Abstract
Heat shock protein 83 (HSP83) is homologous to the chaperone HSP90. It has pleiotropic functions in Drosophila melanogaster, including the control of longevity and fecundity, and facilitates morphological evolution by buffering cryptic deleterious mutations in wild populations. In the pea aphid Acyrthosiphon pisum, HSP83 expression is moderately induced by bacterial infection but upregulated more strongly in response to heat stress and fungal infection. Stress-inducible heat shock proteins are of considerable evolutionary and ecological importance because they are known to buffer environmental variation and to influence fitness under non-optimal conditions. To investigate the functions of HSP83 in viviparous aphids, we used RNA interference to attenuate its expression and studied the impact on complex parameters. The RNA interference (RNAi)-mediated depletion of HSP83 expression in A. pisum reduced both longevity and fecundity, suggesting this chaperone has an evolutionarily conserved function in insects. Surprisingly, HSP83 depletion reduced the number of viviparous offspring while simultaneously increasing the number of premature nymphs developing in the ovaries, suggesting an unexpected role in aphid embryogenesis and eclosion. The present study indicates that reduced HSP83 expression in A. pisum reveals both functional similarities and differences compared with its reported roles in holometabolous insects. Its impact on aphid lifespan, fecundity, and embryogenesis suggests a function that determines their fitness. This could be achieved by targeting different client proteins, recruiting distinct co-chaperones or transposon activation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Heat shock proteins (HSPs) are evolutionarily conserved chaperones whose predominant function is to prevent the misfolding and denaturation of proteins caused by environmental stressors such as heat, toxins, or pathogens (Johnson 2012). Their functions in Drosophila melanogaster are associated with the buffering of environmental variations, determining fitness under non-optimal conditions, and are therefore of significant evolutionary and ecological relevance (Sorensen et al. 2003). HSPs have been assigned to five families based on homology and molecular mass. The HSP90 family is particularly relevant in the context of evolutionary biology because one member (HSP90) acts as a capacitor for morphological evolution in D. melanogaster (Rutherford and Lindquist 1998) by buffering phenotypic variance producing altered phenotypes in response to environmental stressors. The silencing of HSP90 generates variation by transposon-mediated “canonical” mutagenesis (Specchia et al. 2010).
The pleiotropic roles of HSP90 family members in D. melanogaster are associated with spermatogenesis, oogenesis, and embryogenesis (Ding et al. 1993; Yue et al. 1999; Song et al. 2007; Pisa et al. 2009) as well as the buffering of cryptic deleterious mutations in wild populations, longevity, and fecundity (Chen and Wagner 2012). In the beetle Tribolium castaneum, another holometabolous model insect, HSP83, which belongs to the HSP90 family, is expressed in the whole body as well as in the oocytes where it is specifically located in the follicle cells. There it is differently expressed during different stages of oogenesis (Xu et al. 2010) and in response to heat shock (Xu et al. 2009). The latter suggests that HSP90 family members may regulate physiological processes in response to, e.g., environmental signals (Erlejman et al. 2014). In the whole body of T. castaneum, the expression of HSP90 reaches its highest levels during the larval and pre-pupal phases and the attenuation of its expression negatively affects compound eye development in larvae, suggesting that members of the HSP90 family are essential for normal post-embryonic development (Knorr and Vilcinskas 2011). A phylogenetic analysis of arthropod HSP90 genes reveals that the sequences cluster according to their taxonomic order, with holometabolous and hemimetabolous species showing clear separation (Knorr and Vilcinskas 2011).
In response to heat shock, the hemimetabolous whitefly Bemisia tabaci shows no differential expression of members of the HSP90 family (Lü and Wan 2011). In the aphid species Acyrthosiphon pisum, the first hemimetabolous insect with a completely sequenced genome (The International Aphid Genomic Consortium 2010), Gerardo et al. (2010) demonstrated that HSP83 expression is induced fivefold in response to heat stress. The latter study also showed only minor differences in expression levels between untreated controls and aphids exposed to environmental stress or pathogens. To determine whether HSP90 family members show overlapping or diverse functions in holometabolous and hemimetabolous insects, we investigated the direct impact of HSP83 expression on reproduction in the pea aphid A. pisum, as previously shown for HSP90 in D. melanogaster (Chen and Wagner 2012). Aphids have evolved complex life cycles including the alternation of sexual and asexual reproduction, with an unusual (autosome-like) inheritance of the X chromosome (The International Aphid Genomic Consortium 2010).
