (Applied Biosystems, Foster City, CA, USA) following the manufacturer’s protocol. Reactions began with an initial incubation at 50 °C for 2 min and 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s, and 60 °C for 60 s. The Livak method (Livak and Schmittigen 2001) was employed to calculate gene relative expression levels.
Statistical analysis
The data were evaluated by Duncan’s multiple test or the t test in the ANOVA program of SAS (SAS Institute, Cary, NC, USA). Differences were considered significant at P < 0.05.
Results
Data mining and molecular characterization of ACL genes
A sweet orange genome sequence was recently published (Xu et al. 2012). Genomic DNA analysis revealed at least three ACL sequences, two of which (Cs6g01210 and Cs7g08950) encode the ACL α subunit and one (Cs9g02230) that encodes the ACL β subunit. Each had four or five putative transcripts in the genome database. However, PCR amplification using primers HXM1, HXM3, and HXM9, designed based on the sequence of different region among putative transcripts (Table 1), and subsequent sequencing indicated that only Cs6g01210.2, Cs7g08950.2, and Cs9g02230.2 existed in the citrus fruit flesh. The coding DNA sequences were used to query the other two citrus genomic databases at Phytozome, each of which contained one transcript per gene (orange1.1g014514 and Ciclev10013702, orange1.1g014493 and Ciclev10031633, and orange1.1g007327 and Ciclev10004572, respectively). Multiple alignments revealed strong similarities among: Cs6g01210.2, orange1.1g014514, and Ciclev10013702; Cs7g08950.2, orange1.1g014493, and Ciclev10031633; and Cs9g02230.2, orange1.1g007327, and Ciclev10004572 (Table 2).
The ORFs of Cs6g01210.2, Cs7g08950.2, and Cs9g02230.2 were successfully amplified by reverse transcription-PCR with primers of HXM16, HXM17, and HXM19 (Table 1), respectively. Sequence analysis showed that they had high similarity (>99 %) with their respective sequences in the genomic database. The three ACL genes were named CitACLα1, CitACLα2, and CitACLβ1, respectively. The transcript IDs are listed in Table 2. CitACLα1, CitACLα2, and CitACLβ1 had ORFs that encoded proteins containing 423, 423, and 608 amino acid residues, respectively. CitACLα1 and CitACLα2 had 82.5 % amino acid identity. Theoretically, CitACLα1 encodes a ~46.60 kDa peptide with pI 5.52, CitACLα2 a ~46.63 kDa peptide with pI 5.57, and CitACLβ1 a ~65.95 Da peptide with pI 7.62 (Table 2).
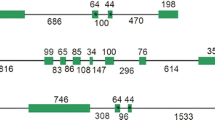
Full-length cDNA and gDNA sequences of ACL genes were downloaded from the C. sinensis genome database (citrus.hzau.edu.cn/orange/), and their exon and intron structures (from start to stop codon) were further analyzed by GSDS (gsds.cbi.pku.edu.cn) (Guo et al. 2007). Twelve exons and eleven introns existed in the genes of both ACL α subunits; the exon sizes were the same, but the intron sizes were different. CitACLβ1 had 16 exons and 15 introns (Fig. 1). In addition, PLACE scanning revealed 124 putative cis-regulatory elements (PcREs) in the promoter regions (+strand) of the three ACL genes, with their numbers and/or distribution varying among the three genes. Only 33 PcREs existed in the promoter regions of the three citrus ACL genes, 13, 22, and 24, in the CitACLα1, CitACLα2, and CitACLβ1 promoter regions, respectively. Thirteen existed only in the CitACLα1 and CitACLα2 promoter regions, five only in CitACLα1 and CitACLβ1, and fourteen only in CitACLα2 and CitACLβ1 (Table S1).
The deduced amino acid sequences of the two citrus ACL α subunit genes and the citrus ACL β subunit gene were aligned with the ACLs of Homo sapiens and Saccharum officinarum (Fig. 2). CitACLα1 and CitACLα2 had 88.7 and 80.9 % amino acid identity, respectively, with S. officinarum ACL α1, while CitACLβ1 had 92.9 % amino acid identity with S. officinarum ACL β1. Additional analysis indicated that CitACLα1 had 81.8, 80.9, and 89.8 % amino acid identity with Arabidopsis thaliana ACL α1, α2, and α3, respectively; CitACLα2 had 85.3, 84.4, and 79.0 % amino acid identity with A. thaliana ACL α1, α2, and α3, respectively; and CitACLβ1 had 92.9 and 93.8 % amino acid identity with A. thaliana ACL β1 and β2, respectively (Table S2). Multiple alignments indicated that putative ATP-grasp conserved and citrate binding amino acid sites (Fig. 2A) were found in citrus ACLα, whereas a histidine phosphorylating amino acid site and potential ATP-binding, phosphorylating, and CoA-binding domains (Fig. 2B) were found in CitACLβ1. In addition, citrus ACL subunits α and β aligned with H. sapiens ACL N- and C-termini, respectively. A region of about 60 amino acids between citrus subunits α and β was absent relative to H. sapiens ACL (Fig. 2C).
