Abstract
Hox genes form clusters. Invertebrates and Amphioxus have only one hox cluster, but in vertebrates, they are multiple, i.e., four in the basal teleost fish Polyodon and tetrapods (HoxA, B, C, D), but seven or eight in common teleosts. We earlier completely sequenced the entire hox gene loci in medaka fish, showing a total of 46 hox genes to be encoded in seven clusters (hoxAa, Ab, Ba, Bb, Ca, Da, Db). Among them, hoxAa, hoxAb and hoxDa clusters are presumed to be important for fin-to-limb evolution because of their key role in forelimb and pectoral fin development. In the present study, we compared genome organization and nucleotide sequences of the hoxAa and hoxAb clusters to these of tetrapod HoxA clusters, and found greater similarity in hoxAa case. We then analyzed expression of Abd-B family genes in the clusters. In the trunk, those from the hoxAa cluster, i.e., hoxA9a, hoxA10a, hoxA11a and hoxA13a, were expressed in a manner keeping the colinearity rule of the hox expression as those of tetrapods, while those from the hoxAb cluster, i.e., hoxA9b, hoxA10b, hoxA11b and hoxA13b, were not. In the pectoral fins, the hoxAa cluster was expressed in split domains and did not obey the rule. By contrast, those from the hoxAb and hoxDa clusters were expressed in a manner keeping the rule, i.e., an ancestral pattern similar to those of tetrapods. It is plausible that this differential expression of the two clusters is caused by changes occurred in global control regions after cluster duplications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hox genes encode transcription factors that play key roles in body planning (Gehring et al. 1994). In Drosophila, the Hox cluster is termed the HOM complex, containing 11 homeotic genes. The HOM genes have the remarkable feature called ‘colinearity’, that is, they are expressed along the body axis corresponding to the chromosomal positions (Levine et al. 1983; Harding et al. 1985). Homologues of the HOM genes have been identified in a number of animal species and vertebrates that maintain HOM homologues as clusters as well as the colinearity rule (Izpisúa-Belmonte et al. 1991; Krumlauf 1992). Their Hox genes have been classified into 13 paralogy groups, numbered HoxA1 through to HoxA13 (Scott 1992).
In the evolutionary lineage of vertebrates, Amphioxus with archaic vertebrate features carries only one hox cluster (Ferrier et al. 2000), whereas mammals and the basal ray-finned fish Polyodon possess four Hox clusters that are termed HoxA, B, C and D (Duboule and Dollé 1989, Metscher et al. 2005), due to two episodes of genome duplication (Hart et al. 1987). Further, it is known that common ray-finned fishes (teleosts) including zebrafish, pufferfish and medaka fish have experienced an additional duplication resulting in seven or eight clusters (Amores et al. 1998, 2004; Naruse et al. 2000; Kurosawa et al. 1999, 2006). These are termed hoxAa, hoxAb, hoxBa, hoxBb, hoxCa, hoxDa and hoxDb in the medaka (Kurosawa et al. 2006).
The duplicated Hox clusters have acquired new or modified regulation features, causing an increase in developmental complexity and capacity for diversification (Holland 1999; Holland and Garcia-Fernàndez 1996). In mice, paralogous Hox genes have expressional differences, suggesting that duplication allows expression changes depending on cis-regulatory regions (Kessel and Gruss 1990; Burke et al. 1995).
We earlier determined the structure of the entire hox gene loci in medaka fish, revealing that 46 hox genes in total are encoded in seven clusters, and suggested that the third round of duplication of hox clusters might have occurred before their divergence from zebrafish (Kurosawa et al. 2006). The hoxAa, hoxAb and hoxDa clusters are important for fin-to-limb evolution, playing key roles in forelimb and pectoral fin development (Dollé et al. 1989; Sordino et al. 1995). In the present study, we first compared genome organization and nucleotide sequences of hoxAa and Ab clusters with those known to date for tetrapods and teleosts. Then, we analyzed expression patterns of Abd-B family genes in the clusters in the trunk and pectoral fin buds to allow speculation on the regulatory basis of differential expression of the duplicated genes.
Materials and methods
Fish strains
Orange–red variety Japanese medaka fish (Oryzias latipes) were originally obtained from a vendor in Nagoya. These fish were raised and maintained in our laboratory under standard conditions of 14/10 h day/night cycle at 27–28°C. Embryos at 1, 2, and 3 days past fertilization (dpfs) were sampled and used for in situ hybridization. These dpfs correspond to stages 21–22, 26–27, and 29–30 of the stage map of medaka embryo development published by Iwamatsu (2004).
