Abstract
Cryptosporidium parvum DNA from 175 neonatal calves on 16 farms in eight eastern states in the United States was subtyped by sequence analysis of the 60-kDa glycoprotein gene to determinate the parasite genetic diversity. Six subtypes of the IIa subtype family were found. Subtype IIaA15G2R1, which is the predominant C. parvum subtype in calves in many parts of the world, was identified in 77% of the C. parvum DNA from calves. Several farms had more than one C. parvum subtype and a few calves had infections with mixed subtypes. Distribution of subtypes differed geographically. Diversity of C. parvum in calves in eastern United States was lower than that previously seen in Michigan and southern Ontario. The high prevalence of one subtype in calves worldwide and frequent detection of this subtype in humans suggests that parasite fitness probably plays an important role in transmission of cryptosporidiosis among cattle and in zoonotic infections.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
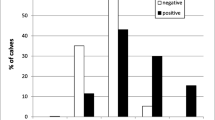
Cattle are infected with at least four Cryptosporidium species: Cryptosporidium parvum, Cryptosporidium bovis, Cryptosporidium andersoni, and the Cryptosporidium deer-like genotype (Xiao et al. 2004; Fayer et al. 2006). A recent study showed that these Cryptosporidium spp. in cattle are age-related (Santin et al. 2004). C. parvum, the only prevalent zoonotic species in cattle, is responsible for about 85% of the Cryptosporidium infections in preweaned calves but only 1% of the Cryptosporidium infections in postweaned calves. Postweaned calves and older cattle are mostly infected with C. bovis, C. andersoni, and the Cryptosporidium deer-like genotype (Santin et al. 2004; Fayer et al. 2006). These findings clearly demonstrate that only neonatal calves are an important source of zoonotic cryptosporidiosis in humans. Neonatal calves are also the age group of cattle mostly affected by cryptosporidiosis in terms of prevalence of infection and the associated morbidity and mortality (Fayer et al. 1997).
Despite the clinical and public health importance of C. parvum, little is known about its transmission dynamics in cattle. Currently, the maintenance of the parasites on cattle farm and the role of herd-to-herd transmission in cryptosporidiosis epidemiology are not known. Recently, researchers have used highly discriminatory subtyping techniques to study the transmission of bovine C. parvum infections in a geographic area (Mallon et al. 2003a,b; Tanriverdi et al. 2006; Tanriverdi and Widmer 2006). These tools are very useful in tracking infection sources and examining transmission dynamics. The most commonly used subtyping tool is based on sequence analysis of the 60-kDa glycoprotein gene (GP60), which enables the identification of many subtype families of C. parvum and Cryptosporidium hominis and several subtypes within each subtype family (Strong et al. 2000; Peng et al. 2001, 2003a,b; Sulaiman et al. 2001, 2005; Glaberman et al. 2002; Leav et al. 2002; Alves et al. 2003, 2006; Sturbaum et al. 2003; Wu et al. 2003; Zhou et al. 2003; Chalmers et al. 2005; Abe et al. 2006; Trotz-Williams et al. 2006). Of the two major C. parvum subtype families, the IIa subtype family is zoonotic, seen in both humans and calves, whereas the IIc subtype family is anthroponotic and is found only in humans (Alves et al. 2003; Xiao and Ryan 2004).
In the present study, C. parvum specimens from cattle farms in eight eastern states in the United States were subtyped and compared with results previously obtained from cattle farms in Michigan and neighboring Ontario, Canada.
Materials and methods
Specimens
All specimens used in this study were diagnosed as positive for Cryptosporidium by microscopy of immunofluorescence-stained fecal materials. They were further identified as C. parvum by a small subunit (SSU) rRNA-based nested PCR and DNA sequencing as previously described (Santin et al. 2004). A total of 189 C. parvum-positive specimens from neonatal calves less than 8 weeks old were used in the study. They were collected in 1996 and 1997 in four dairy farms in Ohio and in 2002 on two farms each in New York, Maryland, Virginia, North Carolina, and Florida and one farm each in Vermont and Pennsylvania (Table 1). At least 15 specimens were collected from each farm, but only those positive for C. parvum were used in the present study.
DNA extraction and PCR
Total DNA was extracted from Cryptosporidium oocysts, concentrated by cesium chloride or sucrose floatation, using a QIAamp DNA Mini Kit (Qiagen, Valencia, CA, USA). A fragment of the GP60 gene of approximately 850 bp was amplified by nested PCR as previously described, using 1 or 2 μl of the extracted DNA in primary PCR (Alves et al. 2003). To reduce PCR inhibition, 400 ng/μl of nonacetylated bovine serum albumin (Sigma, St. Louis, MO, USA) was used in primary PCR. Each DNA was analyzed by PCR twice with the aim to generate at least one positive GP60 product for each C. parvum specimen.
