Abstract
Main conclusion
This article provides an overview of the interactions between Phytophthora effectors and plant immune system components, which form a cross-linked complex network that regulates plant pathogen resistance.
Abstract
Pathogens secrete numerous effector proteins into plants to promote infections. Several Phytophthora species (e.g., P. infestans, P. ramorum, P. sojae, P. capsici, P. cinnamomi, and P. parasitica) are notorious pathogens that are extremely damaging to susceptible plants. Analyses of genomic data revealed that Phytophthora species produce a large group of effector proteins, which are critical for pathogenesis. And, the targets and functions of many identified Phytophthora effectors have been investigated. Phytophthora effectors can affect various aspects of plant immune systems, including plant cell proteases, phytohormones, RNAs, the MAPK pathway, catalase, the ubiquitin proteasome pathway, the endoplasmic reticulum, NB-LRR proteins, and the cell membrane. Clarifying the effector–plant interactions is important for unravelling the functions of Phytophthora effectors during pathogenesis. In this article, we review the effectors identified in recent decades and provide an overview of the effector-directed regulatory network in plants following infections by Phytophthora species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants are constantly attacked by pathogens during their growth and development. Unlike animals, plants are sessile organisms unable to physically escape from pathogens. Consequently, they have evolved an advanced and sophisticated immune system to perceive pathogens and prevent infections. The plant immune system consists of two related layers: microbe- and pathogen-associated molecular pattern (MAMP and PAMP)-triggered immunity (PTI) and effector-triggered immunity (ETI) (Jones and Dangl 2006; Dodds and Rathjen 2010).
In the first layer of immunity, the pattern-recognition receptors (PRRs) on the plant cell membrane perceive PAMPs/MAMPs and activate PTI, which can temporarily prevent further pathogen infections (Jones and Dangl 2006; Couto and Zipfel 2016a, b). A number of PAMPs/MAMPs with conserved structures, which are common in microbes but not in host plants, have been identified in bacterial, fungal, and oomycete pathogens, including a bacterial flagellum peptide comprising 22 conserved amino acids (flg22) (Felix et al. 1999; Taguchi et al. 2003; Chinchilla et al. 2006), bacterial elongation factor Tu (EF-Tu) (Zipfel et al. 2006), Nep1-like proteins (NLPs) (Fellbrich et al. 2002; Qutob et al. 2006), cellulose-binding elicitor lectins (CBELs) (Gaulin et al. 2006), INF1 (Heese et al. 2007; Kamoun et al. 1997, 1998), and the glycoside hydrolase protein PsXEG1 (Ma et al. 2017).
In the second layer of immunity, pathogens that have penetrated plant cells secrete effector molecules that interfere with or inhibit PTI to enhance infections (Jones and Dangl 2006; Win et al. 2012). Bacteria use the type III secretion system (T3SS) to directly inject effectors into plant cells, whereas fungi and oomycetes introduce effectors into plant cells through appressoria or haustoria (Galan et al. 2014; Panstruga and Dodds 2009; Wang et al. 2017, 2018). Effectors can be categorized as apoplastic or cytoplasmic depending on their location in plants (Whisson et al. 2007). Apoplastic effectors accumulate in the intercellular spaces and interact with extracellular defense-related factors, whereas cytoplasmic effectors are transported into the cytoplasm via known and unknown mechanisms and are localized in various plant subcellular regions (Sharpee and Dean 2016).
To decrease their susceptibility to pathogens, plants have evolved resistance proteins (R proteins) that directly or indirectly interact with effectors and usually induce a hypersensitive response (HR) (Boller and He 2009). The HR, which is a form of programmed cell death (PCD), is often associated with the resistance of plants to pathogens at the infection site, and prevents pathogenic microorganisms from further infecting the plant (Jones and Dangl 2006; Dangl et al. 2013). The main R proteins are nucleotide-binding leucine-rich repeat (NB-LRR) proteins (Lee and Yeom 2015).
