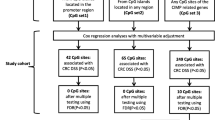
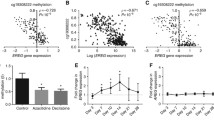
Abstract
CpG island methylator phenotype (CIMP) refers to a subset of colorectal cancers (CRCs) that are characterized by concordant hypermethylation of multiple CpG island loci. CIMP+ CRCs have peculiar clinicopathological features. However, controversy exists over prognostic implications of CIMP in CRCs. We analyzed 320 cases of CRCs for their CIMP status using the MethyLight assay and determined clinicopathological features and prognostic implications of CIMP alone or in combination with microsatellite instability (MSI). With methylation of five or more markers among eight markers examined, CIMP+ tumors were significantly associated with female gender, proximal tumor location, poor differentiation, nodal metastasis, more advanced cancer, BRAF mutations, MSI, and poor prognosis (all P values <0.05). Ogino’s combined eight-marker panel outperformed the Ogino and the Laird five-marker panels in detecting these features. Of the four molecular subtypes generated by the combination of CIMP and MSI status, the CIMP+/MSI− subtype showed the worst clinical outcome (P = 0.0003). However, poor prognosis of CIMP+/MSI− subtype was found to be attributed to BRAF mutation. In conclusion, the CIMP+/MSI− subtype tends to present with distinct clinicopathological and molecular features and shows the worst clinical outcome among the four molecular subtypes of CRCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epigenetics describes transmission through cell division of heritable changes in phenotype that do not involve DNA sequence changes. The underlying mechanisms for epigenetic transmission include DNA methylation, histone modification, and transmitted chromatin structure. Among these, the alteration of DNA methylation patterns is known to be a key component for altered gene expression associated with human cancers. Promoter CpG island hypermethylation is found in virtually all tissue types of human cancers and acts as an important mechanism for inactivation of tumor suppressor genes and tumor-related genes [1, 2]. Promoter CpG island hypermethylation and its associated histone modifications render the chromatin structure of a gene promoter into a closed compact structure inaccessible to transcription factors, which results in the inactivation of gene transcription [3]. Human cancer cells have both genetic changes and epigenetic changes in tumor-related genes. Recent studies have demonstrated that promoter CpG island hypermethylation is more frequent than genetic changes in human colorectal cancers [4, 5], which suggests that promoter CpG island hypermethylation is a potential mechanism of colorectal carcinogenesis.
In addition to two known molecular pathways in colorectal carcinogenesis, which involve chromosomal instability (CIN) and microsatellite instability (MSI), a third epigenetic instability pathway has been proposed by Dr. Issa’s group [6]. The CpG island methylator phenotype (CIMP) refers to a subset of colorectal cancers (CRCs) that occur through the epigenetic instability pathway. CIMP+ CRCs are characterized by widespread hypermethylation of promoter CpG island loci, which results in the inactivation of the involved genes. In the study of Weisenberger et al. [7], the presence of CIMP+ CRCs has been well documented; these tumors are distinct from CIMP− CRCs because of the higher methylation frequencies or higher methylation levels of the examined CpG island loci. A growing number of studies has consistently demonstrated close associations between CIMP+ CRCs and proximal colon location, MSI, and a high frequency of BRAF mutation, regardless of the methodology and CIMP marker panels used [7–11]. However, a marked controversy exists over the prognostic implications of CIMP: Some studies suggest an adverse effect of CIMP on survival of CRC patients [12–14], whereas other studies suggest little prognostic value of CIMP [15]. Furthermore, Ogino et al. [16] reported that CIMP was an independent predictor of good prognosis in colon cancers, which is contrary to previous findings. These differences may be related to differences in the methodology and CIMP marker panel used to determine CIMP status in these studies. The association of good prognosis with CIMP+ tumors was produced using MethyLight technology, whereas poor prognosis in CIMP+ tumors was seen using methylation-specific polymerase chain reaction (PCR) or combined bisulfite restriction analysis. Except for the study by Ogino et al. [16], no data are available regarding the prognostic implications of CIMP+ CRCs determined by MethyLight technology.
