Abstract
Leishmaniasis is a worldwide epidemic disease caused by the genus Leishmania, which is still endemic in the west and northwest areas of China. Some viewpoints of the traditional taxonomy of Chinese Leishmania have been challenged by recent phylogenetic researches based on different molecular markers. However, the taxonomic positions and phylogenetic relationships of Chinese Leishmania isolates remain controversial, which need for more data and further analysis. In this study, the heat shock protein 70 (HSP70) gene and cytochrome b (cyt b) gene were used for phylogenetic analysis of Chinese Leishmania isolates from patients, dogs, gerbils, and sand flies in different geographic origins. Besides, for the interesting Leishmania sp. in China, the ultrastructure of three Chinese Leishmania sp. strains (MHOM/CN/90/SC10H2, SD, GL) were observed by transmission electron microscopy. Bayesian trees from HSP70 and cyt b congruently indicated that the 14 Chinese Leishmania isolates belong to three Leishmania species including L. donovani complex, L. gerbilli, and L. (Sauroleishmania) sp. Their identity further confirmed that the undescribed Leishmania species causing visceral Leishmaniasis (VL) in China is closely related to L. tarentolae. The phylogenetic results from HSP70 also suggested the classification of subspecies within L. donovani complex: KXG-918, KXG-927, KXG-Liu, KXG-Xu, 9044, SC6, and KXG-65 belong to L. donovani; Cy, WenChuan, and 801 were proposed to be L. infantum. Through transmission electron microscopy, unexpectedly, the Golgi apparatus were not observed in SC10H2, SD, and GL, which was similar to previous reports of reptilian Leishmania. The statistical analysis of microtubule counts separated SC10H2, SD, and GL as one group from any other reference strain (L. donovani MHOM/IN/80/DD8; L. tropica MHOM/SU/74/K27; L. gerbilli MRHO/CN/60/GERBILLI). The ultrastructural characteristics of Leishmania sp. partly lend support to the phylogenetic inference that Chinese Leishmania sp. is in close relationship with reptilian Leishmania.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leishmaniases, one of the world’s most neglected diseases, are caused by obligate protozoan parasite from genus Leishmania and transmitted via sandfly. More than 98 countries worldwide are currently threatened by leishmaniasis (WHO 2010). The clinical syndromes of this disease in human range from cutaneous Leishmaniasis (CL, most common form) to acute visceral Leishmaniasis (VL, kala-azar). These different clinical syndromes usually correlate with different pathogenic species. Thus, accurate species discrimination is crucially important for prognosis and treatment choice of the disease. It is now commonly accepted that genus Leishmania comprises three subgenera, i.e., L. (Leishmania) and L. (Viannia), and L. (Sauroleishmania) (Schönian et al. 2010). Thereinto, 31 Leishmania species are known to be parasites of mammals and 20 species are pathogenic for human beings (Akhoundi et al. 2016). Sauroleishmania was proposed as a subgenus for Leishmania that infects lizards (Saf’janova 1982), which was supported by researches on biological criteria and analyses of different Leishmania gene markers (Croan et al. 1997; Fraga et al. 2010; Noyes et al. 1997; Orlando et al. 2002). While there are still a few species within the subgenus Sauroleishmania without specific taxonomic positions.
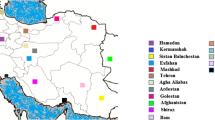
In China, leishmaniasis is mainly endemic in the western and northwestern regions, such as Xinjiang Uygur Autonomous Region (Xinjiang province), Gansu province, and Sichuan province (Zheng et al. 2009). Previous researchers divided the epidemic focus of leishmaniasis in China into plain foci, hill foci, and desert foci according to different geographical origins, causative agents, and clinical manifestations (Lu et al. 1994). As a large country with a vast population and complex ecological environment, China gains an intricate taxonomy and evolution of Leishmania. By analyzing kDNA or nDNA, Hu et al. (1992) and Lu et al. (2001) suggested that the differences at genetic level did exist in Leishmania isolates from different areas in China. Then, our recent studies further revealed the heterogeneity of Chinese Leishmania isolates since several species, including L. donovani complex, L. gerbilli, L. tropica, and L. turanica, have been identified (Cao et al. 2011; Yang et al. 2010, 2013). Moreover, a novel and undescribed Leishmania species may exist in China (Cao et al. 2011; Guan et al. 2012; Sun et al. 2012). Nevertheless, the phylogenetic relationships and taxonomy of Chinese Leishmania have not been completely clarified, especially identification within species complex. In Xinjiang, although L. donovani and L. infantum have been confirmed as the causative agents of VL (Wang et al. 2010a), their distribution is still not clear. Considering that VL is the main form endemic in China, accurate identification of the causative agent is essential to effective control of the disease. Therefore, to get a more reliable phylogenetic inference of Chinese Leishmania, further studies are still necessary.
