Abstract
The Tsc/Rheb signaling pathway plays critical roles in the control of growth and cell cycle. Studies in fission yeast have also implicated its importance in the regulation of amino acid uptake. Disruption of tsc2 +, one of the tsc + genes, has been shown to result in decreased arginine uptake and resistance to canavanine. A similar effect is also seen with other basic amino acids. We have identified a permease responsible for the uptake of basic amino acids by genetic complementation and disruption. SPAC869.11 (termed Cat1 for cationic amino acid transporter) contains 12 predicted transmembrane domains and its overexpression in wild type fission yeast leads to the increased uptake of basic amino acids and sensitivity to canavanine. Disruption of cat1 + in the Δtsc2 background interfered with the suppression of the canavanine-resistant phenotype of Δtsc2 mutants by a dominant negative Rheb. In Δtsc2 mutant strains, the amount of Cat1 was not altered, but instead was mislocalized. This mislocalization was suppressed by the expression of dominant negative Rheb. In addition, we found that the loss of the E3 ubiquitin ligase, Pub1, also restores proper localization. These results provide a crucial link between Tsc/Rheb signaling and the regulation of the basic amino acid permease in fission yeast.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
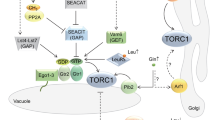
Rheb G-protein belongs to a unique family of the Ras superfamily GTP-binding proteins (Aspuria and Tamanoi 2003; Aspuria et al. 2007). Rheb is conserved from yeast to human and possesses unique structural features including the presence of arginine at residue 15 that corresponds to glycine-12 in Ras. Rheb is downregulated by the complex of Tsc1 and Tsc2 proteins that acts as a GTPase activating protein (GAP) for Rheb. Mutations in the TSC1 or TSC2 gene result in tuberous sclerosis complex (TSC), a genetic disorder that is associated with the appearance of benign tumors at various sites in the body including the kidneys, lungs and brain (Gomez et al. 1999; Crino et al. 2006).
Studies in yeast have led to the idea that Tsc/Rheb signaling regulates uptake of basic amino acids such as arginine and lysine. Our initial observation was made using budding yeast (Urano et al. 2000). In this organism, disruption of the RHB1 gene encoding the budding yeast Rheb led to the increased uptake of arginine and lysine and increased sensitivity to canavanine and thialysine, toxic analogues of arginine and lysine, respectively. Recent studies have shown that fission yeast is an ideal system to further investigate Tsc/Rheb/TOR signaling. Fission yeast has both tsc1 + and tsc2 + genes and, like mammalian cells, these gene products form a complex that act to downregulate Rheb (Matsumoto et al. 2002). Loss of Tsc function in fission yeast by disrupting tsc1 + or tsc2 + leads to decreased uptake of arginine and a dramatic increase in resistance to canavanine (van Slegtenhorst et al. 2004). The pool of arginine and lysine is decreased in the mutants. The canavanine-resistant phenotype of Δtsc mutants is reversed by the expression of dominant negative Rheb protein. Inhibition of Rheb function by the inhibition of protein farnesyltransferase leads to hypersensitivity to canavanine and increased uptake of arginine (Yang et al. 2001). On the other hand, hyperactivation of Rheb causes resistance to canavanine and thialysine (Urano et al. 2006). These results suggest that the Tsc/Rheb signaling pathway negatively regulates the uptake of arginine and, presumably, lysine.
To further understand the regulation of basic amino acid uptake by the Tsc/Rheb signaling pathway, we sought to identify the permease responsible for the uptake of basic amino acids in fission yeast. To accomplish this, we developed a two-step assay that utilized both budding yeast and fission yeast. We first took advantage of the availability of a budding yeast mutant defective in Can1, a permease for arginine (Grenson et al. 1966; Ahmad and Bussey 1986). Since this mutant is resistant to canavanine, we sought to identify a fission yeast gene whose expression in the Δcan1 mutant restores canavanine sensitivity. The candidate permease genes were then disrupted in fission yeast. Finally overexpression in the wild type fission yeast was used to confirm the involvement of the putative permease in the uptake of basic amino acids. This led to the identification of SPAC869.11 as a fission yeast permease responsible for the uptake of basic amino acids (cationic amino acid transporter, Cat1). Characterization of Cat1 showed that its amount is unchanged in the Δtsc2 mutant strain; rather its intracellular localization is affected.
