Abstract
Purpose
The fat mass- and obesity-associated (FTO) gene on chromosome 16q12.2 shows an intimate association with obesity and body mass index. Recently, research into the FTO gene and its expression product has attracted widespread interest due to the identification of FTO as an N6-methyladenosine (m6A) demethylase. FTO primarily regulates the m6A levels of downstream targets via their 3′ untranslated regions. FTO not only plays a critical role in obesity-related diseases but also is involved in the occurrence, development and prognosis of many types of cancer, such as acute myeloid leukaemia, glioblastoma and breast cancer. Currently, studies indicate that FTO is a crucial component of m6A modification, it regulates cancer stem cell function, and promotes the growth, self-renewal and metastasis of cancer cells. In this review, we summarized and analysed the data regarding the structural features and biological functions of FTO as well as its association with different cancers and possible molecular mechanisms.
Methods
We systematically reviewed the related literatures regarding FTO and its demethylation activity in many pathologic and physiological processes, especially in cancer-related diseases based on PubMed databases in this article.
Results
Mounting evidence indicated that FTO plays a critical role in occurrence, progression and treatment of various cancers, even acting as a cancer oncogene in acute myeloid leukaemia, research on which is no longer restricted to metabolic diseases such as obesity and diabetes.
Conclusion
Considering FTO’s critical role in many diseases, FTO may become a new promising target for the diagnosis and treatment of various diseases in the near future, especially for specific types of cancers, such as acute myeloid leukaemia, glioblastoma and breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Basic information for FTO
FTO gene discovery
In 1994, Van der Hoeven et al. (1994) first discovered a deletion, up to 1.6 Mb, on chromosome 8 in the fused-toe (Ft) mutant mouse that was created by insertional mutagenesis. A follow-up study reported that the deletion comprises three genes of unknown function (FTS, FTM and FTO) and another three genes from the Iroquois gene family (IRX3, IRX5 and IRX6) (Anselme et al. 2007). In 2007, during a study of type 2 diabetes mellitus (T2DM) in Europe, Frayling et al. (2007) found a cluster of single-nucleotide polymorphisms (SNPs) in the first intron of the FTO gene, which was associated with obesity as it affected the body mass index (BMI) in genome-wide association studies (GWAS), and the gene was later officially named the fat mass- and obesity-associated protein (FTO). The FTO gene exists only in vertebrates and marine algae, while no expression is observed in plants, fungi or invertebrate animals (Robbens et al. 2008). At present, studies have demonstrated that FTO is located in both the nucleus and cytoplasm and that a mobile fraction shuttles between both cellular compartments, possibly via a mechanism mediated by one of the Exportin 2 (XPO2) family of proteins (Gulati et al. 2014). Subsequent accumulating evidence suggested that the FTO gene was closely related to the occurrence and development of obesity (Dina et al. 2007; Scuteri et al. 2007; Zhang et al. 2010). However, the study of FTO is not limited to the obesity field. Recent research shows that this protein is likely to predispose patients to the onset and development of certain tumours, such as acute myeloid leukaemia (AML).
Structure of FTO
The FTO gene is 410.50 kb, contains 9 exons and is located on chromosome 16q12.2. This gene is widely expressed in many tissues at various development phases, especially in the brain regions (Frayling et al. 2007). As research has progressed, the understanding of the FTO gene, its expression products, its structure and its function have been further developed. Gerken et al. (2007) first showed that the FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase that catalyses the demethylation of 3-methylthymine in single-stranded DNA using Fe(II) and 2-oxoglutarate (2-OG) while producing succinate, formaldehyde and carbon dioxide. The FTO protein, consisting of an amino-terminal AlkB-like domain and a carboxy-terminal domain with a novel fold, which shows high sequence conservation, was identified as an AlkB-like DNA/RNA demethylase that has high affinities for 3-methylthymidine (3-meT) in single-stranded DNA and 3-methyluracil (3-meU) in single-stranded RNA (Gerken et al. 2007; Han et al. 2010; Sanchez-Pulido and Andrade-Navarro 2007; Jia et al. 2008a). Compared with other AlkB members, an extra loop in the FTO structure covers one side of the conserved jelly-roll motif, which allows it to specifically compete with unmethylated double-stranded DNA for binding to FTO (Han et al. 2010). The evidence suggests that this special structure plays a significant role in limiting the binding of FTO to double-stranded nucleic acids. Subsequent ground-breaking research indicated that the FTO protein has a high affinity for N6-methyladenosine (m6A) in messenger RNA (mRNA) and shows efficient demethylation activity (Jia et al. 2011). This FTO mRNA demethylase activity is currently gaining increased notoriety, which is drawing additional attention to its roles in various diseases.
Regulation of the FTO gene in obesity
At present, the FTO gene is generally considered a susceptibility gene that is strongly linked to obesity. In 2007, through a GWAS analysis among 490,032 autosomal single-nucleotide polymorphisms in Britons, 10 SNPs were identified (represented by rs9939609) in the FTO gene region on chromosome 16 (Frayling et al. 2007). This study found that adults carrying the rs9939609 A allele weighed approximately 1.5 kg more than adults without the allele and that the dominant homozygous allele increased the average weight by almost 3 kg, with a 1.67-fold increased odds ratio of obesity, compared to those without this risk allele (Frayling et al. 2007). Meanwhile, Dina et al. (2007) found that the FTO gene may contribute to early-onset and severe obesity in children and adults of European ancestry. In fact, the SNP sites in FTO show diverse characteristics in people from different regions, such as Africa, Spain, Australia, India, Japan and China (Chang et al. 2008; Hotta et al. 2008; Gonzalez-Sanchez et al. 2009; Fang et al. 2010; Grant et al. 2008; Ningombam et al. 2018). In general, the prevalence of the risk allele is obviously lower in Asian populations than that in European populations. Strangely, another study reported that three FTO variants (rs8050136, rs9939609, and rs9930506) seemed to be unrelated to obesity in the Chinese Han population (Li et al. 2008). The discrepancies in this study remain to be further confirmed by additional evidence.
In regard to the proposed mechanisms through which the FTO genetic polymorphisms are involved in triggering obesity, current evidence indicates that the correlation between FTO genetic polymorphisms and obesity is mostly driven by increased energy intake rather than by energy expenditure, as the carriers of the risk allele tend to consume high-calorie diets. The FTO gene is highly expressed in the hypothalamus, which is the critical region that regulates feeding and energy expenditure, implying that it may modulate the function of the feeding centre (Frayling et al. 2007; Gerken et al. 2007; McTaggart et al. 2011). Leptin functions to stimulate metabolism and facilitate weight loss. It was proven that some FTO obesity risk alleles (rs17817449 and rs9939609) are connected with lower serum levels of the satiety-enhancing adipokine leptin (Qi et al. 2008; Benedict et al. 2014), while other researchers take an opposing view of the related evidence (Arrizabalaga et al. 2014; Do et al. 2008). Therefore, whether leptin plays a role in the effect of FTO polymorphisms on obesity has not been thoroughly elucidated to date. Moreover, a decline in exercise levels was observed to enhance the risk of obesity. Andreasen et al. (2008) found that lack of physical activity might fortify the effect of the FTO rs9939609 polymorphism on body fat accumulation. To summarize, the association between the FTO gene and obesity may depend on appetite-regulating hormones and physical activity, but the related concrete mechanism is complicated and warrants further research.