In order to discriminate Leishnamia species conveniently and accurately, various genetic markers have been developed, such as ribosomal DNA (rDNA) genes, kinetoplast DNA (kDNA) genes, and nuclear genes (Dávila and Momen 2000; Mahboudi et al. 2001; Victoir et al. 1998). In our previous studies, traditional viewpoints on taxonomy and cladistics hypotheses of Chinese Leishmania isolates have been challenged based on different molecular markers (Cao et al. 2011; Guan et al. 2012; Yang et al. 2010). More effective markers are still needed to clarify the species status and phylogeny of Leishmania in China. HSP70 gene, a Leishmania antigen gene on chromosome, is considered the optimal marker for species delimitation and phylogenetic inference of Leishmania, which showed a nearly perfect congruence with multilocus enzyme electrophoresis (MLEE) typing (widely accepted gold standard in Leishmania typing) (Fraga et al. 2010; Van der Auwera and Dujardin 2015). Cyt b gene, which is encoded on the kDNA maxicircle, is another widely used marker that has successfully separated most tested species (Asato et al. 2009; Luyo-Acero et al. 2004). Considering different selection pressure and evolutionary rate on different genes, analyses combining gene markers from nucleus with kinetoplast would be reliable for phylogeny. The recent study (Zhang et al. 2013a) has constructed phylogenetic trees based on HSP70 and lack gene of some Chinese Leishmania strains; unfortunately, the analysis lacked the comparison with the representative Leishmania sequences from other regions of the world and the controversial Chinese Leishmania sp. was absent. Additionally, some researches have confirmed that more isolates and data would affect the phylogenetic results (Botilde et al. 2006; Kuwahara et al. 2009). Thus, more isolates from diverse regions would add to our understanding of species identification and phylogenetic relationships of Chinese Leishmania isolates.
Studies on morphology of Leishmania have been carried out extensively in the middle of twentieth century and was overlooked gradually with the booming up of molecular biology. Since all amastigotes and promastigotes look very alike by optical microscope (Bray 1974), results from electron microscopy seem to be more significant. Pioneering scientists did a lot of research on the ultrastructure of Leishmania and found that the differences on the number of subpellicular microtubules may be of value to differential taxonomy (Garnham 1962; Garnham 1971; Lewis 1975; Sanyal and Sen Gupta 1967). Since the Chinese Leishmania sp. without ascertainable taxonomic status, it is necessary to collect more comprehensive information that related to classification, including ultrastructural characteristics.
In this study, to further evaluate the phylogenetic relationships and obtain reliable species identification of Chinese Leishmania, 14 cyt b sequences and 10 HSP70 sequences of Leishmania isolates were obtained for phylogenetic analyses in conjunction with some representative Leishmania sequences retrieved from GenBank. In addition, for the interesting Leishmania sp. isolates in China, we further characterized the ultrastructure of three Chinese Leishmania sp. strains (MHOM/CN/90/SC10H2; SD; GL) by transmission electron microscopy in order to obtain additional information to help us to understand the novel species.
Materials and methods
Parasites and cultures
Fourteen Chinese Leishmania isolates and three WHO reference strains were used in this study and listed in Table 1. The promastigotes were cultivated in medium 199 supplemented with 15% heat-inactivated fetal bovine serum (HIFBS). For axenic cultivation, 100 U/mL penicillin (Sigma), and 100 μg/mL streptomycin (Sigma) were added in and were put at 25 °C, pH 5.8.
DNA extraction, amplification, and sequencing protocols
The promastigotes in logarithmic phase (approximately 1–5 × 109/mL) were harvested by centrifuging at 3300×g for 10 min at 4 °C and washed with PBS for three times. Genomic DNA was extracted using a standard sodium dodecyl sulfate-proteinase K procedure (Sambrook and Russell 2001). The cyt b fragment was amplified using primers LCBF1 and LCBR2 (Luyo-Acero et al. 2004). The PCR procedure were described as follows: initial denaturation at 94 °C for 3 min followed by 35 cycles of 94 °C for 30 s, 58 °C for 30 s, 72 °C for 1.5 min, and a final extension at 72 °C for 10 min. The primers HSP70sen and HSP70ant (Garcia et al. 2004) were used to amplify HSP70 segments. For HSP70, the PCR conditions were 95 °C for 5 min followed by 30 cycles of 95 °C for 60 s, 60 °C for 60 s, 72 °C for 1.5 min, with a final extension at 72 °C for 10 min. The PCR products were checked using 1.5% agarose gel electrophoresis and purified using a DNA purification kit (TIANGEN, Beijing, China) per the protocol of the manufacturer. Purified products were then sent to Tsingke Biological Technology Co., Ltd. (Chengdu, China) for gene sequencing using the same PCR primers on ABI 3730 automated sequencer (Applied Biosystems, Inc.). Obtained gene sequences were finally uploaded to GenBank (accession numbers are listed in Table 1).
Sequence alignment and phylogenetic analyses
A group of cyt b sequences were retrieved from GenBank, including 30 sequences of genus Leishmania and 1 sequence of Trypanosoma brucei (Tables 1 and 2). For HSP70, a total of 47 sequences of Leishmania with 1 sequence of Trypanosoma cruzi were retrieved from GenBank (Tables 1 and 2). The sequences were multiple-aligned using Clustal X v.1.83 (Thompson et al. 1997) with its default option. To generate a consensus tree, the base composition heterogeneity was evaluated using chi-squared (χ 2) tests executed in PAUP* 4.0b10 (Swofford 2002). Then, the phylogenetic trees reconstruction of Leishmania were implemented applying Bayesian inference (BI) with the MrBayes v.3.2 program (Ronquist and Huelsenbeck 2003). Gaps were treated as missing data. Trypanosoma cruzi and Trypanosoma brucei were used to root the trees. The most appropriate model (cyt b: GTR + I + G; HSP70: TrN + I + G) for Bayesian analyses was selected by combined utilization of the program PAUP and Modeltest 3.7 (Posada and Crandall 1998). Four Markov chains were proceeded for four million generations, and trees were sampled for every 200 generations. The first one million sample trees were discarded, and the remaining trees were used for construction of 50% majority-rule consensus tree and calculation of posterior probabilities of clades. The results of Bayesian analyses were visualized using Treeview v1.6.6 (Page 1996).
