Abstract
Notch signaling is a crucial cog in early development of euechinoid sea urchins, specifying both non-skeletogenic mesodermal lineages and serotonergic neurons in the apical neuroectoderm. Here, the spatial distributions and function of delta, gcm, and hesc, three genes critical to these processes in euechinoids, are examined in the distantly related cidaroid sea urchin Eucidaris tribuloides. Spatial distribution and experimental perturbation of delta and hesc suggest that the function of Notch signaling in ectodermal patterning in early development of E. tr ibuloides is consistent with canonical lateral inhibition. Delta transcripts were observed in t he archenteron, apical ectoderm, and lateral ectoderm in gastrulating e mbryos of E. tribuloides. Perturbation of Notch signaling by either delta morpholino or treatment of DAPT downregulated hesc and upregulated delta and gcm, resulting in ectopic expression of delta and gcm. Similarly, hesc perturbation mirrored the effects of delta perturbation. Interestingly, perturbation of delta or hesc resulted in more cells expressing gcm and supernumerary pigment cells, suggesting that pigment cell proliferation is regulated by Notch in E. tribuloides. These results are consistent with an evolutionary scenario whereby, in the echinoid ancestor, Notch signaling was deployed in the ectoderm to specify neurogenic progenitors and controlled pigment cell proliferation in the dorsal ectoderm.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In multicellular organisms the Notch signaling pathway functions to regulate cell fates by mediating short-range, cell-cell interactions when cells are directly apposed with or in close proximity to one another (Artavanis-Tsakonas et al. 1999). The Notch/LIN-12 protein family comprises single-pass transmembrane receptors that, when tickled by their Delta/Serrate/LAG-2 ligands bound to membranes of neighboring cells, causes release and subsequent intracellular processing of the Notch intracellular domain (NICD) by γ-secretase (Kopan and Ilagan 2009). Once cleaved, the NICD translocates to the nucleus and interacts with members of the Suppressor of Hairless(SuH)/LAG-1 protein family to facilitate DNA binding and alter gene expression. In canonical Notch signaling, members of the Hairy and Enhancer of Split (HES) protein family are present in signal-receiving cells as principal targets of Notch, by which they mediate transcriptional repression of genes expressed in signal-sending, neighboring cells. Notch signaling has been shown to act in diverse metazoan developmental contexts, from the maintenance of neural precursors and neurogenic placodes to the specification of mesenchymal cell lineages, as well as the spacing of sensory organs (Bray 2016).
In early development of euechinoid sea urchins, Notch signaling is deployed as a patterning mechanism in endomesodermal and ectodermal domains. In the endomesoderm of the euechinoid Lytechinus variegatus (Lv), early cleavage stage embryos exhibit uniform levels of Notch protein until early blastula stage, at which time Notch becomes concentrated in a ring of cells near the vegetal plate (Sherwood and McClay 1997). Subsequent experiments revealed that Notch segregates non-skeletogenic mesoderm (NSM) from cells that will become anterior endoderm (Sherwood and McClay 1999). Correspondingly, the absence of Delta ligand, which is expressed specifically in NSM-adjacent skeletogenic mesenchyme (SM), results in complete loss of NSM-precursor lineages that later become pigment cells and other mesenchymal cell types (Sweet et al. 2002). Similarly in Strongylocentrotus purpuratus (Sp), another camarodont euechinoid, perturbation of Notch signaling results in loss of activation of NSM-specific regulatory genes, e.g., glial cells missing (gcm), as well as loss of small micromere-specific genes in neighboring small micromeres (Materna et al. 2013a; Materna et al. 2013b; Ransick and Davidson 2006). Downregulation of NSM regulatory genes results in the failure to clear endodermal regulatory genes from mesodermal domains (Croce and McClay 2010; Materna and Davidson 2012; Peter and Davidson 2011). Data from other camarodont euechinoid taxa, viz. Paracentrotus lividus (Pl) and Hemicentrotus pulcherrimus (Hp) (Ohguro et al. 2011; Rottinger et al. 2006), as well as an irregular euechinoid, Astriclypeus manni (Takata and Kominami 2011), suggest that this early developmental capacity of Notch signaling to specify NSM is a highly conserved feature of euechinoids, dating back at least to the divergence of irregular and non-irregular echinoids more than 210 mya (Smith et al. 2006).
In the ectoderm and later in the endoderm of the euechinoid embryo, Notch signaling functions in a highly conserved role of recruiting neurogenic progenitor cells in the animal pole domain (APD) and ciliary band (CB) (Burke et al. 2014). In Sp and Pl, delta transcripts are expressed during gastrulation in isolated cells of the apical neuroectoderm (ANE) (Lapraz et al. 2009; Wei et al. 2011; Yaguchi et al. 2012; Yaguchi et al. 2011). Perturbation of Notch results in ectopic patches of serotonergic neurons in those domains, suggesting that Delta/Notch functions to laterally inhibit signal-receiving cells. Indeed, Notch-mediated lateral inhibition recruits neuronal progenitors in APD and in lateral ectoderm (LE), where bilateral postoral neurogenic cells are recruited from CB (Garner et al. 2016; Mellott et al. 2017). Moreover, in the closely related euechinoid Hp and the distantly related irregular euechinoid Scaphechinus mirabilis (Sm), delta transcripts are also expressed at mesenchyme blastula in APD, as well as bilaterally in isolated cells in LE, indicating that these echinoids also may possess an ANE domain and bilateral postoral cells (Yamazaki et al. 2010). Hence, in euechinoids, Notch signaling functions as a conserved spatial regulatory mechanism in at least three distinct domains during embryogenesis: in the endomesoderm for cell fate specification and lineage segregation, in the APD to regulate proliferation of neuronal progenitors, and in LE to recruit neuronal progenitors from CB.
