Abstract
Schistosomiasis, caused by blood flukes belonging to several species of the genus Schistosoma, is a serious and widespread parasitic disease. Accurate and rapid differentiation of these etiological agents of animal and human schistosomiasis to species level can be difficult. We report a real-time PCR assay coupled with a high-resolution melt (HRM) assay targeting a portion of the nuclear 18S rDNA to detect, identify, and distinguish between four major blood fluke species (Schistosoma japonicum, Schistosoma mansoni, Schistosoma haematobium, and Schistosoma mekongi). Using this system, the Schistosoma spp. was accurately identified and could also be distinguished from all other trematode species with which they were compared. As little as 10−5 ng genomic DNA from a Schistosoma sp. could be detected. This process is inexpensive, easy, and can be completed within 3 h. Examination of 21 representative Schistosoma samples from 15 geographical localities in seven endemic countries validated the value of the HRM detection assay and proved its reliability. The melting curves were characterized by peaks of 83.65 °C for S. japonicum and S. mekongi, 85.65 °C for S. mansoni, and 85.85 °C for S. haematobium. The present study developed a real-time PCR coupled with HRM analysis assay for detection and differential identification of S. mansoni, S. haematobium, S. japonicum, and S. mekongi. This method is rapid, sensitive, and inexpensive. It has important implications for epidemiological studies of Schistosoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Schistosomiasis is a neglected tropical disease caused by blood-dwelling trematodes of the genus Schistosoma (Phylum Platyhelminthes, Class Trematoda). It has a high prevalence in subtropical regions of Africa, the Americas, and Asia (Gryseels 2012). This helminthiasis has been ranked second only to malaria among parasitic diseases, affects about 200 million people and causes about 41,000 deaths annually worldwide (Gryseels et al. 2006; WHO 2008). Out of the 21 currently recognized Schistosoma species, Schistosoma japonicum (China and Southeast Asia; dwells in peri-intestinal venules), Schistosoma mansoni (Africa, Arabia, and South America; dwells in peri-intestinal venules), and Schistosoma haematobium (Africa and Arabia; dwells in the perivesical plexus) are the main species responsible for human infections and are hence have public health significance as well as imposing an economic burden (Clerinx and Van Gompel 2011). Additionally, Schistosoma mekongi (dwells in peri-intestinal venules) is found in communities along the Mekong River in Cambodia and Lao People’s Democratic Republic. Although primarily of local importance, it is associated with high mortality rates (Muth et al. 2010; Meltzer and Schwartz 2013). Two of these species, S. japonicum and S. mekongi, can also infect some wild and domesticated animals as their reservoir hosts. Their zoonotic nature complicates control efforts (Hotez et al. 2007; Clerinx and Van Gompel 2011).
Nowadays, people are moving around the world more than ever before, responding to the need for social and economic development. Travel to a variety of tropical destinations by expatriates from nonendemic areas exposes them to risk of infection by several Schistosoma species (Li et al. 2015). The specific diagnosis of schistosomiasis in humans is central to control of this disease and to understand the intricacies of its epidemiology. Traditionally, microscopic, biochemical, and serological techniques have been applied for diagnosis of Schistosoma infections, but most of them have limitations in the specific identification, especially in the case of mixed infections and hybrids (Huyse et al. 2009; Zhao et al. 2012a). Increasing evidence from experimental and human studies has shown that polymerase chain reaction (PCR) and PCR-coupled methods improve the diagnosis of Schistosoma infection as well as the identification of Schistosoma species (Gobert et al. 2005; Sandoval et al. 2006; Kato-Hayashi et al. 2010; Lv et al. 2015). Although these techniques are useful and effective, the electrophoresis or sequencing analysis can be, in some cases, quite time consuming and expensive, rendering such tools impracticable for large-scale molecular epidemiological studies (Lymbery and Thompson 2012).
