Abstract
Invasive bladder cancer is diverse, and includes several named histomorphologies that differ from conventional urothelial carcinoma, termed “histologic variants.” By transcriptional analysis, bladder cancers can be divided into luminal and basal subtypes. In this paper, we study associations between markers of transcriptional subtypes and variant histology in a retrospective cohort of 309 cystectomy specimens. Histology slides were methodically reviewed for all cases, and variant histology was documented. Immunohistochemistry for FOXA1 (luminal marker) and CK14 (basal maker) was performed on histologic variants and their associated conventional urothelial carcinomas. Invasive carcinoma was present in 270 of the cystectomy specimens, 35% of which contained a histologic variant. Squamous carcinomas expressed higher CK14 levels than micropapillary, nested, and plasmacytoid carcinomas (p < 0.001, Kruskal-Wallis), keeping with the basal character of squamous carcinoma. Likewise, squamous carcinomas expressed lower FOXA1 levels than micropapillary, nested, and plasmacytoid carcinomas (p < 0.001, Kruskal-Wallis), keeping with the luminal character of micropapillary carcinoma, and suggesting that nested and plasmacytoid cancers have luminal character. FOXA1 was expressed at lower levels in conventional urothelial carcinoma associated with squamous carcinoma than conventional urothelial carcinoma associated with micropapillary carcinoma (p = 0.0072, Wilcoxon rank sum). CK14 expression did not differ between conventional urothelial carcinomas associated with squamous versus micropapillary carcinoma (p = 0.89, Wilcoxon rank sum). Instead, CK14 expression was higher in squamous carcinoma than conventional urothelial carcinoma present in the same bladder (p = 0.014, Wilcoxon rank sum, paired). Overall, the findings show that squamous and micropapillary cancers have different expression patterns of CK14 and FOXA1 and suggest that they arise from distinct precursors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Invasive bladder cancer is histologically diverse. While the majority of invasive bladder cancers have the morphology of conventional urothelial carcinoma (conventional UC; also known as transitional cell carcinoma), distinct variants include micropapillary, squamous, plasmacytoid, nested, and other rarer ones [1, 2]. Several of these “histologic variants” are inherently aggressive, and management thus differs from conventional UC. For example, early stage (pT1) micropapillary cancer has high rates of metastasis and death compared to early stage conventional UC, and some advocate radical cystectomy rather than transurethral resection and intravesical therapy [3, 4], though this area is somewhat controversial [5,6,7]. As several histologic variants are inherently aggressive, there is potential clinical value in understanding their underlying biology.
Molecular studies have revealed that bladder cancer may be subtyped based on transcriptional profile, and the resulting transcriptional subtypes may inform on prognosis and chemotherapy response [8,9,10,11,12]. Though studies differ in the number of named transcriptional subtypes and the genes that define them, it has been consistently found that bladder cancer may be broadly categorized into two groups: a “luminal” group, which expresses markers of urothelial differentiation including Forkhead box A1 (FOXA1), GATA-binding protein 3 (GATA3), and the uroplakins, and a “basal” group, which expresses high molecular weight keratins, such as cytokeratin 14 (CK14), and other markers of more primitive, basal urothelial cells [8, 13].
Variant histology associates with transcriptional subtype. Specifically, tumors with micropapillary histology tend to be luminal, while tumors with squamous histology tend to be basal [8, 13, 14]. Bladder cancers with variant histology often have a separate component of conventional UC, which presumably arises from the same precursor as the histologic variant.
In the study presented here, we investigate associations between markers of transcriptional subtype and four histologic variants of bladder cancer: squamous, micropapillary, nested, and plasmacytoid. Specifically, we study patterns of mutual exclusivity among histologic variants and evaluate expression of FOXA1 (luminal marker) and CK14 (basal marker) in histologic variants and their associated conventional UCs.
Material and methods
Case selection and slide review
A consecutive series of specimens was collected from the pathology archives of Penn State Hershey Medical Center, including all radical cystectomies and cystoprostatectomies performed for bladder cancer from May 2001 to February 2014 (n = 309; excluding cases for which slides were not available). Pathology slides were assembled and all sections of bladder and prostate were methodically re-reviewed by study pathologists (JIW and GC). All diagnostic slides were reviewed by a subspecialized urologic pathologist with fellowship training (JIW) to confirm the diagnosis.