Sequence analysis
The Hox clusters and their DNA sequences used in this study were as follows: mouse HoxA cluster (Mus musculus HoxA cluster: Mmu–HoxA), accession nos. AC015583.34, AC091106.17, and AC113985.11 in GenBank DNA Database; mouse HoxD cluster (Mmu–HoxD), accession nos. AL928644.12 and AL928733.14; mouse HoxA amino acid sequences, HoxA2, A9, A10, A11, and A13, accession nos. P31245, P09631, P31310, P31311, and Q62424; mouse HoxD9 amino acid sequence, accession no. P28357. Those for O. latipes hoxAa, hoxAb, hoxDa, hoxDb clusters (Ola–HoxAa, Ola–HoxAb, Ola–HoxDa and Ola–HoxDb) were obtained from AB232918, AB232919, AB232923, and AB232924, respectively (Kurosawa et al. 2006).
To elucidate similar regions in the two hox clusters, we compared them with the Pip-Maker program (Schwartz et al. 2000), which computes alignments of similar regions in two or more DNA sequences. The mouse HoxA cluster sequence was employed as the template to which the others were compared with the program MultiPipMaker (http://pipmaker.bx.psu.edu/pipmaker/). The results summarized as “percent identity plots,” or “pips” for short.
Probes
Polymerase chain reaction (PCR) primer sequences for production of probes were selected from each gene sequence. To avoid cross hybridization between paralogous genes, we searched for low homology regions by National Center for Biotechnology Information blast and multiple alignment programs, such as ClustalW (http://www.ddbj.nig.ac.jp/search/clustalw-j.html) and GENETYX-MAC (Genetyx, Tokyo). For example, to prepare probes not cross hybridizing between hoxA9a and hoxA9b that are highly homologous with each other, we identified unique sequences from hox coding sequences with accession nos. AB232918 and AB232919 by the alignment software. All Abd-B genes in hoxAa, Ab and Da clusters were well amplified by PCR and cloned, but hoxD9b gene was missed to amplified. This missing may be caused on very weak or no expressions in 2 and 3 dpfs. Medaka myoD, used for the purpose of staining somites in the trunk and muscle mass in the fin buds, was isolated from cDNA library by using PCR, and then sequenced (accession no. AB288366). Medaka shh and dHAND sequences, posterior markers of pectoral fin buds, were searched from golw_scaffold Hd-rR 200506 in the Medaka genome database (http://dolphin.lab.nig.ac.jp/medaka) based on the partial shh mRNA sequence (AB007129) and dHAND of other organisms. The primer sequences designed for selective amplification of respective hox genes are listed in Table 1.
Total RNA was isolated from medaka 2 and 3 dpf embryos using ISOGEN (Nippon Gene). Total cDNA was generated from total RNA using SuperScript II-Oligo dT, following recommended protocols (Invitrogen), and DNA fragments for probes were amplified by PCR using the primers (Table 1). The products were cloned into pGEM-T Easy (Invitrogen) and checked for correct sequences by sequencing, and then transcribed to digoxigenin (DIG)-labeled RNA with T7 or SP6 polymerase (Roche). DIG-labeled antisense RNA probes and sense RNA probes for controls were used for in situ experiments.
Sample fixation, hybridization, and staining
Embryos at 1, 2, and 3 dpf were fixed with 4% paraformaldehyde at 4°C overnight, dehydrated with 100% MeOH and stored at −20°C in 100% MeOH until use. Whole-mount in situ hybridization (WISH) was carried out by a modified Inohaya’s method (Inohaya et al. 1995, 1999). Simply, dehydrated embryo samples were washed in phosphate-buffered saline (PBS)-0.1% Tween20 (PBST), and treated with 10 μg/ml proteinase K solution for an appropriate time. After refixing with 4% paraformaldehyde, they were washed three times in PBST, then incubated in prehybridization buffer (50% formamide, 5× saline-sodium citrate (SSC), 50 mg/ml heparin, 0.1% Tween20). 1–2 ml of hybridization buffer with a 1/250 vol of DIG-labeled antisense RNA probe were added to each sample, followed by incubation overnight at 55–58°C. After hybridization, samples were washed four times with 50% formamide-2× SSC—0.1% Tween20, two times with 2× SSC—0.1% Tween20, and two times with 0.2× SSC—0.1% Tween20 at 68°C. After three washes with PBST, samples were blocked with 5% sheep-serum/PBST and bound to DIG-antibody including alkaline phosphatase (Roche) overnight at 4°C. Samples were washed six times with PBST and twice with alkaline phosphatase buffer (0.1 M Tris pH 9.5, 50 mM MgCl2, 0.1 M NaCl, 0.1% Tween20) then stained with NBT/BCIP solution at 4°C or room temperature. When samples were moderately stained, reactions were stopped by PBST washing, and refixation was performed again with 4% paraformaldehyde overnight. Finally, samples were passed through a glycerol series, and stored in 80% glycerol at 4°C until use.