Materials and methods
Yeast strains, media and reagents
Budding yeast cells were grown in YPD or SD with the appropriate supplements (Sherman 1991). Fission yeast cells were grown in yeast extract complete medium with 225 μg/μl adenine, leucine, histidine, uracil (YES) or Edinburgh minimal media (EMM) + 225 μg/μl adenine at 30°C (Moreno et al. 1991). Transformations were performed using the lithium acetate method (Kanter-Smoler et al. 1994). Yeast strains used are listed in Table 1. The tsc2 +, spac869.11c +, spac359.03c +, sbpb2b2.01c + and pub1 + genes were disrupted by the PCR-based method using the kanamycin resistance cassette kanMX, URA cassette, and the hygromycin B resistance cassette hphMX (Bahler et al. 1998; Sato et al. 2005). The 3xHA-hphMX and GFP-hphMX cassettes were also integrated into the spac869.11 + C-terminus via the same PCR-based method. Canavanine and thialysine were purchased from Sigma and Research Organics, respectively. [3H]arginine (40 Ci/mmol), [3H]lysine (60 Ci/mmol), [3H]histidine (40 Ci/mmol), [3H]proline (20 Ci/mmol), and [3H]leucine (72.5 Ci/mmol) were obtained from American Radiolabeled Chemicals.
Constructs
p416a-ADH-fission yeast permeases were created by a PCR-cloning approach. The putative Schizosaccharomyces pombe permeases were amplified from a genomic DNA prep from the wild-type strain FY972 using primers with XhoI and XbaI restriction sites and cloned into the p416a-ADH expression vector. pREP41-c869.11 +-Myc was also created by a PCR-cloning approach by using primers with SalI and BamHI restriction sites and cloned into the thiamine repressible pREP41 expression vector.
Amino acid uptake assays
Amino acid uptake assays were performed essentially as previously described with some minor modifications (Urano et al. 2000). Cells were grown to mid log phase (OD600 0.4–0.8) in EMM with appropriate supplements, collected, and washed with dH2O. Cells were resuspended in 1.2 ml of media without amino acids to an OD600 2.0. 4.9 μl of [3H]amino acid and 50 μM cold amino acid were then added. Two hundred-microliter aliquots were taken at the indicated time points, injected into 5 ml of dH2O, and immediately filtered and washed twice in a vacuum manifold. Filtration was performed on glass fiber filter circles (Fisher Scientific). Filters were dried under a heat lamp and counted in a Beckman LS-6500 scintillation counter using Econosafe scintillation solution (Resource Product International).
Northern blot analysis
Ten micrograms of total RNA was run on a 4% formaldehyde gel at 100 V for 1 h and transferred to a nylon membrane for 3 h in 10× SSC. Probes for spac869.11c +and tub1 + were PCR-amplified from cDNA, cleaned via a Quick Spin column (Roche Diagnostics) and labeled with [α32P] dATP (American Radiolabeled Chemicals) using standard methods. Hybridizations were performed in Quickhyb buffer (Stratagene).
Western blot analysis
Whole cell extracts were made essentially as described previously with minor modifications (Umebayashi and Nakano 2003). Fifty milliliters of cells were grown to an OD600 1.0–2.0, collected, and washed with dH2O. The cells were resuspended in 250 μl of lysis buffer (20 mM Tris–HCl, pH 7.4, 1 mM EDTA, 1.6% SDS, 6 M urea) containing a protease inhibitor mixture [1 mM PMSF and 1× protease inhibitor cocktail (Roche diagnostics)]. Cells were lysed with glass beads and large cell debris and unbroken cells were removed by centrifugation for 10 min at 3,000 rpm. Protein concentration was determined using the Bradford method (Bio-Rad protein assay). Fifty micrograms of the sample was subjected to SDS-PAGE and immunoblotting with anti-HA antibody (HA11 from BabCO) or anti-PCNA antibody (Ab1 from Oncogene).
Fluorescence microscopy
For Cat1-GFP localization, cells were grown to midlog phase in EMM medium. The cells were then spotted directly onto poly-l-lysine slides and visualized using a Zeiss Microscope. Images were captured using AxioVision software.
Results
The Δtsc2 mutant cells have a defect in basic amino acid uptake, which can be suppressed by the expression of dominant negative RhebD60K.