The high-resolution melt (HRM) technique uses a single closed tube in which PCR is followed by precise dissociation (melting) behavior determination of double-stranded DNA amplicons. The accuracy of the melting curve is maximized by acquiring fluorescence data over small temperature increments (as low as 0.02 °C) (Price et al. 2007). Samples can be discriminated according to sequence, length, GC content, or strand complementarity, down to single base-pair changes. The intercalating double-stranded DNA-binding dyes used do not inhibit PCR reactions at the concentrations necessary for them to fully saturate the target DNA duplexes and ensures the accuracy of experimental results. Such a probe-free assay is easier and more cost-effective and, unlike conventional methods, it has no manual post-PCR processing, thus avoiding the risk of contamination with PCR products. This method greatly simplifies the operation and reduces the analysis time, making it suitable for broad application. In the area of parasitology, it has been adapted recently for the genotyping of apicomplexan parasites (Al-Mohammed 2011; Costa et al. 2011; Ngui et al. 2012; Zhang et al. 2012; Higuera et al. 2013; Salim et al. 2013).
In the present study, we established a simple and effective real-time PCR assay coupled with HRM analysis, using the 18S ribosomal DNA (rDNA) region as the genetic marker, for the detection and differentiation of the four main Schistosoma spp. infecting humans, and discuss the value of such a tool in rapid screening and detection of closely related species in a laboratory.
Materials and methods
Parasites and isolation of genomic DNA
Twenty-five worm samples were originated from endemic areas in Puerto Rico, Cameroon, Egypt, China, Philippines, Japan, and Cambodia (Table 1). No specific permission was required for the collection of intermediate host snails of Schistosoma spp. Field studies did not involve any endangered or protected species. No ethical approval was required for the present study because all the parasite samples used in the present study had been prepared and used in previous studies (Sugiyama et al. 1997; Li et al. 2011; Zhao et al. 2010; 2012b). All S. japonicum worms in China were collected from rabbits exposed to cercariae from infected snails (Oncomelania hupensis) in our laboratory (Li et al. 2011; Zhao et al. 2010; 2012b), while S. japonicum from the Philippines and Japan were isolated from experimentally infected mice (Sugiyama et al. 1997). For S. mekongi, S. mansoni, and S. haematobium, worms were collected from rabbits or mice experimentally infected with cercariae from the relevant intermediate snail hosts (Neotricula aperta, Biomphalaria glabrata, and Bulinus spp., respectively). These worms were provided by Dr. Hiroshi Ohmae, Department of Parasitology, National Institute of Infectious Diseases, and by Dr. Mona S. Mahmoud in the Department of Parasitology and Animal Diseases, National Research Center, Dokki, Giza, Egypt. All the Schistosoma specimens were fixed in 70 % molecular grade ethanol and stored at −20 °C until used for DNA extraction. Additionally, some adults of common zoonotic parasites, namely Fasciola hepatica, F. gigantica, Clonorchis sinensis, and Opisthorchis viverrini were used as “controls” for the evaluation of the specificity of the specific PCR-coupled HRM assay.
Total genomic DNA (gDNA) of each parasite was extracted using SDS/proteinase K treatment, column-purified (Wizard® SV Genomic DNA Purification System, Promega) and eluted into 60 μL H2O according to the manufacturer’s recommendations. An RNAase treatment step was also used. Then, the quality of parasite gDNA was confirmed by amplification of the first internal transcribed spacer of ribosomal DNA (van Herwerden et al. 1998).
