Abstract
Aldehyde dehydrogenase isoforms, ALDH1A1 and ALDH3A1, are associated with poor clinical outcome and resistance to chemotherapy in a wide variety of human malignancies. So far, their expression and prognostic significance in hepatocellular carcinoma (HCC) remains unknown. The aim of our study was to investigate their expression in HCC, and to correlate this to clinical, pathological and molecular features. ALDH1A1 and ALDH3A1 expression was first evaluated by microarray analysis in a series of 60 HCCs and five tumour-free liver tissue samples. Our findings related to ALDH3A1 were further validated by immunohistochemistry in a series of 81 HCCs and 23 hepatocellular adenomas (HCA). Microarray analysis showed no difference in ALDH1A1 expression between HCCs and tumour-free liver tissue. In contrast, ALDH3A1 was strongly upregulated in a subset of HCCs characterised by activation of the Wnt/ß-catenin pathway and CTNNB1 mutations. Using immunohistochemistry, we confirmed that high ALDH3A1 expression is associated with nuclear staining for ß-catenin and strong homogeneous staining for glutamine synthetase, two classical Wnt/ß-catenin pathway activation markers. Consistent with this finding, in tumour-free liver tissue, ALDH3A1 expression was observed in centrilobular hepatocytes, in which the Wnt/ß-catenin pathway is known to be physiologically activated. We also observed higher ALDH3A1 expression in CTNNB1-mutated HCA when compared with other subtypes. No correlation between ALDH3A1 expression and patient survival or tumour recurrence was observed.
In conclusion, ALDH3A1 is a marker of activation of the Wnt/ß-catenin pathway in HCC, HCA and tumour-free liver tissue. Further studies may help to elucidate the potential role of ALDH3A1 in HCC development and resistance to chemotherapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aldehyde dehydrogenase superfamily is a group of nicotinamide-adenine dinucleotide phosphate (NADPH+)-dependent enzymes that catalyse the oxidation of endogenous and exogenous aldehyde substrates to their corresponding carboxylic acids [1–3]. Endogenous aldehydes are generated during metabolism of amino acids, alcohol, lipids or vitamins, whilst exogenous aldehydes, whether intermediates or products, are derived from the metabolism of a wide variety of environmental agents or drugs (cigarette smoke, vehicle exhaust fumes, cytotoxic drugs) [1].
Expression of two ALDH isoforms, ALDH1A1 and ALDH3A1, and/or ALDH overall enzymatic activity has also been reported to represent cancer stem cell markers in various malignancies, including breast, lung, upper urinary tract and endometrioid adenocarcinoma [4–7]. Luo et al. [6] have demonstrated that ALDH1A1 silencing by siRNA or shRNA in melanoma cell lines leads to cell cycle arrest, apoptosis, and reduced tumourigenesis in vivo. Downregulation of expression of ALDH3A1 has been shown to affect cell growth, proliferation and motility in lung and hepatocellular carcinoma (HCC) cell lines [1, 8–10]. ALDH1A1 and/or ALDH3A1 expression also confers resistance to a wide variety of cytotoxic drugs, including gemcitabine, temozolomide and cyclophosphamide [1, 11–14].
Expression of ALDH1A1, which has been reported in numerous human tumours, is associated with poor clinical outcome in breast and lung adenocarcinoma [4, 5, 13]. So far, very few studies have investigated the expression of ALDH1A1 and ALDH3A1 in HCC [15], a malignancy associated with strong resistance to chemotherapy and adverse prognosis.
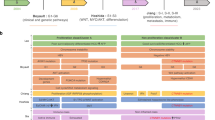
HCC is a heterogeneous disease for which we have formerly established a robust genomic and transcriptomic classification linked to different clinical and molecular features [16–18]. HCC groups G1 to G3 are characterised by overexpression of genes involved in cell and cycle proliferation, high rates of chromosomal instability, TP53 mutations (G2–G3) and HBV infection (G1–G2) [20]. HCC groups G4 to G6 are characterised by chromosomal stability. Groups G5 and G6 are strongly related to activating mutations of CTNNB1, encoding ß-catenin which is the most frequently mutated gene in HCC [17–19]. These mutations result in strong Wnt/ß-catenin pathway activation, which plays a critical role in hepatic carcinogenesis [17, 19].
