Abstract
Purpose
To screen potential serological biomarkers and develop decision tree classifications of chronic hepatitis B, liver cirrhosis (LC) and hepatocellular carcinoma (HCC), respectively, with high prediction score for improving diagnosis of liver diseases.
Methods
The total serum samples were randomly divided into three training sets (41 HBV and 35 health; 36 LC and 35 health; 39 HCC and 35 health) and three testing groups (34 HBV and 38 health; 18 LC and 52 health; 42 HCC and 47 health). Selected WCX2 protein chip capture followed by SELDI-TOF-MS analysis was applied to generate the serum protein profiles. Subsequently serum protein spectra were normalized and aligned by Ciphergen SELDI Software 3.1.1 with Biomarker Wizard including baseline subtraction, mass accuracy calibration, automatic peak detection. Once the intensities of selected significant peaks from the training data set were transferred to further BPS analysis, an optimized classification tree with sequence-decision was established to divide training data set into disease group and control group successfully. A double blind test was employed to determine the clinical sensitivity and clinical specificity of three models.
Results
After comparative analysis of SELDI based serum protein profile between the cases of disease and healthy, a HCC decision tree classification with sensitivity of 94.872% and specificity of 94.286%; a LC decision tree classification with sensitivity of 91.667% and specificity of 94.286% and a HBV decision tree classification with sensitivity of 95.122% and specificity of 94.286% were produced by BPS respectively. When three decision tree models were challenged by the double-blind test samples, clinical sensitivity and clinical specificity of these models were predicted in diagnosis of three liver diseases (HCC: 90.48 and 89.36%; cirrhosis: 100 and 86.5%; HBV: 85.29 and 84.21%).
Conclusion
SELDI-based decision tree classifications showed great advantages over conventional serological biomarkers in the diagnosis of chronic hepatitis B, LC as well as HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Despite some significant improvement in diagnosis and treatment of human liver diseases over last decades, chronic hepatitis B, liver cirrhosis (LC) and hepatocellular carcinoma (HCC) still intimidate the public health and remain unchanged rank in high burden diseases in China. Previous epidemiological studies have shown that there were more than 350 million of hepatitis B virus (HBV) carriers in the world. Most of them (120 million) were in china (Purcell 1993; Lai et al. 2003). HBV, a leading cause of chronic hepatitis, was linked to the development of LC. Approximately 10–20% of the high HBV infection cases were reported to form LC (Ganem and Prince 2004) and 55–85% of LC cases finally develop to HCC with poor prognosis (Zhu et al. 2004). Liver cirrhosis and HCC accounted for 1 million deaths annually (Lee 1997). HCC was the third frequent cause of cancer death worldwide, second in China (Parkin et al. 2001; Pisani et al. 1999). It was estimated that 52% HCC occurred on a background of HBV and 25% HCC were associated with HCV infection (Bosch et al. 1999), As reported, more than 90% of HCC were found to have HBV background in high incidence area of china (Gao et al. 2005). Presently chronic hepatitis B, hepatitis C and LC are all recognized as the major risk factors for HCC worldwide. In addition, about 10% of patients dying with cirrhosis were identified to have unsuspected HCC at postmortem (Seow et al. 2001), and 3–6% of explanted livers containing small unsuspected HCC was reported in liver transplantation (Figueras et al. 1997). These data mentioned above not only demonstrated that there existed strong association in etiology and pathological mechanism among these three liver diseases, but also indicated that it was a challengeable to screen ideal biomarkers for early diagnosis or differential diagnosis only relying on traditional approaches, especially in these multifactor and multigene diseases.
As a key organ with biological importance, the abnormal function of liver would cause a great alteration in serum protein profile undoubtedly. Such serum proteins as HBV marker (HBVM) (including HBsAg, anti-HBS, HBeAg, anti-HBE and anti-HBC), hyaluronic acid (HA), type III procollagen peptide, laminin, type IV collagen, DCP and AFP, have been verified to be the indicators to assess pathological initiation, development and prognosis of liver injury. Although these traditional serum biomarkers can play a role in clinical diagnosis and surveillance of these diseases, pretreatment liver biopsy is still considered as only gold standard for evaluating the pathological grade of liver injury. This phenomenon is in part related to their poor sensitivity and specificity. Even though AFP, a well-known biomarker in serum for HCC diagnosis and prognosis in clinic, its sensitivity was reported from 39 to 64% and specificity ranged from 76 to 91%. This means that up to 40% patients bearing HCC have normal AFP level (false negative) and about 9–15% patients with hepatitis and LC have elevating level of AFP (false positive) (Schwegler et al. 2005). So screening a set of reliable, reproducible and noninvasive new biomarkers will have significant impacts on the clinic diagnosis and treatment of liver diseases.