The euechinoid taxa described above are members of one great branch of a two pronged lineage of extant sea urchins—the other lineage being the cidaroid echinoids. These two sister groups, cidaroids and euechinoids, diverged from each other at least 268 mya (Thompson et al. 2015). Studies focusing on the function and spatiotemporal deployment of regulatory genes in early development of cidaroids are beginning to reveal the degree to which developmental gene regulatory networks change in deep time (Bishop et al. 2013; Erkenbrack 2016; Erkenbrack et al. 2016; Erkenbrack and Davidson 2015; Yamazaki et al. 2014). For instance, it is now clear that the endomesodermal role of Notch signaling in early development of the cidaroid echinoids Eucidaris tribuloides (Et) and Prionocidaris baculosa (Pb) exhibits important functional differences when contrasted to euechinoids. In blastular stages of Et, delta transcripts are present in micromere-descendants, a homologous cell type to euechinoid SM and that also expresses the SM regulatory gene alx1. Neighboring signal-receiving cells are comprised of NSM that abut the SM. After the onset of delta expression, NSM begin to accumulate transcripts of the HES-1 protein family member hesc. When delta translation is disrupted, expression of hesc is extinguished in SM-neighboring cells, and the SM domain expands into surrounding NSM (Erkenbrack and Davidson 2015). Rather than a cell lineage segregation mechanism as in euechinoids, a canonical Notch lateral inhibition mechanism is deployed whereby delta is confined to SM by Notch-mediated HES protein family members. Thus, the endomesodermal role of Notch diverged after the cidaroid-euechinoid split.
To date, no studies have addressed these mechanisms in the distantly related cidaroid lineage. The study by Erkenbrack and Davidson (2015) showed delta and hesc transcripts in the ectoderm of Et at early gastrula stage, hinting that Notch signaling also may play a role during ectodermal patterning in Et. Here, I elaborate on that preliminary observation by presenting spatial and perturbation data on three core regulatory genes—delta, hesc, and gcm—involved in euechinoid neurogenic ectodermal progenitor specification and mesenchymal pigment cell specification during gastrulation of Et. These results suggest that the predominant mechanism of Notch signaling in Et is via canonical lateral inhibition. The spatial confinement of delta transcripts is under control of this mechanism in the archenteron, the APD, and in isolated cells in LE. These cells in LE are likely homologous to euechinoid postoral cells in CB. The regulatory mechanisms installing hesc transcripts in the ectoderm are complex, consisting of one that is tightly linked to the spatial localization of delta, and one that is independent of delta expression. Perturbation of both delta and hesc results in ectopic expression of delta. Delta perturbation or disruption of Notch downregulates zygotic hesc. Intriguingly, ectodermally confined gcm expression was sensitive to perturbation of Notch and hesc, increasing the abundance of gcm transcripts, gcm-positive cells, and pigment cells. These results suggest that the spatial confinement and proliferation of gcm in LE during Et gastrulation are Notch-dependent. Lastly, employing a comparative developmental framework (Table 1), these data paint a picture of the evolution of Notch signaling in early development of echinoderms.
Materials and methods
Animals, cloning, whole-mount in situ hybridization, qPCR
Animals were obtained from SeaLife, Inc. (KP Aquatics, FL) and were maintained at room temperature. Embryos were cultured in Millipore-filtered seawater (MFSW) in temperature controlled conditions at 22 °C. Full-length or partial coding sequences for delta, hesc, and gcm were obtained by PCR from a cDNA library comprised of mixed developmental time points. Primer sequences for qPCR and for whole-mount in situ hybridization (WMISH) probe construction for delta, hesc, and gcm have been previously described (Erkenbrack 2016; Erkenbrack and Davidson 2015). Full-length or partial PCR products were subsequently cloned into E. coli by standard procedures. Plasmids were isolated and sequenced for confirmation of the correct insert. Detailed protocols for WMISH and qPCR were previously described (Erkenbrack 2016; Erkenbrack and Davidson 2015).
Morpholino perturbations
A morpholino microinjection protocol modified from Materna (2017) was carried out as previously described (Erkenbrack and Davidson 2015). Morpholinos targeting delta and hesc translation start sites were synthesized by GeneTools (Philomath, OR, USA), and their sequences are 5′-ATAACATATAGCACGCCGAGAAGGC-3′ and 5′-AATCACAAGGTAAGACGAGGATGGT-3′, respectively. Injection solutions of each morpholino were 1 mM, and approximately 10 pL was injected into each zygote. Embryos were then cultured to the desired time point and processed either for qPCR via RNA isolation (RNeasy; Qiagen) and subsequent cDNA synthesis (iScript cDNA Synthesis; Bio-Rad) or for WMISH by overnight fixation in MOPS-buffered MFSW with 4% paraformaldehyde. WMISH and qPCR analyses were replicated twice in the case of morpholinos and three times in the case of DAPT treatment (Supplementary Table 1).
Treatment of embryos with DAPT
For treatment with N-[(3,5-difluorophenyl)acetyl]-L-alanyl-2-phenyl]glycine-1,1-dimethyl ester (DAPT) (cat no. 2634; Tocris Biosciences), embryos developing in MFSW were transferred at 2 h postfertilization (hpf) into an equal volume of MFSW containing 20 μM DAPT, giving a final concentration of 10 μM DAPT. Embryos were allowed to develop to the desired stage and fixed either for qPCR or WMISH as described above.