In the present study, we used MethyLight technology and analyzed 320 CRC cases for CIMP and MSI status and characterized the clinicopathological and molecular features of the CIMP+ CRCs. We then assessed the independent effect of CIMP and the combinatorial effect of CIMP and MSI on patient outcome.
Materials and methods
Tissue samples
Formalin-fixed, paraffin-embedded archival tissues from 320 CRC patients were retrieved from the Department of Pathology, Seoul National University Hospital (Seoul, Korea). These patients had undergone curative surgery at Seoul National University Hospital between 1999 and 2002. The selection was solely on the availability of archival tissue blocks for the study, and we did not exclude patients with a family history of CRCs. However, CRC patients treated with neoadjuvant therapy were excluded from the study. Clinicopathologic information including age, sex, histological differentiation, tumor location, tumor stage, and overall survival were obtained from these 320 patients. Tumor staging was based on the pTNM staging system of the American Joint Committee on Cancer (AJCC). The histological differentiation was determined using the World Health Organization criteria. This study was approved by the Institutional Review Board.
DNA extraction and bisulfite modification
Through light microscopic examination, we marked tumor areas where tumor cells occupied 50% or more of all cells and represented the main histology and differentiation of the tumor. Non-tumorous portions in matched CRC were obtained from normal colorectal mucosa and were confirmed by microscopy to be tumor free. Ten serial 10-μm-thick histologic slides of formalin-fixed tumor and normal tissue blocks were used for manual microdissection. Dissected tissue samples were subjected to tissue lysis using proteinase K lysis buffer. Sodium bisulfite conversion of genomic DNA was performed as described [17].
DNA methylation analysis
DNA methylation analyses were performed using MethyLight as previously described [18]. We quantified DNA methylation in eight CIMP markers—CACNA1G, CDKN2A (p16), CRABP1, IGF2, MLH1, NEUROG1, RUNX3, and SOCS1. The oligonucleotide sequences of the primers and probes used have been described [7]. The PCR conditions were as follows: initial denaturation at 95°C for 10 min followed by 50 cycles of 95°C for 15 s and 60°C for 1 min. M.SssI-treated genomic DNA was used as a reference sample for complete methylation to determine the percentage of fully methylated alleles (percentage of methylated reference (PMR)) at a particular locus. The PMR value was calculated by dividing the GENE/ALU ratio of a sample by the GENE/ALU ratio of the M.SssI-treated human genomic DNA sample and multiplying by 100 [19]. We considered a CpG island locus methylated if the PMR value was >4.
Optimal panel for CIMP determination
For the MethyLight-based determination of CIMP, three kinds of CIMP marker panels were available: Dr. Ogino’s original five-marker panel (CACNA1G, CRABP1, MLH1, NEUROG1, and p16) [8], Dr. Laird’s five-marker panel (CACNA1G, IGF2, NEUROG1, RUNX3, and SOCS1) [7], and Dr. Ogino’s combined eight-marker panel (CACNA1G, CRABP1, IGF2, MLH1, NEUROG1, p16, RUNX3, and SOCS1) [21]. To determine which CIMP marker panel was optimal for CIMP diagnosis, the three CIMP marker panels were screened against 196 CRC cases. For the five-marker panels, CRC cases were considered CIMP+ if at least three of the markers were methylated; for the combined eight-marker panel, cutoff values of five and six markers were each tested for CIMP determination. CIMP+ tumors defined by the three CIMP marker panels were compared regarding their associations with previously known clinicopathological features of CIMP+ CRCs, including poor prognosis, older age, female predominance, proximal colon location, poor differentiation, high frequency of BRAF mutations, and high frequency of MSI. Overall, the combined eight-marker panel with a cutoff value of five outperformed the five-marker panels and Dr. Ogino’s combined eight-marker panel with a cutoff value of six in most comparisons based on the accuracy of association (Fig. 1 and Supplementary Table 1). Thus, we used the combined eight-marker panel with a cutoff value of five for subsequent determination of CIMP.