Development of high throughput proteomics technology provides a novel pathway to large-scale screening and identifying biomarkers in body fluid (serum, plasma, urine, etc.) and tissue. For comparative proteomics study, 2-DE followed by mass spectrometry analysis as core technology always maintain dominant status in finding the differential displayed proteins and identifying the novel biomarkers these years. Applying this classic technology strategy, a remarkable finding associated with carcinogenesis, 14-3-3γ protein, was identified by integrating 2D-DIGE with nano-LC–MS/MS (Lee et al. 2005). In another study, two up-regulated proteins PCNA and stathmin1 in HCC tumor tissue were also manifested to be related to HCC carcinogenesis after comparative analysis of expression profile of ten tumorous tissues and their paired adjacent non-tumor tissues, (Li et al. 2005). Because the tissue samples were obtained under invasive procedure, the application of these discovered novel biomarkers were limited partly in clinical diagnosis. While there are great advantages in protein separation and viewing for 2-DE based technique pathway, it is not somewhat suitable to screen biomarkers in numerous clinical samples and large-scale clinical testing analysis. Recently another promising proteomics technology, SELDI-TOF-MS (surface enhanced laser desorption ionization -time of flight-mass spectrometry), was introduced to remedy shortcoming of 2-DE. SELDI based technique performance was similar as the clinical assay and was prone to clinical application. Up to now, SELDI based technique coupled with bioinformatics has been successfully applied to screen and verify the cases of breast (Li et al. 2002), prostate (Adam et al. 2002), pancreatic (Yu et al. 2005), ovarian (Zhang et al. 2004) and liver cancer (Schwegler et al. 2005) by establishing decision tree classification, as well as to identify new biomarker of HCC (Paradis et al. 2005).
The aim of this study was to find potential serum biomarkers and to establish the diagnostic decision tree classification by comparative analysis of SELDI serum protein profiles between three diseases and health controls.
Materials and methods
Serum sample collection
Serum samples from HCC patient (n = 81) and health control (n = 87) were collected with informed consent from Liver Cancer Institute and Department of Clinical Laboratory Medicine of Zhongshan Hospital, Fudan University. All HCC patients were diagnosed according to clinical combined criteria including image evidence (focal lesion >2 cm with arterial hypervascularization), serum AFP level >400 ng/ml, and pathological confirmation (Bruix et al. 2001). Health controls with no history of liver disease were proved to be normal biochemical function of liver and without any viral hepatitis. Samples of LC patients (n = 54) were provided friendly by Department of Infection of Shanghai East Hospital. The diagnosis of these patients was mainly depended on clinical history, physical examination, laboratory finding and ultrasonographic and/or computed tomographic imaging with or without liver biopsy (Erlinger and Benhamou 1999). Serum samples were collected in patients with chronic HBV hepatitis (n = 75) with consent from Shandong Provincial Hospital based on HBsAg (+), HBeAg (+) and HBVDNA (+). All samples were randomly divided into three training groups (41 HBV and 35 health; 36 LC and 35 health; 39 HCC and 35 health) and three testing groups (34 HBV and 38 health; 18 LC and 52 health; 42 HCC and 47 health).
All serum samples were separated as routine laboratory procedure and were aliquoted in 10 μl each then stored at −80°C until used.
Reproducibility analysis of SELDI protein spectra
To evaluate the reproducibility of serum SELDI based mass spectra, several independent protein peaks of same serum sample were randomly selected to calculate the CV (coefficient of variance) of intensity and CV of M/Z in intra-assay and inter-assay, respectively.
SELDI serum protein profile
Four kinds of protein chips with different chemical modified surface (cationic, anionic, copper mental binding and hydrophobic, Ciphergen Biosystems, Fremont, CA) were initially selected to determine which chip could produced the best serum protein profile. The weak cation exchange protein chip (WCX2) was accepted and applied in present study for presence of more different protein fingerprint peaks.
Three microliters of serum samples were diluted in 6 μl U9 sample buffer (9 M urea, 2% CHAPS, 50 mM Tris–HCl, 1% DTT, pH 9.0, Sigma) and vortexed for 30 min at 4°C. After that, 90 μl of above samples were further added in 108 μl WCX2 binding buffer (50 mM sodium acetate, pH 4.0, Sigma). A total of 100 μl of diluted serum samples were added to each spot on pre-equilibrated WCX2 protein chips packed in a processor and incubated under shaking at 500 rpm/h for 60 min. Subsequently unbinding proteins on chips were washed out, and then further washed two times with WCX2 binding buffer and rinsed quickly with HPLC grade water one time, respectively. Finally 1 μl of a saturated solution of the EAM sinapinic acid (Ciphergen Biosystems) in 50% acetonitrile and 0.5% trifluoroacetic acid (Sigma) was loaded on each spot and air-dried for future detection.