Transmission electron microscopy
Besides the three undescribed Leishmania sp. isolates MHOM/CN/90/SC10H2, SD, and GL, three WHO reference strains (L. donovani MHOM/IN/80/DD8; L. tropica MHOM/SU/74/K27; L. gerbilli MRHO/CN/60/GERBILLI) were chosen for comparison of ultrastructural characteristics. The promastigotes in logarithmic phase, washed with PBS, was prefixed in 3% glutaraldehyde solution. Then, the promastigotes was postfixed in 1% osmium tetroxide (OsO4), dehydrated in acetone of gradient concentration, and embedded in epoxy resin. The ultrathin sections were stained with uranyl acetate and lead citrate. Sections were examined with a transmission electron microscope (TEM; HITACHI, H-600IV, Japan). Electron micrograph were taken of clearly visible cells. The longitudinal sections were used for ultrastructural observation. The transverse sections of each of six specimens were selected for counting the numbers of subpellicular microtubules.
Statistical analysis
The microtubule counts were analyzed by IBM SPSS Statistics 20.0 (IBM Corp., USA) using one-way analysis of variance, followed by Scheffe post hoc comparison. Any P value of less than or equal to 0.05 was regarded as statistical significance. All tables and pictures were created by using Microsoft Office 2013 (Microsoft Corp., USA).
Results
Sequence alignment and analyses
There were 14 cyt b gene fragments and 10 HSP70 gene fragments obtained through PCR. The GenBank accession numbers of these newly obtained sequences of Chinese isolates are listed in Table 1, of which KX061904-KX061917 were for cyt b and KX061893-KX061902 were for HSP70.
A total of 773 aligned sites in the matrix of cyt b, 324 were variable sites, with 193 parsimony-informative. Percentage of four DNA bases were as follows: A, 26.8; C, 7.8; G, 15.6; T, 49.8. The cyt b gene fragment of Leishmania is AT rich (76.6%). The heterogeneity test showed that there were no significant differences in base composition bias among cyt b data (χ 2 = 66.42, df = 150, P = 1.00). For HSP70, 235 of the 1262 aligned characters were variable, including 116 that were parsimony-informative. Percentage of four DNA bases were as follows: A, 21.5; C, 30.4; G, 34.7; T, 13.5. The HSP70 gene fragment of Leishmania is CG rich (65.0%). The heterogeneity test of HSP70 data also showed that there were no significant differences in base composition bias (χ 2 = 15.83, df = 195, P = 1.00).
Phylogenetic analyses
In the BI tree inferred from the aligned matrix of cyt b (Fig. 1), three clades were recovered with high support (PP = 0.97, 1.0, and 1.0, respectively), corresponding to the three subgenera Vanina, Leishmania, and Sauroleishmania. Chinese Leishmania isolates in this study were distributed in three subclades, i.e., L. donovani complex, L. major complex, and Leishmania sp. Ten isolates (KXG-918, KXG-927, KXG-Liu, KXG-Xu, 9044, SC6, KXG-65, 801, WenChuan, Cy) from different foci of China were clustered with five reference sequences of L. donovani/L. infantum and other three sequences from China (Yang et al. 2013), then together formed the monophyletic clade of Leishmania donovani complex (PP = 1.00). In this clade, however, L. donovani and L. infantum were not reciprocally monophyletic groups. The strain Ejni-154 isolated from Rhombomys opimus in Inner Mongolia was clustered with the WHO L. gerbilli reference strain MRHO/CN/60/Gerbilli (PP = 1.00) and joined by some reference sequences of Leishmania major and Leishmania turanica, which formed a subclade L. major complex (PP = 1.0). The isolates GL and SC10H2, from Gansu and Sichuan respectively, were clustered with the other five Chinese Leishmania isolates which termed Leishmania sp. before (Yang et al. 2013) and formed a strong clade with one sequence of L. tarentolae (PP = 1.00).