More importantly, obesity is a risk factor for certain cancers, which suggests that FTO may be an important link between obesity and tumourigenesis. Obesity is considered to be closely related to more than 10 types of cancer, including colon cancer, post-menopausal breast cancer, endometrial cancer, pancreatic cancer, and advanced-stage prostate cancer (Pischon et al. 2008). An updated meta-analysis has demonstrated that FTO rs11075995 is associated with breast cancer and that this connection depends on BMI (Kang et al. 2017). The main factors connecting obesity to cancer are classified into three functional classes, including the insulin-IGF-1 axis, sex hormones, and adipokines (such as leptin and adiponectin), which are all involved in the secretion and regulation of adipose tissue (Pischon and Nimptsch 2016). Obesity-related inflammation may increase tumour burden and promote tumour growth, development and metastasis (Deng et al. 2016). Recently, FTO was shown to affect obesity and breast cancer, possibly through similar mechanisms, by regulating the Iroquois-related homeobox 3 (IRX3) gene expression level (Akbari et al. 2018). In fact, there is no clear molecular mechanism through which FTO affects obesity-mediated tumourigenesis and tumour progression. However, FTO seems to increase the risk of some cancers by promoting obesity.
Epigenetic research on the FTO gene
Epigenetic regulation of FTO gene expression
In recent years, research on the role of epigenetic regulation in various biological functions and in the pathogenesis of diseases has gained extensive attention. Epigenetic mechanisms, including DNA methylation, play critical roles in the regulation of gene expression. In mammals, more than 70% of the CpG (5′-C-phosphate-G-3′) cytosine residues in promoter regions are methylated (Jabbari and Bernardi 2004; Saxonov et al. 2006). Generally, CpG islands are hypomethylated in normal cells. It has been demonstrated that SNPs in intron 1 of FTO are associated with increased levels of FTO expression that affect the primary transcript levels, at least in blood cells and skin fibroblasts (Berulava and Horsthemke 2010). A number of studies have found that variations in DNA methylation at clusters of CpG methylation sites correlate with the susceptibility to T2DM. Toperoff et al. (2012, 2015) demonstrated that a CpG site in the first intron of the FTO gene exhibited small but significant hypomethylation in T2DM patients compared to controls; the same results were also found in peripheral blood leukocytes. It is notable that decreased methylation of FTO CpG11 sites correlates with increased FTO mRNA expression (Liu et al. 2016). Therefore, the data are convincing that the hypomethylation of specific CpG sites in the FTO gene promotes FTO expression (Melnik 2015) and that increased FTO expression is connected with the occurrence of some metabolic diseases and cancers. FTO expression is influenced by many factors. Previous research has demonstrated that milk functions as an epigenetic regulator by promoting the expression of FTO via transfer of exosomal miRNA-29, which downregulates the expression DNA methyltransferases (DNMT) and suppresses DNA CpG methylation (Melnik 2015). Moreover, FTO expression is regulated by essential amino acids (Gulati and Yeo 2013). In the presence of amino acids, FTO acts as a nutrient sensor that, via its demethylation activity, induces decreased activation of the mammalian target of the rapamycin complex 1 (mTORC1) signalling pathway, which is vital for the regulation of mRNA translation rates and cell growth (Yeo 2014; Gulati et al. 2013). Follow-up studies found that the role of FTO in nutrient regulation seems to be connected to its cellular localization, as it shuttles between the nucleus and cytoplasm, possibly via a mechanism mediated by XPO2 (Gulati et al. 2014). Recently, Aas et al. (2017) identified an N-terminal nuclear localization signal (NLS) and a C-terminal nuclear transport signal in FTO that participate in nucleocytoplasmic shuttling; however, this behaviour is not influenced by short-term amino acid starvation or by manipulation of autophagy. However, the mechanisms underlying FTO nucleocytoplasmic shuttling and its physiological functions remain to be further elucidated. Mechanistically, recent studies have found that FTO facilitates the activation of mTOR via increasing the mRNA level of TSC1, through which it is involved in insulin defects related to Alzheimer’s disease (Li et al. 2018).
Recently, it has been proposed that FTO may be associated with the regulation of telomere length (TL), through which it affects ageing and stress-related diseases including obesity, T2DM and certain types of cancer (Zhou et al. 2017). Shorter telomeres may be connected to increased BMI and adiposity (Tzanetakou et al. 2012). The 2-OG-dependent dioxygenase catalytic activity of FTO may regulate gene transcription or TL regulation by affecting nucleic acid demethylation (Zhou et al. 2017; Lister et al. 2009). However, further exploration is needed to understand the correlations between epigenetic modification by FTO and TL regulation.
The alternative splicing effects of FTO play a crucial role in mRNA processing events. The connection between FTO binding, activity and pre-mRNA processing mediated by m6A have been demonstrated (Bartosovic et al. 2017). Bartosovic et al. (2017) discovered that FTO preferentially binds to pre-mRNAs and mediates m6A removal in introns. Substantial changes in pre-mRNA splicing occurred after FTO knockout, exhibiting a clear prevalence of exon-skipping events that were negatively correlated with METTL3 knockdown, revealing the involvement of m6A. Moreover, extensive pre-mRNA back-splicing generates numerous circular RNAs (circRNAs) rich in consensus m6A motifs in the human transcriptome. This m6A-driven translation requires initiation factor eIF4G2 and the m6A reader YTHDF3 and is enhanced by the methyltransferase METTL3/14 and inhibited by the demethylase FTO (Yang et al. 2017). With respect to FTO degradation, related studies are lacking to a great extent. It has been previously described that ubiquitin proteasome-mediated degradation of FTO can be restrained by overexpression of protein kinase Cβ (PKCβ) (Tai et al. 2017). A recent study showed that FTO may undergo active ubiquitination on the evolutionarily conserved Lys-216 residue, routing FTO to proteasomal degradation (Zhu et al. 2018). These findings provide new insights into FTO protein turnover, but the precise mechanism requires further investigation.
Association between the FTO gene and RNA m6A demethylation
Chemical modification plays a critical role in DNA, RNA and histone metabolism by introducing or removing various groups, such as methyl, phosphate and acetyl groups. These modifications enrich the functions and genetic polymorphisms of DNA to a great extent. The N6-methyladenosine (m6A) modification is one of the most common modifications in messenger RNAs (mRNAs) and long non-coding RNAs (lncRNAs) in eukaryotes, accounting for approximately 80% of all methylations, and this process is regulated by methyltransferases (METTL3, METTL14 and WTAP), RNA-binding proteins (YTH Domain) and demethylases (FTO and ALKBH5) (Zhao et al. 2017; Wei et al. 1975; Desrosiers et al. 1974; Bokar et al. 1997; Liu et al. 2014; Ping et al. 2014; Wang et al. 2014; Liao et al. 2018). m6A is enriched in the 3′-untranslated regions (3′-UTRs) around the stop codon and the start codon (Schwartz et al. 2013; Meyer et al. 2012; Luo et al. 2014), and such modifications play critical roles in RNA splicing, degradation, translation and RNA–protein interactions (Wang et al. 2014, 2015; Yang et al. 2017a; Liu et al. 2015; Alarcon et al. 2015). Therefore, the m6A modification is engaged in widespread fundamental processes, including cell differentiation, tissue development, stem cell differentiation, circadian cycles, and fertility (Geula et al. 2015; Wang et al. 2014a; Zheng et al. 2013; Fustin et al. 2013).
