Abstract
Herpes simplex virus 1 (HSV-1) is the archetypal member of the alphaherpesvirus with a large genome encoding over 80 viral proteins, many of which are involved in virus–host interactions and show immune modulatory capabilities. In this study, we demonstrated that the HSV-1 UL42 protein, a DNA polymerase processivity factor, was a novel antagonism of the canonical NF-κB signaling pathway. UL42 was shown to significantly suppress TNF-α mediated NF-κB activation. Co-immunoprecipitation experiment revealed that UL42 bound to the NF-κB subunits p65 and p50. Fluorescence microscopy demonstrated that UL42 abolished nuclear translocation of p65 and p50 upon TNF-α-stimulation. But the inhibiting capacity of UL42 2R/2A (R279A, R280A) and UL42 3R/3A (R113A, R279A and R280A) mutants were less than wild type UL42. Also UL42 bound to the Rel homology domain of the NF-κB subunit p65 and p50. Notably, the N-terminal of UL42 was sufficient to interact with p65 and p50 and abolished NF-κB reporter gene activity. Thus, it was first time we demonstrated that HSV-1 UL42 appeared to prevent NF-κB-dependent gene expression by retaining p65 and p50 in the cytoplasm, and UL42-dependent transcriptional activation were inherently coupled to promote HSV-1 lytic replication, which also may contribute to immune evasion and pathogenesis of HSV-1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The innate immune responses to viruses involve activation of pattern recognition receptors (PRRs) [1–3] and transcriptional induction of type I interferons (IFNs) and proinflammatory cytokines [4]. The transcription factor NF-κB plays a pivotal role in many cellular events, such as innate and adaptive immunity, inflammation [5–8].
Tumor necrosis factor-alpha (TNF-α) is a multifunctional cytokine that plays a key role in innate immunity by inducing the expression of a variety of genes that are involved in inflammatory responses. TNF-α binds to its receptor TNF-R1 resulting in recruitment of the adaptor protein TNF receptor death domain (TRADD), and then TRADD recruits TNFR-associated factor 2 (TRAF2), and receptor-interacting protein 1 (RIP1) to the receptor complex, TRAF2 mediates K63-linked poly-ubiquitination of RIP1. Ubiquitinated RIP1 further recruits TGF-β-activated kinase 1 (TAK1) and subsequently activates the IκB kinase (IKK) complex, leading to the phosphorylation and degradation of IκBα, at last, activation of NF-κB [9].
The mammalian NF-κB family comprises five members: p65/RelA, RelB, p50/NF-κB1, p52/NF-κB2 and c-Rel. All family members share a structurally conserved N-terminal region, named Rel homology domain (RHD), which is critical for protein dimerization, DNA binding, interaction with IκB (inhibitor of NF-κB) and nuclear translocation [10, 11]. Rel proteins (p65/RelA, RelB, and c-Rel) contain a C-terminal transactivation domain, which is lacking in p50 and p52. The predominant form of NF-κB is a heterodimer of p65 and p50 subunits [12, 13]. There exist two major NF-κB-activating signaling pathways: the canonical NF-κB pathway and the noncanonical NF-κB pathway.
HSV-1 is a large DNA virus known to encode several gene products that enable viral evasion of the host innate immune response [14, 15]. Several studies have shown that HSV-1 encodes proteins to disturb NF-κB pathway. ICP0, an immediate-early gene product of HSV-1, antagonizes TLR-driven inflammatory cytokine response following viral infection [16, 17]. HSV-encoded late gene product γ134.5 protein inhibits activation of NF-κB in CD8+ Dendritic Cells (DCs) [18]. Vhs, a tegument protein, which is carried in the virion and delivered into DCs blocks the early replication-independent activation of NF-κB during HSV-1 infection [19].
HSV-1 UL42 is one of seven proteins that are required for the replication of the HSV-1 genome is found in a 1:1 complex with the HSV-1 DNA polymerase UL30, and UL42 increases the processivity of the polymerase along the DNA template during replication [20, 21]. Previous studies identified four conserved arginines (113, 182, 279, and 280) on a basic surface of UL42 and postulate that DNA binding by UL42 is mediated by this surface, and substitution of any of these arginines with alanines results in reduced DNA binding ability of UL42 and also impaired for viral replication and DNA synthesis [22–24].
We hypothesized that HSV-1 might evolve strategies to interfere with TNF-α-induced NF-κB activation to evade the immune responses of the host. To test this hypothesis, we examined whether TNF-α-induced NF-κB activation could be modulated by HSV-1. In the current study, we demonstrate the ability of HSV-1 UL42 protein to inhibit TNF-α-mediated NF-κB activation. Further study indicates that p65 and p50 highly likely serves as the target for UL42 protein.
Materials and methods
Cells, antibodies and cytokines
HEK 293T cells and Vero cells were grown in Dulbecco’s modified Eagle medium (DMEM) (Gibco-BRL) supplemented with 10 % fetal bovine serum (FBS) and 100 U/ml of penicillin and streptomycin. HeLa cells were maintained in Eagle’s minimum essential medium (MEM) (Gibco-BRL) supplemented with 10 % FBS. The wild-type (WT) HSV-1 F strain was propagated in Vero cells and titrated as described previously [35].
Mouse anti-Myc (isotype IgG1), anti-Flag (isotype IgG2b) and anti-hemagglutinin (HA) (isotype IgG2b) monoclonal antibodies (mAbs) were purchased from ABmart (Shanghai, China). Mouse monoclonal IgG1 and IgG2b isotype control antibodies were purchased from eBioscience Inc. (San Diego, CA). Rabbit anti-p65 polyclonal antibody (pAb) and Rabbit anti-p50 polyclonal antibody (pAb) were purchased from Proteintech (Wuhan, China). Rabbit anti-UL42 polyclonal antibody (pAb) was purchased from GL Biochem, and mouse anti-β-actin mAb was purchased from Santa Cruz Biotechnology (Santa Cruz, CA). Recombinant human TNF-α was purchased from Biovision (San Francisco, CA).