Criteria for variant histology classification
Documented variant morphologies included micropapillary, squamous, nested, and plasmacytoid. Thresholds for assigning a variant pattern were strict, including only cases fitting World Health Organization descriptions [2]. Micropapillary was strictly defined as classic micropapillary. That is, prior studies have shown that invasive bladder cancers may be composed of admixed large and small cancer nests with retraction, similar to classic micropapillary carcinoma [15, 16]. However, these are irreproducibly considered micropapillary by pathologists [15], and were thus not considered micropapillary in the current study. Squamous differentiation was strictly considered tumor with definite keratin formation or intercellular bridges. Nested cases were strictly considered invasive small nests of carcinoma with cytologically bland nuclei. Plasmacytoid cases were strictly considered as cancers composed of diffusely infiltrative single cells, similar in appearance to lobular breast cancer. For consistency in nomenclature, we include all cancers with a given variant morphology under that same category, irrespective of other cancer morphologies in the specimen. For example, “squamous bladder cancer” describes carcinoma with squamous morphology, irrespective of the presence of conventional UC within the specimen.
This study also sought to compare conventional UC to cancer with variant histology within the same bladder. As such, for cases with a variant histology, we only considered a focus as conventional UC if the focus was obviously spatially and morphologically distinct from the histologic variant. If conventional UC and histologic variant were tightly admixed), and did not consider any of the focus as conventional UC. For example, in bladders containing squamous bladder cancer, we only considered a separate focus as conventional UC if (1) it was spatially distinct from the squamous cancer, and (2) its morphology was qualitatively different from the squamous cancer (i.e., it did not merely look like the squamous cancer with the keratin subtracted).
Selection of markers for luminal and basal subtype
Single markers were chosen to represent luminal and basal subtypes. Prior study has identified FOXA1 as a marker of the luminal subtype [17], and we have shown that overexpression of this transcription factor operates in a cooperative manner to drive the luminal expression subtype [18], while genetic ablation of this transcription factor results in squamous metaplasia [19]. FOXA1 was chosen as our luminal marker because it is a strong nuclear marker, and our group has extensive experience with this gene [19,20,21,22]. We chose CK14 as the basal marker, as this is a known marker of basal urothelium and stem cells [23], and is overexpressed in bladder cancers of the basal subtype [8].
To determine the accuracy of FOXA1 and CK14 as markers of luminal and basal subtype, respectively, we collected gene expression and transcriptional subtyping data for 127 cases of muscle invasive bladder cancer from The Cancer Genome Atlas (TCGA) data from the Genomic Data Commons (https://portal.gdc.cancer.gov/). TCGA type 1 and 2 tumors were considered luminal, and type 3 and 4 tumors were considered basal. An expression matrix was created, log2 transformed, and scaled by Z-score. FOXA1 and CK14 expression were compared between luminal and basal cancers (see the “Results” section).
Tissue microarray construction
Tissue microarrays (TMAs) were created that included histologic variants and conventional UCs. Specifically, histology slides were reviewed and areas of histologic variant and conventional UC selected and circled. Corresponding tissue blocks (formalin fixed paraffin embedded tissues) were retrieved and aligned to histology slides. Single 3 mm punches were taken from tissue blocks from areas of interest to include histologic variants and conventional UCs. Paraffin recipient blocks were punched with 3 mm cores, and donor cores were placed in recipient blocks using a manual arrayer. The resulting blocks was baked at 42 °C for 40 min, then allowed to cool. Multiple 4 μm sections were taken from the final TMA blocks.
