Abstract
Cystic echinococcosis caused by the larval stages of Echinococcus granulosus sensu lato s.l is endemic in Turkey with a high public health impact particularly in rural areas. The aim of this study was to investigate the genetic variation and population structure of E. granulosus s.s using metacestode isolates removed from surgically confirmed patients originating from several regions in Turkey and to investigate the occurrence of autochthonous transmission. Using DNA extracted from a total of 46 human-derived CE isolates, we successfully analysed an 827-bp fragment within the cox1 mitochondrial gene and confirmed the causative agent of human cystic echinococcosis in patients included in this study to be Echinococcus granulosus s.s (G1 and G3 genotypes). The haplotype parsimony network consisted of 28 haplotypes arranged within three main clusters and the neutrality indices were both negative and significant indicating negative selection or population expansion. The assessment carried out in this study using GenBank nucleotide sequence data from Turkey for sheep and cattle hosts demonstrated the importance of autochthonous transmission with sheep, cattle and humans harbouring the same haplotypes. Further studies are required to investigate the biological significance, if any, of E. granulosus s.s haplotypes and the genetic variability of CE from human patients using longer nucleotide sequences and a larger sample set.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cystic echinococcosis (CE) caused by the larval stages of Echinococcus granulosus sensu lato s.l is one of the most important, globally distributed zoonotic diseases. E. granulosus s.l is a cryptic species complex that includes Echinococcus granulosus sensu stricto (s.s) (genotypes G1–G3), Echinococcus felidis, Echinococcus equinus (G4), Echinococcus ortleppi (G5) and Echinococcus canadensis (G6/G7, G8, G10) (Nakao et al. 2007; Thompson 2008; Hüttner et al. 2008; Romig et al. 2015). E. granulosus s.l circulates among domestic and/or wild carnivores and herbivores, especially dogs and livestock animals (sheep, goats, cattle, camels, pigs) (Thompson 2008). Humans are accidental hosts infected through the ingestion of eggs excreted in dog faeces but do not contribute to the perpetuation of the life cycle.
High CE incidence has been reported from several countries neighbouring Turkey. For example, incidence rates of 5.71 (Jordanova et al. 2015), 4.5 (Abdulhameed et al. 2018) and 1.18–3 (Rokni 2008; Tavakoli et al. 2008) cases per 100,000 inhabitants were documented in recent years from Bulgaria, Basrah (Iraq) and Iran respectively. CE is endemic in Turkey with a high public health impact in rural areas where the local population is actively engaged in animal breeding and remains in close contact with ruminants and dogs (Altintas 2003). Although CE is a notifiable disease in Turkey, data on human infection is principally obtained from hospital records. The incidence rate of CE from 1990 to 2005 was estimated to be 0.8–2 per 100,000 (Yolasıgmaz et al. 2006; Altintas 2008).
Hydatidosis in ruminants is widespread and has been reported from many regions in Turkey (Altintas 2003; Umur 2003; Yildiz and Gurcan 2003; Yildiz and Tunçer 2005; Esatgil and Tuzer 2007; Utuk et al. 2008; Kose and Sevimli 2008; Simsek and Eroksuz 2009; Beyhan and Umur 2011). Meat production losses due to CE for 2008 were US$89.2 million and nationwide annual losses attributed to CE in sheep, cattle and goats were estimated to be US$ 54.1, 32.4 and 2.7 million respectively (Sariozkan and Yalcin 2009). Losses relating to offal condemnation estimated for a 6-month period during 2012 in Bursa Province were US$4015 and US$12,321 for sheep and cattle respectively (Yibar et al. 2015). The domestic dog is presumed to be the most important definitive host and is responsible for contamination of the environment with Echinococcus eggs. Variable prevalence rates of canine echinococcosis from different regions following necropsy (reviewed by Altintas 2003), coproantigen ELISA (Guzel et al. 2008) and parasitological faecal examination (Öter et al. 2011) have been recorded. Recent reports have used molecular methods to identify E. granulosus in dog faeces from Turkey (Acioz 2008; Utuk et al. 2008; Kuru et al. 2013; Öge et al. 2017).
