Abstract
Cranial neurogenic placodes have been considered vertebrate innovations. However, anterior neural plate border (ANB) cells of ascidian embryos share many properties with vertebrate neurogenic placodes; therefore, it is now believed that the last common ancestor of vertebrates and ascidians had embryonic structures similar to neurogenic placodes of vertebrate embryos. Because BMP signaling is important for specifying the placode region in vertebrate embryos, we examined whether BMP signaling is also involved in gene expression in the ANB region of ascidian embryos. Our data indicated that Admp, a divergent BMP family member, is mainly responsible for BMP signaling in the ANB region, and that two BMP-antagonists, Noggin and Chordin, restrict the domain, in which BMP signaling is activated, to the ANB region, and prevent it from expanding to the neural plate. BMP signaling is required for expression of Foxg and Six1/2 at the late gastrula stage, and also for expression of Zf220, which encodes a zinc finger transcription factor in late neurula embryos. Because Zf220 negatively regulates Foxg, when we downregulated Zf220 by inhibiting BMP signaling, Foxg was upregulated, resulting in one large palp instead of three palps (adhesive organs derived from ANB cells). Functions of BMP signaling in specification of the ANB region give further support to the hypothesis that ascidian ANB cells share an evolutionary origin with vertebrate cranial placodes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cranial neurogenic placodes in vertebrate embryos are ectodermal thickenings at the anterior border of the neural plate. These placodes give rise to cells in various organs in the vertebrate head, including the adenohypophysis, olfactory epithelium, lens, inner ear, and petrosal and nodose ganglia (Schlosser 2014; Singh and Groves 2016; Steventon et al. 2014). It has been proposed that acquisition of these placodes remodeled rostral structures in the vertebrate lineage (Gans and Northcutt 1983).
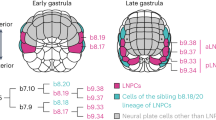
Ascidians belong to the sister group of vertebrates (Delsuc et al. 2006; Putnam et al. 2008), and previous studies have shown that the anterior neural plate border (ANB) region of ascidian embryos shares an evolutionary origin with vertebrate placodes (Abitua et al. 2015; Cao et al. 2019; Horie et al. 2018; Ikeda et al. 2013; Liu and Satou 2019, 2020; Manni et al. 2005, 2004; Mazet et al. 2005). At the gastrula stage, two rows of ANB cells (four cells in each row), which express Foxc, are formed between the neural plate and non-neural ectoderm (Ikeda et al. 2013; Wagner and Levine 2012) (Fig. 1A). At the neurula stage, ANB cells divide once to yield four rows, each of which contains four cells. Cells in the most posterior row express transcription factor genes, Six1/2 and Foxg (Abitua et al. 2015; Liu and Satou 2019). The most anterior row also expresses Foxg, but not Six1/2. The second and third rows express Emx, which is repressed by Foxg in the most anterior and posterior rows (Liu and Satou 2019; Wagner et al. 2014). The MAPK pathway is specifically activated in the most anterior and posterior rows, because it is negatively regulated by Ephrina.d in the middle two rows. This MAPK pathway specifically activates Foxg in the first and fourth rows. After the next division, the first and second anterior rows both include eight cells, and Zf220 begins to be expressed in four cells in each row. Foxg expression becomes rarely visible, but soon after it is reactivated in descendant cells of the most anterior row. Because Zf220 negatively regulates Foxg, Foxg expression becomes restricted in four cells, which do not express Zf220, in the first row at the middle tailbud stage (Liu and Satou 2019). Thus, several distinct cell populations are specified in the ANB region. Among them, Foxg( +)/Six1/2(-) cells gives rise to the protrusive parts of the palps, which are adhesive organs required for metamorphosis. Foxg(-) cells contribute to basal parts of the palps. Foxg( +)/Six1/2( +) cells give rise to the oral siphon primordium (Abitua et al. 2015; Liu and Satou 2019; Wagner et al. 2014).
BMP signaling is activated in ANB cells. (A) Schematic illustration of the anterior border of the neural plate in ascidian late gastrula and early neurula embryos. Because embryos are bilaterally symmetrical, names of individual cells are indicated only in the left half, and are prefixed with IDs shown next to the illustration, e.g., the left upper cell of the late gastrula embryo is called a9.36. Vertical bars indicate sister cell relationships. (B, C) Immunostaining of (B) a late gastrula embryo and (C) an early neurula embryo to detect phosphorylated Smad1/5/9 (pSmad1/5/9). ANB cells were marked by Foxc > GFP expression, detected with an anti-GFP antibody. Note that GFP was detected only on the right side because of mosaic incorporation in (C). Right images are overlaid in pseudocolor. The brightness and contrast of these photographs were adjusted linearly. ANB cells are enclosed by broken lines. Note that strong signals are observed in the anterior medial ANB cells and relatively weak signals are also observed in the remaining ANB cells. Dorsal views are shown. The scale bar in (B) represents 50 μm
A previous study showed that misexpression of Bmp2/4 or a constitutively active form of a BMP receptor in ANB cells downregulates expression of Six1/2 (Abitua et al. 2015). Therefore, BMP signaling may be involved in formation and pattering of the ANB region of ascidian embryos as in formation of the pre-placodal region of vertebrate embryos, in which BMP signaling is essential to form placodes (Ahrens and Schlosser 2005; Brugmann et al. 2004; Esterberg and Fritz 2009; Glavic et al. 2004; Kwon et al. 2010; Litsiou et al. 2005). In the present study, we examined how BMP signaling contributes to patterning of ANB cells in ascidian embryos to explore developmental mechanisms of the last common ancestor of vertebrates and ascidians.