We investigated ALDH1A1 and ALDH3A1 expression by gene expression microarray analysis in a series of 60 HCCs (‘microarray series’) and validated our results related to ALDH3A1 by immunohistochemistry in an additional series of 81 HCCs with available paraffin-embedded tissue (‘validation series’). ALDH1A1 and ALDH3A1 expression were correlated to HCC transcriptomic subgroups and CTNNB1-activating mutations. In the validation series, we compared immunohistochemical expression of ALDH3A1 to that of the Wnt/ß-catenin pathway activation markers nuclear ß-catenin staining and diffuse strong staining for glutamine synthetase (GS). Finally, we investigated ALDH3A1 expression in tumour-free liver tissue and in a series of 23 hepatocellular adenomas (HCA), and correlated this to patient prognosis and tumour reccurence.
Material and methods
Patients and samples
For the microarray series, 60 frozen tumour samples and 5 nontumour liver samples were collected from patients surgically treated in three French departments of surgery from 1992 to 1999, as previously described [16].
For the validation series, 81 HCCs cases treated by surgical resection were retrieved from the archives of the Departments of Pathology of Henri Mondor (Créteil, France) and Bordeaux (France) University Hospitals. As required by French law, informed consent was obtained from all patients. The following items were systematically recorded: age (< or ≥60 years), sex, liver disease etiology, follow-up (n = 78, mean 33 months), tumour size (< or ≥50 mm), Edmondson grade and vascular invasion.
Main clinical and pathological features are listed in Table 1.
The 23 HCA cases included six CTNNB1 mutated, ten inflammatory, five HNF1A-inactivated and two unclassified HCAs, according to molecular classification [20–23] (Table 2).
Immunohistochemistry
All immunostainings were performed on whole sections. For ALDH3A1, after deparaffinisation and rehydratation, the sections were placed into a boiling target retrieval solution (DiaPath Citrate Buffer pH 6) for 40 min, and left to cool down without cover for 20 min. Slides were then processed on an automated immunostainer (Dako AutoStainer). Endogenous peroxidase was blocked with H2O2 (10 min) and nonspecific background staining with goat serum (20 min). Sections were further incubated with a primary anti-ALDH3A1 (Santa Cruz Biotechnology, clone B-8; sc-137168, dilution 1/200) antibody for 1 h at room temperature and washed with phosphate-buffered saline solution. After incubation with the Envision + System-labelled polymer–horseradish peroxidase (HRP) kit (Dako/Cytomation) and staining with 3-diaminobenzidine (Dako) as chromogen, the slides were counterstained with Mayer's haematoxylin, dehydrated and coverslipped. Samples with at least 50 % of stained tumour cells were classified ‘ALDH3A1 high’; samples with less than 50 % of stained tumour cells were classified ‘ALDH3A1 low’.
Immunohistochemical staining for GS (BD Biosciences-Pharmigen, ref 610517, dilution 1/1,000) and ß-catenin (BD Biosciences-Pharmigen, ref 610153, dilution 1/4,000) was performed on an automated immunostainer (Leica Bond-Max), according to the manufacturer's instructions. GS expression was scored as follows: 0 (no staining), 1 (focal or diffuse and heterogeneous staining) and 2 (diffuse and strong staining). For ß-catenin, nuclear staining was considered as positive if at least one tumour cell had a stained nucleus.
Mutation screening
Mutation status for CTNNB1 had been performed previously for the microarray series [16]. For the validation series, all HCC samples were sequenced for CTNNB1 (exons 2 to 4), using the same method [16]. All mutations were confirmed by sequencing a second independent amplification product on both strands.
Statistical analysis
Differences in microarray expression of ALDH1A1 and ALDH3A1 between tumour subgroups were assessed using Student's t test. Correlations between immunohistochemical scores, clinical and molecular data was analysed using Fisher's exact test. Kaplan–Meier disease-specific survival and tumour recurrence analysis was performed and illustrated using GraphPad Prism 5 software and survival curve comparison analyses were performed using the log–rank (Mantel–Cox) test. p Values < 0.05 were considered statistically significant.
Results
ALDH1A1 and ALDH3A1 expression by microarray
Expression of ALDH1A1 by gene expression array in HCC samples was not significantly different from that in tumour-free liver tissue samples (n = 5; mean intensity 2,352 vs. 2,420, p = 0.9). ALDH1A1 expression was lower in G1 HCC than in tumour-free liver tissue (mean intensity 1,100 vs. 2,352, p = 0.02); no other significant difference was observed between ALDH1A1 expression in other subgroups and nontumour livers (data not shown).