The protein chips were analyzed using PBSII mass spectrometry system (Ciphergen Biosystems). Before chip processing, mass accuracy was calibrated daily using the all-in-one peptide molecular mass standard (Ciphergen Biosystems). The chips were detected according to following optimized parameters: laser intensity 185; detector sensitivity 8; focus by optimization center; Mass Deflector 1,000 Da; data acquisition parameters 22. delta to 4. transients per to 7 ending position to 82 by Seldi Quantitation; high mass 50,000 Da, molecular mass range 2,000–20,000 Da. After all raw data were automatically collected, these data were further normalized and aligned by Ciphergen SELDI Software 3.1.1 with Biomarker Wizard including baseline subtraction, mass accuracy calibration, automatic peak detection. Protein peaks ranging from 2,000 to 30,000 Da were determined in term of S/N > 5 and minimum threshold for cluster >20%.
Decision tree classification and double blind test
Biomarker Pattern Software (BPS, Ciphergen Biosystems) is a multivariate tool, which strength was to find a pattern of protein markers. Once the intensities of selected peaks identified by Biomarker Wizard Software from training data set were transferred to BPS, a root classification tree was generated firstly. Subsequently various tree classification trees were test and screened by adjusting parameter set. Finally a classification tree with sequence-decision (decision tree classification) was established to divide all the training cases into disease group and control group depend on selected peak intensity. If the corresponding peak intensity of a sample was lower than or equal to the threshold of node, this sample was classified in left sub-node. Otherwise, the sample would be classified in right sub-node. Such sequence-decision continued to perform until entered to the terminal node.
Three different testing groups were parallel used to test clinical efficiency of decision tree classification of three diseases respectively. Protein peaks from the testing samples were normalized and selected with Ciphergen SELDI Software 3.1.1 and Biomarker Wizard as the same as data mining of training set, and then the identity of testing samples were determined with decision tree classification of BPS based on peak selected intensities. Through comparative analysis of above result and corresponding clinical data, the prediction value of decision tree classification (including clinical sensitivity, clinical specificity and positive predictive value) was presented.
Results
Reproducibility of SELDI system
The reproducibility of SELDI system was demonstrated by the changes of mass location (M/Z) and intensity of protein peaks from array to array on one chip (intra-assay) and between chips (inter-assay). Eight independently protein spectra of same healthy serum and ten peaks with relative intensities above 10 and highest amplitude <10% were randomly used to determine intra-assay reproducibility of SELDI system. The intra-assay CV for intensity and M/Z were 17.46 and 0.024%, respectively. Nine independently protein spectra of same healthy serum and nine peaks with relative intensities above 10 and highest amplitude <10% were applied to calculate inter-assay reproducibility of SELDI system. The inter-assay CV for intensity and M/Z were 17.74 and 0.0237% respectively. The protein spectra with little variation for one sample between one chip and different chips suggested SELDI system in present study could be used to screen different protein peaks in serum protein profiles between healthy and patients.
Data analysis and HCC decision tree classification and double blind test
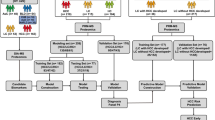
The mass spectra produced using SELDI-TOF-MS from 74 samples in the training set (39 HCC and 35 health) were firstly analyzed with Ciphergen SELDI Software 3.1.1 and Biomarker Wizard. Sum of 120 protein peaks in the area between 2,000 and 30,000 Da were identified under the condition of signal-to-noise >5 and minimum threshold for cluster >20%. Seventy-three of these protein peaks were found to be statistic significant difference between HCC and health control (P < 0.05). In the help of BPS power screening and optimizing in multivariate combination, a decision tree classification with five terminal nodes was built up to separate training cases into HCC bearing group and health control (Fig. 1). Two up-regulated discriminating proteins (23,381 and 28,040.6) and one down-regulated protein (2,018.5) in HCC were integrated to form the decision tree model (Fig. 1). Peak 23,381, the most important protein peak, was located at node 1. If the intensity of peak 23,381 was lower than or equal to 0.197, these samples went to node 2 (including 96.97% class 1 sample and 3.03% class 0 sample) and other samples accessed to node 3 (including 92.68% class 0 sample and 7.32% class 1 sample). Subsequently samples at next nodes were further classified into corresponding groups according to the intensity threshold of other discriminating protein peaks. After sequence-decision, all samples reached terminal node, meanwhile a sensitivity and specificity of this mode was presented 94.872% (37/39) and 94.286% (33/35), respectively, by BPS analysis.