The 50% majority-rule consensus tree inferred from Bayesian inference of cyt b dataset using MrBayes v. 3.2. The numbers at the nodes represent Bayesian posterior probabilities; brackets indicate Leishmania species, complexes, and subgenera with reference to descriptions based on multilocus enzyme electrophoresis (Lainson et al. 1987; Rioux et al. 1990; Cupolillo et al. 1994; Cupolillo et al. 2000); red-marked: strains that genes were sequenced in this study; the strains that from China were tagged with asterisk (color figure online)
The BI tree based on the aligned matrix of HSP70 (Fig. 2) showed that three subgenera Vanina, Leishmania, and Sauroleishmania were clearly divided and eight Chinese isolates in this study were distributed in three clades: L. donovani complex, L. gerbilli, and Leishmania sp. As well as cyt b, L. donovani and L. infantum were not reciprocally monophyletic groups. Nevertheless, the L. infantum strains were found as a separate subgroup within L. donovani complex in the BI tree (PP = 0.9). Two strains KXG-918 and KXG-927, which were isolated from Phlebotomus major wui in Xinjiang, were clustered with the WHO reference strain DD8 and other L. donovani strains. The other two strains Cy and WenChuan, which were isolated from canine in Sichuan and Gansu of China respectively, were clustered with other Chinese Leishmania isolates and FN395031, FN395032, HF586393, and HF586408 identified as L. infantum before (Fraga et al. 2010; Van der Auwera et al. 2013). The isolates SC10H2, SD, and GL were still clustered with other strains of Leishmania sp. and lizard Leishmania and then formed a clade with nonpathogenic L. tarentolae (PP = 1) instead of any other human pathogenic Leishmania species.
The 50% majority-rule consensus tree inferred from Bayesian inference of HSP70 dataset using MrBayes v. 3.2. The numbers at the nodes represent Bayesian posterior probabilities; brackets indicate Leishmania species, complexes, and subgenera with reference to descriptions based on multilocus enzyme electrophoresis (Lainson et al. 1987; Rioux et al. 1990; Cupolillo et al. 1994; Cupolillo et al. 2000); red-marked: strains that genes were sequenced in this study; the strains that from China were tagged with asterisk (color figure online)
Ultrastructure characterization under transmission electron microscope
Through transmission electron microscopy, the promastigotes of three Leishmania sp. isolates showed roughly identical intracellular structures with other three reference strains, including a centrally located nucleus, a rod-like kinetoplast, branched mitochondrial system, and anterior flagellum. Unexpectedly, the Golgi apparatus has not been observed in the three isolates SC10H2, SD, and GL (Fig. 3b showed the Golgi absence in SC10H2). While, in the other reference strains (L. donovani MHOM/IN/80/DD8; L. tropica MHOM/SU/74/K27; L. gerbilli MRHO/CN/60/GERBILLI), the Golgi was well developed. The extensive membrane laminations and unequal-sized vesicles located near the kinetoplast could be observed (Fig. 3a showed the Golgi in DD8).
The plasma membrane of promastigote is double layered, and lying immediately beneath it are longitudinally arranged subpellicular microtubules. Table 3 shows the counting results of microtubules of the six strains. Then, the results were analyzed using one-way analysis of variance, and the calculated F value is 405.186, which suggested significant differences in the mean number of microtubules among the six strains (P < 0.01). Followed multiple comparisons test set SC10H2, SD and GL as one group (P > 0.05) which apart from DD8 (P < 0.01), K27 (P < 0.01) and GERBILLI (P < 0.01) respectively.
Discussion
With reconstruction of phylogenetic relationships of Chinese Leishmania based on HSP70 and cyt b gene sequences, the species identification of Chinese isolates were analyzed, compared, and correlated with their geographical origins. The ultrastructure of the interesting Chinese Leishmania sp. isolates were also observed to further understand the novel species. This study brings further insight into the taxonomic positions and phylogenetic relationships of Chinese Leishmania.
Using the cyt b gene, most of the strains analyzed in this study (KXG-918, KXG-927, KXG-Liu, KXG-Xu, 9044, SC6, KXG-65, 801, WenChuan, Cy), which were collected from the plain, hill, and desert foci of China, were clustered with other strains of L. donovani complex (Fig. 1). However, L. infantum cannot be discriminated from L. donovani according to cyt b data. While HSP70 gene is more powerful to identify Leishmania at the species, intra- and supra-species level (Fraga et al. 2010), which was confirmed in this study. As can be seen in Fig. 2, the canine isolates Cy and WenChuan, along with other six Chinese isolates, 801, 8801, KS6, 1101, 1102, and WDD23 (accession numbers JX970993, JX970997, JX970996, JX312705, and JX312712), were clustered with other identified L. infantum strains from Europe (Fraga et al. 2010; Van der Auwera et al. 2013) and formed a subclade within the L. donovani complex. Although mentioned six Chinese isolates were classified as L. donovani according to lack gene analysis (Zhang et al. 2013b), the phylogenetic tree based on lack gene have not reflected a clear classification of subspecies within L. donovani complex. Therefore, the two canine isolates and the other six Chinese isolates are suggested to be L. infantum in this study. Since the canine isolate WenChuan was from Sichuan, this result indicated that the canine leishmaniasis (CanL) in Sichuan may be caused by L. infantum, which is consistent with the recent studies (Shang et al. 2011; Wang et al. 2011). Moreover, considering the infected dogs are the main reservoir host of human VL, L. infantum should be the causative agents of VL in Sichuan province. Likewise, Cy, 1101, 1102, and WDD23 were isolated from domestic dogs in Gansu, and 8801 from VL patients in Gansu as well. Thus, we inferred that L. infantum might be the pathogen that cause the VL in Gansu, and dogs are likely the principal source of infection to human. These inference was supported by the research of Zhang and Zhang (2010). While, about the identification of isolate 801, which are from VL patients in Kashi of Xinjiang, it remains controversial. That 801 are suggested to be L. infantum based on HSP70 analysis in this study is in accord with multilocus microsatellite typing (MLMT) investigation (Alam et al. 2014), but differs from the previous study of ITS1 sequences (Wang et al. 2010a; Yang et al. 2010) which considered that the isolate 801 belong to L. donovani. This divergence may be due to the short history that L. infantum has differentiated from L. donovani and/or the different efficacy of the two gene markers on phylogenetic analysis. It now appears that L. infantum, which is one of the causative agents of VL, is primarily distributed in western mountainous areas and plain of northwestern China, including Sichuan, Gansu, and Xinjiang provinces. This point is shared by the previous report (Lun et al. 2015).
