Abstract
The major aim of molecular cancer research is the development of new therapeutic strategies and compounds that target directly the genetic and biochemical causes of malignant transformation. Therapeutic genes, antibodies and their derivatives, but also small molecular weight compounds, have been used for this purpose. Small peptides might be able to complement these agents because of their ability to recognize specific protein domains and thus to interfere with enzymatic functions or protein-protein interactions. A variation of the yeast-two-hybrid procedure allows to select specifically binding peptides, so called peptide aptamers, from a peptide library of high complexity. This selection procedure can be adapted to any protein or protein fragment as a bait construct and selects for the intracellular interaction between the bait of choice and the peptide aptamer prey. Peptide aptamers thus selected can be cloned, provided with a protein transduction domain, expressed in bacteria and introduced into cancer cells. There they might disrupt protein-protein interactions crucial for cancer cell growth or survival. We introduce an example in which the Stat3 arm of EGF receptor signaling is selectively inhibited by a peptide aptamer and causes the growth arrest of EGF receptor-dependent tumor cells. The aptamer constructs can be supplemented with additional functional domains to enhance their inhibitory effects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Most impressive advances have been made in the past years concerning the molecular description of intracellular signaling pathways, their aberrations in tumor cells and their contributions to the phenotypic alterations encountered in cancer. It is an important goal of molecular cancer research to integrate these insights into therapeutic strategies and utilize essential differences in the genetic and biochemical properties of normal and cancer cells for therapeutic purposes (Liem et al. 2002). The targeting of drugs at such molecules might result in a high therapeutic index and minimize the undesirable side effects of cancer treatment on normal cells.
The genetic alterations found in cancer cells are manifold. Dominantly acting oncogenes may acquire their gain of function through gene amplifications and resulting protein overexpression or through mutations affecting their primary protein sequence and their inherent functional properties. Tumor suppressor genes exert their effects through the loss of function, which can be the consequence of gene deletions or disabling mutations in their amino acid sequence. The multitude of molecular mechanisms altering oncogene or tumor suppressor gene functions is reflected in the numerous strategies attempted to exploit these defects for therapeutic purposes.
Small molecules have been derived that target inappropriately expressed or activated protein kinases. The market introduction of Bcr/abl and EGFR tyrosine kinase inhibitors best exemplifies these developments (Traxler 2003; Dancey and Sausville 2003). These successful attempts to target specific molecular alterations of cancer cells was preceded by the development of a specific inhibitor of the HER2 receptor. This growth factor receptor is overexpressed in many human adenocarcinomas, and its action can be suppressed by the monoclonal antibody herceptin (Yarden and Sliwkowski 2001). The effects of monoclonal antibodies might be further enhanced by the combination with additional functional domains. Recombinant immunotoxins were initially conceived for the elimination of lymphocytes from allogenic bone marrow transplants (Vallera et al. 1983; Vallera et al. 1983; Filipovich et al. 1984) by conjugating potent toxins of plant or bacterial origin to monoclonal antibodies. Today, technical advances allow the construction of single-chain derivatives of monoclonal antibodies and fuse them to the enzymatically active domains of bacerial toxins. This results in recombinant proteins that target cells expressing defined surface antigens with very high specificity (Wels et al. 1992; Myklebust et al. 1994). ErbB2 and EGFR specific immunotoxins have been tested systemically in animal models and strongly reduced the growth of tumors and metastatic lesions (Schmidt et al. 1999; Azemar et al. 2000).
In addition to small molecules, antibodies and antibody-based recombinant constructs, other classes of molecules have proven to be valuable in preclinical experiments and in early clinical developments. Antisense oligonucleotides, therapeutic gene constructs and RNAi are being evaluated. In this report we will discuss a new approach to develop recombinant proteins for cancer therapy. This approach is based on the derivation of small peptides as ligands for predetermined proteins or protein domains. These peptide aptamers have the potential to inhibit intrinsic protein functions, such as enzymatic activities, or to interfere with protein-protein interactions, which might represent essential steps in signaling cascades.
What are peptide aptamers?