During the last decade, several reports on the molecular confirmation of the causative agents of CE in ruminants have been published (Utuk et al. 2008; Šnàbel et al. 2009; Simsek and Eroksuz 2009; Beyhan and Umur 2011; Simsek et al. 2011a; Eroğlu et al. 2016). Using various DNA-based molecular approaches, these studies have shown that E. granulosus s.s (G1 and G3 genotypes) is the most prevalent species affecting sheep, cattle, goats, camels and pigs in Turkey. Similarly, several reports on the molecular identification of human CE causative agents in Turkey confirmed the predominance of E. granulosus s.s (G1 and G3 genotypes) (Utuk et al. 2008; Ergin et al. 2010; Simsek et al. 2011b; Eryildiz and Şakru 2012; Bakal et al. 2015; Eroğlu et al. 2016). In addition, E. canadensis G6 and G7 were reported from humans and sheep (Šnàbel et al. 2009; Simsek et al. 2011b).
The aim of this study was to investigate the genetic variation and population structure of E. granulosus s.s using metacestode isolates removed from surgically confirmed patients originating from several regions in Turkey. This work also involves the investigation of the occurrence of autochthonous transmission based on the comparison of our human CE nucleotide sequences with those reported from Turkish ruminants and deposited in GenBank.
Materials and methods
Ethics statement
The current study was approved by the Ethical Committee of the Faculty of Medicine, Hacettepe University, Ankara, Turkey (GO 14/293-37).
Sample collection, DNA extraction, PCR amplification and sequencing
Hydatid material (cyst fluid and/or germinal layer) retrieved between November 2014 and December 2016 from 46 percutaneously treated CE patients was included in this study. Demographic features such as sex, age and origin of patients were retrieved from patients’ clinical records.
Genomic DNA extracted from cyst material using the Qiagen DNeasy Blood and Tissue DNA Extraction Kit (Qiagen, Hilden, Germany) was used to amplify an 828-bp fragment within the cytochrome c oxidase subunit 1 (cox 1) mitochondrial gene using primers F/COI (5′ TTGAATTT-GCCACGTTTGAATGC 3′) and R/COI (5′ GAACCTAACGACATAACATAATGA 3′) (Nakao et al. 2000). Amplified PCR products were visualised under ultraviolet light (Syngene G: Box gel documentation system, Cambridge Biosciences) following electrophoresis on 1.5% (w/v) ethidium bromide-stained agarose gels (Cleaver Scientific Limited) and purified using QIAquick PCR Purification Kit (Qiagen) according to the manufacturer’s instructions. Bi-directional commercial sequencing was achieved using the PCR primers (Macrogen EZ-Sequence, Amsterdam, The Netherlands). FinchTV viewer (Geospiza, Seattle, WA, USA) was used to examine chromatograms and ascertain the quality of generated nucleotide sequences. A BLAST algorithm search (http://www.ncbi.nlm.nih.gov/BLAST/) was performed to compare our sequenced data with that present on the NCBI database. In addition, an alignment of a 366-bp fragment of our generated sequences with reference sequences published for E. granulosus G1-G3 genotypes (Bowles et al. 1992) was also conducted.
Nucleotide sequence data analysis
This was carried out using published methodology (Boufana et al. 2014). In brief, nucleotide DNA sequences were aligned using ClustalX2 software (Larkin et al. 2007) and transported into DnaSP 5 (Librado and Rozas 2009) to determine DNA polymorphisms and haplotype distributions. The best-fit nucleotide substitution model using the Akaike Information Criterion (AIC) (TrN + 1) was obtained through the application of Modeltest 3.7 (Posada and Crandall 1998) executed in Paup 4.0 (Swofford 1998). Population diversity indices such as haplotype numbers (hn), haplotype (hd) and nucleotide diversities (πd) were calculated in Arlequin version 3.5. (Excoffier and Lischer 2010) which was also used to test for population expansion or bottleneck using Tajima’s D (Tajima 1989) and Fu’s Fs (Fu 1997). Hapview (Salzburger et al. 2011) was used to generate parsimony haplotype networks and maximum likelihood trees were constructed by DNAML program in PHYLIP (Felsenstein 1989).