Materials and Methods
Animals and gene identifiers
Adult Ciona robusta (also called Ciona intestinalis type A) were obtained from the National Bio-Resource Project for Ciona intestinalis. This animal is excluded from legislation regulating scientific research on animals in Japan. cDNA clones were obtained from our EST clone collection (Satou et al. 2005). Identifiers (Satou et al. 2022; Stolfi et al. 2015b) for genes examined in the present study are as follows: CG.KY21.Chr12.158 for Foxc, CG.KY21.Chr8.693 for Foxg, CG.KY21.Chr2.381 for Admp, CG.KY21.Chr4.346 for Bmp2/4, CG.KY21.Chr2.933 for Bmp5/6/7/8, CG.KY21.Chr12.737 for Noggin, CG.KY21.Chr6.382 for Chordin, CG.KY21.Chr3.541 for Six1/2, and CG.KY21.Chr13.415 for Zf220.
Functional assays
Dorsomorphin, an inhibitor of BMP signaling (Wako, #044–33751), was applied to embryos at a concentration of 50 μM. The upstream sequence of Foxc [nucleotide positions 1,011,273 to 1,013,312 on chromosome 12 of the HT version of the assembly (Satou et al. 2019)], which directs expression in ANB cells (Liu and Satou 2019; Wagner and Levine 2012), was used for misexpression of Noggin and Chordin. To mark ANB cells, the same upstream sequence was used to direct GFP expression. These constructs were introduced into fertilized eggs by electroporation (Corbo et al. 1997). An antisense morpholino oligonucleotide against Admp, which was used previously (Imai et al. 2012, 2006; Waki et al. 2015) (5’-TATCGTGTAGTTTGCTTTCTATATA-3’), was introduced into eggs by microinjection. All functional assays were performed at least twice using different batches of embryos.
In situ hybridization and immunostaining
For whole-mount in situ hybridization, digoxigenin (DIG)-RNA probes were synthesized by in vitro transcription with T7 RNA polymerase. Embryos were fixed in 4% paraformaldehyde in 0.1 M MOPS-NaOH (pH 7.5) and 0.5 M NaCl at 4 °C overnight and then stored in 80% ethanol. After washing with a phosphate-buffered saline containing 0.1% tween 20 (PBST), embryos were treated with 2 μg/mL Proteinase K for 30 min at 37 °C, washed again with PBST, and fixed with 4% paraformaldehyde for 1 h at room temperature. Embryos were then incubated in 6 × saline sodium citrate buffer (SSC), 50% formamide, 5 × Denhardt’s solution, 100 μg/mL yeast tRNA, and 0.1% tween 20 for 1 h at 50 °C. After this pre-hybridization step, specific RNA probes were added and incubated for 48 h at 50 °C. Embryos were treated with RNase A, and incubated in 0.5 × SSC, 50% formamide, and 0.1% tween 20 for 15 min at 50 °C twice. Embryos were further incubated in 0.5% blocking reagent (Roche) in PBST for 30 min, and then in 1:2000 alkaline-phosphatase-conjugated anti-DIG antibody (Roche). For chromogenic detection, embryos were further washed with 0.1 M NaCl, 50 mM MgCl2, and 0.1 M Tris–HCl (pH 9.5). Then NBT and BCIP were used for detection. For fluorescent detection, we used the TSA plus system (Perkin Elmer, #NEL753001KT).
For immunostaining, embryos were fixed with 3.7% formaldehyde, and treated with 3% H2O2 for 30 min to quench endogenous peroxidase activity. Then they were incubated overnight with an anti-phosphoryrated-Smad1 antibody (1:1000, Abcam, #ab97689) in Can-Get-Signal-Immunostain Solution B (TOYOBO, #NKB-601). The signal was visualized with a TSA Kit (Invitrogen, #T30953) using HRP-conjugated goat anti-rabbit IgG and Alexa Fluor 555 tyramide. GFP protein was similarly detected with an antibody against GFP (1:300, MBL, #M048-3). Fluorescence intensities were quantified with ImageJ software (Schneider et al. 2012), and represented as relative intensities, which were calculated by dividing sums of pSmad1/5/9 signals of four ANB cells with sums of DAPI signals of the same cells.
Results
BMP signaling is activated in ANB cells
To understand exactly how BMP signaling influences patterning of ANB cells, we first examined whether BMP signaling is activated in ANB cells by an antibody against phosphorylated Smad1, which was previously used to detect phosphorylated Smad1/5/9 (pSmad1/5/9) in ascidian embryos (Waki et al. 2015). To mark ANB cells, we introduced a reporter construct containing a fusion gene of GFP and the Foxc upstream regulator sequence (Foxc > GFP). At the gastrula stage, immunostaining signals for pSamd1/5/9 were detected in all ANB cells marked with GFP, although the medial two cells in the anterior row (a9.40 cells) showed much stronger signals than the other ANB cells (Fig. 1B). At the early neurula stage, these cells divide along the anterior–posterior axis to make four rows. The central cells of the anterior two rows (a10.79 and a10.80, which are daughter cells of a9.40) showed stronger signals than the remaining ANB cells (Fig. 1C).