Top panel Diagram of hepatocellular carcinoma (HCC) decision tree classification for HCC patients and healthy controls. The squares represent the parent nodes and descendant nodes labeled with number and the broken line frames denote terminal nodes. If the corresponding peak intensity of a sample was lower than or equal to the threshold of each node (class0 HCC, class1 healthy control), this sample was classified in left side sub-node. Otherwise, the sample would be classified in right side sub-node. Lower panel Two up-regulated discriminating proteins (23,381 and 28,040.6) and one down-regulated protein (2,018.5) in HCC in comparison with health control. HCC serum protein spectra of HCC, N serum protein spectra of health control
The testing set (42 HCC and 47 health) was applied to assess clinical efficiency of HCC decision tree classification. Serum protein profiles of these samples were exactly collected and treated as the same as those above mentioned including spot reading, normalization and protein peak selection. Once intensity of protein peaks were transferred to BPS for challenging HCC model, the identity of double-blind samples would be determined. By comparative analysis of result above and corresponding clinical data, clinical sensitivity of HCC decision tree classification (38/42, 90.48%), clinical specificity of HCC decision tree classification (42/47, 89.36%), and positive predictive value (88.37%) were calculated.
Liver cirrhosis decision tree classification and double blind test
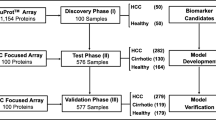
After standardizing the mass spectra of training set (36 cirrhosis and 35 health) in the case of signal-to-noise >5 and minimum threshold for cluster >20%, 94 among 144 protein peaks between 2,000 and 30,000 Da. were discovered to be significant difference between HBV and health control (P < 0.05). Follow on similar pathway of constructing model above-described, a decision tree classification with three terminal nodes, which yielded a sensitivity of 91.667% and specificity of 94.286%, was established to separate training samples into cirrhosis group and health control group (Fig. 2). One down-regulated discriminating protein peak of 2,049.74 while another up-regulated protein peak of 3,166.05 in cirrhosis in comparison with health control were shown at the node of this model (Fig. 2). When the double-blind testing set (18 LC and 52 health) was applied to challenge this model, a clinical sensitivity of 100% (18/18), a clinical specificity of 86.5% (45/52) and positive predictive value of 72% were predicted.
Top panel Diagram of cirrhosis decision tree classification for cirrhosis patients and healthy controls. The parent nodes and descendant nodes labeled with number were shown as squares and terminal nodes were shown as the broken line frames. If the corresponding peak intensity of a sample was lower than or equal to the threshold of each node (class0 cirrhosis, class1 healthy control), this sample was classified in left side sub-node. Otherwise, the sample would be classified in right side sub-node. Lower panel One discriminating protein peak of 2,049.74 was down-regulated while another protein peak of 3,166.05 was up-regulated in cirrhosis in comparison with health control. C serum protein spectra of cirrhosis, N serum protein spectra of health control
HBV decision tree classification and double blind test
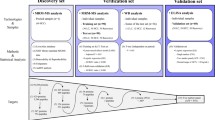
Seventy-two of 124 protein peaks were demonstrated to be significant difference between HBV and health control (P < 0.05) after mass spectra analysis of 76 training group samples (41 HBV and 35 health) under same parameter set above-described. As the intensities of these peaks were transferred to BPS, HBV decision tree classification with four terminal nodes was determined with a sensitivity of 95.122% and a specificity of 94.286% (Fig. 3). One discriminating protein with the M/Z of 22,842.3 at node1 was up-regulated in HBV and another two discriminating proteins with M/Z value of 2,957.83 and 2,049.57 appeared to be down-regulated in HBV (Fig. 3). A clinical sensitivity of 85.29% (29/34), the specificity of 84.21% (32/38) and a positive predictive value of 82.86% for HBV decision tree classification were obtained in double blind test.
