Abstract
Purpose
Inter-individual variations in treatment efficacy may be influenced by polymorphisms in DNA repair genes. We investigated the association of 3 functional polymorphisms in the nucleotide excision repair (NER) pathway with survival outcome of 95 patients with metastatic breast cancer (MBC) treated with DNA-damaging chemotherapy.
Methods
ERCC1 8092 C/A, ERCC2 Asp312Asn and ERCC2 Lys751Gln were determined using Taqman-based genotyping assays. Genotype associations with breast cancer-specific survival (BCSS) and progression-free survival (PFS) were evaluated using Kaplan–Meier estimates and hazard ratios calculated using Cox regression analysis. Tests for trend were conducted by calculating P-values for the HR coefficient in proportional hazards regression models.
Results
ERCC2 Lys751Gln was significantly associated with BCSS (median: 24.8 months for AA/AC combined and 14.2 months for CC, HR: 1.9 (95% CI 1.06–3.26)). Median BCSS decreased with increasing number of designated adverse genotypes for the 3 polymorphisms (Ptrend = 0.003). Risk estimates for PFS were nonsignificantly elevated and were significantly elevated for BCSS for patients with 2 (HR = 2.21, 95% CI: 1.04–4.72) or 3 (HR = 6.67, 95% CI: 2.19–20.29) adverse genotypes. In treatment subgroup analysis, risk estimates for BCSS were significantly elevated for patients with 3 adverse genotypes treated with cyclophosphamide, mitoxantrone and vinblastine (HR: 11.9, 95% CI 1.77–79.51) and Ptrend = 0.02 for increasing number of adverse genotypes. Risk of progression was significantly increased for patients with 1 adverse genotype treated with cyclophosphamide, mitoxantrone and carboplatin (HR: 3.5, 95% CI 1.19–10.6) and Ptrend = 0.02 for increasing number of adverse genotypes.
Conclusion
Polymorphisms in NER pathway may impact survival outcome for patients with MBC following treatment with DNA-damaging chemotherapy. These results provide support for a polygenic pathway approach for assessing the prognostic and predictive potential of polymorphisms in treatment outcome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Metastatic breast cancer (MBC) is diagnosed in approximately 5% of new patients, and 30% of previously treated patients will develop recurrent metastatic disease (Gonzalez-Angulo et al. 2007). Presently, few effective treatments for MBC are available, and most patients show either poor or limited responses to treatment. As a result, MBC is considered incurable, and the primary treatment objectives are disease control and palliation. The 5-year survival rate for patients with MBC is approximately 20%, and the median survival duration varies from 12 to 24 months (Gonzalez-Angulo et al. 2007).
There is considerable inter-individual variation in response and survival following chemotherapy and radiation treatment of cancer. DNA damage from these treatments can initiate a number of cellular pathways involved in DNA repair, cell cycle control, metabolism and apoptosis. Single nucleotide polymorphisms (SNPs) in genes in these pathways may significantly affect DNA repair efficiency (Au et al. 2004; Abdel-Rahman and El-Zein 2000; Duell et al. 2000), drug toxicity and clinical outcomes following treatment with cancer drugs and/or radiation (Booton et al. 2006; Ambrosone et al. 2006; Gurubhagavatula et al. 2004; Kamikozuru et al. 2008). The influence of individual gene polymorphisms on treatment outcome may be moderate, but reports from our laboratory (Bewick et al. 2006, 2008) and others (Gurubhagavatula et al. 2004; Ambrosone et al. 2005; Nagle et al. 2007; Nowell et al. 2005) suggest that combinations of polymorphisms in several genes may significantly impact clinical responses and survival.
The nucleotide excision repair (NER) pathway is important for DNA repair, removing adducts caused by UV irradiation and environmental agents such as tobacco smoke (polycyclic aromatic hydrocarbons). This pathway is also the main mechanism for the repair of bulky DNA adducts generated by platinum drugs (Altaha et al. 2004) and cyclophosphamide (Andersson et al. 1996) and may be involved in the repair of nonbulky DNA lesions that result from oxidative damage (Hutsell and Sancar 2005). In tumor tissue, low levels of NER gene expression are associated with increased survival for a number of cancers following platinum-based treatments (Azuma et al. 2007; Handra-Luca et al. 2007; Lee et al. 2007; Simon et al. 2007).