Primers, Real-time PCR, and HRM analysis
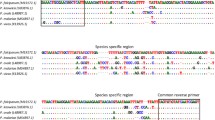
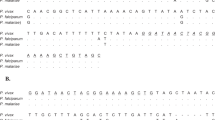
18S rDNA sequence has been identified as good genetic marker for identification and phylogenetic studies of Schistosoma members (Johnston et al. 1993; Attwood et al. 2002; Webster et al. 2006). The 18S rDNA sequence is also quite conserved within a Schistosoma species (Yu et al. 2000; Li et al. 2008). Our recent study also demonstrated that 35 S. japonicum isolates from mainland China, the Philippines, and Japan have almost identical 18S rDNA sequences, with only one nucleotide variation (Chen et al. 2011). Therefore, a portion of the 18S rDNA of four Schistosoma species was chosen as the genetic marker. A pair of primers for real-time PCR coupled with HRM analysis was designed specific for the V4 region (∼280 bp) of the nuclear 18S rDNA of four Schistosoma species (based on GenBankTM accession numbers AY157226, AY157228, SMU65657, Z11976). Primer names and sequences are as follows: M182U (5′-GGTAACTCCAGCTCCAAAAGCGTAT-3′) and M182D (5′-TTGTTCAAAGTAAAGATGCCGTCCG-3′). The real-time PCR conditions were optimized empirically. Finally, each PCR reaction (20 mL in volume) was performed in 10 μL 2× HRM PCR master mix (QIAGEN), 0.7 μM of each primer, and 1–50 pg genomic DNA in a Rotor-Gene 6000 (HRM)™ (QIAGEN) under the following thermocycling conditions: 5 min at 95 °C to activate HotStarTaq Plus DNA Polymerase, then 40 cycles of three steps: 10 s denaturation at 95 °C, 30 s annealing at 55 °C, and 10 s extension at 72 °C. After amplification, a melt step in the range 78 to 89 °C in 0.2 °C increments, pausing for 2 s per step, was initiated. The increase in Evagreen fluorescence was monitored in real time during the PCR, and the subsequent decrease during the melt phase was measured by acquiring each step to the green channel (470-nm excitation and 510-nm emission) of the Rotor-Gene 6000. Genotypes were then determined by examining normalized and difference melt plots using the Corbett Rotor-Gene 6000 software (Rotor-Gene Q Series Software 2.0.2) with a confidence limit specified as 80 % (a score above this signifying that a sample can be confidently assigned to a particular genotype, and a score below this indicating a sequence variation).
Cycle threshold (Ct) values and amplification efficiency were assessed to identify outliers (samples of poor quality) that were then filtered out of subsequent HRM analysis. Ct values would be ideally similar for samples and reference. Therefore, amplicons from randomly chosen S. japonicum, S. mansoni, S. haematobium, and S. mekongi individuals were purified and built into recombinant plasmids to establish standard references (namely Sjap-B, Sman-B, Shae-B, Smek-B, respectively, Table 1). Each plasmid stock was diluted so as to give a reaction Ct value between 10 and 20 when used as a standard. To examine their consistency, all standard references of each Schistosoma species were tested in triplicate and repeated on several different days using similar reagents and DNA concentrations. After determining the optimized reaction system and conditions, all samples under test were diluted (ddH2O dilution) or concentrated (DNAmate, TaKaRa, D605A) to match the Ct value of the standards. Any DNA with a Ct ≥ 30 was removed, because it indicated too little starting template or sample degradation. Reactions with amplification efficiency different from the standards or with efficiency less than 1.4 were omitted from evaluation as outliers (Poláková et al. 2008).
The identity of each amplicon was further confirmed by sequencing (Sangong Biotech, China). The acquired sequences were aligned with those of published Schistosoma species from GenBankTM, and phylogenetic analyses (MEGA5.2) (Tamura et al. 2011) were conducted subsequently to verify the accuracy of the genotype results obtained by the established HRM analysis.
Assessment of specificity and sensitivity of the detection assay
To evaluate the specificity of the designed HRM assay, some of the common zoonotic trematodes which are genetically closely related to Schistosoma and used in previous studies (Zhao et al. 2010, 2012b), namely, Fasciola hepatica, F. gigantica, Clonorchis sinensis, and Opisthorchis viverrini, were chosen as “controls.” For the sensitivity analysis, the concentration of DNA in each Schistosoma sample was determined using a spectrophotometer, and then tested in tenfold serial dilutions. The detection limit was determined as the smallest genomic DNA amount at which Ct value <30.
Validation of the HRM assay for identification and differentiation of isolates belongs to the four Schistosoma species
After optimization, 21 representative specimens representing four Schistosoma species from 15 geographical localities in seven endemic countries were randomly chosen to assess the capacity of the real-time PCR coupled with HRM assay (Table 1).