In this study, two other strains, KXG-918 and KXG-927, which were isolated from Phlebotomus major wui in Karamay, Xinjiang, along with the strain KXG-65(accession number JX021437) also isolated from Phlebotomus major wui in the same area, were clustered with the reference strain DD8 and identified as L. donovani. Meanwhile, we found that the strains KXG-XU and KXG-LIU (HSP70 gene sequence JX021433 and JX021432), isolated from CL patients in Karamay, Xinjiang, also belong to L. donovani according to HSP70 BI tree, which was in accord with the study based on ISSR-PCR by Wang et al. (2010b). Accordingly, since the five strains KXG-918, KXG-927, KXG-65, KXG-XU, and KXG-LIU were all isolated in Karamay, Xinjiang, and grouped together based on HSP70 data, it can be inferred that L. donovani is the pathogen of CL in Karamay and Phlebotomus major wui is the vector. Whereas this phylogenetic inference challenged the previous determination that the L. infantum is the pathogen of CL in Karamay based on gene hybridization and animal inoculation (Guan et al. 1994; Wang et al. 1996; Yang et al. 1997). Generally, L. donovani cause VL and there have been few reports of CL caused by L. donovani except in Sri Lanka, where CL by L. donovani has been prevalent recently (Sanayaka et al. 2014). In China, cutaneous leishmaniasis has not been reported except in the single isolated endemic site, i.e., Karamay in Xinjiang. Given that Xinjiang is the major epidemic region and has an intricate prevalence of leishmaniasis, to be clear about the types of endemic Leishmania, more isolates would be needed, especially from Kashi and Karamay.
Using transmission electron microscope, there were no significant differences in the basic ultrastructure of the six strains, but a clear Golgi apparatus had not yet been observed in the three isolates SC10H2, SD, and GL. Few studies have reported the Leishmania species which lack of Golgi apparatus before. Nevertheless, according to the ultrastructural study on Leishmania promastigotes from reptiles by Lewis (1975), the Golgi apparatus in L. adleri and L. agamae were not well developed and had not been found in L. hoogstraali. The three Leishmania strains, L. adleri, L. agamae, and L. hoogstraali, all belong to L. (Sauroleishmania) (Akhoundi et al. 2016). Kazemi et al. (2008) compared ultrastructure of promastigotes of lizard Leishmania and Leishmania major and found that there was no Golgi apparatus for lizard Leishmania promastigotes. Lewis also considered the true biologic dissimilarities must be reflected by the differences of development of the Golgi apparatus among strains, since his previous study suggested a positive correlation between the development degree of Golgi and the ability for promastigotes to invade and get into host cells (Lewis 1974). Additionally, we also analyzed the microtubule counts of the six strains, and found that the three isolates SC10H2, SD, GL formed one group and showed significant differences comparing with any other reference strain (P < 0.01). Previously, Gardener et al. (1977) found that the number of microtubules are various among different Leishmania species. Lewis (1975) considered that the microtubule counts in classification would be valuable to judge to which species the Leishmania strains does not belong rather than the opposite. Those above leaded us to speculate that the three isolates SC10H2, SD, and GL may be distinct from the other strains in this study (DD8, K27, and GERBILLI), and close to the reptile-infecting Leishmania that belong to Sauroleishmania. This conjecture was just in accord with the phylogeny inferred from cyt b and HSP70 gene data. Of course, we cannot rule out the possibility that the Golgi in the three strains has been changed in form or so underdeveloped that it has not been observed. The study of microtubule counts was also restricted by the amount of samples. Thus, to obtain a relatively accurate assessment, a wide range of sample sizes would be necessary.