Plasmid construction
All enzymes, except for T4 DNA ligase (New England BioLabs, MA), used for cloning procedures were purchased from Takara (Dalian, China). To construct UL42-HA, the ORF of UL42 was amplified from plasmid UL42-EYFP as previously described [25] and cloned into the BamHI and EcoRI sites of the pCMV-HA vector (Beyotime, Shanghai, China). Commercial reporter plasmids include NF-κB-Luc (Stratagene, La Jolla, CA) and pRL-TK plasmid (Promega). Gift plasmids include the following: pcDNA3-NIK [26], pFlag-p65 [27], pFlag-p50 [28], pFlag-IKKβ [29], pHA-IKKα [30], pGL3-pIL-6-Luc and pXP2-pIL-8-Luc [31].
Transfection and dual-luciferase reporter (DLR) assay
HEK 293T cells were co-transfected with 1,000 ng expression plasmid, 500 ng NF-κB-Luc reporter plasmid and 50 ng of pRL-TK Renilla luciferase reporter plasmid to normalize transfection efficiency, as indicated by standard calcium phosphate precipitation [32, 33]. At 24 h post-transfection, cells were mock treated or treated with recombinant human TNF-α (10 ng/ml) for 6 h, and then luciferase assays were performed as previously described [34] with a luciferase assay kit (Promega, Madison, WI).
Western blot analysis
Western blot (WB) analysis was performed as previously described [35]. Briefly, whole-cell extracts were subjected to 10 % SDS-PAGE and transferred to PVDF membranes, followed by blocking with 5 % nonfat milk in Tris-buffered saline-Tween (TBST) and probed with the indicated primary antibodies at 37 °C for 2 h. After washing with TBST, the membrane was incubated with alkaline phosphatase (AP)-conjugated goat anti-rabbit IgG or goat anti-mouse IgG. Protein bands specific to the antibody were developed by 5-bromo-4-chloro-3-indolylphosphate (BCIP)-nitroblue tetrazolium (NBT) and terminated by distilled water.
Co-immunoprecipitation assay
Co-immunoprecipitation (co-IP) assays were performed as previously described [35]. Briefly, HEK 293T cells (~5 × 106) were co-transfected with 5 μg of each of the indicated expression plasmids carrying Flag, Myc or HA tags. Transfected cells were harvested at 36 h post-transfection and lysed on ice with 700 μl of lysis buffer. For each immunoprecipitation (IP), a 0.5 ml aliquot of lysate was incubated with 0.5 μg of the anti-HA mAb, anti-Myc mAb or anti-Flag mAb or nonspecific mouse monoclonal antibody (IgG1 isotype matched with anti-Myc mAb; IgG2b isotype matched with anti-HA and anti-Flag mAbs) and 30 μl of a 1:1 slurry of Protein A/G Plus-agarose (Santa Cruz, CA) for at least 4 h or overnight at 4 °C. The beads were washed four times with 1 ml of lysis buffer containing 500 mM NaCl and then subjected to WB analysis. All co-IP assays were repeated at least two times, similar data were obtained, and a typical blot was shown.
Immunofluorescence assay
Immunofluorescence assays were performed as described previously [36]. In brief, HeLa cells were transfected with indicated plasmids for 24 h and then fixed in 4 % paraformaldehyde. Cells were incubated with rabbit anti-p65 pAb (diluted 1:50) or rabbit anti-p50 pAb (diluted 1:50) with mouse anti-HA mAb and anti-Myc mAb (diluted 1:2,000), followed by incubation with tetramethyl rhodamine isocyanate (TRITC)-conjugated goat anti-rabbit IgG (Pierce) and fluorescein isothiocyanate (FITC)-conjugated goat anti-mouse IgG (Sigma-Aldrich). After each incubation step, cells were washed extensively with PBS. Samples were analyzed with fluorescence microscopy (Zeiss, Germany).
Results
Suppression of TNF-α-mediated NF-κB activation by HSV-1 UL42
The canonical NF-κB activation pathway can be initiated by various stimuli, including TNF-α, a key cytokine regulating immune functions as well as inflammatory responses. In order to verify whether UL42 protein affected NF-κB activity, we performed dual-luciferase reporter gene assays with cells treated with TNF-α. HA-tagged UL42 was co-expressed in HEK 293T cells in the presence of NF-κB-Luc reporter genes. TNF-α treatment resulted in strong induction of NF-κB activity (Fig. 1a). In contrast, ectopic expression of UL42 significantly inhibited TNF-α-mediated activation of NF-κB promoter activity (Fig. 1a). Additionally, UL42 inhibited NF-κB promoter activity in a dose-dependent manner (Fig. 1b).
HSV-1 UL42 inhibits TNF-α mediated NF-κB activation. HEK 293T cells were transfected with 500 ng of NF-κB promoter reporter plasmid NF-κB-luc (a), together with Renilla luciferase plasmid pRL-TK (50 ng) and pCMV-HA empty vector or plasmids encoding the indicated viral proteins (500 ng). At 24 h post-transfection, cells were treated with or without 10 ng/ml recombinant human TNF-α and incubated for an additional 6 h, followed by cell lysis. NF-κB-driven luciferase activity was determined by a dual-luciferase assay. (b) As in (a), except an increase amount of UL42-HA expression plasmid as indicated was used. The expression of UL42 was analyzed by WB using anti-HA and anti-β-actin (as a control) mAbs. The data represent means plus standard deviations for three replicates. Statistical analysis was performed using Student’s t test. **P < 0.01
The three arginine residues (113, 279, and 280) of UL42 is important for the inhibition of TNF-α-mediated NF-κB activation
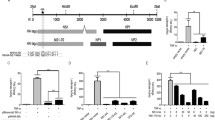
The order of UL42 four arginine mutants’ effects on long-chain DNA synthesis (R280A > R279A ≅ R113A > R182A) were identical to the order of their effects on DNA binding [22]. To determine whether the arginine residues were also responsible for UL42 inhibiting TNF-α-mediated activation of NF-κB, UL42 arginine residue mutants (UL42 2R/2A, UL42 3R/3A) expression plasmids were generated by substitution of alanine for 279R and 280R residues or 113R, 279R and 280R residues (Fig. 2a), we also constructed the UL42(1-390aa) expression plasmid, which encompassed these three arginine residues (Fig. 2a).