Large family of Phytophthora effectors
Phytophthora is a genus in the class Oomycota (Stramenopila) (Beakes et al. 2012). Because of similarities in morphological structures, such as mycelia and spores, oomycetes were considered to be fungi. However, a phylogenetic analysis revealed oomycetes are more closely related to diatoms and brown algae (Thines et al. 2014). Approximately 60% of oomycetes are plant pathogens, including the species in the genus Phytophthora (Thines et al. 2014). A previous study confirmed there are more than 100 Phytophthora species, several of which are notorious pathogens that can severely damage crops, including vegetables, and forests (Érsek and Ribeiro 2010). For example, Phytophthora infestans is responsible for late blight in potato and tomato, which annually cause tremendous yield losses worldwide (Zadoks 2008). Additionally, Phytophthora ramorum is the causative agent of sudden oak death (SOD), which kills oak trees and other tree species, resulting in considerable economic losses in North America and Europe (Grünwald et al. 2008). Phytophthora sojae, Phytophthora parasitica, and Phytophthora capsici cause major losses to soybean, tobacco, and pepper production, respectively, and consequently substantial economic losses (Tyler 2007; Meng et al. 2014; Hausbeck and Lamour 2004). Phytophthora species exhibit a hemibiotrophic lifestyle, initially existing as biotrophs before becoming necrotrophs (Fawke et al. 2015). The jasmonate (JA) and salicylate (SA) signaling pathways in plants mediate a universal defense response. Specifically, the SA and JA pathways generally initiate the resistance to biotrophic and necrotrophic pathogens, respectively (Zhang et al. 2017). An earlier investigation determined that SA contributes to the resistance of plants to P. sojae (Liu et al. 2014).
The genomes of P. infestans, P. sojae, P. ramorum, P. parasitica, and P. capsici have been fully sequenced (Haas et al. 2009; Tyler et al. 2006; Liu et al. 2016; Lamour et al. 2012) (Table 1). The generated genomic data suggest that Phytophthora species produce many effectors (Table 1), which have been divided into two main types, namely, RxLR and CRN effectors (Haas et al. 2009). The N terminus of RxLR effectors contains a conserved arginine–any amino acid–leucine–arginine (R–x–L–R) motif. This motif is usually followed by a small, conserved glutamic acid–glutamic acid–arginine (EER) domain. The conserved RxLR-dEER domain is closely related to the transport of effectors into cells (Jiang et al. 2008; Win and Kamoun 2007). The CRN effectors (crinkling- and necrosis-inducing proteins), which were originally identified in P. infestans, have a highly conserved N-terminal LFLAK domain (Haas et al. 2009). The C-terminal regions of RxLR and CRN effectors are relatively diverse.
The considerable abundance of effectors produced by Phytophthora species is important for overcoming plant immune systems. In recent decades, many Phytophthora effectors have been identified, and their targets and functions in plant cells have been analyzed. Researchers have confirmed that Phytophthora effectors influence various aspects of plant immune systems (Dou and Zhou 2012; Whisson et al. 2016; Wang et al. 2019). The effectors and plant immune system components form a complex network that modulates plant resistance. We herein review the Phytophthora effector proteins identified in recent decades regarding their current known plant targets and the relationships among these targets. Thus, we provide an overview of the plant–pathogen interaction network involving multiple plant immune system components. This review is relevant for characterizing the mechanisms underlying pathogenicity and plant resistance.
Manipulating cell protease-mediated immunity
Proteases, which can catalyze the hydrolysis of their target proteins, have been detected in almost all plants, animals, and microorganisms (Mosolov and Valueva 2005). Many resistance-related proteases are present in plant apoplasts and are important components of plant immune systems (Habib and Majid 2007). For example, P69B is a tomato subtilisin-like apoplastic serine protease, whereas Phytophthora-inhibited protease 1 (PIP1), Rcr3pim, and C14 are papain-like extracellular apoplastic cysteine proteases (Tornero et al. 1997; Jorda et al. 1999; Kruger et al. 2002). All of these proteases are important for plant resistance responses, and both SA and BTH (i.e., SA analog) can induce P69B expression.