Genes involved in the NER pathway code for at least 16 proteins that form a complex nucleotide excision repairosome (Reed 2005). Key genes of the NER pathway include ERCC1 (excision repair cross-complementation group 1) and ERCC2 (excision repair cross-complementation group 2, XPD). ERCC2, a 5′ DNA helicase, is a component of the multi-subunit transcription factor TFIIH that unwinds the DNA duplex in the immediate area of the lesion. The endonuclease, ERCC1, is involved in excising the damage to facilitate repair.
Studies suggest that polymorphisms in the NER pathway influence survival outcome in lung (Gurubhagavatula et al. 2004), esophageal (Bradbury et al. 2009), colorectal (Le Morvan et al. 2007) and bone (Caronia et al. 2009) cancer following various DNA-damaging treatments. Functional polymorphisms in these NER genes have been reported that may affect DNA repair capacity and influence the efficacy of treatments using DNA-damaging agents (Wolfe et al. 2007; Au et al. 2003; Lunn et al. 2000). In the ERCC1 gene, the ERCC1 8092 C/A (rs3212986) polymorphism occurs with relatively high frequency. This polymorphism is located in the 3′-untranslated region and may affect mRNA stability (Chen et al. 2000). It is associated with survival outcome for patients with lung and esophageal cancer treated with cisplatin (Zhou et al. 2004; Wu et al. 2008; Bradbury et al. 2009).
A number of studies have examined two common polymorphisms in the ERCC2 gene, ERCC2 Asp312Asn (rs1799793) and ERCC2 Lys751Gln (rs13181). These polymorphisms are associated with variations in adduct levels (Benhamou and Sarasin 2005), chromosomal aberrations (Au et al. 2004; Affatato et al. 2004) and mRNA expression levels in lymphocytes (Wolfe et al. 2007).
Few studies have examined the relationship of these polymorphisms with survival outcome for patients with breast cancer. Therefore, we investigated the association of ERCC1 8092 C/A (rs3212986), ERCC2 Asp312Asn and ERCC2 Lys751Gln with breast cancer-specific survival (BCSS) and progression-free survival (PFS) in 95 patients with metastatic breast cancer (MBC). These patients were enrolled in studies of high-dose chemotherapy (HDC) and autologous stem cell transplantation (ASCT).
Materials and methods
Patient population
The patient population consisted of women that were eligible and had participated in the clinical trials of high-dose chemotherapy and autologous stem cell transplantation. Eligibility criteria for these clinical trials included female patients with histologically proven breast cancer. Age eligibility was restricted to women between 18 and 56 years. Patients had metastatic (stage IV) breast cancer not previously treated with chemotherapy or treated with adjuvant chemotherapy or hormonal therapy and who had progressive disease at least 6 months after completion of adjuvant chemotherapy. Patients were hormone receptor negative or were hormone receptor positive and had progressed on hormone therapy or had rapidly progressing disease where response to hormone treatment could not be awaited.
Patient characteristics, HDC treatment regimens, follow-up and survival information were obtained from clinical research records. These are listed in Table 1 and have been previously described (Bewick et al. 2006, 2008). The clinical trials and study were approved by the Research Ethics Board, Sudbury Regional Hospital, Sudbury, Ontario, and informed signed consent was obtained from all patients.
Analysis of polymorphisms
Apheresis blood product, peripheral blood and bone marrow samples were prospectively collected during treatment, frozen and stored for future use. DNA was isolated from samples using the DNA Blood MiniKit (Qiagen, Mississauga, Ontario) following the manufacturer’s protocol. The National Cancer Institute SNP500Cancer database (http://snp500cancer.nci.nih.gov/snplist.cfm) provided information on polymorphisms (target sequence, frequency estimates and referenced, validated 5′-nuclease assays). Genotypes were determined using TaqMan® SNP genotyping assays and the ABI Prism 7900HT Sequence Detection System (Applied Biosystems, Foster City, Ca.). For quality control purposes, 25% of the samples were randomly selected and reanalyzed at least twice for each polymorphism: the results were 100% concordant. Assignment of genotypes was performed independently by two investigators blinded to the survival endpoints.