Based on HSP70 and cyt b analyses, the three strains SD, GL, and SC10H2 formed a clade which was most closely related to the nonpathogenic L. tarentolae with several other Chinese isolates. The topological structure is consistent with our anterior studies on ITS1 (Yang et al. 2010), COII (Cao et al. 2011), SSU rRNA, and 7SL RNA (Guan et al. 2012). Meanwhile, the ultrastructural characteristics of three Leishmania sp. isolates were also inclined to the results of phylogenetic analyses in this study. However, the three strains were all isolated in VL patients from Shandong, Gansu, and Sichuan provinces, respectively, and the strain SC10H2 was confirmed to share high homology with the pathogens isolated from CanL in Beichuan County, Sichuan, China (Sun et al. 2012). These seem to be contradictory with former studies which indicated that Sauroleishmania is not pathogenic to human (Belova 1971). Thus, our results further confirmed that a novel species of Leishmania causing CanL and kala-azar does exist in China and closely related to L. tarentolae. Although a growing number of information about Chinese Leishmania sp. has been obtained, however, the epidemiology and distribution of the Leishmania sp. are still unclear. It is known that sandfly is the key to Leishmania natural cycle. A recent study has found that Leishmania DNA obtained from Sergentomyia dentate is highly similar to Leishmania sp. (KJ667095, 99.3%) and L. tarentolae (98%) based on HSP70 gene analysis, which may be an evidence for natural cycle of Sauroleishmania agents in western Turkey (Özbel et al. 2016). Considering that Chinese Leishmania sp. gene segments have been detected in canines in Sichuan (Sun et al. 2012), it is necessary to investigate the Leishmania infection rate of sandflies in these endemic areas to track new evidence to understand the natural cycle of Leishmania sp. In addition, more recent Leishmania sp. isolates, valid L. tarentolae type strains, and abundant gene information of Chinese Leishmania sp. would add to our understanding of its definite taxonomic status and evolution.
In conclusion, based on cyt b and HSP70 sequence data, the 14 Chinese Leishmania isolates were found to belong to three Leishmania species including L. donovani complex, L. gerbilli, and L. (Sauroleishmania) sp. This result further confirmed that the undescribed Leishmania species causing visceral Leishmaniasis (VL) in China is closely related to L. tarentolae. The HSP70 results also suggested the classification of subspecies within L. donovani complex: KXG-918, KXG-927, KXG-Liu, KXG-Xu, 9044, SC6, and KXG-65 belong to L. donovani; Cy, WenChuan, and 801 were proposed to be L. infantum. Meanwhile, the ultrastructural characteristics of Leishmania sp. isolates partly lent support to the phylogenetic inference that Chinese Leishmania sp. is in close relationship with reptilian Leishmania. There are still many unknowns about Chinese Leishmania to be resolved. Further study on a wide range of epidemiological survey about patients, sandflies, and other reservoirs would be needed to get the whole picture of Leishmania in China.
References
Akhoundi M, Kuhls K, Cannet A, Votýpka J, Marty P, Delaunay P, Sereno D (2016) A historical overview of the classification, evolution, and dispersion of Leishmania parasites and sandflies. Plos Neglect Trop D 10(3):e0004349
Alam MZ, Nakao R, Sakura T, Kato H, Qu JQ, Chai JJ, Chang KP, Schönian G, Katakura K (2014) Genetic diversity of Leishmania donovani/infantum complex in China through microsatellite analysis. Infect Genet Evol 22(3):112–119
Asato Y, Oshiro M, Myint CK, Yamamoto Y, Kato H, Marco JD, Mimori T, Gomez EA, Hashiguchi Y, Uezato H (2009) Phylogenic analysis of the genus Leishmania by cytochrome b gene sequencing. Exp Parasitol 121(4):352–361
Belova EM (1971) Reptiles and their importance in the epidemiology of leishmaniasis. Bull. Wld Hlth Org 44:553–560
Botilde Y, Laurent T, Tintaya WQ, Chicharro C, Cañavate C, Cruz I, Kuhls K, Schönian G, Dujardin JC (2006) Comparison of molecular markers for strain typing of Leishmania infantum. Infect Genet Evol 6:440–446
Bray RS (1974) Leishmania. Annu Rev Microbiol 28:189–217
Cao DP, Guo XG, Chen DL, Chen JP (2011) Species delimitation and phylogenetic relationships of Chinese Leishmania isolates reexamined using kinetoplast cytochrome oxidase II gene sequences. Parasitol Res 109(1):163–173
Croan DG, Morrison DA, Ellis JT (1997) Evolution of the genus Leishmania revealed by comparison of DNA and RNA polymerase gene sequences. Mol Biochem Parasit 89(2):149–159
Cupolillo E, Grimaldi G Jr, Momen H (1994) A general classification of New World Leishmania using numerical zymotaxonomy. AmJTrop Med Hyg 50(3):296–311
Cupolillo E, Medina-Acosta E, Noyes H, Momen H, Grimaldi G Jr (2000) A revised classification for Leishmania and Endotrypanum. Parasitol Today 16(4):142–143