The Tsc/Rheb signaling pathway has been shown to regulate arginine uptake (Yang et al. 2001; van Slegtenhorst et al. 2004; Urano et al. 2006). To determine whether this is applied to all basic amino acids, we examined the uptake of lysine, arginine and histidine in Δtsc2 mutant cells. In these mutant cells, Rheb is activated because Tsc2 of the Tsc1/Tsc2 complex that acts as a GAP is missing. As can be seen, there is a three- to fivefold decrease in the uptake of basic amino acids in Δtsc2 mutant cells compared with the control cells that have tsc2 + expressed in the mutant (Fig. 1a). This point was further confirmed by the use of canavanine and thialysine, as Δtsc2 mutant cells were resistant to these toxic amino acid analogues (Fig. 8).
The decreased uptake of basic amino acids in Δtsc2 mutant cells was reversed by the expression of a dominant negative Rheb. This is shown in Fig. 1c, where we examine the uptake of lysine, arginine and histidine in Δtsc2 mutant cells expressing a dominant negative Rheb mutant, RhebD60K (rhb1D60K+ construct) (Tabancay et al. 2003). As can be seen, expression of dominant negative Rheb causes an increase in the uptake of these basic amino acids. In addition, expression of dominant negative Rheb restores sensitivity of Δtsc2 mutant cells to canavanine (Fig. 8).
Δtsc2 Cells defect in basic amino acid uptake and resistance to canavanine can be suppressed by the expression of dominant negative RhbD60K. a Δtsc2 cells transformed with pREP1-tsc2 + or pREP1 were assayed for the uptake of arginine, lysine and histidine. Assays were done in triplicate. b Δtsc2 cells transformed with pREP1-tsc2 + or pREP1 were spotted onto EMM + Ade plates either with canavanine or thialysine. c Δtsc2 cells transformed with pREP41-rhb1D60K + or pREP41 were assayed for the uptake of arginine, lysine and histidine. Assays were done in triplicate
Identification of SPAC869.11 as a cationic amino acid permease in fission yeast
In order to understand how Rheb signaling controls the uptake of basic amino acids, we sought to identify the cationic amino acid permease in fission yeast. We first examined putative permeases with sequence similarity to Saccharomyces cerevisiae Can1p according to the BLAST search of the Sanger Center Schizosaccharomyces pombe database. Twelve amino acid permeases were identified by this analysis and the result of this analysis is shown in Fig. 2. As can be seen, the overall percent identity among these putative permeases is rather low. Based upon homology, it is possible to come up with a dendrogram for these putative permeases. However, it is difficult to choose a particular permease that is highly similar to Can1p from this analysis.
Amino acid sequence analysis using DNAStar was performed on ScCan1 and 12 hypothetical Schizosaccharomyces pombe amino acid permeases. a Percent identity table shows low amino acid identity between the Schizosaccharomyces pombe amino acid permeases and ScCan1. b Phylogenetic tree based upon the divergence from ScCan1
We decided to experimentally identify the fission yeast permease responsible for the uptake of basic amino acids using a two-step assay described in Fig. 3. In this approach, we first take advantage of the expected functional similarity between the fission yeast permease and the budding yeast Can1p. Disruption of the budding yeast CAN1 gene leads to a defect in arginine uptake, highlighted by a resistance to the toxic arginine analogue, canavanine. Therefore, we initially screened putative Schizosaccharomyces pombe amino acid permease ORFs for their ability to convert resistance of the Δcan1 mutant to a canavanine-sensitive phenotype. In the second step, these candidates were disrupted in Schizosaccharomyces pombe and it was examined whether the disruption caused canavanine resistance. To overexpress these fission yeast ORFs in Saccharomyces cerevisiae Δcan1 strain and assay for restoration of canavanine sensitivity, the ORFs were placed under the control of the ADH promoter. As summarized in Fig. 4, expression of spb2b2b2.01 +, spbc359.03 + or spac869.11 + restored canavanine sensitivity to the Δcan1 mutant. To further characterize the three Schizosaccharomyces pombe basic amino acid permease candidates, we disrupted each of them by the kanMX or URA4 cassette in the fission yeast strain SP812. These strains were then assayed for their resistance to canavanine. Disruption of spac869.11 + was able to confer resistance to canavanine, whereas the disruption of spb2b2b2.01 + and spbc359.03 + did not (Fig. 5a). Similar results were obtained with thialysine, when spac869.11 + was disrupted (unpublished observation). This was also furthered by the analysis of double-disrupted strains, wherein canavanine resistance was observed when spac869.11 + was disrupted.