Peptide aptamers are short oligopeptides that are based on the notion that specific binders to any protein or protein domain can be obtained from a library of random peptides of sufficiently high complexity. They are comparable to antibodies with respect to the principle that oligopeptides can assume a conformation that allows the specific interaction and recognition of any target structure. Differently from antibodies, the target recognition domain is composed of a only one single oligopeptide of about 20 amino acids.
The prerequisites for the detection and isolation of a peptide aptamer that specifically recognizes a target structure of choice are the derivation of a peptide library of very high complexity and an appropriate selection system.
A very large number of different peptides can be obtained through the construction of combinatorial libraries. The synthesis of a DNA sequence that encodes 20 amino acids is performed in a way so that each of the 20 positions can be occupied by any of the 20 amino acids resulting in a complexity of several billion different peptide sequences. The next step encompasses the molecular cloning of these synthetic DNA sequences into a vector in which the peptide sequences become embedded in a scaffold protein and that allows the expression of the peptide sequences as a part of this scaffold protein in yeast cells. The scaffold protein can be selected as to display the peptide sequence in a conformationally constrained fashion.
A variation of the yeast-two-hybrid selection can now be employed to find appropriate peptides with specific binding ability to predetermined targets. A yeast bait line is constructed expressing a gene construct in which the target protein or target protein domain is fused to a GAL4 DNA binding domain. This yeast line is transfected with a high complexity library that expresses the divergent peptide aptamers in the scaffold protein fused to the GAL4 transactivation domain. Appropriate selection marker genes, induced through the action of the reconstituted binary GAL4 transcription factor, allow the selection of individual yeast cell clones. In these cells the interaction between the GAL4 DBD bait protein and the GAL4 TAD prey protein is established by a specific peptide aptamer (Colas 2000).
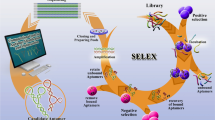
The aptamer selection system is depicted in Fig. 1. It shows the interaction between the protein of interest and a peptide aptamer. The aptamer is inserted into the active loop of the scaffold protein thioredoxin A. Different scaffold proteins for the display of conformationally constrained peptides have been investigated. The use of the green fluorescent protein (GFP) as a scaffold permits the analysis of library diversity and expression levels in cells and the enrichment of the libraries for sequences with predetermined characteristics, such as high expression of correctly folded protein (Abedi et al. 1998). Other proteins that have been described as scaffolds suitable for the presentation of peptides (Klevenz et al. 2002) include a catalytically inactive derivative of Staphylococcus nuclease (Norman et al. 1999), the protease inhibitor eglin C (Cohen et al. 1998), the Streptomyces tendea α-amylase inhibitor tendamistat (McConnell and Hoess 1995) and the cellular transcription factor Sp1 (Cheng et al. 1997). In these experiments peptide aptamers have been selected as ligands for many different proteins.
Screening of peptide in a yeast-two-hybrid screening system. Upon interaction between bait (Gal4 DNA binding domain fused with target protein) and prey (Gal4 transactivation domain fused with thioredoxin peptide library) a complete Gal4 transcription factor is restored. This allows expression of selection marker genes, which guaranties growth under selective conditions
Functional properties of peptide aptamers
There are only a few examples of small molecular weight compounds able to disrupt specific protein-protein interactions (Blaskovitch et al. 2003, Ren et al. 2003). This is a common property of peptide aptamers, and they can be used in a dominant fashion to influence the function of target proteins and cause cellular phenotypes (Hoppe-Seyler and Butz 2000). Proteins of different cellular localizations and functions have been targeted by peptide aptamers, and inhibitory actions have been observed. An aptamer selected as a specific ligand for the kinase Cdk2, an enzyme important for the progression of the cell cycle, was shown to be able to inhibit Cdk2 enzymatic activity (Colas, Cohen et al. 1996) and to block cell cycle progression (Cohen et al. 1998). A peptide aptamer specific for the transcription factor E2F, also centrally involved in the regulation of the cell cycle, blocked its DNA-binding capacity and caused a G1 cell cycle arrest (Fabbrizio et al. 1999).
