Abstract
Yeast Ssb proteins (Ssbp) are ribosome-associated Hsp70 chaperones that function in translation. Elevated levels of Ssbp enhance the ability of over-expressed Hsp104 chaperone to eliminate the yeast [ PSI +] prion, while depletion of Ssbp reduces this effect. Millimolar concentrations of guanidine in the growth medium cure yeast cells of prions by inactivating Hsp104. Guanidine is also toxic to yeast, irrespective of the status of Hsp104 and [ PSI +]. Strains that lack Ssbp are hypersensitive to guanidine toxicity. Here we show that ssb – cells have normal numbers of [ PSI +] "seeds", but can be cured of [ PSI +] using one-sixth of the guanidine concentration required to eliminate [ PSI +] from SSB cells. Correspondingly, the level of intracellular guanidine was eight-fold higher in ssb – cells than in wild-type cells, which explains all effects of Ssbp depletion on susceptibility to guanidine. The sensitivity of wild-type cells to the effects of guanidine also correlated with guanidine uptake, which was enhanced at low temperature. Guanidine sensitivity of strains mutated in any of 16 ABC membrane transporters, which are implicated in multidrug resistance, was normal. We found that an erg6mutant that has an altered membrane lipid composition was hypersensitive to guanidine toxicity, but the lipid composition of ssb – cells was identical to that of wild-type cells. Our results suggest that Ssbp depletion does not affect prion seed regeneration, and that elevated guanidine uptake by ssb – cells may be due to increased retention rather than to an alteration in active or passive transport of the compound.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Ssb1 and Ssb2 proteins of S. cerevisiae, collectively referred to as Ssbp, are 99% identical, and are functionally redundant ribosome-associated chaperones. They are referred to as heat shock cognates as they share 60% identity with the Ssa Hsp70 family (Werner-Washburne et al. 1989; Craig et al. 1995), although Ssbp expression is not induced by conditions that efficiently induce bona-fide heat shock proteins. In fact, expression of Ssbp is repressed at elevated temperatures and induced by low temperature (Werner-Washburne et al. 1989). Ssbp was shown to be associated with ribosomes through an interaction with the nascent polypeptide chain, which led to the proposal that the proteins act as chaperones that facilitate correct folding of newly synthesized polypeptides and assist in transport of nascent peptides through the ribosome (Nelson et al. 1992; Pfund et al. 1998).
The term prion was coined to refer to infectious proteins (Prusiner 1982). The known prions are transmissible amyloid forms of cellular proteins that propagate by converting the native protein into the same abnormal prion form. A prion-like mechanism has been shown to underlie the inheritance of the non-Mendelian genetic elements [ PSI +] and [ URE3] in yeast (Wickner 1994). The [ PSI +] element is a prion form of the Sup35 protein (eRF3), a eukaryotic release factor that plays an essential role in translation termination (Stansfield et al. 1995; Zhouravleva et al. 1995). In [ PSI +] cells, recruitment of much of the soluble form of Sup35p into prion aggregates reduces the efficiency of translation termination, which in turn causes a nonsense suppressor phenotype that can be used to monitor the [ PSI +] state.
Much work on yeast prions has focused on protein chaperones and their effects on prion stability. Thus, the chaperones Hsp104, Hsp70, and Hsp40 have been implicated in yeast prion propagation (Chernoff et al. 1995,1999; Newnam et al. 1999; Jung et al. 2000; Kushnirov et al. 2000; Moriyama et al. 2000; Chacinska et al. 2001). Hsp104, a stress-response chaperone that solubilizes protein aggregates, is required for yeast prion propagation but can eliminate [ PSI +] if overproduced under non-stress conditions. A mutation in Ssa1p, a constitutively expressed cytosolic Hsp70 whose expression is further induced by stress, can destabilize [ PSI +] (Jung et al. 2000; Jones and Masison 2003), and deleting both SSB1and SSB2reduces the [ PSI +]-curing effect of HSP104over-expression (Chernoff et al. 1999). In addition, over-expression of the SSB1or the HSP40gene can cure cells of [ PSI +] (Kushnirov et al. 2000; Chacinska et al. 2001).
Growing yeast in low concentrations (3–5 mM) of guanidine causes them to lose [ PSI +] by specifically inactivating Hsp104 (Tuite et al. 1981; Ferreira et al. 2001; Jung and Masison 2001; Jung et al. 2002; Ness et al. 2002). Guanidine is toxic to yeast at concentrations above 3–5 mM. The reason for this toxicity is unknown, but it is independent of both [ PSI +] and Hsp104 (Jung et al. 2002). Earlier work reported that cells lacking Ssbp are hypersensitive to guanidine toxicity (Chernoff et al. 1999), leading to the suggestion that guanidine specifically targets co-translational protein folding or other processes involving Ssbp.