A total of 59 cox 1 mitochondrial nucleotide sequences of E. granulosus s.s (G1 genotype) from Turkish intermediate hosts (KU925352–KU925384, KU925386–KU925387, KU925389–KU925394, KU925396–KU925398, KU925400–KU925412) (sheep n = 22; cattle = 35) in addition to two shared sequences (KU925385, KU925395; sheep and cattle) (Kinkar et al. 2016) deposited in GenBank (http://www.ncbi.nlm.nih.gov) were also included in this study. These were aligned with the human CE sequences generated in this study, trimmed to equal lengths (827 bp) and used to create a new dataset (n = 105) for a human/animal parsimony haplotype analysis to investigate autochthonous transmission.
Results
Demographics of patients
Of the 46 patients included in this study, 56.5% (26/46) were from central Anatolia (Ankara, n = 16; Aksaray, n = 2; Çankırı, n = 2; Eskișehir, n = 1; Karaman, n = 1; Kırıkkale, n = 1; Sivas, n = 1; Yozgat, n = 2) (Fig. 1). The remaining patients originated from eastern Anatolia (Ağrı, n = 1; Bitlis, n = 2; Hakkari, n = 1; Kars, n = 1; Malatya, n = 1; Van, n = 1), the Black Sea Region (Corum, n = 3; Tokat, n = 1), Marmara (Istanbul, n = 1; Balikesir, n = 2), south-eastern Anatolia (Şanliurfa, n = 4), the Mediterranean region (Hatay, n = 1) and Aegean (Kütahya, n = 1). Our sample consisted of 29 female (63%) and 17 (36.9%) male patients with a mean and median age of 40.4 and 40.5 respectively (range 4–78 years).
Echinococcus species
Using DNA extracted from a total of 46 human-derived CE isolates, we successfully analysed an 827-bp fragment within the cox1 mitochondrial gene. The use of BLAST algorithm confirmed the causative agent of human cystic echinococcosis in patients included in this study to be E. granulosus s.s. Most isolates (39/46, 84.8%) belonged to E. granulosus G1 genotype whereas the remaining 7 isolates (15.2%) were identified as E. granulosus G3 genotype (Bowles et al. 1992).
DNA polymorphisms, diversity indices and parsimony network
Within the 827 bp analysed nucleotide sequences, we detected a total of 28 polymorphic sites of which 7 were parsimony-informative. No indels or gaps were observed. The overall haplotype and nucleotide diversities are shown in Table 1. The neutrality indices were both negative and significant (Tajima’s D − 2.1263; p < 0.01; Fu’s Fs − 26.80; p < 0.001) indicating negative selection or population expansion.
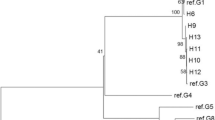
The haplotype parsimony network consisted of 28 haplotypes arranged within three main clusters separated by one to three mutational steps (Fig. 2). The E. granulosus s.s dominant cluster with TUK04 as its main haplotype had 27/46 human CE isolates which were identified in patients from central Anatolia (n = 15), the Black Sea Region (n = 3), Marmara (n = 2), eastern Anatolia (n = 5), south-eastern Anatolia (n = 1) and the Mediterranean region (n = 1). The nucleotide sequences of haplotype TUK04 were 100% identical to those of the globally distributed ‘ancestral’ E. granulosus haplotype (Accession no. AB491414) first described by Nakao et al. (2010). TUK20 was the main haplotype in the second most dominant cluster (12/46) which gave 100% identity to E. granulosus G1 (Accession no. KU925431) and encompassed isolates from patients from central Anatolia (n = 7), the Black Sea Region (n = 1), eastern Anatolia (n = 1), south-eastern Anatolia (n = 2) and the Aegean Region (n = 1). It was also 100% identical to the second most common haplotype recently reported from Kurdistan, Iraq (Accession no. MF004305) (Hassan et al. 2017). The third cluster within this network of E. granulosus s.s (haplotypes TUK24–TUK28) (7/46) occurred in CE patients from central Anatolia (n = 4), Marmara (n = 1), south-eastern Anatolia (n = 1) and eastern Anatolia (n = 1) and gave a 99% homology to E. granulosus G3 genotype (Accession no. KY766903) (Fig. 2).