Results
Comparison of mouse HoxA, medaka hoxAa and hoxAb organization
A comparison of the genomic organization of Ola–hoxAa and Ola–hoxAb with that of Mmu–HoxA is illustrated in Fig. 1a. Sizes of Mmu–HoxA, Ola–hoxAa and Ola–hoxAb are ca. 100, 60 and 20 kb, respectively. Mmu–HoxA contains 11 Hox genes and Evx1 in the cluster, and Ola–hoxAa contains 10 homologous hox genes and evx1 in the same order. The single difference between Mmu–HoxA and Ola–hoxAa organization is the lack of an Mmu–HoxA6 homolog in the latter. On the other hand, Ola–hoxAb does not contain hoxA1b, hoxA3b, hoxA4b, hoxA5b, hoxA6b and hoxA7b genes, featuring more gene loss than the Ola–hoxAa cluster. Thus, genome size, gene order and orientation have been conserved between Mmu–HoxA and Ola–hoxAa, but not Ola–hoxAb.
Comparison of mouse HoxA/HoxD, medaka hoxAa and hoxAb/hoxDa and hoxDb clusters. a Organization of the HoxA and HoxD clusters in mouse Mus musculus (Mmu) and of the hoxAa/Ab and hoxDa/Db clusters in the medaka Oryzias latipes (Ola). Ovals indicate positions of hox and evx1 genes. Filled ovals are for Abd-B family genes and open ovals are for other genes. Horizontal arrows indicate the direction of transcription. A square in Mmu–HoxD indicates the GCR (Spitz et al. 2003). Two squares in Ola–HoxDa indicate two blocks of GCR-like region (see the details in the text). b Pip output of the comparison of Mmu–HoxA, Ola–hoxAa and Ola–hoxAb. Shadowed backgrounds indicate coding regions. Horizontal arrows indicate the direction of transcription, and filled boxes indicate exons. Horizontal small open boxes are for CpG/GpC ratios between 0.6 and 0.75, and small gray boxes for ratios over 0.75
Similarities of hoxA protein sequences in the clusters are summarized in Table 2. Exon2 sequences of the Abd–B family show striking similarity between Mmu–HoxA and Ola–hoxAa proteins (72.5–94.7% identity, 82. 9% overall), while lower similarity is present between Mmu–HoxA and Ola–hoxAb proteins (53.2–90.7% identity, 73.0% in total). With exon1 protein sequences, the respective figures are 57.2 and 44.3%, again suggesting Mmu–HoxA to be more related to Ola–hoxAa than to Ola–hoxAb.
A comparison of the genomic organization of Ola–hoxDa and Ola–hoxDb with that of Mmu–HoxD is illustrated in Fig. 1a. D9 orthologues are the only Abd–B genes found in both medaka clusters. D10, D11, and D12 orthologues are situated in Ola–hoxDa, but D13 genes have not been found in medaka yet. The similarity between Ola–hoxD9a and Mmu–HoxD9 proteins (exon1, 45.3%; exon2, 88.6% identity) is higher than the similarity between Ola–hoxD9b and Mmu–HoxD9 (exon1, 18.6%; exon2, 79.7% identity).
Pip outputs from the comparisons of Mmu–HoxA, Ola–hoxAa and Ola–hoxAb sequences are shown in Fig. 1b. All medaka exons corresponding to mouse exons exhibit high similarity scores. Not only coding regions, but also several conserved noncoding sequences previously described in the literature (Santini et al. 2003) show high similarity, such as proximal 5′ upstream regions of HoxA2, HoxA5 genes and HoxA10 regions (Fig. 1b). Among them, the hoxA9b gene and its intron region are showing high similarity in exon2 (84.6%, Table 2) and intron1 regions (Fig. 1b) but low in exon1 (35.8%, Table 2).
Spitz et al. (2003) identified a global control region (GCR) of HoxD, which is highly conserved between a pufferfish Tetraodon and Homo sapiens (Fig. 1a). The GCR of 40 kb in size is a cluster of global enhancers capable of controlling transcription of several genes, even examples unrelated in structure or function. To confirm the presence of GCR around Ola–hoxDa, we compared genomic regions flanking Mmu–HoxD and Ola–hoxDa with the PipMaker program. Two blocks of high homology regions corresponding to the GCR were thereby detected 84 kb upstream of Ola–hoxD12a, at the physical map site of medaka version1.0 scaffold24 in the Medaka genome database (Fig. 1a). In the Ola–hoxDb cluster and in its 500 kb upstream region, we could not identify the GCR-like structure.
Expression of medaka Abd-B genes in the trunk
Expression analysis of medaka Abd-B genes in embryos at 1, 2, and 3 dpfs was carried out by WISH experiments. The staining pattern of the somites by myoD is shown in Fig. 2a–c (i). At 1 dpf, three Abd-B genes in Ola–hoxAa, i.e., hoxA9a, hoxA10a, hoxA11a but not hoxA13a, were found to be clearly expressed in the trunk (Fig. 2a; boxes a, b, c, and d). With Ola–hoxAb, two genes hoxA9b and hoxA10b demonstrated clear expression in the trunk (Fig. 2a; boxes e and f), while hoxA11b and hoxA13b were lacking (Fig. 2a; boxes g and h).