We have uncovered phenotypes caused by Ssbp deficiency that are unrelated to effects upon [ PSI +], but which affect the interpretation of [ PSI +]-related phenotypes. In agreement with earlier results (Chernoff et al. 1999), we find that Ssbp deficiency sensitizes cells to the toxic effects of guanidine hydrochloride. We further show that this enhanced sensitivity is due to an increase in guanidine uptake, which in turn allows Ssbp depleted cells to be cured of [ PSI +] using a much lower concentration of added guanidine. We also find that Ssbp depletion in [ psi –] cells modestly increases nonsense suppression and variably affects mRNA abundance, which may suggest roles for Ssbp in transcription or nonsense mRNA metabolism.
Materials and methods
Strains, media and genetic methods
The [ PSI +] strains 642 ( MATα kar1 ade2-1 SUQ5 his-11,153 leu2-3,1122 lys2 trp1Δ1 ura3-52) and GB1/2 (as 642, but ssb1:: HIS3 ssb2:: KanMX4) were constructed in this laboratory and are isogenic. The [ psi –] variants of these strains were obtained by guanidine curing (see below). Strain 642 was derived from a cross between strain RW1590 ( MATα kar1-1 ade2-1 SUQ5; obtained from Reed Wickner, NIH, Bethesda, Md.), and strain JN54 (Nelson et al. 1992) ( MAT a his3-11,15 leu2-3,112 lys2 trp1Δ1 ura3-52). SSB1and SSB2were disrupted in strain 642 with KanMX and HIS3, respectively, by transformation using DNA fragments amplified by PCR (Wach et al. 1994). Media were prepared as described by Sherman (1991), except that 1/2YPD contains 0.5% yeast extract, 2% peptone and 2% dextrose. YPAD is similar but contains 1% yeast extract and 400 mg/l adenine. YPD/G3 is 1/2YPD with 3 mM guanidine hydrochloride. Genetic methods used were described by Jung et al. (2000). The presence of [ PSI +] was verified by assaying for curing by guanidine, and by monitoring either its transmission by cytoplasmic transfer (cytoduction) or its dominant phenotype when crossed followed by guanidine treatment of the resulting diploids.
Guanidine curing
Guanidine curing of [ PSI +] was done as described previously (Jung et al. 2000). Briefly, cells from [ PSI +] cultures were grown at 30°C in liquid YPAD containing guanidine at the indicated concentrations. Cells were maintained in log phase by continued dilution into fresh guanidine-containing medium. Aliquots were removed periodically and spread on 1/2YPD plates at a density of approximately 400 colonies per plate. After incubating for 3 days at 30°C followed by 3 days at 25°C, the resulting colonies (~1,000/time point) were scored as [ PSI +] (white) or [ psi –] (red). Red/white sectored colonies were scored as [ PSI +].
Measurement of guanidine uptake
Cells (OD600=0.2) were shaken in YPAD in the presence of 1 mM guanidine and 10 μl of [14C]guanidine (0.1 mCi/ml, 55 mCi/mmol; American Radiolabeled Chemicals) at 30°C for 2 h. GB1/2 cells carrying pSSB1 were first grown overnight in medium lacking uracil, diluted into YPAD to an OD600 of 0.05, and grown to OD600=0.1 before adding guanidine. Equal numbers of cells were harvested, washed twice with 1 ml of water and suspended in 1 ml of TE (pH 8.0). Then 3 ml of scintillation fluid was added and [14C]guanidine was measured in an LKB scintillation counter.
β-Galactosidase (read through) assays
β-Galactosidase was assayed as described previously using the W4 series of plasmids (Bonetti et al. 1995). Briefly, 1-ml aliquots of cells, grown at 30°C to an OD600 value of 0.75, were washed in water and suspended in 1 ml of assay buffer. Cells were permeabilized by adding 50 μl of chloroform and 20 μl of 0.1% SDS. Reactions were initiated by the addition of 200 μl of ONPG (4 mg/ml) and stopped by the addition of 500 μl of 1 M sodium carbonate. β-Galactosidase activity was determined by measuring OD420 (Guarente 1983).
Northern analysis
Cells were grown under the inducing conditions used for β-galactosidase assays, and RNA was isolated using the FastRed RNA kit (Bio101). Aliquots (10 μg) of RNA were loaded on a 1% agarose gel containing 0.66 M formaldehyde, fractionated by electrophoresis, and transferred to Genescreen Plus (DuPont) nylon membrane as recommended by the manufacturer. The membrane was probed at high stringency using a 2.9-kb PvuII fragment of lacZ (Bonetti et al. 1995). The membrane was then stripped by boiling in 0.1xSSC/1% SDS, and probed with a 450-bp fragment of the yeast ACT1gene as a loading control. RNA amounts were quantified using a BASII phosphorimager (Fuji).