Viral proteins have been used as targets for aptamer interference. A peptide aptamer selected for binding to the herpes simplex virus type 16 protein E6 resulted in the induction of apoptosis of HPV-positive cells (Butz et al. 2000). The same group isolated an aptamer that specifically inhibits hepatitis B virus capsid formation and replication by binding to the hepatitis virus core protein (Butz et al. 2001). Aptamers have been selected that bind specifically to cellular oncogenes. An aptamer has been studied that distinguishes between allelic forms of H-Ras and can inhibit the EGF-induced interaction of H-Ras with C-Raf1 (Xu and Luo 2002). The aptamer TRIPα can specifically block the activity of the rho-guanine nucleotide exchange factor leading to reversion of the neurite retraction phenotype in PC12 cells (Schmidt et al. 2002).
We have selected peptide aptamers that specifically recognize the intracellular domain of growth factor receptors, e.g., the EGFR, or transcription factors such as Stat proteins (Levy and Darnell 2002). The peptide aptamer KDI1, specific for the intracellular domain of the epidermal growth factor receptor (EGFR), has most interesting properties. Upon introduction into cells, it caused a showed strong reduction in EGF-induced cellular proliferation and soft agar colony formation. The aptamer did not summarily block the EGF receptor tyrosine kinase activity, but selectively interfered with the EGF-induced phosphorylation of the tyrosine residues 845, 1068 and 1148, as well as the phosphorylation of tyrosine 317 of p46 Shc. In addition it prevented the recruitment of c-Src to the EGF receptor, the activation of Stat3 by phosphorylation at tyrosine 705 and Stat3 dependent transactivation (Buerger et al. 2003). Since other EGFR-mediated signaling pathways, e.g., the MAP kinase pathway and the PI3 kinase pathway, were unaffected, we assume that the elimination of Stat3 signaling is sufficient to block EGF induced proliferation (Fig. 2). Transduction of a short synthetic peptide aptamer sequence not embedded into the scaffold protein resulted in the same impairment of EGF-induced Stat3 activation.
Binding of peptide aptamer KDI1 inhibits c-Src-mediated signaling from the EGF receptor. A Upon EGF binding the receptor dimerizes, which activates the intrinsic kinase activity. In addition c-Src is recruited to the receptor, phosphorylating tyrosine 845, which enhances the kinase activity. C-Src also phosphorylates pre-associated Stat3 molecules, which dimerize and translocate into the nucleus, where they induce the transcription of target genes. The intrinsic kinase activity of the receptor phosphorylates tyrosine residues in the c-terminal part of the receptor that serves as a binding site for signaling molecules that activate the MAPK and Akt pathway. B The peptide aptamer binds to a region within the kinase domain of the EGFR, so that c-Src can no longer be recruited to the activated receptor. This prevents enhancement of the intrinsic receptor kinase activity through phosphorylation of tyrosine 845 and activation of Stat3
These data show that peptide aptamers can have most promising properties. They can be selected for many different targets by a simple screening procedure and interfere with enzymatic activity, protein-protein interactions and selected aspects of proliferative signaling (Hoppe-Seyler and Butz 2000).
Cellular delivery of peptide aptamers
Peptide aptamers are able to specifically inhibit the functions of cellular proteins and therefore have the potential to be used as therapeutic agents. A prerequisite for their intracellular action is the delivery into cells. Several methods can be employed to achieve this aim (Hoppe-Seyler and Butz 2000).
It is possible to integrate the peptide aptamer sequence into a gene construct and deliver it into cells through transfection or viral infection. This approach will result in the intracellular transcription and translation of the peptide aptamer sequence and is most valuable for experiments in which the interaction specificity and the functional properties of aptamer sequences are studied in cultured cells. It is limited, however, in its therapeutic value by the inefficiency of delivery of gene constructs in vivo, problems hampering many other gene therapy strategies.
To circumvent the introduction of genetic material into cells, it is conceivable to synthesize peptide aptamers or produce them as recombinant proteins in bacteria and introduce them as purified proteins into cells. Although this approach is more "drug like," there are obstacles to be overcome. These include the protease sensitivity of peptides and proteins, the inability of these molecules to cross cellular membranes and their potential immunogenicity. However, progress is being made and delivery systems, e.g., the encapsulation of proteins into polymers or lipids that protect therapeutic proteins from degradation and allow their controlled release, have been developed (Langer 1998). Peptides and proteins can be stabilized by chemical modifications, e.g., the modification of chemical bonds and alterations of amino acid side chains.