The three arginine residues (113, 279, and 280) of UL42 is important for its inhibition of TNF-α-mediated NF-κB activation. a Schematic representation of wild type, arginine residues substitution mutants of UL42 and the N-terminal 390 aa of UL42. b HEK 293T cells were co-transfected with pRL-TK plasmid and NF-κB-Luc reporter plasmid along with empty vector or plasmids encoding UL42 WT, UL42(1-241aa),UL42 2R/2A mutant or UL42 3R/3A mutant. At 24 h post-transfection, cells were treated with or mock treated with recombinant human TNF-α (10 ng/ml), and luciferase activity was analyzed as described in Fig. 1a. (c, d and e) As in (b), except an increase amount of UL42(1-390aa) and UL42 2R/2A mutant or UL42 3R/3A mutant expression plasmids as indicated were used. The expression of UL42, UL42(1-390aa), UL42 2R/2A mutant and UL42 3R/3A mutant were analyzed by WB using anti-HA, anti-Flag, anti-Myc and anti-β-actin (as a control) mAbs. The data represent means plus standard deviations for three replicates. Statistical analysis was performed using Student’s t test. **P < 0.01; *P < 0.05
The three plasmids were transfected into HEK 293T cells and examined for their ability to inhibit TNF-α-induced NF-κB activation, respectively. In agreement with Fig. 1, WT UL42 and the UL42(1-390aa) strongly inhibited TNF-α-induced NF-κB activation. However, UL42 2R/2A and UL42 3R/3A inhibited TNF-α-induced NF-κB activity were less than WT UL42 and the UL42(1-390aa) (Fig. 2b). Additionally, UL42(1-390aa) inhibited NF-κB promoter activity in a dose-dependent manner (Fig. 2c), but increased amount of UL42 2R/2A and UL42 3R/3A expression plasmids have a certain influence on NF-κB promoter activity (Fig. 2d, e). Taken together, the results suggested that three arginine residues (113, 279, 280), which resided on the positively charged surface of UL42, might play a significant role in inhibiting TNF-α-induced NF-κB activation.
UL42 prevents the production of NF-κB-regulated innate cytokines
To determine whether UL42 was required to downregulate NF-κB-regulated cytokines, we performed IL-6 and IL-8 reporter assays using a reporter with an IL-6 or IL-8 promoter driving the expression of a luciferase gene. HEK 293T cells were treated with TNF-α, the luciferase activities in both pIL-6-Luc- and pIL-8-Luc-transfected cells were unaffected by the empty vector control, however, UL42 and UL42(1-390aa) significantly inhibited IL-6 and IL-8 luciferase activity, UL42 2R/2A and UL42 3R/3A have little influence on IL-6 luciferase activity (Fig. 3a). But UL42 2R/2A and UL42 3R/3A have a certain influence on IL-8 luciferase activity (Fig. 3b). These results suggested that the three arginine residues (113, 279, and 280) of UL42 have an important role to inhibit NF-κB-targeted innate immune genes expression.
UL42 inhibits the NF-κB-mediated innate cytokines expression. HEK 293T cells were co-transfected with pcDNA empty vector and UL42-HA or UL42(1-390aa)-Flag or UL42 2R/2A mutant or UL42 3R/3A mutant expression plasmid and px-Luc (where x is IL-6 or IL-8, as indicated) along with pRL-TK as an internal control. At 24 h post-transfection, the cells were treated with recombinant human TNF-α (10 ng/ml) for 4 h and luciferase activity was measured. The data represent means plus standard deviations for three replicates. Statistical analysis was performed using Student’s t test. **P < 0.01. *P < 0.05
UL42 inhibits NF-κB signaling pathway at p65 and p50 level
To determine at what level in the pathway UL42 blocked NF-κB activity, increasing amounts of HA-tagged UL42 expressing plasmid and expression plasmids of the canonical NF-κB signaling pathway components including TRADD, TAK1, IKKβ, and p65 were co-transfected into HEK 293T cells. All expression constructs resulted in a 10–100 fold induction of the NF-κB-Luc reporter activity (Fig. 4a–h). NF-κB promoter activation driven by all of the expression constructs was inhibited by UL42 in a dose-dependent manner (Fig. 4a–d). The activation driven by TRADD, TAK1, and IKKβ was inhibited more than 95 % (Fig. 4a–c) and the activation driven by p65 was inhibited up to 75 % (Fig. 4d). UL42 also inhibited the TRAF2, RIP1, NIK, IKKα induced NF-κB-Luc reporter activity (Fig. S1). These results indicated that UL42 inhibited the NF-κB signaling at or downstream of p65. Indeed, p65 is a crucial transcription factor in NF-κB signaling pathway, and the heterdimeric of p65/p50 is a hallmark of the early activation of the antiviral response [5].
