Abstract
Main conclusion
Using genome-wide SNP association mapping, a total of 77 and 7 loci were identified for rice bacterial blight and bacterial leaf streak resistance, respectively, which may facilitate rice resistance improvement.
Abstract
Bacterial blight (BB) and bacterial leaf streak (BLS) caused by Gram-negative bacteria Xanthomonas oryzae pv. oryzae (Xoo) and X. oryzae pv. oryzicola (Xoc), respectively, are two economically important diseases negatively affecting rice production. To mine new sources of resistance, a set of rice germplasm collection consisting of 895 re-sequenced accessions from the 3000 Rice Genomes Project (3 K RGP) were screened for BB and BLS resistance under field conditions. Higher levels of BB resistance were observed in aus/boro subgroup, whereas the japonica, temperate japonica and tropical japonica subgroups possessed comparatively high levels of resistance to BLS. A genome-wide association study (GWAS) mined 77 genomic loci significantly associated with BB and 7 with BLS resistance. The phenotypic variance (R2) explained by these loci ranged from 0.4 to 30.2%. Among the loci, 7 for BB resistance were co-localized with known BB resistance genes and one for BLS resistance overlapped with a previously reported BLS resistance QTL. A search for the candidates in other novel loci revealed several defense-related genes that may be involved in resistance to BB and BLS. High levels of phenotypic resistance to BB or BLS could be attributed to the accumulation of the resistance (R) alleles at the associated loci, indicating their potential value in rice resistance breeding via gene pyramiding. The GWAS analysis validated the known genes underlying BB and BLS resistance and identified novel loci that could enrich the current resistance gene pool. The resources with strong resistance and significant SNPs identified in this study are potentially useful in breeding for BB and BLS resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The ancient domesticated crop, rice (Oryza sativa L.) is one of the most widely cultivated grain crops all over the world and contributes significantly to global food security (Khush 2005). Diseases caused by bacterial, fungal, and viral pathogens pose continuous threats to crop production and lead to significant yield losses worldwide (Ou 1985). Bacterial blight (BB) and bacterial leaf streak (BLS) caused by Gram-negative bacteria Xanthomonas oryzae pv. oryzae (Xoo) and X. oryzae pv. oryzicola (Xoc), respectively, are two important diseases with frequent outbreak in tropical and temperate regions resulting in considerable damage to rice production annually (Niño‐Liu et al. 2006). BB and BLS can cause severe yield loss of up to 50% and 32%, respectively, depending on the rice variety, growth stage, the geographic location and environmental conditions (Niño‐Liu et al. 2006). Chemical control is the most common management practices to mitigate BB and BLS, but increases the cost and leads to environmental risks. Deployment of host resistance is widely accepted as the most effective strategy to relive threats due to these diseases. Identification and characterization of loci or genes for disease resistance are indispensable for both deeply understanding the genetic architecture of phenotypic variation and efficiently developing resistant rice varieties.
Host resistance to BB has been extensively studied over last several decades. To date, at least 46 BB resistance (R) genes resistant to various strains of Xoo have been identified in rice (Neelam et al. 2020; Jiang et al. 2020; Chen et al. 2020). Sixteen of them including Xa1, Xa2/Xa31, Xa3/Xa26, Xa4, xa5, Xa10, xa13, Xa14, Xa21, Xa23, xa25, Xa27, xa41, Xa45, CGS-Xo111 and Xa7 were cloned (Song et al. 1995; Yoshimura et al. 1998; Iyer and McCouch 2004; Sun et al. 2004; Gu et al. 2005; Chu et al. 2006; Xiang et al. 2006; Yang et al. 2006; Liu et al. 2011; Tian et al. 2014; Hutin et al. 2015; Wang et al. 2015a; Hu et al. 2017; Ji et al. 2020; Zhang et al. 2020a, b; Chen et al. 2021; Luo et al. 2021). Some of these genes have been widely applied in rice breeding programs, such as Xa3/Xa26 and Xa4, which play an important role in rice breeding and production in China (Gao et al. 2010; Hu et al. 2017). However, the rapid genetic evolution of the bacteria often leads to frequent breakdown of R genes after several years of large-scale commercial use (Vera-Cruz et al. 2000). The breakdown of resistance contributed by Xa3/Xa26 and Xa4 has been observed in many rice-growing regions (Wang et al. 2005). To effectively control the disease, continuous efforts of enriching the pool of R genes and pyramiding multiple R genes into elite varieties are crucial.
Compared to BB, knowledge on genetic mapping of BLS resistance loci is very limited due to lack of highly resistant rice varieties. A few of quantitative trait loci (QTL) were reported (Tang et al. 2000; Zheng et al. 2005; Chen et al. 2006; Bossa‐Castro et al. 2018). A QTL, qBlsr5a, with relatively larger effect on the short arm of chromosome 5 was finely mapped to a 30-kb interval (Xie et al. 2014). Coincidentally, the BB resistance gene xa5, encoding a small (γ) subunit of the conserved general transcription factor TFIIA, was located in the same region and proved to be responsible for the qBlsr5a effect (Xie et al. 2014, 2019; Yuan et al. 2016). Additionally, a recessive locus bls1 and a dominant locus Xo1 conferring high-level race-specific resistance to BLS were identified from Guangxi common wild rice and American heirloom rice variety Carolina Gold Select, respectively (He et al. 2012; Triplett et al. 2016). It is interesting that a non-host R gene, Rxo1, encoding a nucleotide-binding and leucine-rich repeat domain (NLR) protein, was isolated from maize and confers qualitative resistance to BLS in rice (Zhao et al. 2005). However, strong rice resistance against Xoc is rarely reported and the genetic basis of resistance remains unintelligible.
Quantitative trait locus mapping in bi-parental population is a conventional linkage-based approach extensively used to identify loci for traits of interest. This approach has some limitations, such as low allelic diversity and low recombination events in many cases. Construction of a mapping population is also labor-intensive and time-consuming. In contrast, the genome-wide association studies (GWAS) are based on linkage disequilibrium (LD) to dissect the genetic basis of complex traits in a large collection of germplasm accessions with broad diversity, accumulating more recombination events than bi-parental mapping populations and resulting in higher resolution (Nordborg and Tavaré 2002; Yu and Buckler 2006). The availability of explosive genome sequences and high-density SNP information for rice in combination with recent advances in study on the molecular interactions between plant and pathogens enable GWAS a very powerful tool for identification of loci involved in rice disease resistance. Genetic architecture of rice resistance to diverse diseases, including blast, bacterial blight, bakanae and false smut, has been investigated through GWAS in several studies (Wang et al. 2014; Kang et al. 2016; Mgonja et al. 2016; Raboin et al. 2016; Zhu et al. 2016; Dilla-Ermita et al. 2017; Zhang et al. 2017; Li et al. 2018, 2019; Long et al. 2020). Many candidate genes were identified through GWAS and some of them were cloned and characterized (Li et al. 2019; Liu et al. 2020).