We used nucleotide sequences of E. granulosus s.s (G1 genotype) derived from sheep (n = 24) and cattle (n = 37) hosts originating from Erzurum and Elazig provinces in eastern Turkey (Kinkar et al. 2017) to investigate autochthonous transmission through the generation of a network to include sequences from these animals and those generated from human CE patients in this study. This network consisted of 39 haplotypes with 18/24 (75%) (sheep) and 29/37 (cattle) (78.4%) isolates respectively sharing 7/39 haplotypes with the human CE isolates from this study (data not shown).
Discussion
In the current study, we used CE material derived from patients originating from seven Turkish regions from which human CE was previously reported. A retrospective study based on the examination of hospital, health directorship and Ministry of Health records from 2001 to 2005 identified 14,789 CE cases from central Anatolia (38.57%), Aegean Region (16.94%), Mediterranean region (16.09%), Marmara (13.13%), eastern Anatolia (6.8%), Black Sea Region (5.70%) and south-eastern Anatolia (2.75%) (Yazar et al. 2008). In addition, a study conducted to determine the etiological agent responsible for CE in patients originating from most of these regions showed that infection was due to E. granulosus G1 genotype (Ergin et al. 2010). Most CE cases described in this study were from female patients, which is consistent with results previously reported from Turkey (Hakverdi et al. 2008; Ozekinci et al. 2009).
CE infection in all 46 patients included in this study was confirmed to have been caused by E. granulosus s.s (G1 and G3 genotypes). Our results are consistent with the predominance of CE caused by G1 and G3 genotypes in Turkey (Schneider et al. 2008; Utuk et al. 2008; Šnàbel et al. 2009; Ergin et al. 2010; Simsek et al. 2011b; Eryildiz and Şakru 2012; Bakal et al. 2015; Eroğlu et al. 2016). Although E. granulosus G2 genotype was not detected in this study, it had been reported from two water buffaloes from the Black Sea Region of Turkey (Beyhan and Umur 2011). Interestingly, a re-appraisal of the G1–G3 genotype cluster using 112 metacestodes from sheep and cattle intermediate hosts from Turkey did not find evidence to support the continued presence of G2 genotype within E. granulosus s.s (Vural et al. 2008) and other authors have considered it as a microvariant of G3 genotype (Šnàbel et al. 2009). Further, a recent study questioned the validity of G2 genotype within the E. granulosus species complex whereas G1 and G3 were reported to be valid mitochondrial genotypes, but no difference was found between them at the nuclear level (Kinkar et al. 2017).
It is not surprising that E. granulosus s.s is the most the dominant causative species responsible for CE in humans and animals in Turkey. The Middle East has been described as the ‘ancestral seat’ of E. granulosus s.s (Eckert et al. 2001; Nakao et al. 2010). In addition, E. granulosus s.s is a cosmopolitan species that owes its ubiquitous existence to its ability to parasitize a wide range of animal hosts. Further, 88.44% of 1661 molecularly confirmed CE infections were caused by E. granulosus s.s (Alvarez Rojas et al. 2014). CE human infections in Turkey due to E. granulosus s.s. have been reported in adults as well as paediatric patients (Ergin et al. 2010; Simsek et al. 2011b; Bakal et al. 2015). Other human CE cases from Turkey include those caused by E. canadensis G6, which was reported in two surgically confirmed patients using histopathological tissues, retrieved from an Elazig Province university hospital (Simsek et al. 2011b). In addition, E. canadensis G7 was molecularly confirmed from Selçuk-Izmir Province (Šnàbel et al. 2009) and Edirne in the Thrace region (Eryildiz and Şakru 2012). Additional studies are required to further investigate the epidemiology of E. canadensis in Turkey and identify potential risk factors particularly in rural areas. In a recent study, living in endemic rural areas in association with free roaming dogs with access to infected offal were identified as significant risk factors for acquiring CE infection (Possenti et al. 2016).