Immunohistochemistry assays
Immunohistochemistry (IHC) was performed as previously described [18]. Briefly, slides were deparaffinized and rehydrated through a series of graded alcohols and washed in deionized water for 5 min. Antigen retrieval was performed by placing slides in 1% antigen unmasking solution (Vector Labs, Burlingame, CA) and heating slides for 20 min on high power in a pressure cooker (Cuisinart CPC-600FR). Steam was released in short bursts to prevent boiling and preserve tissue integrity. Slides were cooled to room temperature and washed three times for 10 min in PBS (pH 7.4). All incubations were performed at room temperature unless otherwise noted. Endogenous peroxidases were blocked by incubation in 3% hydrogen peroxide in methanol for 20 min, and slides were again washed three times for 10 min in phosphate-buffered saline (PBS). Sections were incubated in PBS containing horse serum (Vector Labs) for 1 h to reduce nonspecific antibody binding and then incubated overnight with primary antibody at 4 °C in a humidified chamber. Primary antibodies used for IHC include goat anti FOXA1 (1:1000; Santa Cruz Biotechnology, Santa Cruz, CA) and mouse anti Cytokeratin 14 (CK14; 1:200; Vector Laboratories, Burlingame CA). Following overnight incubation, slides were washed three times for 10 min in PBS and sections were incubated in biotinylated secondary antibody diluted in PBS containing horse serum (1:200; Vector Labs) for 1 h. Specific antibody binding was visualized using Vectastain Elite ABC Peroxidase kit (Vector Labs) according to the manufacturer protocol with diaminobenzidine substrate buffer as the chromogen Dako/Agilent.
Immunohistochemistry evaluation
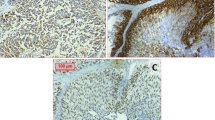
FOXA1 expression was scored with the Allred scoring system [24]. In short, nuclear expression intensity was graded 0 to 3 (least to most intense), and area with positive expression was assigned a score from 0 to 5 (0 = none, 1 = 1%, 2 = 10%, 3 = 33%, 4 = 66%, 5 = 100% cells positive). The intensity and area scores were added to give a final score, with a maximum possible score of 8 and a minimum of 0. CK14 expression was graded similarly. Cytoplasmic expression intensity was graded 0 to 3 (least to most intense), and area with positive expression was assigned a score from 0 to 5 (0 = none, 1 = 1%, 2 = 10%, 3 = 33%, 4 = 66%, 5 = 100% cells positive). The intensity and area scored were added to give a final score, with a maximum possible score of 8 and a minimum of 0. A single pathologist scored all tumors (JIW). Examples are FOXA1 and CK14 immunohistochemistry are presented in Fig. 1.
H&E-stained sections of histologic variants (a–d; ×200). a Squamous carcinoma—invasive carcinoma with definite keratin formation. b Micropapillary carcinoma—small cell nests with surrounding retraction and reverser polarization. c Nested carcinoma—invasive cancer nests with banal nuclei. d Plasmacytoid carcinoma—invasive single cells. Immunohistochemistry (e, f; ×400). FOXA1 expression in conventional urothelial carcinoma; this was assigned Allred score of 5 (area) + 3 (intensity) = 8. CK14 expression in conventional urothelial carcinoma; with assigned score of 5 (area) + 3 (intensity) = 8
Statistical analysis
The Wilcoxon rank sum test, Kruskal-Wallis test, and Fisher exact test were used utilized as designated; p < 0.05 was considered significant. Fisher’s exact test and Wilcoxon rank sum test were performed using the stats package in the R programming language, version 3.1.1 [25]. The Kruskal-Wallis test and follow-up tests of individual significance were performed with the pgirmess package in R [26]. Heatmaps were constructed with the ggplot2 package in R [27].
Results
Diagnostic performance of FOXA1 and CK14 in distinguishing luminal from basal bladder cancer in TCGA data set
In keeping with previous findings, differential expression of FOXA1 and CK14 was detected between luminal and basal cancers in TCGA data (p < 0.001 for both, Wilcoxon rank sum; FOXA1 higher in luminal, CK14 higher in basal; Fig. S1). By receiver operating characteristic (ROC) analysis [28], FOXA1 expression separated luminal from basal cancers with area under the curve (AUC) of 0.88, and CK14 expression separated luminal from basal cancers with AUC of 0.79. FOXA1 and CK14 were independently associated with transcriptional subtype in multivariate analysis (logit, p < 0.001 for FOXA1 and CK14). A score including FOXA1 and CK14 (see below, equation based on logit model) demonstrated AUC of 0.92 for prediction of transcriptional subtype. We thus considered FOXA1 and CK14 to be reasonable markers of luminal and basal subtypes, respectively.
