Abstract
Main conclusion
A maize receptor kinase controls defense response to fungal pathogens by regulating jasmonic acid and antimicrobial phytoalexin production.
Abstract
Plants use a range of pattern recognition receptors to detect and respond to biotic threats. Some of these receptors contain leucine-rich repeat (LRR) domains that recognize microbial proteins or peptides. Maize (Zea mays) has 226 LRR-receptor like kinases, making it challenging to identify those important for pathogen recognition. In this study, co-expression analysis with genes for jasmonic acid and phytoalexin biosynthesis was used to identify a fungal induced-receptor like protein kinase (FI-RLPK) likely involved in the response to fungal pathogens. Loss-of-function mutants in fi-rlpk displayed enhanced susceptibility to the necrotrophic fungal pathogen Cochliobolus heterostrophus and reduced accumulation of jasmonic acid and the anti-microbial phytoalexins -kauralexins and zealexins- in infected tissues. In contrast, fi-rlpk mutants displayed increased resistance to stem inoculation with the hemibiotrophic fungal pathogen Fusarium graminearum. These data indicate that FI-RLPK is important for fungal recognition and activation of defenses, and that F. graminearum may be able to exploit FI-RLPK function to increase its virulence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants such as the major crop, maize (Zea mays), are hosts to a wide range of pathogens that infect different parts of the plant and cause numerous diseases that can negatively impact yield (Munkvold and White 2016). Two common fungal pathogens of maize are: Cochliobolus heterostrophus, a necrotrophic pathogen that is the causal agent of southern leaf blight, eliciting lesions on leaves and other aboveground organs of the plant; and Fusarium graminearum, a hemibiotrophic pathogen and the causal agent of seedling blight and Gibberella stalk and ear rots (Munkvold and White 2016; Yang et al. 2017).
Plants have active immune systems that recognize and respond to pathogen attack. The responses in maize to fungal pathogens include an induction of the plant hormone jasmonic acid, activation of signaling cascades and the transcription of antifungal genes. A subset of these antifungal genes encodes enzymes that produce antimicrobial phytoalexins that can act to limit fungal growth (Schmelz et al. 2011; Christensen et al. 2018; Ding et al. 2020). Two major classes of phytoalexins in maize are the labdane-related diterpenoid kauralexins and the sesquiterpenoid zealexins. The committed step for kauralexin synthesis is the bi-cyclization of geranylgeranyl diphosphate into ent-copalyl diphosphate by the ent-copalyl diphosphate synthase, Anther ear 2 (An2, GRMZM2G044481) (Harris et al. 2005; Vaughan et al. 2015). While the committed step for zealexin biosynthesis is the formation of bicyclic olefin, β-macrocarpene, from farnesyl diphosphate by the terpene synthase tps6 (GRMZM2G127087) (Ding et al. 2020). Ent-copalyl diphosphate and β-macrocarpene are then converted into kauralexin A1 and zealexin A1, respectively, by cytochrome P450s. These phytoalexins are then further modified into suites of related antimicrobial compounds (Schmelz et al. 2011; Christensen et al. 2018; Ding et al. 2020). These phytoalexins are important for antifungal defenses in maize as evidenced by the increased susceptibility of the kauralexin-deficient an2 mutants to C. heterostrophus (Christensen et al. 2018). The enzymes and pathways responsible for the production of these phytoalexins are slowly being revealed, yet little is known about how the induction/production of these compounds is regulated in response to specific pathogens.
What is known, is that plants recognize specific pathogens using a combination of membrane bound pattern-recognition receptors (PRRs) that recognize conserved epitopes from the pathogens, and intercellular nucleotide binding domain leucine-rich repeat receptors (NLR) that recognize pathogen effector proteins. These receptors form immune complexes that consist of a variety of receptor-like kinases (RLK) and receptor-like proteins that act to induce signaling cascades leading to activation of defenses (Tang et al. 2017).
PPRs have been identified in several plant species, with a focus on model plants such as Arabidopsis and tomato (Boutrot and Zipfel 2017). Yet the identity and function of many PRRs is still elusive. In maize, for example, genome analysis revealed the presence of 151 NLR genes and 226 leucine-rich repeat (LRR)-RLK genes that may function in defense activation and signaling (Song et al. 2015). The specific function of most of these genes is unknown. The challenge, therefore, is to uncover which members of these large gene families are part of anti-fungal immune receptor complexes, identify which pathogens they recognize and understand what defense mechanisms they regulate. One approach to do this is, to use co-expression analysis to find candidate genes that co-express with known anti-fungal defense components. Co-expression is often an indicator of shared function and can be used to home in on candidates in large gene families that are involved in a specific function (Block et al. 2014; Latimer et al. 2018; van Dam et al. 2018). This technique works by analyzing the expression levels of a large array of genes in publicly available RNAseq and microarray datasets and ranking the genes based on similar expression patterns to the gene of interest across all the available tissues and treatments (Obayashi et al. 2018). This holistic view of co-expression facilitates the identification of candidate genes involved in specific pathways/processes; the candidates can then be tested using traditional molecular genetic approaches.