UL42 targets p65 and p50 to inhibit NF-κB signaling. The HEK 293T cells were co-transfected with NF-κB-Luc, pRL-TK and TRADD (a), TAK1 (b), IKKβ (c), p65 and (d) expression plasmids along with indicated amount of UL42 expression plasmid. Luciferase activity was analyzed as described in Fig. 1a. Cell lysates were analyzed by immunoblotting (IB) with tag-specific Abs (bottom panels) to detect the expression of Myc-TRADD, Flag-TAK1, Flag-IKKβ and Flag-p65. The UL42 expression was detected by anti-HA mAb. IB of β-actin was used to verify equal loading of protein in each lane. The data represent means plus standard deviations for three replicates. Statistical analysis was performed using Student’s t test. **P < 0.01
UL42 interacts with endogenous p65 and p50
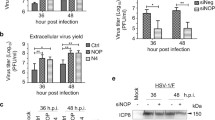
The aforementioned results demonstrated that UL42 inhibited NF-κB signaling pathway at p65/p50 level. In order to clarify the molecular mechanism of UL42 to suppress NF-κB activation, we analyzed the NF-κB subunits p65 and p50 for their potential interaction with viral protein UL42 in co-IP experiments. UL42-HA and either Flag-p65 or Flag-p50 expression plasmids were co-transfected into HEK 293T cells, and co-IP/WB analysis was performed with anti-HA and anti-Flag mAbs. Both ectopically expressed p65 and p50 were efficiently co-IP with UL42 by anti-HA mAb (Fig. 5a, d). We further investigated the interaction between UL42 and endogenous p65 or p50, UL42-HA expression plasmid was transfected into HEK 293T cells and at 24 h post-transfection, and the cells treated with or without 10 ng/ml TNF-α for 8 h. Co-IP/WB analysis was performed with anti-HA and antibodies against p65 or p50, both endogenous p65 and p50 proteins were efficiently co-IP with UL42 by anti-HA mAb (Fig. 5b, c, e, f) in the presence of TNF-α or not. To determine whether UL42 interacts with p65 and p50 under physiological conditions, we performed co-IP experiments using endogenous proteins. In WT HSV-1-infected cells, the UL42 protein was easily immunoprecipitated by p65 or p50 using anti-p65 or anti-p50 pAbs (Fig. 5g, h), respectively, but not by control antibody IgG (Fig. 5g, h). These results demonstrated that UL42 interacts with endogenous p65 and p50 under physiological conditions.
UL42 interacts with p65 and p50. The HEK 293T cells were co-transfected with UL42-HA and Flag-p65 expression plasmids (a), UL42-HA and Flag-p50 expression plasmids (d). At 36 h after transfection, Cells were harvested and lysed, the samples were then subjected to immunoprecipitation assays using anti-HA mAb (IP: HA) or nonspecific mouse monoclonal antibody (IgG2b). Immunoprecipitated samples (IP) were separated by 10 % SDS-PAGE, and proteins were transferred onto a PVDF membrane. IB were probed with the indicated Ab. (b, c, e, f) HEK 293T cells were transfected with UL42-HA expression plasmid. At 24 h post-transfection, cells were mock treated (b, e) or treated with (c, f) recombinant human TNF-α(10 ng/ml) and incubated for an additional 8 h, cells harvested and lysed, the samples were then subjected to immunoprecipitation assays using anti-HA mAb (IP: HA) or nonspecific mouse monoclonal antibody (IgG2b). Immunoprecipitated samples (IP) were separated by 10 % SDS-PAGE, and proteins were transferred onto a PVDF membrane. IB was probed with the indicated Ab. (g, h) UL42 interacts with endogenous p65 and p50 in HSV-1-infected cells. HEK293T cells were infected with WT HSV-1 at an MOI of 10 for 16 h. The cells were then lysed, and the extracts were subjected to immunoprecipitation using anti-p65 pAb (IP: p65), anti-p50 pAb (IP: p50) or control IgG. Precipitates were analyzed by Western blotting (i, j). The HEK 293T cells were co-transfected with UL42-HA and p65 RHD-Flag expression plasmids (i), UL42-HA and p50 RHD-Flag expression plasmids (j). At 36 h after transfection, Cells were harvested and lysed, the samples were then subjected to immunoprecipitation assays using anti-Flag mAb (IP: Flag) or nonspecific mouse monoclonal antibody (IgG2b). Immunoprecipitated samples (IP) were separated by 10 % SDS-PAGE, and proteins were transferred onto a PVDF membrane. IB was probed with the indicated Ab
To determine if UL42 protein bound to the N-terminal RHD of p65 or p50, which is responsible for dimerization, DNA binding and nuclear import [37], we constructed the fragment of p65 spanning amino acid 21–186 (p65 RHD) and the fragment of p50 spanning amino acid 44–242 (p50 RHD), UL42-HA and either Flag-p65 RHD or Flag-p50 RHD expression plasmids were co-transfected into HEK 293T cells, and co-IP/WB analysis was performed with anti-Flag and anti-HA mAbs. UL42 was efficiently co-IP with p65 RHD or p50 RHD by anti-Flag mAb (Fig. 5i, j). Considering these things together, we observed that UL42 bound with the RHD of p65 and p50.
UL42 prevents nuclear translocation of p65 and p50
Nuclear translocation of p65/p50 is crucial for the transcription of NF-κB. Immunofluorescence was carried out to investigate whether UL42 prevented the nuclear translocation of p65 and p50 (Fig. 6). In mock-treated Hela cells, p65 or p50 localized exclusively to the cytoplasm, however, p65 or p50 translocated into the nucleus in the TNF-α-stimulated cells. Ectopic expression of UL42 prevented the nuclear translocation of p65 or p50 induced by TNF-α, whereas UL42 2R/2A and UL42 3R/3A mutants did not (Fig. 6a, b). These results demonstrated that UL42 bound to RHD of p65 and p50 were sufficient to prevent the nuclear accumulation of NF-κB and thereby precluded its transcriptional activity.