Expression of medaka Abd-B genes in the trunk. Expression of the Abd-B genes in medaka embryos were carried out by WISH experiments. Black arrows and arrow heads indicate the anterior limit of gene expression in the CNS and the paraxial mesoderm, respectively. Scale bars at the right bottom (h) are for 0.5 mm. Expression at a 1 dpf, b 2 dpf, and c 3 dpf. Figures (i) at each dpf are expressions of myoD, a somite marker, and red arrows in magnified views are applied every five somite steps. d Schematic illustration of Abd-B gene expression in the trunk. Genes for hoxAa are on the left and for hoxAb on the right. Dashed lines on the top of several bars indicate incomplete boundaries to be caused by the difficulty of precise positions. Gray ovals with numbers indicate somite levels
Figure 2b and c illustrates expression of these genes at 2 and 3 dpf, respectively, hoxA9a and hoxA10a in Ola–hoxAa being strongly (Fig. 2b and c, boxes a and b), but hoxA11a only weakly present at both stages (Fig. 2b and c, box c). HoxA13a was found in tail ends at both stages (Fig. 2b and c, box d), and strongly in the cloaca area at 3 dpf (Fig. 2c, box d). In Ola–hoxAb, hoxA9b and hoxA10b could be shown to be expressed clearly (Fig. 2b and c, boxes e and f), whereas hoxA11b and hoxA13b were not expressed or only weakly (Fig. 2b and c, boxes g and h).
The difference between hoxA9a and hoxA9b expression was seen at all the developmental stages (1, 2, and 3 dpfs) examined. At 2 and 3 dpfs, the anterior boundary of hoxA9a was around somite 1 of the spinal cord on the dorsal side (Fig. 2b and c, box a), whereas that for hoxA9b was around somite level 5, with a shifting down to the posterior side in comparison with the hoxA9a boundary (black arrows in Fig. 2b and c, box e). An equivalent shift was also observed on the ventral side of 2 and 3 dpfs embryos (arrow heads in Fig. 2b and c, boxes a and e).
Dorsal expression boundaries of paralogous group 10 genes, hoxA10a and hoxA10b, were similar and found around somite level 7 at 2 and 3 dpfs (Fig. 2b and c, boxes b and f).
There was no or weak expression of hoxA11b and hoxA13b in all stages examined, while paralogous hoxA11a and hoxA13a were expressed. Because of weak staining, the precise boundaries of the hoxA11a expression were difficult to decide at 2 and 3 dpfs. However, these boundaries were more posterior than the boundaries of hoxA9a/A9b and hoxA10a/A10b. In contrast, the hoxA13a expression appeared strong in the tail ends and the cloaca area at 2 and 3 dpfs (Fig. 2a–c, boxes c and d).
Expression of medaka Abd-B genes in pectoral fin buds
shh and dHAND expression
shh and dHAND are typical posterior markers in fin and limb bud development (Riddle et al. 1993; Akimenko and Ekker 1995; Charité et al. 2000) and were found to be expressed in the posterior end of the pectoral fin buds at stage 3 dpf (Figs. 3a and 4a), but not before. The fin buds are not well developed at stage 1 and 2 dpfs, and also, Abd-B genes in hox clusters were not or only weakly expressed (data not shown).
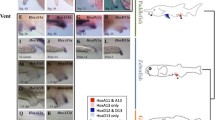
Expression of medaka Abd-B genes in pectoral fin buds—lateral views. Expression of the Abd-B genes in medaka embryos at 3 dpf was assessed by WISH experiments. In figure of pectoral fin buds (a, b–d; b, a–l), the anterior is to the left and the distal end is to the top. Scale bars in the bottom right of figures (a, d; b, l) indicate 0.05 mm. a Lateral view of marker gene expression. shh and dHAND are posterior markers in pectoral fin buds. myoD is a muscle mass marker in pectoral fin buds. Total view of shh expression (a), and magnified views of shh, dHAND, and myoD expression in pectoral fin buds (b, c, d). b hoxAa, Ab, and Da expression in pectoral fin buds. A, anterior; Di, distal; Pr, proximal; Po, posterior
Expression of medaka Abd-B genes in pectoral fin buds—overhead views. Overhead views of the same samples in Fig. 3. The anterior is to the left and the ventral side is to the top. Scale bars in the bottom right of figures (a, d; b, l) indicates 0.05 mm. a Overhead view of marker gene expression. shh and dHAND are posterior markers of fin buds. myoD is a muscle mass marker in pectoral fin buds. Total view of shh expression (a), and magnified views of shh, dHAND, and myoD expression in pectoral fin buds (b, c, d). b hoxAa, hoxAb, and hoxDa expressions. A, anterior; Do, dosal; V, ventral; Po, posterior
hoxDa expression
Expression patterns of Abd-B genes in Ola–hoxDa in lateral and dorsal views at 3 dpf are summarized in Figs. 3b and 4b, respectively. When the positions of shh and dHAND expression were used as posterior markers, hoxD9a and hoxD10a are expressed in distal–medial portion with moderate posterior restriction (Figs. 3b and 4b, boxes i and j), whereas hoxD11a and hoxD12a exhibited strong posterior restriction (Figs. 3b and 4b, boxes k and l). Furthermore, genes located more 3′ downstream on the chromosome (Fig. 1a), such as hoxD9a and hoxD10a, were found to have more advanced boundaries from anterior to posterior sites, clearly following the colinearity rule of hox expression. The boundary order was hoxD9a > hoxD10a > hoxD11a > hoxD12a (Figs. 3b, 4b, and 5), along the anterior–posterior axis of the fin buds (Fig. 5). Thus expression of the hoxDa genes, such as hoxD9a, hoxD10a, hoxD11a and hoxD12a, was shown to be similar to those of orthologous genes in mammals (Dollé et al. 1989), aves (Yokouchi et al. 1991) and zebrafish (Sordino et al. 1995), indicating maintenance of “orthodox” HoxD expression.