The attenuation of gene expression by RNA interference (RNAi) is a powerful method for the functional analysis of genes in A. pisum (Mutti et al. 2006; Jaubert-Possamai et al. 2007; Will and Vilcinskas 2013). We therefore attenuated HSP83 expression in viviparous A. pisum by microinjecting the aphids with the corresponding double-stranded RNA (dsRNA). Several fitness parameters were observed in the injected insects to determine the effect of HSP83 attenuation on longevity, fecundity, and embryogenesis.
Material and methods
Aphid and plant rearing
The rearing of A. pisum clone LL01 and the cultivation of the host plant Vicia faba var. minor were carried out as previously described (Will and Vilcinskas 2015). During the experiments, aphids were kept on detached, mature V. faba leaves under controlled environmental conditions (Mutti et al. 2006; Will and Vilcinskas 2015).
RNAi-mediated attenuation of HSP83 expression
The RNAi-mediated suppression of HSP83 expression was carried out as previously described (Will and Vilcinskas 2015). Briefly, the Ambion MEGAscript T7 Kit (Applied Biosystems, Austin, TX) was used to prepare dsRNA according to the manufacturer’s protocol. Gene-specific primers including the T7 polymerase promoter sequence at the 5′ end were used to synthesize a 530-bp HSP83 (GenBank XM_001943137.3) dsRNA template (forward primer 5′-TAA TAC GAC TCA CTA TAG GGA GAG TGA GCC GCA TCA AGC CTA AC-3′, reverse primer 5′-TAA TAC GAC TCA CTA TAG GGA GAT ATC AGC CTC GGC CTT CTG TC-3′). We excluded the presence of sequence overlaps >19 bp with other A. pisum genes to avoid off-target effects. The QIAquick PCR Purification Kit (Quiagen, Hilden, Germany) was used for template preparation, and dsRNA was produced using the Ambion MEGAscript RNAi kit (Applied Biosystems). Primers were designed with Primer3 (Rozen and Skaletsky 2000) and were purchased from Sigma-Aldrich (Taufkirchen, Germany). Control aphids were injected with equivalent concentrations of dsRNA encoding the insect metalloproteinase inhibitor IMPI (GenBank gbAY330624.1) from the greater wax moth Galleria mellonella (Clermont et al. 2004; Wedde et al. 2007). This sequence is not present in insects other than the Lepidoptera (Mylonakis et al. 2016).
We injected 8-day-old apterous L4 nymphs with ∼50 ng dsRNA in a total volume of 6.9 nl under a stereomicroscope using a Nanoliter 2000 injector with a Sys-Micro4 controller (World Precision Instruments, Berlin, Germany). Glass microcapillaries for injection were prepared using a PN-30 puller (Narishige International Limited, London, UK). Prior to injection, aphids were immobilized with their dorsal thorax on a vacuum holder (van Helden and Tjallingii 2000). The dsRNA was applied to aphids with an injection rate of 2 nl/s according to Mutti et al. (2006). The experiment was carried out three times, each replicate with 15 aphids per group.
Quantification of HSP83 expression by real-time PCR
Total RNA was extracted from the aphids 1, 3, and 6 days after RNAi treatment. The 3 × 5 aphids per treatment were collected and RNA was extracted using Direct-zol™ RNA MiniPrep with TRI-Reagent® (Zymo Research, Freiburg, Germany). Complementary DNA was synthesized using 1 μg of total RNA, oligo(dT)18 primers, and the First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, Waltham, MA) according to the manufacturer’s recommendations. Real-time PCR was performed on a StepOnePlus system (Applied Biosystems) using gene-specific TaqMan Gene Expression assays (Thermo Fisher Scientific). The assay was carried out according to the manufacturer’s protocol using custom TaqMan gene expression assays, including the HSP83 gene (GenBank XM_001943137.3) and the reference ribosomal protein L32 (rpl32) gene (GenBank NM_001126210.2). To ensure reproducibility, gene expression was tested in triplicate. Data were analyzed using the ΔΔCq method in REST (Pfaffl 2001; Pfaffl et al. 2002).