Statistical analysis
For each polymorphism, median survival was calculated for each genotype and genotype pairs assessed in codominant, dominant and recessive models. The final model was selected by combining the genotypes with similar median survivals for each polymorphism. Fisher’s exact test was used to evaluate the association between patient and tumor characteristics (age group, ER status, PR status, site and number of metastatic sites) and genotype or genotype combination for each polymorphism. Analyses included two endpoints: breast cancer-specific survival (BCSS) and progression-free survival (PFS). BCSS was defined as time (months) from study registration until death from metastatic disease or censorship (alive at the end of the follow-up time period). Survival information was abstracted from clinical research records. Progression-free survival (PFS) was defined as time (months) from study registration until documented progression of metastatic disease or censorship (not progressed during follow-up time period). In general, patients were assessed every 2–3 months for the first 2 years and then every 4–6 months thereafter until evidence of progression.
Survival curves were generated using the Kaplan–Meier product limit estimate of the survivorship function. Equality of survivorship function was assessed using the log-rank test. The Cox proportional hazard regression model defined hazard ratios (HR) and 95% confidence intervals (CI). All models were adjusted for age. Genotypes were assigned as “adverse” based on examination of median survival, HR and survival curves of various models of BCSS. Tests for trend were conducted by calculating the P-value for the coefficient in a proportional hazards regression models using the adverse variable coded as an ordinal variable. A “risk” score was obtained by summing the number of assigned adverse genotypes for each individual and analyzed for groups of patients with 0, 1, 2 and 3 adverse genotypes. Statistical analysis was done using Stata version 10.1 (Stata Corporation, College Station, TX).
Results
Patient characteristics
The median age at study entry was 45 years (range, 19–56). Disease progression occurred in 91 patients (96%), and 91 patients (96%) died due to MBC, 4 patients remained alive at final follow-up. The time at risk for disease progression ranged from 1.6 to 111.7 months. The time at risk for death ranged from 4.8 to 111.7 months. For the total group (n = 95), the median PFS was 10.4 months and the median BCSS was 22.4 months.
The characteristics of the patient group are shown in Table 1. Genotypes were not associated with ER status, PR status, bone, liver, lung and “other” metastases or number of metastatic sites with the exception of the ERCC2 Asp312Asn GG/AG genotype group that was associated with lymph node metastases compared to patients with the AA genotype (P = 0.01).
Patients were assigned to 4 major groups based on differences in HDC treatment regimens (a fifth group contained only 1 patient) (Table 1). There were no significant differences in PFS and BCSS (Kaplan–Meier survival and log-rank analysis of survivorship function) between major treatment groups, patients receiving 1 or 2 (n = 6) HDC treatments or patients treated with or without carboplatin.
In univariate analysis of patients with known hormone receptor status, there were significant differences for PFS (P = 0.02) and BCSS (P = 0.005) by estrogen receptor status (n = 84) but not by progesterone receptor status (n = 80, PFS P = 0.27, BCSS: P = 0.09). Survival differences were significant for patients with metastatic site(s) that included liver (PFS: P < 0.001; BCSS: P < 0.001). Patients with site(s) of metastases that included bone had significantly longer survival than any other metastatic sites (n = 53, PFS: P = 0.03; BCSS: P = 0.02).
Patients with more than one metastatic site (n = 41) had significantly decreased PFS (P = 0.002) but not BCSS (P = 0.15) compared to the combined patient group with unassessable disease (UD) or 1 metastatic site.