Patient demographics and tumor stage
Of the 309 consecutive cases in our in-house cohort, 270 had invasive carcinoma. No patient had schistosomiasis infection per history. Clinical, demographic, and staging data are presented in Table 1.
Variant morphology prevalence and mutual-exclusivity
Variant histology of at least one type was identified in 35% of invasive cases (94/270). These included squamous (52 cases, 19%), micropapillary (27 cases, 10%), plasmacytoid (8 cases, 3%), and nested (7 cases, 3%), as shown in Table 2. Some cases had more than one variant pattern.
Two cases had concurrent squamous and micropapillary carcinoma, and one case had both plasmacytoid and micropapillary carcinoma. Patterns of mutual-exclusivity were identified among several variant patterns. Most notably, squamous carcinoma was largely mutually exclusive of micropapillary carcinoma, which approached statistical significance (p = 0.074, Fisher’s exact test, one-sided). Nested and plasmacytoid morphologies appeared mutually exclusive of the others, though there were too few cases to reasonably perform test statistics. Concurrent non-invasive papillary UC was more common in micropapillary carcinoma than squamous carcinoma (p = 0.05, Fisher), though concurrent flat carcinoma did not differ in prevalence among these variants (p = 1).
Of histologic variants, 70 cases were available for immunohistochemistry (39 squamous, 19 micropapillary, 5 nested, 7 plasmacytoid). Of these, 56 had not received neoadjuvant chemotherapy (38 squamous, 11 micropapillary, 2 nested, 5 plasmacytoid). Patterns of mutual exclusivity among variants and their expression of FOXA1 and CK14 are presented in Fig. 2.
Morphologic variants and Molecular subtype, with FOXA1 expression (a) and CK14 expression (b). Each row is a patient, and each column is a histologic pattern. The rows (patients) are in the same order for a and b. Conventional urothelial carcinoma was only included for a patient if it was spatially and morphologically distinct from the histologic variant
FOXA1 and CK14 expression in histologic variants
Expression of FOXA1 was lower in squamous carcinoma than micropapillary, plasmacytoid, and nested carcinomas (Fig. 3a; p < 0.001, Kruskal Wallis; multiple comparison p < 0.05 for squamous versus micropapillary, plasmacytoid, and nested; other multiple comparisons p > 0.05). Expression of CK14 was likewise higher in squamous carcinoma than micropapillary, plasmacytoid, and nested carcinoma (Fig. 3b; p < 0.001, Kruskal Wallis; multiple comparison p < 0.05 for squamous versus micropapillary, plasmacytoid, and nested; other multiple comparisons p > 0.05).
When including only cases that had not received neoadjuvant chemotherapy, all significant associations above were maintained (p < 0.05) except for FOXA1 and CK14 expression in squamous versus plasmacytoid carcinoma on multiple comparisons testing (p > 0.05).
FOXA1 and CK14 expression in conventional urothelial carcinoma associated with histologic variants
Conventional UC associated with squamous carcinoma had lower FOXA1 expression than conventional UC associated with micropapillary carcinoma (Fig. 4a; p = 0.0072, Wilcoxon rank sum test; cases with both micropapillary and squamous morphology excluded). This association was held when removing patients who had received neoadjuvant chemotherapy (all p = 0.019). In contrast, expression of CK14 did not differ between conventional UC associated with squamouscarcinoma and conventional UC associated with micropapillary carcinoma (Fig. 4b; p = 0.89, Wilcoxon rank sum test).
FOXA1 and CK14 expression in histologic variant versus conventional urothelial carcinoma in the same bladder
In bladders with both squamous carcinoma and conventional UC, CK14 was higher in the squamous component (Fig. 5; p = 0.014, Wilcoxon rank sum test, paired), while FOXA1 expression did not differ between the squamous and conventional components (p = 0.10, Wilcoxon rank sum test, paired). The former association held when removing patients who had received neoadjuvant chemotherapy (p = 0.014, Wilcoxon rank sum test, paired). In bladders with both micropapillary carcinoma and conventional UC, neither FOXA1 nor CK14 expression differed between the two components (p = 0.28 and p = 0.71, respectively, Wilcoxon rank sum test, paired).
