Abstract
Main conclusion
Application of the recently developed CRISPR/Cas tools might help enhance cereals’ growth and yield under biotic and abiotic stresses.
Abstract
Cereals are the most important food crops for human life and an essential source of nutrients for people in developed and developing countries. The growth and yield of all major cereals are affected by both biotic and abiotic stresses. To date, molecular breeding and functional genomic studies have contributed to the understanding and improving cereals’ growth and yield under biotic and abiotic stresses. Clustered, regularly inter-spaced, short palindromic repeats (CRISPR)/CRISPR-associated protein (Cas) system has been predicted to play a major role in precision plant breeding and developing non-transgenic cereals that can tolerate adverse effects of climate change. Variants of next-generation CRISPR/Cas tools, such as prime editor, base editor, CRISPR activator and repressor, chromatin imager, Cas12a, and Cas12b, are currently used in various fields, including plant science. However, few studies have been reported on applying the CRISPR/Cas system to understand the mechanism of biotic and abiotic stress tolerance in cereals. Rice is the only plant used frequently for such studies. Genes responsible for biotic and abiotic stress tolerance have not yet been studied by CRISPR/Cas system in other major cereals (sorghum, barley, maize and small millets). Examining the role of genes that respond to biotic and abiotic stresses using the CRISPR/Cas system may help enhance cereals’ growth and yield under biotic and abiotic stresses. It will help to develop new and improved cultivars with biotic- and abiotic-tolerant traits for better yields to strengthen food security. This review provides information for cereal researchers on the current status of the CRISPR/Cas system for improving biotic and abiotic stress tolerance in cereals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
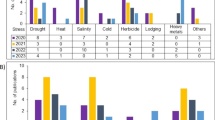
Cereals are considered staple food crops in both developed and developing countries. Its whole grain contains rich sources of vitamins, minerals, carbohydrates, fats, oils, and protein. Cereal foods reduce the risk of certain diseases, including diabetes, colon cancer, coronary heart disease, and diverticular disease (Sarwar et al. 2013; Maharajan et al. 2021a). More than 50% of the world’s daily caloric intake is derived directly from cereal consumption (Awika 2011). The growth and yield of cereals are severely affected by biotic (fungi, bacteria, and viruses) and abiotic (drought, salinity, heat, heavy metals, nutrient deficiency, and cold) stresses. These stresses reduce cereals’ productivity by up to > 75% depending on the cereal and geographic location. Among cereal diseases, blast, sheath blight, sheath rot, brown leaf spot, bacterial leaf blight, tungro virus, bakanae disease, leaf scald, false smut, and grain discoloration are more prevalent in all cereals, particularly in rice (Oryza sativa) (Sumit et al. 2020). Among these, bacterial leaf blight disease is caused by Xanthomonas oryzae pv. Oryzae most severely affects rice growth and yield. Approximately, 50% of rice yield is estimated to be lost due to this disease worldwide (Fiyaz et al. 2022). In wheat (Triticum aestivum), leaf streak and black chaff diseases are caused by X. translucens pv. undulosa. These diseases reduce annual wheat production by up to 40% (Tambong 2022). Fungal diseases such as tan spot, fusarium head blight, stripe rust, septoria leaf blotch, spot blotch, and powdery mildew are the most common fungal diseases of cereals, reducing yield by 15–20% and in extreme cases, up to 60% (Rozewicz et al. 2021). Among abiotic stresses, drought stress reduces the morphological traits (leaf size and width, shoot length, plant height), physiological and biochemical processes, including respiration, photosynthesis, carbohydrates, and nutrient metabolism of maize (Zea mays)(Zhang et al. 2018a), barley (Hordeum vulgare) (Sallam et al. 2019), sorghum (Sorghum bicolor) (Queiroz et al. 2019) and other cereals (Rakkammal et al. 2022; David et al. 2021). Salinity is another major stress and poses a threat to crop production. About 20% of the agricultural land worldwide is affected by saline soils, increasing daily (Shrivastava and Kumar 2015). Salinity stress reduced wheat yield by more than 45% (Ali et al. 2009). It also reduces the grain per spike, 1000-grain weight, and wheat yield (Hasan et al. 2015). Nutrient deficiency (macro and micronutrient deficiency) reduces plant biomass, growth, and nutrient contents in below and above-ground traits and yield by > 50% in various cereals, including rice (Muthukumararaja and Sriramachandrasekharan 2012), wheat (Plenet et al. 2000; Bagci et al. 2007; Maharajan et al. 2021b), sorghum(Afzal et al. 2012), barley (Shafi et al. 2011), maize (Hong and Jin 2007), and small millets (Maharajan et al. 2019; Ceasar et al. 2014; Roch et al. 2020). These reports indicate that both biotic and abiotic stresses considerably reduce cereal production. Researchers try to develop new cultivars tolerant to biotic and abiotic stresses through various biotechnological approaches, including genome editing.
Genome editing is a technique used to create defined/desired changes in the genetic composition of an organism. The main scope of this technology is the use of site-based nuclei that precisely target specific DNA sequences. There are various tools such as zinc-finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), and clustered, regularly inter-spaced, short palindromic repeats (CRISPR)/CRISPR-associated protein (Cas) system are used for genome editing. Genome editing using the CRISPR/Cas system was first reported for model plants in 2013 (Feng et al. 2013). Since then, genomes of many plants have been edited through CRISPR/Cas system. All approaches other than the CRISPR tool are expensive and include complex procedures for construct design and delivery. This review summarizes the details of CRISPR/Cas-based tools and their applications. We also enlist detailed information on the CRISPR/Cas system to improve cereals’ biotic and abiotic stress tolerance. We also draw insights and future direction on harnessing the outputs of functional genomics studied for the potential application of the CRISPR/Cas system to improve biotic and abiotic stress tolerance in cereals. This review will help cereal researchers to know about the application of various CRISPR/Cas tools to improve the growth and yield of cereals under biotic and abiotic stresses.
Effect of biotic and abiotic stresses in cereals
Plant biotic stresses are caused by various organisms such as bacteria, fungi, insects, and viruses. Drought, salinity, heat, cold, nutrient deficiency, and heavy metals stresses are major factors causing abiotic stresses in plants. Several reports have proved that biotic and abiotic stresses severely reduce the growth and yield of cereals. Bacterial leaf blight disease reduced rice yield and grain quality at different growth stages (panicle formation, booting stage, and milk stage) (Noh et al. 2007). Fusarium stalk rot and charcoal rot disease reduced total seed weight, 100-seed weight, and seeds per panicle of sorghum (Bandara et al. 2017). In cereals, blast disease is the most devastating disease caused by Magnaporthe oryzae. Blast disease affects all cereals’ leaf, stem, collar, node, neck, and panicle growth. It has also reduced the growth and yield of all economically important cereals such as rice (Kihoro et al. 2013), foxtail millet (Setaria italica) (Sharma et al. 2014), finger millet (Eleusine coracana) (Gashaw et al. 2014), barley (Aghnoum et al. 2019), and wheat (Cruz and Valent 2017). Yellow dwarf virus reduced the yield of barley and wheat by > 40% (Edwards et al. 2001; Perry et al. 2000). In rice, drought stress reduced yield and yield-related traits such as, panicle length, number of grains, and spikelet per panicle, spikelet fertility, number of ear bearing tillers and biological yield per plant (Singh et al. 2012). Drought stress reduced barley yield (56.8%) due to a reduction in the number of spikes, tillers, and grains per plant (Samarah 2005). The grain number of wheat was reduced by > 51% due to pollen sterility under drought stress (Dong et al. 2017). Drought stress also reduces the morphological traits (leaf size and width, shoot length, plant height), physiological and biochemical processes, including respiration, photosynthesis, carbohydrates, nutrient metabolism, and growth promoters in maize (Wang et al. 2019a), barley (Alghabari and Ihsan 2018), sorghum (Maharajan et al. 2021c) and finger millet (David et al. 2021). Many researchers reported that salinity stress decreases spikelet number, kernel weight, grain yield, and the number of fertile tillers in wheat (Saddiq et al. 2021; Izadi et al. 2014). In rice, salinity stress inhibits seed germination and leaf area development; reduces plant growth, biomass, and grain yield components (Hussain et al. 2017). Salinity stress also reduced the concentration of phosphorus, potassium, fat, protein, and fiber in wheat grain (Abbas et al. 2013). Under salt stress, panicle sterility is a major issue during grain development in rice (Pruthi et al. 2022). For example, several studies have shown that salt stress causes panicle sterility in rice, which leads to decreased grain setting, pollen-bearing capacity, and stigmatic surface (Gerona et al. 2019). The salt stress reduced the panicle length, the number of florets, the number of tillers, and the 1000-grain weight of rice (Rahman et al. 2015; Aref and Rad 2012). The number of grains per panicle, 1000-grain weight, panicle length, and total rice yield decreased under cold stress at the heading and flowering stages (Li et al. 2022). Macro- and micronutrients are essential for cereals’ growth and development. Nutrient deficiency altered root architecture and reduced plant biomass and yield in all cereals (Maharajan et al. 2018, 2019; Krishna et al. 2017). All these studies confirmed that biotic and abiotic stresses reduced grain yield and their components in cereals.