Because the cell membrane is impermeable for most peptides and proteins, the introduction of therapeutic proteins into tissues or organs proved to be rather difficult and is limited by the size and biochemical properties of the protein to be delivered. Until recently, it was not possible to introduce proteins into cells unless they were bioactive peptides of small size and highly lipophilic. In 1988, an exception to these limitations was discovered. The HIV TAT protein was found to be able to permeate the cell membrane from outside the cell (Green and Loewenstein 1988; Frankel and Pabo 1988). This functional property of the HIV TAT protein was delimited to a short stretch of amino acids and shown to be transferable to heterologous protein through gene fusion. In 1994 Fawell et al. introduced heterologous proteins into cells that were linked to a 38 amino acid fragment of the HIV TAT protein (Fawell et al. 1994). Subsequently, other protein transduction domains (Fig. 3) have been identified. Examples are the protein transduction domains present in the antennapedia protein of Drosophila (Joliot et al. 1991) and the HSV VP22 protein of the herpes simplex virus (Elliott and O'Hare 1997; Prochiantz 2000).
Amino acid sequences of protein transduction domains (PTD) so far characterized. All PTDs contain numerous basic amino acids like arginine and lysine, which mediate contact with the cell membrane. The minimal TAT transduction domain encompasses the basic residues 47–55, whereas residues 267–to 300 mediate transduction via the VP22 protein. In the antennapedia protein (Antp), the third α-helix (aa 43–58) is responsible for transduction. Modified from (Schwarze and Dowdy 2000)
All protein transduction domains (PTD) are characterized by numerous basic amino acids, arginines and lysines (Fig. 3). These amino acids are probably responsible for the interaction with the membrane lipid bilayer. Attempts have been made to derive synthetic sequences with optimised protein transduction characteristics. Wender et al. showed that a peptide consisting of nine L-arginines is about 20 times more efficient in the transduction of linked proteins or peptides than the naturally occurring HIV TAT domain. A D-arginine oligomer transduces proteins even 100 times better into Jurkat cells.
Mechanistic considerations have been entertained, and it was proposed that the guanidinium part of arginine is more important for the function than the charge or the backbone structure (Wender et al. 2000). Mai et al. could show that the efficiency of transduction is very much dependent on the cell line utilized and demonstrated at the same time that poly-lysine sequences are more efficient transduction domains than the TAT or poly-arginine sequences (Mai et al. 2002). The mechanism that promotes protein transduction via PTD into cells has not been totally clarified yet. Transduction can occur at 37°C and at 4°C and is non-saturable. This led to the suggestion that protein transduction does not occur via classical receptor-, transporter- or endocytosis-mediated mechanisms (Derossi et al. 1996). Tyagi et al. have proposed that heparin sulfate proteoglycanes on the cell surface might serve as receptors for transduced proteins (Tyagi et al. 2001). Cells that do not express such molecules or cells treated with the appropriate glycosamine glycane lyases are not able to take up TAT proteins or TAT protein fusions.
PTD have been employed to transport various coupled macromolecules into cells (Rouselle 2000). The cargo can consist of peptides, proteins or even of DNA. Size does seem to limit the introduction of these fusions or conjugates into cells. A small protein of 20 kDa has been transduced into cells and was found to be biologically active immediately following its transfer. A protein of 120 kDa has been found to require a period of about 5 min following introduction before intracellular biological activity could be shown (Schwarze et al. 1999). A fusion protein of the HIV TAT PTD and β-galactosidase was expressed in bacteria and purified in a partially denatured state. The protein was present intracellularly 15 min after addition to the cells. Enzymatic activity, however, was only detected after a period of 2 h. This has led to the suggestion that transduced proteins might require refolding and the assumption of a proper conformation before they become active. This might require cellular chaperones (Schwarze et al. 1999).