Results and discussion
Alignment of sequences from Schistosoma spp. showed that the ribosomal 18S gene is highly conserved within a species but exhibits some consistent interspecific differences (Johnston et al. 1993). One pair of primers (M182U and M182D), based on conserved portions of the sequences of the V4 region of the 18S rDNA, generated an amplicon from all tested Schistosoma species that could be used for detection and specific identification of these. The primers did not amplify host DNA. Samples representing four Schistosoma species were randomly chosen to establish the standard reaction. All samples amplified a product of about 280 bp with a good Ct value around 10 to 20. Unique HRM profiles were produced for each of the four Schistosoma species (Fig. 1). Nucleotide sequence analyses and phylogenetic analysis of the amplicons confirmed the identity of each species (not shown).
The specificity of the detection assay was examined by comparison with some other common zoonotic trematodes. Amplicons could be produced from all worm species using these primers, but visual assessment of the conventional and normalized curves could distinguish all species, with the genotypes of other trematodes represented as “variation” after analysis of the melt curve by the software (Fig. 2, Table 2). The assay sensitivity was assessed by using tenfold serial dilutions of predefined Schistosoma genomic DNA (recombinant plasmids). Besides, there was a good linear correlation coefficient (R 2 = 0.992) between the log concentrations of purified DNA (1–10-6 ng) and the Ct value (Fig. 3). The lower limit of detection was 10-5 ng (Fig. 3), which was four orders of magnitude more sensitive than conventional agarose electrophoresis (Li et al. 2011; Zhao et al. 2012b) and three orders of magnitude higher than polyacrylamide gel detection (Zhao et al. 2010). Our detection assay displayed a high degree of sensitivity and reproducibility.
Specificity of HRM analysis of normalized and temperature-shifted melting curves of Schistosoma spp. and other common zoonotic trematodes. Sjap S. japonicum, Smek S. mekongi, Sman S. mansoni, Shae S. haematobium, NC negative control. Melting peaks of trematodes, Bin A includes S. japonicum and S. mekongi, Tm, 83.65 °C; Bin B represents S. mansoni, Tm, 85.65 °C; Bin C represents S. haematobium, Tm, 85.85 °C; Bin D includes C. sinensis and O. viverrini, Tm, 85.78; and Bin E includes Fasciola hepatica and Fasciola gigantica, Tm, 86.30 °C
Assessment of sensitivity of the HRM assay for detection of Schistosoma spp. a Lanes 1–7 represent serial dilutions of genomic DNA of S. japonicum, 1–7: dilutions 1 to 10-6 ng, respectively. b A linear correlation is shown between the Ct and the tenfold diluted DNA of Schistosoma spp. NC negative control
To assess the capacity of HRM curve analysis for the detection and identification of specimens of different species, 21 individual specimens of Schistosoma from different endemic regions were analyzed: S. mansoni (n = 3), S. haematobium (n = 3), S. mekongi (n = 4), and S. japonicum (n = 11). Visual assessment of the conventional and normalized curves could distinguish all species (Fig. 4, Table 2) with high confidence values. No signal was detected in the negative controls. The melting curves were characterized by peaks of 83.65 °C for S. japonicum and S. mekongi, 85.65 °C for S. mansoni, and 85.85 °C for S. haematobium (Fig. 4). Although S. japonicum and S. mekongi melting peaks were identical to each other, their melting profiles are significantly different. In S. mekongi, a minor peak on the left shoulder of the major peak is a distinctive feature (Fig. 4). These are known to be closely related species from Asia (Chen et al. 2011, 2015), so the similarity is not surprising. S. mansoni and S. haematobium are both from Africa and rather distantly related to the Asian species (Chen et al. 2011; Zhao et al. 2012c).
Real-time PCR and HRM analyses for all Schistosoma spp. samples. a Normalized melting curves for Schistosoma spp. isolates. b Melting peaks of Schistosoma spp. Bin A includes S. japonicum and S. mekongi, Tm, 83.65 °C; Bin B includes S. mansoni, Tm, 85.65 °C; and Bin C includes S. haematobium, Tm, 85.85 °C. Sjap S. japonicum, Smek S. mekongi, Sman S. mansoni, Shae S. haematobium, NC negative control
Detection of characteristic eggs of parasitic worms, including Schistosoma species, is the “gold standard” for diagnosis. However, direct detection of eggs is not possible in prepatent infections, and difficult in low-endemicity areas, hence rendering specific diagnosis difficult especially in travelers and expatriates with mixed infections (Meltzer and Schwartz 2013).