Results
Spatial distribution of mRNAs encoding delta, gcm, and hesc during E. tribuloides gastrulation
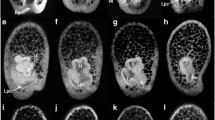
As the archenteron extends during early gastrulation in Et, and just prior to the ingression of mesenchyme, delta transcripts are observed in the archenteron and in isolated cells in LE (Fig. 1a) (Erkenbrack and Davidson 2015). At 36 hpf, as mesenchyme invades the blastocoel, delta transcripts are seen in APD, in addition to the archenteron and LE (Fig. 1a). By 50 h, delta increases in ectoderm that will become CB. At 28 h, prior to ingression of mesenchyme, gcm is detected just below the anterior end of the archenteron, as well as in isolated cells in LE (Fig. 1b) (Erkenbrack 2016; Erkenbrack and Davidson 2015). By mid-gastrula gcm-positive cells are in the blastocoel and in LE (Fig. 1b). At 50 h, gcm is seen in the future CB and is in a small cluster in the anterior archenteron (Fig. 1b). Hesc transcripts are detected in a complex distribution throughout the ectoderm at 28 h, as well as in the archenteron (Fig. 1c). Independent of its expression in the NSM during late blastula stages, hesc is installed gradually in the ectoderm beginning from mid-blastula stage to early gastrula, when it is observed in a mottled pattern (Erkenbrack and Davidson 2015). At 28 h, hesc is conspicuously absent from particular anterior regions of the embryo and from a ring of 5–6 cell diameters surrounding the blastopore (Fig. 1c). At 36 h, hesc transcripts are detected in the archenteron and are more localized in the APD and in LE (Fig. 1c). By 50 h, hesc transcripts are seen in the future CB and in the archenteron and are cleared from the most apical region of the APD as well as the area near the blastopore (Fig. 1c). The spatial distribution of hesc transcripts in Et agrees well with those observed in the closely related, Pacific-dwelling cidaroid Pb (Yamazaki et al. 2014).
Spatial distribution of mRNAs encoding delta, gcm, and hesc during E. tribuloides gastrulation. a At early gastrula, delta is expressed at the anterior end of the archenteron and in a few isolated ectodermal cells near the boundary of endoderm and ectoderm. As mesenchyme ingresses at 36 h, delta is detected as at 28 h but also in isolated cells at the apical plate. By mid-gastrula at 50 h, delta is detected in cells just below the larval equator, as well as the apical plate and archenteron. b At early gastrula, gcm is detected in a patch of cells just under the anterior end of the archenteron and in isolated cells in the ectoderm. By 36 h, gcm is seen also in mesenchymal cells ingressing into the blastocoel. Apical views of embryos at this stage show the restriction of gcm to one side of the archenteron and its expression in isolated ectodermal cells more near the blastopore. At mid-gastrula, gcm is detected in cells at the tip of the archenteron as well as in scattered ectodermal cells including the larval equator. c Hesc at early gastrula is detected in the endoderm, ectoderm, and mesoderm. At this stage, hesc is spatially distributed throughout much of the ectoderm with isolated patches cleared of its expression. At 36 h, hesc is broadly expressed as at 28 h, with some cells in the mesoderm and the region near the blastopore void of its expression. By 50 h, hesc is detected in cells near the region where the ciliary band will form in the apical plate and between the larval equator and blastopore, as well as in the archenteron. Arrows show detected staining. Bars show regions of the embryo cleared of transcripts. Dashed lines indicate regions of the larva that were also imaged from the apical view shown immediately to the right and with a corresponding number. AV apical view, RV archenteron view
Notch signaling regulates delta and locally regulates hesc expression
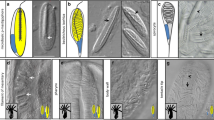
Previous work indicated that transcription of delta increases in pregastrular Et embryos injected with delta morpholino, suggesting that either delta is increasing in the micromere descendants or that its domain of expression is expanding (Erkenbrack and Davidson 2015). As with the SM-specific regulatory gene alx1, the domain of delta indeed expands into the NSM prior to gastrulation (Fig. S1(a)). Later, at 36 h, delta continues to increase in embryos injected with delta morpholino (Fig. 2a). Spatially, delta morpholino resulted in ectopic delta expression in the APD, LE, and archenteron (Figs. 2b, S1(b), and S2(a)). This was also the case for embryos cultured to 50 hpf (Figs. 2c and S1(c)). Moreover, the Notch-signaling responsive regulatory gene hesc showed spatially specific downregulation in the archenteron and the ectoderm at 26 h (Figs. 2d and S1(d)). Correspondingly, disruption of NICD cleavage by culturing embryos in the presence of the γ-secretase inhibitor DAPT increased delta mRNA and induced ectopic expression of delta (Figs. 2e, g and S3(a)). As observed in delta MASO background, hesc continued to show localized downregulation in the archenteron and depressed expression in ectoderm in DAPT-treated embryos (Figs. 2g and S3(b)). These results suggest (1) that Notch signal-receiving cells express a repressor, likely hesc, that functions by lateral inhibition to confine delta transcripts to Notch signal-sending cells, and (2) that there likely exists a local activator of delta in these regions where ectopic expression is occurring.
Perturbation of Notch signaling by blocking translation of Delta ligand or by culturing embryos in the presence of DAPT increases the abundance and results in ectopic expression of delta and gcm. a Delta morpholino (MASO) increases RNA abundance of delta and gcm. b Spatial distribution of delta and gcm at 36 h in delta MASO background and uninjected controls. Whereas the number of delta-positive cells increases in both the apical ectoderm and the endodermal-ectodermal boundary, the number of gcm-positive cells only increases in the latter region. c At 50 h, embryos injected with delta MASO show well-defined regions of delta expression, expanding domains in the mesoderm, apical ectodermal, and endodermal-ectodermal boundary. d Expression of hesc is extinguished specifically in the archenteron in the presence of delta MASO. e Abundance of delta and gcm RNA increases in embryos cultured in the presence of 10 μM DAPT. f At 26 h, the number of ectodermal gcm-positive cells increases in the presence of DAPT. g At 50 h, DAPT-treated embryos exhibit increases in both delta- and gcm-positive cells. This treatment mirrors that seen in delta MASO. The expression of hesc is reduced in the mesoderm at the archenteron. RV archenteron view. Red dashed line in qPCR graphs indicates a difference of 0.7 cycles or 1.6-fold change
Ectopic expression of gcm is observed in the lateral ectoderm in delta morpholino background
In Et, lineage tracing studies and WMISH have indicated that SM is the first mesenchyme to ingress, and that this only occurs well after the archenteron has extended into the blastocoel at early gastrula (Erkenbrack and Davidson 2015; Urben et al. 1988; Wray and McClay 1988). These lines of evidence make the previously observed presence of gcm-positive cells in LE all the more interesting (Erkenbrack 2016), since in euechinoids gcm-positive cells come solely from Notch signal-receiving cells in the endomesoderm (Ransick and Davidson 2006). However, since we do not know the source of gcm-positive cells in Et, we cannot definitively say whether these cells in LE are of mesodermal origin or ectodermal origin. Delta transcripts are also present in LE at this time and show ectopic expression upon perturbation of Notch signaling (Figs. 1a and 2b). Similar to delta, gcm expression was increased in delta MASO background (Fig. 2a) and exhibited ectopic expression in LE at 36 h (Figs. 2b and S2(b)). Embryos cultured in DAPT also showed increased gcm abundance and ectopic expression at three different time points (Figs. 2e–g and S3(c,d)). Importantly, these data do not indicate whether the spatial expression of gcm is also changing in the archenteron.