ScCan1 complementation assay. Budding yeast Δcan1 mutant cells were transformed with putative fission yeast amino acid permeases that are placed under control of the ADH promoter. a Three positive transformants (spbpb2b2.01 +, spbc359.03 + and spac869.11 +), a negative transformant (spac869.10 +), and a vector control were streaked onto SD–URA with and without canavanine. Strains that complement ScCan1 restore sensitivity to canavanine. b Spotting of three positives (spbpb2b2.01 +, spbc359.03 + and spac869.11 +) on a plate containing canavanine. Two clones each were tested for spbpb2b2.01 + and spbc359.03 +
Loss of SPAC869.11 results in canavanine resistance and a defect in basic amino acid uptake. a Schizosaccharomyces pombe strains with spac869.11 +, spac359.03 +, and spbpb2b2.01 + disrupted with the kanMX or URA cassette were serially diluted and spotted onto EMM + Ade plates with and without 60 mg/l canavanine. Δspac869.11 as well as Δspac869.11 double-mutant cells were resistant to canavanine. b and c Arginine and lysine uptake assays were performed on Δspac869.11 and Δspac359.03 cells. Δspac869.11 Δspbc359.03 double-mutant cells were also assayed for their ability to take up arginine. Cells lacking spac869.11 + have a defect in arginine and lysine uptake. Wild type and Δtsc2 cells were used as positive and negative controls, respectively. Assays were done in triplicate
Because canavanine resistance is likely due to a defect in arginine uptake, the wild-type strain as well as the Δtsc2, Δspbc359.03, and Δspac869.11 strains were assayed for their ability to take up [3H]arginine. Cells with spac869.11 + disrupted, including the double mutant, exhibited a defect in arginine uptake, similar to the Δtsc2 mutants compared to wild type (Fig. 5b). On the other hand, loss of spbc359.03 + had no effect. Disruption of spac869.11 + also had a major effect on lysine uptake (Fig. 5c). As shown in Fig. 5c, Δspac869.11 cells failed to take up [3H]lysine and the level of decreased uptake was similar to that seen with the tsc2 + disruption strain. These uptake defects appear to be specific for arginine and lysine, as the uptake of non-basic amino acids such as tyrosine, proline, and leucine was not decreased and instead slightly increased (data not shown). From these results, we conclude that the spac869.11 + gene encodes the major permease specifically involved in arginine and lysine uptake in fission yeast.
Features of SPAC869.11
spac869.11 + encodes a protein of 580 amino acids, which contains the PROSITE amino acid permeases signature. While the amino acid sequences of SPAC869.11 and Saccharomyces cerevisiae Can1 are only 14.8% identical, we found that they are structurally similar. Using a prediction server of transmembrane topology, ConPredII (Arai et al. 2004), we deduced the predicted transmembrane architectures of SPAC869.11 and Saccharomyces cerevisiae Can1. As can be seen in Fig. 6, the structures of these permeases are predicted to be very similar. SPAC869.11 has 12 predicted transmembrane domains with an intracellular 87 amino acid N-terminal tail and 47 amino acid C-terminal tail. This overall structure is strikingly similar to that of the Saccharomyces cerevisiae Can1. Interestingly, both transporters have a conserved glutamate residue in the periplasmic loop 7, which has been speculated to be involved in cationic substrate recognition (Regenberg and Kielland-Brandt 2001).
Transmembrane architecture analysis of SPAC869.11 and ScCan1. The amino acid sequences of both permeases were analyzed by ConPredII (Arai et al. 2004), a transmembrane prediction software program. In and out designate intracellular and extracellular matrices, respectively. The conserved glutamate residue is marked by an asterisk
Overexpression of spac869.11 + leads to an increase in the uptake of basic amino acids
We overexpressed spac869.11 + in the wild-type strain SP812 and measured the rate of uptake of arginine, lysine and histidine. Overexpressing spac869.11 + using the pREP41 or pREP81 promoter was sufficient to cause a marked increase in the amount of arginine, lysine and histidine uptake (Fig. 7a). The increase in arginine uptake occurred in a dose-dependent matter, as we observed higher uptake when using a stronger promoter (data not shown). In contrast, the overexpression of spac869.11 + did not alter the uptake of proline, thus implying specificity for basic amino acids. The overexpression of spac869.11 + also led to the increase in sensitivity to canavanine (Fig. 7b). As can be seen, the strain overexpressing spac869.11 + grew significantly less on a plate containing canavanine, compared to the control strain carrying the vector. This, along with the disruption data, suggested that spac869.11 + is a major Schizosaccharomyces pombe arginine transporter, which we have designated as cat1 + (cationic amino acid transporter).