Discussion
Based on the use of a limited panel of markers, our findings support the concept that squamous bladder cancer tends to be basal (low FOXA1, high CK14), while micropapillary cancer tends to be luminal (high FOXA1, low CK14). These findings are consistent with prior studies [8, 14]. The data also show that nested and plasmacytoid cancers tend to express high levels of FOXA1, suggesting that these variants exhibit a luminal characteristics. While more studies will be required to test this hypothesis, the current report is the first to suggest this possibility to the best of our knowledge.
This study also provides two pieces of evidence that precursors to micropapillary carcinoma and squamous carcinoma tend to be biologically different. First, micropapillary carcinoma tended to be mutually exclusive of squamous cell carcinoma, though this only approached statistical significance. Second, conventional UCs differed in FOXA1 expression depending on their associated histologic variant. That is, conventional UCs associated with squamous carcinoma tended to be FOXA1-low, while conventional UCs associated with micropapillary carcinoma tended to be FOXA1-high. The combined findings suggest that a luminal precursor gives rise to micropapillary carcinoma, while a basal precursor gives rise to squamous carcinoma. This would also keep with the findings that luminal cancers tend to arise with noninvasive papillary urothelial carcinoma [8] and that squamous carcinoma in situ of the bladder is often associated with invasive carcinoma with squamous differentiation [29]. It is also possible that the precursors are invasive conventional UC. Specifically, luminal conventional UC may give rise to micropapillary carcinoma, while basal conventional UC may give rise to squamous carcinoma.
We initially expected that CK14 expression would inversely correlate with FOXA1 expression, and would thus be overexpressed in both squamous bladder cancers and their associated conventional UCs. We did not observe this. Instead, we found that while squamous bladder cancers expressed higher CK14 than micropapillary, nested, and plasmacytoid bladder cancers, conventional UC associated with squamous cancer expressed similar CK14 levels to conventional UC associated with micropapillary cancer. Furthermore, cancer with squamous morphology expressed significantly higher CK14 than conventional UC found in the same bladder. This keeps with the previous finding that CK14 is a marker of squamous differentiation in transitional cell carcinoma [30], and indicates it is a marker of squamous differentiation, not a marker of a general basal subtype.
Bladder cancer is molecularly diverse, which our study highlights. Indeed, TCGA consortium showed invasive bladder cancer is among the most genetically diverse cancer types they have studied [31]. Although transcriptional profiling divides bladder cancers into broad luminal and basal subtypes, there is much diversity within these broad subtypes. For example, a study from Lund University named seven expression subtypes [12]. One subtype was called “squamous-like” and characteristically overexpressed CK14, similar to the squamous cancers in the present study. The cumulative findings thus suggest that expression profiles in bladder cancer fall into two large groups—luminal and basal—with transcriptional diversity in these large groups. By showing that CK14 is overexpressed in squamous cancer compared to conventional UC in the same bladder, we provide evidence that specific subtypes may evolve within the same bladder.
The findings from the present study raise many questions. First, while our findings suggest micropapillary cancers tend to arise from a precursor common to luminal conventional UC, it is unclear what drives this morphologic changes. Further work is needed to clarify this issue and to understand what drives its aggressive clinical behavior. Second, it is interesting that CK14 appeared to be an accurate marker of basal subtype per our analysis of TCGA data, yet was specific for squamous carcinoma in our immunohistochemical analysis. The TCGA study did not dissect tumor with squamous morphology from conventional morphology in their analysis, but instead included all tumors of all morphologies in the expression analysis [12]. It is thus probable that squamous morphology was common in their basal tumors, as squamous differentiation is present in up to 40% of invasive bladder cancers [1, 2]. Including such tumors may have driven the high CK14 expression seen in this study. Also, we detected protein expression, while the TCGA study detected RNA expression. This difference in targets may have contributed to our differing results.