Genome sequences of rice are becoming increasingly available. The 3000 Rice Genomes Project (3 K RGP) was launched and successfully accomplished by the scientists from Chinese Academy of Agricultural Sciences (CAAS), the Beijing Genomics Institute (BGI)-Shenzhen and the International Rice Research Institute (IRRI). A total of 3010 accessions originating from 89 countries were re-sequenced using the Illumina NGS technology, with an average depth of 14.3×, which provides a valuable resource for rice genomics research and breeding (The 3,000 rice genomes project 2014; Wang et al. 2018). The objectives of this study were to identify novel resistance sources and loci for BB and BLS resistance. Thus, we evaluated the BB and BLS resistance of 895 re-sequenced accessions, a subset of the germplasm selected from the 3 K RGP, to multiple strains under natural conditions. Then, GWAS was performed to detect associations between the disease reactions and high-density SNP markers. The results may facilitate genetic improvement of rice resistance against BB and BLS.
Materials and methods
Plant materials
A total of 895 geographically diverse O. sativa germplasm accessions included in the 3 K RGP (The 3,000 rice genomes project 2014) were provided by the International Rice Research Institute (IRRI). These accessions, including indica (371), japonica (60), temperate japonica (143), tropical japonica (181), intermediate (38), aus/boro (79), and basmati/sadri (23), displayed obvious variation in many agronomic traits. Based on the feasibility or practicability of the field experiments, we randomly selected 407, 271 and 528 of the 3 K RGP accessions for evaluation of BB resistance at the seedling, BB resistance at the adult, and BLS resistance at the adult stage, respectively (Table S1).
Evaluation of BB and BLS resistance under field conditions
All of the tested accessions were planted in field at the experimental station of the Huazhi Bio-tech Company Ltd., Changsha, China (28° 18′ N and 113° 18′ E). BB and BLS resistance evaluations were performed in the cropping season of 2017 and 2018, respectively. The experimental field was bordered with plastic film with 2 m high to prevent the spread of inoculum. The investigation of BB resistance was conducted in 2017 using the randomized complete block design with two replications, while the observation of BLS resistance was carried out in 2018 using the randomized complete block design with one replication. To evaluate the seedling-stage BB resistance of 407 accessions, 50 seeds per accession were sown in a seedling nursery. The seedlings were inoculated with the Philippine strain PXO99 (race P6) and Chinese strain FuJ (race C8) at the three to four-leaf stage, using the method described in Kauffman et al. (1973). Briefly, the bacterial strains were cultured on peptone sucrose agar medium at 30 °C for 3 days, and each inoculum was prepared by suspending the bacterial mass in sterile water at a concentration of OD600 = 1.0. Twenty healthy plants of each accession were selected and 1–2 uppermost fully expanded leaves of each plant were inoculated using scissors dipped in bacterial suspensions to clip 1–2 cm of the leaf tip. Plant reactions to disease infection were evaluated three weeks after inoculation by measuring lesion length (LL, cm). For each replication, the average LL per accession was calculated based on five longest lesions from at least three individual plants. The average LL of two replications for each accession was used for normalization through rankTransPheno function in the FRGEpistasis R package and subsequent association analysis. To evaluate the adult-stage BB resistance of 271 accessions, 8 healthy plants per accession at the tillering or booting stage (about 60 days after sowing) were inoculated with the Philippine strains PXO99 (race P6), PXO61 (race P1) and Chinese strains FuJ (race C8), YuN24 (race C9), and disease reactions were investigated with the same method used at the seedling stage. Seven more Xoo stains from China and Philippines were used for analysis of resistance spectra. To evaluate BLS resistance of 528 accessions, eight healthy plants per accession were inoculated with Chinese strains HAB8-47 and HNB1-19 through the penetration method using a needleless syringe at the tillering or booting stage (about 60 days after sowing). One inoculation per leaf was performed in two uppermost fully expanded leaves of the main stem. LL was measured three weeks after inoculation. The mean value of six longest lesions from at least three individual plants for each accession was used for normalization and subsequent association analysis.
The resistance level of accessions was scored based on the LL. For BB, accessions with LL < 5 cm, 5 cm ≤ LL < 10 cm, 10 cm ≤ LL < 15 cm, and LL ≥ 15 cm were rated as resistant (R), moderately resistant (MR), moderately susceptible (MS), and susceptible (S), respectively (Dilla-Ermita et al. 2017). For BLS, we used the criterion described by Wonni et al. (2015) with some modifications. Accessions with LL < 0.5 cm, 0.5 cm ≤ LL < 1.0 cm, 1.0 cm ≤ LL < 1.5 cm, and LL ≥ 1.5 cm were rated as resistant (R), moderately resistant (MR), moderately susceptible (MS), and susceptible (S), respectively.
SNP genotyping data and population structure
The SNP genotyping data of the tested accessions were retrieved from Rice SNP-Seek Database (https://snp-seek.irri.org) (Mansueto et al. 2017). SNP markers with a minor allele frequency (MAF) < 5% and call rate < 20% were removed from the dataset. The most widely applied method for analysis of population structure is principal components analysis (PCA). PCA-based methods use markers (SNPs in most cases) to infer orthogonal axes of continuous variation, called principal components that reduce the data to few variables explaining most of the variation of the genetic information. Here, PCA was performed using the program SMARTPCA (Patterson et al. 2006). First five principal components (PCs) explained the most variations were used as covariates to correct population structure in association studies and first two principal components (PCs) were visualized using the package matplotlib of Python (Hunter 2007).
Genome-wide association analysis
GEMMA software and the mixed linear model (MLM) with the control of structure (Q matrix consisting of the first five PCs) and kinship (K matrix) were used to determine the association between SNP markers and the observed phenotypic traits (Zhou and Stephens 2012). To reduce the false-positive rate, SNP-trait associations were adjusted by Bonferroni correction. After Bonferroni correction, the cutoff for statistical significance was determined to be P < 4.57 × 10–6, 4.17 × 10–6, 4.37 × 10–6 for BLS, BB at the adult stage and BB at the seedling stage, respectively (Lander and Kruglyak 1995). Manhattan plots and QQ plots were generated using the package matplotlib of Python (Hunter 2007). Phenotypic variation (R2) explained by multiple SNPs in each significant locus was estimated by stepwise regression using R software (Zhou et al. 2017).
Analysis and annotation of significant association signals
We used R package Big-LD with threshold 0.6 of r2 for LD block partitioning (Kim et al. 2018). LD blocks containing significantly associated SNPs were defined as the candidate genomic loci. The SNP with most significant association in a block was determined as the lead SNP. Genes in the blocks with significant SNPs were considered as the candidates for BB and BLS resistance. Gene-based SNP annotation was performed using the IRGSP-1.0 (Ensembl release 41) of the rice genome in the program SnpEff (Cingolani et al. 2012). Information of the reported BB resistance genes near the hits was retrieved from the literature, the Q-TARO (http://qtaro.abr.affrc.go.jp/) and funRiceGenes database (http://funricegenes.ncpgr.cn/) (Yamamoto et al. 2012; Yao et al. 2018). The Rice Annotation Project (RAP) identification numbers for the candidate genes involved in BB and BLS resistance are summarized in Table S6.