RT-PCR assays
Because the adenine biosynthetic pathway is repressed by excess adenine, total RNA was isolated from cells grown in 1/2YPD. RT-PCR assays were performed as described by Song et al. (2002). The ADE2-specific primers A (5´-TTCCTGTGGAAACAAGCCAGT-3´) and B (5´-GTGACGCAAGCATCAATGGT-3´) amplify a 490-bp segment of ade2-1extending from position 371 to 860 (where position 1 is the A of the initiator ATG). Competitor ADE2RNA, which lacks bases 391–445 and competes for the same ADE2-specific primers, was prepared using an RT-PCR competitor construction kit (Ambion). The Qiagen Quantitech RT-PCR kit was used with antisense specific primers to convert mixtures of total yeast RNA (750 ng) and serially diluted competitor RNA to cDNA, which was then co-amplified by PCR in the same reaction with the ADE2-specific primer pair. Products were resolved on 2% agarose gels and stained with ethidium bromide.
Results
Cells lacking Ssbp are hypersensitive to guanidine toxicity
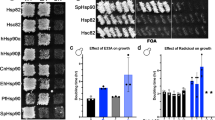
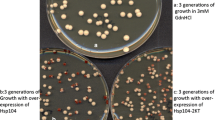
Cells lacking Ssbp ( ssb –) grow more slowly than wild-type cells, a phenotype that is likely to be due to a reduction in translation rates (Nelson et al. 1992). Whether [ PSI +] or [ psi –], the doubling times for our wild-type (642) and ssb – (GB1/2) strains in liquid YPAD at 30°C were 120 and 180 min, respectively. On plates without guanidine GB1/2 cells required incubation for two additional days to form colonies equivalent in size to those of the wild type. In agreement with observations made by others (Chernoff et al. 1999), our ssb – strain was hypersensitive to the toxic effects of guanidine. In liquid YPAD with 1 mM guanidine, the doubling times of strains 642 and GB1/2 were 110 and 260 min, respectively. Moreover, GB1/2 cells were unable to form colonies on 3 mM guanidine, a concentration routinely used to cure yeast prions (Fig. 1; see below). Guanidine concentrations of about 10 mM were required to inhibit growth of the isogenic wild-type strain to the same extent (data not shown). GB1/2 cells carrying a plasmid containing SSB1 (Ohba 1994) had wild type growth rate and guanidine sensitivity, indicating that the reduced growth and guanidine hypersensitivity were due to lack of Ssbp alone (data not shown).
Cells lacking Ssbp are hypersensitive to curing of [ PSI +] by guanidine
When yeast cells are grown in the presence of 3–5 mM guanidine replication of inheritable prion particles, or "seeds", is inhibited (Eaglestone et al. 2000). Non-replicating seeds are then randomly distributed among dividing cells until they become diluted to the point where additional cell division gives rise to [ psi –] cells. The length of the lag in appearance of [ psi –] cells thus provides an estimate of the number of [ PSI +] seeds present per cell before the addition of guanidine. Typical [ PSI +] cells have an average of about 60 seeds per cell and thus require four to five cell divisions in the presence of guanidine before [ psi –] cells appear.
To determine whether cells lacking Ssbp were also hypersensitive to the [ PSI +] curing effect of guanidine, strains 642 and GB1/2 were grown on 1/2YPD medium containing 0.25 mM, 0.5 mM, 0.75 mM or 1 mM guanidine. Cells from small colonies (1–2 mm diameter) were suspended in water and spread onto 1/2YPD, and the resulting colonies were scored for the presence of [ PSI +]. For strain 642, [ PSI +] was completely stable on all plates. [ PSI +] was also stable in GB1/2 cells on plates containing 0.25 mM guanidine. However, among GB1/2 cells grown on plates with 0.5 mM guanidine, 50–70% were [ psi –]. Although the cured cells were less red than 642 [ psi –] cells (Fig. 1, see below), they were confirmed to be [ psi –] by transfer of cytoplasm into a [ psi –] strain. Thus Ssbp deficiency causes hypersensitivity to curing of [ PSI +] by guanidine.
The rate of [ PSI +] loss more accurately defines sensitivity to the curing effect of guanidine, and can also provide an estimate of the number of transmissible prion particles, or [ PSI +] "seeds", present per cell (Eaglestone et al. 2000). Guanidine was added to liquid cultures, and the fraction of cells in the population that remained [ PSI +] was monitored as the cells divided. For wild-type cells, 1 mM guanidine did not cure [ PSI +] and 3 mM guanidine had the maximal curing effect (Fig. 2A). For strain GB1/2, 0.25 mM guanidine had no effect on [ PSI +] but curing kinetics very similar to those seen in wild type cells grown with 3 mM guanidine were obtained using 0.5 mM, 0.75 mM and 1.0 mM guanidine (Fig. 2B). Thus, for yeast cells lacking Ssbp a wild type curing profile was seen with about sixfold less added guanidine. Nevertheless, the lag before the appearance of [ psi –] cells, which is a direct reflection of the starting number of seeds per cell, was similar for 642 and GB1/2. Thus, [ PSI +] cells lacking Ssbp are capable of maintaining a normal number of [ PSI +] seeds per cell.