Studies related to the m6A modification have stagnated for some time due to the lack of information on the relevant demethylases. However, the role of m6A modification in mRNA has recently gained renewed attention due to the breakthrough discoveries of two RNA demethylases, the fat mass- and obesity-associated protein (FTO) and the alkylation repair homologue protein 5 (ALKBH5) (Jia et al. 2011; Zheng et al. 2013). These discoveries indicate that the modification is reversible and dynamic, similar to the epigenetic modifications of DNA and histone proteins. He et al. (2011) found that siRNA-mediated knockdown of FTO enhanced m6A levels in mRNA and that upregulated expression of the FTO gene suppressed m6A methylation, revealing the demethylation activity of FTO. These researchers subsequently found that FTO partially co-localized with nuclear speckles, providing evidence of m6A in nuclear RNA as a physiological substrate of FTO (Jia et al. 2011). An earlier study found that the FTO protein targets certain m6A sites in mRNA, which may be an effect of the extra loop covering one side of the conserved jelly-roll motif, as revealed through its crystal structure (Han et al. 2010). Recently, Zou et al. (2016) indicated that m6A serves as a ‘conformational marker’ by inducing different conformational outcomes in RNAs that depend on sequence context, which then regulate the substrate specificity of the m6A demethylases. Nevertheless, these data fail to explain the differences in the substrate selectivity between FTO and ALKBH5; thus, more convincing evidence is needed.
While both FTO and ALKBH5 belong to the AlkB family of dioxygenases, FTO has a high affinity for 3-meT in ssDNA and 3-meU in ssRNA, while ALKBH5 only catalyses the N-demethylation of m6A ssRNA and ssDNA (Gerken et al. 2007; Xu et al. 2014; Jia et al. 2008). Their reaction pathways and functions appear to be different. FTO-mediated m6A demethylation generates two intermediates, N6-hydroxymethyladenosine (hm6A) and N6-formyladenosine (f6A). However, ALKBH5 converts m6A to hm6A without any intermediate (Chen et al. 2014; Fu et al. 2013). Furthermore, the FTO gene is widely expressed in many tissues including the hypothalamus, skeletal muscle and adipose tissue (Fan et al. 2009). The expression product of ALKBH5 mainly exists in the testis and lung, where it plays critical roles in RNA metabolism and impaired fertility (Zheng et al. 2013). It is important to note that the complexities of the demethylation-regulating activities of FTO and ALKBH5 remain to be further elucidated owing to the uncertain regulatory mechanisms of many physiological and pathological processes.
The role of FTO as a specific eraser of m6A tags on RNA has recently been challenged (Mauer and Jaffrey 2018; Mauer et al. 2017). In 2017, Mauer et al. (2017) proposed a new viewpoint on the actual substrate of FTO. They suggested that FTO preferentially demethylates m6Am rather than m6A and that m6Am tends to stabilize mRNA. Their findings showed that the demethylation activity of FTO was approximately 100 times higher towards m6Am than towards m6A. Moreover, the depletion of FTO showed no obvious effect on m6A levels, while the m6Am levels were more clearly enhanced in response to the changes in FTO levels. These results implied that the substrate of FTO was likely to be m6Am, which starkly differed from the results of a previous study by He et al., who discovered that m6A in mRNA serves as the substrate of FTO (Jia et al. 2011; Fu et al. 2013). Notably, the total amount of internal m6A is at least tenfold higher than that of m6Am in different cell lines; thus, the absolute level of m6A demethylation may be higher (Wei et al. 1975, 2018). The latest research conducted by He et al. revealed that the distribution of FTO varies between the nucleus and cytoplasm in different cell lines and that m6A seems to be the primary polyadenylated RNA substrate of FTO in the nucleus, while FTO preferentially targets m6Am in the cytoplasm (Wei et al. 2018).
The involvement of demethylation by FTO in many physiological and pathological processes, including RNA modification, transcriptome regulation and translation, is being gradually revealed. Wu et al. (2017) discovered that FTO-dependent demethylation, regulated by the AMPK pathway, suppresses mRNA m6A methylation and lipid accumulation in skeletal muscle cells. Recently, another study indicated that FTO reduced the mitochondrial content and thus increased the fat deposition in the liver by downregulating m6A levels (Kang et al. 2018). These results provide evidence for an association between FTO and fat metabolism. In addition, as an m6A demethylase, FTO is reported to play a key role in various cancers, such as leukaemia, breast cancer, glioblastoma, a topic that is now gaining attention (Li et al. 2017; Kaklamani et al. 2011; Cui et al. 2017) and is described below.
Briefly, m6A modification, which is regulated by various regulatory proteins, is universal and complicated in eukaryotes. FTO, which serves as a critical component of m6A modification, is related to the dysregulation of pathways that may underlie serious diseases, although the exact functions and concrete mechanisms remain to be explored.
Role of the FTO gene in various cancers
Although studies related to the role of the FTO gene in cancer are in the early stages, increasing evidence shows an association between FTO and the occurrence, development and prognosis of various cancers. Considering that cancer is always multifactorial, the role of FTO tends not to be the sole cause, and the recognition of the relevant pathogenesis is challenging. Through its function as an m6A demethylase, FTO seems to be attracting more attention in the cancer field because of the growing awareness of the role of the m6A modification in cancer. In this review, we describe the recent progress in understanding the function of FTO in cancer.
FTO SNPs related to different types of cancer
Several single-nucleotide polymorphisms (SNPs) in the FTO gene within a strong linkage disequilibrium block appear to be involved in many cancers. Considering the complexity of the samples and underlying causes, associations between FTO SNPs and the risk of some cancers are always controversial. The rs9939609, rs6499640, rs19079260, and rs8050136 variants in the FTO gene have been identified as being associated with endometrial cancer risk, and rs9939609 is also related to susceptibility to pancreatic cancer (Delahanty et al. 2011; Huang et al. 2017; Lurie et al. 2011). The results of two studies showed no significant association between FTO rs9939609 and the risk of prostate cancer (Salgado-Montilla et al. 2017; Lewis et al. 2010). Interestingly, it has been demonstrated that an inverse association between diabetes and prostate cancer exists (Jian Gang et al. 2015; Kasper and Giovannucci 2006) and that FTO was one of the susceptibility markers of this connection; however, the relevant mechanism remains unclear (Machiela et al. 2012). The role of FTO in breast cancer has been described by many studies. It is generally acknowledged that obesity increases the risk of breast cancer, which is associated with poorer prognosis, greater tumour size, faster metastasis, and higher malignancy (Trentham-Dietz et al. 1997; Renehan et al. 2008; Hernandez-Caballero and Sierra-Ramirez 2015; Calle and Kaaks 2004). Compared with adjacent breast tissues, FTO expression is highly expressed in breast cancer tissues, especially in those that are hormone receptor (HR)-negative and show HER2 amplification (Tan et al. 2015). Previously, it was shown that SNPs located in intron 1 of the FTO gene played a vital role in breast cancer, especially rs1477196, rs9939609, rs7206790 and rs8047395 (Kaklamani et al. 2011). Garcia-Closas et al. (2013) discovered that another SNP, rs11075995, showed activity in both normal and triple-negative breast cancer cells, which was correlated to patient BMI. However, the genetic polymorphisms in the FTO gene vary by ethnicity. The FTO SNPs rs1477196 and rs9939609 have no relationship with increased risk of breast cancer in Iranian women (Mojaver et al. 2015). Furthermore, whether the FTO-associated increase in the risk of breast cancer is mediated by obesity in different breast cancer subtypes is largely unknown.