Overexpression of cat1 + in wild type cells increases basic amino acid uptake and canavanine sensitivity. a spac869.11 + was overexpressed under the control of the pREP41 promoter in wild type cells. Cells were assayed for their ability to take up [3H] arginine, lysine, histidine, and proline. Cells overexpressing spac869.11 + had a significantly higher uptake of basic amino acids. Assays were done in triplicate. b The same cells were spotted onto EMM + Ade plates with and without 10 mg/l canavanine to test for sensitivity
Cat1 is required for RhebD60K suppression of Δtsc2 mutants’ resistance to canavanine
With the identification of a permease responsible for the uptake of basic amino acids, we returned our attention to the situation in Δtsc2 mutant cells. Based upon the results in Fig. 1d, we asked whether cat1 + is required for the ability of dominant negative Rheb to restore canavanine sensitivity in Δtsc2 mutant cells. As shown in Fig. 8, the canavanine resistance of Δtsc2 mutants is reversed by the expression of rhb1D60K + to a level comparable to that seen with the Δtsc2 mutant cells expressing tsc2 + on a plasmid. In contrast, when the cat1 + gene is disrupted, rhb1D60K + expression did not lead to the restoration of canavanine sensitivity in Δtsc2 mutant cells. Therefore, these results are consistent with the idea that Cat1 functions downstream of Rheb to regulate the uptake of arginine.
Cat1 expression is unchanged but its localization is altered in Δtsc2 mutant cells
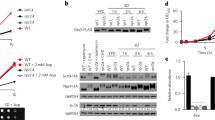
The above results suggest that there is a decrease in the function of Cat1 in Δtsc2 mutant cells. To further characterize this point, we first examined expression of the cat1 + gene by performing Northern analysis of wild type and Δtsc2 mutant cell lysates using a full-length cat1 + probe. tub1 + was used as a loading control. A single band of 1.7 kb was detected and the intensity of the bands between wild type and the Δtsc2 mutant was similar (Fig. 9a). Thus, cat1 + expression is unaffected by the disruption of the tsc2 + gene. We then performed Western analysis to examine Cat1 protein level. A 3xHA epitope tag was chromosomally integrated in the C-terminus of cat1 +. Anti-HA antibody was used to examine the level of Cat1 protein. On a gel, a band was observed at roughly 60 kD, the expected size of Cat1. As shown in Fig. 9b, total amount of the permease protein is not altered in Δtsc2 mutant cells.
Loss of tsc2 + does not affect cat1 + expression, but results in the mislocalization of Cat1. a Northern analysis of cat1 + in wild type and Δtsc2 mutant cells. tub1 + is used as a loading control. b Western analysis of Cat1-3xHA in wild type and Δtsc2 mutant cells. PCNA is used as a loading control. c Fluorescence microscopy of Cat1-GFP in wild type, Δtsc2 mutant cells transformed with or without pREP1-RhebD60K +, and Δtsc2Δpub1 cells
Since protein levels are not affected, we examined if Cat1 was altered in its intracellular localization in Δtsc2 mutant cells. This was examined using strains with a GFP tag chromosomally integrated into the C-terminus of cat1 +. Cat1-GFP is functional as assessed by the strain’s sensitivity to canavanine (data not shown). In wild-type cells, Cat1-GFP was predominantly found on the periphery of cell tips with minor internal staining (Fig. 9c). This is similar to the plasma membrane ABC transporter, Pmd1 (Iwaki et al. 2006). In Δtsc2 mutant cells, however, there was a more dispersed and mostly cytoplasmic fluorescence of the Cat1 protein. This punctate staining is similar to that of the golgi protein Ynd1 (Matsuyama et al. 2006). Therefore, Cat1 appears to be mislocalized in the Δtsc2 mutant. Since RhebD60K could revert the Δtsc2 mutant’s defect in arginine uptake (Fig. 8), we assessed whether Cat1 localization was being restored. Indeed, the expression of RhebD60K could restore proper Cat1-GFP localization to the cell tips in the Δtsc2 mutant (Fig. 9c).