Assaying longevity, fecundity, and embryogenesis
Survival assays and reproduction assays were conducted separately using 15 aphids per group in each test. Aphids placed on a leaf in an agar plate were checked each day and nymphs were removed. Plates were kept in a climate cabinet under the conditions described by Will and Vilcinskas (2015). Images of whole animals were taken 5, 7, and 12 days after injection (dai) using a MZ16FA stereo microscope (Leica, Wetzlar, Germany). Ovaries of four to five living aphids from each treatment (untreated, impi dsRNA and hsp83 dsRNA) were dissected 12 days after injection in insect Ringer’s solution (9 g NaCl, 0.25 g MgCl2 × 6H2O, 0.2 g KCl, 1 g glucose in 1 l H2O, pH 6.8). Dissected ovaries were observed under a MZ16FA stereo microscope, and digital images were analyzed to determine the number of ovary follicles and their developmental stage according to Schmidtberg and Vilcinskas (2016).
Image analysis for coloring and body plan area
The quality of images of whole animals and dissected ovaries was improved for brightness and contrast using Photoshop CS v5.1 (Adobe Systems Inc., San Jose, CA, USA). Images forming part of an image set (images from one experiment) were treated in the same manner. RGB images from adults/embryos were transformed to an 8-bit gray scale, pixel gray values were measured and a whole body/embryo mean was calculated using ImageJ v1.42q (Wayne Rosband, National Institute of Health, USA). To compensate for color changes in adults that naturally occur during the aging of aphids, relative brightness was calculated whereas the mean gray value of untreated control animals at each time point was set to 1. The gray value of embryos from untreated mothers was set to 1 as well. The relative brightness of adults/embryos of the microinjected control group (impi dsRNA) and HSP83-depleted aphids was calculated in relation to the gray value of untreated adults/embryos. We analyzed images of 15 untreated adult aphids and 14 adult dsRNA-injected aphids (impi dsRNA and hsp83 dsRNA). Color determination of embryos was based on the measurement of nine late-stage embryos (embryo stage ≥18) from three different ovaries per treatment.
Statistical analysis
Survival analysis was carried out using the Kaplan-Meier log-rank test in Sigma Plot v11. Reproduction, embryogenesis, coloring, and body plan area data were compared by analysis of variance (ANOVA). The level for statistical significance was set to p = 0.05.
Results
Effect of attenuated HSP83 expression on longevity and fecundity
Kaplan-Meier log-rank of survival data from untreated viviparous A. pisum individuals was compared with those injected with dsRNA encoding either hsp83 or an unrelated control gene, the insect metalloproteinase inhibitor impi, which is specific for lepidopterans (Mylonakis et al. 2016). Aphids in the untreated and the injected control groups survived for a maximum of ∼35 days, whereas those injected with hsp83 dsRNA survived for a maximum of ∼22 days (Fig. 1). Depleted HSP83 expression in viviparous A. pisum individuals also significantly reduced the number of nymphs born per aphid and per day compared with the untreated and impi controls (Fig. 2). Aphids in the hsp83 dsRNA group produced a mean of 27 nymphs during the experiment, which was significantly lower (p < 0.001) than both control groups. The same result emerged independently when we assessed the total number of nymphs born per aphid per day (Fig. 2a) or born per aphid throughput the experiment (Fig. 2b).