In summary, certain FTO SNPs are associated with cancer susceptibility, possibly by regulating related gene expression. This association provides evidence for us to understand the roles of FTO in cancer, even if uncovering the relevant relationships is not easy. Although numerous loci associated with different cancers have been identified, further functional studies are needed to explore the concrete mechanisms.
FTO functions as a cancer oncogene
Studies on the role of FTO in cancer are rapidly accumulating. It was recently reported that FTO plays a critical oncogenic role in haematopoietic cell transformation and acute myeloid leukaemia (AML) as an m6A demethylase (Li et al. 2017). Li et al. (2017) found that the FTO gene, which is mediated by certain oncogenic proteins, is highly expressed in certain subtypes of AMLs, including t(11q23)/MLL-rearranged, t(15;17), FLT3-ITD, and/or NPM1-mutated AMLs. FTO improves the viability and proliferation of human AML cells and exerts anti-apoptosis effects (Li et al. 2017). Moreover, by targeting the critical downstream genes (ASB2 and RARA) and by reducing their m6A levels, mainly in the UTRs, FTO reinforces leukaemic oncogene-mediated cell transformation and leukaemogenesis and inhibits the progression of all-trans-retinoic acid (ATRA)-induced t(15;17) AML cells towards granulocytic and monocytic differentiation (Li et al. 2017). Therefore, using targeted FTO signalling inhibitors in combination with an all-trans-retinoic acid (ATRA) treatment may become an effective therapeutic strategy for leukaemia. Studies have emphasized the critical role of m6A modification in cancer and suggested that FTO acts as an oncoprotein. Nevertheless, a Cancer Genome Atlas Research Network (TCGA) AML study suggests that ALKBH5, also an m6A demethylase, shows frequent copy number loss that results in non-carcinogenic effects in AML, in contrast to FTO (Kwok et al. 2017). Approximately 10–20% of AML patients present recurrent somatic mutations in isocitrate dehydrogenase 1/2 (IDH1/2), leading to high levels of R-2-hydroxyglutarate (R-2HG) (Mardis et al. 2009). Recently, Su et al. (2018) discovered that R-2HG exerts effects in AML by suppressing FTO function, thereby promoting the accumulation of m6A on the MYC and CEBPA transcripts, decreasing their mRNA stability and expression, and eventually exhibiting antitumour activity. The discovery of this FTO/m6A/MYC/CEBPA signalling pathway offers insight into the concrete role of FTO in AML.
Cancer stem cells (CSCs) are widely considered to drive tumour progression through self-renewal and multipotent differentiation. A study performed by Cui et al. (2017) indicated a key role for m6A modification in the self-renewal and tumourigenesis of glioblastoma stem cells (GSCs), the tumour heterogeneity and treatment resistance of which tend to be the main causes of refractory glioblastoma (Huang et al. 2010; Satchi-Fainaro et al. 2012; Bao et al. 2006). FTO, as an m6A demethylase, exerts a positive effect on glioblastoma progression. The FTO inhibitor MA2 markedly suppresses GSC growth and self-renewal in vitro and lengthens the survival time of GSC-grafted mice, while the knockdown of METTL3 or METTL14, the key elements of the m6A methyltransferase complex, promotes GSC growth and self-renewal (Cui et al. 2017). Meanwhile, Zhang et al. (2017) discovered that ALKBH5 also maintained GSC proliferation and tumourigenesis by maintaining the methylation and expression of FOXM1. These findings emphasize a critical oncogenic role for FTO in CSCs by regulating the occurrence and development of the related tumour. However, to date, research into the effect FTO on CSCs is scarce.
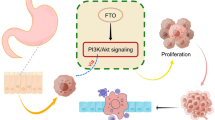
Effect of FTO on the growth, proliferation and metastasis of cancer
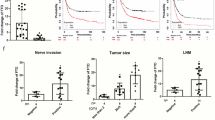
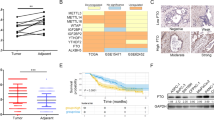
As mentioned above, FTO enhances the growth and proliferation of human AML cells and clearly promotes leukaemic cell transformation and immortalization (Li et al. 2017). Moreover, another AML study revealed that FTO activity is inhibited by R-2HG, which decreases the stability of MYC/CEBPA transcripts by increasing m6A mRNA levels, thereby suppressing leukaemia-cell proliferation and promoting cell-cycle arrest (Su et al. 2018). In addition, FTO functions in cancers such as endometrial cancer and gastric cancer. FTO is connected to the PI3K/AKT and MAPK signalling pathways in oestrogen-driven endometrial cancer, where it induces cancer cell growth and invasion (Zhang et al. 2012). In addition, it has been demonstrated that oestrogen promotes FTO nuclear localization and increases endometrial cancer cell proliferation via the mTOR signalling pathway, which is involved in energy metabolism; however, how ERα mediates FTO nuclear accumulation is unknown (Zhu et al. 2016). A recent study on gastric cancer revealed that FTO was also highly expressed in tumour regions, which promotes the occurrence of gastric cancer by enhancing the proliferation and migration of gastric cancer cell lines and lymph node metastasis (Xu et al. 2017). However, studies related to the molecular mechanism of this aspect of gastric cancer are lacking. Due to the significant role of FTO in the growth, proliferation and metastasis of cancer cells, research into the molecular and cellular mechanisms are urgently needed for improving cancer treatments and prognoses.
Impact of FTO on chemo-radiotherapy resistance and prognosis
FTO is also involved in chemo-radiotherapy resistance and may give rise to poor cancer prognoses. Cancer stem cells (CSCs) are often a crucial factor regulating treatment resistance. Cui et al. (2017) discovered that m6A RNA modification exerted a critical effect on the self-renewal and tumourigenesis of glioblastoma stem cells (GCSs). The FTO inhibitor MA2 extended the lifespan of GSC-engrafted mice, implying that m6A modification may be a target for suppressing tumour progression and reversing resistance to radiotherapy and chemotherapy in glioblastoma. In AML, all-trans-retinoic acid (ATRA) treatment is an important treatment regimen (Hu et al. 2009; Wang and Chen 2008). Li et al. (2017) discovered that FTO suppressed all-trans-retinoic acid (ATRA)-triggered AML cell differentiation by decreasing m6A levels in some mRNA transcripts, including ASB2 and RARA, indicating that FTO plays a positive role in the poor prognosis conferred by AML. Recently, it was illustrated that FTO participates in regulating chemo-radiotherapy resistance in cervical squamous cell carcinoma (CSCC) (Zhou et al. 2018). As an m6A demethylase, FTO confers chemo-radiotherapy resistance in CSCC by targeting β-catenin, an EMT marker that indicates resistance to anticancer treatment and is upregulated by excision repair cross-complementation group 1 (ERCC1), a downstream modulator of FTO (Zhou et al. 2018). In addition, high expression of FTO is reported to be closely associated with the poor prognosis conferred by gastric cancer by promoting gastric cancer cell migration and invasion and lymph node metastasis (Xu et al. 2017). In fact, studies on how FTO regulates chemo-radiotherapy resistance are still in the initial stages. Further functional studies are needed to verify the roles of FTO in different cancers, especially for a deep understanding of the related mechanisms. Accurate knowledge of the function of FTO and m6A modifications in cancer is of great significance for making accurate diagnoses, understanding therapy sensitivity and improving the prognoses of the related diseases.