Statistical analysis
All the phenotypic screening experiments were performed with at least three replications. The mean values and standard errors were calculated using Microsoft Excel 2007. Statistical significance was tested by the one-way ANOVA followed by the Duncan’s multiple range test using SPSS software. Significance was declared when P < 0.05 and indicated by different letters.
Results
Phenotypic evaluation of bacterial blight and bacterial leaf streak resistance
To evaluate the BB and BLS resistance of the selected germplasm accessions from 3 K RGP, four representative X. oryzae pv. oryzae (Xoo) strains (FuJ, YuN24, PXO99 and PXO61) and two X. oryzae pv. oryzicola (Xoc) strains (HAB8-47 and HNB1-19) were used for artificial inoculation under natural conditions in 2017 and 2018, respectively. Phenotypic reactions to BB infection at the seedling stage (BB-ss), BB and BLS at the adult stage (BB-as and BLS-as) were summarized in the supplementary tables (Tables S2, S3, S4). Distributions of disease reactions to all the BB strains at both seedling and adult stages were skewed toward susceptibility. A considerable proportion of tested accessions was susceptible/moderately susceptible to BB strains and higher proportions were observed for strains PXO99-ss and PXO99-as (85.4% and 95.5%), indicating that PXO99 was more virulent to the tested rice accessions in this study at both growth stages (Fig. 1a). However, some accessions showed high-level resistance. Three accessions, AUS 295 (IRGC 127184, aus/boro, Bangladesh), IR 70027-8-2-2-3-2 (IRGC 126028, indica, Philippines), SADA AUS (IRGC 127776, aus/boro, India), were resistant to the two Xoo strains at the seedling stage (Table S2). And three accessions, AUS 295 (IRGC 127184, aus/boro, Bangladesh), SAITA (IRGC 127778, aus/boro, Bangladesh), UCP 122 (IRGC 127871, aus/boro, Bangladesh), showed resistance to all four Xoo strains at the adult stage (Table S3). It is noteworthy that both SAITA and UCP 122 displayed resistance to FuJ and moderate resistance to PXO99 at the seedling stage, and AUS 295 was resistant to all the Xoo strains at both growth stages. We further investigated the resistance spectra and influence of plant development on BB resistance of the three accessions AUS 295, UCP 122 and SAITA. The results indicated that these three accessions conferred all-growth-stage resistance to FuJ and broad-spectrum resistance to 11 tested Xoo strains at the adult stage (Figs. S1, S2). For BLS, over half the tested accessions were resistant/moderately resistant to Xoc strains HAB8-47-as (55.9%) and HNB1-19-as (57.1%) (Fig. 1a). Twenty tested accessions including seven from tropical japonica subgroup, seven from indica subgroup, four from japonica subgroup, one from intermediate, and one from aus/boro subgroup, showed resistance to both of the two BLS strains (Table S4).
We then compared the disease severity among the seven subgroups of the observed germplasm accessions. The analysis revealed that 71.88%, 27.27%, 42.50%, 47.50%, 18.92% and 12.50% of the tested accessions were resistant or moderately resistant to FuJ-ss, PXO99-ss, FuJ-as, PXO61-as, YuN24-as and PXO99-as, respectively, in the aus/boro subgroup which was more resistant than other six subgroups against Xoo strains. But 100.00%, 100.00%, 96.30%, 96.00%, 100.00% and 100.00% of the tested accessions were susceptible or moderately susceptible to FuJ-ss, PXO99-ss, FuJ-as, PXO61-as, YuN24-as and PXO99-as, respectively, in the intermediate which was the most susceptible subgroup (Fig. 1b).
As to the resistance to BLS strains, out of the tested accessions in japonica, 68.42% and 76.19% were resistant or moderately resistant to HAB8-47 and HNB1-19, respectively. Among the tested accessions in temperate japonica, 69.23% and 68.00% were resistant or moderately resistant to HAB8-47 and HNB1-19, respectively. In the tropical japonica subgroup, 70.48% and 73.33% of the tested accessions were resistant or moderately resistant to HAB8-47 and HNB1-19, respectively. Thus, the japonica, temperate japonica and tropical japonica subgroups were relatively resistant to BLS strains. But the 60.00% and 71.43% of the tested accessions in basmati/sadri subgroup were susceptible or moderately susceptible to HAB8-47 and HNB1-19, respectively, and thus was the most susceptible subgroup (Fig. 1b).
Genome-wide association analysis
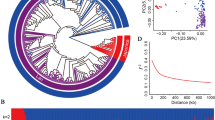
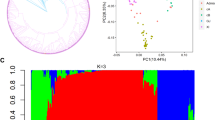
To exclude the possibility of population stratification causing false-positive association, we conducted principal component analysis (PCA) of the 407, 271 and 528 accessions for phenotypic scores of BB at the seedling and adult stages, and BLS at the adult stage, respectively (Fig. 2). PCA showed that the randomly selected sample accessions were evenly distributed among all the 3 K RGP germplasm accessions, suggesting that these three sets of sample accessions could represent the genetic variance of this unprecedented genomics resource in rice. To identify genomic loci associated with BB and BLS resistance, we performed GWAS using the SNP datasets and the phenotypic data. Totally, 823 SNPs clustering in 84 genomic loci were found to be associated with the observed phenotypic traits (Fig. 3; Table 1; Table S4). Among these loci, 77 across 12 chromosomes were significantly associated to BB resistance explaining 0.4–30.2% of the phenotypic variation, including 24 (10 and 14 associated with resistance to FuJ-ss and PXO99-ss, respectively) with resistance at the seedling stage, and 53 (13, 30, 6 and 4 associated with resistance to FuJ-as, PXO61-as, YuN24-as and PXO99-as, respectively) with resistance at the adult stage. These loci explained together 30.51% (FuJ-ss), 31.99% (PXO99-ss), 67.41% (FuJ-as), 74.48% (PXO61-as), 11.76% (YuN24-as), and 33.21% (PXO99-as) of the phenotypic variation for resistance reaction, respectively. Several hotspots of significantly associated SNPs were identified. A total of 343 SNPs at 4 loci (BBRAL8, BBRAL30, BBRAL7, BBRAL68) significantly associated with BB resistance were clustered in an approximately 245-kb region (259,484–504,974 bp) on chromosome 5, accounting for 41.70% of all the associated SNPs. A cluster of 5 resistance-associated loci, BBRL39, BBRL40, BBRL41, BBRL42 and BBRL43, spanned a region of approximately 462-kb interval (27,364,589–27,827,470 bp) on chromosome 11, and contained 202 SNPs. Moreover, an interval encompassing 84-kb (2,733,731–2,818,547 bp) on chromosome 12 harbored the locus BBRAL13 containing 61 SNPs.