Dávila AMR, Momen H (2000) Internal-transcribed-spacer (ITS) sequences used to explore phylogenetic relationships within Leishmania. Ann Trop Med Parasitol 94:651–654
Fraga J, Montalvo AM, De Doncker S, Dujardin JC, Van der Auwera G (2010) Phylogeny of Leishmania species based on the heat-shock protein 70 gene. Infect Genet Evol 10(2):238–245
Garcia L, Kindt A, Bermudez H, Llanos-Cuentas A, De Doncker S, Arevalo J, Tintaya WQ, Dujardin JC (2004) Culture-independent species typing of neotropical Leishmania for clinical validation of a PCR-based assay targeting heat shock protein 70 genes. J Clin Microbiol 42:2294–2297
Gardener PJ, Shchory L, Chance ML (1977) Species differentiation in the genus Leishmania by morphometric studies with the electron microscope. Ann Trop Med Parasi 71(2):147–155
Garnham PCC (1962) Cutaneous leishmaniasis in the new world with special reference to Leishmania mexicana. Sci Rep 1st Sup Sanita 2:76–82
Garnham PCC (1971) The genus Leishmania. B World Health Organ 44(4):477–489
Guan LR, Yang YQ, Xu YX, Qu JQ, Zuo XP, Wang G, Lu HG, Zhong L, Chang KP (1994) Leishmaniasis in Karamay XIV. Identification of promastigote isolates from naturally infected phlebotomus major wui. Chin J Parasitol Paras Dis 12(4):257–261 (In Chinese with English abstract)
Guan W, Cao DP, Sun K, Xu JN, Zhang JR, Chen DL, Chen JP (2012) Phylogenic analysis of Chinese Leishmania isolates based on small subunit ribosomal RNA (SSU rRNA) and 7 spliced leader RNA (7SL RNA). Acta Parasitol 57(2):101–113
Hu X, Lu H, Luo P, Wang Z, Yi T (1992) Identification of Leishmania donovani isolates from different kala-azar foci in China by kDNA hybridization. Chin Med Sci J 7:63–66 (in Chinese with English abstract)
Kazemi B, Heidari MH, Naderi M, Piryaei A, Nazari-Pouya MR (2008) Study on ultrastructure of Leishmania major and lizard Leishmania. J Cell Anim Biol 2:129–133
Kuwahara K, Kato H, Gomez EA, Uezato H, Mimori T, Yamamoto Y, Calvopiña M, Cáceres AG, Iwata H, Hashiguchi Y (2009) Genetic diversity of ribosomal RNA internal transcribed spacer sequences in Lutzomyia species from areas endemic for New World cutaneous leishmaniasis. Acta Trop 112:131–136
Lainson R, Shaw JJ, Peters W, Killick-Kendrick R (1987) Evolution, classification and geographical distribution. In: The leishmaniases in biology and medicine. London
Lewis DH (1974) Infection of tissue culture cells of low phagocytic ability by Leishmania mexicana mexicana. Ann Trop Med Parasit 68(3):327–336
Lewis DH (1975) Ultrastructural study of promastigotes of Leishmania from reptiles. J Eukaryot Microbiol 22(3):344–352
Lu HG, Zhong L, Guan LR, Qu JQ, Hu XS, Chai JJ, Xu ZB, Wang CT (1994) Separation of Chinese Leishmania isolates into five genotypes by kinetoplast and chromosomal DNA heterogeneity. AmJTrop Med Hyg 50:763–770
Lu DM, Hu XS, Qiao ZD (2001) Analysis of Leishmania species and strains from China by RAPD technique. Chin J Parasitol Paras Dis 19(5):290–293 (in Chinese with English abstract)
Lun ZR, Wu MS, Chen YF, Wang JY, Zhou XN, Liao LF, Chen JP, Chow LMC, Chang KP (2015) Visceral leishmaniasis in China: an endemic disease under control. Clin Microbol Rev 28(4):987–1004
Luyo-Acero GE, Uezato H, Oshiro M, Takei K, Kariya K, Katakura K, Gomez-Landires E, Hashiguchi Y, Nonaka S (2004) Sequence variation of the cytochrome b gene of various human infecting members of the genus Leishmania and their phylogeny. Parasitology 128(05):483–491
Mahboudi F, Abolhassan M, Yaran M, Mobtaker H, Azizi M (2001) Identification and differentiation of Iranian Leishmania species by PCR amplification of kDNA. Scand J Infect Dis 33:596–598
Noyes HA, Arana BA, Chance ML, Maingon R (1997) The Leishmania hertigi (kinetoplastida; trypanosomatidae) complex and the lizard Leishmania: their classification and evidence for a neotropical origin of Leishmania-endotrypanum clade. J Eukaryot Microbio 44(5):511–517
Orlando TC, Rubio MAT, Sturm NR, Campbell DA, Floeter-Winter LM (2002) Intergenic and external transcribed spacers of ribosomal RNA genes in lizard-infecting Leishmania: molecular structure and phylogenetic relationship to mammal-infecting Leishmania in the subgenus Leishmania (Leishmania). Mem Inst Oswaldo Cruz 97(5):695–701
Özbel Y, Karakuş M, Arserim SK, Kalkan ŞO, Töz S (2016) Molecular detection and identification of Leishmania spp. in naturally infected phlebotomus tobbi and sergentomyia dentata in a focus of human and canine leishmaniasis in western Turkey. Acta Trop 155:89–94
Page RDM (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12(4):357–358
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14(9):817–818
Rioux JA, Lanotte G, Serres E, Pratlong F, Bastien P, Perieres J (1990) Taxonomy of Leishmania. Use of isoenzymes. Suggestions for a new classification. Ann Parasitol Hum Comp 65(3):111–125
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19(12):1572–1574