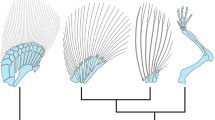
Comparison of expression patterns of Abd-B genes in the pectoral fin buds. a Schematic lateral view. The anterior is to the left and the distal end is to the top. In the medaka (Ola), hoxAa expression is drawn with horizontal stripes and hoxAb and Da expression with vertical stripes. Patterns for zebrafish Danio rerio (Dre) are after Sordino et al. (1996) and Neumann et al. (1999; see the details in the text). Those for the ray-finned fish, Polyodon spathula (Psp), are after Metscher et al. (2005). Question marks indicate expression patterns not yet examined in Dre or not identified in Psp. A, anterior; Di, distal; Pr, proximal; Po, posterior. b Schematic overhead view. The anterior is to the left and the ventral side to the top. In pectoral fin buds of the medaka (Ola), regions drawn with horizontal and vertical stripes are as in a. In wings of the chicken Gallus gallus (Gga), expression of HoxA proteins examined by Hox-protein-specific antibodies is categorized into two types; the mesenchyme type (vertical stripes) and the dorso-ventral muscle mass type (horizontal stripes). The latter shows a split pattern (Yamomoto et al. 1998). These two type expressions overlap in overhead views (cross patterns). A, anterior; Do, dosal; V, ventral; Po, posterior
hoxAa and hoxAb expression
Expression of the Abd-B genes in Ola–hoxAa, such as hoxA9a, hoxA10a, and hoxA11a, was localized in proximal portions of pectoral fin buds, hoxA13a being lacking. Lateral and overhead views are shown in Figs. 3b and 4b with a summary in Fig. 5. Thus, the colinearity rule was not followed for Ola–hoxAa expression. Note here that genes in Ola–hoxAa were found to be in split parts; one in the dorsal and the other in the ventral region (Figs. 4b and 5b). These split expressions were similar to that of myoD, a muscle mass marker (Fig. 4a, box d).
Ola–hoxAb forms such as hoxA9b, hoxA10b, and hoxA11b were localized in distal–posterior portions of buds as shown in lateral views (Fig. 3b, boxes e–h). HoxA13b exhibited stronger posterior restriction than other hoxAb members (Fig. 3b, box h). Overhead views are also summarized in Fig. 5b. The boundaries advanced from posterior to anterior and distal to proximal as recognized in lateral (Figs. 3b and 5a) and overhead views (Figs. 4b and 5b), respectively, in the order of hoxA10b > hoxA9b > hoxA11b > hoxA13b.
Discussion
Expressional boundaries of duplicated Abd-B genes in the trunk
To facilitate comparison of expression of duplicated Abd-B genes in the central nervous system (CNS), we here show schematic diagrams of their expression boundaries on somite levels (Fig. 2d). These diagrams were based on comparisons between hox and myoD expressions at equivalent stages. It was found that genes more 3′ downstream on the chromosome have more anterior boundaries, especially in the Ola–hoxAa case (Fig. 2d). On the other hand, expression of Ola–hoxAb does not obey the rule. It is clear that, at least in the trunks, expression of Ola–hoxAa, i.e., hoxA9a, hoxA10a, hoxA11a and hoxA13a, agrees with the colinearity rule, but that Ola–hoxAb is an exception. As described in zebrafish (Sordino et al. 1996), the expression boundaries of medaka Abd-B hoxA situate more anteriorly than in mouse. When comparing between medaka and zebrafish, Ola–hoxA10a and A10b seem to have more anterior expression boundaries than Dre–hoxA10 (Prince et al. 1998), and Ola–hoxA9a expression is found striking more anterior than Dre–hoxA9a (accession no. Y14538; there listed as Dre–hoxx9). The Ola–hoxA13a expressions at the posterior tip and in the cloaca area are similar to the expression of Dre–hoxA13b (accession no. Y07699; there listed as Dre–hoxA13) and Dre–hoxD13 (Sordino et al. 1996; van der Hoeven et al. 1996a). In contrast, Ola–hoxA13b, the orthologue of the Dre–hoxA13b gene, was not expressed in these regions. Based on these differences between medaka and zebrafish, we hypothesize that hox duplications allowed expressional differentiation and contributed to teleost evolution.