Genome-editing tools
Genome editing is the process of changing the genetic code of an organism. In conventional genome-editing systems, enzymes were guided by proteins to cut DNA at the specific and targeted location and create the double-standard break (DSB). DSB repair occurs by either Non-Homologous End Joining (NHEJ) or homology-directed repair (HDR). The NHEJ produces random mutations (gene knockout), while HDR uses additional DNA to generate a desired sequence within the genome (gene knockin). However, apart from simple breaks, various other forms of genome editing have evolved from CRISPR/Cas. Three types of tools, including ZFNs, TALENs, and CRISPR/Cas, are used for genome-editing technology. ZFNs are custom-designed and targetable DNA cleavage proteins that cut DNA sequences at specific sites (Carroll 2011). They facilitate targeted gene editing by inducing the DSB in DNA to replace the gene by homologous recombination. ZFN contains two domains: DNA binding and cleaving (Gupta et al. 2012). The DNA binding domain recognizes a unique 6-base pair in the DNA sequence, while the DNA cleaving domain consists of a FokI nuclease (Asmamaw and Zawdie 2021). These two domains are linked together to form a zinc-finger protein. When both domains are fused, they form a highly specific genomic scissor. The ZFNs are expensive, difficult to handle, and time-consuming, thus limiting their widespread use, especially in high-throughput studies.
CRISPR is a family of repetitive DNA sequences found in the genomes of archaea (84%) and bacteria (45%). It was first detected downstream of the alkaline phosphatase isozyme gene in Escherichia coli (Ishino et al. 1987). It is formerly known as short, regularly spaced repeats (SRSRs) and helps to detect and destroy the DNA of the virus. The CRISPR system uses small guide RNAs (gRNAs) for sequence-specific interference with invading nucleic acids. CRISPR is an array of short repeated sequences (repeats) separated by unique sequences (spacers). The spacers and repeats are derived from the virus’s nucleic acid and plasmids. Some of the proteins involved in the CRISPR mechanism are called CRISPR/Cas, which can search, cut, and finally transform phage DNA in a specific way. Cas is a protein with an enzymatic activity that plays a distinct role in the DNA sequences and CRISPR arrays with nuclease activity. In general, CRISPR/Cas mechanism can be separated into three steps (1) insertion of unique sequences into the CRISPR locus (spacer acquisition or adaptation); (2) transcription of CRISPR locus and processing of gRNA (expression or gRNA biogenesis); (3) detection and degradation of nucleic acids by gRNA and Cas proteins (target interference) (Cong et al. 2013; Mali et al. 2013; Liu et al. 2017). The CRISPR/Cas system has been classified into two classes (Classes I and II) (Makarova and Koonin 2015; Makarova et al. 2020). Both classes of CRISPR systems could be used for genome editing; however, the class II system is more desirable for genome editing because the methods of class II systems are due to a much-simplified protocol. The class II CRISPR/Cas systems are categorized by three signature proteins, Cas1, 2, and 9, which include three subtypes (II-A, II-B, and II-C). Of the three proteins, Cas9 is the most widely used CRISPR system for genome editing in various organisms. The recently developed CRISPR/Cas12a and Cas12b come under class VA and VB, respectively.
Mechanism and variants of CRISPR/Cas system
CRISPR/Cas system is a user-friendly genome-editing tool with many advantages over other genome-editing systems. It contains two essential components: gRNA and Cas protein. Cas protein is an RNA-dependent DNA endonuclease that forms a complex with gRNA to target the specific location in the genome (Jinek et al. 2012). The idea of using gRNA and exploiting CRISPR as a genome-editing tool was introduced by the collaborative work of Emmanuelle Charpentier and Jennifer A. Doudna (Jinek et al. 2012), for which they were also awarded the Nobel Prize in 2020. The gRNA is made of two parts, CRISPR RNA (crRNA), and trans-activating CRISPR RNA (tracrRNA). The length of crRNA is 18–20 base pairs which bind to the target DNA by attaching to the target sequence, whereas tracrRNA is a long extension of the loops that act as a binding scaffold for the Cas protein. The designed gRNA activates Cas9 and recognizes the target sequence in the gene of interest by its 5ʹcrRNA complement base pair component. In general, the widely used Cas9 from Streptococcus pyogenes (SpCas9) primarily recognizes the PAM sequence at 5ʹ-NGG-3" (N can be any nucleotide base). Once gRNA detects a target site with the suitable PAM, it subsequently forms the RNA–DNA hybrid. The complementary and non-complementary strands of target DNA are cut by HNH and RuvC domains of Cas9, respectively. Many researchers have used the CRISPR/Cas system to improve plant growth under various biotic and abiotic stresses (Ceasar et al. 2016; Hillary and Ceasar 2019; Altpeter et al. 2016; Krishna et al. 2022a). Therefore, several reports are now available to review and predict this method for crop improvement. There were some limitations with the first generation CRISPR/Cas9 system, especially due to off-target effects caused by the wild Cas9 from S. pyogenes, randomly digested DNA at off-target sites (Yang et al. 2021). Several improvements were made to overcome this problem, and many high-fidelity Cas9 with engineered residues were introduced, such as HF-Cas9, eSpCas9, HypaCas9, and Sniper Cas9, with reduced off-target effects (Lee et al. 2018, 2019; Chen et al. 2017; Hu et al. 2018).
To improve the CRISPR/Cas9 system, Qi et al. (2013) developed the dead Cas9 (dCas9) system. In dCas9, H840A and D10A mutations were introduced in HNH and RuvC domains of Cas9, respectively, to inactivate the nuclease activity (Qi et al. 2013). The dCas9 system cannot cleave DNA but can target and bind DNA with the same specificity when guided by gRNA. Many gene-editing technologies have emerged based on dCas9. Currently, the dCas9 system is used in various molecular genetic applications, such as regulation of gene expression, base editing, primer editing, chromatin topology, epigenome editing, chromatin imaging, and reducing the off-target effects (Fig. 1). The CRISPR-dCas9 components, dCas9, and effectors (activator and repressor) help to enhance the efficiency of transcriptional regulation (Xu and Qi 2019). When the dCas9 protein is fused with transcriptional activators (VP64), it regulates/activates the gene’s transcription, a process called CRISPR activator (CRISPRa). When dCas9 binds to the target site of the gene, it effectively inhibits the expression of the target gene. This process is called CRISPR interference (CRISPRi). In general, gRNAs play a significant role in CRISPRa and CRISPRi because dCas9 activators and repressors are guided by gRNA, which may enhance or inhibit the transcription of the target gene (Moradpour and Abdulah 2020). When the target gene is activated or suppressed by dCas9 activators or repressors, gRNAs must be targeted to the gene’s promoter region of interest.
Applications of various CRISPR/Cas tools to improve the biotic and abiotic stress tolerance in cereals. When the dCas9 is fused with appropriate repressors, activators, and epigenetic engineering, it helps to find the accurate role of each gene by transcriptional regulation, as indicated in the image. The dCas9 fused with various modulators targeting many potential genes might help enhance cereals’ biotic and abiotic stress tolerance. Abbreviations: ABE, Adenine base editors (facilitates A to G substitution in the DNA); AID, cytidine deaminase (enables C to T/G substitution); APOBEC1, cytidine deaminase enzyme (enables C to T substitution); CBE, cytosine base editors (facilitates C to T substitution in the DNA); DNMTs, DNA methyltransferases (induces site-specific promoter methylation, which results in gene silencing and is heritable across mitotic division); EZH2, histone methyltransferase (facilitates gene silencing); GFP, green fluorescent protein (gene visualization), HDAC3, histone deacetylase 3 (induces locus-specific histone deacetylation that results in gene silencing); LSD1, histone demethylases (allows transcriptional silencing); KRAB, Kruppel associated box domains of Kox1 (transcriptional repression); p65AD, p65 activation domain VPR which is a combination of VP64, p65 and Rta (enhances the transcriptional activity); SMYD3, SET and MYND domain-containing protein 3 (facilitates the trimethylation of histone)
CRISPR/Cas-mediated base editing system allows direct conversion of one target DNA base into another base without DSB. Therefore, base editing technology does not require DSB, donor DNA templates, and HDR. Two base editors (cytosine base editor (CBE) and adenine base editor (ABE)) have been developed through protein engineering (Molla and Yang 2019). All four types of transition mutations (C-T, G-A, A-G, and T-C) can be installed or corrected at target positions in DNA without making DSB by CBE and ABE. The dCas9 fused with ABE allows A-T to G-C conversion, while dCas9 fused with CBE allows C-G to T-A change (Bharat et al. 2020). This editing will be reflected in the proteins, so hence it will be highly useful in introducing the point mutations in proteins and will help to alter the functions of the protein in plants. Many base editor systems have also been introduced, such as dual base editor/saturated targeted endogenous mutagenesis editor (STEME), trans-version base editor, PAMless base editor and multiplex base editor (Azameti and Dauda 2021). STEME allows C-G to T-A and A-T to G-C conversion. It consists of four enzymes such as APOBEC3A (cytidine deaminase), ecTadA (adenosine deaminase), D10A (nCas9) and uracil DNA glycosylase inhibitor (UGI). The STEME converts cytidines and adenosines to uridine and inosines, respectively. Manipulating DNA by CRISPR/Cas system (including ABE and CBE) requires the recognition of PAM sequences. Therefore, the CRISPR/Cas system is highly constrained. To overcome this constraint, the PAMless base editor was created by Walton et al. (2020), called as SpRY-base editor. They have engineered specific variants of the SpCas9 enzyme, named SpG and SpRY, that can potentially edit any gene independent of PAMs requirement. Multiplex base editing modifies two or more specific loci in a genome with high precision (Abdelrahman et al. 2021). Prime editing has many advantages over other methods due to its precise sequence deletion, addition, and substitution (Hillary and Ceasar 2022). The prime editing consists of a SpCas9 nickase (HB40A), reverse transcriptase fusion protein, and prime editing gRNA (pegRNA) instead of a deaminase (Wang et al. 2022). The development of prime editing was a breakthrough because it does not require a PAM sequence near the target site. In addition, it can perform not only all 12 types of point mutations but also involves insertions (up to 44 bp) and deletions (up to 80 bp). Prime editing showed less off-target activity than other CRISPR/Cas systems. Thus, it may help to improve the biotic and abiotic stress tolerance in cereals.