Summary of comparative analysis among CIMP marker panels. Red bars indicate the presence of methylation in CIMP marker columns, and gray bars indicate CIMP+ status in four differently defined CIMP columns. Blue bars indicate older age (>61 years) in the age column, female in the gender column, proximal colon in the location column, poor differentiation in the differentiation column, higher stage (III, IV) in the stage column, MSI+ status in the MSI column, presence of mutant forms of KRAS in the KRAS column, and presence of mutant forms of BRAF in the BRAF column
Mutation analysis of KRAS codons 12 and 13 and BRAF codon 600
KRAS codon 12 and 13 mutations were assayed by PCR–restriction fragment length polymorphism (RFLP) analysis, and suspected cases were confirmed by direct sequencing of the KRAS gene [20]. BRAF mutations were assayed using PCR–RFLP analysis and confirmatory sequencing as described [20].
Microsatellite analysis
The MSI status of each tumor was determined based on an examination of microsatellite markers (D2S123, D5S346, D17S250, BAT25, and BAT26). We classified MSI status as follows: MSI+ tumors had instability at two or more microsatellite markers, and MSI− tumors had instability at no more than one marker.
Statistical analysis
All statistical analyses were conducted using SPSS software version 12.0 (SPSS, Inc., Chicago, IL, USA). Pearson’s chi-square test was used to test associations between clinicopathologic or molecular variables and CIMP+ tumors defined by different CIMP marker panels. Pearson’s chi-square test was used to compare the frequency of each clinicopathological or genetic parameter in CRCs by four molecular subtypes. To compare the means of numeric variables among three or more groups, an analysis of variance test was used. Survival was measured from the date of resection of CRC to the date of death or the last clinical review before August 29, 2007. The average follow-up time (from surgery to death or the last follow-up) was 63.5 months (range, 1–99 months). Crude overall survival rates were assessed using the Kaplan–Meier log-rank test, and using Cox proportional-hazards regression models, multivariate analysis was performed in a backward manner to determine association between survival and CIMP, adjusting for variable suggested to be prognostic factors on univariate analysis. Hazard ratios and 95% confidence interval (95% CI) for death were computed using Cox survival modeling. All reported P values are two-sided, and P values of less than 0.05 were considered to indicate significance.
Results
Distinct clinicopathological features of the CIMP+/MSI− subtype
To study clinicopathological features of CIMP and MSI, we analyzed an additional 124 CRC cases, giving a total of 320 cases that were classified into four molecular subtypes (CIMP+/MSI+, CIMP+/MSI−, CIMP−/MSI+, and CIMP−/MSI−) using the eight-marker panel with a cutoff of five. The CIMP−/MSI− subtype was the most common, comprising 78.8% of CRCs, whereas the CIMP+/MSI+ subtype was the least common, comprising 3.8% of CRCs (Table 1). When these four molecular subtypes were compared regarding their associations with several clinicopathological features, the CIMP+/MSI− subtype (7.8%) showed close associations with proximal colon location, frequent poor differentiation, frequent nodal metastasis, frequent distant metastasis, high cancer stage, high frequency of BRAF mutation, and low frequency of KRAS mutation (Table 1). In survival analysis, the CIMP+/MSI− subtype showed the worst clinical outcome, whereas the CIMP−/MSI+ subtype showed the best clinical outcome (Fig. 2). These differences were statistically significant (Kaplan–Meier log-rank test, P < 0.001). These results suggest that CIMP+/MSI− CRCs have distinct clinicopathological features.
The relationship between the degree of methylation and survival
A close association of CIMP+ with poor prognosis for MSI− CRC patients was still found after introduction, in the survival analysis, of CIMP-low group and resultant separation of CIMP− CRCs into CIMP-0 (lack of methylation in eight markers) and CIMP-low (with one to four methylated markers; Fig. 3). CIMP-low CRC patients showed clinical outcomes better than those of CIMP+ (or CIMP-high) CRC patients but worse than those of CIMP-0 CRC patients. When clinicopathological or molecular features were correlated with methylation status in MSI− CRCs, CIMP-low CRCs were similar to CIMP-0 CRCs in the examined features except for KRAS mutation (Table 2). KRAS mutation was more common in CIMP-low CRCs (45%) than in CIMP-0 (24%) or CIMP-high (14%) CRCs (P = 0.001).