In summary, these findings indicate that the relevant modulators involved in m6A modification have notable impacts on various cancers. As seen below, it can be concluded that FTO plays critical roles in tumour germination, development and chemo-radiotherapy resistance, exerting its effects to a large extent via its m6A demethylase activity (Fig. 1). Furthermore, FTO serves as an oncogene in AML and glioblastoma. As FTO was only recently identified as a demethylase, much remains to be explored with regard to its functions and mechanisms in different cancers.
Conclusions
The FTO gene was initially identified as a susceptibility gene strongly linked to obesity. It has been demonstrated that FTO plays critical roles in fat metabolism, energy regulation and other physiological activities and that it exerts positive effects on the occurrence and development of lipid metabolism-related diseases, including obesity and T2DM. As studies progressed, it was discovered that FTO is an m6A demethylase that regulates the m6A levels in mRNA, which are involved in many fundamental physiological processes. More importantly, FTO plays key roles in the occurrence, progression and treatment of various cancers, such as leukaemia, breast cancer, and glioblastoma. FTO exhibits both cancer-inhibiting and -promoting activities. Nevertheless, comprehensive and systematic studies on the exact roles and molecular mechanisms underlying the roles of FTO in cancer are incomplete. Therefore, finding more convincing evidence to define the detailed roles of FTO is desperately needed to analyse and understand the similarities and differences between various cancers. Considering FTO’s critical role in many diseases, FTO may become a new promising target for the diagnosis and treatment of various diseases in the near future, especially for specific types of cancers, such as AML, glioblastoma and breast cancer.
Abbreviations
- FTO:
-
Fat mass and obesity-associated
- BMI:
-
Body mass index
- T2DM:
-
Type 2 diabetes mellitus
- SNPs:
-
Single-nucleotide polymorphisms
- m6A:
-
N6-methyladenosine
- GWAS:
-
Genome-wide association studies
- XPO2:
-
Exportin 2
- UTRs:
-
Untranslated regions
- 2-OG:
-
2-oxoglutarate
- 3-meT:
-
3-methylthymidine
- AML:
-
Acute myeloid leukaemia
- MLL:
-
Mixed lineage leukaemia
- 3-meU:
-
3-methyluracil
- CSC:
-
Cancer stem cell
- DNMT:
-
DNA methyltransferases
- IRX3:
-
Iroquois-related homeobox 3
- circRNAs:
-
Circular RNAs
- mRNAs:
-
Messenger RNAs
- lncRNAs:
-
Long non-coding RNAs
- METTL14:
-
Methyltransferase-like 14
- METTL3:
-
Methyltransferase-like 3
- WTAP:
-
Wilms’ tumour 1-associating protein
- ALKBH5:
-
AlkB homologue 5
- ASB2:
-
Ankyrin-repeat SOCS box-containing protein 2
- RARA:
-
Retinoic acid receptor alpha
- ATRA:
-
All-trans-retinoic acid
- R-2HG:
-
R-2-hydroxyglutarate
- IDH1/2:
-
Isocitrate dehydrogenase ½
- CEBPA:
-
CCAAT enhancer-binding protein alpha
- GSCs:
-
Glioblastoma stem cells
- FOXM1:
-
Forkhead box transcription factor M1
- ERCC1:
-
Excision repair cross-complementation group 1
References
Aas A, Isakson P, Bindesboll C, Alemu EA, Klungland A, Simonsen A (2017) Nucleocytoplasmic shuttling of FTO does not affect starvation-induced autophagy. PLoS One 12(3):e0168182
Akbari ME, Gholamalizadeh M, Doaei S, Mirsafa F (2018) FTO gene affects obesity and breast cancer through similar mechanisms: a new insight into the molecular therapeutic targets. Nutr Cancer 70(1):30–36
Alarcon CR, Lee H, Goodarzi H, Halberg N, Tavazoie SF (2015) N6-methyladenosine marks primary microRNAs for processing. Nature 519(7544):482–485
Andreasen CH, Stender-Petersen KL, Mogensen MS, Torekov SS, Wegner L, Andersen G, Nielsen AL, Albrechtsen A, Borch-Johnsen K, Rasmussen SS et al (2008) Low physical activity accentuates the effect of the FTO rs9939609 polymorphism on body fat accumulation. Diabetes 57(1):95–101
Anselme I, Laclef C, Lanaud M, Ruther U, Schneider-Maunoury S (2007) Defects in brain patterning and head morphogenesis in the mouse mutant Fused toes. Dev Biol 304(1):208–220
Arrizabalaga M, Larrarte E, Margareto J, Maldonado-Martin S, Barrenechea L, Labayen I (2014) Preliminary findings on the influence of FTO rs9939609 and MC4R rs17782313 polymorphisms on resting energy expenditure, leptin and thyrotropin levels in obese non-morbid premenopausal women. J Physiol Biochem 70(1):255–262
Bao S, Wu Q, McLendon RE, Hao Y, Shi Q, Hjelmeland AB, Dewhirst MW, Bigner DD, Rich JN (2006) Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 444(7120):756–760
Bartosovic M, Molares HC, Gregorova P, Hrossova D, Kudla G, Vanacova S (2017) N6-methyladenosine demethylase FTO targets pre-mRNAs and regulates alternative splicing and 3′-end processing. Nucleic Acids Res 45(19):11356–11370
Benedict C, Axelsson T, Soderberg S, Larsson A, Ingelsson E, Lind L, Schioth HB (2014) Fat mass and obesity-associated gene (FTO) is linked to higher plasma levels of the hunger hormone Ghrelin and lower serum levels of the satiety hormone leptin in older adults. Diabetes 63(11):3955–3959
Berulava T, Horsthemke B (2010) The obesity-associated SNPs in intron 1 of the FTO gene affect primary transcript levels. Eur J Hum Genet EJHG 18(9):1054–1056
Bokar JA, Shambaugh ME, Polayes D, Matera AG, Rottman FM (1997) Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3(11):1233–1247
Calle EE, Kaaks R (2004) Overweight, obesity and cancer: epidemiological evidence and proposed mechanisms. Nat Rev Cancer 4(8):579–591
Chang YC, Liu PH, Lee WJ, Chang TJ, Jiang YD, Li HY, Kuo SS, Lee KC, Chuang LM (2008) Common variation in the fat mass and obesity-associated (FTO) gene confers risk of obesity and modulates BMI in the Chinese population. Diabetes 57(8):2245–2252
Chen W, Zhang L, Zheng G, Fu Y, Ji Q, Liu F, Chen H, He C (2014) Crystal structure of the RNA demethylase ALKBH5 from zebrafish. FEBS Lett 588(6):892–898
Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, Sun G, Lu Z, Huang Y, Yang CG et al (2017) m(6)A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep 18(11):2622–2634
Delahanty RJ, Beeghly-Fadiel A, Xiang YB, Long JR, Cai QY, Wen WQ, Xu WH, Cai H, He J, Gao YT et al (2011) Association of obesity-related genetic variants with endometrial cancer risk: a report from the Shanghai endometrial cancer genetics study. Am J Epidemiol 174(10):1115–1126