Ectopic expression of delta and gcm upon perturbation of Hesc
The data described thus far suggest that Notch signal-receiving cells come to express the obligate repressor Hesc, which in turn confines delta and gcm transcripts in different regions of the embryo to isolated ectodermal cells. Therefore, delta and gcm should respond in a similar fashion to hesc morpholino. At 36 h, hesc, gcm, and hesc transcripts increased in Et embryos injected with hesc morpholino (Fig. 3a). At 36 and 50 h, embryos injected with hesc MASO induced ectopic expression of delta transcripts in APD as well as in LE (Figs. 3b, c and S4(a)). Similarly gcm transcripts exhibited ectopic expression in LE (Figs. 3b, d and S4(b,c)). These results suggest that Notch signal-receiving cells restrict the spatial distribution of delta and gcm to isolated cells in the archenteron and ectoderm. In the case of gcm, ectopic expression is induced, minimally, in LE; whether or not a similar effect occurs where gcm is expressed in the archenteron will require additional experimental data.
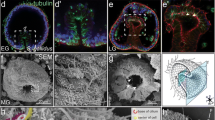
Perturbation of hesc results in increased abundance of delta and gcm transcripts, as well as supernumerary pigment cell formation. a Microinjection of hesc MASO increases delta, gcm and hesc RNA transcripts at 36 h. b Embryos injected with hesc MASO exhibit ectopic delta-positive cells in the apical ectoderm as well as near the endodermal-ectodermal boundary. An increase in gcm-positive cells is observed near the vegetal lateral clusters in the ectoderm at 36 h. c, d At 50 h, the spatial domains of delta and gcm expand in the apical ectoderm and near the vegetal lateral clusters, respectively, in hesc MASO background. e Microinjection of hesc MASO resulted in embryos with increased pigment cell (PC) numbers, on average, relative to uninjected control. ***p < 0.01, Mann-Whitney U test. f Example embryos in uninjected control and hesc MASO background showing pigment cells at 120 h
Induction of supernumerary pigment cells in hesc perturbation background
In euechinoids, Delta presentation in SM directly regulates specification of NSM via Notch-mediated regulatory control of gcm and other NSM regulatory genes (Materna and Davidson 2012; Ransick and Davidson 2006; Ransick and Davidson 2012; Sherwood and McClay 1999; Sweet et al. 2002). Perturbation of either Notch signaling or gcm translation produces albino embryos (Ransick and Davidson 2006; Sherwood and McClay 1999). As seen above, in Et the early transcription of gcm does not depend on Notch for its activation, rather its spatial confinement depends on Notch. Lastly, I sought to determine if changes in gcm abundance affected pigment cell specification. Intriguingly, embryos cultured in the presence of hesc morpholino exhibited supernumerary pigment cells (Fig. 3e, f), suggesting that hesc perturbation increased the number of gcm-positive cells. These data do not indicate whether supernumerary pigment cells are coming from gcm-positive cells in LE or the archenteron (Fig. 4). On average, there were 15–30 additional pigment cells in embryos injected with hesc MASO (Fig. 3e), which is consistent with the hypothesis that a small population of cells in the embryo are transfated to become pigment cells.
Models and hypothetical scenarios for the installation and function of Notch signaling components and gcm in early development of Eucidaris tribuloides. a Installation of zygotic hesc transcripts comes in two forms in E. tribuloides from mid-blastula to mid-gastrula. Expression of hesc (dark gray) abutting delta-positive cells (light blue) is sensitive to Notch perturbation, whereas most hesc expression in the ectoderm are insensitive to Notch perturbation (light gray) and is installed independent of Notch. b Ontogenetic source of gcm-positive cells in the lateral ectoderm of E. tribuloides. Three scenarios are shown: in the first, transcription factors (TFs) in the lateral ectoderm activate gcm expression independently of its mesodermal activity; in the second, mesodermally derived gcm-positive cells migrate directly through the endoderm to arrive at LE; in the third, a pregastrular ingression of mesodermally derived gcm-positive cells migrates through the blastocoel to LE
Discussion
Spatial installation of hesc in E. tribuloides and Hesc regulation of delta
In euechinoids, Hesc functions globally throughout the early embryo to repress SM specific genes, e.g., delta, until another repressor, Pmar1, relieves that repression specifically in the large micromeres, whose ontogenetic descendants are SM (Oliveri et al. 2008; Revilla-i-Domingo et al. 2007). This double-negative gate circuitry is a highly conserved feature of euechinoid sea urchins (Minokawa 2017; Thompson et al. 2017). Perturbation of Hesc activity results in global expression of delta transcripts (Revilla-i-Domingo et al. 2007). In contrast, it was previously reported that Hesc perturbation in Et does not result in global expression of the SM-specific gene alx1 (Erkenbrack and Davidson 2015). Here, we see delta responding similarly to Hesc perturbation as did alx1 in previous reports, suggesting that there is no global driver of SM gene expression in Et. Furthermore, these data argue for two distinct modes of spatial regulation of hesc transcription: a Notch-mediated mode in the endomesoderm, the APD, and LE, and a second Notch-independent mode of regulation that is installed in the ectoderm from late blastula and beyond (Fig. 4a). Given Hesc is a repressor of delta transcription and since hesc transcripts are present throughout most of the ectoderm, it is still not clear how delta transcription is activated if Hesc is uniformly expressed. Thus, in order for delta expression to be turned on in isolated APD and LE cells, hesc must either be kept transcriptionally silent in particular regions of the ectoderm or, if hesc is globally expressed, regions of the ectoderm must be cleared of Hesc. Unfortunately, the data presented here do not allow for resolution of this issue, and the only thing that is clear is that Hesc perturbation does not expand the domain of delta and alx1 throughout the embryo. One possible solution is that Hesc protein is regulated locally by posttranslational modifications, as phosphorylation of HES proteins is known to interfere with their DNA-binding activity (Popovic et al. 2014; Strom et al. 1997). Thus, even if hesc transcripts are expressed throughout the ectoderm, their mere presence does not allow us to rule out posttranslational regulation of Hesc protein.