As phytoalexins have been shown to play important roles in anti-fungal defenses and the phytohormone jasmonic acid is known to regulate antifungal defenses in an array of plant species (Antico et al. 2012), we used co-expression analysis with known jasmonic acid and phytoalexin biosynthesis genes to identify a candidate LRR-RLK associated with these anti-fungal defense related responses in maize. We isolated loss-of-function mutants in this gene and revealed that this LRR-RLK is indeed an important component of the defense response of maize against fungal pathogens, and it can regulate fungal induced jasmonic acid accumulation and the activation of phytoalexin biosynthesis.
Materials and methods
Co-expression analysis
The top 300 co-expressed genes for each query gene were determined by Logit Score (MR) which is a monotonic transformation (negative logit) of the mutual rank (MR) index using the data available in ATTED II (https://atted.jp/) (Obayashi et al. 2018). The number of overlapping genes in the top co-expressors for different query genes was determined using lists of the 300 genes plus the query gene on the Venn Diagram tool on (http://bioinformatics.psb.ugent.be/webtools/Venn/).
Isolation of maize mutants
Data available on the Maize Genetics and Genomics Database (https://maizegdb.org) was analyzed to identify two maize lines in the BzW22 inbred background with transposons inserted in the fi-rlpk gene GRMZM2G132212. These lines were obtained from the Maize Genetics Cooperation Stock Center (maizecoop.cropsci.uiuc.edu) and named fi-rlpk1 (mu1039318) and fi-rlpk2 (mu1067999). They are from the UniformMu collection and contain a Mutator transposable element (McCarty et al. 2013). The insertions were confirmed to be present in the predicted coding sequence of the first exon of FI-RLPK using the transposon-specific PCR primer TIR6 (5ʹ-AGAGAAGCCAACGCCAWCGCCTCYATTTCGTC-3ʹ) in combination with gene-specific primers for fi-rlpk: mu1039318fw (5ʹ-GGACCACAACAACTTCACCG-3ʹ), mu1039318rev (5ʹ-TCTGGAGGTTCCATATCCCG-3ʹ), mu1067999fw (5ʹ-ATAGCCCAGCTACCGTCCTT-3ʹ) and mu1067999rev (5ʹ-TCCATTGTTCTCCAACGACA-3ʹ). The resulting PCR bands were sequenced to confirm the transposon insertion sites. A representative genotyping gel and the location of the transposon insertion sites are shown in Fig. S1. The homozygous mutant lines were established by self-pollinating genotyped, homozygous mutants.
Leaf inoculation with Cochliobolus heterostrophus
The C. heterostrophus was cultured on plates containing 30% V8 juice (v/v) in water and 20 g L−1 agar, at pH 6.0 at 25 °C under a 12 h day/night cycle for 3–4 weeks. Spores were harvested in 0.1% (v/v) Tween-20 and adjusted to 1 × 106 spores/mL. The fourth leaf of each 2-week-old maize plant was spot-inoculated with 10 μL of the spore suspension applied at each of four spots, two either side of the mid vein at a 2 cm spacing. Mock inoculations were performed using 0.1% (v/v) Tween-20 alone. Post inoculation the plants were immediately placed at 100% humidity for 24 h then returned to the greenhouse with 12 h supplemental lighting using a combination of 400-Watt high pressure sodium and metal halide bulbs. The greenhouse temperature range was 25–40 °C. Tissue was collected at the indicated times by cutting the leaf at 0.5 cm from either side of the inoculation site. Lesion area was determined at 4 days post-inoculation by photographing the leaves and measuring lesion area using manual lesion edge detection and ImageJ software (Rasband 1997). A total of 24 lesions were imaged per treatment, per experiment. Collected tissue was flash frozen in liquid nitrogen at the relevant time post inoculation and used for subsequent analyses. Each experiment was repeated twice with comparable results and data from one representative experiment shown in each figure.
Stem inoculation with Fusarium graminearum
The F. graminearum was cultured on plates containing 30% V8 juice (v/v) in water and 20 g L−1 agar, at pH 6.0 at 25 °C under a 12 h day/night cycle for 12–14 days. Spores were harvested in 0.1% Tween 20 (v/v) and adjusted to 1 × 106 spores/mL. For stem inoculations, a 1 cm long incision was made into the stack node of the stem of each 30-day-old plants and 0.2 mL of spore suspension placed in the incision. Each stem was then wrapped in Parafilm® to maintain high relative humidity around the inoculation site. Plants were grown in the standard greenhouse conditions described above. Infected tissue was collected at the indicated time points by splitting the stem and removing the tissue to a depth of 2 mm on either side of the infection site and a length of 1 cm above and below the infection site, using a razor blade. Collected tissue was flash frozen in liquid nitrogen and used for subsequent analyses. Each experiment was repeated twice with comparable results and data from one representative experiment shown in each figure.