Principal component analysis plots for the first two components of rice 3 K RGP accessions. PCA analysis of 407 rice accessions for evaluation of BB resistance at the seedling satge (a), 271 rice accessions for evaluation of BB resistance at the adult satge (b) and 528 rice accessions for evaluation of BLS resistance at the adult satge (c)
Genome-wide association scan of rice BB and BLS resistance. Manhattan plots for eight Xoo and Xoc strains: a FuJ-ss; b PXO99-ss; c FuJ-as; d PXO61-as; e PXO99-as; f YuN24-as; g HAB8-47-as; h HNB1-19-as. Negative log10-transformed P values from a genome-wide scan are plotted against position on each of 12 chromosomes. Black horizontal dashed line indicates the genome-wide significance threshold. The red arrows indicate that the identified loci are co-localized with previously mapped or cloned resistance Xa genes or QTL. Quantile–quantile plot for the eight strains: i FuJ-ss; j PXO99-ss; k FuJ-as; l PXO61-as; m PXO99-as; n YuN24-as; o HAB8-47-as; p HNB1-19-as
In contrast to BB, only 17 significantly associated SNPs in 7 loci including 1 and 6 for resistance to HAB8-47 and HNB1-19, respectively, were detected. These loci could explain 0.65% and 25.11% of the phenotypic variance for reaction to infection of BLS strains HAB8-47 and HNB1-19, respectively. The locus BLSRAL2 demonstrated the highest level of associations with 6 SNPs.
Validation of significant SNP-resistance associations for BB and BLS
To understand effects of the allelic variation on BB or BLS resistance, we selected 5 loci associated with the highest number of SNPs on Chromosome 5, 11, and 12. The most significantly associated SNP on each locus was considered as the candidate resistance allele. It is shown that there are five SNPs significantly associated with the resistance in the natural population (Fig. 4a). Among the 7 subgroups of rice accessions, R alleles in BBRAL8, BBRAL13 and BBRAL30 are more abundant in aus/boro accessions. We found 35.90%, 30.00% and 33.33% of accessions from aus/boro subgroup carry the R alleles in BBRAL8, BBRAL13 and BBRAL30, respectively. In BBRAL39 and BBRAL41, indica accessions harbors more R alleles than other subgroups (Fig. 4b). Additionally, we investigated the average number of R alleles of the five loci in the subgroups of accessions. The analysis indicated that aus/boro had the highest frequency of R alleles (1.05), whereas 0.35, 0.51, 0.04, 0.16, 0.15, 0.05 were observed in basmati/sadri, indica, intermediate, japonica, temperate japonica and tropical japonica, respectively. Furthermore, we analyzed the SNP haplotypes of the 84 associated loci in the rice accessions that displayed broader spectrum and high levels of resistance to BB or BLS strains. The results showed that the average frequency of the R alleles for 3 BB-resistant accessions is 77.1% and 20 BLS-resistant accessions is 67.2% (Fig. 5). These results demonstrate that the R alleles are highly enriched in these resistant accessions which are valuable resistant resources for rice BB and BLS resistance breeding.
Candidate genes involved in the BB and BLS resistance
Out of the 7 loci associated with resistance to BLS, one was co-localized with a previously reported QTL, qXO-5-3 (Table 1) (Bossa‐Castro et al. 2018). Only one significant SNP was detected in this locus (BLSRAL3). A total of 20 genes were identified in BLSRAL3 based on haplotype block structure analysis and gene annotation information (Table S6). Interestingly, one of them, Os05g0439400, encoding ubiquitin E3 ligase, was proved to positively regulate rice resistance to BB (Ishikawa et al. 2014). Another locus, BLSRAL2, which contains 6 SNPs shows highly significant and the strongest association with BLS resistance. None of BLS-resistant QTLs has been previously located in this region, suggesting that BLSRAL2 represents a new genomic region associated with BLS resistance. Among the 121 genes identified in this locus, Os02g0759400, a RING-H2-type zinc finger protein gene, could be induced by pathogen infections (Meng et al. 2006), which may be involved in the rice disease resistance.
Among the 77 loci associated with resistance to BB, 7 were co-localized with the known Xa genes (Table 1). BBRAL8, which contains the most significant SNP explaining up to 30.15% of the phenotypic variation, overlapped with the well characterized BB resistance gene xa5 (Os05g0107700). This xa5 encoding the transcription factor IIA gamma subunit 5 (TFIIAγ5) is a widely used recessive R gene in rice resistance breeding programs. Similarly, BBRAL30 and BBRAL68 were localized in the same genomic region. The lead SNP in BBRAL30 explained 29.39% of the phenotypic variation. In contrast, the only significant SNP in BBRAL68 explained 1.84% of the phenotypic variation and this locus may play minor role in the resistance to PXO99-ss. The results agreed with previous studies, that PXO99 was compatible on xa5, whereas PXO61 and FuJ were incompatible (Liu et al. 2007). In the hotspot region on chromosome 11, 5 loci (BBRL39, BBRL40, BBRL41, BBRL42 and BBRL43) involving 202 SNPs significantly associated with BB resistance spanned an approximately 462-kb genomic region, which was overlapped with 4 Xa genes, including Xa4, Xa22(t), Xa35(t) and Xa36(t). In addition to Xa4, which encodes a cell wall-associated kinase and improves the BB resistance by strengthening the cell wall (Hu et al. 2017), other three genes were not yet isolated (Wang et al. 2003; Guo et al. 2010; Miao et al. 2010). Twenty-five annotated genes were identified in the region including three NB-ARC genes (Os11g0686900, Os11g0686500, Os11g0678400) (Table S6). NB-ARC and NBS-LRR encoding genes are the most prevalent and ancient R gene families in plants. Moreover, 61 SNPs significantly associated with BB resistance were clustered in hotspot region (BBRAL13) on chromosome 12, which was mapped far from any currently known BB resistance gene and likely represent a novel resistance locus. Eight functionally annotated genes were found in this region. However, none of them were reported to be related to rice disease resistance (Table S6). These candidate genes need to be functionally verified and annotated.
Discussion
The main objectives of the current study were to identify novel germplasm with high-level and broad-spectrum resistance against Xoo and Xoc, and novel loci for BB and BLS resistance by GWAS approach. We randomly selected and constructed three sets of rice accessions, which were inoculated with 2 Xoo strains at the seedling stage, 4 Xoo strains and 2 Xoc strains at the adult stage, respectively. Most of the tested accessions showed moderate susceptibility or susceptibility against all the Xoo strains at both seedling and adult stages. Among the inoculated Xoo strains, PXO99 showed the highest virulence. We found 85.4% and 95.5% of the accessions were susceptible or moderately susceptible to PXO99 at the seedling stage and adult stage, respectively. However, over half of the tested accessions were resistant or moderately resistant to the two Xoc strains. Among the seven rice subgroups, aus/boro accessions showed relatively high frequency of BB resistance to all the strains at both seedling and adult stages. This result agreed with the previous reports that aus/boro accessions were eltite resistant resources for BB (Sidhu et al. 1978; Dilla-Ermita et al. 2017; Li et al. 2018). Dilla-Ermita et al. (2017) reported that 96% of the tested aus genotypes exhibited resistance to both strains of race 9, PXO339 and PXO349. Similarly, aus subgroup was proved to be more resistant to Xoo race C1 in another study (Li et al. 2018). Based on the phenotypic screening, we obtained three and 20 accessions exhibiting high-level resistance against BB and BLS, respectively. The three BB resistant accessions conferred not only all-growth-stage resistance, but also broad-spectrum resistance. (Tables S2, S3; Figs. S1, S2).