Deng T, Lyon CJ, Bergin S, Caligiuri MA, Hsueh WA (2016) Obesity, inflammation, and cancer. Annu Rev Pathol 11:421–449
Desrosiers R, Friderici K, Rottman F (1974) Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci USA 71(10):3971–3975
Dina C, Meyre D, Gallina S, Durand E, Korner A, Jacobson P, Carlsson LM, Kiess W, Vatin V, Lecoeur C et al (2007) Variation in FTO contributes to childhood obesity and severe adult obesity. Nat Genet 39(6):724–726
Do R, Bailey SD, Desbiens K, Belisle A, Montpetit A, Bouchard C, Perusse L, Vohl MC, Engert JC (2008) Genetic variants of FTO influence adiposity, insulin sensitivity, leptin levels, and resting metabolic rate in the Quebec Family Study. Diabetes 57(4):1147–1150
Fan B, Du ZQ, Rothschild MF (2009) The fat mass and obesity-associated (FTO) gene is associated with intramuscular fat content and growth rate in the pig. Anim Biotechnol 20(2):58–70
Fang H, Li Y, Du S, Hu X, Zhang Q, Liu A, Ma G (2010) Variant rs9939609 in the FTO gene is associated with body mass index among Chinese children. BMC Med Genet 11:136
Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM, Perry JR, Elliott KS, Lango H, Rayner NW et al (2007) A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316(5826):889–894
Fu Y, Jia G, Pang X, Wang RN, Wang X, Li CJ, Smemo S, Dai Q, Bailey KA, Nobrega MA et al (2013) FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nat Commun 4:1798
Fustin JM, Doi M, Yamaguchi Y, Hida H, Nishimura S, Yoshida M, Isagawa T, Morioka MS, Kakeya H, Manabe I et al (2013) RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155(4):793–806
Garcia-Closas M, Couch FJ, Lindstrom S, Michailidou K, Schmidt MK, Brook MN, Orr N, Rhie SK, Riboli E, Feigelson HS et al (2013) Genome-wide association studies identify four ER negative-specific breast cancer risk loci. Nat Genet 45(4):392–398 (398e391–392)
Gerken T, Girard CA, Tung YC, Webby CJ, Saudek V, Hewitson KS, Yeo GS, McDonough MA, Cunliffe S, McNeill LA et al (2007) The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318(5855):1469–1472
Geula S, Moshitch-Moshkovitz S, Dominissini D, Mansour AA, Kol N, Salmon-Divon M, Hershkovitz V, Peer E, Mor N, Manor YS et al (2015) m(6)A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science 347(6225):1002–1006
Gonzalez-Sanchez JL, Zabena C, Martinez-Larrad MT, Martinez-Calatrava MJ, Perez-Barba M, Serrano-Rios M (2009) Variant rs9939609 in the FTO gene is associated with obesity in an adult population from Spain. Clin Endocrinol 70(3):390–393
Grant SF, Li M, Bradfield JP, Kim CE, Annaiah K, Santa E, Glessner JT, Casalunovo T, Frackelton EC, Otieno FG et al (2008) Association analysis of the FTO gene with obesity in children of Caucasian and African ancestry reveals a common tagging SNP. PLoS One 3(3):e1746
Gulati P, Yeo GS (2013) The biology of FTO: from nucleic acid demethylase to amino acid sensor. Diabetologia 56(10):2113–2121
Gulati P, Cheung MK, Antrobus R, Church CD, Harding HP, Tung YCL, Rimmington D, Ma M, Ron D, Lehner PJ et al (2013) Role for the obesity-related FTO gene in the cellular sensing of amino acids. Proc Natl Acad Sci USA 110(7):2557–2562
Gulati P, Avezov E, Ma M, Antrobus R, Lehner P, O’Rahilly S, Yeo GS (2014) Fat mass and obesity-related (FTO) shuttles between the nucleus and cytoplasm. Biosci Rep 34(5):621–628
Han Z, Niu T, Chang J, Lei X, Zhao M, Wang Q, Cheng W, Wang J, Feng Y, Chai J (2010) Crystal structure of the FTO protein reveals basis for its substrate specificity. Nature 464(7292):1205–1209
Hernandez-Caballero ME, Sierra-Ramirez JA (2015) Single nucleotide polymorphisms of the FTO gene and cancer risk: an overview. Mol Biol Rep 42(3):699–704
Hotta K, Nakata Y, Matsuo T, Kamohara S, Kotani K, Komatsu R, Itoh N, Mineo I, Wada J, Masuzaki H et al (2008) Variations in the FTO gene are associated with severe obesity in the Japanese. J Hum Genet 53(6):546–553
Hu J, Liu YF, Wu CF, Xu F, Shen ZX, Zhu YM, Li JM, Tang W, Zhao WL, Wu W et al (2009) Long-term efficacy and safety of all-trans retinoic acid/arsenic trioxide-based therapy in newly diagnosed acute promyelocytic leukemia. Proc Natl Acad Sci USA 106(9):3342–3347
Huang Z, Cheng L, Guryanova OA, Wu QL, Bao SD (2010) Cancer stem cells in glioblastoma-molecular signaling and therapeutic targeting. Protein Cell 1(7):638–655
Huang XY, Zhao J, Yang MY, Li M, Zheng JM (2017) Association between FTO gene polymorphism (rs9939609 T/A) and cancer risk: a meta-analysis. Eur J Cancer Care (Engl) 26(5):e12464
Jabbari K, Bernardi G (2004) Cytosine methylation and CpG, TpG (CpA) and TpA frequencies. Gene 333:143–149
Jia G, Yang CG, Yang S, Jian X, Yi C, Zhou Z, He C (2008a) Oxidative demethylation of 3-methylthymine and 3-methyluracil in single-stranded DNA and RNA by mouse and human FTO. FEBS Lett 582(23–24):3313–3319
Jia GF, Yang CG, Yang SD, Jian X, Yi CQ, Zhou ZQ, He C (2008b) Oxidative demethylation of 3-methylthymine and 3-methyluracil in single-stranded DNA and RNA by mouse and human FTO. FEBS Lett 582(23–24):3313–3319
Jia GF, Fu Y, Zhao X, Dai Q, Zheng GQ, Yang Y, Yi CQ, Lindahl T, Pan T, Yang YG et al (2011) N6-Methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 7(12):885–887
Jian Gang P, Mo L, Lu Y, Runqi L, Xing Z (2015) Diabetes mellitus and the risk of prostate cancer: an update and cumulative meta-analysis. Endocr Res 40(1):54–61
Kaklamani V, Yi N, Sadim M, Siziopikou K, Zhang K, Xu Y, Tofilon S, Agarwal S, Pasche B, Mantzoros C (2011) The role of the fat mass and obesity associated gene (FTO) in breast cancer risk. BMC Med Genet 12:52
Kang Y, Liu F, Liu Y (2017) Is FTO gene variant related to cancer risk independently of adiposity? An updated meta-analysis of 129,467 cases and 290,633 controls. Oncotarget 8(31):50987–50996
Kang H, Zhang Z, Yu L, Li Y, Liang M, Zhou L (2018) FTO reduces mitochondria and promotes hepatic fat accumulation through RNA demethylation. J Cell Biochem 119(7):5676–5685
Kasper JS, Giovannucci E (2006) A meta-analysis of diabetes mellitus and the risk of prostate cancer. Cancer Epidemiol Biomark Prev Publ Am Assoc Cancer Res 15(11):2056–2062 (Cosponsored by the American Society of Preventive Oncology)
Kwok CT, Marshall AD, Rasko JE, Wong JJ (2017) Genetic alterations of m(6)A regulators predict poorer survival in acute myeloid leukemia. J Hematol Oncol 10(1):39