Metabolite analysis
Ergosterol was extracted in MeCl2, derivatized with methyl-N-(trimethylsilyl) trifluoroacetamide and analyzed by gas chromatography-mass spectrometry (GC–MS) as described in Christensen et al. (2018). For jasmonic acid, kauralexin, and zealexin quantification, samples were solvent extracted, methylated, collected on a polymeric adsorbent using vapor phase extraction, and analyzed using GC–MS as previously described (Schmelz et al. 2004). Total zealexin was determined by combining the amounts of the major zealexins (zealexin A1 and zealexin B1). Total kauralexin was determined by combining the amounts of the major kauralexins (kauralexin A1, kauralexin A2, kauralexin A3, kauralexin B1, kauralexin B2 and kauralexin B3).
Gene expression analysis
Total cDNA was made from DNAse I treated RNA, using oligo dT primers and the Retroscript® kit from Ambion (Carlsbad, CA, USA) according to the manufacturer’s instructions. Relative expression levels were determined by quantitative Real Time PCR (qRT-PCR) using SSoAdvanced Universal SYBR Green Supermix® (Biorad, Hercules, CA, USA) on a CFX-96 thermocycler® (Biorad) using gene specific primers for FI-RLPK (5ʹ-CTCGACTCGGAGCTCAATGC-3ʹ and 5ʹ-GTGTACAC-GGACTCGGGAGC-3ʹ) and the geometric mean of fpgs (5ʹ-ATCTCGTTGGGGATGTCTTG-3ʹ and 5ʹ-AGCACCGTTCAAATGTCTCC-3ʹ) and ubcp (5ʹ-CAGGTGGGGTATTCTTGGTG-3ʹ and 5ʹ-ATGTTCGGGTGGAAAACCT-3ʹ) as reference genes according to the 2(ΔΔcq) method (Manoli et al. 2012).
Statistical analysis
Gene expression, metabolite concentration, and lesion area data were determined to be statistically different using t-tests with P ≤ 0.05 for C. heterostrophus and F. graminearum treatments.
Results
Co-expression analysis was used to identify a candidate leucine rich repeat receptor-like kinase associated with jasmonic acid and phytoalexin production
To find candidate LRR-RLK genes that are potentially involved in regulating the production of defense related antimicrobial phytoalexins such as zealexin A1 and kauralexin A1 (Fig. 1a), the ATTED II co-expression database, which compiles and analyzes multiple publicly available microarray and RNAseq datasets (Obayashi et al. 2018), was mined with known maize phytoalexin or jasmonic acid biosynthesis genes. The LRR-RLK gene GRMZM2G132212, hereafter referred to as fungal induced-receptor like protein kinase (FI-RLPK), was found among the top 200 co-expressors of the jasmonic acid biosynthesis gene linoleate 13S-lipoxygenase 8 (lox8, GRMZM2G104843) (Acosta et al. 2009), the kauralexin biosynthetic gene an2 and the zealexin biosynthetic gene tps6. The only other LRR-RLK with such high co-expression with the phytoalexin biosynthesis genes was BRASSINOSTEROID INSENSITIVE 1-associated receptor kinase 1 (BAK1, GRMZM2G145440), a known PRR co-receptor in Arabidopsis (Yashuda et al. 2017).
Co-expression of FI-RLPK in maize with jasmonic acid and phytoalexin biosynthetic genes. a Structures of the phytoalexins zealexin A1 and kauralexin A1. b A Venn diagram showing the overlap between the top 300 co-expressed genes of FI-RLPK and the top 300 co-expressed genes of each of the jasmonic acid biosynthetic enzymes opr7 and lox8. c A Venn diagram showing the overlap between the top 300 co-expressed genes of FI-RLPK and those of kauralexin biosynthetic gene an2 and the zealexin biosynthetic gene tps6. d A Venn diagram showing the overlap between the top 300 co-expressed genes of FI-RLPK, an2, and the PRR co-receptor bak1
To assess the degree of overlap in the co-expression of these pathways, the number of shared genes within the top 300 co-expressors was analyzed (Table S1). For overlap with the jasmonic acid biosynthesis pathway, the number of shared genes between FI-RLPK, lox8 and the jasmonate biosynthesis gene 12-oxophytodienoate reductase 7 (opr7, GRMZM2G148281) (Yan et al. 2012) was determined (Fig. 1b). Interestingly, the two jasmonic acid biosynthesis genes shared only 17 (6%) of the top co-expressors, likely due to redundant functions between opr7 and the 12-oxophytodienoate reductase 8 (opr8). Yet FI-RLPK shared 23 (8%) of the top co-expressors with opr7 and 61 (20%) with lox8, indicating that FI-RLPK is functionally associated with jasmonic acid biosynthesis pathways.
For the phytoalexins, the top 300 co-expressors of an2 and tps6 were compared with those of FI-RLPK (Fig. 1c). The strong functional association and shared enzymes of the terpenoid phytoalexin biosynthesis pathways is reflected in the zealexin biosynthetic gene tps6 sharing 213 (70%) of its top co-expressors with the kauralexin biosynthetic gene an2. In comparison, FI-RLPK shared 89 (30%) of the top co-expressors with tps6, 83 (28%) with an2 and 68 (23%) with both an2 and tps6. These data support the hypothesis that FI-RLPK is functionally associated with phytoalexin production. Furthermore, FI-RLPK shared 75 (25%) of its top 300 co-expressors with the PRR co-receptor bak1 (Fig. 1d), and 59 (20%) with both an2 and bak1.