Admp is expressed in ANB cells
Next, we tried to identify a BMP ligand responsible for activation of BMP signaling in the ANB region. In the ascidian genome, four BMP ligand genes are encoded (Hino et al. 2003). Among them, Bmp3 is not expressed at the gastrula or neurula stages (Imai et al. 2004), and Bmp2/4 is activated under control of Admp (Imai et al. 2012; Waki et al. 2015). Therefore, we determined precise identities of cells that expressed the remaining two ligand genes, Admp and Bmp5/6/7/8, by in situ hybridization. Admp was expressed in all ANB cells that express Foxc at the late gastrula stage (Fig. 2AB). Expression of Admp was also observed in all ANB cells of early neurula embryos, in which the most anterior and posterior rows of ANB cells are marked by Foxg expression (Fig. 2CD). On the other hand, Bmp5/6/7/8 began to be expressed weakly in the most anterior row of ANB cells at the early neurula stage (Fig. 2EF). This gene was also expressed in an anterior row of neural plate. There was one row of cells between the posterior row of ANB cells and the row of the neural plate cells with Bmp5/6/7/8 expression. Expression of Bmp5/6/7/8 in the anterior ANB cells was transient, and disappeared by the middle neurula stage (Fig. 2G).
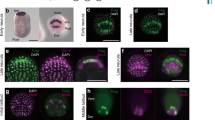
Admp and BMP5/6/7/8 are expressed in ANB cells. (A-G) In situ hybridization for (A, C) Admp (magenta) expression at (A) the late gastrula stage and (C) early neurula stage, and for (E, G) Bmp5/6/7/8 (magenta) expression at (E) the early neurula stage and (G) the middle neurula stage. Foxc in (A) marks ANB cells (green). Foxg in (C) and (E) marks the most anterior and posterior rows of the ANB region (green). Six1/2 in (G) marks the most posterior row of the ANB (green), although this gene is also expressed in the two flanking cells on both sides of the ANB. The results shown in (A), (C), and (E) are illustrated in (B), (D), and (F), respectively. Dorsal views are shown. (H, I) Optical slices of pSmad1/5/9 immunostaining of (H) an unperturbed control early neurula embryo and (I) an early neurula embryo injected with the Admp MO (green). Nuclei are stained with DAPI (gray). ANB cells are marked by arrowheads. Photographs are pseudocolored, and lateral views are shown. (J) Quantification of fluorescence intensities of signals for pSmad1/5/9 in the ANB region of control unperturbed embryos and Admp morphants. Each dot represents relative signal intensities of a line of four medial ANB cells in an embryo, and bars represent median values. Relative intensities were calculated by dividing sums of pSmad1/5/9 signals of four ANB cells with sums of DAPI signals of the same cells. As we used tyramide signal amplification, signal levels might not be amplified linearly. However, a Wilcoxon’s rank sum test indicated that signal levels are significantly different between the control and experimental specimens. The scale bar in (A) represents 50 μm
Because Admp, but not Bmp5/6/7/8, was expressed in ANB cells of late gastrula embryos, we reasoned that Admp is mainly responsible for activation of BMP signaling in the ANB region. To confirm this hypothesis, we injected an antisense morpholino oligonucleotide against Admp. Signal intensity of immunostaining for pSmad1/5/9 was greatly reduced in Admp morphant embryos, indicating a primary role of Admp in activating BMP signaling in this region (Fig. 2H–J). Because Admp is expressed in early embryos (Imai et al. 2004, 2006; Oda-Ishii et al. 2016; Pasini et al. 2006; Tokuoka et al. 2021), we cannot completely rule out the possibility that Admp expressed in early embryos indirectly regulates BMP signaling in the ANB cells. It is also possible that Admp expressed in cells other than ANB cells activates BMP signaling in the ANB region, because Admp is a secreted molecule. However, the knockdown result and the observation that Admp was expressed in all ANB cells agreed with the above hypothesis that Admp is mainly responsible for activation of BMP signaling in the ANB region.
Chordin and Noggin are expressed in cells lateral and posterior to the adjacent ANB region
The expression pattern of Admp was not perfectly consistent with the pattern of pSmad1/5/9 signals (compare Fig. 1 to Fig. 2). Therefore, we examined expression patterns of genes encoding secreted BMP-antagonists, which can act non-cell-autonomously. At the late gastrula stage, Chordin was expressed on both sides of the neural plate, as previously reported (Abitua et al. 2015; Hudson and Yasuo 2005). At this stage, the most anterior cells with Chordin expression were adjacent to the lateral ANB cells (Fig. 3A). Noggin was expressed in the neurula plate and the expression domain was adjacent to the posterior row of ANB cells (Fig. 3B). At the early neurula stage, the lateral and posterior boundaries of the ANB region are still flanked with cells expressing Chordin and Noggin, respectively (Fig. 3C D). Thus, expression patterns of Chordin and Noggin in the ANB region account for the immunostaining pattern for pSmad1/5/9. Indeed, overexpression of Chordin and Noggin in the ANB region using the Foxc regulatory region weakened the level of pSmad1/5/9 staining (Fig. 3EF). Note that the pSmad1/5/9 staining was also weakened in cells that do not express Foxc because Chordin and Noggin are secreted antagonists.