DNA sequencing and phylogenetic analysis
PCR products were sequenced in both directions with the forward and reverse primers used in secondary PCR and an intermediary sequencing primer (5′-GAGATATATCTTGTTGCG-3′). In most cases, two PCR products from each specimen were sequenced. The nucleotide sequences obtained in this study were aligned with reference sequences retrieved from the GenBank using the program ClustalX (ftp://ftp-igbmc.u-strasbg.fr/pub/ClustalX/). The recently proposed nomenclature was used in naming C. parvum subtypes (Sulaiman et al. 2005).
Results
GP60 sequences
Of the 189 DNA extractions from neonatal calves previously diagnosed to be infected with C. parvum, 175 generated GP60 PCR products of the expected size in nested PCR. Those that failed in GP60 PCR mostly had inconsistent amplification in repeated SSU rRNA PCR analyses, indicative of low levels of infections. The secondary GP60 PCR products were all sequenced successfully with the forward and reverse primers used in the secondary PCR and the intermediary sequencing primer. The alignment of sequences obtained with reference sequences downloaded from GenBank indicated that all sequences obtained from the study belonged to the C. parvum subtype family IIa. Almost all sequences were identical to each other in the nonrepeat region, except that some sequences had two copies of the sequence ACATCA immediately after the trinucleotide repeats instead of one copy. In the trinucleotide repeat region, all sequences obtained had two copies of the TCG repeat and 11–19 copies of the TCA repeat.
GP60 IIa subtypes
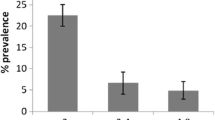
Altogether, six C. parvum IIa subtypes were found in the neonatal calves. They were named based on the number of TCA and TCG repeats in the trinucleotide repeat region and the ACATCA sequence, using the recently proposed nomenclature system (Sulaiman et al. 2005). The most common subtype was IIaA15G2R1 subtype, which had 15 copies of the TCA repeat (represented by the letter A), two copies of the TCG repeat (represented by the letter G), and one copy of the ACATCA sequence (represented by the letter R). This subtype was seen in 135 of the 175 GP60-positive animals. A similar subtype but with two copies of the ACATCA sequence, IIaA15G2R2, was seen in 11 of the 175 positive animals. The other four subtypes were IIaA11G2R1, IIaA17G2R1, IIaA18G2R1, and IIaA19G2R1, seen in 11, 10, 7, and 4 animals, respectively (Table 1).
Frequency of GP60 subtypes on farms
The subtype IIaA15G2R1 was also the most widely distributed C. parvum, seen on 14 of the 16 study farms and in all states except Florida (Table 1). Other subtypes were much more geographically restricted, with IIaA15G2R2 and IIaA11G2R1 each seen on three farms in two states and IIaA17G2R1 on two farms in two states. In contrast, IIaA18G2R1 and IIaA19G2R1 were seen on only one farm in Florida. Over half of the farms had only one C. parvum subtype in calves. However, one farm each in Florida and Ohio and two farms in New York each had two C. parvum subtypes, and the farm in Vermont had four subtypes in calves. Two calves in Vermont and one calf in Florida were concurrently infected with two C. parvum subtypes (Table 1).
Discussion
Multiple C. parvum subtypes were identified in DNA extracts from feces collected from neonatal calves. Six IIa subtypes were identified from calves on 16 farms in eastern United States. Previously, the transmission of C. parvum in calves was examined by GP60 sequence typing with modest numbers of specimens only in Michigan, southern Ontario, and Portugal (Alves et al. 2003, 2006; Peng et al. 2003b; Trotz-Williams et al. 2006). Seven, six, and two IIa subtypes were found in 36, 34, and 72 bovine specimens subtyped, respectively (Table 2). Considering previous results from Michigan and Ontario and the much larger sample size in the present study, the finding of only six IIa subtypes in calves on diverse farms in eastern United States is surprising. It is possible that C. parvum genetic diversity is greater in northern states and provinces in North America, as the number of GP60 IIa subtypes seen in Michigan and Ontario in previous studies and Vermont in the present study is higher than the number of subtypes found in other states. As in Michigan (Peng et al. 2003b), several farms in the present study had more than one C. parvum subtypes in calves at the time of sampling. In addition, two calves in Vermont and one calf in Florida had concurrent infection of mixed IIa subtypes (Table 1).