Loss of [PSI +] after addition of guanidine. A Percentage of wild-type (WT, strain 642) [PSI +] cells present at various times after the addition of guanidine to final concentrations of 0–1 mM (circles), 2 mM (squares), 3 mM (triangles) at 30°C, or 1 mM at 20°C (diamonds). B Similar plot for strain GB1/2 (ssb –) grown at 30°C with no guanidine (circles), 0.25 mM (squares), 0.5/0.75 mM (triangles), 1 mM (diamonds)
Because of the slow growth of GB1/2 cells, we tested whether prolonged generation time might enhance the curing effect of guanidine in wild-type cells. We did this by assaying the effects of 1 mM guanidine on strain 642 at 20°C; at this temperature this strain grows at a rate (~200 min/cell division) similar to that of GB1/2 in 1 mM guanidine at 30°C. Although less efficient than 2 mM guanidine at 30°C, 1 mM guanidine cured strain 642 of [ PSI +] at 20°C (Fig. 2A). We also observed infrequent (~0.1%) loss of [ PSI +] from untreated 642 cells during prolonged growth at 20°C. Thus, decreasing the growth temperature results in increased sensitivity to curing of [ PSI +] by guanidine and spontaneous mitotic loss of [ PSI +].
Hypersensitivity to guanidine is caused by increased uptake of the compound
The difference in guanidine sensitivity could easily be explained if the GB1/2 cells accumulated more guanidine intracellularly than do wild-type cells. We tested this hypothesis by using radiolabeled guanidine in cultures containing a final concentration of 1 mM unlabeled guanidine. Indeed, GB1/2 cells were found to accumulate approximately eightfold more of the label than the wild type (Table 1). GB1/2 cells with a single-copy plasmid containing the SSB1gene had a level of guanidine uptake similar to that of wild-type cells, indicating that the increased uptake in GB1/2 cells was due to lack of Ssbp alone. In cultures exposed to 0.5 mM guanidine intracellular guanidine concentrations were about half those attained when 1 mM guanidine was used, indicating that guanidine uptake was essentially linear across this concentration range (data not shown). Wild type cells took up about twice as much guanidine when grown at 20°C compared with 30°C (Table 1), which is consistent with the finding that they are cured of [ PSI +] in the presence of lower concentrations of added guanidine at low temperature. Wild-type cells were also more sensitive to the toxic effects of guanidine at 20°C, producing fewer viable cells per generation in the presence of 3 mM guanidine at this temperature (data not shown). Thus, differences in accumulation of intracellular guanidine adequately explain all of the guanidine hypersensitivity phenotypes.
Assuming the volume of a haploid yeast cell to be 70 μm3 (Sherman 1991), we estimated the intracellular concentration of guanidine in strain 642 grown at 30°C in 1 mM guanidine to be 19 mM. GB1/2 cells contain about 150 mM guanidine when grown under the same conditions.
Mutation of ERG6but not of genes for ABC transporters causes guanidine hypersensitivity
Possible causes of the increased guanidine accumulation in cells lacking Ssbp include defects in membrane pumping systems or in the membranes themselves. An interaction has been demonstrated genetically between the Ssbp Hsp40 co-chaperone Zuo1p (Yan et al. 1998) and the pleiotropic drug resistance factor Pdr13p, which is a protein with weak homology to Hsp70 that is involved in the transcriptional regulation of ATP-binding-cassette (ABC) proteins (Hallstrom et al. 1998; Michimoto et al. 2000). ABC proteins are a universally conserved family whose members include membrane transporters that have been implicated in metal and multidrug resistance (Higgins 2001). Taking a candidate gene approach we assayed the guanidine sensitivity of 16 mutants, each lacking one ABC protein (a total of 29 ABC proteins are encoded in the yeast genome). Except for the mating pheromone exporter encoded by STE6, those tested included all of the full length transmembrane homologs that are implicated in metal and mutlidrug resistance (Decottignies et al. 1997). None of the mutants tested was significantly more sensitive to guanidine than our wild-type strain (data not shown). Thus, it appears unlikely that Ssbp deficiency indirectly increases guanidine uptake by affecting ABC membrane transporters.