Chordin and Noggin are expressed in cells next to the ANB cells. (A-D) In situ hybridization for (A) Foxc (green) and Chordin (magenta), (B) Foxc (green) and Noggin (magenta) at late gastrula stage, and for (C) Foxg (green) and Chordin (magenta), and (D) Foxg (green) and Noggin (magenta) at the early neurula stage. Nuclei were stained with DAPI (gray). Photographs are pseudocolored Z-projected image stacks. Dorsal views are shown. Note that the anterior row of the ANB is not visible in (D). (E) Optical slices of pSmad1/5/9 immunostaining of an unperturbed control early neurula embryo and early neurula embryos injected with Foxc > Chordin or Foxc > Noggin. ANB cells are marked with arrowheads. Lateral views are shown. The scale bar in (A) represents 50 μm. (F) Quantification of fluorescence intensities of signals for pSmad1/5/9 in the ANB region of unperturbed control embryos and embryos injected with Foxc > Chordin or Foxc > Noggin. Each dot represents relative signal intensities of a line of four medial ANB cells in an embryo, and bars represent median values. Relative intensities were calculated by dividing sums of pSmad1/5/9 signals of four ANB cells with sums of DAPI signals of the same cells. As we used tyramide signal amplification, signal levels might not be amplified linearly. However, Wilcoxon’s rank sum tests indicated that signal levels are significantly different between the control and experimental specimens
Expression of Foxg and Six1/2 in the ANB region requires BMP signaling
To examine whether BMP signaling is necessary for gene expression in the ANB region of neurulae, we misexpressed Noggin in ANB cells using the upstream regulatory sequence of Foxc (Foxc > Noggin). Foxg expression was clearly downregulated in the posterior row of 82% of these experimental embryos, and also in the anterior row of 7% of embryos (Fig. 4AB). Next, embryos introduced with the Foxc > Noggin construct were treated with dorsomorphin, which is an inhibitor of BMP signaling and has been used in this animal (Ohta and Satou 2013; Waki et al. 2015), from the early gastrula stage. Because DNA constructs introduced by electroporation may not be expressed in all ANB cells, we expected BMP signaling levels to be further reduced by dorsomorphin treatment. All of these embryos lost Foxg expression in the posterior row, and 31% of these embryos additionally lost Foxg expression in the anterior row (Fig. 4B). These results indicate that BMP signaling is required for Foxg expression in the ANB region, although Foxg expression is more sensitive to downregulation of BMP signaling in the posterior row than in the anterior row.
Suppression of BMP signaling activity affects expression of Foxg and Six1/2 at the neurula stage. (A) A dorsal view of in situ hybridization for Foxg in an embryo in which Foxc > Noggin was introduced by electroporation. Expression of Foxg in the posterior row (white arrowhead) was lost, while expression in the anterior row was not affected in this embryo (black arrowhead). (B) Percentages of embryos that normally expressed Foxg, or lost Foxg expression. (C) A dorsal view of in situ hybridization for Six1/2 in an embryo in which Foxc > Noggin was introduced by electroporation. Expression of Six1/2 in the ANB region was lost or greatly reduced (white arrowheads). (D) Percentages of embryos that normally expressed or lost Six1/2 expression in the posterior row of the ANB region of normal and embryos electroporated with Foxc > Noggin. (E) Percentages of embryos that normally expressed or lost Six1/2 expression in one or more ANB cells in embryos microinjected with Foxc > Noggin at different concentrations. (F) Percentages of embryos that ectopically expressed Six1/2 in embryos injected with Foxc > Noggin at different concentrations. (G) A dorsal view of an embryo in which Foxc > Noggin was injected at 20 ng/μL, expressed Six1/2 ectopically (arrowheads). The scale bar in (A) represents 50 μm
Next, we examined Six1/2 expression in the above two types of experimental embryo. In 93% of embryos electroporated with Foxc > Noggin, Six1/2 expression was reduced (Fig. 4CD). Notably, 18% of embryos lost Six1/2 expression completely in the ANB region, indicating that BMP signaling is required for Six1/2 expression in the ANB region. On the other hand, in the experiment in which we additionally treated embryos with dorsomorphin, Six1/2 expression was lost in only 63%. We did not find any embryos that completely lost Six1/2 expression in the ANB region, raising the possibility that Six1/2 is also expressed in cells with a very low level or absence of BMP signaling.
To confirm this hypothesis, we injected various concentrations of Foxc > Noggin into eggs. While less than 10% of these embryos lost Six1/2 expression in the most posterior cells at low concentrations (1 or 5 ng/μL), 24 to 31% of embryos lost Six1/2 expression at high concentrations (10, 15, or 20 ng/μL) (Fig. 4E). The percentage of embryos that lost Six1/2 expression is slightly lower at 20 ng/μL than at 10 or 15 ng/μL. These observations are consistent with the hypothesis that BMP signaling is necessary for Six1/2 expression in the posterior row, but a very low level or absence of BMP signaling can also induce Six1/2 expression.