GWAS approach has recently been widely used to dissect the genetic architecture of rice resistance to diverse pathogens. However, most of the studies focused on rice blast disease, caused by the fungal pathogen Magnaporthe oryzae (Wang et al. 2014, 2015b; Shinada et al. 2015; Kang et al. 2016; Mgonja et al. 2016; Raboin et al. 2016; Zhu et al. 2016; Lin et al. 2018; Li et al. 2019; Lu et al. 2019; Liu et al. 2020; Zhang et al. 2020a, b). There are few reports on application of GWAS to identify loci associated with BB and BLS resistance (Dilla-Ermita et al. 2017; Zhang et al. 2017; Bossa‐Castro et al. 2018; Li et al. 2018; Kim and Reinke 2019). Using an indica/aus panel and an indica subset, 15 loci associated with resistance to Xoo strains from the Philippines were detected (Dilla-Ermita et al. 2017). Zhang et al. (2017) identified twelve loci containing 121 significantly associated signals in an indica population consisting 172 accessions. Li et al. (2018) identified 15 loci associated with the resistance against Xoo race C1 using 267 rice accessions from Rice Diversity Panel 1 (RDP1). Additionally, researchers used multi-parent advanced generation intercross (MAGIC) populations to map the QTL or genes conferring resistance to BB and BLS through combining GWAS with QTL mapping (Kim et al. 2019). To our knowledge, the present work represents the first report on identification of resistant loci against BB at the seedling stage and BLS using natural population through a GWAS approach. In this study, a total of 84 loci were identified, including 7 for BLS resistance, 24 for BB resistance at the seedling stage, and 53 for BB resistance at the adult stage. Among them, one BLSAL and seven BBRALs were overlapped with previously reported genes/QTLs. On chromosome 5, a hotspot region containing 343 SNPs significantly associated with BB resistance, was co-localized with an extensively studied and used gene xa5 (Jiang et al. 2020). It is interesting that, the frequency distribution of resistant alleles in this region was higher in aus/boro subgroup (35.90%) than any other subgroups. Our resluts agreed wtih the earlier studies that xa5 is prevalent in aus/boro (Garris et al. 2003). Therefore, xa5 is the most probable candidate gene for this region. However, the single xa5 gene would not confer resistance to the Xoo strains PXO99 and YuN24. Thus, other R genes may exist in the accessions with broad-spectrum resistance, such as Aus 295, UCP122 and SAITA. In another hotspot region on chromosome 11, 5 BBRALs involving 202 associated SNPs were overlapped with 4 Xa genes, including Xa4, Xa22(t), Xa35(t) and Xa36(t). To date, over 20 QTL conferring BLS resistance have been reported (Bossa-Castro et al. 2018; Wang et al. 2020). Two of them, qBlsr5a (xa5) and Xo1 were successfully isolated (Xie et al. 2014; Ji et al. 2020; Read et al. 2020). Among seven associated loci identified, only BLSRAL3 was found to co-localize with a fomerly described QTL qXO-5–3 on chromosome 3. Interestingly, qXO-5–3 was identified to confer resistance to both Xoo and Xoc strains (Bossa-Castro et al. 2018).
Analysis of the associated loci in these resistant accessions indicated that the high-level resistance was due to the accumulation of R alleles. It suggested that pyramiding of the resistant alleles may achieve effective resistance to BB and BLS. In addition, the lead SNPs at the five loci in hotspot regions were analyzed across all the tested accessions. The aus/boro subgroup contains more R alleles, and thus demonstrates high frequency of BB resistance.
Artificial inoculation of Xoo and Xoc strains under field conditions allowed effective identification of resistant accessions to BB and BLS. Application of GWAS in this study revealed 7 and 77 SNP loci associated with BLS and BB resistance, respectively. This study provides new insights into the genetic architecture of BLS resistance and BB resistance at two growth stages in rice. The findings in our study may facilitate the rice breeding for BB and BLS resistance. Future studies will focus on fine mapping of these resistant loci, validating the effects and functional characterization of the candidate genes.
Author contribution statement
JHP, XLL and YZY conceived and designed the study. NJ, YL, YLS, ZWL, YLX, ZZH, YTW and YL conducted the field experiments. JF, QZ and KW conducted the GWAS. NJ, JF, QZ, YL and JHP wrote the manuscript. All authors read and approved the final version of the manuscript.
References
Bossa-Castro AM, Tekete C, Raghavan C, Delorean EE, Dereeper A, Dagno K, Koita O, Mosquera G, Leung H, Verdier V, Leach JE (2018) Allelic variation for broad-spectrum resistance and susceptibility to bacterial pathogens identified in a rice MAGIC population. Plant Biotechnol J 16:1559–1568. https://doi.org/10.1111/pbi.12895
Chen C, Zheng W, Huang X, Zhang D, Lin X (2006) Major QTL conferring resistance to rice bacterial leaf streak. Agric Sci China 5:216–220. https://doi.org/10.1016/S1671-2927(06)60041-2
Chen S, Wang C, Yang J, Chen B, Wang W, Su J, Feng A, Zeng L, Zhu X (2020) Identification of the novel bacterial blight resistance gene Xa46(t) by mapping and expression analysis of the rice mutant H120. Sci Rep 10:1–11. https://doi.org/10.1038/s41598-020-69639-y
Chen X, Liu P, Mei L, Chen L, Liu H, Shen S, Ji Z, Zheng X, Zhang Y, Gao Z, Zeng D, Qian Q, Ma B (2021) Xa7, a new executor R gene that confers durable and broad-spectrum resistance to bacteria-blight disease in rice. Plant Commun. https://doi.org/10.1016/j.xplc.2021.100143
Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen JL, Zhang Q, Wang S (2006) Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev 20:1250–1255. https://doi.org/10.1101/gad.1416306
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6:80–92. https://doi.org/10.4161/fly.19695