The present study describes differential expression patterns for paralogous genes in medaka hoxAa and hoxAb. Our finding that the anterior boundary of hoxA9b was shifted by 3 or 4 somite levels to the posterior side, compared with that of hoxA9a, whereas that of hoxA10b seem almost equal to that of hoxA10a. Whereas hoxA11a and hoxA13a were expressed, hoxA11b and hoxA13b expression was not detectable. These variations in expression of each paralogous genes are probably caused by a local regulatory changes (Gérard et al. 1993; van der Hoeven et al. 1996b). Belting et al. (1998) and Anand et al. (2003) earlier claimed that expressional boundary changes were dependent on differences in local enhancers for the HoxC8 gene. Furthermore, it is reported in a pufferfish Takifugu rubripes that paralogous genes, hoxA2a and hoxA2b, show differential expression patterns caused by local cis-regulatory changes (Tümpel et al. 2006).
The colinearity rule is explained by the presence of the GCR, previously identified in mouse hoxD cluster (Spitz et al. 2003). However, the report of hox random integration in the genome supports that local control regions make some endogenous aspects of the hox expressions (van der Hoeven et al. 1996b). We hypothesize that lack of some parts of the GCR for the trunk caused the colinearity break and that local control regions contribute to the hoxA9b and A10b expressions. Conservations of group 9–13 in both clusters may be explained by the GCR specificity for fin buds.
Difference of expression patterns between duplicated Abd-B genes in pectoral fin buds
In Fig. 5, to facilitate comparisons, WISH results for zebrafish Danio rerio (Dre) from Sordino et al. (1996) and Neumann et al. (1999), for the basal ray-finned fish Polyodon spathula (Psp) from Metscher et al. (2005) and expression patterns of Hox proteins from a chick Gallus gallus (Gga) described by Yokouchi et al. (1991) are included. Note here that hoxA expression of the zebrafish in the pectoral fin bud reported by Sordino et al. (1996) and Neumann et al. (1999) should be revised here in their nomenclature. At the time of their publication, only one hoxA cluster in zebrafish was identified, and the extra duplication had not been assumed. Our homology comparisons suggest that the hoxA sequences belong to Dre–hoxAb (data not shown). Furthermore, it has been reported that the Dre–hoxAa cluster lacks not only hoxA6a but also hoxA2a, hoxA7a and A10a, with more loss of genes than with the medaka hoxAa cluster (Amores et al. 1998; Kurosawa et al. 2006). Thus, in the present paper, we used the results by Sordino et al. (1996) and Neumann et al. (1999) with Dre–hoxA9b for hoxA9, Dre–hoxA10b for hoxA10, Dre–A11b for hoxA11, and Dre–hoxA13b for hoxA13.
It is well known that muscles in the pectoral fin buds feature distinct two parts, ventral and dorsal muscle masses. Our result of the myoD expression exhibited these muscle masses in the medaka (Figs. 4a, box d; and 5b) and was equal to the results of the zebrafish (Neyt et al. 2000; Haines et al. 2004). Haines et al. (2004) proposed that these muscle masses are equal to those shown in chick wing buds by Dietrich et al. (1999). Thus, the dorsal and ventral region hoxAa expression documented here reflects the two muscle masses in the pectoral fin buds. In wings of the chicken G. gallus (Gga), expression of Gga–HoxA is also categorized in two types; the mesenchyme type (vertical stripes in Fig. 5) and dorso-ventral muscle mass type (horizontal stripes). The latter shows a split pattern (Yamomoto et al. 1998). As in the case of Gga–HoxA, Ola–hoxAa expression well corresponds to the positions of ventral and dorsal muscle masses in the pectoral fin buds. On the other hand, Ola–hoxAb expression corresponds to the mesenchyme tissue. Bruce et al. (2001) claimed that biochemical functions of duplicate hoxB5 genes, i.e., Dre–hoxB5a and Dre–hoxB5b, are not changed, and the ancestral function of HoxB5 must be divided (subfunctionalization). The Ola–hoxAa and Ola–hoxAb, which are duplicates of HoxA, well share the expression patterns of HoxA in Gga, suggesting such subfunctionalization of the ancestral gene. Most importantly, the present study demonstrated this subfunctionalization happened at the level of clusters, not at the level of paralogous genes. Namely, all genes on the hoxAa cluster are expressed in muscles and those on the hoxAb cluster in mesenchymes. Although GCR-like structures have not been identified in these clusters, the result strongly suggests that the GCR changed its character after duplication, thus leading to this tissue-specific expression of each cluster.