We also tested the guanidine sensitivity of an erg6 mutant, which displays an altered membrane lipid composition and is hypersensitive to several toxic compounds. The erg6mutant was considerably more sensitive to guanidine than wild-type cells, forming only micro-colonies on 3 mM guanidine after 5–7 days at 30°C (data not shown). It was slightly less sensitive than strain GB1/2, which did not form colonies on 3 mM guanidine. In agreement with our other results, the level of intracellular guanidine was threefold higher in the erg6mutant than in wild type (data not shown). Prompted by this result, we examined the lipid composition of our experimental strains. Lipids (sterols) of wild-type and GB1/2 cells were extracted by alkaline saponification of whole cells (Parks et al. 1985) and analyzed by reversed-phase HPLC (Xu et al. 1988). Both strains had essentially identical lipid profiles, suggesting that it is also unlikely that Ssbp depletion increases guanidine accumulation by altering membrane permeability.
Deletion of SSB increases nonsense suppression and affects nonsense mRNA abundance
The presence of [ PSI +] allows the weak suppressor tRNA SUQ5 ( SUP16) to suppress the UAA nonsense allele ade2-1in our strains (Cox 1965). Non-suppressed [ psi –] cells require adenine for growth and form red colonies when adenine is limiting (i.e. on 1/2YPD medium) due to the accumulation of a pigmented form of the substrate of Ade2p (Silver et al. 1969). Suppressed [ PSI +] cells grow in the absence of adenine, and are white on 1/2YPD. Any condition that affects the stability or translation efficiency of the mRNA can affect nonsense suppression and influence color development or growth in the absence of adenine.
GB1/2 [ psi –] colonies are less red than those of strain 642, and [ PSI +] colonies of GB1/2 also appeared whiter than 642 [ PSI +] colonies (Fig. 1). These differences probably reflect increased suppression of ade2-1, which could be achieved primarily in two ways: by increasing the frequency of read through of the ochre nonsense mutation within the ade2-1gene, or by increasing the steady-state level of ade2-1mRNA, increasing the number of transcripts in which read through may occur. We quantified translation read through by using reporter plasmids encoding E. coli β-galactosidase fused to a 6-residue leader peptide containing a tryptophan codon (UGG) or one of the three stop codons at the fourth position (Bonetti et al. 1995). To avoid contributions to nonsense suppression by [ PSI +] (Firoozan et al. 1991), [ psi –] cells were used. The frequency of read through of the UAA codon was higher than that of the other stop codons in both wild-type and GB1/2 transformants (Table 2), which was expected because of the presence of the weak SUQ5ochre-suppressing tRNA. Read through of all three stop codons was reproducibly 50% higher in GB1/2 cells, despite ssb – cells having a much-reduced number of translating ribosomes compared to wild-type cells (Nelson et al. 1992). Because SSB expression is reduced when cells are grown on galactose compared to glucose (Norbeck et al. 1997), we repeated the experiment using β-estradiol as an indirect inducer of the GAL1promoter in the W4 plasmids (Louvion et al. 1993). We saw the same reproducible 50% increase in read through of nonsense codons (Table 2).
To determine if an effect of Ssbp depletion on mRNA abundance contributed to this increased nonsense suppression, we measured steady-state levels of lacZ mRNA in the galactose-induced cells that were used to quantify read through. GB1/2 cells contained three-fold more UAA nonsense mRNA and five-fold more UAG and UGA mRNA compared to wild type cells (Fig. 3A). Results of RT-PCR analysis of lacZ mRNA qualitatively reproduced this difference (data not shown). The GB1/2 cells also had about twice as much of the UGG (sense) transcript relative to the actin mRNA control, suggesting that at least part of the increased abundance of all of the mRNAs in this strain was due to increased transcription. This elevated abundance may explain the increased nonsense suppression of lacZ mRNA in GB1/2 cells.
Relative abundance of nonsense mRNA is affected in GB1/2 cells. A Northern analysis of total RNA from wild type (642) and ssb – (GB1/2) cells. The ratio of lacZ mRNA to ACT1mRNA is indicated. The relative abundance of nonsense mRNA is indicated as a percentage of UGG mRNA. B RT-PCR analysis of ade2-1mRNA. Mixtures of 750 ng of total yeast RNA and increasing amounts of competitor (comp.) RNA were reverse transcribed and amplified by PCR. The ratio of PCR products obtained reflects the ratio of ade2-1mRNA to competitor mRNA. The calculated copy numbers of competitor sequence included in the reactions are indicated
As a more physiologically relevant assay, we compared the levels of ade2-1mRNA in wild-type and GB1/2 cells by the more sensitive competitive RT-PCR assay (see Materials and methods). In contrast to the results with the highly expressed galactose-induced lacZ mRNA, we found that the chromosomally expressed ade2-1mRNA was about six-fold less abundant in GB1/2 cells (Fig. 3B).