Transduced proteins have been shown to be able to affect important cellular signaling pathways. A dominant-negative version of IκB (TAT-IκB46–317) has been introduced into osteoclasts as a TAT-PTD fusion protein. This protein was able to inhibit osteoclast differentiation (Abu-Amer et al. 2001). The cell cycle inhibitor p27 kip has been introduced into cells and shown to induce migration (Nagahara et al. 1998). The EGFR-specific peptide aptamer KDI1 described above was produced as a membrane traversing protein by provision of a poly-arginine sequence. When this fusion protein was added to A431 human tumor cells, it inhibited Stat3 activation and cellular proliferation, very much like a gene construct encoding the peptide aptamer transfected into these cells.
The PTD-β-galactosidase protein has also been evaluated in vivo. Mice were injected intraperitoneally with the protein and the presence of the transduced protein was assayed in different tissues. It was found to be present in liver, kidney, lung, heart and spleen. Injected protein even crossed the blood-brain barrier and showed enzymatic activity in brain tissue (Schwarze et al. 1999). Other proteins introduced into cells via protein transduction domains exhibit therapeutic effects in vivo. A TAT-Bcl-xL fusion protein has been introduced into primary cultures of neurones and into mice. Mice with cerebral ischemia were intraperitoneally injected with the fusion protein. A decrease in the extent of apoptosis in brain cells mediated by the recombinant Bcl-xL protein was shown when compared to untreated animals (Cao et al. 2002). Injection of an antimicrobial peptide fused to a PTD into solid tumors significantly induced tumor apoptosis and a reduction of tumour volume (Mai et al. 2001). These examples show that the method of protein transduction is well suited to study signal transduction pathways through specific inhibitors, but might also serve as a therapeutic tool (Ford et al. 2001; Kau and Silver 2003).
Arming peptide aptamers with additional functions
The distinctive functional property of peptide aptamers is their ability to bind to a protein target. Since the bait construct ussually comprises about 50 amino acids, the ligand property of the aptamer does not assure that it also interferes with functional aspects of the intact target protein. It was shown that only a fraction of the selected aptamers exhibited inhibitory functions. Many protein are organized into functional domains that work with surprising autonomy. This observation can be used for the construction of recombinant proteins with novel capabilities. The combination of functional domains that naturally occur in totally unrelated proteins offers most interesting perspectives.
Specific ligands for intracellular proteins have been exploited. A single chain antibody derivative specific for the erbB2 growth factor receptor has been linked with an endoplasmatic reticulum retention signal. The expression of this recombinant protein in cells caused the retention of newly synthesized erbB2 receptor in the ER and prevented its localisation in the periplasmic membrane (Beerli et al. 1994).
We have exploited the specific binding function of peptide aptamers for the targeted degradation of individual intracellular proteins. Many crucial signaling pathways depend upon the presence of key proteins, and the selective degradation of individual components can drastically alter the cellular phenotype. For this purpose we have made use of protein domains that specifically interact with the protein degradation components of the cell. The SOCS box domain of the SOCS proteins (suppressors of cytokine signaling) promotes the assembly of an E3 ligase complex. The SH2 domain interacts with tyrosine phosphorylated signaling molecules and marks them as substrates for ubiquitination and degradation by the 26 S proteasome (Ungureanu et al. 2002; Zhang et al. 1999). Therefore, the SOCS box domain can be used as a functional domain to target proteins bound to the same molecule for degradation. We could show that the replacement of the SH2 domain of the SOCS protein with a peptide aptamer changes the substrate specificity and causes the degradation of the protein for which the peptide aptamer serves as a ligand. EGFR or Stat3 specific peptide aptamers have been successfully employed (Buerger, Nagel-Wolfrum and Groner, unpublished results).
Colas et al. took a similar approach to manipulate the function of target proteins via peptide aptamers comprising functional domains. Aptamers fused to the catalytic domain of a ubiquitin ligase (HECT domain) specifically transferred ubiquitin moieties to the target protein Cdk2 in vivo, while other aptamers carrying a nuclear localization sequence transported their targets into the nucleus (Colas et al. 2000). These experiments indicate that recombinant fusion proteins containing aptamer recognition moieties will become useful in many respects. The possibility to inhibit enzymatic functions, to interfere with protein-protein interactions, to change the subcellular localization of proteins and to target individual proteins for degradation will become most useful for functional studies of genes and for therapeutic purposes.