Survival analysis of aphids treated with hsp83 dsRNA. HSP83-attenuated aphids were compared to control groups using the Kaplan-Meier log-rank test. Survival was significantly reduced for aphids in the hsp83 dsRNA treatment group compared with untreated (nt) aphids (p < 0.01) and aphids injected with impi dsRNA (p < 0.001). There was no significant difference between the control groups (p > 0.05)
Influence of HSP83 attenuation on the reproduction of A. pisum. a Reproduction of aphids treated with hsp83 dsRNA decreases more rapidly and ends at an earlier point compared to the control groups. b Lifetime reproduction of aphids treated with hsp83 dsRNA is significantly reduced compared to both controls (p < 0.001). There was no significant difference between the control groups (p > 0.05). c The body plan area of aphids treated with hsp83 dsRNA is marginally reduced but is not significantly affected compared with untreated (nt) aphids and aphids treated with impi dsRNA (p > 0.05). A significant difference between the groups is indicated in the graph by different letters
Effect of attenuated HSP83 expression on aphid embryogenesis and eclosion
Using the gene-specific TaqMan Gene Expression assay, we observed attenuated HSP83 expression levels after 1 day, hsp83 dsRNA. However, the attenuation of gene expression was not significant, and the expression increased slightly 3 and 6 days after the injection of (Supplementary Fig. S1). HSP83 depletion did not appear to affect L4 (8-day-old) nymphs, which developed into reproductive adults. However, the attenuated expression of HSP83 resulted in the eclosion of many premature nymphs (Fig. 3). These died a few hours after eclosion and their antennae and legs remained folded. In the untreated control group, 1 % of eclosed nymphs were premature and the phenomenon was only observed on day 11. In the control group injected with impi dsRNA, 2–9 % of eclosed nymphs were premature and the eclosions occurred between days 6 and 10 after injection. But in the group injected with hsp83 dsRNA, the proportion of premature eclosed nymphs increased from 16 to ∼80 % between days 9 and 12 after treatment, the time during which the reproduction phase of the hsp83 dsRNA-injected aphids is completed (Fig. 2a). Remarkably, when observing embryos through the integument of the ovary 12 dai, we observed many more translucent eyes representing developing embryos in the group treated with hsp83 dsRNA compared to the two control groups (Fig. 4a–c). Dissection of ovaries from four hsp83 dsRNA-injected adults and from five adult aphids from each control group revealed that aphids injected with hsp83 dsRNA contained no embryos at or before developmental stage 6, whereas embryos were present in both control groups. In addition, the second developmental phase (embryo stages 7–13) differed significantly between the HSP83-depleted aphids and those from control groups. A striking characteristic of aphids treated with hsp83 dsRNA was the presence of embryos at later developmental stages (embryo stage ≥18) that are detached from the ovarioles and lie free inside the hemocoel (Fig. 4f). These are not present in either of the control groups (Table 1; Fig. 4d, e). Effects were only considered to be HSP83-dependent when significant differences were observed between the HSP83-depleted aphids and both control groups.
Impact of HSP83 attenuation on the percentage of premature nymphs. In comparison to the untreated control (a), control aphids injected with impi dsRNA (b) produce a small proportion (maximum 9 %) of premature nymphs 6–10 days after injection. In contrast, aphids injected with hsp83 dsRNA (c) show a rapid increase in the percentage of premature nymphs from day 9 after injection until reproduction stopped (12 days after injection)
Influence of HSP83 attenuation on aphid phenotype. Untreated (a) and impi control (b) adult apterous female aphids are bright green, and a small number of embryo eye spots are visible through the cuticle. Scale bars = 2 mm. Aphids treated with hsp83 dsRNA (c) are dark green, and more eye spots (arrowheads) can be seen compared to the control groups. Scale bar = 2 mm. Dissected ovaries of untreated (d) and impi control aphids (e) contain embryos at developmental stage 6 and earlier (see Table 1). Scale bar = 1 mm. In contrast, these developmental stages are absent in ovaries from aphids treated with hsp83 dsRNA, and some late-stage embryos are not attached to ovaries (f). Scale bar = 1 mm
Effect of attenuated HSP83 expression on aphid color
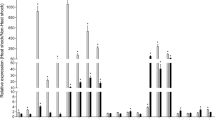
The attenuated expression of HSP83 caused the injected adults (Fig. 5a) to become significantly darker in color (Fig. 4c) during the observation period, with a mean relative brightness of 0.95 (5 dai; p = 0.047), 0.94 (7 dai; p = 0.024), and 0.84 (12 dai; p = 0.002), compared to untreated aphids whose relative brightness was set to 1 at each time point. This was also observed for the embryos (Fig. 5b) inside HSP83-attentuated adults, with a mean relative brightness of 0.68 (12 dai; p < 0.001). Between the control groups of untreated and impi dsRNA-injected animals, there were no significant differences in coloring at any time point for adults (5 dai: p = 0.31; 7 dai: p = 0.488; 12 dai: p = 0.582) or embryos (12 dai: p = 0.161).