Alignment of the deduced amino acid sequence of CitACLα (A) and CitACLβ (B) with the deduced amino acid residues of H. sapiens and S. officinarum ACLs. Identical and similar amino acid residues are shown in black and gray, respectively. The triangles, diamonds, and stars below the residues show the ATP-grasp conserved, citrate binding, and histidine phosphorylating site, respectively; underlines labeled i, ii, and iii indicate potential ATP-binding, phosphorylated, and CoA-binding sites, respectively (Langlade et al. 2002; Li et al. 2012). The accession numbers of other ACLs are: HsACL (AAH06195.1), SoACLα1 (AFN22056.1), and SoACLβ1 (AFO64345.1). C Subunit organization of C. sinensis, S. officinarum, and H. sapiens ACLs
Organ-/tissue-specific expressions of ACL genes
ACL gene expression profiles were examined in different organs/tissues, including fruit JS of 70 DAF, FL, ML, seedling FR, and ST (Fig. 3). The highest transcript level of CitACLα1 was observed in FL (nearly three times that in FR), while the lowest was observed in JS (half that in FR); in the stem, the CitACLα1 transcript level was similar to that in FR (Fig. 3A). Unlike CitACLα1, CitACLα2 transcript levels were similar in ST and FR and in the other three tissues. However, the transcript levels in ST and FR were about twice those in ML, FL, and JR (Fig. 3B). Similar to CitACLα1, the highest transcript level of CitACLβ1 was observed in FL, but it was nearly 30 times that in FR. In ST, the CitACLβ1 transcript was also highly expressed and was about 15 times higher than in FR. In addition, the CitACLβ1 transcript level in ML and JS was similar to that in FR (Fig. 3C).
Relative transcript levels of the three citrus ACL genes in seedling fibrous roots (FR) and stems (ST), mature leaves (ML), flowers (FL), and fruit juice sacs (JS). Data represent the mean ± SE of three biological replicates. Different lowercase letters between any two organs or tissues for each gene indicate significant differences (P < 0.05) by Duncan’s test
Citrate content and ACL gene expression in ripening fruits of different cultivars
Citrate contents and three ACL gene expressions were further analyzed at the T1 and T2 developmental stages of different citrus cultivar fruits (Fig. 4). As shown in Fig. 4A, citrate content at T2 was significantly lower than at T1 in ‘Niuher’, ‘Owari’, ‘Egan No. 1’, and ‘Iyokan’ fruits; in ‘HB pummelo’, citrate content at T2 was significantly higher than at T1, while citrate was undetectable at both stages in acid-free pumello.
Analysis of citric acid content (A) and relative expression levels in ripening fruits of ‘Niuher’ orange (B; NH), ‘Owari’ Satsuma mandarin (C; OW), ‘HB pummelo’ (D; HB), acid-free pummelo (E; AFP), ‘Egan No. 1’ Ponkan (F; EG), and ‘Iyokan’ (G; IY). T1 and T2 refer to the two sampling times. Error bars represent standard errors from three independent replicates. Asterisks on the bars indicates significant differences (P < 0.05) between fruits at the two harvesting times by the t test (LSD)
The expression patterns of the three ACL genes at the two fruit developmental stages varied among the six cultivars. In ‘Niuher’ orange, CitACLα1 and CitACLβ1 transcript levels increased significantly over time, whereas that of CitACLα2 did not change notably (Fig. 4B) as the fruits ripened. In ‘Owari’ mandarin, the three ACL genes showed similar expression patterns, with significant increases from T1 to T2. In CitACLβ1, the three transcript levels at T2 were more than 28 times higher than at T1 (Fig. 4C). Interestingly, the expression patterns of CitACLα1, CitACLα2, and CitACLβ1 in ‘HB pummelo’ (Fig. 4D) and acid-free pummelo (Fig. 4E) were similar to those in ‘Owari’ mandarin (Fig. 4C) as the fruit ripened, except that the CitACLα1 transcript level increased only slightly in ‘HB pummelo’ (Fig. 4D). Unlike in other cultivars, the CitACLα1, CitACLα2, and CitACLβ1 transcript levels decreased significantly from T1 to T2 in fruit JS of ‘Egan No. 1’ and ‘Iyokan’; notably, the CitACLβ1 transcript level was undetectable at T2 in fruit JS of both cultivars (Fig. 4F, G).