UL42 blocks TNF-α-induced p65 and p50 nuclear translocation. HeLa cells were transfected with empty vector, HA-tagged UL42 WT, Myc-tagged UL42 2R/2A mutant or Myc-tagged UL42 3R/3A mutant expression plasmid. At 24 h post-transfection, cells were treated with recombinant human TNF-α (10 ng/ml) or mock treated for 30 min as indicated. Cells were stained with mouse anti-HA or mouse anti-Myc mAb and rabbit anti-p65 pAb or anti-p50 pAb. FITC-conjugated goat anti-rabbit (green) and TRITC-conjugated goat anti-mouse (red) were used as the secondary antibody. Cell nuclei (blue) were stained with Hoechst 33258. The images were obtained by fluorescence microscopy using a ×40 objective (color figure online)
The N-terminus of UL42 is required and sufficient for p65 and p50 binding and suppression of p65/p50-mediated NF-κB activation
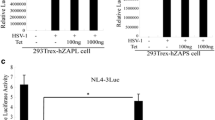
As the three arginine residues (113, 279, and 280) in the N-terminus of UL42 played an important role in inhibiting TNF-α-mediated NF-κB activity, we reasoned that p65 and p50 might interact with the N-terminus of UL42. To verify this, HEK 293T cells were transfected with UL42(1-390aa)-Flag expression plasmid, at 24 h post-transfection, the cells treated with or without 10 ng/ml TNF-α for 8 h. As results, UL42(1-390aa) was efficiently co-IP with endogenous p65/p50 (Fig. 7a–d). To determine whether UL42(1-390aa) really interacted with p65 and p50, we performed co-IP experiments using endogenous proteins. In UL42(1-390aa)-Flag transfected cells, the UL42(1-390aa) protein was easily immunoprecipitated by p65 or p50 using anti-p65 or anti-p50 pAbs, respectively (Fig. S2), but not by control antibody IgG (Fig. S2). These results demonstrated that UL42(1-390aa) interacted with endogenous p65 and p50.
The N-terminus of UL42 is required and sufficient for p65/p50 binding and suppression of p65/p50-mediated NF-κB Activation. The HEK 293T cells were transfected with UL42(1-390aa)-Flag expression plasmid. At 24 h after transfection, cells were mock treated (a, c) or treated with (b, d) recombinant human TNF-α (10 ng/ml) and incubated for an additional 8 h, cells harvested and lysed, the samples were then subjected to immunoprecipitation assays using anti-Flag mAb (IP: Flag) or nonspecific mouse monoclonal antibody (IgG2b). Immunoprecipitated samples (IP) were separated by 10 % SDS-PAGE, and proteins were transferred onto a PVDF membrane. IB were probed with the indicated Ab. e The HEK 293T cells were co-transfected with NF-κB-Luc, pRL-TK, Flag-p65 and Flag-p50 expression plasmids along with indicated amount of UL42-HA, UL42(1-390aa)-Flag, UL42 2R/2A-Myc or UL42 3R/3A-Myc expression plasmids. Luciferase activity was analyzed as described in Fig. 1a. Cell lysates were analyzed by IB with tag-specific Abs (bottom panels) to detect the expression of Flag-p65, Flag-p50, UL42-HA, UL42(1-390aa)-Flag, UL42 2R/2A-Myc and UL42 3R/3A-Myc. IB of β-actin was used to verify equal loading of protein in each lane. The data represent means plus standard deviations for three replicates. Statistical analysis was performed using Student’s t test. **P < 0.01; *P < 0.05
To clarify whether binding of the UL42(1-390aa) is also sufficient for inhibition of NF-κB transcriptional activity, p65 and p50 were overexpressed in HEK 293T cells along with UL42-HA, UL42(1-390aa)-Flag, UL42 2R/2A-Myc or UL42 3R/3A-myc. A dose-dependent reduction of NF-κB-luc activity confirmed that binding of UL42(1-390aa) was sufficient for its inhibition (Fig. 7e). In contrast, expression of the UL42 2R/2A and UL42 3R/3A mutants had a certain effect on p65/p50-mediated luciferase activity. But the inhibiting capacity was less than WT UL 42 and UL42(1-390aa). In summary, the N-terminus of UL42 was sufficient for binding to p65 and p50, and also the three arginine residues (113, 279, 280) of the UL42 protein played an significant role in inhibiting p65/p50-mediated NF-κB activation.
Discussion
HSV-1 infection can cause proinflammatory cytokine expression with elevate levels of proinflammatory and inflammatory cytokines, including TNF-α [38, 39]. TNF-α activates the NF-κB signaling pathway that regulates expression of many cellular factors playing important roles in innate immune responses and inflammation in infected hosts [40]. In this study, we report that HSV-1 UL42 protein can directly interact with p65 and p50 by inhibiting p65/p50 nuclear translocation and thus inhibits TNF-α-induced NF-κB activation.
We found that UL42 could inhibit TRADD, TRAF2, TAK1, RIP1, NIK, IKKα, IKKβ or p65-mediated activation of NF-κB-Luc promoter activity, implying that UL42 might act at or downstream of the p65/p50 level. We therefore hypothesized that UL42 may interact with p65 or p50 to inhibit NF-κB promoter activity. To validate our hypothesis, co-IP analysis was employed to determine whether UL42 could directly interact with p65 or p50 under physiological conditions. As expected, UL42 interacted with both p65 and p50. Thus, HSV-1 UL42 could inhibit the canonical NF-κB signaling pathway by interacting with p65 and p50. We also found that UL42 could inhibit NIK and IKKα induced NF-κB promoter activity, suggesting that UL42 might be an antagonism of the noncanonical NF-κB signaling pathway.