We also found that loss of the E3 ubiquitin ligase Pub1 restores proper localization of Cat1 in the Δtsc2 mutant. Cat1-GFP localization was examined in the Δpub1Δtsc2 double mutant and was found properly localized to the periphery of cell tips as well as mild intracellular punctate staining (Fig. 9c). Based on these results, it appears that Pub1 is required for the control of Tsc2 in Cat1-GFP localization.
Discussion
In this paper, we report the identification of a fission yeast permease for cationic amino acids. This permease, Cat1, is capable of replacing the function of the arginine permease, Can1p, in Saccharomyces cerevisiae. Disruption of cat1 + in fission yeast causes dramatic decrease in the uptake of arginine or lysine and resistance to canavanine and thialysine. Overexpression of cat1 + in fission yeast results in the significant increase in the uptake of arginine, lysine and histidine. These results provide convincing evidence for the assignment of spac869.11 + as a major permease responsible for the uptake of basic amino acids in fission yeast.
The predicted overall structure of Cat1 is very similar to that of budding yeast Can1p. Both N-terminal and C-terminal sequences are predicted to be located inside the cell and a crucial residue speculated to be involved in the recognition of cationic amino acids is present in the same periplasmic loop (Regenberg and Kielland-Brandt 2001). Thus, the basic mechanism of the function of these permeases may be similar between the fission yeast and budding yeast proteins. On the other hand, there are intriguing differences between the two permeases. In budding yeast, separate permeases are used for the uptake of arginine, lysine and histidine (Can1p, Lyp1p and Hip1p for arginine, lysine and histidine, respectively). In contrast, our study suggests that a single permease can take up all three basic amino acids in fission yeast. In support of this idea, the uptake of lysine into fission yeast cells is competed by l-arginine, l-histidine and d-lysine (Sychrova et al. 1989).
Cat1 appears to be the major route for the uptake of arginine in the presence of ammonium. Interestingly, preliminary data suggest that when ammonium is depleted, Δtsc2 cells as well as Δcat1 are no longer canavanine resistant (unpublished observation). These results may suggest that another permease that can take up arginine can function in the absence of ammonium. It has been speculated that there are two systems involved in the uptake of arginine (Fantes and Creanor 1984). System I is the major route for the uptake of arginine, while system II is inhibited in the presence of ammonium in the medium. In support of this idea, the double-disrupted Δcat1 Δspbc359.03 strain is still canavanine-resistant under ammonium depletion (unpublished observation). This may suggest that Cat1 and SPBC359.03 are components of system I and system II, respectively. This idea is consistent with our observation that SPBC359.03 could confer canavanine sensitivity to the Saccharomyces cerevisiae Δcan1 mutant when overexpressed. In addition, this gene product is most similar to Cat1.
Our results show that the Cat1 permease is enriched toward the growing ends of fission yeast, possibly reflecting its increased utilization at the growing ends. Similar localization of another fission yeast permease, SPBC359.03, has also been reported (Matsumoto et al. 2002). This intracellular distribution of both permeases is altered significantly in Δtsc2 cells; Cat1 and SPBC359.03 staining appear dispersed as intracellular dots. Thus, the Tsc/Rheb signaling pathway regulates the localization of these permeases. On the other hand, we did not detect any significant changes in the expression of cat1 + mRNA and the total amount of Cat1 permease. It has been previously reported that the transcription of isp5 +, spac869.10 +, and spap7g5 + is decreased in Δtsc2, Δtor1, and in rapamycin-treated cells (van Slegtenhorst et al. 2004; Weisman et al. 2005, 2007). In addition, we have observed the induction of expression of a variety of putative amino acid permeases upon inhibition of Tor2 (Matsuo et al. 2007). Therefore, it appears that there are transcriptional as well as post-translational mechanisms used to regulate the uptake of amino acids in fission yeast.