Top panel Diagram of HBV decision tree classification for HBV patients and healthy controls. The squares represent the parent nodes and descendant nodes labeled with number and the broken line frames denote terminal nodes. If the corresponding peak intensity of a sample was lower than or equal to the threshold of each node (class0 HBV, class1 healthy control), this sample was classified in left side sub-node. Otherwise, the sample would be classified in right side sub-node. Lower panel One discriminating protein of 22,842.3 at node1 was up-regulated in HBV and other two discriminating proteins of 2,957.83 and 2,049.57 appeared to be down-regulated in HBV in comparison with health control. HBV serum protein spectra of hepatitis B, N serum protein spectra of health control
Discussion
The serum protein profile could be changed constantly as a consequence of the perfusion of organs systems underlying pathological status in which there are some released proteins or small proteolytical or enzymatical peptides and their derivatives. So serum was believed to provide a rich pool of disease associated proteins. However, separation and identification of serological biomarker was laborious and difficulty relying on classical approach and screening strategy. Although some identified serological biomarkers were shown to be definitely significance in diagnosis and prognosis, its clinical evaluation remained dismal due to the relative lower sensitivity and specificity. From the other point of view, the single molecular diagnostic model or screening method was not enough to cove all the disease states, and the potential valuable candidates of disease biomarker might be lower molecular or lower abundant proteins in serum. More evidences suggested that the most uncharacterized serological proteins appeared to be closely associated with pathological process (Tirumalai et al. 2003; Petricoin and Liotta 2004). Recently mass spectrometry-based technology with optical performance was widely applied for discovery of serological biomarkers with low molecular weight. Being a new exemplification for biomarker screening and disease detection, SELDI-based analysis have now been accepted for clinical routine diagnosis trial in large commercial reference laboratories after it has been shown to be useful in the discovery of potential diagnostic markers for disease (Petricoin and Liotta 2004).
To screen and identify ideal serological biomarkers for pathological states of liver injury (chronic hepatitis B, LC and hepatocellular carcinoma) always attracted much attentions in science and medicine. Although a lot of studies were carried out on it, current serological biomarkers were unable to be well qualified for clinical diagnosis requirement. At present, the diagnostic accuracy of HCC was still relatively lower in early diagnosis, prediction or epidemiological screening only with conventional serological biomarkers and without liver biopsy or image analysis. Because modestly elevated AFP was occasionally detected in serum of nonmalignant chronic liver disease, mainly in chronic hepatitis and LC, elevated AFP in serum cannot specifically reflect pathological states of HCC, thereby reduce the diagnosis efficient of AFP in HCC diagnosis (Schwegler et al. 2005; Poon et al. 2003). Another serological biomarker DCP (des-gamma carboxyprothrombin) was reported to be better than AFP in differentiating HCC from nonmalignant chronic liver disease by a cross-sectional case control study, a DCP value more than125 mAU/ml among American patients with underlying chronic liver disease was associated with a high probability of HCC (sensitivity and specificity approximately 90%) (Marrero et al. 2003). However, other studies on DCP evaluation implied that DCP was not superior to AFP or no significant difference between DCP and AFP (Marrero et al. 2003; Grazi et al. 1995; Brunello et al. 1993). The value of DCP in HCC diagnosis still need to be evaluated in future study. In this study, a serum decision tree classification with five terminal nodes was built up to distinguish between HCC and healthy successful. A best clinical sensitivity of 90.48% and a specificity of 89.36% were yielded, which was higher than prediction score of AFP (sensitivity39–64%; specificity 76–91%) and the most common imaging modality-ultrasound (sensitivity 71–78%; specificity 90–93%) (Marrero et al. 2003). Schwegler et al. (2005) analyzed the serum protein profiling of liver disease progression to HCC by SELDI-TOF-MS and got the decision tree classifications for different sets of liver diseases versus normal samples as well as HCV versus HCV-HCC. In the case of all HCC versus normal, a sensitivity of 82% and a specificity of 90% were documented. It was also better than AFP, but slightly lower than our results. A possible consideration was that the sum of serum protein fingerprint peaks on IMAC-Cu protein chip was much less than that on WCX protein chip under the same sample loading, no more different protein peaks can be selected for further building a better decision tree classification. Another similar study with serum fractionation prior to loading on protein chip and different data mining (two-way hierarchical clustering analysis and ANN) showed that ANN model with a high sensitivity of 92% and a high specificity of 90% could well differentiate HCC from CLD patient, even when serum AFP concentrations were within the diagnostic gray region (Poon et al. 2003). This promising result definitely indicated that SELDI-based decision tree classification also possessed obvious advantages on solving difficult clinical diagnosis problem such as HCC serum with negative AFP and CLD serum with elevated AFP.