The effect of BRAF mutation on the prognostic effect of CIMP
In the present study, MSI− CRCs showed significantly different clinical outcomes depending on CIMP status, and particularly, the clinical outcomes of CIMP+/MSI− CRCs were worse than those of CIMP−/MSI− CRCs (P < 0.001). In order to identify whether the prognostic effect of CIMP was related to the presence of BRAF mutation, we stratified MSI− CRCs into four subtypes according to CIMP and BRAF status: CIMP+/BRAF+, CIMP+/BRAF−, CIMP−/BRAF+, and CIMP−/BRAF−. CIMP+/BRAF+ CRCs showed worse clinical outcome than that of CIMP+/BRAF− CRCs, which was similar to that of CIMP−/BRAF− CRCs. This result clearly indicates that the worse clinical outcome of CIMP+/MSI− CRCs was clearly associated with the presence of BRAF mutation (Fig. 4).
Overall survival according to CIMP, MSI, and mutations of KRAS and BRAF
Survival was analyzed in 318 patients; two patients were excluded because of loss to follow-up. Univariate relationships between clinicopathogical or molecular factors and the overall survival are summarized in Table 3. The factors showing significant associations with poor survival were poor differentiation, high cancer stage (AJCC), and older age (>61 years), whereas the factor significantly associated with good prognosis was MSI+ status. However, CIMP+ status was not a statistically significant factor. When we separated our study cases into colon cancer group and rectal cancer group (Table 4 and Supplementary Table 2), CIMP+ status was a significant factor in colon cancer group. For colon cancer group, besides CIMP+ status, older age, high cancer stage, MSI+ status, and BRAF mutation were significant factor associated with prognosis of colon cancer patients and differentiation was marginally significant. To determine association between CIMP and survival, multivariate analysis was performed with adjustment for above five variables, revealing that CIMP+ status was not an independent prognostic factor for colon cancers.
Discussion
MSI and CIN are two important mechanisms of genetic instability in CRCs, but they cannot account for all CRCs because a proportion of CRCs have neither MSI nor CIN [22, 23]. A potential mechanism capable of filling the gap is CIMP, which is characterized by widespread hypermethylation of multiple promoter CpG island loci. CIMP has now been recognized as a potential alternative mechanism for genetic instability driving molecular diversity in CRCs. CIMP appears to overlap MSI, because a considerable proportion of sporadic MSI+ CRCs arise as a consequence of CIMP-related hypermethylation of MLH1. Recent studies suggested that CIMP and CIN are mutually exclusive pathways [24, 25].
Since the introduction of the CIMP concept to the molecular carcinogenesis of CRCs, many investigators have attempted to characterize the clinicopathologic and molecular features of CIMP+ CRCs using their own CIMP marker panels and cutoff values. Despite the lack of a reference marker panel, associations with proximal colon location, MSI, and a high frequency of BRAF mutation have been virtually consistent findings for CIMP+ CRCs. However, inconsistent findings among an accumulating series of studies include older age, female predominance, high frequency of KRAS mutation, poor differentiation, and poor clinical outcomes. Furthermore, the proportion of CIMP+ tumors in all CRCs varied from 48% and 12%. These inconsistencies are related to the fact that laboratories have developed their own tools to define CIMP, and the variety of approaches has made it difficult to compare results from different groups. It is necessary to define uniform criteria for CIMP. The technology for performing quantitative methylation analysis and producing high-throughput analysis, e.g., MethyLight or pyrosequencing, is likely to be adopted as the reference technology for CIMP diagnosis. For the MethyLight technology, the eight-marker panel appears to be superior to the five-marker panels based on the findings of the present study and that of Ogino et al. [21]. However, for the pyrosequencing methodology, CIMP marker panels have not been as thoroughly evaluated.