Next, we counted embryos with ectopic expression in the same specimens. While ectopic expression was hardly observed at low concentrations, it was observed in almost all embryos at 15 and 20 ng/μL (Fig. 4FG). This observation is consistent with the hypothesis that a very low level or lack of BMP signaling can induce Six1/2 expression.
BMP signaling activity is required for proper palp formation
Larval palps are derived from the anterior three rows of ANB cells at the neurula stage. The most anterior row gives rise to protrusive structures and the middle two rows give rise to basal cells surrounding the protrusions (Cao et al. 2019; Ikeda et al. 2013; Liu & Satou 2019; Wagner & Levine 2012; Wagner et al. 2014). Normal larvae have three protrusions (two are seen in Fig. 5A). Larvae with the misexpression construct of Noggin developed one large protrusion (Fig. 5B), suggesting that BMP signaling is involved in palp formation.
BMP signaling activity is required for proper palp formation. (A) Morphology of the trunk of larvae developed from (A) a control unperturbed egg and (B) an egg electroporated with the Foxc > Noggin construct. Two of three palp protrusions are visible in this larva. We examined 16 control larvae and 24 experimental larvae, and all of them exhibited phenotypes represented by these photographs. (C-F) In situ hybridization for Foxg in (C) a control unperturbed embryo, (D) an embryo electroporated with the Foxc > Noggin construct, (E) a control DMSO-treated embryo, and (F) an embryo treated with 50 μM dorsomorphin at the middle tailbud stage. We examined 45 embryos electroporated with Foxc > Noggin, and 101 embryos treated with dorsomorphin. Among them, 93% and 75% of embryos exhibited the phenotypes represented in (D) and (F), respectively. (G, H) Double fluorescence in situ hybridization for Foxg (green) and Zf220 (magenta) in (G) a control embryo, (H) an embryo treated with 50 μM dorsomorphin. In (H), the experimental embryo lost Zf220 in the central region, and ectopically expressed Foxg. The expression pattern of Foxg and Zf220 in the most anterior row of normal embryos are depicted on the right of (G). These most-anterior-row cells are daughters of a10.72 and a10.80 as illustrated. Note that Zf220 is also expressed in the second row (Liu and Satou 2019), which are not depicted. Photographs from C to H are anterior views, and the dorsal side of the trunk is down. (I) Percentages of embryos that normally expressed Zf220 (black), lost Zf220 expression in the central cells (gray), and lost it in both of the central and flanking cells (white) in the anterior row of unperturbed control embryos and embryos treated with dorsomorphin. We examined Zf220 expression by non-fluorescence in situ hybridization in addition to double fluorescence in situ hybridization shown in (G) and (H). The scale bars in (A), (C), and (G) represent 50 μm
We previously reported that larvae in which Foxg is misexpressed in all ANB cells also develop one large palp (Liu and Satou 2019). Therefore, we examined whether misexpression of Noggin induced ectopic Foxg expression in tailbud embryos. In normal development, Foxg expression becomes rarely visible at the early tailbud stage, but soon after it is reactivated in three clusters of cells, which are descendants of the most anterior row cells (Fig. 5C), each of which gives rise to a palp protrusion, as previously shown (Liu and Satou 2019). On the other hand, in embryos with the Noggin misexpression construct, Foxg was expressed ectopically in gaps between these clusters, and the Foxg expression domain became a single arc (Fig. 5D). Similarly, in tailbud embryos treated with dorsomorphin from the middle neurula stage, Foxg was expressed ectopically in a single arc, while it was normally expressed in control embryos treated with DMSO (Fig. 5E, F).
At the tailbud stage, Zf220 represses Foxg in cells between the clusters where Foxg is normally expressed (Liu and Satou 2019). We therefore hypothesized that Zf220 might be downregulated in embryos in which BMP signaling was inhibited. In normal embryos, Zf220 was expressed in cells between the clusters of cells with Foxg expression (Fig. 5G), as previously shown (Liu & Satou 2019). On the other hand, expression of Zf220 in the intervening cells was downregulated in almost all (94%) embryos treated with dorsomorphin from the gastrula stage (Fig. 5H). Thus, BMP activity is required for Zf220 expression in these intervening cells. Zf220 is also expressed in the both ends of the anterior rows, and this expression was also affected in 52% of the embryos (Fig. 5I). Intervening cells that lost Zf220 are derived from a10.80 in the most anterior row and a10.79 in the second row, and these parental cells showed strong pSmad1/5/9 signals at the early neurula stage (Fig. 1DE). On the other hand, flanking cells, in which Zf220 expression was less affected, are derived from a10.72 in the most anterior row and a10.71 in the second row, and these parental cells showed weak pSmad1/5/9 signals at the early neurula stage (Fig. 1DE). Thus, expression of Zf220 in medial cells is under control of BMP signaling, and also weakly regulated by BMP signaling in the flanking cells.
Discussion
At the late gastrula and early neurula stages, BMP signaling is activated in ANB cells. Our data indicate that Admp activates BMP signaling in the ANB region, and that Chordin and Noggin restrict the region with BMP signaling activity to the ANB region and prevent it from expanding to the neural plate. Because Bmp2/4 is activated under control of Admp (Imai et al. 2012; Waki et al. 2015), it is possible that Bmp2/4 also acts in this region. Although these factors create a graded pattern of BMP signaling, the difference in intensity of BMP signaling within the ANB region is not important for Foxg and Six1/2 expression at the late gastrula stage, because a low level of BMP signaling is sufficient for expression of these genes. In addition, since Foxg expression is restricted to the most anterior and posterior rows of the ANB region by the action of the MAPK signaling, BMP signaling may not be required for positional information, and may have a permissive role.