Genotypes, PFS and BCSS
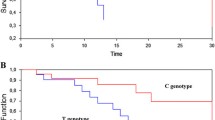
The genotype frequencies and HRs for PFS and BCSS are shown in Table 2. Significant differences for BCSS were observed for the ERCC2 Lys751Gln polymorphism. Median survival of patients with the ERCC2 Lys751Gln AA/AC combined genotypes was 24.8 months compared to 14.2 months for patients with the ERCC2 Lys751Gln CC genotype (HR:1.9, 95% CI 1.06–3.26). ERCC1 8092 C/A and ERCC2 Asp312Asn polymorphisms were not significantly associated with PFS or BCSS, and the ERCC2 Lys751Gln polymorphism was not significantly associated with PFS. ERCC2 Lys751Gln CC, ERCC2 Asp312Asn AA and ERCC1 8092C/A CC were assigned as adverse genotypes based on the analysis of HRs, Kaplan–Meier survival curves and associations reported in other studies (Le Morvan et al. 2007; Petty et al. 2007; Chew et al. 2009). Table 3 shows the effect on BCSS and PFS with increasing number of adverse genotypes. Median BCSS and PFS progressively decreased for patients with increasing number of adverse genotypes with significant test of trend for BCSS (P = 0.003) (Table 3). The risk of death (BCSS) was increased for patients with 1 adverse genotype (HR: 1.53, 95% CI 0.94–2.51) and was significantly increased for patients with 2 adverse genotypes (HR: 2.21, 95% CI 1.04–4.72) and with 3 adverse genotypes (HR: 6.67, 95% CI 2.19–20.29) when compared to patients with no adverse genotypes (P trend = 0.002) (Table 4). For PFS, results were not significant but were suggestive of increased risk of progression with increasing number of adverse genotypes (P trend = 0.06) (Table 5). Fig. 1 shows the Kaplan–Meier survival curves for BCSS for patients grouped according to the number of adverse genotypes.
Subgroup analysis of BCSS and PFS by treatment
Results of studies suggest that polymorphisms in the NER pathway may be important predictors of survival following the treatment of patients with platinum drugs. Therefore, we explored the influence of these polymorphisms in three major HDC treatment groups: patients treated with cyclophosphamide, mitoxantrone and carboplatin (n = 21); patients treated with cyclophosphamide, mitoxantrone and paclitaxel (n = 42); and patients treated with cyclophosphamide, mitoxantrone and vinblastine (n = 27). For BCSS, the results, shown in Table 4, suggest that for all treatment groups, patients with 1 or more adverse genotype were at increased risk and results were significant for patients with 3 adverse genotypes and treated with cyclophosphamide, mitoxantrone and vinblastine and P trend = 0.02 (Table 4). For PFS, the results, shown in Table 5, suggest that patients with 1 or more adverse genotypes treated with cyclophosphamide, mitoxantrone and carboplatin (n = 21) were at increased risk of progression with a significant test of trend (P trend = 0.02).
Discussion
Polymorphisms in genes involved in DNA repair and drug-related pathways may significantly influence clinical outcome and thus may help identify patients that can benefit from various treatments. This study examined the association of 3 polymorphisms in two genes in the NER pathway with survival outcome for patients with MBC treated with HDC using DNA-damaging drugs. In univariate analysis, results were suggestive of associations of the polymorphisms with increased risk of PFS and BCSS but only the ERCC2 Lys751Gln polymorphism was significantly associated with BCSS (HR:1.9, 95% CI1.06–3.26). In combined analyses of all 3 polymorphisms, median PFS and BCSS progressively decreased for patients with increasing number of adverse genotypes and the risk of death (BCSS) was significantly increased for patients with 2 or more adverse genotypes. Further analysis of treatment subgroups suggests that the risk of death was increased for patients in each of the three major treatment subgroups and was significant for the patient group with 3 adverse genotypes that received cyclophosphamide, mitoxantrone and vinblastine.
Overall, these results are biologically plausible and consistent with the role of the NER pathway in the repair of platinum-DNA adducts (Wang and Lippard 2005) and DNA damage resulting from cyclophosphamide (Andersson et al. 1996) and mitoxantrone (Durr et al. 1983). In addition, some studies have reported associations between polymorphisms in NER genes and cancer cell sensitivity in response to taxanes and to vinca alkaloids such as vinblastine (Le Morvan et al. 2006), although these may be mediated indirectly (Zhang et al. 1999; Reed et al. 1995). It is also possible that the association of these polymorphisms with BCSS overall and for treatment subgroups may indicate a potential role as prognostic factors. It is also interesting that these polymorphisms were strongly associated with PFS (Table 5) for the subgroup of patients that received carboplatin. This suggests that these polymorphisms may be predictive of outcome for platinum therapy.