A conserved role for Hesc in restricting delta and gcm during ectoderm development
One scenario suggested by these data is that delta and gcm are co-expressed in LE. This would not be the first instance of this in the echinoid clade. The simultaneous transcriptional activation of delta and gcm has been previously observed during early development of the irregular echinoid Sm (Yamazaki et al. 2010; Yamazaki and Minokawa 2016), as well as the likely co-expression of delta and gcm in the micromeres of the spatangoid echinoid Echinocardium cordatum (Yamazaki and Minokawa 2015). Indeed, it has been reported that Hesc also functions upstream of delta and gcm during NSM and SM specification in Sm and Hp (Yamazaki and Minokawa 2016). These studies showed that Hesc perturbation resulted in expanding domains of delta and gcm, as well as an increase in the number of pigment cells in these echinoids. Here, in the cidaroid Et, we also see an increase in pigment cells as well as expanding domains of spatial expression of delta and gcm. From an evolutionary perspective, these data argue for conserved gene regulatory network (GRN) circuitry linking delta, hesc, and gcm in both SM and NSM specification since the divergence of cidaroids and euechinoids. Furthermore, similarities in the regulatory connectivity of these three genes in three distantly related taxa suggest that canonical Notch lateral inhibition likely plays a widespread role in echinoid NSM specification.
Gcm in pigment cell specification GRN cicruitry in echinoids
The transcriptional regulation of gcm is one of the most thoroughly dissected cis-regulatory loci in the Sp developmental GRN (Ransick and Davidson 2006; Ransick and Davidson 2012). Gcm is a crucial regulator of pigment cell specification, the regulatory state of which is locked down by an intergenic feedback circuit involving Gcm, Gatae, and Six1/2 (Ransick and Davidson 2012). During NSM specification in euechinoids, gcm activation and transcription depend entirely upon Delta presentation in SM and subsequent NICD nuclearization (Ransick and Davidson 2006). Data reported here and elsewhere (Erkenbrack 2016) show gcm-positive cells near the endodermal-ectodermal boundary prior to mesenchymal ingression in Et. One question remaining is whether or not the gcm-positive cells in the LE are truly pigment cells. This question can likely be addressed by surveying the regulatory state of those cells for gatae and six1/2. Unfortunately, no data are published for six1/2 in Et. However, there are data for gatae (Erkenbrack 2016). At the time of invagination, gatae is expressed in endodermal and mesodermal domains and expression is not observed near the ectoderm until after mesenchyme begins to ingress into the blastocoel. Therefore, if we assume that the intergenic stabilization loop Gcm-Gatae-Six1/2 is a marker for pigment cell specification, then it is plausible that the gcm activity we are seeing in the ectoderm is not related to a pigment cell lineage in Et. Indeed, the data from Erkenbrack (2016) show that gcm and gatae may be expressed in an overlapping domain at the tip of the archenteron in Et, which may correlate with its mesenchymal-derived pigment cell lineage. Double in situ hybridization combined with perturbation of Gcm, Gatae, and Six1/2 are needed in Et to resolve this interesting question.
The ontogenetic origin of gcm-positive cells in the E. tribuloides ectoderm
The ontogenetic source of gcm-positive cells populating LE is unclear. In camarodont euechinoids, gcm-positive pigment cell precursors arise from blastocoelar ingression of NSM and their subsequent migration from the archenteron to ectoderm at mid-gastrula stage (Gibson and Burke 1985). The most likely scenario in Et is that gcm- and gatae-positive cells ingressing at the tip of the archenteron are the source of all of the pigment cells in Et. If that is the case, then perhaps gcm serves a distinct regulatory function in LE of Et, as well as in LE of the closely related cidaroid Pb (Yamazaki et al. 2014). Still, we would like to know whether these gcm-positive cells in LE are mesodermally or ectodermally derived. Unfortunately, the scope of this study could not include experiments that specifically addressed this question. However, there are a few scenarios that can be framed around it based on what is seen in non-camarodont euechinoids (Fig. 4b). The presence of a single population of gcm-positive cells around the time of gastrulation in Et (Erkenbrack 2016) suggests that either gcm-positive cells begin to migrate to LE shortly after gastrulation begins or gcm transcription begins in LE independently of its NSM activation. As previously mentioned, lineage tracing studies make it clear that mesenchymal ingression in Et does not occur until the archenteron has passed two thirds of the way into the blastocoel (Urben et al. 1988; Wray and McClay 1988; Wray and McClay 1989); hence, while it is unlikely that gcm-positive cells are ingressing before skeletogenic mesenchyme in Et, it is still a possibility. This hypothesis could be called the furtive ingression hypothesis and, importantly, cannot be ruled out. Another scenario is direct migration from NSM through the endoderm without ingressing into the blastocoel (Fig. 4b). Two observations make this scenario plausible: (1) it is known that pigment cell precursors in Hp invade the ectoderm at the apical plate and then begin their migration through the aboral ectoderm without ingressing (Kominami et al. 2001) and (2) in the euechinoids Toxopneustes pileolus and Anthocidaris crassispina, populations of pigment cells in euechinoids migrate directly from the vegetal plate in the direction of the apical plate through the aboral ectoderm without ingressing (Takata and Kominami 2004). However, it is unclear how a few cells of a large population of gcm-positive cells would become primed to migrate to LE. The simplest interpretation is that gcm turns on independently in LE where the neuronal progenitors arise in the future CB domain, and that the spatial localization is due to convergence of numerous signaling pathways near the endodermal-ectodermal boundary (Range 2014).