Our results showed that Six1/2 expression in the ANB region normally requires BMP signaling, because reduction of BMP signaling levels by misexpression of Noggin resulted in loss of Six1/2 expression. However, because Six1/2 expression was seen in embryos in which BMP signaling levels were further reduced by additional treatment with dorsomorphin, it is likely that Six1/2 expression is also induced at a very low level or even an absence of BMP signaling. This is consistent with expression of Six1/2 in cells flanking the ANB region. In these flanking cells, Chordin is expressed and pSmad1/5/9 was scarcely detected. In addition, because a previous study showed that overexpression of Bmp2/4 suppressed Six1/2 expression (Abitua et al. 2015), it is likely that Six1/2 expression is induced at modest, or very low (or negligible) levels of BMP signaling, and is suppressed at a high level or between the modest and very low levels.
At the late neurula stage, Zf220 begins to be expressed in descendants of cells that show strong signals for pSmad1/5/9, and Zf220 expression in these descendants is under control of BMP signaling. Although it remains to be clarified how Zf220 is expressed in only some of these descendant cells, it is possible that BMP signaling has an instructive role in activating Zf220 in these cells. Zf220 is also expressed in descendants of cells that showed weak signals for pSmad1/5/9. However, in these cells, Zf220 expression was less sensitive to downregulation of BMP signaling; therefore, regulation of Zf220 in these cells differs from that in medial cells.
Thus, BMP signaling positively regulates gene expression in the ANB region, which is thought to share an evolutionary origin with vertebrate cranial placodes (Abitua et al. 2015; Graham and Shimeld 2013; Ikeda et al. 2013; Ikeda and Satou 2017; Liu and Satou 2019; Manni et al. 2005, 2004; Mazet et al. 2005; Wagner and Levine 2012). In vertebrate embryos, it has been proposed that ectodermal cells are induced by BMP, giving rise to epidermis, pre-placodal ectoderm, neural crest, and neural plate at different BMP concentrations, although specification of pre-placodal ectoderm and neural crest have different requirements for additional signaling molecules and for BMP signaling at different times (Ahrens and Schlosser 2005; Brugmann and Moody 2005; Glavic et al. 2004; Kwon et al. 2010; Steventon et al. 2014; Tribulo et al. 2003). Thus, ANB cells of ascidians and pre-placodal cells of vertebrates commonly use BMP signals for specification.
In ascidian embryos, there are cell populations that arguably share their evolutionary origins with vertebrate neural crest cells (Abitua et al. 2012; Stolfi et al. 2015a; Waki et al. 2015). It has not been determined whether BMP signaling is involved in specification of these neural-crest-like cells in ascidian embryos. However, in ascidian embryos, cell lineages of these neural-crest-like cells are clearly distinct from the cell lineage of the ANB. This means that there is no cell population that has potential to become both ANB cells and neural-crest-like cells in ascidian embryos; therefore, it is unlikely that a gradient of BMP signaling is required in this animal to specify a cell population that has bipotential to give rise to these two cell populations. Nevertheless, the requirement of BMP signaling for gene expression in the ANB region of ascidian embryos further supports the hypothesis that ANB cells share an evolutionary origin with vertebrate cranial placodes.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Abitua PB, Wagner E, Navarrete IA, Levine M (2012) Identification of a rudimentary neural crest in a non-vertebrate chordate. Nature 492:104–107. https://doi.org/10.1038/nature11589
Abitua PB, Gainous TB, Kaczmarczyk AN, Winchell CJ, Hudson C, Kamata K, Nakagawa M, Tsuda M, Kusakabe TG, Levine M (2015) The pre-vertebrate origins of neurogenic placodes. Nature 524:462–465. https://doi.org/10.1038/nature14657
Ahrens K, Schlosser G (2005) Tissues and signals involved in the induction of placodal Six1 expression in Xenopus laevis. Dev Biol 288:40–59. https://doi.org/10.1016/j.ydbio.2005.07.022
Brugmann SA, Moody SA (2005) Induction and specification of the vertebrate ectodermal placodes: precursors of the cranial sensory organs. Biol Cell / Under Auspices Eur Cell Biol Organ 97:303–319. https://doi.org/10.1042/BC20040515
Brugmann SA, Pandur PD, Kenyon KL, Pignoni F, Moody SA (2004) Six1 promotes a placodal fate within the lateral neurogenic ectoderm by functioning as both a transcriptional activator and repressor. Development 131:5871–5881. https://doi.org/10.1242/dev.01516