Mean expression of ALDH3A1 was higher in HCC samples than in tumour-free liver tissue, but this was not statistically significant (mean = 363 vs. 68, p = 0.37). ALDH3A1 was strongly over-expressed in G5 and G6 subgroups of HCC (p < 0.001; Fig. 1a). ALDH3A1 expression was not correlated with other clinical or pathological parameters.
ALDH3A1, GS and ß-catenin expression in the validation series
As most G5 and G6 HCC carry an activating CTNNB1 mutation, we investigated ALDH3A1 expression in relation to CTNNB1 mutational status in a separate set of tumours (validation set) including 81 HCC treated by resection in Créteil and Bordeaux hospitals. Immunohistochemical expression of ALDH3A1 was negative in 31 cases (39 %), low in 17 cases (21 %) and high in 33 cases (40 %; Fig. 2a). Both cytoplasmic and nuclear staining of ALDH3A1 was observed (Fig. 2b). ALDH3A1 expression was not significantly correlated with any clinical or pathological features.
In 28 cases (34 %), we identified a somatic activating CTNNB1 mutation (Table 3). High ALDH3A1 expression strongly correlated with CTNNB1 mutations (p < 0.0001). Sensitivity and specificity of high ALDH3A1 expression for prediction of CTNNB1 mutation were 71 % (20/28) and 75 % (40/53), respectively. Glutamine synthase over-expression (score 2), which is a reliable marker of ß-catenin activation in hepatocytes, was observed in 45 % of cases and significantly associated with CTNNB1 mutation with a 78 % (22/28) sensitivity and 73 % (39/53) specificity (Fig. 3). Nuclear ß-catenin staining, a sign of ß-catenin activation, was observed in 20 out of 81 cases (24 %) with a sensitivity and specificity for predicting CTNNB1 mutations of 53 % (15/28) and 89 % (47/53), respectively (Fig. 3).
A strong positive correlation was observed between high ALDH3A1 expression and score 2 GS expression (p < 0.0001) and nuclear ß-catenin staining (p < 0.0001). Of the 12 tumours with high ALDH3A1 expression but CTNNB1 wild type, eight cases had a GS score of 2, and five cases also featured nuclear ß-catenin staining. No correlation was observed between ALDH3A1 expression and other clinical or pathological parameters.
Expression of ALDH3A1 was further investigated in 23 HCA including six CTNNB1 mutated, five HNF1A inactivated, ten inflammatory and two unclassified HCAs. We observed a higher percentage of positive cells in the CTNNB1-mutated HCA than in other subtypes (p = 0.03; Fig. 4). However, the total number of positive hepatocytes was lower in CTNNB1-mutated HCA than in mutated HCC, and two CTNNB1-mutated HCA were ALDH3A1-negative. Finally, no significant differences were observed between inflammatory, unclassified or HNF1A inactivated HCA.
ALDH3A1 expression in nontumour liver
Next, we analysed ALDH3A1 expression in tumour-free liver tissue samples remote from HCC in 71 cases (METAVIR F0 n = 3; F1 n = 13; F2 n = 13; F3 n = 29; F4 n = 23). In liver tissue without extensive fibrosis (F0–F1; n = 16), five cases featured striking centrilobular expression (Fig. 5a, b), ten cases did not show any ALDH3A1 expression, and one case exhibited few clusters of positive cells without a particular pattern. In liver tissue with extensive fibrosis (F2 to F4), we observed a few cell clusters expressing ALDH3A1, most frequently located around fibrotic septa.
ALDH3A1 expression, disease specific survival rate and tumour recurrence
There were no significant differences in disease-specific survival according to the level of ALDH3A1 expression (p = 0.53, Fig. 6a). A nonsignificant trend towards earlier tumour recurrence in ALDH3A1 high tumours was observed (p = 0.06, Fig. 6b).