The subtype IIaA15G2R1 was responsible for 77% of C. parvum infections in calves in this study, but for a much lower percentage in Michigan and Ontario (Table 2). In contrast, 85% of C. parvum infections in calves and other ruminants in Portugal had this subtype. On most farms where IIaA15G2R1 subtype was present few other subtypes were found, whereas other subtypes were found mostly on farms where IIaA15G2R1 did not predominate (Table 1). These findings parallel the low subtype diversity found in calves in Portugal where IIaA15G2R1 is the major subtype and the high subtype diversity in Michigan and Ontario where IIaA15G2R1 is less frequent (Table 2).
The high prevalence of IIaA15G2R1 in calves in the United States and Canada may be due to the fitness of the parasite, as it is also the predominant IIa subtype in calves and humans all over the world (Table 2). So far, it has been seen with high frequency in the United States, Canada, Portugal, Slovenia, Australia, Kuwait, and Japan (Table 2). The only possible exception is the United Kingdom, where it was seen less frequently in humans who were infected with many other IIa subtypes (Table 2) (Glaberman et al. 2002; Chalmers et al. 2005). Many widely used laboratory isolates of C. parvum belong to this subtype, such as the Iowa, UCP, Tamu, KSU1, Texas, Maine Apple Cider, and Glasgow isolates (Table 2). It is interesting to note that subisolates of the Iowa isolate, passaged in different laboratories over many years, differ from each other by one nucleotide in the GP60 gene; the parasite from Pleasant Hill Farm has a GP60 sequence identical to IIaA15G2R1 seen worldwide, whereas the parasite from the University of Arizona has a “G” to “T” nucleotide change after the trinucleotide repeats, which results in an amino acid change and is not seen in any other IIa subtypes (Sturbaum et al. 2003).
Despite the high prevalence of IIaA15G2R1 worldwide, there is geographic segregation in some C. parvum subtypes. The distribution of IIa subtypes in calves and humans in Michigan and neighboring southern Ontario is very similar, and most of the IIa subtypes there have 16 copies of the TCA repeat. Several subtypes common in Michigan and Ontario, such as IIaA16G2R1 and IIaA16G3R1, were not seen in this study (Table 2). Likewise, IIaA18G2R1 and IIaA19G1 were seen in calves on only one farm in Florida in this study, and many of the IIa subtypes seen in humans in Northern Ireland were not seen in other areas (Table 2). Even though an earlier multilocus typing study in Scotland showed no geographic isolation of C. parvum subtypes (Mallon et al. 2003b), results of a recent similar study in Turkey and Israel demonstrated the occurrence of geographically unique C. parvum subtypes in calves (Tanriverdi et al. 2006).
The C. parvum subtype IIaA15G2R1 is also the most commonly identified zoonotic C. parvum infection in humans. It was found in humans in the United States, United Kingdom, Portugal, Slovenia, Australia, Japan, and Kuwait (Table 2). It is likely the major subtype responsible for zoonotic cryptosporidiosis in areas with intensive animal husbandry, even though an endemicity of IIaA15G2R1 was found in children in Kuwait City (Sulaiman et al. 2005). Three other subtypes seen in calves in this study, IIaA15G2R2, IIaA17G2R2, and IIaA19G2R1, were also previously reported in humans (Table 2), indicating that all IIa subtypes have the potential to infect humans. The anthroponotic C. parvum IIc subtype family, which is the most common type of C. parvum in humans in most countries (Alves et al. 2003; Xiao et al. 2003; Xiao and Ryan 2004), were expectedly not seen in calves in this and any previous studies. The C. parvum IId subtype family previously seen in a few calves and humans in Portugal was not detected in US calves (Alves et al. 2006), demonstrating that this group of C. parvum is also not a major zoonotic pathogen in North America.
In addition to the studies of transmission of C. parvum in calves, GP60 subtyping was also used in tracking the source of contamination in waterborne and foodborne outbreaks of cryptosporidiosis (Glaberman et al. 2002; Xiao et al. 2003; Chalmers et al. 2005; Blackburn et al. 2006; Wheeler et al. 2006). Even though in several occasions a direct linkage at the subtype level between outbreak cases and the implicated food or water has been made (Glaberman et al. 2002; Blackburn et al. 2006), the power of the approach was compromised by the lack of baseline data on the distribution of C. parvum subtypes in the same geographic area. A noticeable exception is the investigation of an outbreak of cryptosporidiosis in Ohio in 2003 associated with consumption of ozonated apple cider in Blackburn et al. (2006). In this outbreak, two C. parvum subtypes, IIaA15G2R1 and IIaA17G2R1, were found in eight outbreak cases, and the IIaA17G2R1 subtype was further found in epidemiologically implicated apple cider. Existing data collected from dairy farms several years ago, reported now in the present study, have already demonstrated the occurrence of these two C. parvum subtypes in calves in Ohio, supporting the conclusion of environmental investigation that cattle might be a source of contamination of apples used in making the cider.