Cao C, Lemaire LA, Wang W, Yoon PH, Choi YA, Parsons LR, Matese JC, Wang W, Levine M, Chen K (2019) Comprehensive single-cell transcriptome lineages of a proto-vertebrate. Nature 571:349–354. https://doi.org/10.1038/s41586-019-1385-y
Corbo JC, Levine, M, Zeller, RW (1997) Characterization of a notochord-specific enhancer from the Brachyury promoter region of the ascidian Ciona intestinalis Development 124(3) 589–602. https://doi.org/10.1242/dev.124.3.589
Delsuc F, Brinkmann H, Chourrout D, Philippe H (2006) Tunicates and not cephalochordates are the closest living relatives of vertebrates. Nature 439:965–968. https://doi.org/10.1038/nature04336
Esterberg R, Fritz A (2009) dlx3b/4b are required for the formation of the preplacodal region and otic placode through local modulation of BMP activity. Dev Biol 325:189–199. https://doi.org/10.1016/j.ydbio.2008.10.017
Gans C, Northcutt RG (1983) Neural Crest and the Origin of Vertebrates - a New Head. Science 220:268–273. https://doi.org/10.1126/science.220.4594.268
Glavic A, Honore SM, Feijoo CG, Bastidas F, Allende ML, Mayor R (2004) Role of BMP signaling and the homeoprotein iroquois in the specification of the cranial placodal field. Dev Biol 272:89–103. https://doi.org/10.1016/j.ydbio.2004.04.020
Graham A, Shimeld SM (2013) The origin and evolution of the ectodermal placodes. J Anat 222:32–40. https://doi.org/10.1111/j.1469-7580.2012.01506.x
Hino K, Satou Y, Yagi K, Satoh N (2003) A genomewide survey of developmentally relevant genes in Ciona intestinalis. VI. Genes for Wnt, TGFbeta, Hedgehog and JAK/STAT signaling pathways. Dev Genes Evol 213:264–272. https://doi.org/10.1007/s00427-003-0318-8
Horie R, Hazbun A, Chen K, Cao C, Levine M, Horie T (2018) Shared evolutionary origin of vertebrate neural crest and cranial placodes. Nature 560:228–232. https://doi.org/10.1038/s41586-018-0385-7
Hudson C, Yasuo H (2005) Patterning across the ascidian neural plate by lateral Nodal signalling sources. Development 132:1199–1210. https://doi.org/10.1242/dev.01688
Ikeda T, Satou Y (2017) Differential temporal control of Foxa.a and Zic-r.b specifies brain versus notochord fate in the ascidian embryo. Development 144:38–43. https://doi.org/10.1242/dev.142174
Ikeda T, Matsuoka T, Satou Y (2013) A time delay gene circuit is required for palp formation in the ascidian embryo. Development 140:4703–4708. https://doi.org/10.1242/dev.100339
Imai KS, Hino K, Yagi K, Satoh N, Satou Y (2004) Gene expression profiles of transcription factors and signaling molecules in the ascidian embryo: towards a comprehensive understanding of gene networks. Development 131:4047–4058. https://doi.org/10.1242/dev.01270
Imai KS, Levine M, Satoh N, Satou Y (2006) Regulatory blueprint for a chordate embryo. Science 312:1183–1187. https://doi.org/10.1126/science.1123404
Imai KS, Daido Y, Kusakabe TG, Satou Y (2012) Cis-acting transcriptional repression establishes a sharp boundary in chordate embryos. Science 337:964–967. https://doi.org/10.1126/science.1222488
Kwon HJ, Bhat N, Sweet EM, Cornell RA, Riley BB (2010) Identification of Early Requirements for Preplacodal Ectoderm and Sensory Organ Development. PLoS genetics 6:e1001133. https://doi.org/10.1371/journal.pgen.1001133
Litsiou A, Hanson S, Streit A (2005) A balance of FGF, BMP and WNT signalling positions the future placode territory in the head. Development 132:4051–4062. https://doi.org/10.1242/dev.01964
Liu B, Satou Y (2019) Foxg specifies sensory neurons in the anterior neural plate border of the ascidian embryo. Nat Commun 10:4911. https://doi.org/10.1038/s41467-019-12839-6
Liu B, Satou Y (2020) The genetic program to specify ectodermal cells in ascidian embryos. Dev Growth Differ 62:301–310. https://doi.org/10.1111/dgd.12660
Manni L, Lane NJ, Joly JS, Gasparini F, Tiozzo S, Caicci F, Zaniolo G, Burighel P (2004) Neurogenic and non-neurogenic placodes in ascidians. J Exp Zool. Part b, Mol Dev Evol 302:483–504. https://doi.org/10.1002/jez.b.21013
Manni L, Agnoletto A, Zaniolo G, Burighel P (2005) Stomodeal and neurohypophysial placodes in Ciona intestinalis: insights into the origin of the pituitary gland. J Exp Zool. Part b, Mol Dev Evol 304:324–339. https://doi.org/10.1002/jez.b.21039
Mazet F, Hutt JA, Milloz J, Millard J, Graham A, Shimeld SM (2005) Molecular evidence from Ciona intestinalis for the evolutionary origin of vertebrate sensory placodes. Dev Biol 282:494–508. https://doi.org/10.1016/j.ydbio.2005.02.021
Oda-Ishii I, Kubo A, Kari W, Suzuki N, Rothbacher U, Satou Y (2016) A maternal system initiating the zygotic developmental program through combinatorial repression in the ascidian embryo. PLoS genetics 12:e1006045. https://doi.org/10.1371/journal.pgen.1006045