Expression of the receptor-like kinase FI-RLPK is induced in response to fungal attack
As jasmonic acid, kauralexins and zealexins are known to be induced in maize in response to fungal pathogens (Christensen et al. 2018), the expression of FI-RLPK was assessed in maize inoculated with the fungal pathogens C. heterostrophus and F. graminearum (Fig. 2). Leaf inoculation with C. heterostrophus led to a rapid 70-fold induction of FI-RLPK expression, peaking around 1-day post infection, then slowly declining over the subsequent two days (Fig. 2a). Stem inoculation with F. graminearum also led to a 50-fold induction of FI-RLPK expression (Fig. 2b). The kinetics of this induction differed from that seen in response to C. heterostrophus in that it slowly increased after inoculation rather than peaking early. The induction following inoculation with F. graminearum was around 20-fold at 2 days post infection and 50-fold at 3 days post infection. FI-RLPK expression is therefore induced in response to attack by these two fungal pathogens.
Expression of FI-RLPK in maize is induced in response to fungal pathogens. a Relative expression of FI-RLPK in leaves of 2-week-old plants inoculated with Cochliobolus heterostrophus. b Relative expression of FI-RLPK in stems of 4-week-old plants inoculated with Fusarium graminearum. *Samples significantly different from that of the non-treated tissue at P ≤ 0.05 by a t test, n = 4. c Location of Uniform mu transposon insertions in the FI-RLPK gene are given in bp with ATG as position 1 and stop as position 3346 bp. Thick bars show exon and thin lines introns and UTR. d Domain structure of the FI-RLPK protein, Leucine Rich Repeat (LRR), transmembrane domain (TM) and kinase domain
FI-RLPK mediates resistance to C. heterostrophus
To functionally test the role of this candidate RLPK in mediating the response to fungal pathogens, two maize mutants with transposons in the first exon of FI-RLPK were isolated from the Uniform Mu collection (Supplementary Fig. S1). The mutants were named fi-rlpk1 and fi-rlpk2 (Fig. 2c) and the transposon insertions occur at 891 bp and 1240 bp after the start site and fall within the coding sequence for the leucine rich repeat domains of the protein (Fig. 2c–d), rendering the maize lines loss-of-function mutants. The two mutant lines and their corresponding wild type (BzW22) were each spot inoculated on their leaves with C. heterostrophus spores, and the lesion areas quantified at 4 days post-inoculation. Significantly larger disease lesions were caused by C. heterostrophus on leaves of both fi-rlpk1 and fi-rlpk2 plants when compared to leaves of the wild type control plants (Fig. 3a, b). Measurement of the fungal sterol ergosterol—as an estimate of fungal biomass within the plant tissue—revealed significantly more ergosterol in inoculated leaves of fi-rlpk1 and fi-rlpk2 plants at 4 days post-inoculation compared to that in wild type plants (Fig. 3c). These data show that the fi-rlpk mutants were significantly less resistant to C. heterostrophus than the wild type plants.
Phenotype of fi-rlpk mutants in response to infection with Cochliobolus heterostrophus. Leaves of 2-week-old wild type (WT) and mutant maize plants were non-treated (NT) or inoculated with C. heterostrophus (C. het) and fungal growth, symptoms and plant defense metabolite levels assessed at 4 days post infection. Jasmonic acid levels were assessed at 24 h post infection. a Symptoms, b lesion area, and c levels of the fungal sterol, ergosterol were measured. d The phytohormone jasmonic acid and the phytoalexins. e Kauralexins. f Zealexins. Data from one representative experiment is shown (Bars reflect ± standard error of the mean, and asterisks denote treatments significantly different from those seen in the WT plants (P ≤ 0.05) using a t-test, number of plants per treatment in this experiment = 6)
To determine at what stage in the resistance response FI-RLPK functions, the amounts of the defense related plant hormone jasmonic acid were measured in non-treated and C. heterostrophus inoculated leaves of wild type and fi-rlpk mutant plants at 1 day post inoculation. Jasmonic acid levels were significantly lower in C. heterostrophus infected fi-rlpk1 and fi-rlpk2 plants compared to those in the wild type plants (Fig. 3d). These data suggest that jasmonic acid is involved in FI-RLPK-mediated resistance to C. heterostrophus and that FI-RLPK functions prior to jasmonic acid induction. There was no significant impact of the fi-rlpk mutants on levels of jasmonic acid in the leaves of non-treated plants (Fig. 3d).