Discussion
In several reports, ALDH1A1 and ALDH3A1 have been associated with resistance to chemotherapy and adverse clinical outcome in a wide variety of human malignancies. We present a large study on the expression of ALDH1A1 and ALDH3A1 in a series of human HCCs. We show that ALDH3A1 expression is upregulated in a subset of HCCs with CTNNB1 mutations activating the Wnt/ß-catenin pathway. In the microarray series of 60 samples analysed for mRNA expression and in the validation set of 81 HCCs studied by immunochemistry, our results show that ALDH3A1 over-expression and CTNNB1 mutations are strongly associated in HCC. Moreover, high ALDH3A1 expression by immunohistochemistry had a sensitivity and specificity for predicting CTNNB1 mutations comparable to that of other classical immunohistochemical markers (ß-catenin nuclear staining and GS over-expression). Interestingly, of the 12 cases with high ALDH3A1 expression but no CTNNB1 mutation, eight cases had a GS score of 2, and five cases also featured nuclear ß-catenin staining. These observations suggest that in those cases high ALDH3A1 expression reflects activation of the Wnt/ß-catenin pathway caused by a different mechanism.
Consistent with our findings in HCC, percentage of neoplastic cells expressing ALDH3A1 was higher in CTNNB1-mutated HCA, even if only one mutated case was classified as ‘ALDH3A1 high’, with two cases being completely negative. This finding casts doubt on the use of ALDH3A1 expression as marker of CTNNB1 mutation in HCA.
The Wnt/ß-catenin pathway plays a key role in liver zonation and differentiation [19, 24] and is strongly activated in centrilobular areas [19]. The striking centrilobular pattern of ALDH3A1 expression observed in nonfibrotic tumour-free liver tissue is also consistent with a physiological link between ALDH3A1 and activation of the Wnt/ß-catenin pathway. Functional studies are needed to confirm that ALDH3A1 transcription is directly or indirectly controlled by the Wnt/ß-catenin pathway.
Overall ALDH enzymatic activity, which partly relies on expression of ALDH1A1 and ALDH3A1, is considered as a reliable marker of cancer stem cells [1, 5, 25–27]. In our series, CTNNB1-mutated HCC that also overexpressed ALDH3A1 were well differentiated, with intra-tumour cholestasis. As reported before, these HCC often developed in a noncirrhotic liver [28]. In our series, 31 HCC did not express ALDH3A1, but in 33 cases, more than 50 % of the cells were positive. As cancer stem cells presumably represent a very small proportion of tumour cells, this finding might argue against the validity of ALDH3A1 expression as a marker of cancer stem cells in HCC.
In view of their detoxifying functions, ALDH isoforms are considered to be involved in tumour resistance to various anticancer drugs [11, 13, 14, 29, 30]. Hu et al. [31] reported that ALDH3A1 knock down enhances sensitivity to paclitaxel, doxorubicin and 4-hydroxycyclophosphamide in a breast cancer cell line. Such experiments in HCC, modulating ALDH3A1 expression, might clarify whether or not ALDH3A1 contributes to strong resistance to chemotherapy, in particular, in CTNNB1-mutated HCC.
We found a trend towards an association of high ALDH3A1 expression with early recurrence, which suggests that ALDH3A1 expression might be a reflection of tumour aggressiveness. In view of the on-going discussion on the relationship between WNT/ß-catenin pathway activation and survival [32–36], ALDH3A1 expression might play a role in assessment of prognosis but this needs to be further validated in a larger series of patients.
In conclusion, we show that ALDH3A1 expression is strongly upregulated in a subset of HCC with CTNNB1 mutations activating the Wnt/ß-catenin pathway. Further studies will be needed to elucidate the potential role of ALDH3A1 in HCC development and resistance to chemotherapy.