Though we identified a small number of plasmacytoid cancers, their pattern of FOXA1 and CK14 expression was qualitatively different from the other cancer types. That is, half of our plasmacytoid cancers expressed high levels of both FOXA1 and CK14 (Fig. 2). This was uncommon in the other histologic variants. The finding suggests that plasmacytoid cancers may have unique transcriptional features, not easily classified by the luminal-basal dichotomy.
A weakness of the present study, and indeed a weakness of studying histologic variants in general, is distinction between conventional UC and histologic variants. These may often be admixed, and distinction may be subjective. In the present study, we attempted to ameliorate this problem by requiring that conventional UC and histologic variant are spatially distinct and appear qualitatively different. In conclusion, this study adds evidence that the biology underlying transcriptional subtypes relates to histologic variants of bladder cancer. Specifically, pathways of morphologic evolution tend to differ for luminal versus basal carcinoma. As several histologic variants are inherently aggressive, our findings inform on the clinical relevance of studying transcriptional subtypes.
References
Amin MB (2009) Histological variants of urothelial carcinoma: diagnostic, therapeutic and prognostic implications. Mod Pathol 22 Suppl 2:S96–S118. doi:10.1038/modpathol.2009.26
Grignon DJ, Al-Ahmadie H, Algaba F, Amin MB, Comperat E, Dryskjot L, Epstein JI, Hansel DE, Knuchel R, Lloreta J, Lopez-Beltran A, McKenney JK, Netto GJ, Paner G, Reuter VE, Shen SS, Van der Kwast T (2016) Infiltrating Urothelial Carcinoma. In: Moch H, Humphrey PA, Ulbright TM, Reuter VE (eds) WHO classificaiton of tumours of the urinary system and male genital organs, 4th edn. International Agencyfor Research on Cancer (IARC), Lyon
Kamat AM, Dinney CP, Gee JR, Grossman HB, Siefker-Radtke AO, Tamboli P, Detry MA, Robinson TL, Pisters LL (2007) Micropapillary bladder cancer: a review of the University of Texas M. D. Anderson Cancer Center experience with 100 consecutive patients. Cancer 110(1):62–67. doi:10.1002/cncr.22756
Kamat AM, Gee JR, Dinney CP, Grossman HB, Swanson DA, Millikan RE, Detry MA, Robinson TL, Pisters LL (2006) The case for early cystectomy in the treatment of nonmuscle invasive micropapillary bladder carcinoma. J Urol 175(3 Pt 1):881–885. doi:10.1016/S0022-5347(05)00423-4
Sui W, Matulay JT, James MB, Onyeji IC, Theofanides MC, RoyChoudhury A, DeCastro GJ, Wenske S (2016) Micropapillary bladder cancer: insights from the National Cancer Database. Bladder Cancer 2(4):415–423. doi:10.3233/blc-160066
Spaliviero M, Dalbagni G, Bochner BH, Poon BY, Huang H, Al-Ahmadie HA, Donahue TF, Taylor JM, Meeks JJ, Sjoberg DD, Donat SM, Reuter VE, Herr HW (2014) Clinical outcome of patients with T1 micropapillary urothelial carcinoma of the bladder. J Urol 192(3):702–707. doi:10.1016/j.juro.2014.02.2565
Gaya JM, Palou J, Algaba F, Arce J, Rodriguez-Faba O, Villavicencio H (2010) The case for conservative management in the treatment of patients with non-muscle-invasive micropapillary bladder carcinoma without carcinoma in situ. Can J Urol 17(5):5370–5376
Cancer Genome Atlas Research N (2014) Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507(7492):315–322. doi:10.1038/nature12965