Lewis SJ, Murad A, Chen LN, Smith GD, Donovan J, Palmer T, Hamdy F, Neal D, Lane JA, Davis M et al (2010) Associations between an obesity related genetic variant (FTO rs9939609) and prostate cancer risk. PLoS One 5(10):e13485
Li H, Wu Y, Loos RJ, Hu FB, Liu Y, Wang J, Yu Z, Lin X (2008) Variants in the fat mass- and obesity-associated (FTO) gene are not associated with obesity in a Chinese Han population. Diabetes 57(1):264–268
Li Z, Weng H, Su R, Weng X, Zuo Z, Li C, Huang H, Nachtergaele S, Dong L, Hu C et al (2017) FTO plays an oncogenic role in acute myeloid leukemia as a N(6)-methyladenosine RNA demethylase. Cancer Cell 31(1):127–141
Li H, Ren Y, Mao K, Hua F, Yang Y, Wei N, Yue C, Li D, Zhang H (2018) FTO is involved in Alzheimer’s disease by targeting TSC1-mTOR-Tau signaling. Biochem Biophys Res Commun 498(1):234–239
Liao SH, Sun HB, Xu C (2018) YTH domain: a family of N-6-methyladenosine (m(6)A) readers. Genom Proteom Bioinf 16(2):99–107
Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo QM et al (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462(7271):315–322
Liu J, Yue Y, Han D, Wang X, Fu Y, Zhang L, Jia G, Yu M, Lu Z, Deng X et al (2014) A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem Biol 10(2):93–95
Liu N, Dai Q, Zheng G, He C, Parisien M, Pan T (2015) N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518(7540):560–564
Liu ZW, Zhang JT, Cai QY, Zhang HX, Wang YH, Yan HT, Wu HM, Yang XJ (2016) Birth weight is associated with placental fat mass- and obesity-associated gene expression and promoter methylation in a Chinese population. J Matern Fetal Neonatal Med 29(1):106–111
Luo GZ, MacQueen A, Zheng G, Duan H, Dore LC, Lu Z, Liu J, Chen K, Jia G, Bergelson J et al (2014) Unique features of the m6A methylome in Arabidopsis thaliana. Nat Commun 5:5630
Lurie G, Gaudet MM, Spurdle AB, Carney ME, Wilkens LR, Yang HP, Weiss NS, Webb PM, Thompson PJ, Terada K et al (2011) The obesity-associated polymorphisms FTO rs9939609 and MC4R rs17782313 and endometrial cancer risk in non-Hispanic white women. PLoS One 6(2):e16756
Machiela MJ, Lindstrom S, Allen NE, Haiman CA, Albanes D, Barricarte A, Berndt SI, Bueno-de-Mesquita HB, Chanock S, Gaziano JM et al (2012) Association of type 2 diabetes susceptibility variants with advanced prostate cancer risk in the Breast and Prostate Cancer Cohort Consortium. Am J Epidemiol 176(12):1121–1129
Mardis ER, Ding L, Dooling DJ, Larson DE, McLellan MD, Chen K, Koboldt DC, Fulton RS, Delehaunty KD, McGrath SD et al (2009) Recurring mutations found by sequencing an acute myeloid leukemia genome. New Engl J Med 361(11):1058–1066
Mauer J, Jaffrey SR (2018) FTO, m(6) Am, and the hypothesis of reversible epitranscriptomic mRNA modifications. FEBS Lett 592(12):2012–2022
Mauer J, Luo X, Blanjoie A, Jiao X, Grozhik AV, Patil DP, Linder B, Pickering BF, Vasseur JJ, Chen Q et al (2017) Reversible methylation of m(6)Am in the 5′ cap controls mRNA stability. Nature 541(7637):371–375
McTaggart JS, Lee S, Iberl M, Church C, Cox RD, Ashcroft FM (2011) FTO is expressed in neurones throughout the brain and its expression is unaltered by fasting. PLoS One 6(11):e27968
Melnik BC (2015) Milk: an epigenetic amplifier of FTO-mediated transcription? Implications for Western diseases. J Transl Med 13:385
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149(7):1635–1646
Mojaver M, Mokarian F, Kazemi M, Salehi M (2015) Specific TaqMan allelic discrimination assay for rs1477196 and rs9939609 single nucleotide polymorphisms of FTO gene demonstrated that there is no association between these SNPs and risk of breast cancer in Iranian women. Adv Biomed Res 4:136
Ningombam SS, Chhungi V, Newmei MK, Rajkumari S, Devi NK, Mondal PR, Saraswathy KN (2018) Differential distribution and association of FTO rs9939609 gene polymorphism with obesity: a cross-sectional study among two tribal populations of India with East-Asian ancestry. Gene 647:198–204
Ping XL, Sun BF, Wang L, Xiao W, Yang X, Wang WJ, Adhikari S, Shi Y, Lv Y, Chen YS et al (2014) Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res 24(2):177–189
Pischon T, Nimptsch K (2016) Obesity and risk of cancer: an introductory overview. Recent Results Cancer Res Fortschritte der Krebsforschung Progres dans les recherches sur le cancer 208:1–15
Pischon T, Nothlings U, Boeing H (2008) Obesity and cancer. Proc Nutr Soc 67(2):128–145
Qi L, Kang K, Zhang CL, van Dam RM, Kraft P, Hunter D, Lee CH, Hu FB (2008) Fat mass- and obesity-associated (FTO) gene variant is associated with obesity longitudinal analyses in two cohort studies and functional test. Diabetes 57(11):3145–3151
Renehan AG, Tyson M, Egger M, Heller RF, Zwahlen M (2008) Body-mass index and incidence of cancer: a systematic review and meta-analysis of prospective observational studies. Lancet 371(9612):569–578
Robbens S, Rouze P, Cock JM, Spring J, Worden AZ, Van de Peer Y (2008) The FTO gene, implicated in human obesity, is found only in vertebrates and marine algae. J Mol Evol 66(1):80–84
Salgado-Montilla JL, Rodriguez-Caban JL, Sanchez-Garcia J, Sanchez-Ortiz R, Irizarry-Ramirez M (2017) Impact of FTO SNPs rs9930506 and rs9939609 in prostate cancer severity in a cohort of Puerto Rican Men. Arch Cancer Res 5(3):148
Sanchez-Pulido L, Andrade-Navarro MA (2007) The FTO (fat mass and obesity associated) gene codes for a novel member of the non-heme dioxygenase superfamily. BMC Biochem 8:23
Satchi-Fainaro R, Ferber S, Segal E, Ma L, Dixit N, Ijaz A, Hlatky L, Abdollahi A, Almog N (2012) Prospective identification of glioblastoma cells generating dormant tumors. PLoS One 7(9):e44395
Saxonov S, Berg P, Brutlag DL (2006) A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc Natl Acad Sci USA 103(5):1412–1417
Schwartz S, Agarwala SD, Mumbach MR, Jovanovic M, Mertins P, Shishkin A, Tabach Y, Mikkelsen TS, Satija R, Ruvkun G et al (2013) High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155(6):1409–1421
Scuteri A, Sanna S, Chen WM, Uda M, Albai G, Strait J, Najjar S, Nagaraja R, Orru M, Usala G et al (2007) Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genet 3(7):e115
Su R, Dong L, Li C, Nachtergaele S, Wunderlich M, Qing Y, Deng X, Wang Y, Weng X, Hu C et al (2018) R-2HG exhibits anti-tumor activity by targeting FTO/m(6)A/MYC/CEBPA signaling. Cell 172(1–2):90–105 (e123)