In recent years, with the increased numbers of travelers and expatriates worldwide, the potential risk of Schistosoma spp. coinfection in endemic areas or discovery of infection in returning travelers and immigrants has attracted broad attention. Development of a rapid and easy screening method for identification and differentiation of species of blood flukes is a problem to be solved. Many PCR and real-time PCR methods (Kato-Hayashi et al. 2010; Cnops et al. 2012; Ten et al. 2008) have been previously reported but showed limitations for rapid interspecies identification. Marx et al. (2000) used a genome-wide melting curve scanning method to distinguish between S. mansoni, S. japonicum, and S. haematobium. Then, Kongklieng et al. (2013) made real-time PCR with high-resolution melting analysis to differentiate S. japonicum and S. mekongi. Compared with traditional melting curve analysis of Marx et al. (2000), the HRM detection assay established in the present study provides more information than just the calculated melting temperature (Tm), permitting differentiation on the basis of curve shape, even when they define the same Tm values (Jin et al. 2012). In addition, the detection method for two schistosoma species established by Kongklieng et al. (2013) showed limited detecting scale compared with the present study which made rapid detection and identification of four major Schistosoma species. Furthermore, the entire process from PCR to HRM curve analysis can be completed within 3 h, which is a great time saving.
In conclusion, the present study developed a real-time PCR coupled with HRM analysis assay for detection and differential identification of S. mansoni, S. haematobium, S. japonicum, and S. mekongi. This method has the advantage of being rapid, sensitive, and inexpensive. It has important implications for epidemiological studies and could be applicable to diagnosis of a wide range of microorganism species, especially when these are closely related.
References
Al-Mohammed HI (2011) Genotypes of Giardia intestinalis clinical isolates of gastrointestinal symptomatic and asymptomatic Saudi children. Parasitol Res 108:1375–1381
Attwood SW, Upatham ES, Meng XH, Qiu DC, Southgate VR (2002) The phylogeography of Asian Schistosoma (Trematoda: Schistosomatidae). Parasitology 125:99–112
Chen F, Li J, Sugiyama H, Weng YB, Zou FC, Lin RQ, Yuan ZG, Song HQ, Zhu XQ, Zhao GH (2011) Comparative analysis of 18S and 28S rDNA sequences of Schistosoma japonicum from mainland China, the Philippines and Japan. J Anim Vet Adv 10:2010–2015
Chen F, Li J, Sugiyama H, Zhou DH, Song HQ, Zhao GH, Zhu XQ (2015) Genetic variability among Schistosoma japonicum isolates from the Philippines, Japan and China revealed by sequence analysis of three mitochondrial genes. Mitochondrial DNA 26:35–40
Cnops L, Tannich E, Polman K, Clerinx J, Van Esbroeck M (2012) Schistosoma real-time PCR as diagnostic tool for international travellers and migrants. Trop Med Int Health 17:1208–1216
Clerinx J, Van Gompel A (2011) Schistosomiasis in travellers and migrants. Travel Med Infect Dis 9:6–24
Costa JM, Cabaret O, Moukoury S, Bretagne S (2011) Genotyping of the protozoan pathogen Toxoplasma gondii using high-resolution melting analysis of the repeated B1 gene. J Microbiol Methods 86:357–363
Gryseels B (2012) Schistosomiasis. Infect Dis Clin North Am 26:383–397
Gryseels B, Polman K, Clerinx J, Kestens L (2006) Human schistosomiasis. Lancet 368:1106–1118
Gobert GN, Chai M, Duke M, McManus DP (2005) Copro-PCR based detection of Schistosoma eggs using mitochondrial DNA markers. Mol Cell Probes 19:250–254