Responses of citrate content and ACL gene expression to mild drought and ABA injection
At 15 days after film mulching, the soil water content was 18.5 % beneath the film mulch and 20.1 % in control soil. Moreover, citrate content in the fruit JS was significantly increased by film mulching (Fig. 5A1), but the expressions of the three ACL genes were significantly lower. The CitACLα1 transcript level in the film-mulched fruits was about half of that in control fruits, while those of CitACLα2 and CitACLβ1 were about 70 % and one-third of that in control fruits, respectively (Fig. 5A2).
Similar to the film-mulching treatment, the citrate content in ABA-treated fruits was significantly higher, about 120 % of that in control fruits (Fig. 5B1). However, the two CitACLα transcript levels responded differently to ABA treatment than to film mulching. The CitACLα1 and CitACLα2 transcript levels in ABA-treated fruits were similar to their respective levels in control fruits, although CitACLα2 transcript levels were slightly decreased by ABA injection. Similar to the film-mulching treatment, the CitACLβ1 transcript level was significantly lower in ABA-treated fruits, about half of that in control fruits (Fig. 5B2).
Discussion
As an important cytosolic enzyme in the regulation of fatty acids, cholesterol biosynthesis, and histone acetylation, ACL has been well characterized in animals, including its tissue distribution, subcellular localization, crystal structure, and enzymatic and genetic properties (Chypre et al. 2012). Although the enzymatic activity of ACL in plants was first assayed more than 30 years ago (Nelson and Rinne 1975), it was only molecularly characterized in Arabidopsis (Fatland et al. 2002), lupin (Langlade et al. 2002), and sugarcane (Li et al. 2012) in the past decade. Plant ACL consists of two distinct subunits (ACLα and ACLβ) encoded by separate genes. For example, ACLα and ACLβ are encoded by two different genes in lupin (Langlade et al. 2002) and sugarcane (Li et al. 2012), whereas in Arabidopsis three genes encode ACLα and two encode ACLβ (Fatland et al. 2002). Here, we confirmed that two putative ACL genes encode ACLα and one encodes ACLβ in the current citrus genome databases (Table 2). CitACLα1 and CitACLα2 encode 423 amino acid residues, while CitACLβ1 encodes 608 residues (Table 2), similar to other plant ACLα and ACLβ genes (Fatland et al. 2002; Langlade et al. 2002; Li et al. 2012). Only 17.5 % amino acid difference was found between the two citrus ACLα proteins, more than that between Arabidopsis ACLα1 and ACLα2 but slightly less than that between Arabidopsis ACLα1 or ACLα2 and ACLα3 (Fatland et al. 2002). Moreover, the two ACLα gene structures had the same exon sizes, but different intron sizes (Fig. 1). The citrus ACL genes were highly similar to other plant ACLα and ACLβ genes and had putative functional domains (Fig. 2) as described by Li et al. (2012).
Plant ACL is a heteromeric enzyme composed of two distinct subunits (ACLα and ACLβ) (Fatland et al. 2002). Interestingly, the transcript level of CitACLβ1 was not identical with that in CitACLα1 or CitACLα2 in different tissues or organs (Fig. 3) and in response to different treatments (Fig. 5), as found in other plants (Fatland et al. 2002; Langlade et al. 2002; Li et al. 2012). This variance in transcript profiles of the three ACL genes may be due to differences in cis-regulatory elements of their promoter regions (Table S1). Moreover, different combinations of subunits may give rise to enzymes with different kinetic properties, and the varying gene expressions may reflect different roles of ACL. In addition, a report in sugarcane showed that ACL transcripts in leaf and root were induced by drought stress and ABA treatment (Li et al. 2012). We found that transcript levels of ACL genes in fruit juice sacs were significantly reduced by MDS (Fig. 5A2), and ABA injection significantly reduced only the transcript level of CitACLβ1 (Fig. 5B2). Their divergent responses may vary by organ or other factors and should be further studied.