To promote infections, Phytophthora pathogens secrete effectors that inhibit protease activities. The P. infestans effectors EPI1 (Tian et al. 2004), EPI10 (Tian et al. 2005), EPIC2B (Tian et al. 2007; Song et al. 2009; Kaschani et al. 2010; Kaschani and Van Der Hoorn 2011), EPIC1 (Kaschani et al. 2010; Kaschani and Van Der Hoorn 2011), and AVRblb2 (Bozkurt et al. 2011) function as protease inhibitors. Additionally, EPI1 and EPI10 interact with tomato P69B to inhibit its function (Tian et al. 2004, 2005). Moreover, EPI1 and EPI10, which have two and three Kazal-like domains, respectively, are structurally diverse, but they both target the same plant protease, P69B, implying that P69B is an important target of P. infestans (Tian et al. 2004, 2005). In contrast, AVRblb2, which mainly accumulates around haustoria, targets C14 and specifically prevents its secretion into the apoplast (Bozkurt et al. 2011), whereas Rcr3pim helps plants defend against P. infestans strains producing EPIC1 and EPIC2B (Tian et al. 2007; Song et al. 2009; Kaschani et al. 2010; Kaschani and Van Der Hoorn 2011). Interestingly, EPIC2B can interact with and inhibit three proteases (i.e., PIP1, Rcr3pim, and C14), suggesting it considerably promotes P. infestans infections. The P. sojae effector glucanase inhibitor protein 1 (GIP1) can inhibit the release of glucan elicitors from the P. sojae cell wall by forming a complex with endoglucanases (Rose et al. 2002). The above-mentioned examples indicate that various effectors from the same pathogen can target the same plant immune system components. For example, the P. infestans effectors EPI1 and EPI10 both affect P69B, whereas EPIC1 and EPIC2B both target Rcr3, and EPIC2B and AVRb1b2 both target C14. However, the same effectors can also target different components. For instance, EPIC2B can target three immune system components, namely PIP1, Rcr3, and C14, which suggests there is a link between PIP1 and Rcr3. The relationships among these effectors are presented in Fig. 1a.
Phytophthora effectors target plant immune system components. a Targeting of plant components by Phytophthora effectors leading to modifications in SA, H2O2, and callose contents. Interaction indicates a direct interaction between effectors and the targets. Physical association indicates a close correlation, but no direct interaction observed between the effectors and the plant targets. b Interaction network of AVR3a, PsAvh262 and PsCRN108 with their targets, which involves SGT1, HSP90, BiPs, and the HSP promoters. c Diagram of the interactions between AVR2 and BSL1 as well as R2
Manipulating phytohormone-mediated immunity
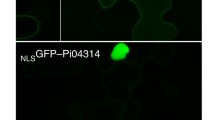
Phytohormone signaling pathways play important roles in plant disease resistance (Robert-Seilaniantz et al. 2011). Specifically, SA, jasmonic acid (JA), ethylene (ET), and auxin form a complex signaling network that regulates plant resistance to pathogenic microbes (Pieterse et al. 2012; Thaler et al. 2012). The P. sojae effector PsIsc1 can hydrolyze host isochorismate and modulate SA metabolism, thereby decreasing SA levels in hosts and suppressing immunity (Liu et al. 2014). The P. infestans effector PexRD24 (also known as Pi04314) interacts with three protein phosphatase 1 catalytic (PP1c) isoforms required for disease development and induces their re-localization from the nucleolus to the nucleoplasm, which decreases JA and SA accumulation and suppresses host immunity (Boevink et al. 2016). Regarding P. parasitica, penetration-specific effector 1 (PSE1), whose transcript is transiently accumulated during the penetration of host roots, can modulate auxin contents at the penetration points and facilitate infections (Evangelisti et al. 2013). In Arabidopsis thaliana, PSE1 also has regulatory functions and is associated with the low auxin concentrations at the root apex (Evangelisti et al. 2013). The relationships among these effectors are presented in Fig. 1a.
A physical interaction network has been constructed for A. thaliana proteins and effector proteins from pathogens, including the Gram-negative bacterium Pseudomonas syringae and the obligate biotrophic oomycete Hyaloperonospora arabidopsidis (Mukhtar et al. 2011). According to this network, the effectors of these two pathogens strongly interact with JAZ proteins, which are important JA signaling components (Mukhtar et al. 2011). Like Phytophthora species, H. arabidopsidis is an oomycete. Accordingly, Phytophthora effector proteins may also influence the JA pathway.