In contrast to the findings of Ogino et al. [26] that CIMP+ colon cancers had a better prognosis than CIMP− colon cancers and that CIMP was an independent prognostic factor, our study showed that CIMP was not an independent prognostic factor for colon cancers. At present, we cannot explain why such a difference exists in the prognosis of CIMP+ tumors between the Ogino study and ours. In contrast to those studies reporting CIMP as an independent prognostic marker of poor prognosis in CRCs [12, 13], our study showed that CIMP was not a prognostic factor in CRCs and not an independent prognostic factor in colon cancers. The association of CIMP+ status with poor prognosis, which was statistically significant in colon cancer patients, remained no longer significant after adjusting for BRAF mutation. Previous studies reporting CIMP as an independent prognostic factor in CRCs did not analyze BRAF mutation in study samples and thus did not take the effect of BRAF mutation into consideration for the analysis of the association between CIMP+ status and prognosis of CRC patients [12, 13]. However, in the study of Barault et al. [14], CIMP+ status was closely associated with poor prognosis of MSI− colon cancers, and this association remained significant in multivariate analysis adjusted for age, stage, and BRAF and KRAS mutational status. The CIMP marker panel and methylation analysis technology used in the study of Barault et al. were classic five-marker panel (MLH1, MINT1, MINT2, MINT31, and p16) and methylation-specific PCR, respectively, and different from the panel and the analysis technology of the present study.
Compared with studies using Western patient populations, the present study showed discrepancies in the following respects: (1) the proportion of CIMP+ tumors in all CRCs, (2) the proportion of CIMP+ tumors in MSI+ CRCs, and (3) the rate of BRAF mutation in overall CRCs. In our study, 12% of CRCs were CIMP+, compared to 18% in the Ogino study [21]. Of MSI+ CRCs, 30% were CIMP+, compared to 71% in the Ogino study [21]. Studies having looked at specific causes of MSI+ CRCs not caused by germline mismatch repair gene mutation reported that majority (83–100%) of these cases have MLH1 methylation [27–30]. Based on such a high frequency of MLH1 methylation in MSI+ CRCs, Western researchers propose that virtually all sporadic MSI+ CRCs occur through the epigenetic instability pathway, with loss of MLH1 gene expression through promoter CpG island hypermethylation. If this contention is true, the following interpretation can be deduced from our findings: either three quarters of MSI+ cases should be hereditary or our methylation assay had a lower sensitivity for detection of CIMP. The latter is unlikely because we tested and corroborated the precision of the MethyLight technology by comparing the results of the MethyLight analysis with those of bisulfite genomic sequencing in cancer cell lines (data not shown). The former possibility is also unlikely because we randomly selected study cases from the surgical files of our department without information about familial history. Our results suggest the possibility that sporadic MSI+ cases develop through mechanisms other than MLH1 hypermethylation, including MLH1 or MSH2 gene mutation. Thus, it is plausible that there are ethnic differences in the causation of sporadic MSI+ CRCs, which explains the discrepancy in the percentage of CIMP positivity in MSI+ CRCs between Western data and the present study.
The rate of BRAF mutation (4.8%) in our CRC cases was similar to the rate (4.2%–9%) in other East Asian CRC studies [31–33], but lower than that in US, West Asian, and European CRC studies (9.5%–20.9%) [15, 34–36]. The rate of BRAF mutation was still low (6.1%, 11/181) when we excluded rectal cancers. However, the rate of KRAS mutation (101/285, 35.4%) was similar to that in other studies, regardless of geographic or ethnic differences (32%–34%) [9, 31, 37, 38]. The sensitivity of BRAF mutation detection could have been influenced by the mutation analysis methodology and use of the formalin-fixed paraffin-embedded tissues. But the former is unlikely because the enriched PCR–RFLP assay provides a highly sensitive means of detection and is capable of detecting one mutant allele in the presence of 1,000 normal alleles. In addition, we repeated BRAF mutation analysis at codon 600 (V600E) using real-time PCR-based allelic discrimination [36], giving the same result [20]. And it is also unlikely that use of the formalin-fixed tissue yielded lower detection of BRAF mutation because no difference was found in the mutation analysis of BRAF between the use of formalin-fixed tissues and the use of methanol-fixed tissues in a preliminary study (data not shown). At present, we cannot provide a satisfactory explanation for the discrepancy in BRAF mutation rates and the similarity of KRAS mutation rates between Western and East Asian CRC cases. Another point to address is the relationship between BRAF mutation and CIMP. One series of studies suggested that BRAF mutation is closely associated with sporadic MSI+ CRCs [39, 40], but this is disputed by recent studies indicating that BRAF mutation is closely associated with CIMP rather than MSI [16, 24]. Our study supports the close association of BRAF mutation with CIMP, because the BRAF mutation rate in CIMP+/MSI− CRCs was similar to that in CIMP+/MSI+ CRCs (33.3% and 37.5%, respectively).