Notch signaling as a conserved mechanism specifying neuronal progenitors in echinoids
Notch-mediated neurogenesis in the APD of euechinoids has become an important model contributing to our understanding of the evolution of neurogenesis in metazoans (Burke et al. 2014). Numerous studies have revealed mechanisms underlying the specification of euechinoid larval neurons (Burke et al. 2014; Garner et al. 2016; Lapraz et al. 2009; Mellott et al. 2017; Wei et al. 2011; Yaguchi et al. 2012; Yaguchi et al. 2011). This study, along with another previous study (Bishop et al. 2013), contributes to the evolutionary story of Notch-mediated ectodermal patterning in cidaroids. In euechinoids, serotonergic neurons develop on the aboral side of the APD, and Notch functions to laterally inhibit adjacent, signal-receiving cells from differentiating into neurons (Yaguchi et al. 2012). Perturbation of Notch in euechinoids increases the number of serotonergic neurons and expands the domains of neural markers in the APD and in postoral cells positioned in CB (Mellott et al. 2017; Yaguchi et al. 2012; Yaguchi et al. 2011). Notably, in Patiria miniata, delta knockdown expands the domain of the neural marker soxc, suggesting a lateral inhibition mechanism is also at play in asteroids (Yankura et al. 2013). In Et, it was previously shown that serotonergic neural markers are known to be bilaterally expressed in the APD and in CB; later in development, these cells develop into clusters of four or five neurons each (Bishop et al. 2013). Here, I show that Notch-mediated lateral inhibition is active precisely in the locations as those previously shown to exhibit neuronal markers. Furthermore, in the case of the delta-positive cells in LE, these cells are positioned in spatially homologous locations in embryos of cidaroids and euechinoids, precisely where CB will later form. These cells are very likely to be components of the CB ganglionic nervous system and are likely to be a homologous cell type to those previously described as neurogenic postoral cells in euechinoids (Burke et al. 2014). Similarly, as in the euechinoids Sp, Pl, and Hp (Materna et al. 2013a; Rottinger et al. 2006; Yaguchi et al. 2012), delta is expressed in Et in clustered cells within APD (Fig. 4a). Upon perturbation of Notch, delta expands, indicating a canonical lateral inhibition mechanism of Notch signaling in Et apical neuroectoderm, which is also the case in euechinoids. Integrating these previous lines of evidence with the results presented here makes a very strong case for conserved ontogenetic deployment and function of Notch signaling in both APD and postoral cells in LE of echinoids.
Notch signaling as a model GRN plugin in early development
Intercellular signals have been described as GRN plugins that can be deployed as switches in distinct spatial embryonic addresses by altering their regulatory control during development (Davidson and Erwin 2006; Erwin and Davidson 2009). Affording support for this notion is the partial survey provided in the above description of Notch signaling in early development of echinoids as well as data from echinoderms more broadly. Comparative analyses of developmental GRNs between cidaroids and euechinoids are revealing how plugins and other GRN subcircuits are rewired and altered over vast geological timeframes (Thompson et al. 2017; Thompson et al. 2015), how the regulatory systems that install them evolve (Erkenbrack 2016; Gao et al. 2015), and how embryonic developmental programs likely functioned in the most recent common ancestors of extant sea urchins (Erkenbrack et al. 2016; Erkenbrack and Petsios 2017). Evidence collected from crown group echinoderms makes an illustrative case study showing how regulatory circuitry coupling Notch signaling and gcm appears to have been shuffled around in this manner during the evolution of eleutherozoan echinoderm lineages over the last 500 million years. Notch signaling components have been deployed in some aspect of specification of all spatial domains in echinoderms—endoderm (Hinman and Davidson 2007), mesodermal SM restriction (Erkenbrack and Davidson 2015), mesodermal NSM specification (Sherwood and McClay 1999), and ectodermal and endodermal specification of serotonergic neurons and neuronal progenitors (Burke et al. 2014; Garner et al. 2016; Mellott et al. 2017; Wei et al. 2011; Yaguchi et al. 2012). This report adds further evidence to the pliant nature of Notch signaling in development of echinoid neuronal and mesodermal cell lineages, showcasing both the conserved hierarchical nature of developmental GRNs as well as their marked plasticity.