Manipulating MAPK-mediated immunity
Mitogen-activated protein kinase (MAPK) cascades, which form a highly conserved signaling pathway in all eukaryotes, can transfer extracellular signals into cells via the protein phosphorylation and dephosphorylation catalyzed by MAPK kinases (MAPKKs) and MAPKK kinases (MAPKKKs) (Rodriguez et al. 2010; Melech-Bonfil and Sessa 2010). The MAPKs are crucial for regulating plant defensive responses to pathogens (Meng and Zhang 2013; Pitzschke et al. 2009). The flg22 bacterial flagellin peptide is a typical MAMP directly recognized by the plant protein FLAGELLIN SENSITIVE2 (FLS2), and this recognition activates the MAPK immunity pathway to induce the expression of numerous defense-related genes, including FRK1 (Chinchilla et al. 2006; Taguchi et al. 2003).
The P. infestans RXLR effector, PexRD2, interacts with the MAPKKKe kinase domain to suppress activity (King et al. 2014). The overexpression of PexRD2 or the silencing of MAPKKKe in Nicotiana benthamiana promotes infection. Eight suppressors of the early flg22-induced immune response (SFIs), which were identified as RXLR effectors in P. infestans, can alter various steps of the MAPK signaling pathway to suppress early-induced immunity (Zheng et al. 2014). A P. sojae RXLR effector, Avh331, manipulates the MAPK signaling pathway to promote the infection of A. thaliana and N. benthamiana. This process is accompanied by significantly decreased H2O2 accumulation and callose deposition (Cheng et al. 2012). The PsCRN63 of P. sojae functions downstream of MAPK cascades to suppress flg22-induced defense responses, such as FRK1 expression, and callose deposition (Li et al. 2016). The relationships among these effectors are presented in Fig. 1a.
Manipulating cell membrane-mediated immunity
The plasma membrane–cell wall continuum in plants is important for defense responses, and disrupted plasma membrane–cell wall adhesions may increase the susceptibility of plants to pathogens (Zhu et al. 1993; Mellersh and Heath 2001). The P. infestans in planta induced-O (IPI-O), which is an RXLR-dEER effector, can disrupt the plasma membrane–cell wall adhesions in A. thaliana through its RGD motif (arginine–glycine–aspartic acid) (Senchou et al. 2004; Gouget et al. 2006; Bouwmeester et al. 2011). However, when the IPI-O RGD motif is mutated to RGE or RGA, the adhesions remain intact. An 80-kDa RGD plasma membrane protein of A. thaliana and the legume-like lectin receptor kinase LecRK-I.9 can interact with IPI-O through the RGD motif. Additionally, LecRK-I.9 can directly or indirectly trigger increases in callose deposition (Fig. 1a). A P. sojae RxLR effector, Avh241, which localizes to the membrane and triggers cell death in multiple plant species (Yu et al. 2012), is essential for the full virulence of P. sojae and functions as a susceptibility inducer when produced in plant cells. However, it does not need to induce cell death to promote susceptibility, suggesting Avh241 interacts with the plant immune system in at least two different ways (Yu et al. 2012).
Manipulating RNA-mediated immunity
In eukaryotic organisms, RNA silencing is a universal gene regulatory mechanism that controls many biological processes (Baulcombe 2004; Sarkies and Miska 2014). Small RNAs comprising 20–30 nucleotides play a central role in this process. Plants produce two types of small RNAs, microRNAs (miRNAs) and small interfering RNAs (siRNAs), which help mediate the plant disease resistance system (Vance and Vaucheret 2001; Huang et al. 2016). Specifically, miRNAs are transcribed from endogenous MIR family genes, whereas siRNAs are derived from invading nucleic acids, such as viruses and transgenes, and from endogenous loci, such as repeats, transposable elements, and genes (Voinnet 2009).