Epigenetic control of the plant’s response to stress is a complex phenomenon. Epigenetic changes not only induce stress but also cause changes in gene expression, which may remain over many generations. CRISPR/Cas-mediated epigenetic engineering could be used to target epigenetic factors (histones) (Pulecio et al. 2017). DNA methyltransferase (DNMT3a and b) fused with dCas9 activates methylation or demethylation at the target site. Histone modification can also be achieved by combining dCas9 and histone modifiers. Gene structure and folding of chromatin within the nucleus are considered important factors for gene expression studies. CRISPR/Cas-mediated chromatin topology modifier system helps to target and manipulate chromatin structure and DNA loop formation, which alters gene expression. The CRISPR/Cas protein and a catalytically inactive or dCas9 system provide a powerful genetic manipulation tool that can steer biological studies based on the ability to achieve targeted modifications, which is critical for phenotypic alteration. Levels of gene expression can be changed through the fusion of dCas9 to transcriptional regulators. Recruitment of these transcriptional modulators to the promoter region close to the transcriptional start site alters the expression of the desired downstream genes. Similarly, fusing dCas9 to epigenetic modulators such as methylation and deacetylation enzymes will establish dCas9-based DNA-chromatin-modifying enzymes for precise epigenome editing directed by gRNA.
Moreover, Cas12a (previously called Cpf1) and Cas12b (previously called C2c1) systems have been identified. Compared to CRISPR/Cas9, CRISPR/Cas12a is better due to its short gRNA nucleotide length and reduced size of the Cas protein (Matres et al. 2021). CRISPR/Cas12a prefers T-rich PAMs and does not require tracrRNA and crRNA because its gRNA only requires crRNA (Haroon et al. 2021). In Cas12a, the DSB are also different from Cas9 because it creates staggered cuts. Identification of the CRISPR/Cas12a system expands the application of genome editing, as it enables the editing of AT-rich regions (such as untranslated and promoter regions) (Zafar et al. 2020). The PAM requirement for Cas12a is “TTTV” or “TTV,” which favors targeting promoters and AT-rich regions in gene coding regions (Gao et al. 2017). However, this system has a drawback because other sites lacking TTTN motifs cannot be identified and edited, thus limiting their use in plants. CRISPR/Cas12b also has an RNA-guided endonuclease like CRISPR/Cas12a. However, CRISPR/Cas12b is smaller in protein size than CRISPR/Cas9 and CRISPR/Cas12a. Like CRISPR/Cas9, CRISPR/Cas12b requires crRNA and tracrRNA for targeting any gene or DNA. CRISPR/Cas12b has the most extended sticky ends of all CRISPR systems, creating DSB with 6–8 nucleotide sticky ends. It can be temperature-inducible; therefore, it helps develop plants’ resistance to high temperatures.
Improving biotic stress tolerance in cereals by CRISPR/Cas system
Cereals are susceptible to a wide range of pathogens that cause several diseases. Different types of pesticides, herbicides, and fungicides control cereals’ diseases, but they all pollute the environment and are toxic to human and animal health. The use of pesticides, herbicides, and fungicides also affects the metabolic pathway of cereals. Hence, CRISPR/Cas technology induces plant resistance against pathogens such as bacteria, viruses, fungi, and insects. Cereal researchers have already targeted some biotic stress-responsive genes by CRISPR/Cas variants, which are discussed below.
Bacterial disease tolerance
Bacteria cause several diseases in cereals, reducing the growth and yield of all cereals. In rice, bacterial blight is a major disease in South Asia and West Africa. Some researchers reported that sucrose family (SWEET) transporters enhanced rice growth and yield when plants are affected by bacterial blight disease. Knockout of the OsSWEET13 gene by CRISPR/Cas9 improved rice growth against bacterial blight disease (Table 1) (Zhou et al. 2015). In another study, three SWEET family genes (SWEET11, 13, and 14) edited by CRISPR/Cas9 system helped to reduce bacterial blight infection and enhance the plant height and panicle length of rice (Oliva et al. 2019). CRISPR/Cas system has yet to target only one family transporter. Several bacterial blights (BB) and resistance (R) family genes were identified and functionally characterized in cereals. Targeting BB and R family genes by CRISPR/Cas variants helps to enhance the cereal’s growth under bacterial disease.
Fungal disease tolerance
Agriculture relies on chemical fungicides to prevent losses from fungal diseases. Developing cereals resistant to fungal diseases without using fungicides help to improve sustainable agriculture and the environment. Wang et al. (2014) edited mildew resistance locus (MLO) genes (TaMLO-A1, B1, and D1) with the help of the CRISPR/Cas9 system to develop powdery mildew wheat-resistant plants. CRISPR/Cas9 system was used to knockout the ethylene-responsive factor 22 (OsERF922) gene in rice, which enabled the development of blast the disease-resistant rice variety without affecting the typical growth of the rice (Wang et al. 2016). Disruption of the subunit exocyst complex 3A (OsSEC3A) gene in rice by the CRISPR/Cas9 system enhanced rice plant growth against blast disease (Ma et al. 2018). Zhang et al. (2017) generated powdery mildew resistance wheat using the enhanced disease resistance1 (EDR1) gene with the help of the CRISPR/Cas9 system. Control of viruses by chemical methods is challenging due to the virulence of pathogenesis. Therefore, targeting the fungal disease tolerance genes by CRISPR/Cas system may inhibit the virulence of pathogens, which helps to control fungi diseases during plant growth.
Insects or pesticide tolerance
Insects are the primary source of virus infection transmission in cereals, so intensive pesticide use is necessary to avoid transmission. The pesticide application is insufficient to control the insects in cereals and not good for human health and the environment. In rice, tungro disease is a severe constraint to rice production throughout tropical Asia, which is caused by the interaction between tungro spherical virus and the tango bacilliform virus. Previously, the translation initiation factor 4 gamma (eIF4G) gene was found to help control tungro spherical viruses in rice (Table 1) (Lee et al. 2010). Gene for eIF4G was mutated by CRISPR/Cas9 system in rice, resulting in the development of rice lines resistant to the tungro spherical virus (Macovei et al. 2018). Among biotic stress, weeds and insects also affect cereals’ growth and yield. Weeds cause maximum damage to cereals. They increase the competition for food, space, shelter, sunlight, water, and fertilizers, inhibiting cereals’ proper growth and development. Many cost-effective herbicides are used against weeds, although repeated applications of the same herbicide can lead to resistance in weeds. A better way to achieve herbicide resistance in plants may be through genome editings, such as base editing and prime editing. We can alter or introduce point mutation in the plants’ genome through the base editor, which helps to develop herbicide-resistant plants. Herbicide-resistance rice plant was developed when acetolactate synthase (OsALS) gene was altered with the help of a base editor (Kuang et al. 2020). In another study, the same OsALS gene was targeted by CBE (editing efficiency 37.5–61.5%) enhanced rice growth against the five herbicides (nicosulfuron, imazapic, pyroxsulam, flucarbazone and bispyribac) (Zhang et al. 2020). Prime editing has been used to develop herbicide-resistance in plants, including cereals (rice and wheat). In cereals, prime editing was first implemented in rice by Butt et al. (2020). Through prime editing, they have targeted OsALS and ideal plant architecture 1 (OsIPA1) genes in rice. Among these, the OsALS gene (editing efficiency 26%) enhanced rice growth against Bispyribac sodium’s herbicide. In addition, the targeted OsIPA1 gene reduced the number of unproductive tillers and improved rice yield. Jiang et al. (2020) have developed herbicide-tolerant maize with higher editing efficiency (53.2%) with the help of two ZmALS genes (ZmALS1 and ZmALS2) by base editing. The development of herbicide-tolerant cereals by genome editing can help to control weeds during cereals cultivation. Functional genomics approaches in cereals characterized the role of many weeds and insect stress-responsive genes. However, the exact role of insect and weed genes in any cereals has yet to be identified by the current CRISPR/Cas system.
Further studies are required to identify the exact role of weed and herbicide-responsive genes, which may help develop non-polluted cereals. In addition, most biotic responsive genes have been functionally characterized only in rice. With rice, researchers should apply the CRISPR/Cas system in other cereals that will help to enhance food production worldwide. Various biotic and abiotic stress-responsive genes have been targeted by CRISPR/Cas variants (ABE, CBE, NG-CBE, SpRY-CBE, and primed editing) to improve tolerance against biotic and abiotic stresses (Table 2). Most of the analysis revealed that all the variants have higher editing efficiency. Therefore, further phenotypic and exploitation of CRISPR/Cas variants may help to develop biotic and abiotic stress resistance cereals.