From clinic point of view, current single serological biomarker including laminin, HA, type IV collagen, type III precollagen peptide, etc., was not satisfactory as diagnostic and prognostic indicator for LC. Even the combination of several serological biomarkers for diagnosis of LC, sensitivity of only 90% at maximum and specificity of about 60% was presented (Oh and Afdhal. 2001). When a gold standard-pretreatment liver biopsy was integrated to assess LC, its accuracy could be improved to 80% (Poniachik et al. 1996). In our study, two differential protein peaks with M/Z of 2,049.74 and 3,166.05 were finally selected to form LC decision tree classification and then confirmed by the identity of 70 double-blind test, a clinical sensitivity of 100% and a specificity of 86.5% were obtained. It was much better than that of classic serological index. The remarkable diagnostic performance of this model displayed a bright perspective in clinical diagnosis of LC with SELDI-based biomarker discovery. Another classification model with high sensitivity and specificity (97.1–100 and 92.3%, respectively) for LC using selected serum protein marker set had also been reported (Xu et al. 2004). High prediction score of classification model proved out its great advantages over traditional serological biomarkers in cirrhosis diagnosis. Meanwhile it also showed that the perfect decision tree classification model was easily to be established in serum samples with similar genetic background and coincident treatment. Zhu et al (2004) analyzed the serum protein spectra derived from 25 LC samples and 45 control samples (20 HCC plus 25 healthy) by SELDI-TOF-MS and BPS analysis, a classification model with two low molecule weight biomarkers of 7,772 and 3,933 (M/Z) was constructed for LC prediction, which was the sensitivity of 80% for all LC patients and a specificity of 81.8% for all non-cirrhotic cohorts through the methods mentioned above. Because control grouping was set as HCC patients and healthy men simultaneously in their study, prediction score of tree model was little lower in comparison with our results.
Diagnosis of HBV infection has long been relied on two sides of serological proof. One side emphasized particularly on serology analysis of HBV related index, such as HBsAg, anti-HBS, HBeAg, anti-HBE, anti-HBC, HBVDNA, anti-HBcIgM, etc. Other side concentrated on the measurement of liver-associated serum enzymes or serum protein profile. The latter could be used for evaluation of biological function states of liver, such as ALT, AST, LDH, AKP, TP, A, G, etc. Unfortunately no single serological biomarkers described above could unequivocally diagnose chronic HBV infection (Chan 2002). Though chronic HBV infection was defined as the persistence of HBsAg for at least 6 months (Lok and McMahon 2001), negative HBsAg in serum could not be exclude HBV infection (Chan 2002). Current serological test for HBsAg was only 1 ng/ml, which was lower than the minimum infections dose of virus (1 pg/ml). Recently HBV DNA has been used increasingly in clinic as one of the important hallmark for diagnosis of chronic HBV infection, monitoring the disease progress as well as assessing curative effect. According to the difference in detection principle, current assays for the quantitative measurement of HBV DNA could be classified into signal amplification assays (liquid phase hybridization, branched-chain DNA signal amplification) and PCR-based DNA amplification. The former’s sensitivity was around 1 pg of DNA (105–106 genome copies) and the latter method could reach 102–103 genome copies (Sablon and Shapiro 2005). However, affected by several factors including reproducibility, considerable skill and high cost, HBV DNA assay is not presently recommended for routine evaluation as well as monitor chronic HBV infection. Such biomarkers focused on pathogen itself. For studies of serum protein expression profile derived from chronic HBV infection, there was almost no more progress in the area of novel serological biomarker on HBV infection. Up to now, only a proteomics study on serum biomarkers of HBV infection was reported by He et al. (2003). By comparison with normal and HBV negative serum samples, at least seven proteins including haptoglobinβ and α2 chain, apolipoproteinA-1 and A-IV, α1 antitripsin, transthyretin and DNA topoisomerase Iiβ were discovered to be significantly changed in HBV infected sera. Though these new findings had potential significance in clinic, a definite conclusion was not presented in their study because of no validation in clinical serum samples. Difference with 2DE based technique pathway of above study, HBV decision tree classification with clinical sensitivity of 85.29% and a clinical specificity of 84.21% was first reported in this study after analysis of serum protein pattern between HBV patients and healthy with SELDI-TOF-MS and BPS software. Because serum proteome of chronic hepatitis B contained double pathological information of virus and liver, screened different serum proteins could reflect real pathological state of HBV infection very well. From this point of view, there might be a bright application in diagnosis of HBV infection for this tree model.
All data and established diagnostic models in the present study were very encouraging for clinical application. The proceeding study now is under process. Consideration have been made to further verification of its specificity, sensitivity and the accurate determination in more large population, to screen the potential patients from the so called healthy case in high risk area of China. It also should be noticed to integrate the clinic data and pathological disease staging and possible grade and to standardize the parameters of given machinery statement, meanwhile it is necessary to identify discriminating proteins used for such diagnostic model and develop the gold standard screening method or the useful early diagnostic kit for the convenience for clinical use.