In conclusion, results of the subtyping study showed the uniqueness of C. parvum transmission on farms in eastern United States, with one predominant subtype in neonatal calves. The widespread occurrence of the subtype in calves in many parts of the world demonstrates the role of genetic fitness of the parasite in the transmission of cryptosporidiosis, and the report of this subtype in humans in many areas raises some concerns about the selection of highly infectious strains by intensive animal husbandry practices. The finding of apparent differences in the distribution of C. parvum subtypes in northern and southern regions in North America is also intriguing. Similar studies in other geographic areas and other agricultural settings would serve to increase our understanding of the transmission dynamics of cryptosporidiosis in cattle and the zoonotic potential of C. parvum.
References
Abe N, Matsubayashi M, Kimata I, Iseki M (2006) Subgenotype analysis of Cryptosporidium parvum isolates from humans and animals in Japan using the 60-kDa glycoprotein gene sequences. Parasitol Res 99:303–305
Alves M, Xiao L, Sulaiman I, Lal AA, Matos O, Antunes F (2003) Subgenotype analysis of Cryptosporidium isolates from humans, cattle, and zoo ruminants in Portugal. J Clin Microbiol 41:2744–2747
Alves M, Xiao L, Antunes F, Matos O (2006) Distribution of Cryptosporidium subtypes in humans and domestic and wild ruminants in Portugal. Parasitol Res 99:287–292
Blackburn BG, Mazurek JM, Hlavsa M, Park J, Tillapaw M, Parrish M, Salehi E, Franks W, Koch E, Smith F, Xiao L, Arrowood M, Hill V, da Silva A, Johnston S, Jones JL (2006) Outbreak of cryptosporidiosis associated with consumption of ozonated apple cider. Emerg Infect Dis 12:684–687
Chalmers RM, Ferguson C, Caccio S, Gasser RB, Abs El-Osta YG, Heijnen L, Xiao L, Elwin K, Hadfield S, Sinclair M, Stevens M (2005) Direct comparison of selected methods for genetic categorisation of Cryptosporidium parvum and Cryptosporidium hominis species. Int J Parasitol 35:397–410
Fayer R, Speer CA, Dubey JP (1997) The general biology of Cryptosporidium. In: Fayer R (ed) Cryptosporidium and cryptosporidiosis. CRC Press, Boca Raton, FL, pp 1–41
Fayer R, Santin M, Trout JM, Greiner E (2006) Prevalence of species and genotypes of Cryptosporidium found in 1–2-year-old dairy cattle in the eastern United States. Vet Parasitol 135:105–112
Glaberman S, Moore JE, Lowery CJ, Chalmers RM, Sulaiman I, Elwin K, Rooney PJ, Millar BC, Dooley JS, Lal AA, Xiao L (2002) Three drinking-water-associated cryptosporidiosis outbreaks, Northern Ireland. Emerg Infect Dis 8:631–633
Leav BA, Mackay MR, Anyanwu A, O’Connor RM, Cevallos AM, Kindra G, Rollins NC, Bennish ML, Nelson RG, Ward HD (2002) Analysis of sequence diversity at the highly polymorphic Cpgp40/15 locus among Cryptosporidium isolates from human immunodeficiency virus-infected children in South Africa. Infect Immun 70:3881–3890
Mallon M, MacLeod A, Wastling J, Smith H, Reilly B, Tait A (2003a) Population structures and the role of genetic exchange in the zoonotic pathogen Cryptosporidium parvum. J Mol Evol 56:407–417
Mallon ME, MacLeod A, Wastling JM, Smith H, Tait A (2003b) Multilocus genotyping of Cryptosporidium parvum Type 2: population genetics and sub-structuring. Infect Genet Evol 3:207–218
Peng MM, Matos O, Gatei W, Das P, Stantic-Pavlinic M, Bern C, Sulaiman IM, Glaberman S, Lal AA, Xiao L (2001) A comparison of Cryptosporidium subgenotypes from several geographic regions. J Eukaryot Microbiol 28S–31S
Peng MM, Meshnick SR, Cunliffe NA, Thindwa BD, Hart CA, Broadhead RL, Xiao L (2003a) Molecular epidemiology of cryptosporidiosis in children in Malawi. J Eukaryot Microbiol 50:557–559 (Suppl)
Peng MM, Wilson ML, Holland RE, Meshnick SR, Lal AA, Xiao L (2003b) Genetic diversity of Cryptosporidium spp. in cattle in Michigan: implications for understanding the transmission dynamics. Parasitol Res 90:175–180