Citrate content differs among cultivars and usually declines gradually during fruit development (Albertini et al. 2006; Baldwin 1993). This decrease occurred in ‘Niuher’, ‘Owari’, ‘Egan No. 1’, and ‘Iyokan’ in the present study (Fig. 4A). ACL is a cytosolic enzyme that converts mitochondria-derived citrate into oxaloacetate and acetyl-CoA (Rangasamy and Ratledge 2000) and has important roles in the biosynthesis of isoprenoids, flavonoids, and malonate derivatives and in the elongation of fatty acids (Chávez-Cabrera et al. 2010; Chypre et al. 2012; Fatland et al. 2005; Xing et al. 2014; Zaidi et al. 2012). Although citrate breakdown can be directly catalyzed by cytosolic aconitase and ACL in the cytosol, ACL’s role in citrate use during fruit ripening was underestimated, because mRNA levels of a putative ACL gene decreased continuously as clementines ripened (Cercós et al. 2006).
The prevalent hypothesis for citrate use or decrease during fruit ripening in most citrus cultivars is that citrate is released from the vacuole into the cytosol and then metabolized into isocitrate by cytosolic aconitase (Cercós et al. 2006; Sadka et al. 2000; Terol et al. 2010). Citrate loss from the cytosol during fruit ripening is considered to be involved mainly in amino acid biosynthesis via the GABA shunt pathway (Cercós et al. 2006; Degu et al. 2011). However, a recent report showed that cold reduced blood orange acidity, suggesting that low temperature may partially shift citrate to flavonoid biosynthesis, since transcripts of ACL were simultaneously induced (Lo Piero et al. 2014). In fact, ACL was considered to be related to citrate use, since its transcript or protein was detected during fruit development and ripening (Cercós et al. 2006; Katz et al. 2007, 2011), although direct proof is still lacking. In the present study, we identified three putative ACL genes from the citrus genome and simultaneously detected changes in citrate and ACL transcript levels in six citrus cultivars at two stages of fruit ripening (Fig. 4). Transcripts of three ACL genes were detectable in the six cultivars. However, the relationships between their transcript levels and citrate content differed among the cultivars. During fruit ripening of ‘Niuher’, ‘Owari’, ‘Egan No. 1’, and ‘Iyokan’ (Fig. 4A), citrate levels decreased significantly, while transcript levels of CitACLα1, CitACLβ1, and/or CitACLα2 were increased in ‘Niuher’ and ‘Owari’ (Fig. 4B, C) but decreased significantly in ‘Egan No. 1’ and ‘Iyokan’ (Fig. 4F, G), similar to clementines (Cercós et al. 2006). In contrast, although citrate content increased significantly in ‘HB pummelo’ and was very low in acid-free pummelo during fruit ripening, transcript levels of CitACLα2, CitACLβ1, and/or CitACLα1 increased (Fig. 4D, E). These results suggested that the increase in ACL transcript levels may be directly related to the citrate decrease in ‘Niuher’ and ‘Owari’ during normal fruit ripening. In the other four cultivars, changes in ACL transcript levels may be related to the synthesis of carbohydrate and/or secondary metabolites (such as isoprenoids, flavonoids, and malonate derivatives) by using ACL catalyzing products (Chávez-Cabrera et al. 2010; Fatland et al. 2005; Lo Piero et al. 2014). In other words, ACL uses some citrate to biosynthesize some metabolites, but does not determine the citrate content in the fruit juice sacs of these four cultivars.
Clearly, MDS increases fruit citrate contents (Jiang et al. 2014; Navarro et al. 2010). This increase may be due to decreases in cytosolic aconitase and isocitrate dehydrogenase activities (Jiang et al. 2014). As mentioned above, the decline in citrate content during normal ‘Owari’ fruit ripening was accompanied by an increase in the three ACL transcript levels (Fig. 4A, C). Under MDS, the increase in citrate content was accompanied by a decrease in the three ACL transcript levels (Fig. 5A), in addition to a decrease in cytosolic aconitase and isocitrate dehydrogenase activities (Jiang et al. 2014). These results suggested that increased ACL gene expression affects citrate accumulation during fruit ripening in some citrus cultivars, such as ‘Owari’ Satsuma mandarin. Namely, ACL plays an obvious role in determining citrate content in some citrus cultivars’ fruits. On the other hand, the present results confirmed the notion that MDS increased citrus fruit citrate content is related with the decrease of citrate utilization which is directly subjected to the decrease of cytosolic aconitase, isocitrate dehydrogenase activities, and ACL expression levels as well.