As FI-RLPK co-expresses with terpenoid phytoalexin biosynthesis genes, the amount of kauralexins (Fig. 3e) and zealexins (Fig. 3f) in C. heterostrophus infected plants was measured. Kauralexins and zealexins were barely detectable in the leaves on non-treated plants. Kauralexin levels in the leaves of wild type plants reached around 300 µg/g in response to C. heterostrophus infection, while zealexins reached around 10 μg/g. Kauralexins were, therefore, the more predominant of the two phytoalexins induced in leaves in response to this pathogen. Total kauralexins and zealexins at 4 days post-inoculation were induced to two- to threefold higher amounts in wild type plants inoculated with C. heterostrophus than in fi-rlpk1 and fi-rlpk2 mutants. These data indicate that FI-RLPK is involved in regulating the induction of phytoalexin production in response to C. heterostrophus infection.
FI-RLPK mediates susceptibility to F. graminearum
To determine if FI-RLPK’s function in meditating fungal resistance in maize is specific to C. heterostrophus, the stems of fi-rlpk mutant and wild type plants were inoculated with F. graminearum. In contrast to their response to C. heterostrophus infections, fi-rlpk1 and fi-rlpk2 plants were more resistant to F. graminearum than the wild type plants, as evidenced by lower concentrations of the fungal sterol ergosterol detected in the mutant plants (Fig. 4a). Interestingly, despite the differences in susceptibility, inoculation with F. graminearum also led to significantly more jasmonic acid induction in wild type plants (~ tenfold) compared to the fi-rlpk1 and fi-rlpk2 plants (~ fivefold) (Fig. 4b). No differences were observed in jasmonic acid levels in non-treated stems of mutant and wild type plants. These data indicate that FI-RLPK mediates fungal induced jasmonic acid production in response to both F. graminearum and C. heterostrophus.
Phenotype of fi-rlpk mutants in response to infection with Fusarium graminearum. Stems of 4-week-old wild type (WT) and mutant maize plants were inoculated with F. graminearum (F.gram) and fungal growth and plant defense metabolite levels assessed at 4 days post-inoculation in infected and non-treated (NT) plants. Metabolites quantified were a the fungal sterol, ergosterol, b the phytohormone jasmonic acid and the phytoalexins, c kauralexins and d zealexins. Data from one representative experiment is shown (Bars reflect ± standard error of the mean, and asterisks denote treatments significantly different from those seen in WT (P ≤ 0.05) using a t-test, number of plants per treatment in this experiment = 6)
In contrast to C. heterostrophus infections, both kauralexins (Fig. 4c) and zealexins (Fig. 4d) were induced to high concentrations (200–400 μg/g) in the stems of wild type plants in response to F. graminearum infection. Furthermore, fi-rlpk1 and fi-rlpk2 plants displayed wild type phytoalexin accumulation in response to F. graminearum infection indicating that FI-RLPK is not a major player in F. graminearum induced phytoalexin accumulation.
Discussion
Our co-expression analysis was used to identify FI-RLPK as a candidate gene that was likely functionally involved with jasmonic acid and phytoalexin production as well as having an association with PPRs. Our qRT-PCR data showed that it was strongly expressed in response to fungal pathogens and our analysis of loss-of-function fi-rlpk mutants in maize demonstrated that it is involved in mediating the defense responses of maize to fungal pathogens.
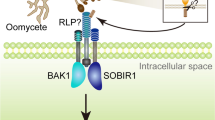
FI-RLPK has an N-terminal protein–protein or protein-peptide binding LRR domain, a transmembrane domain, and a C-terminal serine/threonine protein kinase domain. The LRR domains on these LRR-RLK proteins are often extracellular and the kinase domains intercellular. LRR-RLKs function either as PRRs for protein or peptide ligands or as co-receptors in stress or developmental signaling pathways (Wu et al. 2016). It is likely, therefore, that FI-RLPK is either a PRR that recognizes fungal or damage related peptides or proteins, or it is a co-receptor to such a PRR. However, the relatively high number of LRR domains on FI-RLPK indicates a greater likelihood of FI-RLPK being a PRR than a co-receptor. LRR containing receptor like kinases in cereal crops involved in basal resistance to F. graminearum have been identified in barley (HvLRRK-6H) and wheat (TaLRRK-6D) (Thapa et al. 2018). As is the case for FI-RLPK, the associated immune complexes and ligands for these receptors remain to be identified. Our co-expression data hints at a functional association between FI-RLPK and the co-receptor BAK1. This co-receptor is a common component of several immune complexes (Yashuda et al. 2017) so it is possible that it also forms part of an immune complex with FI-RLPK, though this would need to be confirmed experimentally.
The clearest evidence that FI-RLPK may be a part of an immune complex comes from the interaction of maize with C. heterostrophus, in which the fi-rlpk mutants have enhanced susceptibility to the fungus, and diminished jasmonic acid and phytoalexin production. These data support the hypothesis that FI-RLPK is involved in the recognition of attack by C. heterostrophus and subsequent signaling to induce jasmonic acid production. This jasmonic acid production leads to increased phytoalexins (Huffaker et al. 2011; Schmelz et al. 2011). We have previously demonstrated that kauralexin-deficient an2 mutants are more susceptible to C. heterostrophus (Christensen et al. 2018). The increased susceptibility of the fi-rlpk mutants to this pathogen could, therefore, be due in part to reduced terpenoid phytoalexin production.