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- HBV:
-
Hepatitis B virus
- LC:
-
Liver cirrhosis
- HBVM:
-
HBV marker
- 2D-DIGE:
-
Two-dimensional difference in gel electrophoresis
- LC–MS/MS:
-
Liquid chromatography–tandem mass spectrometry
- SELDI-TOF-MS:
-
Surface enhanced laser desorption ionization-time of flight-mass spectrometry
- BPS:
-
Biomarker pattern software
- CV:
-
Coefficient of variance
- WCX2:
-
Weak cation exchange protein chip
- ANN:
-
Artificial neural networks
- CLD:
-
Chronic liver disease
References
Adam BL, Qu Y, Davis JW, Ward MD, Clements MA, Cazares LH, Semmes OJ, Schellhammer PF, Yasui Y, Feng Z, Wright GL Jr (2002) Serum protein fingerprinting coupled with a pattern-matching algorithm distinguishes prostate cancer from benign prostate hyperplasia and healthy men. Cancer Res 62:3609–3614
Bosch FX, Ribes J, Borras J (1999) Epidemiology of primary liver cancer. Semin Liver Dis 19:271–285
Bruix J, Sherman M, Llovet JM, Beaugrand M, Lencioni R, Burroughs AK, Christensen E, Pagliaro L, Colombo M, Rodes J; EASL Panel of Experts on HCC (2001) Clinical management of hepatocellular carcinoma. Conclusions of the Barcelona-2000 EASL conference. European Association for the Study of the Liver. J Hepatol 35(3):421–430
Brunello F, Marcarino C, Pasquero P, Gastaldi P, Gonella S, Martini S, Calcamuggi G, Emanuelli G (1993) The des-gamma-carboxyprothrombin for the diagnosis of hepatocellular carcinoma. Ital J Gastroenterol 25:9–12
Chan HL (2002) Changing scene in hepatitis B serology interpretation. Hosp Med 63:16–19
Erlinger S, Benhamou JP (1999) Cirrhosis: clinical aspects. In: Bircher J, Benhamou JP, McIntyre N, Rizetto M, Rodes J (eds) Oxford textbook of clinical hepatology, 2nd edn. Oxford University Press, Oxford, pp 629–641
Figueras J, Jaurrieta E, Valls C, Benasco C, Rafecas A, Xiol X, Fabregat J, Casanovas T, Torras J, Baliellas C, Ibanez L, Moreno P, Casais L (1997) Survival after liver transplantation in cirrhotic patients with and without hepatocellular carcinoma: a comparative study. Hepatology 25:1485–1489
Ganem D, Prince AM (2004) Hepatitis B virus infection—natural history and clinical consequences. N Engl J Med 350:1118–1129
Gao JD, Shao YF, Xu Y, Ming LH, Wu ZY, Liu GT, Wang XH, Gao WH, Sun YT, Feng XL, Liang LM, Zhang YH, Sun ZT (2005) Tight association of hepatocellular carcinoma with HBV infection in North China. Hepatobiliary Pancreat Dis Int 4:46–49
Grazi GL, Mazziotti A, Legnani C, Jovine E, Miniero R, Gallucci A, Palareti G, Gozzetti G (1995) The role of tumor markers in the diagnosis of hepatocellular carcinoma, with special reference to the des-gamma-carboxy prothrombin. Liver Transpl Surg 1:249–255
He QY, Lau GK, Zhou Y, Yuen ST, Lin MC, Kung HF, Chiu JF (2003) Serum biomarkers of hepatitis B virus infected liver inflammation: a proteomic study. Proteomics 3:666–674
Lai CL, Ratziu V, Yuen M, Poynard T (2003) Viral hepatitis B. Lancet 362: 2089–2094
Lee WM (1997) Hepatitis B virus infection. N Engl J Med 337:1733–1745
Lee IN, Chen CH, Sheu JC, Lee HS, Huang GT, Yu CY, Lu FJ, Chow LP (2005) Identification of human hepatocellular carcinoma-related biomarkers by two-dimensional difference gel electrophoresis and mass spectrometry. J Proteome Res 4:2062–2069
Li J, Zhang Z, Rosenzweig J, Wang YY, Chan DW (2002) Proteomics and bioinformatics approaches for identification of serum biomarkers to detect breast cancer. Clin Chem 48:1296–1304
Li C, Tan YX, Zhou H, Ding SJ, Li SJ, Ma DJ, Man XB, Hong Y, Zhang L, Li L, Xia QC, Wu JR, Wang HY, Zeng R (2005) Proteomic analysis of hepatitis B virus-associated hepatocellular carcinoma: identification of potential tumor markers. Proteomics 5:1125–1139