Finally, peptide aptamers open new possibilities for drug design. The resolution of the structure of a peptide aptamer bound to its target protein might suggest new lead structures that could inspire synthetic organic chemists to come up with non-peptidic analogues (Berezov et al. 2002).
Abbreviations
- PTD:
-
Protein transduction domain
- Trx:
-
Thioredoxin
- EGFR:
-
Epidermal growth factor receptor
- STAT:
-
Signal transducer and activator of transcription
References
Abedi MR, Caponigro G, Kamb A (1998) Green fluorescent protein as a scaffold for intracellular presentation of peptides. Nucleic Acids Res 26:623–630
Abu-Amer Y, Dowdy SF, Ross FP, Clohisy JC, Teitelbaum SL (2001) Tat fusion proteins containing tyrosine 42-deleted ikappa balpha arrest osteoclastogenesis. J Biol Chem 276:30499–30503
Azemar M, Schmidt M, Arlt F, Kennel P, Brandt B, Papadimitriou A, Groner B, Wels W (2000) Recombinant antibody toxins specific for ErbB2 and EGF receptor inhibit the in vitro growth of human head and neck cancer cells and cause rapid tumor regression in vivo. Int J Cancer 86:269–275
Beerli RR, Wels W, Hynes NE (1994) Intracellular expression of single chain antibodies reverts ErbB-2 transformation. J Biol Chem 269:23931–23936
Berezov A, Chen J, Liu Q, Zhang HT, Greene MI, Murali R (2002) Disabling receptor ensembles with rationally designed interface peptidomimetics. J Biol Chem 277:28330–28339
Blaskovich MA, Sun A, Cantor J, Turkson R, Jove, Sebti SM (2003) Discovery of JSI-124 (cucurbitacin I), a selective Janus kinase/signal transducer and activator of transcription 3 signaling pathway inhibitor with potent antitumor activity against human and murine cancer cells in mice. Cancer Res 63:1270–1279
Buerger C, Nagel-Wolfrum K, Kunz C, Wittig I, Butz K, Hoppe-Seyler F, Groner B (2003) Sequence specific peptide aptamers, interacting with the intracellular domain of the epidermal growth factor receptor, interfere with STAT3 activation and inhibit the growth of tumor cells. J Biol Chem (in press)
Butz K, Denk C, Ullmann A, Scheffner M, Hoppe-Seyler F (2000) Induction of apoptosis in human papilloma virus positive cancer cells by peptide aptamers targeting the viral E6 oncoprotein [In Process Citation]. Proc Natl Acad Sci USA 97:6693–6697
Butz K, Denk C, Fitscher B, Crnkovic-Mertens I, Ullmann A, Schroder CH, Hoppe-Seyler F (2001) Peptide aptamers targeting the hepatitis B virus core protein: a new class of molecules with antiviral activity. Oncogene 20:6579–6586
Cao G, Pei W, Ge H, Liang Q, Luo Y, Sharp FR, Lu A, Ran R, Graham SH, Chen J (2002) In vivo delivery of a Bcl-xL fusion protein containing the TAT protein transduction domain protects against ischemic brain injury and neuronal apoptosis. J Neurosci 22:5423–5431
Cheng X, Boyer JL, Juliano RL (1997) Selection of peptides that functionally replace a zinc finger in the Sp1 transcription factor by using a yeast combinatorial library. Proc Natl Acad Sci USA 94:14120–14125
Cohen BA, Colas P, Brent R (1998) An artificial cell-cycle inhibitor isolated from a combinatorial library. Proc Natl Acad Sci USA 95:14272–14277
Colas P (2000) Combinatorial protein reagents to manipulate protein function. Curr Opin Chem Biol 4:54–59
Colas P, Cohen B, Jessen T, Grishina I, McCoy J, Brent R (1996) Genetic selection of peptide aptamers that recognize and inhibit cyclin- dependent kinase 2. Nature 380:548–550
Colas P, Cohen B, Ferrigno PK, Silver PA, Brent R (2000) Targeted modification and transportation of cellular proteins. Proc Natl Acad Sci USA 97:13720–13725