Tai H, Wang X, Zhou J, Han X, Fang T, Gong H, Huang N, Chen H, Qin J, Yang M et al (2017) Protein kinase Cbeta activates fat mass and obesity-associated protein by influencing its ubiquitin/proteasome degradation. FASEB J 31(10):4396–4406
Tan A, Dang Y, Chen G, Mo Z (2015) Overexpression of the fat mass and obesity associated gene (FTO) in breast cancer and its clinical implications. Int J Clin Exp Pathol 8(10):13405–13410
Toperoff G, Aran D, Kark JD, Rosenberg M, Dubnikov T, Nissan B, Wainstein J, Friedlander Y, Levy-Lahad E, Glaser B et al (2012) Genome-wide survey reveals predisposing diabetes type 2-related DNA methylation variations in human peripheral blood. Hum Mol Genet 21(2):371–383
Toperoff G, Kark JD, Aran D, Nassar H, Ahmad WA, Sinnreich R, Azaiza D, Glaser B, Hellman A (2015) Premature aging of leukocyte DNA methylation is associated with type 2 diabetes prevalence. Clin Epigenet 7:35
Trentham-Dietz A, Newcomb PA, Storer BE, Longnecker MP, Baron J, Greenberg ER, Willett WC (1997) Body size and risk of breast cancer. Am J Epidemiol 145(11):1011–1019
Tzanetakou IP, Katsilambros NL, Benetos A, Mikhailidis DP, Perrea DN (2012) “Is obesity linked to aging?”: adipose tissue and the role of telomeres. Ageing Res Rev 11(2):220–229
van der Hoeven F, Schimmang T, Volkmann A, Mattei MG, Kyewski B, Ruther U (1994) Programmed cell death is affected in the novel mouse mutant Fused toes (Ft). Development 120(9):2601–2607
Wang ZY, Chen Z (2008) Acute promyelocytic leukemia: from highly fatal to highly curable. Blood 111(5):2505–2515
Wang Y, Li Y, Toth JI, Petroski MD, Zhang Z, Zhao JC (2014a) N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat Cell Biol 16(2):191–198
Wang X, Lu ZK, Gomez A, Hon GC, Yue YN, Han DL, Fu Y, Parisien M, Dai Q, Jia GF et al (2014b) N-6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505(7481):117–117+
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, Fu Y, Parisien M, Dai Q, Jia G et al (2014c) N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505(7481):117–120
Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, Weng X, Chen K, Shi H, He C (2015) N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell 161(6):1388–1399
Wei CM, Gershowitz A, Moss B (1975) Methylated nucleotides block 5′ terminus of HeLa cell messenger RNA. Cell 4(4):379–386
Wei J, Liu F, Lu Z, Fei Q, Ai Y, He PC, Shi H, Cui X, Su R, Klungland A et al (2018) Differential m(6)A, m(6)Am, and m(1)A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol Cell 71(6):973–985 (e975)
Wu WC, Feng JE, Jiang DH, Zhou XH, Jiang Q, Cai M, Wang XX, Shan TZ, Wang YZ (2017) AMPK regulates lipid accumulation in skeletal muscle cells through FTO-dependent demethylation of N-6-methyladenosine. Sci Rep Uk 7:41606
Xu C, Liu K, Tempel W, Demetriades M, Aik W, Schofield CJ, Min J (2014) Structures of human ALKBH5 demethylase reveal a unique binding mode for specific single-stranded N6-methyladenosine RNA demethylation. J Biol Chem 289(25):17299–17311
Xu D, Shao WW, Jiang YS, Wang X, Liu Y, Liu XC (2017) FTO expression is associated with the occurrence of gastric cancer and prognosis. Oncol Rep 38(4):2285–2292
Yang Y, Fan XJ, Mao MW, Song XW, Wu P, Zhang Y, Jin YF, Yang Y, Chen LL, Wang Y et al (2017a) Extensive translation of circular RNAs driven by N-6-methyladenosine. Cell Res 27(5):626–641
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, Jin Y, Yang Y, Chen LL, Wang Y et al (2017b) Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res 27(5):626–641
Yeo GS (2014) The role of the FTO (Fat Mass and Obesity Related) locus in regulating body size and composition. Mol Cell Endocrinol 397(1–2):34–41
Zhang G, Karns R, Narancic NS, Sun G, Cheng H, Missoni S, Durakovic Z, Rudan P, Chakraborty R, Deka R (2010) Common SNPs in FTO gene are associated with obesity related anthropometric traits in an island population from the eastern Adriatic coast of Croatia. PLoS One 5(4):e10375
Zhang Z, Zhou D, Lai Y, Liu Y, Tao X, Wang Q, Zhao G, Gu H, Liao H, Zhu Y et al (2012) Estrogen induces endometrial cancer cell proliferation and invasion by regulating the fat mass and obesity-associated gene via PI3K/AKT and MAPK signaling pathways. Cancer Lett 319(1):89–97
Zhang S, Zhao BS, Zhou A, Lin K, Zheng S, Lu Z, Chen Y, Sulman EP, Xie K, Bogler O et al (2017) m(6)A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell 31(4):591–606 (e596)
Zhao BS, Roundtree IA, He C (2017) Post-transcriptional gene regulation by mRNA modifications. Nat Rev Mol Cell Biol 18(1):31–42
Zheng GQ, Dahl JA, Niu YM, Fedorcsak P, Huang CM, Li CJ, Vagbo CB, Shi Y, Wang WL, Song SH et al (2013) ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell 49(1):18–29
Zhou Y, Hambly BD, McLachlan CS (2017) FTO associations with obesity and telomere length. J Biomed Sci 24(1):65
Zhou S, Bai ZL, Xia D, Zhao ZJ, Zhao R, Wang YY, Zhe H (2018) FTO regulates the chemo-radiotherapy resistance of cervical squamous cell carcinoma (CSCC) by targeting-catenin through mRNA demethylation. Mol Carcinogen 57(5):590–597
Zhu Y, Shen J, Gao L, Feng Y (2016) Estrogen promotes fat mass and obesity-associated protein nuclear localization and enhances endometrial cancer cell proliferation via the mTOR signaling pathway. Oncol Rep 35(4):2391–2397
Zhu T, Yong XLH, Xia D, Widagdo J, Anggono V (2018) Ubiquitination regulates the proteasomal degradation and nuclear translocation of the fat mass and obesity-associated (FTO) protein. J Mol Biol 430(3):363–371
Zou S, Toh JDW, Wong KHQ, Gao YG, Hong WJ, Woon ECY (2016) N-6-Methyladenosine: a conformational marker that regulates the substrate specificity of human demethylases FTO and ALKBH5. Sci Rep Uk 6:25677
Acknowledgements
This study was supported by the National Natural Science Foundation of China (China; 31201028, 81872893), the Fundamental Research Fund for the Central Universities (China; 21617462), the Guangzhou Science Technology and Innovation Commission (China; 201707010099), the Medical Scientific Research Foundation of Guangdong Province (China; A2017574) and the Provincial Undergraduates’ Innovation and Entrepreneurship Training Programs (China; 82618257).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Chen, J., Du, B. Novel positioning from obesity to cancer: FTO, an m6A RNA demethylase, regulates tumour progression. J Cancer Res Clin Oncol 145, 19–29 (2019). https://doi.org/10.1007/s00432-018-2796-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-018-2796-0