Santin M, Trout JM, Xiao L, Zhou L, Greiner E, Fayer R (2004) Prevalence and age-related variation of Cryptosporidium species and genotypes in dairy calves. Vet Parasitol 122:103–117
Stantic-Pavlinic M, Xiao L, Glaberman S, Lal AA, Orazen T, Rataj-Verglez A, Logar J, Berce I (2003) Cryptosporidiosis associated with animal contacts. Wien Klin Wochenschr 115:125–127
Strong WB, Gut J, Nelson RG (2000) Cloning and sequence analysis of a highly polymorphic Cryptosporidium parvum gene encoding a 60-kilodalton glycoprotein and characterization of its 15- and 45-kilodalton zoite surface antigen products. Infect Immun 68:4117–4134
Sturbaum GD, Jost BH, Sterling CR (2003) Nucleotide changes within three Cryptosporidium parvum surface protein encoding genes differentiate genotype I from genotype II isolates. Mol Biochem Parasitol 128:87–90
Sulaiman IM, Lal AA, Xiao L (2001) A population genetic study of the Cryptosporidium parvum human genotype parasites. J Eukaryot Microbiol 24S–27S
Sulaiman IM, Hira PR, Zhou L, Al-Ali FM, Al-Shelahi FA, Shweiki HM, Iqbal J, Khalid N, Xiao L (2005) Unique endemicity of cryptosporidiosis in children in Kuwait. J Clin Microbiol 43:2805–2809
Tanriverdi S, Widmer G (2006) Differential evolution of repetitive sequences in Cryptosporidium parvum and Cryptosporidium hominis. Infect Genet Evol 6:113–122
Tanriverdi S, Markovics A, Arslan MO, Itik A, Shkap V, Widmer G (2006) Emergence of distinct genotypes of Cryptosporidium parvum in structured host populations. Appl Environ Microbiol 72:2507–2513
Trotz-Williams LA, Martin DS, Gatei W, Cama V, Peregrine AS, Martin SW, Nydam DV, Jamieson F, Xiao L (2006) Genotype and subtype analyses of Cryptosporidium isolates from dairy calves and humans in Ontario. Parasitol Res 99:346–352
Wheeler C, Vugla D, Thomas G, Beach M, Carnes S, Maier T, Gorman J, Xiao L, Arrowood M, Gilliss D, Werner SB (2006) Outbreak of cryptosporidiosis at a California waterpark: employee and patron roles and the long road toward prevention. Epidemiol Infect (in press)
Wu Z, Nagano I, Boonmars T, Nakada T, Takahashi Y (2003) Intraspecies polymorphism of Cryptosporidium parvum revealed by PCR-restriction fragment length polymorphism (RFLP) and RFLP-single-strand conformational polymorphism analyses. Appl Environ Microbiol 69:4720–4726
Xiao L, Ryan UM (2004) Cryptosporidiosis: an update in molecular epidemiology. Curr Opin Infect Dis 17:483–490
Xiao L, Bern C, Sulaiman IM, Lal AA (2003) Molecular epidemiology of human cryptosporidiosis. In: Thompson RCA, Armson A, Ryan UM (eds) Cryptosporidium: from molecules to disease. Elsevier, Amsterdam, pp 121–146
Xiao L, Fayer R, Ryan U, Upton SJ (2004) Cryptosporidium taxonomy: recent advances and implications for public health. Clin Microbiol Rev 17:72–97
Zhou L, Singh A, Jiang J, Xiao L (2003) Molecular surveillance of Cryptosporidium spp. in raw wastewater in Milwaukee: implications for understanding outbreak occurrence and transmission dynamics. J Clin Microbiol 41:5254–5257
Acknowledgements
We thank Dr. William Shulaw and Dr. Khalid Saeed of The Ohio State University for providing some specimens used in this study. The findings and conclusions in this report are those of the authors and do not necessarily represent the views of the Centers for Disease Control and Prevention.
Author information
Authors and Affiliations
Corresponding author
Additional information
Nucleotide sequence data reported in this paper are available in the GenBank database under the accession numbers DQ630514–DQ630519.
Rights and permissions
About this article
Cite this article
Xiao, L., Zhou, L., Santin, M. et al. Distribution of Cryptosporidium parvum subtypes in calves in eastern United States. Parasitol Res 100, 701–706 (2007). https://doi.org/10.1007/s00436-006-0337-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-006-0337-2