In our study, CIMP+/MSI− CRCs exhibited the worst clinical outcomes among four molecular subtypes, which was attributed to BRAF mutation. However, BRAF mutation without CIMP was not associated with worse clinical outcome, which was consistent with our previous study [20]. Thus, CIMP+/MSI− CRCs with BRAF mutation pursued dismal clinical behavior, and the association of this molecular subtype with worse clinical outcome remained significant in multivariate analysis after adjusting for stage, differentiation, and age (data not shown). Because of the worse clinical outcome, molecular diagnostic tests to identify this subtype will be necessary in clinics, and targeted therapy against BRAF could be considered in the first-line adjuvant therapy.
Lastly, in our study, a close association of KRAS mutation with CIMP-low status was found for MSI− CRCs, which is consistent with the findings of the study of Ogino et al. [41] and furthermore in accordance with the study of Barault et al. using different CIMP marker panel and DNA methylation analysis technology [14]. Except for a strong association of KRAS mutation, CIMP-low tumors were similar to CIMP-0 tumors in several clinicopathological features, including p53 mutation [41], suggesting that CIMP-low tumors might be derived from CIN pathway. Because tubulovillous adenoma or villous adenoma harbors higher frequencies of KRAS mutation and CpG island hypermethylation than those of tubular adenoma [42–44], tubulovillous or villous adenomas might be precursor lesions of CIMP-low CRCs. To support this speculation, a further study analyzing carcinoma–ex adenoma samples for their methylation status in each component will be necessary to identify whether CIMP-low CRCs tend to have tubulovillous or villous adenoma as their contiguous adenoma.
In conclusion, we analyzed 320 CRC cases for CIMP and MSI status and determined the prognostic implications of CIMP status in CRC patients. The CIMP+/MSI− subtype showed the worst clinical outcome among the four possible molecular subtypes of CRCs, and the worse clinical outcome of CIMP+/MSI− subtype was attributed to BRAF mutation.