Intriguingly, when we substituted alanine for R113, R279 and R280, the mutants UL42 2R/2A and UL42 3R/3A had a certain influence on TNF-α and p65/p50 stimulated NF-κB promoter activity, but inhibiting capacity was less than WT UL42 and UL42(1-390aa). The N-terminal of UL42 was sufficient for the specific binding of UL42 to the p65/p50 and the inhibition of canonical NF-κB activation. And also UL42 3R/3A had little influence on p65 and p50 nuclear translocation, so we speculated that arginine residues of UL42 were crucial for UL42 binding to NF-κB promoter and inhibited p65 and p50 bound to NF-κB promoter. Thus, these arginine residues of UL42 not only had an important role in HSV-1 lytic infection, but also were significant for HSV-1 to inhibit host cytokine production.
The relationship between HSV-1 infection and the host NF-κB innate immune response is quite complex, it is well known that the classical NF-κB pathway is significantly activated upon HSV-1 infections [41, 42] and is shown to be involved in up-regulation of several host genes [43], as well as in promoting the progression of the virus replication cycle [42, 44]. However, NF-κB signaling also promotes inflammatory cytokine expression to reduce viral infection. How could HSV-1 selectively promote sufficient NF-κB activity to promote its propagation and avoid the robust innate cytokine response driven by the same NF-κB activity? This apparent contradiction is reconciled by the fact that HSV-1 activates NF-κB through multiple signaling pathways. The early wave involves the UL31 and UL37 virion tegument protein [45, 46] and possibly virion glycoprotein D, glycoprotein B, glycoprotein H and glycoprotein L [47, 48]. UL37 activates NF-κB signaling through interaction with TRAF6, glycoprotein D activates the TNF superfamily receptor HVEM to drive both the canonical and alternative NF-κB pathway; glycoprotein B, glycoprotein H and glycoprotein L are ligands to TLR2, they initiate a signaling cascade which induce NF-κB activation. Thus, HSV-1 may transiently activate NF-κB to start its infection optimally and then inhibit the major part of the innate response using viral gene products expressed during infection. UL42 might be a pivotal regulator in maintaining the balance between suppressing an antiviral response at a tolerable limit and keeping the sufficient activation of NF-κB for viral replication, and consequently contribute to viral virulence.
HSV-1 UL42, as an antagonism of TNF-α-induced NF-κB pathway, may be one of several strategies used by HSV-1 to interrupt the innate immune system. However, more work should be done to reveal the comprehensive tricks by HSV-1 to subvert the host immune response. In conclusion, this is the first study to demonstrate that the UL42 protein suppresses NF-κB-mediated antiviral response via interactions with both RHD domain of p65 and p50. This function of UL42 may benefit the regulation of the delicate balance between the replication process and host defense programs induced by the NF-κB pathway.
References
Akira S, Uematsu S, Takeuchi O (2006) Pathogen recognition and innate immunity. Cell 124(4):783–801. doi:10.1016/j.cell.2006.02.015
Beutler B, Eidenschenk C, Crozat K, Imler JL, Takeuchi O, Hoffmann JA, Akira S (2007) Genetic analysis of resistance to viral infection. Nat Rev Immunol 7(10):753–766. doi:10.1038/nri2174
Medzhitov R (2007) Recognition of microorganisms and activation of the immune response. Nature 449(7164):819–826. doi:10.1038/nature06246
Koyama S, Ishii KJ, Coban C, Akira S (2008) Innate immune response to viral infection. Cytokine 43(3):336–341. doi:10.1016/j.cyto.2008.07.009
Oeckinghaus A, Ghosh S (2009) The NF-kappaB family of transcription factors and its regulation. Cold Spring Harb Perspect Biol 1(4):a000034. doi:10.1101/cshperspect.a000034
Ghosh S, May MJ, Kopp EB (1998) NF-kappa B and Rel proteins: evolutionarily conserved mediators of immune responses. Annu Rev Immunol 16:225–260. doi:10.1146/annurev.immunol.16.1.225
Li Q, Verma IM (2002) NF-kappaB regulation in the immune system. Nat Rev Immunol 2(10):725–734. doi:10.1038/nri910
Bonizzi G, Karin M (2004) The two NF-kappaB activation pathways and their role in innate and adaptive immunity. Trends Immunol 25(6):280–288. doi:10.1016/j.it.2004.03.008
Chen ZJ (2005) Ubiquitin signalling in the NF-kappaB pathway. Nat Cell Biol 7(8):758–765. doi:10.1038/ncb0805-758
Hayden MS, Ghosh S (2004) Signaling to NF-kappaB. Genes Dev 18(18):2195–2224. doi:10.1101/gad.1228704
Gilmore TD (2006) Introduction to NF-kappaB: players, pathways, perspectives. Oncogene 25(51):6680–6684. doi:10.1038/sj.onc.1209954
Baeuerle PA, Baltimore D (1996) NF-kappa B: ten years after. Cell 87(1):13–20
Baldwin AS Jr (1996) The NF-kappa B and I kappa B proteins: new discoveries and insights. Annu Rev Immunol 14:649–683. doi:10.1146/annurev.immunol.14.1.649
Leib DA (2002) Counteraction of interferon-induced antiviral responses by herpes simplex viruses. Curr Top Microbiol Immunol 269:171–185