Improving abiotic stress tolerance in cereals by CRISPR/Cas system
Like biotic stress, abiotic stresses are the most imminent threat to crop production. Several abiotic stress-responsive genes help cereals to survive and produce sufficient yields under abiotic stresses. Various researchers have made substantial efforts to improve crop productivity with the help of abiotic stress-responsive genes through functional genomics approaches. However, the progress has been slow due to its limitations. Genome-editing tools (particularly CRISPR/Cas system) have brought opportunities for precise and efficient manipulation of desired genes to enhance abiotic stress tolerance in cereals. The role of various abiotic stress-responsive genes has already been identified in cereals by CRISPR/Cas system (Table 3). The genome-editing tool has predominantly been applied in rice, and few reports are available in maize, wheat, and barley.
Drought stress tolerance
The role of some drought-responsive genes has been identified in cereals by the CRISPR/Cas system. Stress-activated protein kinase (SAPK) family members of SNF1-related protein kinase 2 (SnRK2) are activated by abscisic acid. The SAPK family members help to enhance plant growth (particularly at initial stages) under various abiotic stresses, including salinity, osmatic, and drought stresses. Lou et al. (2017) generated sapk2 mutants in rice through CRISPR/Cas9 system to characterize the functional properties of SAPK2. They have reported that the sapk2 mutants were more sensitive to drought stress and reactive oxygen species (ROS) than the wild-type plants. This result revealed that SAPK2 is important for responding to drought conditions in rice. In the same study, the additional investigation revealed that the expression level of some abiotic stress-responsive genes was higher in spak mutants. These findings suggest that SAPK2 might be a potential candidate gene for future crop improvement. Developing and cultivating new crops utilizing the SAPK2 locus might help overcome the drought stress in rice. Leaf rolling determines plant architecture, significantly affecting crop yields and subsequent dry matter accumulation. Leaf rolling decreases stomatal conductance and water loss under drought stress. Developing semi-rolled leaf genotypes may provide a source to enhance crop yield under drought stress. Semi-rolled leaf 1 (SRL1) and SRL2 genes play key roles in leaf rolling by severely affecting epidermal growth and structure. CRISPR/Cas9 system was used to develop semi-rolled leaf mutant rice by targeting SRL1 and SRL2 genes (Liao et al. 2019). Knockout of SRL1 and SRL2 genes in rice showed a higher survival rate than the wild type under drought stress (Table 3). Proteomic analysis revealed that 107 proteins were upregulated in the mutant line compared to wild-type plants under drought stress (Liao et al. 2019). They also developed a semi-rolled leaf genotype derived from wild-type and mutant rolled leaves, which increased panicle number, grain number per panicle, and yield per plant under drought stress (Liao et al. 2019). Based on this experiment, we assume that both SRL1 and SRL2 genes contribute to developing unrolled leaf genotypes under drought stress, which help enhance the photosynthesis process.
The drought and salt tolerance (DST) gene encodes a zinc-finger transcription factor that improves leaf width and stomatal closure (by modulation of H2O2 homeostasis) and reduces stomatal density in rice. Knockout of OsDST gene in rice via CRISPR/Cas9 exhibited enhanced flag leaf growth, leaf water retention, and reduced stomatal density under drought stress (Kumar et al. 2020). Therefore, the OsDST gene might play a key role in developing new drought-tolerant Indica rice cultivars in the future. Enhanced response to abscisic acid (ERA) encodes the β-subunit of the farnesyltransferase gene, which regulates abscisic acid signaling and the dehydration response. CRISPR/Cas9 system was used to develop the rice mutant lines to identify the role of the OsERA1 gene under drought stress (Ogata et al. 2020). Knockout of the OsERA1 gene enhanced stomatal conductance under drought stress. This result revealed that the knockout of the OsERA1 gene enhanced drought tolerance in rice. The abscisic acid receptor of the pyrabactin resistance like (PYL) gene regulates plant growth and development under abiotic stresses. Usman et al. (2020) used CRISPR/Cas9 system to develop Ospyl9 mutants in rice to elucidate the role of OsPYL9 under drought stress. The mutant plants showed higher plant height, panicle numbers, panicle length, flag leaf length and width, grain number per panicle, grain weight, grain length and width, and yield per plant under drought stress (Table 3). The result of this study may lay a practical foundation for the development of drought-tolerant and high-yielding rice cultivars in the future. No apical meristem (NAC) genes are involved in plant growth and development under biotic and abiotic stresses. When OsNAC006 was knockout via CRISPR/Cas9 in rice, the mutants enhanced drought sensitivity in rice. MicroRNAs (miRNAs) are involved in various biological processes associated with plant growth, development, and abiotic stress responses. The OsmiR535 modulates plant height, panicle architecture, and grain length under abiotic stresses in rice. Under drought stress, OsmiR535’s function was identified in rice through CRISPR/Cas9 system. The knockout of the OsmiR535 gene significantly increased the survival rate of seedlings under drought stress (Table 3). The epidermal patterning factor-like 9 (OsEPFL9) gene is a developmental gene that helps regulate the leaf’s stomatal density. Knockout of OsEPFL9 gene in rice by CRISPR/Cas12a system reduced stomatal count and increased water use efficiency under drought stress (Yin et al. 2019).
The CRISPR/Cas9 system in maize has exploited the role of only one gene under drought stress. Auxin-regulated gene involved in organ size (ARGOS) members are negative regulators of ethylene responses, regulating ethylene signal transduction and enhancing drought tolerance. Shi et al. (2017) used the HDR pathway to insert the maize native GOS2 promoter into the 5’ untranslated region of the ARGOS8 gene, which helped to develop two mutants (ARGOS8-v1 and ARGOS8-v2). These two mutants were used to develop hybrids, and their growth and yield were evaluated in multi-location fields. The hybrid plants increased the growth and yield of maize under drought stress compared to the wild type (Shi et al. 2017). This study reveals that the CRISPR/Cas system improves maize growth under drought stress and may lead to the development of new cultivars. However, more research must be done to dissect the roles of other key genes responsible for drought tolerance to help develop better maize varieties conferring tolerance to multiple stresses. The CRISPR/Cas system has not been used in other cereals to identify the role of drought-responsive genes. Among the abiotic stresses, many genes and transcription factors have been identified and functionally characterized in most cereals under drought stress. Therefore, targeting other genes and transcription factors by CRISPR/Cas variants in other cereals could help improve the growth and yield of all cereals under drought stress.
Salinity stress tolerance
As in drought stress, the functions of some salt-responsive genes in rice were identified by CRISPR/Cas system. The two-component response regulator (RR) gene encodes the B-type response regulator transcription factor and is involved in cytokinin signal transduction and metabolism. The OsRR22 gene loss of function has significantly increased salt tolerance. Knockout of OsRR22 gene through CRISPR/Cas9 system in rice enhanced the salinity tolerance (Zhang et al. 2019a). The transgenic plants showed significant tolerance to salinity at the seedling stage. Another group used the CRISPR/Cas9 system to characterize the roles of OsRR9 and 10 genes under salt stress (Wang et al. 2019b). They have generated double mutants of osrr9 and 10 genes through multiplexed CRISPR/Cas9 system, which enhanced salinity tolerance. The same study further validated the two potassium transporter (OsHKT1;1 and 2;1) genes in rice leaves under salt stress. Both genes were highly expressed in the leaf of transgenic plants. This result revealed that OsRR9 and 10 genes enhanced tolerance to salinity stress by regulating HKT family transporters. This study indicates that we can develop a new rice cultivar using these genes to grow better under salt stress and potassium deficiency soils. Zhang et al. (2019b) applied CRISPR/Cas9 system to produce a knockout of overly tolerant to salt1 (OsOTS1) (Small Ubiquitin‐like Modifier (SUMO) protease family member) gene in rice. The transgenic lines enhanced salinity tolerance and biomass of shoot and root, indicating that the OsOTS1 gene plays an essential role in salt stress tolerance in rice. These examples demonstrate that CRISPR/Cas system could effectively identify the role of genes that can respond to salt stress in other cereals.
Heat and cold stress tolerance
High temperature affects chloroplast activity, reducing photosynthesis and yields in plants. Qiu et al. (2018) characterized the role of heat sensitive albino1 (HSA1) gene in rice through CRISPR/Cas9. Knockout of HSA1 gene in rice significantly delayed chloroplast activity at high temperatures compared to wild type (Qiu et al. 2018). These results implied that the HSA1 gene plays an essential role in chloroplast activity under heat stress. Malzahn et al. (2019) targeted two genes (rice outermost cell-specific (OsROC5) and dense and erect panicle 1 (OsDEP1)) by CRISPR/Cas12a system in rice and maize, enhancing their growth under higher temperature conditions. Cold stress affects seed germination, seedling growth, tillering, and yield (due to delayed heading and pollen sterility). Developing cold-stress-tolerant cereals by CRISPR/Cas system could help to overcome this problem. Knockout of the annexin 3 (OsAnn3) gene in rice enhanced rice growth under cold stress (Shen et al. 2017). In another report, the role of proline-rich protein 1 (OsPRP1) was identified in rice through CRISPR/Cas9 under cold stress. Knockout of OsPRP1 in mutant rice reduced survival rate, fresh and dry weight of shoot and root, proline, abscisic, and ascorbic acid contents in leaf and root tissues, and activities of antioxidant enzymes under cold stress compared to wild type (Nawaz et al. 2019). This study reveals that the OSPRP1 enhances rice growth under cold stress. Three genes, such as OsPIN5b (a panicle length gene), OsGS3 (a grain size gene), and OsMYB30 (a cold tolerance gene), were targeted by CRISPR/Cas9 system in rice enhanced panicle length and grain size under cold stress (Zeng et al. 2020a). Among CRISPR/Cas variants, CRISPR/Cas12b system has been reported to develop temperature-tolerant cereals. Hence, targeting heat stress-responsive genes by CRISPR/Cas12 system may enable the development of heat tolerance cereals over other CRISPR/Cas variants.