Change of coloring of HSP83-attenuated aphids and their embryos. Adult apterous aphids (a) show a darkening color in response to HSP83 silencing over time. b Embryos of HSP83-attenuated aphids (12 dai) show also a significantly darker coloring than embryos of control groups. A significant difference (p ≤ 0.05) between the groups is indicated in the graph by different letters
Discussion
The postulated evolutionary and ecological role of HSPs predicts that their expression is induced by exposure to environmental stressors and influences fitness parameters such as lifespan and fecundity (Sorensen et al. 2003). However, exposure to mild heat shock or microbial elicitors of immune responses only moderately induced the expression of hsp90 and its homolog hsp83 in the model insects T. castaneum (Freitak et al. 2012) and A. pisum (Gerardo et al. 2010). The HSP90 family also plays a role in insect spermatogenesis, oogenesis, and embryogenesis (Ding et al. 1993; Yue et al. 1999; Song et al. 2007). The developmental roles of HSP90 have recently expanded beyond those known in embryogenesis to encompass functions in post-embryonic development such as the regulation of compound eye formation (Knorr and Vilcinskas 2011). As in the latter study, we also used RNAi-mediated attenuation of HSP expression to explore the functions of hsp83 in hemimetabolous aphids, which have evolved a peculiar life cycle combining the alternation of sexual and asexual reproduction with an unusual (autosome-like) inheritance of the X chromosome (The International Aphid Genomic Consortium 2010). In accordance with our expectations, we found that the injection of hsp83 dsRNA into A. pisum reduced the lifespan, fecundity, and number of viviparous offspring, even though the attenuation of HSP83 expression was not significant. The confirmation of gene knockdown in RNAi experiments is sometimes difficult, particularly if the target gene is expressed at a low level, because it depends on the selected reference genes (Holmes et al. 2010; Baumann et al. 2015).
The negative impact of attenuated HSP83 expression on the survival of A. pisum (Fig. 1a) appears to be in striking agreement with the role of the homologous HSP90 in the longevity of D. melanogaster (Chen and Wagner 2012) suggesting that at least one function of HSP83 is evolutionarily conserved in insects. The proposed role of HSP90 in the fecundity of D. melanogaster (Chen and Wagner 2012) was also observed in A. pisum, where attenuated HSP83 expression significantly inhibited the formation of viviparous offspring compared to untreated controls and controls treated with impi dsRNA. Reduced HSP83 expression also increased dramatically the number of immature eclosed nymphs (Fig. 3). This suggests that HSP83 displays a previously unknown role in embryogenesis. The low number of early-stage embryos in the ovaries of aphids injected with hsp83 dsRNA is presumably caused by the resorption of embryos, occurring under suboptimal environmental conditions, allowing the late-stage embryos to reach maturity (Ward and Dixon 1982).
Members of the HSP90 family are known to participate in signal transduction (Nollen and Morimoto 2002), e.g., by activating steroid receptors (Bohen and Yamamoto 1993). We therefore propose that HSP83 expression regulates embryogenesis and eclosion, which are both strongly influenced by environmental factors (Ward and Dixon 1982; Altincicek et al. 2008). Its function may be mediated by the recently reported interaction with the transcription factor Broad Z7 because Cai et al. (2014) reported that HSP90 associates with the Broad Complex/Tramtrack/Bric-a-brac domain of Broad Z7 to prevent its degradation in the moth Helicoverpa armigera, and Piulachs et al. (2010) showed that Broad plays key roles in embryogenesis of the cockroach Blattella germanica. Therefore, it appears plausible that the downregulation of hsp83 expression to below a specific but unknown threshold could disrupt the interplay between embryonic development and eclosion, leading to the presence of embryos that are detached from the ovarioles and lie free inside the hemocoel of hsp83 dsRNA-injected aphids (Fig. 4f). Interestingly, the observed impact of injected hsp83 dsRNA on embryos suggests the occurrence of parental RNAi because it has been reported that injected or orally delivered dsRNA can cause trans-generational attenuation of gene expression in aphids (Abdellatef et al. 2015).