Three Phytophthora effectors that regulate RNA silencing have been identified (PSR1, PSR2, and PiPSR2) (Qiao et al. 2013, 2015; Xiong et al. 2014). The PSR1 and PSR2 effectors of P. sojae suppress RNA silencing in plants by inhibiting the biogenesis of small RNAs (Qiao et al. 2013). Similarly, PiPSR2, which is a PSR2-like effector produced by P. infestans, also suppresses RNA silencing by inhibiting the biogenesis of small RNAs (Xiong et al. 2014). Additionally, PSR1 interacts with PSR1-interacting protein 1 (PINP1), which regulates the accumulation of both miRNAs and endogenous siRNAs in A. thaliana (Qiao et al. 2015). The PsAvr3c effector from P. sojae interacts with and stabilizes the soybean serine/lysine/arginine-rich proteins GmSKRPs, which contain plant spliceosome components and modulate host pre-mRNA splicing to regulate plant immunity (Huang et al. 2017). In P. infestans, Pi14054 is a novel candidate suppressor affecting the RNA silencing of plant hosts (Vetukuri et al. 2017). Moreover, the P. infestans RXLR effector Pi04089 interacts with StKRBP1, which is a predicted K-homology (KH) class putative RNA-binding protein in Solanum tuberosum, and this interaction occurs in the plant nucleus (Wang et al. 2015). Interestingly, PR1 expression is completely abolished in PsPSR2-expressing plants, which suggests that PsPSR2 interferes with the SA-dependent defense pathway during an infection by Phytophthora species (Xiong et al. 2014) (Fig. 1a).
Manipulating E3 ubiquitin ligase-mediated immunity
The ubiquitin proteasome pathway is the most important protein degradation pathway in eukaryotic organisms, and in plants, it regulates growth, development, and responses to abiotic and biotic stresses (Zhou and Zeng 2017). Ubiquitination involves a series of reactions catalyzed by the E1 ubiquitin-activating enzyme, E2 ubiquitin-binding enzyme, and E3 ubiquitin ligase (Callis 2014). The P. infestans effector AVR3a interacts with the U-box E3 ligase CMPG1 and inhibits its activity, thereby controlling the normal protein degradation system of plants (Bos et al. 2010). Two Avr3a alleles encode distinct 147-amino acid sequences, AVR3aKI and AVR3aEM, which differ in only three amino acids. Moreover, AVR3aKI strongly interacts with and stabilizes CMPG1, whereas AVR3aEM only weakly interacts with CMPG1 (Bos et al. 2010). According to previous reports, AVR3a also interacts with R3a, an intracellular NBS-LRR domain-containing protein. This interaction is dependent on the ubiquitin ligase-associated protein SGT1 and heat-shock protein 90 (HSP90), which can trigger R3a-dependent HR and suppress the cell death induced by the P. infestans elicitin INF1 (Armstrong et al. 2005; Bos et al. 2006) (Fig. 1b).
Manipulating catalase-mediated immunity
Hydrogen peroxide (H2O2), which is a reactive oxygen species (ROS), plays an important role in plant defense responses (Yoshioka et al. 2009; Petrov and Van Breusegem 2012). Catalases in cells convert H2O2 into water and oxygen (Li et al. 2013). The P. sojae effectors PsCRN63/115 manipulate plant PCD by interfering with catalases and perturbing H2O2 homeostasis (Zhang et al. 2015). Additionally, PsCRN63 also suppresses the flg22-induced expression of PTI marker genes and callose deposition (Liu et al. 2011; Li et al. 2016). The PsCRN70 effector of P. sojae, which localizes to the plant cell nucleus and functions as a universal suppressor of the cell death induced by many elicitors, may promote Phytophthora infections by suppressing H2O2 accumulation and the production of defense proteins, including the pathogenesis-related (PR) proteins PR1a and PR2b (markers for SA-dependent defense), ethylene response factor 1 (ERF1), and lipoxygenase (LOX; involved in JA synthesis) (Rajput et al. 2014). The relationships among these effectors are presented in Fig. 1a.