Discussion
We show here that yeast mutants that lack Ssb proteins are hypersensitive to the toxic and prion-curing effects of guanidine. We further show that this hypersensitivity is due to increased uptake of guanidine by these cells, which explains all of the guanidine phenotypes associated with Ssb protein deficiency. In addition, when wild-type cells were exposed to amounts of guanidine expected to produce similar intracellular concentrations as in ssb – cells, similar growth-inhibitory effects were seen. Moreover, erg6mutants, which show a level of guanidine sensitivity intermediate between those of wild-type and ssb – cells, accumulate an intermediate amount of intracellular guanidine. Thus, our results suggest that guanidine toxicity is not due to specific targeting of processes that involve Ssb proteins. The basis for guanidine toxicity to yeast remains unknown, but it is distinct from its prion-curing effect, which is due to inactivation of the non-essential Hsp104 (Jung et al. 2002). There is evidence to suggest that guanidine is a non-specific inhibitor of ATPase activity, and the collective effects of inhibition of many cellular ATPases may account for its toxicity (Pfister and Wimmer 1999).
In wild-type cells the intracellular concentration of guanidine was about twenty-fold higher than that outside the cells. It is possible that this accumulation is due to the activity of membrane transporters, and that ssb – cells accumulated even more guanidine because of indirect effects on a membrane pumping system or on membrane integrity. Our finding that single deletions of all of the predicted membrane transporter class of ABC proteins did not affect sensitivity to guanidine suggests that Ssbp depletion does not affect ABC transporter expression or function. It remains possible, however, that a simultaneous and cumulative effect of Ssbp depletion on the function of several ABC transporters underlies the guanidine hypersensitivity, or that Ssbp depletion indirectly perturbs the function of drug:H+-antiporters, another class of membrane transporters implicated in drug resistance. Although we did find that altered membrane lipid composition resulting from the erg6mutation was associated with guanidine hypersensitivity, the lipid profiles of wild type and ssb – cells were essentially identical. Therefore, the increased guanidine accumulation by ssb – cells is apparently not due to membrane defects. A plausible alternative would be that guanidine is being retained by binding to intracellular targets rather than being actively imported against a gradient, and that Ssbp depletion somehow affects this retention. A genomic approach may be the best way to identify the genes involved guanidine toxicity and how Ssbp depletion affects them.
The lag before appearance of [PSI –] cells in guanidine-treated cultures was similar for wild type and Ssb mutant cells, showing that both strains maintain similar numbers of prion seeds per cell and suggesting that deletion of SSB does not affect the process of [ PSI +] seed regeneration. Because wild-type cells accumulate approximately twice as much guanidine at 20°C as at 30°C, it might be expected that similar curing kinetics would be observed when 2 mM at 30°C and 1 mM at 20°C were used. However, although curing occurs at a comparable rate once [ psi –] cells begin to appear, the number of cell divisions required to produce [ psi –] cells is greater in 1 mM guanidine at 20°C. Even though [ PSI +] was somewhat less stable in wild-type cells at 20°C, this increased lag suggests that wild-type cells had more [ PSI +] seeds at this lower temperature. These inconsistencies probably reflect a shift in the balance of the abundance of different protein chaperones known to affect [ PSI +] stability at the lower temperature. Moreover, although wild-type cells exposed to 2 mM guanidine at 30°C and 1 mM guanidine at 20°C accumulated the compound to similar concentrations intacellularly, at 20°C they were more sensitive to the toxic effect of guanidine. Reduced growth temperature thus had pleiotropic effects with respect to both [ PSI +] propagation and guanidine toxicity.
The increased read through of nonsense lacZ mRNA in cells lacking Ssbp is at least partially due to its increased abundance, which probably resulted from elevated transcription. Also, the relative steady-state abundance of nonsense lacZ mRNA in wild-type cells (15–20% of the sense message) may be considered high in comparison with those of other mRNAs with early nonsense mutations, suggesting that the large amounts of galactose-induced transcripts may strain the capacity of the nonsense-mediated decay (NMD) pathway to eliminate nonsense mRNAs. In contrast to lacZ mRNA, the abundance of the chromosomally expressed ade2-1nonsense mRNA was lower in ssb – cells, raising the possibility that altered translation in these cells may affect NMD processes, or that Ssbp is itself involved in nonsense-mediated decay. By any scenario, if the color differences between wild type and ssb – colonies truly reflect differences in nonsense suppression, then the effect of Ssbp depletion on SUQ5-mediated read through of the UAA nonsense codon in the weakly expressed ade2-1mRNA was greater than that on the abundant lacZ transcripts. Such differences could arise from effects related to the nucleotide sequence context of the nonsense codon in the ade2-1and lacZ mRNAs (Bonetti et al. 1995; Ruiz-Echevarria et al. 1998).