Paladino P, Mossman KL (2009) Mechanisms employed by herpes simplex virus 1 to inhibit the interferon response. J Interferon Cytokine Res 29(9):599–607. doi:10.1089/jir.2009.0074
van Lint AL, Murawski MR, Goodbody RE, Severa M, Fitzgerald KA, Finberg RW, Knipe DM, Kurt-Jones EA (2010) Herpes simplex virus immediate-early ICP0 protein inhibits Toll-like receptor 2-dependent inflammatory responses and NF-kappaB signaling. J Virol 84(20):10802–10811. doi:10.1128/JVI.00063-10
Daubeuf S, Singh D, Tan Y, Liu H, Federoff HJ, Bowers WJ, Tolba K (2009) HSV ICP0 recruits USP7 to modulate TLR-mediated innate response. Blood 113(14):3264–3275. doi:10.1182/blood-2008-07-168203
Jin H, Ma Y, Yan Z, Prabhakar BS, He B (2012) Activation of NF-kappaB in CD8 + dendritic cells ex vivo by the gamma134.5 null mutant correlates with immunity against herpes simplex virus 1. J Virol 86(2):1059–1068. doi:10.1128/JVI.06202-11
Cotter CR, Kim WK, Nguyen ML, Yount JS, Lopez CB, Blaho JA, Moran TM (2011) The virion host shutoff protein of herpes simplex virus 1 blocks the replication-independent activation of NF-kappaB in dendritic cells in the absence of type I interferon signaling. J Virol 85(23):12662–12672. doi:10.1128/JVI.05557-11
Trego KS, Parris DS (2003) Functional interaction between the herpes simplex virus type 1 polymerase processivity factor and origin-binding proteins: enhancement of UL9 helicase activity. J Virol 77(23):12646–12659
Chaudhuri M, Parris DS (2002) Evidence against a simple tethering model for enhancement of herpes simplex virus DNA polymerase processivity by accessory protein UL42. J Virol 76(20):10270–10281
Randell JC, Komazin G, Jiang C, Hwang CB, Coen DM (2005) Effects of substitutions of arginine residues on the basic surface of herpes simplex virus UL42 support a role for DNA binding in processive DNA synthesis. J Virol 79(18):12025–12034. doi:10.1128/JVI.79.18.12025-12034.2005
Jiang C, Hwang YT, Randell JC, Coen DM, Hwang CB (2007) Mutations that decrease DNA binding of the processivity factor of the herpes simplex virus DNA polymerase reduce viral yield, alter the kinetics of viral DNA replication, and decrease the fidelity of DNA replication. J Virol 81(7):3495–3502. doi:10.1128/JVI.02359-06
Jiang C, Hwang YT, Wang G, Randell JC, Coen DM, Hwang CB (2007) Herpes simplex virus mutants with multiple substitutions affecting DNA binding of UL42 are impaired for viral replication and DNA synthesis. J Virol 81(21):12077–12079. doi:10.1128/JVI.01133-07
Xing J, Wang S, Li Y, Guo H, Zhao L, Pan W, Lin F, Zhu H, Wang L, Li M, Zheng C (2011) Characterization of the subcellular localization of herpes simplex virus type 1 proteins in living cells. Med Microbiol Immunol 200(1):61–68. doi:10.1007/s00430-010-0175-9
Jiang X, Takahashi N, Matsui N, Tetsuka T, Okamoto T (2003) The NF-kappa B activation in lymphotoxin beta receptor signaling depends on the phosphorylation of p65 at serine 536. J Biol Chem 278(2):919–926. doi:10.1074/jbc.M208696200
Severa M, Coccia EM, Fitzgerald KA (2006) Toll-like receptor-dependent and -independent viperin gene expression and counter-regulation by PRDI-binding factor-1/BLIMP1. J Biol Chem 281(36):26188–26195. doi:10.1074/jbc.M604516200
Schuhmann KM, Pfaller CK, Conzelmann KK (2011) The measles virus V protein binds to p65 (RelA) to suppress NF-kappaB activity. J Virol 85(7):3162–3171. doi:10.1128/JVI.02342-10
Mercurio F, Zhu H, Murray BW, Shevchenko A, Bennett BL, Li J, Young DB, Barbosa M, Mann M, Manning A, Rao A (1997) IKK-1 and IKK-2: cytokine-activated IkappaB kinases essential for NF-kappaB activation. Science 278(5339):860–866
Park KA, Byun HS, Won M, Yang KJ, Shin S, Piao L, Kim JM, Yoon WH, Junn E, Park J, Seok JH, Hur GM (2007) Sustained activation of protein kinase C downregulates nuclear factor-kappaB signaling by dissociation of IKK-gamma and Hsp90 complex in human colonic epithelial cells. Carcinogenesis 28(1):71–80. doi:10.1093/carcin/bgl094
Gao S, Song L, Li J, Zhang Z, Peng H, Jiang W, Wang Q, Kang T, Chen S, Huang W (2012) Influenza A virus-encoded NS1 virulence factor protein inhibits innate immune response by targeting IKK. Cell Microbiol. doi:10.1111/cmi.12005
Jordan M, Schallhorn A, Wurm FM (1996) Transfecting mammalian cells: optimization of critical parameters affecting calcium-phosphate precipitate formation. Nucleic Acids Res 24(4):596–601
Zhong B, Yang Y, Li S, Wang YY, Li Y, Diao F, Lei C, He X, Zhang L, Tien P, Shu HB (2008) The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29(4):538–550. doi:10.1016/j.immuni.2008.09.003
Zhu H, Zheng C, Xing J, Wang S, Li S, Lin R, Mossman KL (2011) Varicella-zoster virus immediate-early protein ORF61 abrogates the IRF3-mediated innate immune response through degradation of activated IRF3. J Virol 85(21):11079–11089. doi:10.1128/JVI.05098-11
Xing J, Wang S, Lin F, Pan W, Hu CD, Zheng C (2011) Comprehensive characterization of interaction complexes of herpes simplex virus type 1 ICP22, UL3, UL4, and UL20.5. J Virol 85(4):1881–1886. doi:10.1128/JVI.01730-10
Xing J, Wu F, Pan W, Zheng C (2010) Molecular anatomy of subcellular localization of HSV-1 tegument protein US11 in living cells. Virus Res 153(1):71–81. doi:10.1016/j.virusres.2010.07.009