What could account for the mislocalization of the permease? While further work is needed, it is tempting to speculate that control of the ubiquitination machinery is involved in this regulation. In budding yeast, ubiquitination of amino acid permeases by the ubiquitin ligase RSP5 is known to serve as a post-translational modification/signal for intracellular sorting (Hein et al. 1995; Soetens et al. 2001; Blondel et al. 2004). Interestingly, it has been shown that the Schizosaccharomyces pombe Rsp5 homologue, Pub1, is required for the downregulation of leucine uptake (Karagiannis et al. 1999). Therefore, we looked to see if Pub1 was involved in the Tsc2 regulation of arginine uptake. Disruption of pub1 + in a wild-type background resulted in hypersensitivity to canavanine (unpublished observation). In addition, (Δpub1Δtsc2 double mutants were also sensitive to canavanine (unpublished observation). This suggested that perhaps Cat1 mislocalization in the Δtsc2 mutants was being suppressed by the loss of pub1 +. We have shown that the E3 ubiquitin ligase, Pub1, is important for Cat1-GFP mislocalization in Δtsc2 cells. Further work is needed to investigate whether ubiquitination plays a role in the regulation of Cat1 localization by Tsc/Rheb signaling. Also, it is unclear as to where in Cat1 trafficking, anterograde or retrograde transport to and from the plasma membrane, is targeted by Tsc/Rheb signaling. Further studies on how Tsc/Rheb signaling affects transcription and localization of various permeases, as well as its interplay with its downstream effectors, may provide a deeper understanding of how this signaling pathway controls amino acid transport.
References
Ahmad M, Bussey H (1986) Yeast arginine permease: nucleotide sequence of the CAN1 gene. Curr Genet 10(8):587–592
Arai M, Mitsuke H, Ikeda M, Xia JX, Kikuchi T, Satake M, Shimizu T (2004) ConPred II: a consensus prediction method for obtaining transmembrane topology models with high reliability. Nucleic Acids Res 32(Web Server Issue):W390–W393
Aspuria PJ, Tamanoi F (2003) The Rheb family of GTP-binding proteins. Cell Signal 16(10):1105–1112
Aspuria PJ, Sato T, Tamanoi F (2007) The TSC/Rheb/TOR signaling pathway in fission yeast and mammalian cells: temperature sensitive and constitutive active mutants of TOR. Cell Cycle 6(14):1692–1695
Bahler J, Wu JQ, Longtine MS, Shah NG, McKenzie A III, Steever AB, Wach A, Philippsen P, Pringle JR (1998) Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14(10):943–951
Blondel M, Morvan J, Dupre S, Urban-Grimal D, Haguenauer-Tsapis R, Volland C (2004) Direct sorting of the yeast uracil permease to the endosomal system is controlled by uracil binding and Rsp5p-dependent ubiquitylation. Mol Biol Cell 15(2):883–895
Crino P, Nathanson KL, Henske EP (2006) The tuberous sclerosis complex. N Engl J Med 355(13):1345–1356
Fantes P, Creanor J (1984) Canavanine resistance and the mechanism of arginine uptake in the fission yeast Schizosaccharomyces pombe. J Gen Microbiol 130:3265–3273
Gomez MR, Sampson JR, Whittemore VH (1999) Tuberous sclerosis complex. 3rd edn. Oxford University Press, New York
Grenson M, Mousset M, Wiame JM, Bechet J (1966) Multiplicity of the amino acid permeases in Saccharomyces cerevisiae. I. Evidence for a specific arginine-transporting system. Biochim Biophys Acta 127(2):325–338
Hein C, Springael JY, Volland C, Haguenauer-Tsapis R, Andre B (1995) NPl1, an essential yeast gene involved in induced degradation of Gap1 and Fur4 permeases, encodes the Rsp5 ubiquitin-protein ligase. Mol Microbiol 18(1):77–87
Iwaki T, Giga-Hama Y, Takegawa K (2006) A survey of all 11 ABC transporters in fission yeast: two novel ABC transporters are required for red pigment accumulation in a Schizosaccharomyces pombe adenine biosynthetic mutant. Microbiology 152(Pt 8):2309–2321
Kanter-Smoler G, Dahlkvist A, Sunnerhagen P (1994) Improved method for rapid transformation of intact Schizosaccharomyces pombe cells. Biotechniques 16(5):798–800