Dilla-Ermita CJ, Tandayu E, Juanillas VM, Detras J, Lozada DN, Dwiyanti MS, Vera Cruz C, Mbanjo EGN, Ardales E, Diaz MG, Mendioro M, Thomson MJ, Kretzschmar T (2017) Genome-wide association analysis tracks bacterial leaf blight resistance loci in rice diverse germplasm. Rice 10:8. https://doi.org/10.1186/s12284-017-0147-4
Gao J, Zhao J, Xu C, Li X, Wang S (2010) Development of rice germplasms conferring high-level and broad-spectrum resistance to Xanthomonas oryzae pv. oryzae at both seedling and adult stages. Mol Plant Breed 8:420–425. https://doi.org/10.1016/S1872-2067(10)60116-7
Garris AJ, McCouch SR, Kresovich S (2003) Population structure and its effect on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice (Oryza sativa L.). Genetics 165:759–769. https://doi.org/10.1017/S0016672303006396
Gu K, Yang B, Tian D, Wu L, Wang D, Sreekala C, Yang F, Chu Z, Wang GL, White FF, Yin Z (2005) R gene expression induced by a type-III effector triggers disease resistance in rice. Nature 435:1122–1125. https://doi.org/10.1038/nature03630
Guo SB, Zhang DP, Lin XH (2010) Identification and mapping of a novel bacterial blight resistance gene Xa35(t) originated from Oryza minuta. Agric Sci China 43:2611–2618. https://doi.org/10.4028/www.scientific.net/AMM.37-38.1549
He W, Huang D, Li R, Qiu Y, Song J, Yang H, Zheng J, Huang Y, Li X, Liu C, Zhang Y, Ma Z, Yang Y (2012) Identification of a resistance gene bls1 to bacterial leaf streak in wild rice Oryza rufipogon Griff. J Integr Agric 11:962–969. https://doi.org/10.1016/S2095-3119(12)60087-2
Hu K, Cao J, Zhang J, Xia F, Ke Y, Zhang H, Xie W, Liu H, Cui Y, Cao Y, Sun X, Xiao J, Li X, Zhang WS (2017) Improvement of multiple agronomic traits by a disease resistance gene via cell wall reinforcement. Nat Plants 3:17009. https://doi.org/10.1038/nplants.2017.9
Hunter JD (2007) Matplotlib: a 2D graphics environment. Comput Sci Eng 9:90–95. https://doi.org/10.1109/MCSE.2007.55
Hutin M, Sabot F, Ghesquière A, Koebnik R, Szurek B (2015) A knowledge-based molecular screen uncovers a broad-spectrum OsSWEET14 resistance allele to bacterial blight from wild rice. Plant J 84:694–703. https://doi.org/10.1111/tpj.13042
Ishikawa K, Yamaguchi K, Sakamoto K, Yoshimura S, Inoue K, Tsuge S, Kojima C, Kawasaki T (2014) Bacterial effector modulation of host E3 ligase activity suppresses PAMP-triggered immunity in rice. Nat Commun 5:5430. https://doi.org/10.1038/ncomms6430
Iyer AS, McCouch SR (2004) The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol Plant Microbe Interact 17:348–1354. https://doi.org/10.1094/MPMI.2004.17.12.1348
Ji C, Ji Z, Liu B, Cheng H, Liu H, Liu S, Yang B, Chen G (2020) Xa1 allelic R genes activate rice blight resistance suppressed by interfering TAL effectors. Plant Commun 1:100087. https://doi.org/10.1016/j.xplc.2020.100087
Jiang N, Yan J, Liang Y, Shi Y, He Z, Wu Y, Zeng Q, Liu X, Peng J (2020) Resistance genes and their interactions with bacterial blight/leaf streak pathogens (Xanthomonas oryzae) in rice (Oryza sativa L.)—an updated review. Rice 13:3. https://doi.org/10.1186/s12284-019-0358-y
Kang H, Wang Y, Peng S, Zhang Y, Xiao Y, Wang D, Qu S, Li Z, Yan S, Wang Z, Liu W, Ning Y, Korniliev P, Leung H, Mezey J, McCouch SR, Wang GL (2016) Dissection of the genetic architecture of rice resistance to the blast fungus Magnaporthe oryzae. Mol Plant Pathol 17:959–972. https://doi.org/10.1111/mpp.12340
Kauffman HE, Reddy APK, Hsien SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis Rep 57:537–541
Khush GS (2005) What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol 59:1–6. https://doi.org/10.1007/s11103-005-2159-5
Kim SM, Reinke RF (2019) A novel resistance gene for bacterial blight in rice, Xa43(t) identified by GWAS, confirmed by QTL mapping using a bi-parental population. PLoS ONE 14:e0211775. https://doi.org/10.1371/journal.pone.0211775
Kim SA, Cho CS, Kim SR, Bull SB, Yoo YJ (2018) A new haplotype block detection method for dense genome sequencing data based on interval graph modeling of clusters of highly correlated SNPs. Bioinformatics 34:388–397. https://doi.org/10.1093/bioinformatics/btx609
Lander E, Kruglyak L (1995) Genetic dissection of complex traits guidelines for interpreting and reporting linkage results. Nat Genet 11:241–247. https://doi.org/10.1038/ng1195-241
Li C, Su P, Wang D, Peng S, Chen Y, Chen J, Tan X, Zhang D, Wang GL, Liu Y (2018) Dissection of the genetic architecture of rice resistance to Xanthomonas oryzae pv. oryzae using a genomewide association study. J Plant Pathol 166:470–476. https://doi.org/10.1111/jph.12705
Li C, Wang D, Peng S, Chen Y, Su P, Chen J, Zheng L, Tan X, Liu J, Xiao Y, Kang H, Zhang D, Wang GL, Liu Y (2019) Genome-wide association mapping of resistance against rice blast strains in South China and identification of a new Pik allele. Rice 12:47. https://doi.org/10.1186/s12284-019-0309-7
Lin HA, Chen SY, Chang FY, Tung CW, Chen YC, Shen WC, Chen RS, Wu CW, Chung CL (2018) Genome-wide association study of rice genes and loci conferring resistance to Magnaporthe oryzae isolates from Taiwan. Bot Stud 59:32. https://doi.org/10.1186/s40529-018-0248-4
Liu H, Yang W, Hu B, Liu F (2007) Virulence analysis and race classification of Xanthomonas oryzae pv. oryzae in China. J Phytopathol 155:129–135. https://doi.org/10.1111/j.1439-0434.2007.01197.x
Liu Q, Yuan M, Zhou Y, Li X, Xiao J, Wang S (2011) A paralog of the MtN3/salivafamily recessively confers race-specific resistance to Xanthomonas oryzae in rice. Plant Cell Environ 34:1958–1969. https://doi.org/10.1111/j.1365-3040.2011.02391.x
Liu M, Kang H, Xu Y, Peng Y, Wang D, Gao L, Wang X, Ning Y, Wu J, Liu W, Li C, Liu B, Wang GL (2020) Genome-wide association study identifies an NLR gene that confers partial resistance to Magnaporthe oryzae in rice. Plant Biotechnol J 18:1376–1383. https://doi.org/10.1111/pbi.13300