References
Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JP (1998) Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res 72:141–196
Esteller M, Corn PG, Baylin SB, Herman JG (2001) A gene hypermethylation profile of human cancer. Cancer Res 61:3225–3229
Bird AP, Wolffe AP (1999) Methylation-induced repression—belts, braces, and chromatin. Cell 99:451–454
Wood LD, Parsons DW, Jones S et al (2007) The genomic landscapes of human breast and colorectal cancers. Science 318:1108–1113
Schuebel KE, Chen W, Cope L et al (2007) Comparing the DNA hypermethylome with gene mutations in human colorectal cancer. PLoS Genet 3:1709–1723
Toyota M, Ahuja N, Ohe-Toyota M et al (1999) CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci USA 96:8681–8686
Weisenberger DJ, Siegmund KD, Campan M et al (2006) CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet 38:787–793
Ogino S, Cantor M, Kawasaki T et al (2006) CpG island methylator phenotype (CIMP) of colorectal cancer is best characterised by quantitative DNA methylation analysis and prospective cohort studies. Gut 55:1000–1006
Nagasaka T, Koi M, Kloor M et al (2008) Mutations in both KRAS and BRAF may contribute to the methylator phenotype in colon cancer. Gastroenterology 134:1950–1960.e1
Shen L, Toyota M, Kondo Y et al (2007) Integrated genetic and epigenetic analysis identifies three different subclasses of colon cancer. Proc Natl Acad Sci USA 104:18654–18659
Lee S, Cho NY, Yoo EJ, Kim JH, Kang GH (2008) CpG island methylator phenotype in colorectal cancers: comparison of the new and classic CpG island methylator phenotype marker panels. Arch Pathol Lab Med 132:1657–1665
Ward RL, Cheong K, Ku SL et al (2003) Adverse prognostic effect of methylation in colorectal cancer is reversed by microsatellite instability. J Clin Oncol 21:3729–3736
Shen L, Catalano PJ, Benson AB 3rd et al (2007) Association between DNA methylation and shortened survival in patients with advanced colorectal cancer treated with 5-fluorouracil based chemotherapy. Clin Cancer Res 13:6093–6098
Barault L, Charon-Barra C, Jooste V et al (2008) Hypermethylator phenotype in sporadic colon cancer: study on a population-based series of 582 cases. Cancer Res 68:8541–8546
Samowitz WS, Sweeney C, Herrick J et al (2005) Poor survival associated with the BRAF V600E mutation in microsatellite-stable colon cancers. Cancer Res 65:6063–6069
Ogino S, Goel A (2008) Molecular classification and correlates in colorectal cancer. J Mol Diagnostics 10:13–27
Yoo EJ, Park SY, Cho NY et al (2008) Helicobacter pylori-infection-associated CpG island hypermethylation in the stomach and its possible association with polycomb repressive marks. Virchows Arch 452:515–524
Kang GH, Lee S, Cho NY et al (2008) DNA methylation profiles of gastric carcinoma characterized by quantitative DNA methylation analysis. Lab Invest 88:161–170
Weisenberger DJ, Campan M, Long TI et al (2005) Analysis of repetitive element DNA methylation by MethyLight. Nucleic Acids Res 33:6823–6836
Lee S, Cho NY, Choi M et al (2008) Clinicopathological features of CpG island methylator phenotype-positive colorectal cancer and its adverse prognosis in relation to KRAS/BRAF mutation. Pathol Int 58:104–113
Ogino S, Kawasaki T, Kirkner GJ et al (2007) Evaluation of markers for CpG island methylator phenotype (CIMP) in colorectal cancer by a large population-based sample. J Mol Diagnostics 9:305–314
Georgiades IB, Curtis LJ, Morris RM, Bird CC, Wyllie AH (1999) Heterogeneity studies identify a subset of sporadic colorectal cancers without evidence for chromosomal or microsatellite instability. Oncogene 18:7933–7940
Goel A, Arnold CN, Niedzwiecki D et al (2003) Characterization of sporadic colon cancer by patterns of genomic instability. Cancer Res 63:1608–1614
Cheng YW, Pincas H, Bacolod MD et al (2008) CpG island methylator phenotype associates with low-degree chromosomal abnormalities in colorectal cancer. Clin Cancer Res 14:6005–6013
Goel A, Nagasaka T, Arnold CN et al (2007) The CpG island methylator phenotype and chromosomal instability are inversely correlated in sporadic colorectal cancer. Gastroenterology 132:127–138
Ogino S, Nosho K, Kirkner GJ et al (2009) CpG island methylator phenotype, microsatellite instability, BRAF mutation and clinical outcome in colon cancer. Gut 58:90–96