Ohta N, Satou Y (2013) Multiple signaling pathways coordinate to induce a threshold response in a chordate embryo. PLoS genetics 9:e1003818. https://doi.org/10.1371/journal.pgen.1003818
Pasini A, Amiel A, Rothbacher U, Roure A, Lemaire P, Darras S (2006) Formation of the ascidian epidermal sensory neurons: insights into the origin of the chordate peripheral nervous system. PLoS Biol 4:e225. https://doi.org/10.1371/journal.pbio.0040225
Putnam NH, Butts T, Ferrier DE, Furlong RF, Hellsten U, Kawashima T, Robinson-Rechavi M, Shoguchi E, Terry A, Yu JK, Benito-Gutierrez EL, Dubchak I, Garcia-Fernandez J, Gibson-Brown JJ, Grigoriev IV, Horton AC, de Jong PJ, Jurka J, Kapitonov VV, Kohara Y, Kuroki Y, Lindquist E, Lucas S, Osoegawa K, Pennacchio LA, Salamov AA, Satou Y, Sauka-Spengler T, Schmutz J, Shin IT, Toyoda A, Bronner-Fraser M, Fujiyama A, Holland LZ, Holland PW, Satoh N, Rokhsar DS (2008) The amphioxus genome and the evolution of the chordate karyotype. Nature 453:1064–1071. https://doi.org/10.1038/nature06967
Satou Y, Kawashima T, Shoguchi E, Nakayama A, Satoh N (2005) An integrated database of the ascidian, Ciona intestinalis: Towards functional genomics. Zool Sci 22:837–843. https://doi.org/10.2108/zsj.22.837
Satou Y, Nakamura R, Yu D, Yoshida R, Hamada M, Fujie M, Hisata K, Takeda H, Satoh N (2019) A nearly complete genome of Ciona intestinalis type A (C. robusta) reveals the contribution of inversion to chromosomal evolution in the genus Ciona. Genome Biol Evol 11:3144–3157. https://doi.org/10.1093/gbe/evz228
Satou Y, Tokuoka M, Oda-Ishii I, Tokuhiro S, Ishida T, Liu B, Iwamura Y (2022) A Manually Curated Gene Model Set for an Ascidian, Ciona robusta (Ciona intestinalis Type A). Zool Sci 39:253–260. https://doi.org/10.2108/zs210102
Schlosser G (2014) Early embryonic specification of vertebrate cranial placodes. Wires Dev Biol 3:349–363. https://doi.org/10.1002/wdev.142
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675. https://doi.org/10.1038/nmeth.2089
Singh S, Groves AK (2016) The molecular basis of craniofacial placode development. Wiley Interdiscip Rev Dev Biol 5:363–376. https://doi.org/10.1002/wdev.226
Steventon B, Mayor R, Streit A (2014) Neural crest and placode interaction during the development of the cranial sensory system. Dev Biol 389:28–38. https://doi.org/10.1016/j.ydbio.2014.01.021
Stolfi A, Ryan K, Meinertzhagen IA, Christiaen L (2015a) Migratory neuronal progenitors arise from the neural plate borders in tunicates. Nature 527:371–374. https://doi.org/10.1038/nature15758
Stolfi A, Sasakura Y, Chalopin D, Satou Y, Christiaen L, Dantec C, Endo T, Naville M, Nishida H, Swalla BJ, Volff JN, Voskoboynik A, Dauga D, Lemaire P (2015b) Guidelines for the nomenclature of genetic elements in tunicate genomes. Genesis 53:1–14. https://doi.org/10.1002/dvg.22822
Tokuoka M, Maeda K, Kobayashi K, Mochizuki A, Satou Y (2021) The gene regulatory system for specifying germ layers in early embryos of the simple chordate. Sci Adv 7:eabf8210. https://doi.org/10.1126/sciadv.abf8210
Tribulo C, Aybar MJ, Nguyen VH, Mullins MC, Mayor R (2003) Regulation of Msx genes by a Bmp gradient is essential for neural crest specification. Development 130:6441–6452. https://doi.org/10.1242/dev.00878
Wagner E, Levine M (2012) FGF signaling establishes the anterior border of the Ciona neural tube. Development 139:2351–2359. https://doi.org/10.1242/dev.078485
Wagner E, Stolfi A, Choi YG, Levine M (2014) Islet is a key determinant of ascidian palp morphogenesis. Development 141:3084–3092. https://doi.org/10.1242/dev.110684
Waki K, Imai KS, Satou Y (2015) Genetic pathways for differentiation of the peripheral nervous system in ascidians. Nat Commun 6:8719. https://doi.org/10.1038/ncomms9719
Acknowledgements
We thank Reiji Masuda, Shinichi Tokuhiro, Chikako Imaizumi (Kyoto University), Manabu Yoshida (University of Tokyo), and other members working under the National BioResource Project for Ciona (MEXT, Japan) at Kyoto University. We also thank the University of Tokyo for providing experimental animals.
Funding
This research was supported by grants from the Japan Society for the Promotion of Science under the grant numbers 21H02486 and 21H05239 to YS.
Author information
Authors and Affiliations
Contributions
BL and XR performed experiments. BL and YS drafted the original manuscript. BL, XR, and YS reviewed the manuscript draft, revised it, and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Communicated by Hiroki Nishida
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, B., Ren, X. & Satou, Y. BMP signaling is required to form the anterior neural plate border in ascidian embryos. Dev Genes Evol 233, 13–23 (2023). https://doi.org/10.1007/s00427-023-00702-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-023-00702-0