Hydatidosis in animals from Turkey has been reported in sheep, cattle, goats, mouflon, water buffaloes and pigs (Umur 2003; Yildiz and Gurcan 2003; Yildiz and Tunçer 2005; Esatgil and Tuzer 2007; Kose and Sevimli 2008; Simsek and Eroksuz 2009; Beyhan and Umur 2011). The assessment carried out in this study using GenBank nucleotide sequence data from Turkey for sheep and cattle demonstrated the importance of autochthonous transmission concerning these two species where sheep, cattle and humans harboured the same haplotypes. Sheep are known as the most important hosts for the perpetuation and transmission of E. granulosus s.s worldwide (Deplazes et al. 2017) with high cyst fertility and viability reported from many countries. On the other hand, cattle generally harbour sterile cysts of E. granulosus s.s. Cyst fertility for sheep from Kirikkale for example was reported to be 81.53 and 76.47% for liver and lung cysts respectively (Yildiz and Gurcan 2003). The same study found that the mean number of viable liver protoscoleces was 12,400 and that for the lung cysts to be 5800 where the authors emphasised the important role sheep play in CE transmission in Turkey. Cattle cyst fertility, however, was much lower for example 6.6% in Kırıkkale (Yildiz and Tunçer 2005) and 5.42% in Afyonkarahisar district, western Turkey (Kose and Sevimli 2008) whereas as far as is known, no information on the viability of hydatid cysts derived from Turkish cattle has been published.
In summary, this is the first report on E. granulosus s.s haplotypes and population structure of metacestodes derived from human Turkish hosts. The only other known study on genetic diversity using human CE isolates from Turkey determined haplotypes within the E. granulosus s.s species through the identification of E. granulosus variants using RFLP and SSCP profiles (Eryildiz and Şakru 2012).
All three haplotype clusters described in this study occurred in patients from central and eastern Anatolia and Marmara regions. The E. granulosus G3 haplotype cluster (TUK24–TUK28) was observed in CE patients from central, eastern and south-eastern Anatolia and Marmara only, although this may be related to the small sample size for some of these regions. Further studies are required to investigate the biological significance if any, of E. granulosus s.s haplotypes and the genetic variability of CE from human patients using longer nucleotide sequences and a larger sample set.