References
Bonetti B, Fu L, Moon J, Bedwell DM (1995) The efficiency of translation termination is determined by a synergistic interplay between upstream and downstream sequences in Saccharomyces cerevisiae. J Mol Biol 251:334–345
Chacinska A, Szczesniak B, Kochneva-Pervukhova NV, Kushnirov VV, Ter-Avanesyan MD, Boguta M (2001) Ssb1 chaperone is a [PSI+] prion-curing factor. Curr Genet 39:62–67
Chernoff YO, Lindquist SL, Ono B, Inge-Vechtomov SG, Liebman SW (1995) Role of the chaperone protein Hsp104 in propagation of the yeast prion-like factor [psi+]. Science 268:880–884
Chernoff YO, Newnam GP, Kumar J, Allen K, Zink AD (1999) Evidence for a protein mutator in yeast: role of the Hsp70-related chaperone Ssb in formation, stability, and toxicity of the [PSI] prion. Mol Cell Biol 19:8103–8112
Cox BS (1965) [PSI+] a cytoplasmic suppressor of super-suppressor in yeast. Heredity 20:505–521
Craig E, Ziegelhoffer T, Nelson J, Laloraya S, Halladay J (1995) Complex multigene family of functionally distinct Hsp70s of yeast. Cold Spring Harb Symp Quant Biol 60:441–449
Decottignies A, Goffeau A (1997) Complete inventory of the yeast ABC proteins. Nat Genet 15:137–145
Eaglestone SS, Ruddock LW, Cox BS, Tuite MF (2000) Guanidine hydrochloride blocks a critical step in the propagation of the prion-like determinant [PSI(+)] of Saccharomyces cerevisiae. Proc Natl Acad Sci USA 97:240–244
Ferreira PC, Ness F, Edwards SR, Cox BS, Tuite MF (2001) The elimination of the yeast [PSI+] prion by guanidine hydrochloride is the result of Hsp104 inactivation. Mol Microbiol 40:1357–1369
Firoozan M, Grant CM, Duarte JA, Tuite MF (1991) Quantitation of readthrough of termination codons in yeast using a novel gene fusion assay. Yeast 7:173–183
Guarente L (1983) Yeast promoters and lacZ fusions designed to study expression of cloned genes in yeast. Methods Enzymol 101:181–191
Hallstrom TC, Katzmann DJ, Torres RJ, Sharp WJ, Moye-Rowley WS (1998) Regulation of transcription factor Pdr1p function by an Hsp70 protein in Saccharomyces cerevisiae. Mol Cell Biol 18:1147–1155
Higgins CF (2001) ABC transporters: physiology, structure and mechanism—an overview. Res Microbiol 152:205–210
Jones G, Masison DC (2003) S. cerevisiae Hsp70 mutations affect [ PSI +] prion propagation and cell growth differently and implicate Hsp40 and TPR co-chaperones in impairment of [ PSI +]. Genetics 163:495–506
Jung G, Masison DC (2001) Guanidine hydrochloride inhibits Hsp104 activity in vivo: a possible explanation for its effect in curing yeast prions. Curr Microbiol 43:7–10
Jung G, Jones G, Wegrzyn RD, Masison DC (2000) A role for cytosolic Hsp70 in yeast [ PSI +] prion propagation and [ PSI +] as a cellular stress. Genetics 156:559–570
Jung G, Jones G, Masison DC (2002) Amino acid residue 184 of yeast Hsp104 chaperone is critical for prion-curing by guanidine, prion propagation, and thermotolerance. Proc Natl Acad Sci USA 99:9936–9941
Kushnirov VV, Kryndushkin DS, Boguta M, Smirnov VN, Ter-Avanesyan MD (2000) Chaperones that cure yeast artificial [PSI+] and their prion-specific effects. Curr Biol 10:1443–1446
Louvion JF, Havaux-Copf B, Picard D (1993) Fusion of GAL4-VP16 to a steroid-binding domain provides a tool for gratuitous induction of galactose-responsive genes in yeast. Gene 131:129–134
Michimoto T, Aoki T, Toh-e A, Kikuchi Y (2000) Yeast Pdr13p and Zuo1p molecular chaperones are new functional Hsp70 and Hsp40 partners. Gene 257:131–137
Moriyama H, Edskes HK, Wickner RB (2000) [URE3] prion propagation in Saccharomyces cerevisiae: requirement for chaperone Hsp104 and curing by overexpressed chaperone Ydj1p. Mol Cell Biol 20:8916–8922
Nelson RJ, Ziegelhoffer T, Nicolet C, Werner-Washburne M, Craig EA (1992) The translation machinery and 70 kd heat shock protein cooperate in protein synthesis. Cell 71:97–105
Ness F, Ferreira P, Cox BS, Tuite MF (2002) Guanidine hydrochloride inhibits the generation of prion "seeds" but not prion protein aggregation in yeast. Mol Cell Biol 22:5593–5605
Newnam GP, Wegrzyn RD, Lindquist SL, Chernoff YO (1999) Antagonistic interactions between yeast chaperones Hsp104 and Hsp70 in prion curing. Mol Cell Biol 19:1325–1333