Lok AS, McMahon BJ (2001) Chronic hepatitis B. Hepatology 34:1225–1241
Marrero JA, Su GL, Wei W, Emick D, Conjeevaram HS, Fontana RJ, Lok AS (2003) Des-gamma carboxyprothrombin can differentiate hepatocellular carcinoma from nonmalignant chronic liver disease in American patients. Hepatology 37:1114–1121
Oh S, Afdhal NH (2001) Hepatic fibrosis: are any of the serum markers useful?. Curr Gastroenterol Rep 3:12–18
Paradis V, Degos F, Dargere D, Pham N, Belghiti J, Degott C, Janeau JL, Bezeaud A, Delforge D, Cubizolles M, Laurendeau I, Bedossa P (2005) Identification of a new marker of hepatocellular carcinoma by serum protein profiling of patients with chronic liver diseases. Hepatology 41:40–47
Parkin DM, Bray F, Ferlay J, Pisani P (2001) Estimating the world cancer burden: Globocan 2000. Int J Cancer 94:153–156
Petricoin EF, Liotta LA (2004) SELDI-TOF-based serum proteomic pattern diagnostics for early detection of cancer. Curr Opin Biotechnol 15:24–30
Pisani P, Parkin DM, Bray F, Ferlay J (1999) Erratum: estimates of the worldwide mortality from 25 cancers in 1990. Int J Cancer 83:18–29, 870–873
Poniachik J, Bernstein DE, Reddy KR, Jeffers LJ, Coelho-Little ME, Civantos F, Schiff ER (1996) The role of laparoscopy in the diagnosis of cirrhosis. Gastrointest Endosc 43:568–571
Poon TC, Yip TT, Chan AT, Yip C, Yip V, Mok TS, Lee CC, Leung TW, Ho SK, Johnson PJ (2003) Comprehensive proteomic profiling identifies serum proteomic signatures for detection of hepatocellular carcinoma and its subtypes. Clin Chem 49:752–760
Purcell RH (1993) The discovery of the hepatitis viruses. Gastroenterology 104:955–963
Seow TK, Liang RC, Leow CK, Chung MC (2001) Hepatocellular carcinoma: from bedside to proteomics. Proteomics 1:1249–1263
Schwegler EE, Cazares L, Steel LF, Adam BL, Johnson DA, Semmes OJ, Block TM, Marrero JA, Drake RR (2005) SELDI-TOF MS profiling of serum for detection of the progression of chronic hepatitis C to hepatocellular carcinoma. Hepatology 41:634–642
Sablon E, Shapiro F (2005) Advances in molecular diagnosis of HBV infection and drug resistance. Int J Med Sci 2:8–16
Tirumalai RS, Chan KC, Prieto DA, Issaq HJ, Conrads TP, Veenstra TD (2003) Characterization of the low molecular weight human serum proteome. Mol Cell Proteomics 2:1096–1103
Xu XQ, Leow CK, Lu X, Zhang X, Liu JS, Wong WH, Asperger A, Deininger S, Eastwood Leung HC (2004) Molecular classification of liver cirrhosis in a rat model by proteomics and bioinformatics. Proteomics 4:3235–3245
Yu Y, Chen S, Wang LS, Chen WL, Guo WJ, Yan H, Zhang WH, Peng CH, Zhang SD, Li HW, Chen GQ (2005) Prediction of pancreatic cancer by serum biomarkers using surface-enhanced laser desorption/ionization-based decision tree classification. Oncology 68:79–86
Zhang Z, Bast RC Jr, Yu Y, Li J, Sokoll LJ, Rai AJ, Rosenzweig JM, Cameron B, Wang YY, Meng XY, Berchuck A, Van Haaften-Day C, Hacker NF, de Bruijn HW, van der Zee AG, Jacobs IJ, Fung ET, Chan DW (2004) Three biomarkers identified from serum proteomic analysis for the detection of early stage ovarian cancer. Cancer Res 64:5882–5890
Zhu XD, Zhang WH, Li CL, Xu Y, Liang WJ, Tien P (2004) New serum biomarkers for detection of HBV-induced liver cirrhosis using SELDI protein chip technology. World J Gastroenterol 10:2327–2329
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Cui, J., Kang, X., Dai, Z. et al. Prediction of chronic hepatitis B, liver cirrhosis and hepatocellular carcinoma by SELDI-based serum decision tree classification. J Cancer Res Clin Oncol 133, 825–834 (2007). https://doi.org/10.1007/s00432-007-0224-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-007-0224-y