Saf’janova VM (1982) The problem of taxonomy with Leishmania. Ser Protozool Sov Acad Sci Leningr 7:5–109
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Long island, New York
Sanayaka R, Kahawita I, Gamage A, Siribaddana S, Agampodi S (2014) Emergence of cutaneous leishmaniasis in polonnaruwa, Sri Lanka 2008-2011. Tropical Med Int Health 19(2):140–145
Sanyal AB, Sen Gupta PC (1967) Fine structure of Leishmania in dermal leishmanoid. T Roy Soc Trop Med 61(2):211–216
Schönian G, Mauricio I, Cupolillo E (2010) Is it time to revise the nomenclature of Leishmania? Trends Parasitol 26(10):466–469
Shang LM, Peng WP, Jin HT, Xu D, Zhong NN, Wang WL, Wu YX, Liu Q (2011) The prevalence of canine Leishmania infantum infection in Sichuan Province, southwestern China detected by real time PCR. Parasite Vector 4(1):1–5
Sun K, Guan W, Zhang JG, Wang YJ, Tian Y, Liao L, Yang BB, Chen DL, Chen JP (2012) Prevalence of canine leishmaniasis in Beichuan County, Sichuan, China and phylogenetic evidence for an undescribed Leishmania sp. in China based on 7SL RNA. Parasite Vector 5:75
Swofford DL (2002) PAUP 4.0 b10. Phylogenetic analysis using parsimony (and other methods). Sunderland, Massachusetts
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882
Van der Auwera G, Dujardin JC (2015) Species typing in dermal leishmaniasis. Clin Microbiol Rev 28(2):265–294
Van der Auwera G, Maes I, De Doncker S, Ravel C, Cnops L, Van Esbroeck M, Van Gompel A, Clerinx J, Dujardin JC (2013) Heat-shock protein 70 gene sequencing for Leishmania species typing in European tropical infectious disease clinics. Euro Surveill 18(30):20543
Victoir K, Bañuls AL, Arévalo J, Llanos-Cuentas A, Hamers R, Noel S, De Doncker S, Le Ray D, Tibayrenc M, Dujardin JC (1998) The gp63 gene locus, a target for genetic characterization of Leishmania belonging to subgenus Viannia. Parasitology 117:1–13
Wang JY, Qu JQ, Wang RQ, Guan LR, Ren HY, Chang KP (1996) Analysis on homology in several isolates of Leishmania from Karamay Xinjiang. Chin J Parasitol Parasit Dis 14(4):266–269 (in Chinese with English abstract)
Wang JY, Gao CH, Yang YT, Chen HT, Zhu XH, Lu S, Chen SB, Tong SX, Steinmann P, Ziegelbauer K, Zhou XN (2010a) An outbreak of the desert sub-type of zoonotic visceral leishmaniasis in Jiashi, Xinjiang Uygur Autonomous Region, People’s Republic of China. Parasitol Int 59(59):331–337
Wang Y, Yang YT, Wang JY, Bao YF, Guan L, Gao CH, Shi F (2010b) Molecular characterization of Leishmania isolates from China by inter-simple sequence repeat polymerase chain reaction. Parasitol Res 106(6):1385–1394
Wang JY, Ha Y, Gao CH, Wang Y, Yang YT, Chen HT (2011) The prevalence of canine Leishmania infantum infection in western China detected by PCR and serological tests. Parasite Vector 4:69
World Health Organization (2010) Control of leishmaniases. WHO technical report series no. 949. Report of a meeting of the WHO Expert Committee on the Control of Leishmaniases, Geneva, 22–26 March 2010
Yang YQ, Guan LR, Wu JT (1997) Study on the parasitic character of Leishmania infantum in monkey in Karamay area. Endemic Diseases Bulletin 12(1):8–11 (in Chinese with English abstract)
Yang BB, Guo XG, Hu XS, Zhang JG, Liao L, Chen DL, Chen JP (2010) Species discrimination and phylogenetic inference of 17 Chinese Leishmania isolates based on internal transcribed spacer 1 (ITS1) sequences. Parasitol Res 107(5):1049–1065
Yang BB, Chen DL, Chen JP, Liao L, Hu XS, Xu JN (2013) Analysis of kinetoplast cytochrome b gene of 16 Leishmania isolates from different foci of China: different species of Leishmania in China and their phylogenetic inference. Parasite Vector 6:32
Zhang FN, Zhang LP (2010) Analysis on recurrence and death of leishmaniasis cases in Sichuan Province. J Prev Med Inf 26:961–964 (in Chinese with English abstract)
Zhang CY, Lu XJ, Du XQ, Jian J, Shu L, Ma Y (2013a) Phylogenetic and evolutionary analysis of Chinese Leishmania isolates based on multilocus sequence typing. PLoS One 8(4):e63124
Zhang CY, Zhou J, Ding B, Lu XJ, Xiao YL, Hu XS, Ma Y (2013b) Phylogenetic analysis of lack gene sequences for 22 Chinese Leishmania isolates. Infect Genet Evol 17:79–86
Zheng CJ, Wang LY, Xu X, Zhu XH, Wu WP (2009) Visceral leishmaniasis in China during 2004-2007. Chin J Parasitol Paras Dis 27(4):344–346 (in Chinese with English abstract)
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31572240, 81171607, and 81672048) and the National Project of Important Infectious Diseases of China (2008-ZX10004-011).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Yuan, D., Qin, H., Zhang, J. et al. Phylogenetic analysis of HSP70 and cyt b gene sequences for Chinese Leishmania isolates and ultrastructural characteristics of Chinese Leishmania sp.. Parasitol Res 116, 693–702 (2017). https://doi.org/10.1007/s00436-016-5335-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-016-5335-4