Hotez PJ, Molyneux DH, Fenwick A, Kumaresan J, Sachs SE, Sachs JD, Savioli L (2007) Control of neglected tropical diseases. N Engl J Med 357:1018–1027
Huyse T, Webster BL, Geldof S, Stothard JR, Diaw OT, Polman K, Rollinson D (2009) Bidirectional introgressive hybridization between a cattle and human schistosome species. PLoS Pathog 5:e1000571
Higuera SL, Guhl F, Ramirez JD (2013) Identification of Trypanosoma cruzi Discrete Typing Units (DTUs) through the implementation of a High-Resolution Melting (HRM) genotyping assay. Parasit Vectors 6:112
Johnston DA, Kane RA, Rollinson D (1993) Small subunit (18S) ribosomal RNA gene divergence in the genus Schistosoma. Parasitology 107:147–156
Jin D, Luo Y, Zhang Z, Fang W, Ye J, Wu F, Ding G (2012) Rapid molecular identification of Listeria species by use of real-time PCR and high-resolution melting analysis. Fems Microbiol Lett 330:72–80
Kato-Hayashi N, Kirinoki M, Iwamura Y, Kanazawa T, Kitikoon V, Matsuda H, Chigusa Y (2010) Identification and differentiation of human schistosomes by polymerase chain reaction. Exp Parasitol 124:325–329
Kongklieng A, Kaewkong W, Intapan PM, Sanpool O, Janwan P, Thanchomnang T, Lulitanond V, Sri-Aroon P, Limpanont Y, Maleewong W (2013) Molecular differentiation of Schistosoma japonicum and Schistosoma mekongi by real-time PCR with high resolution melting analysis. Korean J Parasitol 51:651–656
Li HJ, Liang YS, Dai JR, Tao YH, Wang W, Qu GL, Wei JY (2008) Homology of 18S small subunit ribosomal RNA gene among species and strains of Schistosoma and sensitivity of PCR assay to detect single cercaria. Chin J Schistosomiasis Control 6:418–422 (in Chinese)
Li J, Chen F, Sugiyama H, Blair D, Lin RQ, Zhu XQ (2015) A specific indel marker for the Philippines Schistosoma japonicum revealed by analysis of mitochondrial genome sequences. Parasitol Res 114:2697–2704
Li J, Zhao GH, Chen F, Song HQ, Zhu XQ, Zhao GH, Li J, Chen F, Lin RQ, Weng YB, Mahmoud MS, Zou FC (2011) A cleaved amplified polymorphic sequence (CAPS) method for the identification of geographical isolates of Schistosoma japonicum in China. Ann Trop Med Parasitol 105:531–536
Lv Z, Wu Z, Zhang L, Ji P, Cai Y, Luo S, Wang H, Li H (2015) Genome mining offers a new starting point for parasitology research. Parasitol Res 114:399–409
Lymbery AJ, Thompson RC (2012) The molecular epidemiology of parasite infections: tools and applications. Mol Biochem Parasitol 181:102–116
Marx KA, Bizzaro JW, Blake RD, Hsien TM, Feng TL (2000) Experimental DNA melting behavior of the three major Schistosoma species. Mol Biochem Parasitol 107:303–307
Meltzer E, Schwartz E (2013) Schistosomiasis: current epidemiology and management in travelers. Curr Infect Dis Rep 15:211–215
Muth S, Sayasone S, Odermatt-Biays S, Phompida S, Duong S, Odermatt P (2010) Schistosoma mekongi in Cambodia and Lao People’s Democratic Republic. Adv Parasitol 72:179–203
Ngui R, Lim YA, Chua KH (2012) Rapid detection and identification of human hookworm infections through high resolution melting (HRM) analysis. PLoS One 7:e41996
Price EP, Smith H, Huygens F, Giffard PM (2007) High-resolution DNA melt curve analysis of the clustered, regularly interspaced short-palindromic-repeat locus of Campylobacter jejuni. Appl Environ Microbiol 73:3431–3436
Poláková KM, Lopotová T, Klamová H, Moravcová J (2008) High-resolution melt curve analysis: initial screening for mutations in BCR-ABL kinase domain. Leuk Res 32:1236–1243
Salim B, Bakheit MA, Sugimoto C (2013) Rapid detection and identification of Theileria equi and Babesia caballi by high-resolution melting (HRM) analysis. Parasitol Res 112:3883–3886