References
Akimenko MA, Ekker M (1995) Anterior duplication of the Sonic hedgehog expression pattern in the pectoral fin buds of zebrafish treated with retinoic acid. Dev Biol 170:243–247
Amores A, Force A, Yan YL, Joly L, Amemiya C, Fritz A, Ho RK, Langeland J, Prince V, Wang YL, Westerfield M, Ekker M, Postlethwait JH (1998) Zebrafish hox clusters and vertebrate genome evolution. Science 282:1711–1714
Amores A, Suzuki T, Yan YL, Pomeroy J, Singer A, Amemiya C, Postlethwait JH (2004) Developmental roles of pufferfish Hox clusters and genome evolution in ray-fin fish. Genome Res 14:1–10
Anand S, Wang WC, Powell DR, Bolanowski SA, Zhang J, Ledje C, Pawashe AB, Amemiya CT, Shashikant CS (2003) Divergence of Hoxc8 early enhancer parallels diverged axial morphologies between mammals and fishes. Proc Natl Acad Sci USA 100:15666–15669
Belting HG, Shashikant CS, Ruddle FH (1998) Modification of expression and cis-regulation of Hoxc8 in the evolution of diverged axial morphology. Proc Natl Acad Sci USA 95:2355–2360
Bruce AE, Oates AC, Prince VE, Ho RK (2001) Additional hox clusters in the zebrafish: divergent expression patterns belie equivalent activities of duplicate hoxB5 genes. Evol Dev 3:127–144
Burke AC, Nelson CE, Morgan BA, Tabin C (1995) Hox genes and the evolution of vertebrate axial morphology. Development 121:333–346
Charité J, McFadden DG, Olson EN (2000) The bHLH transcription factor dHAND controls Sonic hedgehog expression and establishment of the zone of polarizing activity during limb development. Development 127:2461–2470
Dietrich S, Abou-Rebyeh F, Brohmann H, Bladt F, Sonnenberg-Riethmacher E, Yamaai T, Lumsden A, Brand-Saberi B, Birchmeier C (1999) The role of SF/HGF and c-Met in the development of skeletal muscle. Development 126:1621–1629
Dollé P, Izpisúa-Belmonte JC, Falkenstein H, Renucci A, Duboule D (1989) Coordinate expression of the murine Hox-5 complex homeobox-containing genes during limb pattern formation. Nature 342:767–772
Duboule D, Dollé P (1989) The structural and functional organization of the murine HOX gene family resembles that of Drosophila homeotic genes. EMBO J 8:1497–1505
Ferrier DE, Minguillón C, Holland PW, Garcia-Fernàndez J (2000) The amphioxus Hox cluster: deuterostome posterior flexibility and Hox14. Evol Dev 2:284–293
Gehring WJ, Affolter M, Bürglin T (1994) Homeodomain proteins. Annu Rev Biochem 63:487–526
Gérard M, Duboule D, Zákány J (1993) Structure and activity of regulatory elements involved in the activation of the Hoxd-11 gene during late gastrulation. EMBO J 12:3539–3550
Haines L, Neyt C, Gautier P, Keenan DG, Bryson-Richardson RJ, Hollway GE, Cole NJ, Currie PD (2004) Met and Hgf signaling controls hypaxial muscle and lateral line development in the zebrafish. Development 131:4857–4869
Harding K, McGinnis W, Wedeen C, Levine M (1985) Spatially regulated expression of homeotic genes in Drosophila. Science 229:1236–1242
Hart CP, Fainsod A, Ruddle FH (1987) Sequence analysis of the murine Hox-2.2, -2.3 and -2.4 homeo boxes: evolutionary and structural comparisons. Genomics 1:182–195
Holland PW (1999) Gene duplication: past, present and future. Semin Cell Dev Biol 10:541–547
Holland PW, Garcia-Fernàndez J (1996) Hox genes and chordate evolution. Dev Biol 173:382–395
Inohaya K, Yasumasu S, Ishimaru M, Ohyama A, Iuchi I, Yamagami K (1995) Temporal and spatial patterns of gene expression for the hatching enzyme in the teleost embryo, Oryzias latipes. Dev Biol 171:374–385
Inohaya K, Yasumasu S, Yasumasu I, Iuchi I, Yamagami K (1999) Analysis of the origin and development of hatching gland cells by transplantation of the embryonic shield in the fish, Oryzias latipes. Dev Growth Differ 41:557–566
Iwamatsu T (2004) Stages of normal development in the medaka Oryzias latipes. Mech Dev 121:605–618
Izpisúa-Belmonte JC, Falkenstein H, Dollé P, Renucci A, Duboule D (1991) Murine genes related to the Drosophila Abd-B homeotic genes are sequentially expressed during development of the posterior part of the body. EMBO J 10:2279–2289
Kessel M, Gruss P (1990) Murine development control genes. Science 249:374–379
Krumlauf R (1992) Evolution of the vertebrate Hox homeobox genes. Bioessays 14:245–252
Kurosawa G, Yamada K, Ishiguro H, Hori H (1999) Hox gene complexity in medaka fish may be similar to that in pufferfish rather than zebrafish. Biochem Biophys Res Commun 260:66–70