Nutrients (macro- and micro-) stress tolerance
Both macro- and micronutrients play an essential role in plants’ physiological and biochemical processes. Various macro and micronutrients transporters such as nitrate transporter (NRT), phosphate transporters (PHT), potassium transporters (K+ transporters), sulfate transporters (SULTR), ammonium transporters (AMT), iron transporters (IRT), copper transporters (COPT), zinc regulated and iron-regulated transport like proteins (ZIP), copper transporter (COPT), iron-regulated transporter (IRT), cation diffusion facilitator (CDF), ATP-binding cassettes (ABC), heavy metal ATPase (HMA), molybdate transporter type 1 (MOT1), natural resistance-associated macrophage protein (NRAMP) and yellow stripe-like proteins (YSL) have been functionally characterized in various cereals (Maharajan et al. 2022; Krishna et al. 2020, 2022b; Roch et al. 2019). The NRT1 family gene (OsNRT1;1b) has been targeted by base editing (Li et al. 2018; Lu and Zhu 2017). The role of other nutrient transporters has not yet been identified in any cereal through any genome editing tool. More recently, two review articles have discussed improving the plant nutrient transporters with the help of the CRISPR/Cas system (Ceasar et al. 2022; Sathee et al. 2022). The role of antioxidant protein 1 (ATX1) has been identified in rice through CRISPR/Cas9. Knockout of the OsATX1 gene in rice increased the accumulation of Cu in root and older leaves (Zhang et al. 2018b). These results indicate that the OsATX1 plays a key role in facilitating the translocation of Cu from root to shoot and remobilizing Cu from old leaves to developing tissues. Some micronutrients (Cd, Cr, Pb, Al, and Hg) are not involved in any physiological and biochemical processes in plants; therefore, they are considered non-essential micronutrients. These non-essential micronutrients are very toxic for plants, even at very low concentrations. The presence of non-essential micronutrients causes common toxic effects on plants, such as reduced biomass, growth and photosynthesis process, induced chlorosis, altered water balance, and nutrient assimilation, which ultimately cause plant death. A study revealed that knockout of OsNramp5 gene using the CRISPR/Cas9 system in rice reduced Cd accumulations in shoots and roots of mutant plants more than in wild-type plants without affecting yield under Cd contaminated soil (Tang et al. 2017). Knockout of arsenate (As) responsive MYB1 (OsARM1) in rice enhanced plant height and root length under high (40 µM) As stress (Wang et al. 2017). In the same study, the knockout rice plants increased the As concentrations in shoot and root tissues compared to wild-type plants under low (2 µM) As conditions. Based on these results, OsARM1 gene may be involved in the translocation of As from root to shoot (under low As condition) and enhanced plant growth (under high As condition).
Conclusion and future direction
The world population is expected to reach around 10 billion by 2050, and the demand for food will increase. The ever-increasing global population and food demand forced agriculturists, industrialists, soil chemists, scientists, plant breeders, government officials, and farmers to promote integrated and sustainable crop production. The growth and yield of all cereals are severely affected by both biotic and abiotic stresses. Plant breeding and functional genomic fields have played a major role in developing new varieties tolerant to growing under biotic and abiotic stresses. Both these technologies take a long time to develop a new crop. The recently developed CRISPR/Cas technology will enable us to create new crops in a short period due to their simplicity, versatility, accuracy, and sophistication. In cereals (mainly rice), the role of many biotic and abiotic stress-responsive genes and transcription factors have been characterized by CRISPR/Cas system. The role of genes and transcription factors that respond to biotic and abiotic stresses have not yet been identified in other cereals by CRISPR /Cas system. Many other CRISPR/Cas-mediated genome-editing tools such as CRISPRi, CRISPRa, base editors, epigenetic engineering, chromatin imaging, and prime editors have been discovered and used in crop improvement programs. All these tools are becoming popular among plants because of their accuracy and robustness. Identifying the role of each biotic and abiotic stress-responsive gene through CRISPR/Cas-mediated tool may help improve cereals’ growth and yield under biotic and abiotic stresses. Plants developed by CRISPR-based genome-editing tools may also become safe and transgenic-free since marker-free crops could be developed by introducing CRISPR reagents (Cas and gRNA) without conventional transformation and selection under antibiotics. This will also help overcome the hurdles scientists face in commercializing biotech crops. Researchers may focus on these lines to develop new genome-editing methods to develop transgenic crops that are generally accepted by all. Hence, the government should play its part in future policies related to the CRISPR/Cas system and not consider CRISPR-edited transgenic or genetically modified plants for large-scale cultivation. This will help to improve the saline and drought tolerance in cereals strengthening food security.
Author contribution statement
TM and SAC—conceptualized the manuscript. TM, TPAK, KR, and SAC—wrote the manuscript. SAC and MR—assisted, edited, and updated the manuscript.
Data availability
Data sharing does not apply to this article as no datasets were generated or analyzed during the current study.
Abbreviations
- CRISPR/Cas:
-
Clustered, regularly inter-spaced, short palindromic repeats (CRISPR)/CRISPR-associated protein
- CRISPRa:
-
CRISPR activation
- CRISPRi:
-
CRISPR interference
- dCas9:
-
Dead Cas9
- TALENs:
-
Transcription activator-like effector nucleases
- ZFNs:
-
Zinc-finger nucleases
References
Abbas G, Saqib M, Rafique Q, Rahman AU, Akhtar J, Haq MAU, Nasim M (2013) Effect of salinity on grain yield and grain quality of wheat (Triticum aestivum L.). Pak J Bot 50:185–189
Abdelrahman M, Wei Z, Rohila JS, Zhao K (2021) Multiplex genome-editing technologies for revolutionizing plant biology and crop improvement. Front Plant Sci 12:721203
Aghnoum R, Bvindi C, Menet G, Dhoop B, Maciel JLN, Niks RE (2019) Host/nonhost status and genetics of resistance in barley against three pathotypes of Magnaporthe blast fungi. Euphy 215:1–19
Afzal M, Ahmad A, Ahmad AH (2012) Effect of nitrogen on growth and yield of sorghum forage (Sorghum bicolor (L.) Moench cv.) under three cuttings system. Cerc Agron Mol 45:57–64
Alghabari F, Ihsan MZ (2018) Effects of drought stress on growth, grain filling duration, yield and quality attributes of barley (Hordeum vulgare L.). Bangladesh J Bot 47:421–428
Ali A, Basra SMA, Ahmad R, Wahid A (2009) Optimizing silicon application to improve salinity tolerance in wheat. Soil Environ 28:136–144
Altpeter F, Springer NM, Bartley LE, Blechl AE, Brutnell TP, Citovsky V, Stewart CN Jr (2016) Advancing crop transformation in the era of genome editing. Plant Cell 28:1510–1520
Aref F, Rad HE (2012) Physiological characterization of rice under salinity stress during vegetative and reproductive stages. Ind J Sci Technol 5:2578–2586
Asmamaw M, Zawdie B (2021) Mechanism and applications of CRISPR/Cas-9-mediated genome editing. Biologics Tar Ther 15:353
Awika JM (2011) Major cereal grains production and use around the world. In: Joseph MA, Scott B (Eds), Advances in cereal science: implications to food processing and health promotion. ACS Symposium Series, American Chemical Society: Washington, DC, pp 1–13
Azameti MK, Dauda WP (2021) Base editing in plants: applications, challenges, and future prospects. Front Plant Sci 12:1531
Bagci SA, Ekiz H, Yilmaz A, Cakmak I (2007) Effects of zinc deficiency and drought on grain yield of field-grown wheat cultivars in Central Anatolia. J Agron Crop Sci 193:198–206
Bandara YMAY, Weerasooriya DK, Tesso TT, Prasad PVV, Little CR (2017) Stalk rot fungi affect grain sorghum yield components in an inoculation stage-specific manner. Crop Prot 94:97–105
Bharat SS, Li S, Li J, Yan L, Xia L (2020) Base editing in plants: current status and challenges. Crop J 8:384–395
Butt H, Rao GS, Sedeek K, Aman R, Kamel R, Mahfouz M (2020) Engineering herbicide resistance via prime editing in rice. Plant Biotech J 18:2370