The fi-rlpk mutants also had reduced jasmonic acid production during stem infection with F. graminearum compared to the wild type plants. This implies that the role of FI-RLPK in pathogen perception and jasmonic acid induction is likely the same as for C. heterostrophus. Yet the fi-rlpk mutants were less susceptible, rather than more susceptible, to F. graminearum. This contrasting effect on resistance may stem from the different roles of jasmonic acid in regulating the responses to necrotrophic and biotrophic pathogens (Glazebrook 2005). For instance, endogenous application of jasmonic acid to maize can enhance its resistance to Rhizopus microspores and Colletotrichum graminicola, which are both the necrotrophic pathogens (Schmelz et al. 2011). Yet studies in Arabidopsis and wheat revealed a dichotomous role for jasmonic acid in response to the hemibiotrophic fungus F. graminearum, where it promotes susceptibility in early (biotrophic) infection stages and resistance in later (necrotrophic) phases of infection likely via a complex relationship with salicylic acid (Mankandar et al. 2010,2012). As our data show that the fi-rlpk mutants show reduced jasmonic acid production to both pathogens, it is possible that the difference in susceptibility of the mutants could be due to the differing roles of jasmonic acid to the defense response against necrotrophic C. heterostrophus compared to the hemibiotrophic F. graminearum.
A further difference between the responses to the two pathogens was observed during phytoalexin accumulation. As in leaf infections with C. heterostrophus, recognition of the pathogen by FI-RLPK or an FI-RLPK complex led to jasmonic acid production and subsequent phytoalexin induction. Whereas, with stem inoculation F. graminearum is recognized by FI-RLPK and leads to jasmonic acid induction, but this jasmonic acid production is not required for the induction of high levels of phytoalexins during this interaction. These data suggest either there are tissue specific differences in FI-RLPK activity resulting from the presence of different receptor components and/or downstream signaling events, or that a different receptor complex/signaling mechanism regulates the large amounts of phytoalexins produced in response to F. graminearum. There are known differences in the phytoalexin profiles produced during stem infections by C. heterostrophus and F. graminearum, as well as differences in the ability of the phytoalexins to suppress fungal growth. For instance, in a previous study, we showed that the kauralexin deficient an2 maize mutants were more susceptible to C. heterostrophus but not to F. graminearum (Christensen et al. 2018). These differences hint at underlying variation in the effective defense responses of maize to different fungal pathogens.
A possible alternate explanation for the different phenotypes observed in the mutants in response to the different pathogens is that FI-RLPK, or the immune complex it is part of, is a target for effector-mediated susceptibility by one or more F. graminearum-specific effectors. Immune complexes are likely targets for F. graminearum effectors, as three such effectors restrict the immune complex mediated oxidative burst in response to chitin when expressed in the model plant Nicotiana benthamiana (Hao et al. 2020). Further supporting this idea is the fact that the widespread fungal effector NIS1 interacts with and inhibits PRR-associated LRR-RLKs in Arabidopsis thaliana (Irieda et al. 2019). Whether the phenotype observed here is effector mediated susceptibility or serendipitous activation of defense responses that lead to hormone antagonism are interesting questions for future study. Future studies on this receptor to identify the protein or peptide it recognizes would provide insight into the range of fungal pathogens FI-RLPK detects. Identification of other immune receptor proteins that associate with FI-RLPK, as well as potential F. graminearum effector targets, could provide insight into the function of FI-RLPK in plant defense.
Conclusions
The fungal induced-leucine-rich repeat receptor-like protein kinase (FI-RLPK) of maize is a pattern recognition receptor that likely recognizes protein or peptides from various fungal species. The recognition of C. heterostrophus by FI-RLPK leads to the jasmonic acid mediated production of the antimicrobial phytoalexins, zealexins and kauralexins, which act to limit pathogen growth. FI-RLPK is therefore involved in the recognition of, and resistance to, fungal pathogens. However, F. graminearum appears to have evolved the ability exploit FI-RLPK to increase its virulence in maize.
Author contribution statement
AB conceived the original idea and research plans. AB and LdT supervised the experiments. HT and AB isolated the maize mutants. HT, RS and DH performed the fungal experiments. DH performed the gene expression analysis. JM performed phytohormone and phytoalexin quantification. SC performed ergosterol quantification. AB designed the experiments and analyzed the data. AB wrote the article with contributions of all the authors. AB agrees to serve as the author responsible for contact and ensures communication.
Data availability
All data are available upon request from the corresponding author.