References
Abdulhameed MF, Habib I, Al-Azizz SA, Robertson I (2018) A retrospective study of human cystic echinococcosis in Basrah province, Iraq. Acta Trop 178:130–133
Acioz M (2008) The investigation of prevalence of echinococcosis in human by ELISA, in slaughter animals by observation and in dogs by PCR in Muş province. Dissertation, Cunhuriyet University
Altintas N (2003) Past to present: echinococcosis in Turkey. Acta Trop 85:105–112
Altintas N (2008) Parasitic zoonotic diseases in Turkey. Vet Ital 44:633–646
Alvarez Rojas CA, Romig T, Lightowlers MW (2014) Echinococcus granulosus sensu lato genotypes infecting humans—review of current knowledge. Int J Parasitol 44:9–18
Bakal U, Simsek S, Kazez A (2015) Surgical and molecular evaluation of pediatric hydatid cyst cases in eastern Turkey. Korean J Parasitol 53:785–788
Beyhan YE, Umur S (2011) Molecular characterization and prevalence of cystic echinococcosis in slaughtered water buffaloes in Turkey. Vet Parasitol 181:174–179
Boufana B, Lahmar S, Rebai W, Ben Safta Z, Jebabli L, Ammar A, Kachti M, Aouadi S, Craig PS (2014) Genetic variability and haplotypes of Echinococcus isolates from Tunisia. Trans R Soc Trop Med Hyg 108:706–714
Bowles J, Blair D, McManus DP (1992) Genetic variants within the genus Echinococcus identified by mitochondrial DNA sequencing. Mol Biochem Parasitol 54:165–174
Deplazes P, Rinaldi L, Alvarez Rojas CA, Torgerson PR, Harandi MF, Romig T, Antolova D, Schurer JM, Lahmar S, Cringoli G, Magambo J, Thompson RC, Jenkins EJ (2017) Global distribution of alveolar and cystic echinococcosis. Adv Parasitol 95:315–493
Eckert J, Gemmell MA, Meslin FX, Pawlowski ZS (2001) In: WHO/OIE manual on echinococcosis in humans and animals: a public health problem of global concern. Paris OIE. pp. 1–265
Ergin S, Saribas S, Yuksel P, Zengin K, Midilli K, Adas G, Arikan S, Aslan M, Uysal H, Caliskan R, Oner A, Kucukbasmaci O, Kaygusuz A, Torun MM, Kocazeybek B (2010) Genotypic characterisation of Echinococcus granulosus isolated from human in Turkey. Afr J Microbiol Res 4:551–555
Eroğlu F, Genc A, Koltas IS (2016) The genotyping of Echinococcus granulosus isolates with PCR-RFLP methods in Adana Province. Zirve Med J 1:22–25
Eryildiz C, Şakru N (2012) Molecular characterization of human and animal isolates of Echinococcus granulosus in the Thrace Region, Turkey. Balkan Med J 29:261–267
Esatgil MU, Tuzer E (2007) Prevalence of hydatidosis in slaughtered animals in Thrace, Turkey. Türkiye Parazitol Derg 31:41–45
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Res 10:564–567
Felsenstein J (1989) PHYLIP—phylogeny inference package (version 3.2). Cladistics 5:164–166
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Guzel M, Yaman M, Koltas IS, Demirkazik M, Aktas H (2008) Detection of Echinococcus granulosus coproantigens in dogs from Antakya Province, Turkey. Helminthologia 45:150–153
Hakverdi S, Culha G, Canda MS, Yaldiz M, Altintas S (2008) Problem of cystic Echinococcoss in Hatay. Türkiye Parazitol Derg 32:340–342
Hassan ZI, Meerkhan AA, Boufana B, Hama AA, Ahmed BD, Mohammed W, Mero S, Orsten S, Interisano M, Pozio E, Casulli A (2017) Two haplotype clusters of Echinococcus granulosus sensu stricto in northern Iraq (Kurdistan region) support the hypothesis of a parasite cradle in the Middle East. Acta Trop 172:201–207
Hüttner M, Nakao M, Wassermann T, Siefert L, Boomker JDF, Dinkel A, Sako Y, Mackenstedt U, Romig T, Ito A (2008) Genetic characterization and phylogenetic position of Echinococcus felidis Ortlepp, 1937 (Cestoda: Taeniidae) from the African lion. Int J Parasitol 39:861–868