The significant impact on survival outcome with increasing number of adverse genotypes within DNA repair pathways has been reported for other cancers. This includes patients with head and neck cancer following radiation treatment (Carles et al. 2006) and cisplatin treatment (Quintela-Fandino et al. 2006), and also for patients with esophageal cancer (Wu et al. 2006), and NSCLC treated with platinum-based chemotherapy (Wu et al. 2008; Gurubhagavatula et al. 2004). In vitro, the sensitivity of peripheral blood lymphocytes (measured by the number of chromatid breaks) to the mutagen benzo(α)pyrene diol epoxide (BPDE) increased with increasing number of designated adverse alleles in the NER pathway. These polymorphisms, when analyzed individually, had only moderate effects on mutagen sensitivity (Lin et al. 2007).
We used study-defined criteria to assign potential adverse genotypes as presently there are no definitive assignments of adverse genotypes for these polymorphisms and conflicting associations have been reported in the literature. A similar rationale has been used in previous studies (Lin et al. 2007).
Several studies report associations of ERCC2 Lys751Gln AA genotype with decreased DNA repair capacity (Vodicka et al. 2004; Lunn et al. 2000) and therefore it may be associated with a more favorable outcome following DNA-damaging treatment, while others report that the ERCC2 Lys751Gln Gln (C allele) variant is associated with decreased repair (Benhamou and Sarasin 2005; Matullo et al. 2001; Spitz et al. 2001; Hou et al. 2002; Rzeszowska-Wolny et al. 2005). In vitro, cancer cells with the ERCC2 751Gln CC genotype are more sensitive to taxanes and to vinca alkaloids, such as vinblastine (Le Morvan et al. 2006), suggesting that patients with the CC genotype should have better survival following these drug treatments. However, the results of clinical studies suggest that the ERCC2 751Gln CC genotype is predictive of poorer survival for patients with metastatic colorectal cancer treated with the platinum drug, oxaliplatin (Le Morvan et al. 2007), advanced NSCLC patients treated with docetaxel and gemcitabine (Petty et al. 2007) and MBC patients treated with gemcitabine and cisplatin (Chew et al. 2009). Conflicting results are also reported for the ERCC1 8092 C/A polymorphism. The A allele was significantly associated with poorer overall survival for patients with advanced NSCLC treated with platinum-based chemotherapy (Zhou et al. 2004) whereas in other studies, the A allele was associated with significantly improved overall survival for patients with esophageal (Bradbury et al. 2009) and NSCLC (Wu et al. 2008) as well as increased gastrointestinal toxicity for patients with NSCLC (Suk et al. 2005) treated with platinum drugs.
These inconsistent results may be due to differences between studies of cancer type and stage, patient population, combinations of drugs and treatments, types of DNA damage biomarkers examined and their method of assessment. Future research to validate potential biomarkers will be challenging since it is becoming more apparent that DNA repair pathways are complex with cross-functionality between pathways that may be differentially regulated and activated in various tissues and by different drugs and drug combinations (Zhang et al. 2009; Herrera et al. 2009).
We have attempted to follow the guidelines of the reporting recommendations for tumor marker prognostic studies (REMARK) (McShane et al. 2005). This is an exploratory retrospective study using a patient population of women who participated in several clinical trials of HDC and autologous stem cell transplantation. Study information was collected from clinical records rather than prospectively recorded. As such, there are a number of limitations of this study. The collection of information was not standardized and as a result is incomplete or missing for some patients for important clinical variables such as hormone receptor status and post-HDC treatments. In addition, the selection and number of polymorphisms examined was not comprehensive. In future prospective studies, it would be important to examine other potentially functional polymorphisms in NER and other DNA repair and cellular pathways appropriate for the drug combinations used. The sample size of this group of patients is small, limiting interpretation of the results, a limitation noted in similar studies. Although it is possible that the results of this study may be due to the high doses of drugs used, it should be noted that survival outcomes for patients treated with HDC are similar to patients treated with standard-dose chemotherapy regimens containing DNA-damaging agents (Farquhar et al. 2007). Also, the results of studies of patients with other cancers indicate that these and other polymorphisms in the NER pathway are important for the efficacy of DNA-damaging agents in standard-dose treatment regimens (Le Morvan et al. 2007; Gurubhagavatula et al. 2004; Bradbury et al. 2009).