Abbreviations
- An2:
-
Anther ear 2
- BAK1:
-
Brassinosteroid insensitive 1-associated receptor kinase 1
- FI-RLPK:
-
Fungal induced-receptor like protein kinase
- LRR:
-
Leucine rich repeat
- PRR:
-
Pattern-recognition receptor
- RLK:
-
Receptor-like kinase
- TPS6:
-
Terpene synthase 6
References
Acosta IF, Laparra H, Romero SP, Schmelz E, Hamberg M, Mottinger JP, Moreno MA, Dellaporta SL (2009) tasselseed1 is a lipoxygenase affecting jasmonic acid signaling in sex determination of maize. Science 323(5911):262–265. https://doi.org/10.1126/science.1164645
Antico CJ, Colon C, Banks T, Ramonell KM (2012) Insights into the role of jasmonic acid-mediated defenses against necrotrophic and biotrophic fungal pathogens. Front Biol 7:48–56. https://doi.org/10.1007/s11515-011-1171-1
Block A, Widhalm JR, Fatihi A, Cahoon RE, Wamboldt Y, Elowsky C, Mackenzie SA, Cahoon EB, Chapple C, Dudareva N, Basset GJ (2014) The origin and biosynthesis of the benzenoid moiety of ubiquinone (coenzyme Q) in Arabidopsis. Plant Cell 26(5):1938–1948. https://doi.org/10.1105/tpc.114.125807
Boutrot F, Zipfel C (2017) Function, discovery, and exploitation of plant pattern recognition receptors for broad-spectrum disease resistance. Annu Rev Phytopathol 55:257–286. https://doi.org/10.1146/annurev-phyto-080614-120106
Christensen SA, Sims J, Vaughan MM, Hunter C, Block A, Willett D, Alborn HT, Huffaker A, Schmelz EA (2018) Commercial hybrids and mutant genotypes reveal complex protective roles for inducible terpenoid defenses in maize. J Exp Bot 69(7):1693–1705. https://doi.org/10.1093/jxb/erx495
Ding Y, Weckwerth PR, Poretsky E, Murphy KM, Sims J, Saldivar E, Christensen SA, Char SN, Yang B, Tong AD, Shen Z, Kremling KA, Buckler ES, Kono T, Nelson DR, Bohlmann J, Bakker MG, Vaughan MM, Khalil AS, Betsiashvili M, Dressano K, Köllner TG, Briggs SP, Zerbe P, Schmelz EA, Huffaker A (2020) Genetic elucidation of interconnected antibiotic pathways mediating maize innate immunity. Nat Plants 6(11):1375–1388. https://doi.org/10.1038/s41477-020-00787-9
Glazebrook J (2005) Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu Rev Phytopathol 43:205–227. https://doi.org/10.1146/annurev.phyto.43.040204.135923
Hao G, McCormick S, Usgaard T, Tiley H, Vaughan MM (2020) Characterization of three Fusarium graminearum effectors and their roles during Fusarium head blight. Front Plant Sci 11:579553. https://doi.org/10.3389/fpls.2020.579553
Harris LJ, Saparno A, Johnston A, Prisic S, Xu M, Allard S, Kathiresan A, Ouellet T, Peters RJ (2005) The maize An2 gene is induced by Fusarium attack and encodes an ent-copalyl diphosphate synthase. Plant Mol Biol 59(6):881–894. https://doi.org/10.1007/s11103-005-1674-8
Huffaker A, Kaplan F, Vaughan MM, Dafoe NJ, Ni X, Rocca JR, Alborn HT, Teal PE, Schmelz EA (2011) Novel acidic sesquiterpenoids constitute a dominant class of pathogen-induced phytoalexins in maize. Plant Physiol 156(4):2082–2097. https://doi.org/10.1104/pp.111.179457
Irieda H, Inoue Y, Mori M, Yamada K, Oshikawa Y, Saitoh H, Uemura A, Terauchi R, Kitakura S, Kosaka A, Singkaravanit-Ogawa S, Takano Y (2019) Conserved fungal effector suppresses PAMP-triggered immunity by targeting plant immune kinases. Proc Natl Acad Sci USA 116(2):496–505. https://doi.org/10.1073/pnas.1807297116
Latimer S, Li Y, Nguyen TTH, Soubeyrand E, Fatihi A, Elowsky CG, Block A, Pichersky E, Basset GJ (2018) Metabolic reconstructions identify plant 3-methylglutaconyl-CoA hydratase that is crucial for branched-chain amino acid catabolism in mitochondria. Plant J 95(2):358–370. https://doi.org/10.1111/tpj.13955
Makandar R, Nalam V, Chaturvedi R, Jeannotte R, Sparks AA, Shah J (2010) Involvement of salicylate and jasmonate signaling pathways in Arabidopsis interaction with Fusarium graminearum. Mol Plant Microbe Interact 23(7):861–870. https://doi.org/10.1094/MPMI-23-7-0861
Makandar R, Nalam VJ, Lee H, Trick HN, Dong Y, Shah J (2012) Salicylic acid regulates basal resistance to Fusarium head blight in wheat. Mol Plant Microbe Interact 25(3):431–439. https://doi.org/10.1094/MPMI-09-11-0232
Manoli A, Sturaro A, Trevisan S, Quaggiotti S, Nonis A (2012) Evaluation of candidate reference genes for qPCR in maize. J Plant Physiol 169:807–815. https://doi.org/10.1016/j.jplph.2012.01.019