References
Artavanis-Tsakonas S, Rand MD, Lake RJ (1999) Notch signaling: cell fate control and signal integration in development. Science 284(5415):770–776. https://doi.org/10.1126/science.284.5415.770
Bishop CD, MacNeil KE, Patel D, Taylor VJ, Burke RD (2013) Neural development in Eucidaris Tribuloides and the evolutionary history of the echinoid larval nervous system. Dev Biol 377(1):236–244. https://doi.org/10.1016/j.ydbio.2013.03.006
Bray SJ (2016) Notch signalling in context. Nat Rev Mol Cell Biol 17(11):722–735. https://doi.org/10.1038/nrm.2016.94
Burke RD, Moller DJ, Krupke OA, Taylor VJ (2014) Sea urchin neural development and the metazoan paradigm of neurogenesis. Genesis 52(3):208–221. https://doi.org/10.1002/dvg.22750
Croce JC, McClay DR (2010) Dynamics of Delta/Notch signaling on endomesoderm segregation in the sea urchin embryo. Development 137(1):83–91. https://doi.org/10.1242/dev.044149
Davidson EH, Erwin DH (2006) Gene regulatory networks and the evolution of animal body plans. Science 311(5762):796–800. https://doi.org/10.1126/science.1113832
Erkenbrack EM (2016) Divergence of ectodermal and mesodermal gene regulatory network linkages in early development of sea urchins. Proc Natl Acad Sci U S A 113(46):E7202–E7211. https://doi.org/10.1073/pnas.1612820113
Erkenbrack EM, Ako-Asare K, Miller E, Tekelenburg S, Thompson JR, Romano L (2016) Ancestral state reconstruction by comparative analysis of a GRN kernel operating in echinoderms. Dev Genes Evol 226(1):37–45. https://doi.org/10.1007/s00427-015-0527-y
Erkenbrack EM, Davidson EH (2015) Evolutionary rewiring of gene regulatory network linkages at divergence of the echinoid subclasses. Proc Natl Acad Sci U S A 112(30):E4075–E4084. https://doi.org/10.1073/pnas.1509845112
Erkenbrack EM, Petsios E (2017) A conserved role for VEGF signaling in specification of homologous mesenchymal cell types positioned at spatially distinct developmental addresses in early development of sea urchins. J Exp Zool B Mol Dev Evol 328(5):423–432. https://doi.org/10.1002/jez.b.22743
Erwin DH, Davidson EH (2009) The evolution of hierarchical gene regulatory networks. Nat Rev Genet 10(2):141–148. https://doi.org/10.1038/nrg2499
Gao F, Thompson JR, Petsios E, Erkenbrack E, Moats RA, Bottjer DJ, Davidson EH (2015) Juvenile skeletogenesis in anciently diverged sea urchin clades. Dev Biol 400(1):148–158. https://doi.org/10.1016/j.ydbio.2015.01.017
Garner S, Zysk I, Byrne G, Kramer M, Moller D, Taylor V, Burke RD (2016) Neurogenesis in sea urchin embryos and the diversity of deuterostome neurogenic mechanisms. Development 143(2):286–297. https://doi.org/10.1242/dev.124503
Gibson AW, Burke RD (1985) The origin of pigment cells in embryos of the sea urchin Strongylocentrotus purpuratus. Dev Biol 107(2):414–419. https://doi.org/10.1016/0012-1606(85)90323-9
Hinman VF, Davidson EH (2007) Evolutionary plasticity of developmental gene regulatory network architecture. Proc Natl Acad Sci U S A 104(49):19404–19409. https://doi.org/10.1073/pnas.0709994104
Kominami T, Takata H, Takaichi M (2001) Behavior of pigment cells in gastrula-stage embryos of Hemicentrotus pulcherrimus and Scaphechinus mirabilis. Develop Growth Differ 43(6):699–707. https://doi.org/10.1046/j.1440-169X.2001.00605.x
Kopan R, Ilagan MX (2009) The canonical Notch signaling pathway: unfolding the activation mechanism. Cell 137(2):216–233. https://doi.org/10.1016/j.cell.2009.03.045
Lapraz F, Besnardeau L, Lepage T (2009) Patterning of the dorsal-ventral axis in echinoderms: insights into the evolution of the BMP-chordin signaling network. PLoS Biol 7(11):e1000248. https://doi.org/10.1371/journal.pbio.1000248
Materna SC (2017) Using morpholinos to probe gene networks in sea urchin. Methods Mol Biol 1565:87–104. https://doi.org/10.1007/978-1-4939-6817-6_8
Materna SC, Davidson EH (2012) A comprehensive analysis of Delta signaling in pre-gastrular sea urchin embryos. Dev Biol 364(1):77–87. https://doi.org/10.1016/j.ydbio.2012.01.017
Materna SC, Ransick A, Li E, Davidson EH (2013a) Diversification of oral and aboral mesodermal regulatory states in pregastrular sea urchin embryos. Dev Biol 375(1):92–104. https://doi.org/10.1016/j.ydbio.2012.11.033
Materna SC, Swartz SZ, Smith J (2013b) Notch and Nodal control forkhead factor expression in the specification of multipotent progenitors in sea urchin. Development 140(8):1796–1806. https://doi.org/10.1242/dev.091157
Mellott DO, Thisdelle J, Burke RD (2017) Notch signaling patterns neurogenic ectoderm and regulates the asymmetric division of neural progenitors in sea urchin embryos. Development 144(19):3602–3611. https://doi.org/10.1242/dev.151720
Minokawa T (2017) Comparative studies on the skeletogenic mesenchyme of echinoids. Dev Biol 427(2):212–218. https://doi.org/10.1016/j.ydbio.2016.11.011
Ohguro Y, Takata H, Kominami T (2011) Involvement of Delta and Nodal signals in the specification process of five types of secondary mesenchyme cells in embryo of the sea urchin, Hemicentrotus pulcherrimus. Develop Growth Differ 53(1):110–123. https://doi.org/10.1111/j.1440-169X.2010.01233.x
Oliveri P, Tu Q, Davidson EH (2008) Global regulatory logic for specification of an embryonic cell lineage. Proc Natl Acad Sci U S A 105(16):5955–5962. https://doi.org/10.1073/pnas.0711220105
Peter IS, Davidson EH (2011) A gene regulatory network controlling the embryonic specification of endoderm. Nature 474(7353):635–639. https://doi.org/10.1038/nature10100
Popovic M, Wienk H, Coglievina M, Boelens R, Pongor S, Pintar A (2014) The basic helix-loop-helix region of the transcriptional repressor hairy and enhancer of split 1 is preorganized to bind DNA. Proteins 82(4):537–545. https://doi.org/10.1002/prot.24507
Range R (2014) Specification and positioning of the anterior neuroectoderm in deuterostome embryos. Genesis 52(3):222–234. https://doi.org/10.1002/dvg.22759
Ransick A, Davidson EH (2006) cis-regulatory processing of Notch signaling input to the sea urchin glial cells missing gene during mesoderm specification. Dev Biol 297(2):587–602. https://doi.org/10.1016/j.ydbio.2006.05.037