The diverse roles of HSP83 in aphid longevity, fecundity, and embryogenesis may reflect either a distinct pool of client proteins that interact with HSP83 (Erlejman et al. 2014) or the requirement for specific co-chaperones to achieve appropriate HSP83 targeting (Johnson 2012). Multiple isoforms and transcript variants of HSP90 family members such as HSP83, which have been identified in A. pisum and Myzus persicae (cf. AphidBase), appear to act as chaperones for different types of client proteins related to longevity, fecundity, and development (Haslbeck et al. 2012).
Interestingly, the darker color of adult aphids and their embryos in the hsp83 dsRNA group concurs precisely with the proposed epigenetic role of this chaperone in the protection of insects against environmental stress imposed by UV-A (Sang et al. 2012) or heat (Gilbert et al. 2007). Temperature acts on melanin production by modulating a chromatin regulator network, interacting genetically with the transcription factor Bric-a-brac, which is also an HSP83 target (Cai et al. 2014). HSP90 in D. melanogaster and in mammals can target paused RNA polymerases to activate genes in response to environmental stimuli (Sawarkar et al. 2012). Our data suggest that the aphid HSP83 homolog may have a related function.
The collection of altered phenotypes observed in A. pisum following the RNAi-mediated attenuated expression of hsp83 can be explained by an alternative hypothesis based on the occurrence buffered phenotypic variation in response to environmental stimuli. The silencing of HSP90 in D. melanogaster resulted in transposon-mediated mutagenesis (Specchia et al. 2010). However, further research is required to confirm whether the attenuation of HSP83 expression in A. pisum also induces the mobilization of transposable elements, ultimately causing the observed phenotypic variation.
In conclusion, attenuated HSP83 expression in the hemimetabolous aphid A. pisum has revealed functional similarities and differences compared with its reported roles in holometabolous insects such as T. castaneum (Knorr and Vilcinskas 2011). The observed negative impact of reduced HSP83 expression on aphid survival and its complex effects on reproduction and embryogenesis suggest that the protein has pleiotropic roles involving the mediation of environmental stimuli affecting these complex parameters. The resulting functional plasticity could be achieved by targeting different client proteins, by recruiting distinct co-chaperones, or by inducing transposon-mediated mutagenesis. The entity of our results implicates that HSP83 represents another promising target for RNAi-mediated approaches aiming the engineering of aphid-proof crops (Will and Vilcinskas 2013; Abdellatef et al. 2015).
Abbreviations
- dsRNA:
-
double-stranded RNA
- HSP:
-
heat shock protein
- RNAi:
-
RNA interference dai
References
Abdellatef E, Will T, Koch A, Imani J, Vilcinskas A, Kogel KH (2015) Silencing the expression of the salivary sheath protein causes transgenerational feeding suppression in the aphid Sitobion avenae. Plant Biotech J 13:849–857
Altincicek B, Gross J, Vilcinskas A (2008) Wounding-mediated gene expression and accelerated viviparous reproduction of the pea aphid Acyrthosiphon pisum. Insect Mol Biol 17:711–716
Baumann A, Lehmann R, Beckert A, Vilcinskas A, Franta Z (2015) Selection and evaluation of tissue specific reference genes in Lucilia sericata during an immune challenge. PLoS One 10(8):e0135093
Bohen SP, Yamamoto KR (1993) Isolation of Hsp90 mutants by screening for decreased steroid receptor function. Proc Natl Acad Sci U S A 90:11424–11428