Sandoval N, Siles-Lucas M, Perez-Arellano JL, Carranza C, Puente S, López-Abán J, Muro A (2006) A new PCR-based approach for the specific amplification of DNA from different Schistosoma species applicable to human urine samples. Parasitology 133:581–587
Sugiyama H, Kawanaka M, Kameoka Y, Nakamura M (1997) A novel cDNA clone of Schistosoma japonicum encoding the 34,000 Dalton eggshell precursor protein. Int J Parasitol 27:811–817
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Ten HR, Verweij JJ, Vereecken K, Polman K, Dieye L, van Lieshout L (2008) Multiplex real-time PCR for the detection and quantification of Schistosoma mansoni and S. haematobium infection in stool samples collected in northern Senegal. Trans R Soc Trop Med Hyg 102:179–185
van Herwerden L, Blair D, Agatsuma T (1998) Intra- and inter-specific variation in nuclear ribosomal internal transcribed spacer 1 of the Schistosoma japonicum species complex. Parasitology 116:311–317
Webster BL, Southgate VR, Littlewood DT (2006) A revision of the interrelationships of Schistosoma including the recently described Schistosoma guineensis. Int J Parasitol 36:947–955
WHO (2008) The global burden of disease, 2004 update. WHO website. Available: http://www.who.int/healthinfo/global_burden_disease/2004_report_update/en/
Yu XC, Wu GL, Zhang YJ, Wu YQ, Zhu CL, Zhang ZH, Chen BY (2000) Studies of the genetic polymorphism of Schistosoma japonicum (Chinese mainland strains). Yi Chuan Xue Bao 27:498–505 (in Chinese)
Zhang P, Liu Y, Alsarakibi M, Li J, Liu T, Li Y, Li G (2012) Application of HRM assays with EvaGreen dye for genotyping Giardia duodenalis zoonotic assemblages. Parasitol Res 111:2157–2163
Zhao GH, Li J, Lin RQ, Zou FC, Liu W, Yuan ZG, Mo XH, Song HQ, Weng YB, Zhu XQ (2010) An effective sequence characterized amplified region-PCR method derived from restriction site-amplified polymorphism for the identification of female Schistosoma japonicum of zoonotic significance. Electrophoresis 31:641–647
Zhao GH, Li J, Blair D, Li XY, Elsheikha HM, Lin RQ, Zou FC, Zhu XQ (2012a) Biotechnological advances in the diagnosis, species differentiation and phylogenetic analysis of Schistosoma spp. Biotechnol Adv 30:1381–1389
Zhao GH, Li J, Song HQ, Li XY, Chen F, Lin RQ, Yuan ZG, Weng YB, Hu M, Zou FC, Zhu XQ (2012b) A specific PCR assay for the identification and differentiation of Schistosoma japonicum geographical isolates in mainland China based on analysis of mitochondrial genome sequences. Infect Genet Evol 12:1027–1036
Zhao GH, Li J, Mo XH, Li XY, Lin RQ, Zou FC, Weng YB, Song HQ, Zhu XQ (2012c) The second transcribed spacer rDNA sequence: an effective genetic marker for inter-species phylogenetic analysis of trematodes in the order Strigeata. Parasitol Res 111:1467–1472
Acknowledgments
Project support was provided by the “Special Fund for Agro-scientific Research in the Public Interest” (Grant No. 201303037) and the Science Fund for Creative Research Groups of Gansu Province (Grant No. 1210RJIA006) to XQZ, and the Pearl River Science and Technology New Star Project (Grant No. 2014 J2200096) and the President’s Foundation of Guangdong Academy of Agricultural Sciences (Grant No. 201413) to JL.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, J., Zhao, GH., Lin, R. et al. Rapid detection and identification of four major Schistosoma species by high-resolution melt (HRM) analysis. Parasitol Res 114, 4225–4232 (2015). https://doi.org/10.1007/s00436-015-4660-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-015-4660-3