Herman JG, Umar A, Polyak K et al (1998) Incidence and functional consequences of hMLH1 promoter hypermethylation in colorectal carcinoma. Proc Natl Acad Sci USA 95:6870–6875
Kuismanen SA, Holmberg MT, Salovaara R, de la Chapelle A, Peltomaki P (2000) Genetic and epigenetic modification of MLH1 accounts for a major share of microsatellite-unstable colorectal cancers. Am J Pathol 156:1773–1779
Cunningham JM, Christensen ER, Tester DJ et al (1998) Hypermethylation of the hMLH1 promoter in colon cancer with microsatellite instability. Cancer Res 58:3455–3460
Cunningham JM, Kim CY, Christensen ER et al (2001) The frequency of hereditary defective mismatch repair in a prospective series of unselected colorectal carcinomas. Am J Hum Genet 69:780–790
Yuen ST, Davies H, Chan TL et al (2002) Similarity of the phenotypic patterns associated with BRAF and KRAS mutations in colorectal neoplasia. Cancer Res 62:6451–6455
Chang SC, Lin JK, Yang SH et al (2006) Relationship between genetic alterations and prognosis in sporadic colorectal cancer. Int J Cancer 118:1721–1727
Nagasaka T, Sasamoto H, Notohara K et al (2004) Colorectal cancer with mutation in BRAF, KRAS, and wild-type with respect to both oncogenes showing different patterns of DNA methylation. J Clin Oncol 22:4584–4594
Fransen K, Klintenas M, Osterstrom A et al (2004) Mutation analysis of the BRAF, ARAF and RAF-1 genes in human colorectal adenocarcinomas. Carcinogenesis 25:527–533
Vilkin A, Niv Y, Nagasaka T et al (2009) Microsatellite instability, MLH1 promoter methylation, and BRAF mutation analysis in sporadic colorectal cancers of different ethnic groups in Israel. Cancer 115:760–769
Young J, Barker MA, Simms LA et al (2005) Evidence for BRAF mutation and variable levels of microsatellite instability in a syndrome of familial colorectal cancer. Clin Gastroenterol Hepatol 3:254–263
Samowitz WS, Slattery ML, Sweeney C et al (2007) APC mutations and other genetic and epigenetic changes in colon cancer. Mol Cancer Res 5:165–170
Tanaka H, Deng G, Matsuzaki K et al (2006) BRAF mutation, CpG island methylator phenotype and microsatellite instability occur more frequently and concordantly in mucinous than non-mucinous colorectal cancer. Int J Cancer 118:2765–2771
Rajagopalan H, Bardelli A, Lengauer C et al (2002) Tumorigenesis: RAF/RAS oncogenes and mismatch-repair status. Nature 418:934
Oliveira C, Pinto M, Duval A et al (2003) BRAF mutations characterize colon but not gastric cancer with mismatch repair deficiency. Oncogene 22:9192–9196
Ogino S, Kawasaki T, Kirkner GJ, Loda M, Fuchs CS (2006) CpG island methylator phenotype-low (CIMP-low) in colorectal cancer: possible associations with male sex and KRAS mutations. J Mol Diagnostics 8:582–588
Rashid A, Shen L, Morris JS, Issa JP, Hamilton SR (2001) CpG island methylation in colorectal adenomas. Am J Pathol 159:1129–1135
Jass JR, Baker K, Zlobec I et al (2006) Advanced colorectal polyps with the molecular and morphological features of serrated polyps and adenomas: concept of a ‘fusion’ pathway to colorectal cancer. Histopathology 49:121–131
Kakar S, Deng G, Cun L, Sahai V, Kim YS (2008) CpG island methylation is frequently present in tubulovillous and villous adenomas and correlates with size, site, and villous component. Hum Pathol 39:30–36
Acknowledgments
This study was supported by the 21C Frontier Functional Human Genome Project from the Ministry of Science & Technology in Korea (FG09-11-02) and by a grant from the National R&D Program for Cancer Control, Ministry of Health & Welfare, Republic of Korea (0720540).
Conflict of interest statement
We declare that we have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
ELECTRONIC SUPPLEMENTARY MATERIAL
Below is the link to the electronic supplementary material.
Supplementary Table 1
(DOC 43 kb)
Supplementary Table 2
(DOC 58.5 kb)
Rights and permissions
About this article
Cite this article
Kim, J.H., Shin, S.H., Kwon, H.J. et al. Prognostic implications of CpG island hypermethylator phenotype in colorectal cancers. Virchows Arch 455, 485–494 (2009). https://doi.org/10.1007/s00428-009-0857-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-009-0857-0