Cai MJ, Li XR, Pei XY, Liu W, Wang JX, Zhao XF (2014) Heat shock protein 90 maintains the stability and function of transcription factor Broad Z7 by interacting with its Broad-Complex-Tramtrack-Bric-a-brac domain. Insect Mol Biol 23:720–732
Chen B, Wagner A (2012) Hsp90 is important for fecundity, longevity, and buffering of cryptic deleterious variation in wild fly populations. BMC Evol Biol 12:25
Clermont A, Wedde M, Seitz V, Podsiadlowski L, Hummel M, Vilcinskas A (2004) Cloning and expression of an inhibitor against microbial metalloproteinases from insects (IMPI) contributing to innate immunity. Biochem J 382:315–322
Ding D, Parkhurst SM, Halsell SR, Lipshitz HD (1993) Dynamic Hsp83 RNA localization during Drosophila oogenesis and embryogenesis. Mol Cell Biol 13:3773–3781
Erlejman AG, Lagardi M, Toneatto J, Piwien-Pilipuk G, Galigniani MD (2014) Regulatory role of the 90-kDa-heat-shock protein (Hsp90) and associated factors on gene expression. Biochim Biophys Acta 1839:71–87
Freitak D, Knorr A, Vogel H, Vilcinskas A (2012) Gender- and stressor-specific microRNA expression in Tribolium castaneum. Biol Lett 8:860–863
Gerardo N, Altincicek B, Anselme C, Atamian H, Barribeau S, de Vos M, Duncan EJ, Evand JD, Gabaldón T, Ghanim M, Heddi A, Kaloshian I, Latorre A, Moya A, Nakabachi A, Parker BJ, Pérez-Brocal V, Pignatelli M, Rahbé Y, Ramsey JS, Spragg CJ, Tamames J, Tamarit D, Tamborindeguy C, Vincent-Monegat C, Vilcinskas A (2010) Immunity and other defenses in pea aphids, Acyrthosiphon pisum. Genome Biol 11:R21
Gilbert JM, Peronnet F, Schlötterer C (2007) Phenotypic plasticity in Drosophila pigmentation caused by temperature sensitivity of a chromatin regulator network. PLoS Genet 3:e30
Haslbeck V, Kaiser CJ, Richter K (2012) Hsp90 in non-mammalian metazoan model systems. Biochim Biophys Acta 1823:712–721
Holmes K, Williams MC, Chapman EA, Cross MJ (2010) Detection of siRNA induced mRNA silencing by RT-qPCR: considerations for experimental design. BMC Res Notes 3:53
Jaubert-Possamai S, Le Trionnaire G, Bonhomme J, Christophides GK, Rispe C, Tagu D (2007) Gene knockdown by RNAi in the pea aphid Acyrthosiphon pisum. BMC Biotechnol 7:63
Johnson JL (2012) Evolution and function of diverse Hsp90 homologs and cochaperone proteins. Biochim Biophys Acta 1823:607–613
Knorr E, Vilcinskas A (2011) Post-embryonic functions of Hsp90 in Tribolium castaneum include the regulation of compound eye development. Dev Genes Evol 221:357–362
Lü ZC, Wan FH (2011) Using double-stranded RNA to explore the role of heat shock protein genes in heat tolerance in Bemisia tabaci (Gennadius. J Exp Biol 214:764–769
Mutti NS, Park Y, Reese JC, Reeck GR (2006) RNAi knockdown of a salivary transcript leading to lethality in the pea aphid, Acyrthosiphon pisum. Insect Sci 6:38
Mylonakis E, Podsiadlowski L, Muhammed M, Vilcinskas A (2016) Diversity, evolution and medical applications of insect antimicrobial peptides. Phil Trans R Soc B 371:20150290
Nollen EA, Morimoto RI (2002) Chaperoning signaling pathways: molecular chaperones as stress-sensing heat shock proteins. J Cell Sci 115:2809–2816
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29(9, article):e45. doi:10.1093/nar/29.9.e45
Pfaffl MW, Horgan GW, Dempfle L (2002) Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res 30:e36
Pisa V, Cozzolino M, Gargiulo S, Ottone C, Piccioni F, Monti M, Gigliotti S, Talamo F, Graziani F, Pucci P, Verrotti AC (2009) The molecular chaperone Hsp90 is a component of the cap-binding complex and interacts with the translational repressor cup during Drosophila oogenesis. Gene 432:67–74
Piulachs MD, Pagone V, Belles X (2010) Key roles of the Broad-Complex gene in insect embrygenesis. Insect Biochem Mol Biol 40:468–475