In conclusion, these results suggest that combinations of polymorphism in NER may influence survival outcomes in patients with MBC following treatment with DNA-damaging drugs. To our knowledge, only one other study has examined polymorphisms in DNA repair pathways with clinical outcomes for metastatic breast cancer (Chew et al. 2009). Their results also suggest that polymorphisms in DNA repair genes may play an important role in clinical outcomes for patients with MBC following treatment with cisplatin and gemcitabine and similar to this study, the ERCC2 Lys751Gln polymorphism was associated with survival outcome. Future prospective studies examining combinations of polymorphisms in DNA repair and drug-related pathways in patients receiving standard-dose treatments are required to determine their usefulness for the development of personalized treatment strategies for MBC.
References
Abdel-Rahman SZ, El-Zein RA (2000) The 399Gln polymorphism in the DNA repair gene XRCC1 modulates the genotoxic response induced in human lymphocytes by the tobacco-specific nitrosamine NNK. Cancer Lett 159:63–71
Affatato AA, Wolfe KJ, Lopez MS et al (2004) Effect of XPD/ERCC2 polymorphisms on chromosome aberration frequencies in smokers and on sensitivity to the mutagenic tobacco-specific nitrosamine NNK. Environ Mol Mutagen 44:65–73
Altaha R, Liang X, Yu JJ et al (2004) Excision repair cross complementing-group 1: gene expression and platinum resistance. Int J Mol Med 14:959–970
Ambrosone CB, Ahn J, Singh KK et al (2005) Polymorphisms in genes related to oxidative stress (MPO, MnSOD, CAT) and survival after treatment for breast cancer. Cancer Res 65:1105–1111
Ambrosone CB, Tian C, Ahn J et al (2006) Genetic predictors of acute toxicities related to radiation therapy following lumpectomy for breast cancer: a case-series study. Breast Cancer Res 8:R40
Andersson BS, Sadeghi T, Siciliano MJ et al (1996) Nucleotide excision repair genes as determinants of cellular sensitivity to cyclophosphamide analogs. Cancer Chemother Pharmacol 38:406–416
Au WW, Salama SA, Sierra-Torres CH (2003) Functional characterization of polymorphisms in DNA repair genes using cytogenetic challenge assays. Environ Health Perspect 111:1843–1850
Au WW, Navasumrit P, Ruchirawat M (2004) Use of biomarkers to characterize functions of polymorphic DNA repair genotypes. Int J Hyg Environ Health 207:301–313
Azuma K, Komohara Y, Sasada T et al (2007) Excision repair cross-complementation group 1 predicts progression-free and overall survival in non-small cell lung cancer patients treated with platinum-based chemotherapy. Cancer Sci 98:1336–1343
Benhamou S, Sarasin A (2005) ERCC2/XPD gene polymorphisms and lung cancer: a HuGE review. Am J Epidemiol 161:1–14
Bewick MA, Conlon MS, Lafrenie RM (2006) Polymorphisms in XRCC1, XRCC3, and CCND1 and survival after treatment for metastatic breast cancer. J Clin Oncol 24:5645–5651
Bewick MA, Conlon MS, Lafrenie RM (2008) Polymorphisms in manganese superoxide dismutase, myeloperoxidase and glutathione-S-transferase and survival after treatment for metastatic breast cancer. Breast Cancer Res Treat 111:93–101
Booton R, Ward T, Heighway J et al (2006) Xeroderma pigmentosum group D haplotype predicts for response, survival, and toxicity after platinum-based chemotherapy in advanced nonsmall cell lung cancer. Cancer 106:2421–2427
Bradbury PA, Kulke MH, Heist RS et al (2009) Cisplatin pharmacogenetics, DNA repair polymorphisms, and esophageal cancer outcomes. Pharmacogenet Genomics 19:613–625
Carles J, Monzo M, Amat M et al (2006) Single-nucleotide polymorphisms in base excision repair, nucleotide excision repair, and double strand break genes as markers for response to radiotherapy in patients with Stage I to II head-and-neck cancer. Int J Radiat Oncol Biol Phys 66:1022–1030