ABA treatment can increase fruit citrate content (Fig. 5B), probably by activating ABA-regulated transcription factor (Bastías et al. 2011). Previous results suggested that the increase in fruit titratable acid caused by ABA injection was related to the decrease in citrus glutamate decarboxylase 1 transcript level (Liu et al. 2014). We found that the increase in fruit citrate content after ABA injection was also related to the decrease in CitACLβ1 transcript level, which was reduced significantly by ABA injection (Fig. 5B2) along with a significant increase in citrate content (Fig. 5B1).
In conclusion, there are at least three ACL genes in the current citrus genome: two encode the ACLα subunit and one encodes the ACLβ subunit. The three ACL genes have different transcript characteristics in root, stem, leaf, flower, and juice sacs and in response to MDS and ABA treatment, which may be due to differences in the cis-regulatory elements of their promoter regions. Transcripts of the three genes were detectable during fruit ripening of six citrus cultivars, confirming that ACL takes part in citrate use as the fruits ripen. However, their transcript profiles varied among cultivars, suggesting that ACL roles are cultivar dependent. An increase in ACL transcripts may contribute to the decline in citrate as fruits ripen in ‘Owari’ Satsuma mandarin and ‘Niuher’ orange. Moreover, the increased citrate content under MDS and ABA injection may also be attributed to the reduction in ACL β transcripts levels. Based on the present results, the exact role of ACL during fruit development and ripening should be analyzed in the future by adding an ACL inhibitor in vivo and overexpressing the genes. Moreover, the mechanism underlying the response of ACL gene expression to MDS is also worth studying.
References
Albertini M-V, Carcouet E, Pailly O, Gambotti C, Luro F, Berti L (2006) Changes in organic acids and sugars during early stages of development of acidic and acidless citrus fruit. J Agric Food Chem 54:8335–8339
Baldwin EA (1993) Citrus fruit. In: Seymour GB, Taylor JE, Tucker GA (eds) Biochemistry of fruit ripening. Chapman & Hall, London, pp 107–149
Bartolozzi F, Bertazza G, Bassi D, Cristoferi G (1997) Simultaneous determination of soluble sugars and organic acids as their trimethylsilyl derivatives in apricot fruits by gas–liquid chromatography. J Chromatogr A 758:99–107
Bastías A, López-Climent M, Valcárcel M, Rosello S, Gómez-Cadenas A, Casaretto JA (2011) Modulation of organic acids and sugar content in tomato fruits by an abscisic acid-regulated transcription factor. Physiol Plant 141:215–226
Cercós M, Soler G, Iglesias D, Gadea J, Forment J, Talón M (2006) Global analysis of gene expression during development and ripening of citrus fruit flesh. A proposed mechanism for citric acid utilization. Plant Mol Biol 62:513–527
Chávez-Cabrera C, Flores-Bustamante Z, Marsch R, Montes MdC, Sánchez S, Cancino-Díaz J, Flores-Cotera L (2010) ATP-citrate lyase activity and carotenoid production in batch cultures of Phaffia rhodozyma under nitrogen-limited and nonlimited conditions. Appl Microbiol Biotechnol 85:1953–1960
Chypre M, Zaidi N, Smans K (2012) ATP-citrate lyase: a mini-review. Biochem Biophys Res Commun 422:1–4
Crifò T, Puglisi I, Petrone G, Recupero GR, Lo Piero AR (2011) Expression analysis in response to low temperature stress in blood oranges: implication of the flavonoid biosynthetic pathway. Gene 476:1–9
Crifò T, Petrone G, Lo Cicero L, Lo Piero AR (2012) Short cold storage enhances the anthocyanin contents and level of transcripts related to their biosynthesis in blood oranges. J Agric Food Chem 60:476–481
Degu A, Hatew B, Nunes-Nesi A, Shlizerman L, Zur N et al (2011) Inhibition of aconitase in citrus fruit callus results in a metabolic shift towards amino acid biosynthesis. Planta (Berlin) 234:501–513
Fatland BL, Ke J, Anderson MD, Mentzen WI, Cui LW et al (2002) Molecular characterization of a heteromeric ATP-citrate lyase that generates cytosolic acetyl-coenzyme a in Arabidopsis. Plant Physiol 130:740–756