Dancey J, Sausville EA (2003) Issues and progress with protein kinase inhibitors for cancer treatment. Nat Rev Drug Discov 2:296–313
Derossi D, Calvet S, Trembleau A, Brunissen A, Chassaing G, Prochiantz A (1996) Cell internalization of the third helix of the Antennapedia homeodomain is receptor-independent. J Biol Chem 271:18188–18193
Elliott G, O'Hare P (1997) Intercellular trafficking and protein delivery by a herpes virus structural protein. Cell 88:223–233
Fabbrizio E, Le Cam L, Polanowska J, Kaczorek M, Lamb N, Brent R, Sardet C (1999) Inhibition of mammalian cell proliferation by genetically selected peptide aptamers that functionally antagonize E2F activity. Oncogene 18:4357–4363
Fawell S, Seery J, Daikh Y, Moore C, Chen LL, Pepinsky B, Barsoum J (1994) Tat-mediated delivery of heterologous proteins into cells. Proc Natl Acad Sci USA 91:664–668
Filipovich AH, Vallera DA, Youle RJ, Quinones RR, Neville DM Jr, Kersey JH (1984) Ex-vivo treatment of donor bone marrow with anti-T-cell immunotoxins for prevention of graft-versus-host disease. Lancet 1:469–472
Ford KG, Suoberbielle BE, Darling D, Farzaneh F (2001) Protein transduction: an alternative to genetic intervention? Gene Ther 8:1–4
Frankel AD, Pabo CO (1988) Cellular uptake of the tat protein from human immunodeficiency virus. Cell 55:1189–1193
Green M, Loewenstein PM (1988) Autonomous functional domains of chemically synthesized human immunodeficiency virus tat trans-activator protein. Cell 55:1179–1188
Hoppe-Seyler F, Butz K (2000) Peptide aptamers: powerful new tools for molecular medicine. J Mol Med 78:426–430
Joliot A, Pernelle C, Deagostini-Bazin H, Prochiantz A (1991) Antennapedia homeobox peptide regulates neural morphogenesis. Proc Natl Acad Sci USA 88:1864–1868
Kau TR, Silver PA (2003) Nuclear transport as a target for cell growth. Drug Discov Today 8:78–85
Klevenz B, Butz K, Hoppe-Seyler F (2002) Peptide aptamers: exchange of the thioredoxin-A scaffold by alternative platform proteins and its influence on target protein binding Cell Mol Life Sci 59:1993–1998
Langer R (1998) Drug delivery and targeting. Nature 392 [Suppl 6679]:5–10
Liem AA, Chamberlain MP, Wolf CR, Thompson AM (2002) The role of signal transduction in cancer treatment and drug resistance. Eur J Surgical Oncol 28:679–684
Levy DE, Darnell JE (2002) Stats: transcriptional control and biological impact. Nat Rev Cell Biol 3:651–662
Mai JC, Mi Z, Kim SH, Ng B, Robbins PD (2001) A proapoptotic peptide for the treatment of solid tumors. Cancer Res 61:7709–7712
Mai JC, Shen H, Watkins SC, Cheng T, Robbins PD (2002) Efficiency of protein transduction is cell type-dependent and is enhanced by dextran sulfate. J Biol Chem 277:30208–30218
McConnell SJ, Hoess RH (1995) Tendamistat as a scaffold for conformationally constrained phage peptide libraries. J Mol Biol 250:460–470
Myklebust AT, Godal A, Juell S, Pharo A, Fodstad O (1994) Comparison of two antibody-based methods for elimination of breast cancer cells from human bone marrow. Cancer Res 54:209–214
Nagahara H et al (1998) Transduction of full length TAT fusion proteins into mammalian cells: TAT-p27 kip1 induces cell migration. Nature Med 4:1449–1452
Norman TC, Smith DL, Sorger PK, Drees BL, O'Rourke SM, Hughes TR, Roberts CJ, Friend SH, Fields S, Murray AW (1999) Genetic selection of peptide inhibitors of biological pathways. Science 285:591–595
Prochiantz A (2000) Messenger proteins: homeoproteins, TAT and others. Curr Opin Cell Biol 12:400–406