Carroll D (2011) Genome engineering with zinc-finger nucleases. Genetics 188:773–782
Ceasar SA, Hodge A, Baker A, Baldwin SA (2014) Phosphate concentration and arbuscular mycorrhizal colonisation influence the growth, yield and expression of twelve PHT1 family phosphate transporters in foxtail millet (Setaria italica). PLoS ONE 9:e108459
Ceasar SA, Rajan V, Prykhozhij SV, Berman JN, Ignacimuthu S (2016) Insert, remove or replace: A highly advanced genome editing system using CRISPR/Cas9. Bioch Biophy Acta (BBA)-Mol Cell Res 1863:2333–2344
Ceasar SA, Maharajan T, Hillary E, Krishna TPA (2022) Insights to improve the plant nutrient transport by CRISPR/Cas system. Biotechnol Adv 59:107963
Chen JS, Dagdas YS, Kleinstiver BP, Welch MM, Sousa AA, Harrington LB, Doudna JA (2017) Enhanced proofreading governs CRISPR–Cas9 targeting accuracy. Nature 550:407–410
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Cruz CD, Valent B (2017) Wheat blast disease: danger on the move. Trop Plant Pathol 42:210–222
David RHA, Ramakrishnan M, Maharajan T, BarathiKannan K, Babu GA, Daniel MA, Ignacimuthu S (2021) Mining QTL and genes for root traits and biochemical parameters under vegetative drought in South Indian genotypes of finger millet (Eleusine coracana (L.) Gaertn) by association mapping and in silico comparative genomics. Biocatal Agric Biotechnol 32:101935
Dong B, Zheng X, Liu H, Able JA, Yang H, Zhao H, Liu M (2017) Effects of drought stress on pollen sterility, grain yield, abscisic acid and protective enzymes in two winter wheat cultivars. Front Plant Sci 8:1008
Edwards MC, Fetch TG Jr, Schwarz PB, Steffenson BJ (2001) Effect of Barley yellow dwarf virus infection on yield and malting quality of barley. Plant Dis 85:202–207
Feng Z, Zhang B, Ding W, Liu X, Yang DL, Wei P, Zhu JK (2013) Efficient genome editing in plants using a CRISPR/Cas system. Cell Res 23:1229–1232
Fiyaz RA, Shivani D, Chaithanya K, Mounika K, Chiranjeevi M, Laha GS, Sundaram RM (2022) Genetic improvement of rice for bacterial blight resistance: present status and future prospects. Rice Sci 29:118–132
Gao L, Cox DB, Yan WX, Manteiga JC, Schneider MW, Yamano T, Zhang F (2017) Engineered Cpf1 variants with altered PAM specificities. Nat Biotech 35:789–792
Gashaw G, Alemu T, Tesfaye K (2014) Morphological, physiological and biochemical studies on Pyricularia grisea isolates causing blast disease on finger millet in Ethiopia. J Appl Biosci 74:6059–6071
Gerona MEB, Deocampo MP, Egdane JA, Ismail AM, Dionisio Sese ML (2019) Physiological responses of contrasting rice genotypes to salt stress at reproductive stage. Rice Sci 26:207–219
Gupta A, Christensen RG, Rayla AL, Lakshmanan A, Stormo GD, Wolfe SA (2012) An optimized two-finger archive for ZFN-mediated gene targeting. Nat Methods 9:588–590
Haroon M, Wang X, Afzal R, Zafar MM, Idrees F, Batool M, Imran M (2022) Novel plant breeding techniques shake hands with cereals to increase production. Plants 11:1052
Hasan A, Hafiz HR, Siddiqui N, Khatun M, Islam R, Mamun AA (2015) Evaluation of wheat genotypes for salt tolerance based on some physiological traits. J Crop Sci Biotech 18:333–340
Hillary VE, Ceasar SA (2019) Application of CRISPR/Cas9 genome editing system in cereal crops. Open Biotechnol J 13:173–179
Hillary VE, Ceasar SA (2022) Prime editing in plants and mammalian cells: mechanism, achievements, limitations, and future prospects. BioEssays 44:2200032
Hong W, Jin JY (2007) Effects of zinc deficiency and drought on plant growth and metabolism of reactive oxygen species in maize (Zea mays L). Agri Sci China 6:988–995
Hu JH, Miller SM, Geurts MH, Tang W, Chen L, Sun N, Liu DR (2018) Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556:57–63
Hua K, Tao X, Yuan F, Wang D, Zhu JK (2018) Precise A·T to G·C base editing in the rice genome. Mol Plant 11:627–630
Hua K, Tao X, Han P, Wang R, Zhu JK (2019) Genome engineering in rice using Cas9 variants that recognize NG PAM sequences. Mol Plant 12:1003–1014
Hua K, Jiang Y, Tao X, Zhu JK (2020a) Precision genome engineering in rice using prime editing system. Plant Biotechnol J 18:2167–2169
Hua K, Tao X, Liang W, Zhang Z, Gou R, Zhu JK (2020b) Simplified adenine base editors improve adenine base editing efficiency in rice. Plant Biotechnol J 18:770–778
Hussain S, Zhang JH, Zhong C, Zhu LF, Cao XC, Yu SM, Jin QY (2017) Effects of salt stress on rice growth, development characteristics, and the regulating ways: a review. J Int Agric 16:2357–2374
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A (1987) Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol 169:5429–5433
Izadi MH, Rabbani J, Emam Y, Pessarakli M, Tahmasebi A (2014) Effects of salinity stress on physiological performance of various wheat and barley cultivars. J Plant Nutr 37:520–531
Jiang YY, Chai YP, Lu MH, Han XL, Lin Q, Zhang Y, Chen QJ (2020) Prime editing efficiently generates W542L and S621I double mutations in two ALS genes in maize. Genome Biol 21:1–10
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Kihoro J, Bosco NJ, Murage H, Ateka E, Makihara D (2013) Investigating the impact of rice blast disease on the livelihood of the local farmers in greater Mwea region of Kenya. Springerplus 2:1–13
Krishna TPA, Ceasar SA, Maharajan T, Ramakrishnan M, Duraipandiyan V, Al-Dhabi NA, Ignacimuthu S (2017) Improving the zinc-use efficiency in plants: a review. SABRAO J Breed Genet 49:211–230
Krishna TPA, Maharajan T, Roch GV, Ignacimuthu S, Ceasar SA (2020) Structure, function, regulation and phylogenetic relationship of ZIP family transporters of plants. Front Plant Sci 11:662
Krishna TPA, Maharajan T, Ceasar SA (2022a) Application of CRISPR/Cas9 genome editing system to reduce the pre-and post-harvest yield losses in cereals. Open Biotech J 16:1–9
Krishna TPA, Maharajan T, Ceasar SA (2022b) The role of membrane transporters in the biofortification of zinc and iron in plants. Biol Trace Elem Res. https://doi.org/10.1007/s12011-022-03159-w
Kuang Y, Li S, Ren B, Yan F, Spetz C, Li X, Zhou X, Zhou H (2020) Base-editing-mediated artificial evolution of OsALS1 in planta to develop novel herbicide-tolerant rice germplasms. Mol Plant 13:565–572
Kumar VS, Verma RK, Yadav SK, Yadav P, Watts A, Rao MV, Chinnusamy V (2020) CRISPR-Cas9 mediated genome editing of drought and salt tolerance (OsDST) gene in indica mega rice cultivar MTU1010. Phy Mol Biol Plants 26:1099–1110
Lee JH, Muhsin M, Atienza GA, Kwak DY, Kim SM, De Leon TB, Choi IR (2010) Single nucleotide polymorphisms in a gene for translation initiation factor (eIF4G) of rice (Oryza sativa) associated with resistance to Rice tungro spherical virus. Mol Plant Microbe Interact 23:29–38
Lee JK, Jeong E, Lee J, Jung M, Shin E, Kim YH, Kim JS (2018) Directed evolution of CRISPR-Cas9 to increase its specificity. Nat Commun 9:1–10
Lee J, Jung MH, Jeong E, Lee JK (2019) Using Sniper-Cas9 to minimize off-target effects of CRISPR-Cas9 without the loss of on-target activity via directed evolution. J vis Exp 144:e59202
Li J, Sun Y, Du J, Zhao Y, Xia L (2017) Generation of targeted point mutations in rice by a modified CRISPR/Cas9 system. Mol Plant 10:526–529
Li C, Zong Y, Wang Y, Jin S, Zhang D, Song Q, Gao C (2018) Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol 19:1–9
Li C, Zhang R, Meng X, Chen S, Zong Y, Lu C, Qiu JL, Chen YH, Li J, Gao C (2020a) Targeted, random mutagenesis of plant genes with dual cytosine and adenine base editors. Nat Biotechnol 38:875–888
Li H, Li J, Chen J, Yan L, Xia L (2020b) Precise modifications of both exogenous and endogenous genes in rice by prime editing. Mol Plant 13:671–674
Li J, Xu R, Qin R, Liu X, Kong F, Wei P (2021) Genome editing mediated by SpCas9 variants with broad non-canonical PAM compatibility in plants. Mol Plant 14:352–360
Li Z, Qiu Z, Ge H, Du C (2022) Long-term dynamic of cold stress during heading and flowering stage and its effects on rice growth in China. Atmosphere 13:103
Lin Q, Zong Y, Xue C, Wang S, Jin S, Zhu Z, Wang Y, Anzalone AV, Raguram A, Doman JL, Liu DR, Gao C (2020) Prime genome editing in rice and wheat. Nat Biotechnol 38:582–585
Liao S, Qin X, Luo L, Han Y, Wang X, Usman B, Li R (2019) CRISPR/Cas9-induced mutagenesis of semi-rolled Leaf 1, 2 confers curled leaf phenotype and drought tolerance by influencing protein expression patterns and ROS scavenging in Rice (Oryza sativa L.). Agronomy 9:728