References
Muzio G, Maggiora M, Paiuzzi E et al (2012) Aldehyde dehydrogenases and cell proliferation. Free Radic Biol Med 52:735–746. doi:10.1016/j.freeradbiomed.2011.11.033
Chen Y, Thompson DC, Koppaka V et al (2012) Ocular aldehyde dehydrogenases: rotection against ultraviolet damage and maintenance of transparency for vision. Prog Retin Eye Res 33:28–39. doi:10.1016/j.preteyeres.2012.10.001
Kiefer FW, Orasanu G, Nallamshetty S et al (2012) Retinaldehyde dehydrogenase 1 coordinates hepatic gluconeogenesis and lipid metabolism. Endocrinology 153:3089–3099. doi:10.1210/en.2011-2104
Charafe-Jauffret E, Ginestier C, Iovino F et al (2010) Aldehyde dehydrogenase 1-positive cancer stem cells mediate metastasis and poor clinical outcome in inflammatory breast cancer. Clin Cancer Res 16:45–55. doi:10.1158/1078-0432.CCR-09-1630
Li X, Wan L, Geng J et al (2012) Aldehyde dehydrogenase 1A1 possesses stem-like properties and predicts lung cancer patient outcome. J Thorac Oncol 7:1235–1245. doi:10.1097/JTO.0b013e318257cc6d
Luo Y, Dallaglio K, Chen Y et al (2012) ALDH1A isozymes are markers of human melanoma stem cells and potential therapeutic targets. Stem Cells 30:2100–2113. doi:10.1002/stem.1193
Visvader JE, Lindeman GJ (2008) Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nat Rev Cancer 8:755–768
Moreb JS, Baker HV, Chang L-J et al (2008) ALDH isozymes downregulation affects cell growth, cell motility and gene expression in lung cancer cells. Mol Cancer 7:87. doi:10.1186/1476-4598-7-87
Muzio G, Trombetta A, Maggiora M et al (2006) Arachidonic acid suppresses growth of human lung tumor A549 cells through down-regulation of ALDH3A1 expression. Free Radic Biol Med 40:1929–1938. doi:10.1016/j.freeradbiomed.2006.01.020
Muzio G, Canuto RA, Trombetta A, Maggiora M (2001) Inhibition of cytosolic class 3 aldehyde dehydrogenase by antisense oligonucleotides in rat hepatoma cells. Chem Biol Interact 130–132:219–225
Duong H-Q, Hwang JS, Kim HJ et al (2012) Aldehyde dehydrogenase 1A1 confers intrinsic and acquired resistance to gemcitabine in human pancreatic adenocarcinoma MIA PaCa-2 cells. Int J Oncol 41:855–861. doi:10.3892/ijo.2012.1516
Huang C-P, Tsai M-F, Chang T-H et al (2013) ALDH-positive lung cancer stem cells confer resistance to epidermal growth factor receptor tyrosine kinase inhibitors. Cancer Lett 328:144–151. doi:10.1016/j.canlet.2012.08.021
Khoury T, Ademuyiwa FO, Chandrasekhar R et al (2012) Aldehyde dehydrogenase 1A1 expression in breast cancer is associated with stage, triple negativity, and outcome to neoadjuvant chemotherapy. Mod Pathol 25:388–397. doi:10.1038/modpathol.2011.172
Schäfer A, Teufel J, Ringel F et al (2012) Aldehyde dehydrogenase 1A1–a new mediator of resistance to temozolomide in glioblastoma. Neuro-oncology 14:1452–1464. doi:10.1093/neuonc/nos270
Shibuya A, Takeuchi A, Shibata H et al (1994) Immunohistochemical study of hepatocellular carcinoma-specific aldehyde dehydrogenase. Alcohol Alcohol Suppl 29:119–123
Boyault S, Rickman DS, de Reyniès A et al (2007) Transcriptome classification of HCC is related to gene alterations and to new therapeutic targets. Hepatology 45:42–52. doi:10.1002/hep.21467
Nault J-C, Zucman-Rossi J (2011) Genetics of hepatobiliary carcinogenesis. Semin Liver Dis 31:173–187. doi:10.1055/s-0031-1276646
Guichard C, Amaddeo G, Imbeaud S et al (2012) Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat Genet 44:694–698. doi:10.1038/ng.2256
Thompson MD, Monga SPS (2007) WNT/beta-catenin signaling in liver health and disease. Hepatology 45:1298–1305. doi:10.1002/hep.21651
Zucman-Rossi J, Jeannot E, Nhieu JTV et al (2006) Genotype-phenotype correlation in hepatocellular adenoma: new classification and relationship with HCC. Hepatology 43:515–524. doi:10.1002/hep.21068
Nault J-C, Bioulac-Sage P, Zucman-Rossi J (2013) Hepatocellular benign tumors-from molecular classification to personalized clinical care. Gastroenterology 144:888–902. doi:10.1053/j.gastro.2013.02.032