Ren Z, Cabell LA, Schaefer TS, McMurray JS (2003) Identification of a high affinity phosphopeptide inhibitor of Stat3. Bioorg Med Chem Lett 13:633–636
Rouselle C, Clair P, Lefauconnier JM, Kaczorek M, Scherrmann JM, Temsamani J (2000) New advances in the transport of doxorubicin through the blood brain barrier by a peptide vector mediated strategy. Mol Pharmacol 57:679–686
Schmidt M, Maurer-Gebhard M, Groner B, Kohler G, Brochmann-Santos G, Wels W (1999) Suppression of metastasis formation by a recombinant single chain antibody-toxin targeted to full-length and oncogenic variant EGF receptors. Oncogene 18:1711–1721
Schmidt S, Diriong S, Mery J, Fabbrizio E, Debant A (2002) Identification of the first Rho-GEF inhibitor, TRIPalpha, which targets the RhoA-specific GEF domain of Trio. FEBS Lett 523:35–42
Schwarze SR, Dowdy SF (2000) In vivo protein transduction: intracellular delivery of biologically active proteins, compounds and DNA. Trends Pharmacol Sci 21:45–48
Schwarze SR, Ho A, Vocero-Akbani A, Dowdy SF (1999) In vivo protein transduction: delivery of a biologically active protein into the mouse [see comments]. Science 285:1569–1572
Traxler P (2003) Tyrosine kinases as targets in cancer therapy-successes and failures. Expert Opinion Ther Targets 7:215–234
Tyagi M, Rusnati M, Presta M, Giacca M (2001) Internalization of HIV-1 tat requires cell surface heparan sulfate proteoglycans. J Biol Chem 276:3254–3261
Ungureanu D, Saharinen P, Junttila I, Hilton DJ, Silvennoinen O (2002) Regulation of Jak2 through the ubiquitin-proteasome pathway involves phosphorylation of Jak2 on Y1007 and interaction with SOCS-1. Mol Cell Biol 22:3316–3326
Vallera DA, Ash RC, Zanjani ED, Kersey JH, LeBien TW, Beverley PC, Neville DM Jr, Youle RJ (1983) Anti-T cell reagents for human bone marrow transplantation: ricin linked to three monoclonal antibodies. Science 222:512–515
Vallera DA, Youle RJ, Neville DM Jr, Soderling CC, Kersey JH (1983) Monoclonal antibody toxin conjugates for experimental graft-versus-host disease prophylaxis. Reagents selectively reactive with T cells and not murine stem cells. Transplantation 36:73–80
Wels W, Harwerth IM, Mueller M, Groner B, Hynes NE (1992) Selective inhibition of tumor cell growth by a recombinant single-chain antibody-toxin specific for the erbB-2 receptor. Cancer Res 52:6310–6317
Wender PA, Mitchell DJ, Pattabiraman K, Pelkey ET, Steinman L, JB Rothbard (2000) The design, synthesis, and evaluation of molecules that enable or enhance cellular uptake: peptoid molecular transporters. Proc Natl Acad Sci USA 97:13003–13008
Xu CW, Luo Z (2002) Inactivation of Ras function by allele-specific peptide aptamers. Oncogene 21:5753–5757
Yarden Y, MX Sliwkowski (2001) Untangling the ErbB signalling network. Nat Rev Mol Cell Biol 2:127–137
Zhang JG, Farley A, Nicholson SE, Willson TA, Zugaro LM, Simpson RJ, Moritz RL, Cary D, Richardson R, Hausmann G, Kile BJ, Kent SB, Alexander WS, Metcalf D, Hilton DJ, Nicola NA, Baca M (1999) The conserved SOCS box motif in suppressors of cytokine signaling binds to elongins B and C and may couple bound proteins to proteasomal degradation. Proc Natl Acad Sci USA 96:2071–2076
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Buerger, C., Groner, B. Bifunctional recombinant proteins in cancer therapy: cell penetrating peptide aptamers as inhibitors of growth factor signaling. J Cancer Res Clin Oncol 129, 669–675 (2003). https://doi.org/10.1007/s00432-003-0489-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-003-0489-8