Fatland BL, Nikolau BJ, Wurtele ES (2005) Reverse genetic characterization of cytosolic acetyl-CoA generation by ATP-citrate lyase in Arabidopsis. Plant Cell 17:182–203
Guo A-Y, Zhu Q-H, Chen X, Luo J-C (2007) GSDS: a gene structure display server. Yi chuan 29:1023
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucl Acid Res 27:297–300
Jiang N, Jin L-F, Teixeira da Silva JA, Islam MDZ, Gao H-W, Liu Y-Z, Peng S-A (2014) Activities of enzymes directly related with sucrose and citric acid metabolism in citrus fruit in response to soil plastic film mulch. Sci Hortic 168:73–80
Katz E, Fon M, Lee YJ, Phinney BS, Sadka A, Blumwald E (2007) The citrus fruit proteome: insights into citrus fruit metabolism. Planta 226:989–1005
Katz E, Boo KH, Kim HY, Eigenheer RA, Phinney BS et al (2011) Label-free shotgun proteomics and metabolite analysis reveal a significant metabolic shift during citrus fruit development. J Exp Bot 62:5367–5384
Koressaar T, Remm M (2007) Enhancements and modifications of primer design program Primer3. Bioinformatics 23:1289–1291
Langlade NB, Messerli G, Weisskopf L, Plaza S, Tomasi N et al (2002) ATP citrate lyase: cloning, heterologous expression and possible implication in root organic acid metabolism and excretion. Plant Cell Environ 25:1561–1569
Li C-N, Qian N, Tan Q-L, Manoj KS, Li-Tao Y, Li Y-R (2012) Cloning and expression analysis of ATP-citrate lyase genes from sugarcane. Acta Agron Sin 38:2024–2033
Liu YZ, Liu Q, Tao NG, Deng XX (2006) Efficient isolation of RNA from fruit peel and pulp of ripening navel orange (Citrus sinensis Osbeck). J Huazhong Agric Univ 25:300–304
Liu X, Hu X-M, Jin L-F, Shi C-Y, Liu Y-Z, Peng S-A (2014) Identification and transcript analysis of two glutamate decarboxylase genes, CsGAD1 and CsGAD2, reveal the strong relationship between CsGAD1 and citrate utilization in citrus fruit. Mol Biol Rep. doi:10.1007/s11033-11014-13506-x
Livak KJ, Schmittigen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods 25:402–408
Lo Piero A, Lo Cicero L, Puglisi I (2014) The metabolic fate of citric acid as affected by cold storage in blood oranges. J Plant Biochem Biotechnol 23:161–166
Navarro JM, Pérez-Pérez JG, Romero P, Botía P (2010) Analysis of the changes in quality in mandarin fruit, produced by deficit irrigation treatments. Food Chem 119:1591–1596
Nelson DR, Rinne RW (1975) Citrate cleavage enzymes from developing soybean cotyledons incorporation of citrate carbon into fatty acids. Plant Physiol 55:69–72
Popova TN, Pinheiro de Carvalho MA (1998) Citrate and isocitrate in plant metabolism. Biochim Biophys Acta 1364:307–325
Rangasamy D, Ratledge C (2000) Compartmentation of ATP: citrate lyase in plants. Plant Physiol 122:1225–1230
Sadka A, Dahan E, Cohen L, Marsh KB (2000) Aconitase activity and expression during the development of lemon fruit. Physiol Plant 108:255–262
Terol J, Soler G, Talon M, Cercos M (2010) The aconitate hydratase family from Citrus. BMC Plant Biol 10:222
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucl Acid Res 25:4876–4882
Xing S, van Deenen N, Magliano P, Frahm L, Forestier E et al (2014) ATP citrate lyase activity is post-translationally regulated by sink strength and impacts the wax, cutin and rubber biosynthetic pathways. Plant J 79:270–284
Xu Q, Chen LL, Ruan X, Chen D, Zhu A et al (2012) The draft genome of sweet orange (Citrus sinensis). Nat Genet 45:59–66
Zaidi N, Swinnen JV, Smans K (2012) ATP-citrate lyase: a key player in cancer metabolism. Cancer Res 72:3709–3714
Zhou GF, Peng SA, Liu YZ, Wei QJ, Han J, Islam MZ (2014) The physiological and nutritional responses of seven different citrus rootstock seedlings to boron deficiency. Trees 28:295–307
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 31372012).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, XM., Shi, CY., Liu, X. et al. Genome-wide identification of citrus ATP-citrate lyase genes and their transcript analysis in fruits reveals their possible role in citrate utilization. Mol Genet Genomics 290, 29–38 (2015). https://doi.org/10.1007/s00438-014-0897-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-014-0897-2