Jordanova DP, Harizanov RN, Kaftandjiev IT, Rainova IG, Kantardjiev TV (2015) Cystic echinococcosis in Bulgaria 1996−2013, with emphasis on childhood infections. Eur J Clin Microbiol Infect Dis 34:1423–1428
Kinkar L, Laurimae T, Simsek S, Balkaya I, Casulli A, Manfredi MA, Ponce-Gordo F, Varcasia A, Lavikainen A, González LM, Rehbein S, VAN DER Giessen J, Sprong H, Saarma U (2016) High-resolution phylogeography of zoonotic tapeworm Echinococcus granulosus sensu stricto genotype G1 with an emphasis on its distribution in Turkey, Italy and Spain. Parasitology 143:1790–1801
Kinkar L, Laurimäe T, Sharbatkhori M, Mirhendi H, Kia EB, Ponce-Gordo F, Andresiuk V, Simsek S, Lavikainen A, Irshadullah M, Umhang G, Oudni-M'rad M, Acosta-Jamett G, Rehbein S, Saarma U (2017) New mitogenome and nuclear evidence on the phylogeny and taxonomy of the highly zoonotic tapeworm Echinococcus granulosus sensu stricto. Infect Genet Evol 52:52–58
Kose M, Sevimli KF (2008) Prevalence of cystic echinococcosis in slaughtered cattle in Afyonkarahisar. Türkiye Parazitol Derg 32:27–30
Kuru BB, Aypak S, Aysul N (2013) Prevalence of Echinococcus granulosus determined with polymerase chain reaction in dogs in Aydın district. Türkiye Parazitol Derg 37:78–83
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Librado P, Rozas J (2009) DnaDP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Nakao M, Sako Y, Yokoyama N, Fukunaga M, Ito A (2000) Mitochondrial genetic code in cestodes. Mol Biochem Parasitol 111:415–424
Nakao M, McManus DP, Schantz PM, Craig PS, Ito A (2007) A molecular phylogeny of the genus Echinococcus inferred from complete mitochondrial genomes. Parasitology 134:713–722
Nakao M, Yanagida T, Okamoto M, Knapp J, Nkouawa A, Sako Y, Ito A (2010) State-of-the-art Echinococcus and Taenia: phylogenetic taxonomy of human-pathogenic tapeworms and its application to molecular diagnosis. Infect Genet Evol 10:444–452
Öge H, Oge S, Gonenc B, Sarimehmetoglu O, Ozbakis G (2017) Coprodiagnosis of Echinococcus granulosus infection in dogs from Ankara, Turkey. Vet Parasitol 242:44–46
Öter K, Bilgin Z, Tınar R, Tuzer E (2011) Tapeworm infections in stray dogs and cats in İstanbul, Turkey. Kafkas Univ Vet Fak Derg 17:595–599
Ozekinci S, Bakir S, Mizrak B (2009) Evaluation of cystic echinococcosis cases given a histopathologic diagnosis from 2002 to 2007 in Diyarbakir. Türkiye Parazitol Derg 33:232–235
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Possenti A, Manzano-Román R, Sánchez-Ovejero C, Boufana B, La Torre G, Siles-Lucas M, Casulli A (2016) Potential risk factors associated with human cystic echinococcosis: systematic review and meta-analysis. PLoS Negl Trop Dis 10(11):e0005114
Rokni MB (2008) The present status of human helminthic diseases in Iran. Ann Trop Med Parasitol 102:283–295
Romig T, Ebi D, Wassermann M (2015) Taxonomy and molecular epidemiology of Echinococcus granulosus sensu lato. Vet Parasitol 213:76–84
Salzburger W, Ewing GB, Haeseler A (2011) The performance of phylogenetic algorithms in estimating haplotype genealogies with migration. Mol Ecol 20:1952–1963
Sariozkan S, Yalcin C (2009) Estimating the production losses due to cystic echinococcosis in ruminants in Turkey. Vet Parasitol 163:330–334
Schneider R, Gollackner B, Edel B, Schmid K, Wrba F, Tucek G, Walochnik J, Auer H (2008) Development of a new PCR protocol for the detection of species and genotypes (strains) of Echinococcus in formalin-fixed, paraffin-embedded tissues. Int J Parasitol 38:1065–1071
Simsek S, Eroksuz Y (2009) Occurrence and molecular characterization of Echinococcus granulosus in Turkish mouflon (Ovis gmelinii anatolica). Acta Trop 109:167–169