Norbeck J, Blomberg A (1997) Two-dimensional electrophoretic separation of yeast proteins using a non-linear wide range (pH 3–10) immobilized pH gradient in the first dimension; reproducibility and evidence for isoelectric focusing of alkaline (pI>7) proteins. Yeast 13:1519-1534
Ohba M (1994) A 70-kDa heat shock cognate protein suppresses the defects caused by a proteasome mutation in Saccharomyces cerevisiae. FEBS Lett 351:263–266
Parks LW, Bottema CD, Rodriguez RJ, Lewis TA (1985) Yeast sterols: yeast mutants as tools for the study of sterol metabolism. Methods Enzymol 111:333–346
Pfister T, Wimmer E (1999) Characterization of the nucleoside triphosphatase activity of poliovirus protein 2C reveals a mechanism by which guanidine inhibits poliovirus replication. J Biol Chem 274:6992–7001
Pfund C, Lopez-Hoyo N, Ziegelhoffer T, Schilke BA, Lopez-Buesa P, Walter WA, Wiedmann M, Craig EA (1998) The molecular chaperone Ssb from Saccharomyces cerevisiae is a component of the ribosome-nascent chain complex. EMBO J 17:3981–3989
Prusiner SB (1982) Novel proteinaceous infectious particles cause scrapie. Science 216:136–144.
Ruiz-Echevarria MJ, Gonzalez CI, Peltz SW (1998) Identifying the right stop: determining how the surveillance complex recognizes and degrades an aberrant mRNA. EMBO J 17:575–589
Sherman F (1991) Getting started with yeast. Methods Enzymol 194:3–21
Silver JM, Eaton NR (1969) Functional blocks of the ad-1and ad-2mutants of Saccharomyces cerevisiae. Biochem Biophys Res Commun 34:301–305
Song Y, Azakami H, Shamima B, He J, Kato A (2002) Different effects of calnexin deletion in Saccharomyces cerevisiae on the secretion of two glycosylated amyloidogenic lysozymes. FEBS Lett 512:213–217
Stansfield I, Jones KM, Kushnirov VV, Dagkesamanskaya AR, Poznyakovski AI, Paushkin SV, Nierras CR, Cox BS, Ter-Avanesyan MD, Tuite MF (1995) The products of the SUP45 (eRF1) and SUP35genes interact to mediate translation termination in Saccharomyces cerevisiae. EMBO J 14:4365–4373
Tuite MF, Mundy CR, Cox BS (1981) Agents that cause a high frequency of genetic change from [psi+] to [psi-] in Saccharomyces cerevisiae. Genetics 98:691–711
Wach A, Brachat A, Pohlmann R, Philippsen P (1994) New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast 10:1793–1808
Werner-Washburne M, Becker J, Kosic-Smithers J, Craig EA (1989) Yeast Hsp70 RNA levels vary in response to the physiological status of the cell. J Bacteriol 171:2680–2688
Wickner RB (1994) [URE3] as an altered URE2 protein: evidence for a prion analog in Saccharomyces cerevisiae. Science 264:566–569.
Xu SH, Norton RA, Crumley FG, Nes WD (1988) Comparison of the chromatographic properties of sterols, select additional steroids and triterpenoids: gravity-flow column liquid chromatography, thin-layer chromatography, gas-liquid chromatography and high-performance liquid chromatography. J Chromatogr 452:377–398
Yan W, Schilke B, Pfund C, Walter W, Kim S, Craig EA (1998) Zuotin, a ribosome-associated DnaJ molecular chaperone. EMBO J 17:4809–4817
Zhouravleva G, Frolova L, Le Goff X, Le Guellec R, Inge-Vechtomov S, Kisselev L, Philippe M (1995) Termination of translation in eukaryotes is governed by two interacting polypeptide chain release factors, eRF1 and eRF3. EMBO J 14:4065–4072
Acknowledgements
We thank Will Prinz (NIH, Bethesda, MD) for strains and lipid composition analysis, and David Bedwell (University of Alabama, Birmingham) and Masayuki Ohba (Tokyo, Japan) for providing plasmids. We thank Kathy West, Adam West, Damien Hall, and Phil Lee for valuable comments on the manuscript
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. P. Hollenberg
Rights and permissions
About this article
Cite this article
Jones, G.W., Song, Y. & Masison, D.C. Deletion of the Hsp70 chaperone gene SSB causes hypersensitivity to guanidine toxicity and curing of the [ PSI +] prion by increasing guanidine uptake in yeast. Mol Gen Genomics 269, 304–311 (2003). https://doi.org/10.1007/s00438-003-0838-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-003-0838-y