Nault JC, Fabre M, Couchy G et al (2011) GNAS-activating mutations define a rare subgroup of inflammatory liver tumors characterized by STAT3 activation. J Hepatol. doi:10.1016/j.jhep.2011.07.018
Calderaro J, Labrune P, Morcrette G et al (2012) Molecular characterization of hepatocellular adenomas developed in patients with glycogen storage disease type I. J Hepatol. doi:10.1016/j.jhep.2012.09.030
Decaens T, Godard C, de Reyniès A et al (2008) Stabilization of beta-catenin affects mouse embryonic liver growth and hepatoblast fate. Hepatology 47:247–258. doi:10.1002/hep.21952
Huang EH, Hynes MJ, Zhang T et al (2009) Aldehyde dehydrogenase 1 is a marker for normal and malignant human colonic stem cells (SC) and tracks SC overpopulation during colon tumorigenesis. Cancer Res 69:3382–3389. doi:10.1158/0008-5472.CAN-08-4418
Sullivan JP, Spinola M, Dodge M et al (2010) Aldehyde dehydrogenase activity selects for lung adenocarcinoma stem cells dependent on notch signaling. Cancer Res 70:9937–9948. doi:10.1158/0008-5472.CAN-10-0881
Fleischman AG (2012) ALDH marks leukemia stem cell. Blood 119:3376–3377. doi:10.1182/blood-2012-02-406751
Audard V, Grimber G, Elie C et al (2007) Cholestasis is a marker for hepatocellular carcinomas displaying beta-catenin mutations. J Pathol 212:345–352. doi:10.1002/path.2169
Sládek NE, Kollander R, Sreerama L, Kiang DT (2002) Cellular levels of aldehyde dehydrogenases (ALDH1A1 and ALDH3A1) as predictors of therapeutic responses to cyclophosphamide-based chemotherapy of breast cancer: a retrospective study. Rational individualization of oxazaphosphorine-based cancer chemotherapeutic regimens. Cancer Chemother Pharmacol 49:309–321. doi:10.1007/s00280-001-0412-4
Moreb JS, Mohuczy D, Muhoczy D et al (2007) RNAi-mediated knockdown of aldehyde dehydrogenase class-1A1 and class-3A1 is specific and reveals that each contributes equally to the resistance against 4-hydroperoxycyclophosphamide. Cancer Chemother Pharmacol 59:127–136. doi:10.1007/s00280-006-0233-6
Hu G, Chong RA, Yang Q et al (2009) MTDH activation by 8q22 genomic gain promotes chemoresistance and metastasis of poor-prognosis breast cancer. Cancer Cell 15:9–20. doi:10.1016/j.ccr.2008.11.013
Nault J-C, De Reyniès A, Villanueva A et al (2013) A hepatocellular carcinoma 5-gene score associated with survival of patients after liver resection. Gastroenterology 145:176–187. doi:10.1053/j.gastro.2013.03.051
Hsu HC, Jeng YM, Mao TL et al (2000) Beta-catenin mutations are associated with a subset of low-stage hepatocellular carcinoma negative for hepatitis B virus and with favorable prognosis. Am J Pathol 157:763–770
Hoshida Y, Toffanin S, Lachenmayer A et al (2010) Molecular classification and novel targets in hepatocellular carcinoma: recent advancements. Semin Liver Dis 30:35–51. doi:10.1055/s-0030-1247131
Villanueva A, Hoshida Y, Battiston C et al (2011) Combining clinical, pathology, and gene expression data to predict recurrence of hepatocellular carcinoma. Gastroenterology 140:1501–1512.e2. doi:10.1053/j.gastro.2011.02.006
Lachenmayer A, Alsinet C, Savic R et al (2012) Wnt-pathway activation in two molecular classes of hepatocellular carcinoma and experimental modulation by sorafenib. Clin Cancer Res 18:4997–5007. doi:10.1158/1078-0432.CCR-11-2322
Acknowledgements
Authors warmly thank Tumourotheque/Plateforme des Ressources Biologiques of Henri Mondor University Hospital and Réseau des CRB Foie-Inserm. This project was funded by Inserm, the INCA HCC-PAIR project and the ARC 5194 grant.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Calderaro, J., Nault, JC., Bioulac-Sage, P. et al. ALDH3A1 is overexpressed in a subset of hepatocellular carcinoma characterised by activation of the Wnt/ß-catenin pathway. Virchows Arch 464, 53–60 (2014). https://doi.org/10.1007/s00428-013-1515-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-013-1515-0