Long W, Yuan Z, Fan F, Dong D, Pan G, Sun H, Zhang Z, Li N, Li S (2020) Genome-wide association analysis of resistance to rice false smut. Mol Breed 40:46. https://doi.org/10.1007/s11032-020-01130-y
Lu Q, Wang C, Niu X, Zhang M, Xu Q, Feng Y, Yang Y, Wang S, Yuan X, Yu H, Wang Y, Wei X (2019) Detecting novel loci underlying rice blast resistance by integrating a genome-wide association study and RNA sequencing. Mol Breed 39:81. https://doi.org/10.1007/s11032-019-0989-0
Luo D, Huguet-Tapia JC, Raborn RT, White FF, Brendel VP, Yang B (2021) The Xa7 resistance gene guards the susceptibility gene SWEET14 of rice against exploitation by bacterial blight pathogen. Plant Commun. https://doi.org/10.1016/j.xplc.2021.100164
Mansueto L, Fuentes RR, Borja FN, Detras J, Abriol-santos JM, Chebotarov D, Sanciangco M, Palis K, Copetti D, Poliakov A, Dubchak I, Solovyev V, Wing RA, Hamilton RS, Mauleon R, McNally KL, Alexandrov N (2017) Rice SNP-seek database update: new SNPs, indels, and queries. Nucleic Acids Res 45:D1075–D1081. https://doi.org/10.1093/nar/gkw1135
Meng X, Zhao W, Lin R, Wang M, Peng Y (2006) Molecular cloning and characterization of a rice blast-inducible RING-H2 type Zinc finger gene. DNA Seq 17:41–48. https://doi.org/10.1080/10425170500476509
Mgonja EM, Balimponya EG, Kang H, Bellizzi M, Park CH, Li Y, Mabagala R, Sneller C, Correll J, Opiyo S, Talbot NJ, Mitchell T, Wang GL (2016) Genome-wide association mapping of rice resistance genes against Magnaporthe oryzae isolates from four african countries. Phytopathology 106:1359–1365. https://doi.org/10.1094/PHYTO-01-16-0028-R
Miao L, Wang C, Zheng C, Che J, Gao Y, Wen Y, Li G, Zhao K (2010) Molecular mapping of a new gene for resistance to rice bacterial blight. Agric Sci China 43:3051–3058
Neelam K, Mahajan R, Gupta V, Bhatia D, Gill BK, Komal R, Lore JS, Mangat GS, Singh K (2020) High-resolution genetic mapping of a novel bacterial blight resistance gene xa-45(t) identified from Oryza glaberrima and transferred to Oryza sativa. Theor Appl Genet 133:1–17. https://doi.org/10.1007/s00122-019-03501-2
Niño-Liu DO, Ronald PC, Bogdanove AJ (2006) Xanthomonas oryzae pathovars: model pathogens of a model crop. Mol Plant Pathol 7:303–324. https://doi.org/10.1111/j.1364-3703.2006.00344.x
Nordborg M, Tavaré S (2002) Linkage disequilibrium: what history has to tell us. Trends Genet 18:83–90. https://doi.org/10.1016/S0168-9525(02)02557-X
Ou SH (1985) Rice diseases. Commonwealth Agricultural Bureau, Great Britain
Patterson N, Price AL, Reich D (2006) Population structure and eigenanalysis. PLoS Genet 2:e190. https://doi.org/10.1371/journal.pgen.0020190
Raboin LM, Ballini E, Tharreau D, Ramanantsoanirina A, Frouin J, Courtois B, Ahmadi N (2016) Association mapping of resistance to rice blast in upland field conditions. Rice 9:59. https://doi.org/10.1186/s12284-016-0131-4
Read AC, Hutin M, Moscou MJ, Rinaldi FC, Bogdanove AJ (2020) Cloning of the rice Xo1 resistance gene and interaction of the Xo1 protein with the defense-suppressing Xanthomonas effector Tal2h. Mol Plant Microbe Int 33:1189–1195. https://doi.org/10.1094/MPMI-05-20-0131-SC
Shinada H, Yamamoto T, Sato H, Yamamoto E, Hori K, Yonemaru J, Sato T, Fujino K (2015) Quantitative trait loci for rice blast resistance detected in a local rice breeding population by genome-wide association mapping. Breed Sci 65:388–395. https://doi.org/10.1270/jsbbs.65.388
Sidhu JS, Khush GS, Mew TW (1978) Genetic analysis of bacterial blight resistance in seventy-four cultivars of rice, Oryza sativa L. Theor Appl Genet 53:105–111. https://doi.org/10.1007/BF00272687
Song WY, Wang GL, Chen LL, Kim HS, Pi LY, Holsten T, Gardner J, Wang B, Zhai WX, Zhu LH, Fauquet C, Ronald P (1995) A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 270:1804–1806. https://doi.org/10.1111/j.1744-7909.2010.00978.x
Sun X, Cao Y, Yang Z, Xu C, Li X, Wang S, Zhang Q (2004) Xa26, a gene conferring resistance to Xanthomonas oryzae pv. oryzae in rice, encodes an LRR receptor kinase-like protein. Plant J 37:517–527. https://doi.org/10.1046/j.1365-313X.2003.01976.x
Tang D, Wu W, Li W, Lu H, Worland AJ (2000) Mapping of QTLs conferring resistance to bacterial leaf streak in rice. Theor Appl Genet 101:286–291. https://doi.org/10.1007/s001220051481
The 3,000 rice genomes project (2014) The 3,000 rice genomes project. GigaScience 3:7. https://doi.org/10.1186/2047-217X-3-7
Tian D, Wang J, Zeng X, Gu K, Qiu C, Yang X, Zhou Z, Goh M, Luo Y, Murata-Hori M, White FF, Yin Z (2014) The rice TAL effector-dependent resistance protein XA10 triggers cell death and calcium depletion in the endoplasmic reticulum. Plant Cell 26:497–515. https://doi.org/10.1105/tpc.113.119255
Triplett LR, Cohen SP, Heffelfinger C, Schmidt CL, Huerta AI, Tekete C, Verdier V, Bogdanove AJ, Leach JE (2016) A resistance locus in the American heirloom rice variety Carolina Gold Select is triggered by TAL effectors with diverse predicted targets and is effective against African strains of Xanthomonas oryzae pv. oryzicola. Plant J 87:472–483. https://doi.org/10.1111/tpj.13212
Vera-Cruz CMV, Bai J, Ona I, Leung H, Nelson RJ, Mew TW, Leach JE (2000) Predicting durability of a disease resistance gene based on an assessment of the fitness loss and epidemiological consequences of avirulence gene mutation. Proc Natl Acad Sci USA 97:13500–13505. https://doi.org/10.1073/pnas.250271997
Wang C, Tan M, Xu X, Wen G, Zhang D, Lin X (2003) Localizing the bacterial blight resistance gene, Xa22(t), to a 100-kilobase bacterial artificial chromosome. Phytopathology 93:1258–1262. https://doi.org/10.1094/PHYTO.2003.93.10.1258
Wang CL, Qi HX, Pan HJ, Li JB, Fan YL, Zhang Q, Zhao KJ (2005) EST-markers flanking the rice bacterial blight resistance gene Xa23 and their application in marker-assisted selection. Sci Agric Sin 38:1996–2001 (Chinese with English abstract)