Choi W, Porten S, Kim S, Willis D, Plimack ER, Hoffman-Censits J, Roth B, Cheng T, Tran M, Lee IL, Melquist J, Bondaruk J, Majewski T, Zhang S, Pretzsch S, Baggerly K, Siefker-Radtke A, Czerniak B, Dinney CP, McConkey DJ (2014) Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell 25(2):152–165. doi:10.1016/j.ccr.2014.01.009
Chou R, Selph SS, Buckley DI, Gustafson KS, Griffin JC, Grusing SE, Gore JL (2016) Treatment of muscle-invasive bladder cancer: a systematic review. Cancer 122(6):842–851. doi:10.1002/cncr.29843
Damrauer JS, Hoadley KA, Chism DD, Fan C, Tiganelli CJ, Wobker SE, Yeh JJ, Milowsky MI, Iyer G, Parker JS, Kim WY (2014) Intrinsic subtypes of high-grade bladder cancer reflect the hallmarks of breast cancer biology. Proc Natl Acad Sci U S A 111(8):3110–3115. doi:10.1073/pnas.1318376111
Sjodahl G, Lauss M, Lovgren K, Chebil G, Gudjonsson S, Veerla S, Patschan O, Aine M, Ferno M, Ringner M, Mansson W, Liedberg F, Lindgren D, Hoglund M (2012) A molecular taxonomy for urothelial carcinoma. Clin Cancer Res 18(12):3377–3386. doi:10.1158/1078-0432.CCR-12-0077-T
Choi W, Czerniak B, Ochoa A, Su X, Siefker-Radtke A, Dinney C, McConkey DJ (2014) Intrinsic basal and luminal subtypes of muscle-invasive bladder cancer. Nat Rev Urol 11(7):400–410. doi:10.1038/nrurol.2014.129
Guo CC, Dadhania V, Zhang L, Majewski T, Bondaruk J, Sykulski M, Wronowska W, Gambin A, Wang Y, Zhang S, Fuentes-Mattei E, Kamat AM, Dinney C, Siefker-Radtke A, Choi W, Baggerly KA, McConkey D, Weinstein JN, Czerniak B (2016) Gene expression profile of the clinically aggressive micropapillary variant of bladder cancer. Eur Urol 70(4):611–620. doi:10.1016/j.eururo.2016.02.056
Sangoi AR, Beck AH, Amin MB, Cheng L, Epstein JI, Hansel DE, Iczkowski KA, Lopez-Beltran A, Oliva E, Paner GP, Reuter VE, Ro JY, Shah RB, Shen SS, Tamboli P, McKenney JK (2010) Interobserver reproducibility in the diagnosis of invasive micropapillary carcinoma of the urinary tract among urologic pathologists. Am J Surg Pathol 34(9):1367–1376. doi:10.1097/PAS.0b013e3181ec86b3
Shah TS, Kaag M, Raman JD, Chan W, Tran T, Kunchala S, Shuman L, DeGraff DJ, Chen G, Warrick JI (2016) Clinical significance of prominent retraction clefts in invasive urothelial carcinoma. Hum Pathol. doi:10.1016/j.humpath.2016.10.021
Rebouissou S, Bernard-Pierrot I, de Reynies A, Lepage ML, Krucker C, Chapeaublanc E, Herault A, Kamoun A, Caillault A, Letouze E, Elarouci N, Neuzillet Y, Denoux Y, Molinie V, Vordos D, Laplanche A, Maille P, Soyeux P, Ofualuka K, Reyal F, Biton A, Sibony M, Paoletti X, Southgate J, Benhamou S, Lebret T, Allory Y, Radvanyi F (2014) EGFR as a potential therapeutic target for a subset of muscle-invasive bladder cancers presenting a basal-like phenotype. Sci Transl Med 6(244):244ra291. doi:10.1126/scitranslmed.3008970
Warrick JI, Walter V, Yamashita H, Chung E, Shuman L, Amponsa VO, Zheng Z, Chan W, Whitcomb TL, Yue F, Iyyanki T, Kawasawa YI, Kaag M, Guo W, Raman JD, Park JS, DeGraff DJ (2016) FOXA1, GATA3 and PPAR cooperate to drive luminal subtype in bladder cancer: a molecular analysis of established human cell lines. Sci Rep 6:38531. doi:10.1038/srep38531
Reddy OL, Cates JM, Gellert LL, Crist HS, Yang Z, Yamashita H, Taylor JA 3rd, Smith JA Jr, Chang SS, Cookson MS, You C, Barocas DA, Grabowska MM, Ye F, Wu XR, Yi Y, Matusik RJ, Kaestner KH, Clark PE, DeGraff DJ (2015) Loss of FOXA1 drives sexually dimorphic changes in urothelial differentiation and is an independent predictor of poor prognosis in bladder cancer. Am J Pathol 185(5):1385–1395. doi:10.1016/j.ajpath.2015.01.014