McCarty DR, Latshaw S, Wu S, Suzuki M, Hunter CT, Avigne WT, Koch KE (2013) Mu-seq: sequence-based mapping and identification of transposon induced mutations. PLoS ONE 8(10):e77172. https://doi.org/10.1371/journal.pone.0077172
Munkvold GP, White DG (2016) Compendium of corn diseases, 4th edn. American Phytopathological Society, St Paul. https://doi.org/10.1094/9780890544945
Obayashi T, Aoki Y, Tadaka S, Kagaya Y, Kinoshita K (2018) ATTED-II in 2018: A plant coexpression database based on investigation of the statistical property of the mutual rank index. Plant Cell Physiol 59(1):e3. https://doi.org/10.1093/pcp/pcx191
Rasband WS (1997) (1997–2015) ImageJ. National Institutes of Health, Bethesda, Maryland, USA. https://imagej.nih.gov/ij and doi: https://doi.org/10.1186/s12859-017-1934-z
Schmelz EA, Engelberth J, Tumlinson JH, Block A, Alborn HT (2004) The use of vapor phase extraction in metabolic profiling of phytohormones and other metabolites. Plant J 39:790–808. https://doi.org/10.1111/j.1365-313X.2004.02168.x
Schmelz EA, Kaplan F, Huffaker A, Dafoe NJ, Vaughan MM, Ni X, Rocca JR, Alborn HT, Teal PE (2011) Identity, regulation, and activity of inducible diterpenoid phytoalexins in maize. Proc Natl Acad Sci USA 108(13):5455–5460. https://doi.org/10.1073/pnas.1014714108
Song W, Wang B, Li X, Wei J, Chen L, Zhang D, Zhang W, Li R (2015) Identification of immune related LRR-containing genes in maize (Zea mays L.) by genome-wide sequence analysis. Int J Genom. https://doi.org/10.1155/2015/231358
Tang D, Wang G, Zhou J-M (2017) Receptor kinases in plant-pathogen interactions: more than pattern recognition. Plant Cell 29:618–637. https://doi.org/10.1105/tpc.16.00891
Thapa G, Gunupuru LR, Hehir JG, Kahla A, Mullins E, Doohan FM (2018) A pathogen-responsive leucine rich receptor like kinase contributes to Fusarium resistance in cereals. Front Plant Sci 9:867. https://doi.org/10.3389/fpls.2018.00867
van Dam S, Võsa U, van der Graaf A, Franke L, de Magalhães JP (2018) Gene co-expression analysis for functional classification and gene-disease predictions. Brief Bioinform 19(4):575–592. https://doi.org/10.1093/bib/bbw139
Vaughan MM, Christensen S, Schmelz EA, Huffaker A, McAuslane HJ, Alborn HT, Romero M, Allen LH, Teal PE (2015) Accumulation of terpenoid phytoalexins in maize roots is associated with drought tolerance. Plant Cell Environ 38(11):2195–2207. https://doi.org/10.1111/pce.12482
Wu Y, Xun Q, Guo Y, Zhang J, Cheng K, Shi T, He K, Hou S, Gou X, Li J (2016) Genome-wide expression pattern analyses of the Arabidopsis leucine-rich repeat receptor-like kinases. Mol Plant 9(2):289–300. https://doi.org/10.1016/j.molp.2015.12.011
Yan Y, Christensen S, Isakeit T, Engelberth J, Meeley R, Hayward A, Emery RJ, Kolomiets MV (2012) Disruption of OPR7 and OPR8 reveals the versatile functions of jasmonic acid in maize development and defense. Plant Cell 24(4):1420–1436. https://doi.org/10.1105/tpc.111.094151
Yang Q, Balint-Kurti P, Xu M (2017) Quantitative disease resistance: dissection and adoption in maize. Mol Plant 10(3):402–413. https://doi.org/10.1016/j.molp.2017.02.004
Yasuda S, Okada K, Saijo Y (2017) A look at plant immunity through the window of the multitasking coreceptor BAK1. Curr Opin Plant Biol 38:10–18. https://doi.org/10.1016/j.pbi.2017.04.007
Acknowledgements
This work was funded by the United States Department of Agriculture (USDA)-Agricultural Research Service Project number 6036-11210-001-00D to A.B. and S.C., and the USDA- National Institute of Food and Agriculture-Specialty Crop Research Initiative grant 2018-51181-28419 to A.B. and L.d.T. The use of trade name, commercial product or corporation in this publication is for the information and convenience of the reader and does not imply an official recommendation, endorsement or approval by the USDA or the Agricultural Research Service for any product or service to the exclusion of others that may be suitable. USDA is an equal opportunity provider and employer. The authors declare no conflicts of interest with this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Block, A.K., Tang, H.V., Hopkins, D. et al. A maize leucine-rich repeat receptor-like protein kinase mediates responses to fungal attack. Planta 254, 73 (2021). https://doi.org/10.1007/s00425-021-03730-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-021-03730-0