Caronia D, Patino-Garcia A, Milne RL et al (2009) Common variations in ERCC2 are associated with response to cisplatin chemotherapy and clinical outcome in osteosarcoma patients. Pharmacogenomics J 9:347–353
Chen P, Wiencke J, Aldape K et al (2000) Association of an ERCC1 polymorphism with adult-onset glioma. Cancer Epidemiol Biomarkers Prev 9:843–847
Chew HK, Doroshow JH, Frankel P et al (2009) Phase II studies of gemcitabine and cisplatin in heavily and minimally pretreated metastatic breast cancer. J Clin Oncol 27:2163–2169
Duell EJ, Wiencke JK, Cheng TJ et al (2000) Polymorphisms in the DNA repair genes XRCC1 and ERCC2 and biomarkers of DNA damage in human blood mononuclear cells. Carcinogenesis 21:965–971
Durr FE, Wallace RE & Citarella RV (1983) Molecular and biochemical pharmacology of mitoxantrone. Cancer Treat Rev 10 Suppl B:3-11
Farquhar CM, Marjoribanks J, Lethaby A et al (2007) High dose chemotherapy for poor prognosis breast cancer: systematic review and meta-analysis. Cancer Treat Rev 33:325–337
Gonzalez-Angulo AM, Morales-Vasquez F, Hortobagyi GN (2007) Overview of resistance to systemic therapy in patients with breast cancer. Adv Exp Med Biol 608:1–22
Gurubhagavatula S, Liu G, Park S et al (2004) XPD and XRCC1 genetic polymorphisms are prognostic factors in advanced non-small-cell lung cancer patients treated with platinum chemotherapy. J Clin Oncol 22:2594–2601
Handra-Luca A, Hernandez J, Mountzios G et al (2007) Excision repair cross complementation group 1 immunohistochemical expression predicts objective response and cancer-specific survival in patients treated by Cisplatin-based induction chemotherapy for locally advanced head and neck squamous cell carcinoma. Clin Cancer Res 13:3855–3859
Herrera M, Dominguez G, Garcia JM et al (2009) Differences in repair of DNA cross-links between lymphocytes and epithelial tumor cells from colon cancer patients measured in vitro with the comet assay. Clin Cancer Res 15:5466–5472
Hou SM, Falt S, Angelini S et al (2002) The XPD variant alleles are associated with increased aromatic DNA adduct level and lung cancer risk. Carcinogenesis 23:599–603
Hutsell SQ, Sancar A (2005) Nucleotide excision repair, oxidative damage, DNA sequence polymorphisms, and cancer treatment. Clin Cancer Res 11:1355–1357
Kamikozuru H, Kuramochi H, Hayashi K et al (2008) ERCC1 codon 118 polymorphism is a useful prognostic marker in patients with pancreatic cancer treated with platinum-based chemotherapy. Int J Oncol 32:1091–1096
Le Morvan V, Bellott R, Moisan F et al (2006) Relationships between genetic polymorphisms and anticancer drug cytotoxicity vis-a-vis the NCI-60 panel. Pharmacogenomics 7:843–852
Le Morvan V, Smith D, Laurand A et al (2007) Determination of ERCC2 Lys751Gln and GSTP1 Ile105Val gene polymorphisms in colorectal cancer patients: relationships with treatment outcome. Pharmacogenomics 8:1693–1703
Lee HW, Han JH, Kim JH et al. (2007) Expression of excision repair cross-complementation group 1 protein predicts poor outcome in patients with small cell lung cancer. Lung Cancer
Lin J, Swan GE, Shields PG et al (2007) Mutagen sensitivity and genetic variants in nucleotide excision repair pathway: genotype-phenotype correlation. Cancer Epidemiol Biomarkers Prev 16:2065–2071
Lunn RM, Helzlsouer KJ, Parshad R et al (2000) XPD polymorphisms: effects on DNA repair proficiency. Carcinogenesis 21:551–555
Matullo G, Palli D, Peluso M et al (2001) XRCC1, XRCC3, XPD gene polymorphisms, smoking and (32)P-DNA adducts in a sample of healthy subjects. Carcinogenesis 22:1437–1445
McShane LM, Altman DG, Sauerbrei W et al (2005) Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst 97:1180–1184
Nagle CM, Chenevix-Trench G, Spurdle AB et al (2007) The role of glutathione-S-transferase polymorphisms in ovarian cancer survival. Eur J Cancer 43:283–290