Simsek S, Balkaya I, Ciftci AT, Utuk AE (2011a) Molecular discrimination of sheep and cattle isolates of Echinococcus granulosus by SSCP and conventional PCR in Turkey. Vet Parasitol 178:367–369
Simsek S, Kaplan M, Ozercan IF (2011b) A comprehensive molecular survey of Echinococcus granulosus in formalin-fixed paraffin-embedded tissues in human isolates in Turkey. Parasitol Res 109:411–416
Šnàbel V, Altintas N, D’Amelio S, Nakao M, Romig T, Yolasigmaz A, Gunes K, Turk M, Busi M, Hüttner M, Sevcová D, Ito A, Altintas N, Dubinský P (2009) Cystic echinococcosis in Turkey: genetic variability and first record of the pig strain (G7) in the country. Parasitol Res 105:145–154
Swofford DL (1998) PAUP*. Phylogenetic Analysis Using Parsinomy (* and other methods). Version 4. Sinauer, Massachusetts
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tavakoli H, Bayat M, Kousha A (2008) Hydatidosis infection study in human and livestock populations during 2002-2007. Am-Eurasian J Agri Environ Sci 4:473–477
Thompson RCA (2008) The taxonomy, phylogeny and transmission of Echinoccocus. Exp Parasitol 119:439–446
Umur S (2003) Prevalence and economic importance of cystic echinococcosis in slaughtered ruminants in Burdur, Turkey. J Veterinary Med Ser B 50:247–252
Utuk AE, Simsek S, Koroglu E, McManus DP (2008) Molecular genetic characterization of different isolates of Echinococcus granulosus in east and southeast regions of Turkey. Acta Trop 107:192–194
Vural G, Baca AU, Gauci CG, Bagci O, Gicik Y, Lightowlers MW (2008) Variability in the cytochrome c oxidase 1 mitochondrial gene sequence from livestock in Turkey and a re-appraisal of the G1-3 genotype cluster. Vet Parasitol 154:347–350
Yazar S, Ozkan AT, Hökelek M, Polat E, Yilmaz H, Özbilge H, Ustun S, Koltaş IS, Ertek M, Sakru N, Alver O, Cetinkaya Z, Koç Z, Demirci M, Aktaş H, Parsak CK, Ozerdem D, Sakman G, Cengiz ZT, Ozer A, Keklik K, Yemenici N, Turan M, Daştan A, Kaya E, Tamer GS, Girginkardeşler N, Türk M, Sinirtaş M, Evci C, Kiliçturgay S, Mutlu F, Artiş T (2008) Cystic echinococcosis in Turkey from 2001-2005. Türkiye Parazitol Derg 32:208–220
Yibar A, Selcuk O, Senlik B (2015) Major causes of organ/carcass condemnation and financial loss estimation in animals slaughtered at two abattoirs in Bursa Province, Turkey. Prev Vet Med 118:28–35
Yildiz K, Gurcan S (2003) Prevalence of hydatidosis and fertility of hydatid cysts in sheep in Kırıkkale, Turkey. Acta Vet Hung 51(2):181–187
Yildiz K, Tunçer C (2005) Prevalence of hydatid cysts in cattle in the province of Kırıkkale. Türkiye Parazitol Derg 29:247–250
Yolasıgmaz A, Reiterova K, Turk M, Reyhan E, Bozdag AD, Karababa AO, Altintas N, Altintas N (2006) Comparison of serological and clinical findings in Turkish patients with cystic echinococcosis. Helminthologia 43:220–225
Funding
This study was supported by the European Community’s Seventh Framework Programme under the grant agreement 602051 (HERACLES Project; http://www.Heracles-fp7.eu/) and in part by the Overseas Research Fellowship Program (2214) of the Scientific and Technological Research Council of Turkey.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The current study was approved by the Ethical Committee of the Faculty of Medicine, Hacettepe University, Ankara, Turkey (GO 14/293-37).
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Orsten, S., Boufana, B., Ciftci, T. et al. Human cystic echinococcosis in Turkey: a preliminary study on DNA polymorphisms of hydatid cysts removed from confirmed patients. Parasitol Res 117, 1257–1263 (2018). https://doi.org/10.1007/s00436-018-5807-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-018-5807-9