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics: methods and protocols. Humana Press, Totowa, pp. 365–386
Rutherford SL, Lindquist S (1998) Hsp90 as a capacitor for morphological evolution. Nature 396:336–342
Sang W, Ma WH, Qiu L, Zhu ZH, Lei CL (2012) The involvement of heat shock protein and cytochrome genes in response to UV-A exposure in the beetle Tribolium castaneum. J Insect Physiol 58:830–836
Sawarkar R, Sievers C, Paro R (2012) Hsp90 globally targets paused RNA polymerase to regulate gene expression in response to environmental stimuli. Cell 149:807–818
Schmidtberg H, Vilcinskas A (2016) The ontogenesis of the pea aphid Acyrthosiphon pisum. In: Vilcinskas A (ed) Biology and ecology of the aphids. CRC Press, Boca Raton, pp. 14–51
Song Y, Fee L, Lee TH, Wharton RP (2007) The molecular chaperone Hsp90 is required for mRNA localization in Drosophila melanogaster embryos. Genetics 176:2213–2222
Sorensen JG, Kristensen TN, Loeschcke V (2003) The evolutionary and ecological role of heat shock proteins. Ecol Lett 6:1025–1037
Specchia V, Piacentini L, Tritto P, Fanti L, D’Alessandro R, Palumbo G, Pimpinelli S, Bozzetti MP (2010) Hsp90 prevents phenotypic variation by suppressing the mutagenic activity of transposons. Nature 463(7281):662
The International Aphid Genomic Consortium (2010) Genome sequence of the pea aphid Acyrthosiphon pisum. PLoS Biol 8:e1000313
van Helden M, Tjallingii WF (2000) Experimental design and analysis in EPG experiments with emphasis on plant resistance research. In: Walker GP, Backus EA (eds) Principles and applications of electronic monitoring and other techniques in the study of Homopteran feeding behavior. Thomas Say Publications in Entomology, Entomological Society of America, Lanham, pp. 144–171
Ward SA, Dixon AFG (1982) Selective resorption of aphid embryos and habitat changes relative to life-span. J Anim Ecol 51:859–864
Wedde M, Weise C, Nuck C, Altincicek B, Vilcinskas A (2007) The insect metalloproteinase inhibitor gene of the lepidopteran Galleria mellonella encodes two distinct inhibitors. Biol Chem 388:119–127
Will T, Vilcinskas A (2013) Aphid-proof plants: biotechnology-based approaches for aphid control. Adv Biochem Eng Biotechnol 136:179–203
Will T, Vilcinskas A (2015) The structural sheath protein of aphids is required for phloem feeding. Insect Biochem Mol Biol 57:34–40
Xu J, Shu J, Qiu X, Wang Z, Zhao F, Zhang Z, Zhang Q (2009) Effects of heat shock on ovary development and hsp83 expression in Tribolium castaneum (Coleoptera: Tenebrionidae. Insect Biochem Mol Biol 70:204–216
Xu J, Shu J, Zhang Q (2010) Expression of the Tribolium castaneum (Coleoptera: Tenebrionidae) hsp83 gene and its relation to oogenesis during ovarian maturation. J Genet Genomics 37:513–522
Yue L, Karr TL, Nathan DF, Swift H, Srinivasan S, Lindquist S (1999) Genetic analysis of viable Hsp90 alleles reveals a critical role in Drosophila spermatogenesis. Genetics 151:1065–1079
Acknowledgments
The authors acknowledge the generous funding from the Hessen State Ministry of Higher Education, Research and the Arts (HMWK) via the LOEWE center for “Insect Biotechnology and Bioresources.” The authors thank Richard M. Twyman for editing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Claude Desplan
Electronic supplementary material
Fig. S1
(DOCX 273 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Will, T., Schmidtberg, H., Skaljac, M. et al. Heat shock protein 83 plays pleiotropic roles in embryogenesis, longevity, and fecundity of the pea aphid Acyrthosiphon pisum . Dev Genes Evol 227, 1–9 (2017). https://doi.org/10.1007/s00427-016-0564-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-016-0564-1