Karagiannis J, Saleki R, Young PG (1999) The pub1 E3 ubiquitin ligase negatively regulates leucine uptake in response to NH(4)(+) in fission yeast. Curr Genet 35(6):593–601
Matsumoto S, Bandyopadhyay A, Kwiatkowski DJ, Maitra U, Matsumoto T (2002) Role of the Tsc1–Tsc2 complex in signaling and transport across the cell membrane in the fission yeast Schizosaccharomyces pombe. Genetics 161(3):1053–1063
Matsuo T, Otsubo Y, Urano J, Tamanoi F, Yamamoto M (2007) Loss of the TOR kinase Tor2 mimics nitrogen starvation and activates the sexual development pathway in fission yeast. Mol Cell Biol 27(8):3154–3164
Matsuyama A, Arai R, Yashiroda Y, Shirai A, Kamata A, Sekido S, Kobayashi Y, Hashimoto A, Hamamoto M, Hiraoka Y, Horinouchi S, Yoshida M (2006) ORFeome cloning and global analysis of protein localization in the fission yeast Schizosaccharomyces pombe. Nat Biotechnol 24(7):841–847
Moreno S, Klar A, Nurse P (1991) Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol 194:795–823
Regenberg B, Kielland-Brandt MC (2001) Amino acid residues important for substrate specificity of the amino acid permeases Can1p and Gnp1p in Saccharomyces cerevisiae. Yeast 18(15):1429–1440
Sato M, Dhut S, Toda T (2005) New drug-resistant cassettes for gene disruption and epitope tagging in Schizosaccharomyces pombe. Yeast 22(7):583–591
Sherman F (1991) Getting started with yeast. Methods Enzymol 194:3–21
Soetens O, De Craene JO, Andre B (2001) Ubiquitin is required for sorting to the vacuole of the yeast general amino acid permease, Gap1. J Biol Chem 276(47):43949–43957
Sychrova H, Horak J, Kotyk A (1989) Transport of l-lysine in the fission yeast Schizosaccharomyces pombe. Biochim Biophys Acta 978(2):203–208
Tabancay AP Jr, Gau CL, Machado IM, Uhlmann EJ, Gutmann DH, Guo L, Tamanoi F (2003) Identification of dominant negative mutants of Rheb GTPase and their use to implicate the involvement of human Rheb in the activation of p70S6K. J Biol Chem 278(41):39921–39930
Umebayashi K, Nakano A (2003) Ergosterol is required for targeting of tryptophan permease to the yeast plasma membrane. J Cell Biol 161(6):1117–1131
Urano J, Tabancay AP, Yang W, Tamanoi F (2000) The Saccharomyces cerevisiae Rheb G-protein is involved in regulating canavanine resistance and arginine uptake. J Biol Chem 275(15):11198–11206
Urano J, Comiso MJ, Guo L, Aspuria PJ, Deniskin R, Tabancay AP Jr, Kato-Stankiewicz J, Tamanoi F (2006) Identification of novel single amino acid changes that result in hyperactivation of the unique GTPase, Rheb, in fission yeast. Mol Microbiol 58(4):1074–1086
van Slegtenhorst M, Carr E, Stoyanova R, Kruger WD, Henske EP (2004) Tsc1+ and tsc2+ regulate arginine uptake and metabolism in Schizosaccharomyces pombe. J Biol Chem 279(13):12706–12713
Weisman R, Roitburg I, Nahari T, Kupiec M (2005) Regulation of leucine uptake by tor1+ in Schizosaccharomyces pombe is sensitive to rapamycin. Genetics 169(2):539–550
Weisman R, Roitburg I, Schonbrun M, Harari R, Kupiec M (2007) Opposite effects of tor1 and tor2 on nitrogen starvation responses in fission yeast. Genetics 175(3):1153–1162
Yang W, Tabancay AP Jr, Urano J, Tamanoi F (2001) Failure to farnesylate Rheb protein contributes to the enrichment of G0/G1 phase cells in the Schizosaccharomyces pombe farnesyltransferase mutant. Mol Microbiol 41(6):1339–1347
Acknowledgments
We would like to thank the members of the Tamanoi lab for their constructive comments. We would also like to thank Dr Susan Forsburg, Dr Angel Tabancay Jr, and Dr Peter Bradley for their help with the microscopy experiments, Dr Takashi Toda for the hphMX based plasmids, and Dr Tomohiro Matsumoto for the AE502 and SP812 strains. This work is supported by the NIH grant CA41996.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann.
Rights and permissions
About this article
Cite this article
Aspuria, PJ., Tamanoi, F. The Tsc/Rheb signaling pathway controls basic amino acid uptake via the Cat1 permease in fission yeast. Mol Genet Genomics 279, 441–450 (2008). https://doi.org/10.1007/s00438-008-0320-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-008-0320-y