Nowell SA, Ahn J, Rae JM et al (2005) Association of genetic variation in tamoxifen-metabolizing enzymes with overall survival and recurrence of disease in breast cancer patients. Breast Cancer Res Treat 91:249–258
Petty WJ, Knight SN, Mosley L et al (2007) A pharmacogenomic study of docetaxel and gemcitabine for the initial treatment of advanced non-small cell lung cancer. J Thorac Oncol 2:197–202
Quintela-Fandino M, Hitt R, Medina PP et al (2006) DNA-repair gene polymorphisms predict favorable clinical outcome among patients with advanced squamous cell carcinoma of the head and neck treated with cisplatin-based induction chemotherapy. J Clin Oncol 24:4333–4339
Reed E (2005) ERCC1 and clinical resistance to platinum-based therapy. Clin Cancer Res 11:6100–6102
Reed E, Kohn EC, Sarosy G et al (1995) Paclitaxel, cisplatin, and cyclophosphamide in human ovarian cancer: molecular rationale and early clinical results. Semin Oncol 22:90–96
Rzeszowska-Wolny J, Polanska J, Pietrowska M et al (2005) Influence of polymorphisms in DNA repair genes XPD, XRCC1 and MGMT on DNA damage induced by gamma radiation and its repair in lymphocytes in vitro. Radiat Res 164:132–140
Simon GR, Ismail-Khan R, Bepler G (2007) Nuclear excision repair-based personalized therapy for non-small cell lung cancer: from hypothesis to reality. Int J Biochem Cell Biol 39:1318–1328
Spitz MR, Wu X, Wang Y et al (2001) Modulation of nucleotide excision repair capacity by XPD polymorphisms in lung cancer patients. Cancer Res 61:1354–1357
Suk R, Gurubhagavatula S, Park S et al (2005) Polymorphisms in ERCC1 and grade 3 or 4 toxicity in non-small cell lung cancer patients. Clin Cancer Res 11:1534–1538
Vodicka P, Kumar R, Stetina R et al (2004) Genetic polymorphisms in DNA repair genes and possible links with DNA repair rates, chromosomal aberrations and single-strand breaks in DNA. Carcinogenesis 25:757–763
Wang D, Lippard SJ (2005) Cellular processing of platinum anticancer drugs. Nat Rev Drug Discov 4:307–320
Wolfe KJ, Wickliffe JK, Hill CE et al (2007) Single nucleotide polymorphisms of the DNA repair gene XPD/ERCC2 alter mRNA expression. Pharmacogenet Genomics 17:897–905
Wu X, Gu J, Wu TT et al (2006) Genetic variations in radiation and chemotherapy drug action pathways predict clinical outcomes in esophageal cancer. J Clin Oncol 24:3789–3798
Wu X, Lu C, Ye Y et al (2008) Germline genetic variations in drug action pathways predict clinical outcomes in advanced lung cancer treated with platinum-based chemotherapy. Pharmacogenet Genomics 18:955–965
Zhang CC, Yang JM, Bash-Babula J et al (1999) DNA damage increases sensitivity to vinca alkaloids and decreases sensitivity to taxanes through p53-dependent repression of microtubule-associated protein 4. Cancer Res 59:3663–3670
Zhang Y, Rohde LH, Wu H (2009) Involvement of nucleotide excision and mismatch repair mechanisms in double strand break repair. Curr Genomics 10:250–258
Zhou W, Gurubhagavatula S, Liu G et al (2004) Excision repair cross-complementation group 1 polymorphism predicts overall survival in advanced non-small cell lung cancer patients treated with platinum-based chemotherapy. Clin Cancer Res 10:4939–4943
Acknowledgments
We thank the Northern Cancer Research Foundation for their support of this research.
Conflict of interest statement
We declare that we have no conflict of interest in relation to this work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bewick, M.A., Lafrenie, R.M. & Conlon, M.S.C. Nucleotide excision repair polymorphisms and survival outcome for patients with metastatic breast cancer. J Cancer Res Clin Oncol 137, 543–550 (2011). https://doi.org/10.1007/s00432-010-0915-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-010-0915-7