Liu C, Zhang L, Liu H, Cheng K (2017) Delivery strategies of the CRISPR-Cas9 gene-editing system for therapeutic applications. J Control Rel 266:17–26
Lou D, Wang H, Liang G, Yu D (2017) OsSAPK2 confers abscisic acid sensitivity and tolerance to drought stress in rice. Front Plant Sci 8:993
Lu Y, Zhu JK (2017) Precise editing of a target base in the rice genome using a modified CRISPR/Cas9 system. Mol Plant 10:523–525
Ma J, Chen J, Wang M, Ren Y, Wang S, Lei C, Cheng Z (2018) Disruption of OsSEC3A increases the content of salicylic acid and induces plant defense responses in rice. J Exp Bot 69:1051–1064
Macovei A, Sevilla NR, Cantos C, Jonson GB, Slamet-Loedin I, Cermak T, Chadha-Mohanty P (2018) Novel alleles of rice eIF4G generated by CRISPR/Cas9-targeted mutagenesis confer resistance to Rice tungro spherical virus. Plant Biotech J 16:1918–1927
Maharajan T, Ceasar SA, Krishna TPA, Ramakrishnan M, Duraipandiyan V, Naif Abdulla AD, Ignacimuthu S (2018) Utilization of molecular markers for improving the phosphorus efficiency in crop plants. Plant Breed 137:10–26
Maharajan T, Ceasar SA, Krishna TPA, Ignacimuthu S (2019) Phosphate supply influenced the growth, yield and expression of PHT1 family phosphate transporters in seven millets. Planta 250:1433–1448
Maharajan T, Ceasar SA, Krishna TPA, Ignacimuthu S (2021a) Finger millet [Eleusine coracana (L.) Gaertn]: an orphan crop with a potential to alleviate the calcium deficiency in the semi-arid tropics of Asia and Africa. Front Sustain Food Syst 5:684447
Maharajan T, Roch GV, Ceasar SA (2021b) Recent advancements of molecular breeding and functional genomics for improving nitrogen-phosphorus-and potassium-use efficiencies in wheat. In: Hossain MA, Alam M, Seneweera S, Sujay R, Henry R (eds) Molecular breeding in wheat, maize and sorghum: Strategies for improving abiotic stress tolerance and yield. CAB International, Wallingford, pp 170–196
Maharajan T, Krishna TP, Kiriyanthan RMK, Ignacimuthu S, Ceasar SA (2021c) Improving abiotic stress tolerance in sorghum: focus on the nutrient transporters and marker-assisted breeding. Planta 254:1–16
Maharajan T, Ceasar SA, Krishna TPA (2022) Finger Millet (Eleusine coracana (L.) Gaertn): nutritional Importance and Nutrient Transporters. Crit Rev Plant Sci 41:1–31
Makarova KS, Koonin EV (2015) Annotation and classification of CRISPR-Cas systems. In: Lundgren M, Charpentier E, Fineran P (eds) CRISPR, Methods in Molecular Biology. Humana Press, New York, pp 47–75
Makarova KS, Wolf YI, Iranzo J, Shmakov SA, Alkhnbashi OS, Brouns SJ, Koonin EV (2020) Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat Rev Microbiol 18:67–83
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Sci 339:823–826
Malzahn AA, Tang X, Lee K, Ren Q, Sretenovic S, Zhang Y, Qi Y (2019) Application of CRISPR-Cas12a temperature sensitivity for improved genome editing in rice, maize, and Arabidopsis. BMC Biol 17:1–14
Matres JM, Hilscher J, Datta A, Armario Najera V, Baysal C, He W, Slamet-Loedin IH (2021) Genome editing in cereal crops: an overview. Trans Res 30:461–498
Molla KA, Yang Y (2019) CRISPR/Cas-mediated base editing: technical considerations and practical applications. Trends Biotechnol 37:1121–1142
Molla KA, Shih J, Yang Y (2020) Single-nucleotide editing for zebra3 and wsl5 henotypes in rice using CRISPR/Cas9-mediated adenine base editors. aBIOTECH 1:106–118
Moradpour M, Abdulah SNA (2020) CRISPR/dC as9 platforms in plants: strategies and applications beyond genome editing. Plant Biotech J 18:32–44
Muthukumararaja TM, Sriramachandrasekharan MV (2012) Effect of zinc on yield, zinc nutrition and zinc use efficiency of lowland rice. J Agri Tech 8:551–561
Nawaz G, Han Y, Usman B, Liu F, Qin B, Li R (2019) Knockout of OsPRP1, a gene encoding proline-rich protein, confers enhanced cold sensitivity in rice (Oryza sativa L.) at the seedling stage. 3 Biotech 9:1–18
Nilsson L, Muller R, Nielsen TH (2010) Dissecting the plant transcriptome and the regulatory responses to phosphate deprivation. Physiol Plant 139:129–143
Noh TH, Lee DK, Park JC, Shim HK, Choi MY, Kang MH, Kim JD (2007) Effects of bacterial leaf blight occurrence on rice yield and grain quality in different rice growth stage. Res Plant Dis 13:20–23
Ogata T, Ishizaki T, Fujita M, Fujita Y (2020) CRISPR/Cas9-targeted mutagenesis of OsERA1 confers enhanced responses to abscisic acid and drought stress and increased primary root growth under nonstressed conditions in rice. PLoS ONE 15:e0243376
Oliva R, Ji C, Atienza-Grande G, Huguet-Tapia JC, Perez-Quintero A, Li T, Yang B (2019) Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat Biotech 37:1344–1350
Perry KL, Kolb FL, Sammons B, Lawson C, Cisar G, Ohm H (2000) Yield effects of barley yellow dwarf virus in soft red winter wheat. Phytopathol 90:1043–1048
Plenet D, Etchebest S, Mollier A, Pellerin S (2000) Growth analysis of maize field crops under phosphorus deficiency. Plant Soil 223:119–132
Pruthi R, Puram VRR, Ontoy J, Subudhi PK (2022) Genetics of yield component traits under salt stress at flowering stage and selection of salt tolerant pre-breeding lines for rice improvement. Genetica 150(5):273–288
Pulecio J, Verma N, Mejia Ramírez E, Huangfu D, Raya A (2017) CRISPR/Cas9-based engineering of the epigenome. Cell Stem Cell 21:431–447
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA (2013) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152:1173–1183
Qiu Z, Kang S, He L, Zhao J, Zhang S, Hu J, Zhu L (2018) The newly identified heat-stress sensitive albino 1 gene affects chloroplast development in rice. Plant Sci 267:168–179
Queiroz MS, Oliveira CE, Steiner F, Zuffo AM, Zoz T, Vendruscolo EP, Menis FT (2019) Drought stresses on seed germination and early growth of maize and sorghum. J Agri Sci 11:310–318
Rahman MS, Haque MA, Islam MT (2015) Salinity affects flag leaf chlorophyll and yield attributes of rice genotypes. J Biosci Agri Res 4:80–85
Rakkammal K, Maharajan T, Ceasar SA, Ramesh M (2022) Biostimulants and their role in improving plant growth under drought and salinity. Cereal Res Commun. https://doi.org/10.1007/s42976-022-00299-6
Ren B, Liu L, Li S, Kuang Y, Wang J, Zhang D, Zhou X, Lin H, Zhou H (2019) Cas9-NG greatly expands the targeting scope of the genome-editing toolkit by recognizing NG and other atypical PAMs in rice. Mol Plant 12:1015–1026
Ren Q, Sretenovic S, Liu S, Tang X, Huang L, He Y, Liu L, Guo Y, Zhong Z, Liu G, Cheng Y, Zheng X, Pan C, Yin D, Zhang Y, Li W, Qi L, Li C, Qi Y, Zhang Y (2021) PAM-less plant genome editing using a CRISPR-SpRY toolbox. Nat Plants 7:25–33
Roch GV, Maharajan T, Ceasar SA, Ignacimuthu S (2019) The role of PHT1 family transporters in the acquisition and redistribution of phosphorus in plants. Crit Rev Plant Sci 38:171–198
Roch GV, Maharajan T, Krishna TP, Ignacimuthu S, Ceasar SA (2020) Expression of PHT1 family transporter genes contributes for low phosphate stress tolerance in foxtail millet (Setaria italica) genotypes. Planta 252:1–9
Rozewicz M, Wyzińska M, Grabiński J (2021) The most important fungal diseases of cereals—problems and possible solutions. Agronomy 11:714
Saddiq MS, Iqbal S, Hafeez MB, Ibrahim AM, Raza A, Fatima EM, Ciarmiello LF (2021) Effect of salinity stress on physiological changes in winter and spring wheat. Agronomy 11:1193
Samarah NH (2005) Effects of drought stress on growth and yield of barley. Agron Sustain Dev 25:145–149
Sathee L, Barman D, Nagar S, Tripathi S, Jha SK, Chinnusamy V (2022) Genome editing targets for improving nutrient use efficiency and nutrient stress adaptation. Front Genet. https://doi.org/10.3389/fgene.2022.900897
Sallam A, Alqudah AM, Dawood MF, Baenziger PS, Borner A (2019) Drought stress tolerance in wheat and barley: advances in physiology, breeding and genetics research. Int J Mol Sci 20:3137
Sarwar MH, Sarwar MF, Sarwar M, Qadri NA, Moghal S (2013) The importance of cereals (Poaceae: Gramineae) nutrition in human health: a review. J Cereals Oilseeds 4:32–35
Sharma R, Girish AG, Upadhyaya HD, Humayun P, Babu TK, Rao VP, Thakur R (2014) Identification of blast resistance in a core collection of foxtail millet germplasm. Plant Dis 98:519–524
Shafi M, Bakht JEHAN, Jalal FAZAL, Khan MA, Khattak SG (2011) Effect of nitrogen application on yield and yield components of barley (Hordeum vulgare L.). Pak J Bot 43:1471–1475