DeGraff DJ, Clark PE, Cates JM, Yamashita H, Robinson VL, Yu X, Smolkin ME, Chang SS, Cookson MS, Herrick MK, Shariat SF, Steinberg GD, Frierson HF, Wu XR, Theodorescu D, Matusik RJ (2012) Loss of the urothelial differentiation marker FOXA1 is associated with high grade, late stage bladder cancer and increased tumor proliferation. PLoS One 7(5):e36669. doi:10.1371/journal.pone.0036669
Raman JD, Warrick JI, Caruso C, Yang Z, Shuman L, Bruggeman RD, Shariat S, Karam JA, Wood C, Weizer AZ, Remzi M, Haitel A, Bensalah K, Rioux-Leclerq N, Bolenz C, Roscigno M, Krabbe LM, Kapur P, Lotan Y, Margulis V, DeGraff DJ (2016) Altered expression of the transcription factor forkhead box A1 (FOXA1) is associated with poor prognosis in urothelial carcinoma of the upper urinary tract. Urology 94(314):e311–e317. doi:10.1016/j.urology.2016.05.030
Yamashita H, Amponsa VO, Warrick JI, Zheng Z, Clark PE, Raman JD, Wu XR, Mendelsohn C, DeGraff DJ (2017) On a FOX hunt: functions of FOX transcriptional regulators in bladder cancer. Nat Rev Urol 14(2):98–106. doi:10.1038/nrurol.2016.239
Ho PL, Kurtova A, Chan KS (2012) Normal and neoplastic urothelial stem cells: getting to the root of the problem. Nat Rev Urol 9(10):583–594. doi:10.1038/nrurol.2012.142
Harvey JM, Clark GM, Osborne CK, Allred DC (1999) Estrogen receptor status by immunohistochemistry is superior to the ligand-binding assay for predicting response to adjuvant endocrine therapy in breast cancer. J Clin Oncol 17(5):1474–1481. doi:10.1200/JCO.1999.17.5.1474
Team RC (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Giraudoux P (2016) Pgirmess: data analysis in ecology. R package version 1.6.5. https://CRAN.R-project.org/package=pgirmess. Accessed 1 June 2016
Wickham H (2009) Ggplot2: elegant graphics for data analysis. Use R! Springer, New York
Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez JC, Muller M (2011) pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinf 12:77. doi:10.1186/1471-2105-12-77
Warrick JI, Kaag M, Raman JD, Chan W, Tran T, Kunchala S, DeGraff D, Chen G (2016) Squamous dysplasia of the urinary bladder: a consecutive cystectomy series. Int J Surg Pathol 24(4):306–314. doi:10.1177/1066896916629783
Harnden P, Southgate J (1997) Cytokeratin 14 as a marker of squamous differentiation in transitional cell carcinomas. J Clin Pathol 50(12):1032–1033
Hoadley KA, Yau C, Wolf DM, Cherniack AD, Tamborero D, Ng S, Leiserson MD, Niu B, McLellan MD, Uzunangelov V, Zhang J, Kandoth C, Akbani R, Shen H, Omberg L, Chu A, Margolin AA, Van't Veer LJ, Lopez-Bigas N, Laird PW, Raphael BJ, Ding L, Robertson AG, Byers LA, Mills GB, Weinstein JN, Van Waes C, Chen Z, Collisson EA, Cancer Genome Atlas Research N, Benz CC, Perou CM, Stuart JM (2014) Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 158(4):929–944. doi:10.1016/j.cell.2014.06.049
Acknowledgements
Creation of the bladder tissue microarray was supported by the Penn State Milton S. Hershey Medical Center Pathology Research Award Program.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Figure S1
(JPEG 128 kb)
Rights and permissions
About this article
Cite this article
Warrick, J.I., Kaag, M., Raman, J.D. et al. FOXA1 and CK14 as markers of luminal and basal subtypes in histologic variants of bladder cancer and their associated conventional urothelial carcinoma. Virchows Arch 471, 337–345 (2017). https://doi.org/10.1007/s00428-017-2190-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-017-2190-3