Kurosawa G, Takamatsu N, Takahashi M, Sumitomo M, Sanaka M, Yamada K, Nishii K, Matsuda M, Asakawa S, Ishiguro H, Miura K, Kurosawa Y, Shimizu N, Kohara Y, Hori H (2006) Organization and structure of hox gene loci in medaka genome and comparison with those of pufferfish and zebrafish genomes. Gene 370:75–82
Levine M, Hafen E, Garber RL, Gehring WJ (1983) Spatial distribution of Antennapedia transcripts during Drosophila development. EMBO J 2:2037–2046
Metscher BD, Takahashi K, Crow K, Amemiya C, Nonaka DF, Wagner DP (2005) Expression of Hoxa-11 and Hoxa-13 in the pectoral fin of a basal ray-finned fish, Polyodon spathula: implications for the origin of tetrapod limbs. Evol Dev 7:186–195
Naruse K, Fukamachi S, Mitani H, Kondo M, Matsuoka T, Kondo S, Hanamura N, Morita Y, Hasegawa K, Nishigaki R, Shimada A, Wada H, Kusakabe T, Suzuki N, Kinoshita M, Kanamori A, Terado T, Kimura H, Nonaka M, Shima A (2000) A detailed linkage map of medaka, Oryzias latipes: comparative genomics and genome evolution. Genetics 154:1773–1784
Neumann CJ, Grandel H, Gaffield W, Schulte-Merker F, Nüsslein-Volhard C (1999) Transient establishment of anterior–posterior polarity in the zebrafish pectoral fin bud in the absence of sonic hedgehog activity. Development 126:4817–4826
Neyt C, Jagla K, Thisse C, Thisse B, Haines L, Currie PD (2000) Evolutionary origins of vertebrate appendicular muscle. Nature 408:82–86
Prince VE, Joly L, Ekker M, Ho RK (1998) Zebrafish hox genes: genomic organization and modified collinear expression patterns in the trunk. Development 125:407–420
Riddle RD, Johnson RL, Laufer E, Tabin C (1993) Sonic hedgehog mediates the polarizing activity of the ZPA. Cell 75:1401–1416
Santini S, Boore JL, Meyer A (2003) Evolutionary conservation of regulatory elements in vertebrate Hox gene clusters. Genome Res 13:1111–1122
Schwartz S, Zhang Z, Frazer KA, Smit A, Riemer C, Bouck J, Gibbs R, Hardison R, Miller W (2000) PipMaker—a web server for aligning two genomic DNA sequences. Genome Res 10:577–586
Scott MP (1992) Vertebrate homeobox gene nomenclature. Cell 71:551–553
Sordino P, van der Hoeven F, Duboule D (1995) Hox gene expression in teleost fins and the origin of vertebrate digits. Nature 375:678–681
Sordino P, Duboule D, Kondo T (1996) Zebrafish Hoxa and Evx-2 genes: cloning, developmental expression and implication for the functional evolution of posterior Hox genes. Mech Dev 59:165–175
Spitz F, Gonzalez F, Duboule D (2003) A global control region defines a chromosomal regulatory landscape containing the HoxD cluster. Cell 113:405–417
Tümpel S, Cambronero F, Widedemann LM, Krumlauf R (2006) Evolution of cis element in the differential expression of two Hoxa2 coparalogous genes in pufferfish (Takifugu rubripes). Proc Natl Acad Sci USA 103:5419–5424
van der Hoeven F, Sordino P, Fraudeau N (1996a) Teleost HoxD and HoxA genes: comparison with tetrapods and functional evolution of the HOXD complex. Mech Dev 54:9–21
van der Hoeven F, Zákány J, Duboule D (1996b) Gene transpositions in the HoxD complex reveal a hierarchy of regulatory controls. Cell 85:1025–1035
Yamomoto M, Gotoh Y, Tamura K, Tanaka M, Kawakami A, Ide H, Kuroiwa A (1998) Coordinated expression of Hoxa-11 and Hoxa-13 during limb muscle patterning. Development 125:1325–1335
Yokouchi Y, Sasaki H, Kuroiwa A (1991) Homeobox gene expression correlated with the bifurcation process of limb cartilage development. Nature 353:443–445
Acknowledgement
We thank Dr. Keiji Inohaya for the instruction regarding the medaka in situ method, Susumu Hamada, Makiko Tsutsumi, and Rieko Yamamoto for their help in manipulating the medaka Hd–rR BAC library. We also thank Dr. Johannes Martinus Dijkstra for helpful discussions. This work was supported in part by grants from the Ministry of Education, Culture, Sports, Science, and Technology in Japan and by Grant-in-Aid for Special Project Research to H.H. (No. 12202004).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Hollemann
Rights and permissions
About this article
Cite this article
Takamatsu, N., Kurosawa, G., Takahashi, M. et al. Duplicated Abd-B class genes in medaka hoxAa and hoxAb clusters exhibit differential expression patterns in pectoral fin buds. Dev Genes Evol 217, 263–273 (2007). https://doi.org/10.1007/s00427-007-0137-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-007-0137-4