Ransick A, Davidson EH (2012) Cis-regulatory logic driving glial cells missing: self-sustaining circuitry in later embryogenesis. Dev Biol 364(2):259–267. https://doi.org/10.1016/j.ydbio.2012.02.003
Revilla-i-Domingo R, Oliveri P, Davidson EH (2007) A missing link in the sea urchin embryo gene regulatory network: hesC and the double-negative specification of micromeres. Proc Natl Acad Sci U S A 104(30):12383–12388. https://doi.org/10.1073/pnas.0705324104
Rottinger E, Croce J, Lhomond G, Besnardeau L, Gache C, Lepage T (2006) Nemo-like kinase (NLK) acts downstream of Notch/Delta signalling to downregulate TCF during mesoderm induction in the sea urchin embryo. Development 133(21):4341–4353. https://doi.org/10.1242/dev.02603
Sherwood DR, McClay DR (1997) Identification and localization of a sea urchin Notch homologue: insights into vegetal plate regionalization and Notch receptor regulation. Development 124(17):3363–3374
Sherwood DR, McClay DR (1999) LvNotch signaling mediates secondary mesenchyme specification in the sea urchin embryo. Development 126(8):1703–1713
Smith AB, Pisani D, Mackenzie-Dodds JA, Stockley B, Webster BL, Littlewood DT (2006) Testing the molecular clock: molecular and paleontological estimates of divergence times in the Echinoidea (Echinodermata). Mol Biol Evol 23:1832–1851. https://doi.org/10.1093/molbev/msl039
Strom A, Castella P, Rockwood J, Wagner J, Caudy M (1997) Mediation of NGF signaling by post-translational inhibition of HES-1, a basic helix-loop-helix repressor of neuronal differentiation. Genes Dev 11(23):3168–3181. https://doi.org/10.1101/gad.11.23.3168
Sweet HC, Gehring M, Ettensohn CA (2002) LvDelta is a mesoderm-inducing signal in the sea urchin embryo and can endow blastomeres with organizer-like properties. Development 129(8):1945–1955
Takata H, Kominami T (2004) Behavior of pigment cells closely correlates the manner of gastrulation in sea urchin embryos. Zool Sci 21(10):1025–1035. https://doi.org/10.2108/zsj.21.1025
Takata H, Kominami T (2011) Novel population of embryonic secondary mesenchyme cells in the keyhole sand dollar Astriclypeus manni. Develop Growth Differ 53(5):625–638. https://doi.org/10.1111/j.1440-169X.2011.01278.x
Thompson JR, Erkenbrack EM, Hinman VF, McCauley BS, Petsios E, Bottjer DJ (2017) Paleogenomics of echinoids reveals an ancient origin for the double-negative specification of micromeres in sea urchins. Proc Natl Acad Sci U S A 114(23):5870–5877. https://doi.org/10.1073/pnas.1610603114
Thompson JR, Petsios E, Davidson EH, Erkenbrack EM, Gao F, Bottjer DJ (2015) Reorganization of sea urchin gene regulatory networks at least 268 million years ago as revealed by oldest fossil cidaroid echinoid. Sci Rep 5(1):15541. https://doi.org/10.1038/srep15541
Urben S, Nislow C, Spiegel M (1988) The origin of skeleton forming cells in the sea urchin embryo. Roux Arch Dev Biol 197(8):447–456. https://doi.org/10.1007/BF00385678
Wei Z, Angerer RC, Angerer LM (2011) Direct development of neurons within foregut endoderm of sea urchin embryos. Proc Natl Acad Sci U S A 108(22):9143–9147. https://doi.org/10.1073/pnas.1018513108
Wray GA, McClay DR (1988) The origin of spicule-forming cells in a ‘primitive’ sea urchin (Eucidaris tribuloides) which appears to lack primary mesenchyme cells. Development 103(2):305–315
Wray GA, McClay DR (1989) Molecular heterochronies and heterotopies in early echinoid development. Evolution 43(4):803–813. https://doi.org/10.1111/j.1558-5646.1989.tb05178.x
Yaguchi J, Angerer LM, Inaba K, Yaguchi S (2012) Zinc finger homeobox is required for the differentiation of serotonergic neurons in the sea urchin embryo. Dev Biol 363(1):74–83. https://doi.org/10.1016/j.ydbio.2011.12.024
Yaguchi S, Yaguchi J, Wei Z, Jin Y, Angerer LM, Inaba K (2011) Fez function is required to maintain the size of the animal plate in the sea urchin embryo. Development 138(19):4233–4243. https://doi.org/10.1242/dev.069856
Yamazaki A, Furuzawa Y, Yamaguchi M (2010) Conserved early expression patterns of micromere specification genes in two echinoid species belonging to the orders clypeasteroida and echinoida. Dev Dyn 239(12):3391–3403. https://doi.org/10.1002/dvdy.22476
Yamazaki A, Kidachi Y, Yamaguchi M, Minokawa T (2014) Larval mesenchyme cell specification in the primitive echinoid occurs independently of the double-negative gate. Development 141(13):2669–2679. https://doi.org/10.1242/dev.104331
Yamazaki A, Minokawa T (2015) Expession patterns of mesenchyme specification genes in two distantly related echinoids, Glyptocidaris crenularis and Echinocardium cordatum. Gene Expr Patterns 17(2):87–97. https://doi.org/10.1016/j.gep.2015.03.003
Yamazaki A, Minokawa T (2016) Roles of hesC and gcm in echinoid larval mesenchyme cell development. Develop Growth Differ 58(3):315–326. https://doi.org/10.1111/dgd.12277
Yankura KA, Koechlein CS, Cryan AF, Cheatle A, Hinman VF (2013) Gene regulatory network for neurogenesis in a sea star embryo connects broad neural specification and localized patterning. Proc Natl Acad Sci U S A 110(21):8591–8596. https://doi.org/10.1073/pnas.1220903110
Acknowledgements
Experimental data presented here were collected in the dungeon of the late Eric H. Davidson, who, along with Dave Bottjer, enthusiastically supported me and my research efforts. The manuscript was written in the laboratory of Günter P. Wagner, whose support and patience I appreciatively acknowledge. Comments from two anonymous reviewers greatly improved this manuscript. This research was supported by NSF grant IOS1240626. Another one.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Hiroki Nishida
Electronic supplementary material
ESM 1
(DOCX 3564 kb)
Rights and permissions
About this article
Cite this article
Erkenbrack, E.M. Notch-mediated lateral inhibition is an evolutionarily conserved mechanism patterning the ectoderm in echinoids. Dev Genes Evol 228, 1–11 (2018). https://doi.org/10.1007/s00427-017-0599-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-017-0599-y