Wang C, Yang Y, Yuan X, Xu Q, Feng Y, Yu H, Wang Y, Wei X (2014) Genome-wide association study of blast resistance in indica rice. BMC Plant Biol 14:6. https://doi.org/10.1186/s12870-014-0311-6
Wang C, Zhang X, Fan Y, Gao Y, Zhu Q, Zheng C, Qin T, Li Y, Che J, Zhang M, Yang B, Liu Y, Zhao K (2015a) XA23 is an executor R protein and confers broad-spectrum disease resistance in rice. Mol Plant 8:290–302. https://doi.org/10.1016/j.molp.2014.10.010
Wang X, Jia MH, Ghai P, Lee FN, Jia Y (2015b) Genome-wide association of rice blast disease resistance and yield-related components of rice. Mol Plant Microbe Int 28:1383–1392. https://doi.org/10.1094/MPMI-06-15-0131-R
Wang W, Mauleon R, Hu Z et al (2018) Genomic variation in 3010 diverse accessions of Asian cultivated rice. Nature 557:43–49. https://doi.org/10.1038/s41586-018-0063-9
Wang S, Xie X, Zhang Z, Guan H, Mao D, Wu W, Chen Z (2020) Fine mapping of qBlsr3d, a quantitative trait locus conferring resistance to bacterial leaf streak in rice. Crop Sci 60:1854–1862. https://doi.org/10.1002/csc2.20155
Wonni I, Djedatin G, Ouédraogo L, Verdier V (2015) Evaluation of rice germplasm against bacterial leaf streak disease reveals sources of resistance in African varieties. J Plant Pathol Microbiol 6:1–5. https://doi.org/10.4172/2157-7471.1000312
Xiang Y, Cao Y, Xu C, Li X, Wang S (2006) Xa3, conferring resistance for rice bacterial blight and encoding a receptor kinase-like protein, is the same as Xa26. Theor Appl Genet 113:1347–1355. https://doi.org/10.1007/s00122-006-0388-x
Xie X, Chen Z, Cao J, Guan H, Lin D, Li C, Lan T, Duan Y, Mao D, Wu W (2014) Toward the positional cloning of qBlsr5a, a QTL underlying resistance to bacterial leaf streak, using overlapping sub-CSSLs in rice. PLoS ONE 9:e95751. https://doi.org/10.1371/journal.pone.0095751
Xie X, Chen Z, Zhang B, Guan H, Zheng Y, Lan T, Zhang J, Qin M, Wu W (2019) Transcriptome analysis of xa5-mediated resistance to bacterial leaf streak in rice (Oryza sativa L.). bioRxiv. https://doi.org/10.1101/727115
Yamamoto E, Yonemaru J, Yamamoto T, Yano M (2012) OGRO: The overview of functionally characterized genes in rice online database. Rice 5:26. https://doi.org/10.1186/1939-8433-5-26
Yang B, Sugio A, White FF (2006) Os8N3 is a host disease-susceptibility gene for bacterial blight of rice. Proc Natl Acad Sci USA 103:10503–10508. https://doi.org/10.1073/pnas.0604088103
Yao W, Li G, Yu Y, Ouyang Y (2018) funRiceGenes dataset for comprehensive understanding and application of rice functional genes. Gigascience 7:1–9. https://doi.org/10.1093/gigascience/gix119
Yoshimura S, Yamanouchi U, Katayose Y, Toki S, Wang ZX, Kono I, Kurata N, Yano M, Iwata N, Sasaki T (1998) Expression of Xa1, a bacterial blight-resistance gene in rice, is induced by bacterial inoculation. Proc Natl Acad Sci USA 95:1663–1668. https://doi.org/10.1073/pnas.95.4.1663
Yu J, Buckler ES (2006) Genetic association mapping and genome organization of maize. Curr Opin Biotechnol 17:155–160. https://doi.org/10.1016/j.copbio.2006.02.003
Yuan M, Ke Y, Huang R, Ma L, Yang Z, Chu Z, Xiao J, Li X, Wang S (2016) A host basal transcription factor is a key component for infection of rice by TALE-carrying bacteria. Elife 5:e19605. https://doi.org/10.7554/eLife.19605
Zhang F, Wu ZC, Wang MM, Zhang F, Dingkuhn M, Xu JL, Zhou YL, Li ZK (2017) Genome-wide association analysis identifies resistance loci for bacterial blight in a diverse collection of indica rice germplasm. PLoS ONE 12:e0174598. https://doi.org/10.1371/journal.pone.0174598
Zhang M, Wei Z, Yuan X, Wang C, Wang S, Niu X, Xu X, Xu Q, Feng Y, Yu H, Wang Y, Zhu Z, Zhai R, Yang Y, Wei X (2020a) Genetic variation dissection of rice blast resistance using an indica population. Rice Sci 27:255–258. https://doi.org/10.1016/j.rsci.2020.05.001
Zhang B, Zhang H, Li F, Ouyang Y, Yuan M, Li X, Xiao J, Wang S (2020b) Multiple alleles encoding atypical NLRs with unique central tandem repeats (CTRs) in rice confer resistance to Xanthomonas oryzae pv. oryzae. Plant Commun 1:100088. https://doi.org/10.1016/j.xplc.2020.100088
Zhao B, Lin X, Poland J, Trick H, Leach J, Hulbert S (2005) A maize resistance gene functions against bacterial streak disease in rice. Proc Natl Acad Sci USA 102:15383–15388. https://doi.org/10.1073/pnas.0503023102
Zheng J, Li Y, Fang X (2005) Detection of QTL conferring resistance to bacterial leaf streak in rice chromosome 2 (O. sativa L. spp. indica). Sci Agric Sin 38:1923–1925
Zhou X, Stephens M (2012) Genome-wide efficient mixed-model analysis for association studies. Nat Genet 44:821–824. https://doi.org/10.1038/ng.2310
Zhou H, Li P, Xie W, Hussain S, Li Y, Xia D, Zhao H, Sun S, Chen J, Ye H, Hou J, Zhao D, Gao G, Zhang Q, Wang G, Lian X, Xiao J, Yu S, Li X, He Y (2017) Genome-wide association analyses reveal the genetic basis of stigma exsertion in rice. Mol Plant 10:634–644. https://doi.org/10.1016/j.molp.2017.01.001
Zhu D, Kang H, Li Z, Liu M, Zhu X, Wang Y, Wang D, Wang Z, Liu W, Wang GL (2016) A genome-wide association study of field resistance to Magnaporthe oryzae in rice. Rice 9:44. https://doi.org/10.1186/s12284-016-0116-3
Acknowledgements
This work was supported partially by Grants from the National Key Research and Development Program of China (2016YFD0101107, 2016ZX08001-002), Hunan Provincial Key Research and Development Program (2017NK2022), the National Natural Science Foundation of China (31171526), and the Hu-Xiang High Level Talents Recruiting Program.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, N., Fu, J., Zeng, Q. et al. Genome-wide association mapping for resistance to bacterial blight and bacterial leaf streak in rice. Planta 253, 94 (2021). https://doi.org/10.1007/s00425-021-03612-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-021-03612-5