Shimatani Z, Kashojiya S, Takayama M, Terada R, Arazoe T, Ishii H, Teramura H, Yamamoto T, Komatsu H, Miura K, Ezura H, Nishida K, Ariizumi T, Kondo A (2017) Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat Biotechnol 35:441–443
Shen C, Que Z, Xia Y, Tang N, Li D, He R, Cao M (2017) Knock out of the annexin gene OsAnn3 via CRISPR/Cas9-mediated genome editing decreased cold tolerance in rice. J Plant Biol 60:539–547
Shi J, Gao H, Wang H, Lafitte HR, Archibald RL, Yang M, Habben JE (2017) ARGOS 8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol J 15:207–216
Shrivastava P, Kumar R (2015) Soil salinity: a serious environmental issue and plant growth promoting bacteria as one of the tools for its alleviation. Saudi J Biol Sci 22:123–131
Singh CM, Binod K, Suhel M, Kunj C (2012) Effect of drought stress in rice: a review on morphological and physiological characteristics. Trend Biosci 5:261–265
Sumit S, Sinha D, Kumari A (2020) An overview of bacterial leaf blight disease of rice and different strategies for its management. Int J Curr Microbiol App Sci 9:2250–2265
Tambong JT (2022) Bacterial pathogens of wheat: symptoms, distribution, identification, and taxonomy. In: Mahmood RA (ed) Wheat. Intech Open, London, pp 1–22
Tang L, Mao B, Li Y, Lv Q, Zhang L, Chen C, Zhao B (2017) Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield. Sci Rep 7:1–12
Ticconi CA, Lucero RD, Sakhonwasee S, Adamson AW, Creff A, Nussaume L, Abel S (2009) ER-resident proteins PDR2 and LPR1 mediate the developmental response of root meristems to phosphate availability. Proc Nat Acad Sci 106:14174–14179
Usman B, Nawaz G, Zhao N, Liao S, Liu Y, Li R (2020) Precise editing of the ospyl9 gene by rna-guided cas9 nuclease confers enhanced drought tolerance and grain yield in rice (Oryza sativa L.) by regulating circadian rhythm and abiotic stress responsive proteins. Int J Mol Sci 21:7854
Walton RT, Christie KA, Whittaker MN, Kleinstiver BP (2020) Unconstrained genome targeting with near-PAMless engineered CRISPR-Cas9 variants. Science 368:290–296
Wang X, Du G, Wang X, Meng Y, Li Y, Wu P, Yi K (2010) The function of LPR1 is controlled by an element in the promoter and is independent of SUMO E3 Ligase SIZ1 in response to low Pi stress in Arabidopsis thaliana. Plant Cell Physiol 51:380–394
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotech 32:947–951
Wang F, Wang C, Liu P, Lei C, Hao W, Gao Y, Zhao K (2016) Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS ONE 11:e0154027
Wang FZ, Chen MX, Yu LJ, Xie LJ, Yuan LB, Qi H, Chen QF (2017) OsARM1, an R2R3 MYB transcription factor, is involved in regulation of the response to arsenic stress in rice. Front Plant Sci 8:1868
Wang B, Liu C, Zhang D, He C, Zhang J, Li Z (2019a) Effects of maize organ-specific drought stress response on yields from transcriptome analysis. BMC Plant Biol 19:1–19
Wang WC, Lin TC, Kieber J, Tsai YC (2019b) Response regulators 9 and 10 negatively regulate salinity tolerance in rice. Plant Cell Physiol 60:2549–2563
Wang B, Zhong Z, Wang X, Han X, Yu D, Wang C, Zhang Y (2020a) Knockout of the OsNAC006 transcription factor causes drought and heat sensitivity in rice. Int J Mol Sci 21:2288
Wang S, Zong Y, Lin Q, Zhang H, Chai Z, Zhang D, Chen K, Qiu JL, Gao C (2020b) Precise, predictable multi-nucleotide deletions in rice and wheat using APOBEC-Cas9. Nat Biotechnol 38:1460–2146
Wang C, Liu G, Zhang D, Zhang S, Qiu J (2022) Plant prime editing technique: a new genome editing tool for plants. Chinese J Biotech 38:26–33
Xu X, Qi LS (2019) A CRISPR–dCas toolbox for genetic engineering and synthetic biology. J Mol Biol 431:34–47
Xu R, Li J, Liu X, Shan T, Qin R, Wei P (2020a) Development of plant prime-editing systems for precise genome editing. Plant Commun 1:100043
Xu W, Zhang C, Yang Y, Zhao S, Kang G, He X, Song J, Yang J (2020b) Versatile nucleotides substitution in plant using an improved prime editing system. Mol Plant 13:675–678
Xu Z, Kuang Y, Ren B, Yan D, Yan F, Spetz C, Sun W, Wang G, Zhou X, Zhou H (2021) SpRY greatly expands the genome editing scope in rice with highly flexible PAM recognition. Genome Biol 22:6
Yang Y, Xu J, Ge S, Lai L (2021) CRISPR/Cas: advances, limitations, and applications for precision cancer research. Front Med 8:649896
Yin X, Anand A, Quick P, Bandyopadhyay A (2019) Editing a stomatal developmental gene in rice with CRISPR/Cpf1. In: Qi Y (ed) Plant genome editing with CRISPR systems. Humana Press, New York, pp 257–268
Yuan Y, Wu H, Wang N, Li J, Zhao W, Du J, Ling HQ (2008) FIT interacts with AtbHLH38 and AtbHLH39 in regulating iron uptake gene expression for iron homeostasis in Arabidopsis. Cell Res 18:385–397
Yue E, Cao H, Liu B (2020) OsmiR535, a potential genetic editing target for drought and salinity stress tolerance in Oryza sativa. Plants 9:1337
Zafar K, Sedeek KE, Rao GS, Khan MZ, Amin I, Kamel R, Mahfouz MM (2020) Genome editing technologies for rice improvement: progress, prospects, and safety concerns. Front Genome Editing 2:5
Zeng Y, Wen J, Zhao W, Wang Q, Huang W (2020a) Rational improvement of rice yield and cold tolerance by editing the three genes OsPIN5b, GS3, and OsMYB30 with the CRISPR–Cas9 system. Front Plant Sci 10:1663
Zeng D, Li X, Huang J, Li Y, Cai S, Yu W, Li Y, Huang Y, Xie X, Gong Q, Tan J, Zheng Z, Guo M, Liu YG, Zhu Q (2020b) Engineered Cas9 variant tools expand argeting scope of genome and base editing in rice. Plant Biotechnol J 18:1348–1350
Zhang M, Liu B (2017) Identification of a rice metal tolerance protein OsMTP11 as a manganese transporter. PLoS ONE 12:e0174987
Zhang Y, Bai Y, Wu G, Zou S, Chen Y, Gao C, Tang D (2017) Simultaneous modification of three homoeologs of Ta EDR 1 by genome editing enhances powdery mildew resistance in wheat. Plant J 91:714–724
Zhang X, Lei L, Lai J, Zhao H, Song W (2018a) Effects of drought stress and water recovery on physiological responses and gene expression in maize seedlings. BMC Plant Biol 18:1–16
Zhang Y, Chen K, Zhao FJ, Sun C, Jin C, Shi Y, Lian X (2018b) OsATX1 interacts with heavy metal P1B-type ATPases and affects copper transport and distribution. Plant Physiol 178:329–344
Zhang A, Liu Y, Wang F, Li T, Chen Z, Kong D, Luo L (2019a) Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol Breed 39:1–10
Zhang R, Liu J, Chai Z, Chen S, Bai Y, Zong Y, Chen K, Li J, Jiang L, Gao C (2019b) Generation of herbicide tolerance traits and a new selectable marker in wheat using base editing. Nat Plants 5:480–485
Zhang C, Srivastava AK, Sadanandom A (2019c) Targeted mutagenesis of the SUMO protease, Overly Tolerant to Salt1 in rice through CRISPR/Cas9-mediated genome editing reveals a major role of this SUMO protease in salt tolerance. BioRxiv. https://doi.org/10.1101/555706v1
Zhang R, Chen S, Meng X, Chai Z, Wang D, Yuan Y, Chen K, Jiang L, Li J, Gao C (2020) Generating broad-spectrum tolerance to ALS-inhibiting herbicides in rice by base editing. Sci China Life Sci 64:1624–1633
Zhong Z, Sretenovic S, Ren Q, Yang L, Bao Y, Qi C, Yuan M, He Y, Liu S, Liu X, Wang J, Huang L, Wang Y, Baby D, Wang D, Zhang T, Qi Y, Zhang Y (2019) Improving plant genome editing with high-fidelity xCas9 and non-canonical PAM-targeting Cas9-NG. Mol Plant 12:1027–1036
Zhou J, Peng Z, Long J, Sosso D, Liu BO, Eom JS, Yang B (2015) Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant J 82:632–643
Zong Y, Wang Y, Li C, Zhang R, Chen K, Ran Y, Qiu JL, Wang D, Gao C (2017) Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat Biotechnol 35:438–440
Zong Y, Song Q, Li C, Jin S, Zhang D, Wang Y, Qiu JL, Gao C (2018) Effificient C-to-T base editing in plants using a fusion of nCas9 and human APOBEC3A. Nat Biotechnol 36:950–953
Acknowledgements
We thank Rajagiri College of Social Sciences for all the support and help for the research.
Funding
This work was financially supported by Rajagiri College of Social Sciences (Autonomous), Under Seed Money for Faculty Minor Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Communicated by Gerhard Leubner.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Maharajan, T., Krishna, T.P.A., Rakkammal, K. et al. Application of CRISPR/Cas system in cereal improvement for biotic and abiotic stress tolerance. Planta 256, 106 (2022). https://doi.org/10.1007/s00425-022-04023-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-022-04023-w