May MJ, Ghosh S (1997) Rel/NF-kappa B and I kappa B proteins: an overview. Semin Cancer Biol 8(2):63–73. doi:10.1006/scbi.1997.0057
Hu S, Sheng WS, Schachtele SJ, Lokensgard JR (2011) Reactive oxygen species drive herpes simplex virus (HSV)-1-induced proinflammatory cytokine production by murine microglia. J Neuroinflammation 8:123. doi:10.1186/1742-2094-8-123
Li H, Zhang J, Kumar A, Zheng M, Atherton SS, Yu FS (2006) Herpes simplex virus 1 infection induces the expression of proinflammatory cytokines, interferons and TLR7 in human corneal epithelial cells. Immunology 117(2):167–176. doi:10.1111/j.1365-2567.2005.02275.x
Seet BT, Johnston JB, Brunetti CR, Barrett JW, Everett H, Cameron C, Sypula J, Nazarian SH, Lucas A, McFadden G (2003) Poxviruses and immune evasion. Annu Rev Immunol 21:377–423. doi:10.1146/annurev.immunol.21.120601.141049
Patel A, Hanson J, McLean TI, Olgiate J, Hilton M, Miller WE, Bachenheimer SL (1998) Herpes simplex type 1 induction of persistent NF-kappa B nuclear translocation increases the efficiency of virus replication. Virology 247(2):212–222
Amici C, Belardo G, Rossi A, Santoro MG (2001) Activation of I kappa b kinase by herpes simplex virus type 1. A novel target for anti-herpetic therapy. J Biol Chem 276(31):28759–28766. doi:10.1074/jbc.M103408200
Taddeo B, Esclatine A, Roizman B (2002) The patterns of accumulation of cellular RNAs in cells infected with a wild-type and a mutant herpes simplex virus 1 lacking the virion host shutoff gene. Proc Natl Acad Sci U S A 99(26):17031–17036. doi:10.1073/pnas.252588599
Gregory D, Hargett D, Holmes D, Money E, Bachenheimer SL (2004) Efficient replication by herpes simplex virus type 1 involves activation of the IkappaB kinase-IkappaB-p65 pathway. J Virol 78(24):13582–13590. doi:10.1128/JVI.78.24.13582-13590.2004
Liu X, Fitzgerald K, Kurt-Jones E, Finberg R, Knipe DM (2008) Herpesvirus tegument protein activates NF-kappaB signaling through the TRAF6 adaptor protein. Proc Natl Acad Sci USA 105(32):11335–11339. doi:10.1073/pnas.0801617105
Roberts KL, Baines JD (2011) UL31 of herpes simplex virus 1 is necessary for optimal NF-kappaB activation and expression of viral gene products. J Virol 85(10):4947–4953. doi:10.1128/JVI.00068-11
Medici MA, Sciortino MT, Perri D, Amici C, Avitabile E, Ciotti M, Balestrieri E, De Smaele E, Franzoso G, Mastino A (2003) Protection by herpes simplex virus glycoprotein D against Fas-mediated apoptosis: role of nuclear factor kappaB. J Biol Chem 278(38):36059–36067. doi:10.1074/jbc.M306198200
Leoni V, Gianni T, Salvioli S, Campadelli-Fiume G (2012) Herpes simplex virus glycoproteins gH/gL and gB bind Toll-like receptor 2, and soluble gH/gL is sufficient to activate NF-kappaB. J Virol 86(12):6555–6562. doi:10.1128/JVI.00295-12
Acknowledgments
This work was supported by grants from the Major State Basic Research Development Program of China (973 Program) (2011CB504800 and 2010CB530100); the Start-up Fund of the Hundred Talents Program of the Chinese Academy of Sciences (20071010-141); the National Natural Science Foundation of China (81171584). We thank Dr. Rao Anjana (Immune Disease Institute, Harvard Medical School), and Dr. Gangmin Hur (Chungnam National University) for providing plasmids.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
430_2013_295_MOESM1_ESM.tif
Fig S1. UL42 inhibits TRAF2, RIP1, NIK, IKKα-induced NF-κB Activity. The HEK 293T cells were co-transfected with NF-κB-Luc, pRL-TK and TRAF2 (A), RIP1 (B), NIK (C), IKKα and (D) expression plasmids along with indicated amount of UL42 expression plasmid. Luciferase activity was analyzed as described in Fig. 1A. Cell lysates were analyzed by immunoblotting (IB) with tag-specific Abs (bottom panels) to detect the expression of Flag-TRAF2, Flag-TAK1 and HA-IKKα. The UL42 expression was detected by anti-HA mAb. IB of β-actin was used to verify equal loading of protein in each lane. The data represent means plus standard deviations for three replicates. Statistical analysis was performed using Student’s t test. **, P < 0.01. (TIFF 1338 kb)
430_2013_295_MOESM2_ESM.tif
Fig S2. UL42(1-390aa) interacts with p65 and p50. The HEK 293T cells were transfected with UL42(1-390aa)-Flag expression plasmid. At 24 h after transfection, cells harvested and lysed, the samples were then subjected to immunoprecipitation assays using anti-p65 pAb (IP: p65), anti-p50 pAb (IP: p50) or control antibody IgG. Immunoprecipitated samples (IP) were separated by 10 % SDS-PAGE, and proteins were transferred onto a PVDF membrane. IB was probed with the indicated Ab. (TIFF 497 kb)
Rights and permissions
About this article
Cite this article
Zhang, J., Wang, S., Wang, K. et al. Herpes simplex virus 1 DNA polymerase processivity factor UL42 inhibits TNF-α-induced NF-κB activation by interacting with p65/RelA and p50/NF-κB1. Med Microbiol Immunol 202, 313–325 (2013). https://doi.org/10.1007/s00430-013-0295-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-013-0295-0