Manipulating endoplasmic reticulum-mediated immunity
The endoplasmic reticulum (ER) is an important site for the modification and folding of eukaryotic secretory and membrane proteins (Kim et al. 2008; Hetz 2012). Only correctly folded and modified proteins can be transported to the cytoplasm, nucleus, mitochondria, and other organelles to exert their biological functions (Hetz 2012). Protein folding in the ER is mainly mediated by molecular chaperones and cofactors (Saijo 2010). Binding immunoglobulin protein (BiP), which is a member of the large HSP70 family of molecular chaperones, is a high molecular weight chaperone in the ER (Bertolotti et al. 2000). The accumulation of unfolded or incorrectly folded proteins in the ER due to environmental stresses causes the cell to initiate an unfolded protein response (UPR) by regulating a series of downstream genes, such as those encoding molecular chaperones, to promote the correct folding of proteins and the degradation of improperly folded proteins (Saijo 2010). Four important UPR signaling pathways are present in A. thaliana, of which two are mediated by the bZIP family membrane-binding transcription factors bZIP28 and bZIP60, whereas the other two are mediated by the NAC (NAM/ATAF/CUC) family membrane-binding transcription factors NAC062 and NAC089 (Saijo 2010; Yang et al. 2014a, b). Previous research revealed that ER stress responses regulate plant immunity in several ways. For example, the A. thaliana HSP AtBiP2 is involved in the secretion of PR proteins (Wang et al. 2015), and the processing of the pattern-recognition receptor EFR requires BiP (Nekrasov et al. 2009). The fungus Piriformospora indica induces cell death by inhibiting the UPR-related pro-survival machinery and then activating the ER stress-mediated cell death machinery (Qiang et al. 2012). The UPR is crucial for inducing systemic acquired resistance (SAR) against bacterial pathogens as well as abiotic stress tolerance (Moreno et al. 2012).
Two NAC transcription factors, NTP1 and NTP2, are localized in the ER membrane and contribute to the resistance of tobacco against P. infestans. The P. infestans effector Pi03192 interacts with NTP1 and NTP2 and prevents their transmission from the ER into the nucleus, thereby decreasing the pathogenicity of P. infestans towards potato and N. benthamiana (McLellan et al. 2013). Additionally, PsAvh262 from P. sojae can stabilize BiPs and facilitate Phytophthora infections (Jing et al. 2016). The P. sojae CRN effector PsCRN108, which contains a helix–hairpin–helix (HhH) motif, targets HSP promoters in an HSE- and HhH-dependent manner and inhibits HSP expression in response to stress (Song et al. 2015). Moreover, PcAvr3a12 of P. capsici can directly target the ER-localized plant peptidyl-prolyl cis–trans isomerase (PPIase) and facilitate infections (Fan et al. 2018).
The PsCRN108 effector targets HSP promoters and inhibits HSP expression in response to stress, whereas Avh262 affects the ER of plants and interacts with BiPs. Furthermore, AVR3a interacts with R3a, while simultaneously associating with HSP90, which is one of the molecular chaperones of BiPs, and SGT1 (as described in a previous section). These observations imply there are some obvious correlations among AVR3a, PsCRN108, and Avh262 functions in plants (Fig. 1b).
Manipulating NB-LRR protein-mediated immunity
Hundreds of resistance (R) genes, which mainly encode NB-LRR proteins, are important components of plant immune systems (Eitas and Dangl 2010). On the basis of the encoded N-terminal domain, NB-LRR genes can be classified into two groups: toll/interleukin-1 receptor (TIR)–NB-LRRs (TNLs) and coiled-coil (CC)–NB-LRRs (CNLs). Additionally, the N-terminal TIR and CC domains are required for plant defense responses (Lee and Yeom 2015). The RB gene (also known as Rpi-blb1) from the wild potato species Solanum bulbocastanum encodes a CNL protein and mediates broad spectrum resistance to most pathogen strains by detecting IPI-O family effectors (Champouret et al. 2009). Moreover, IPI-O4 can interact with the CC domain of CNLs to suppress the immunity induced by RB (Chen et al. 2012). The P. infestans RXLR effector AVR2 is recognized by the NB-LRR protein R2, which requires the host phosphatase protein BSU-like protein 1 (BSL1) (Saunders et al. 2012) (Fig. 1c).
Manipulating other plant component-mediated immunity
The P. sojae effector Avr3b and the P. infestans effector CRN8 are the only effectors known to be enzymatically active in plant cells (Dong et al. 2011; van Damme et al. 2012). Specifically, Avr3b, an ADP-ribose/NADH pyrophosphorylase identified in soybean varieties containing the disease resistance gene Rps3b, functions as a Nudix hydrolase that impairs host immunity when delivered into the host cells. Moreover, the deletion of the Nudix motif abolishes the enzyme activity (Dong et al. 2011). The accumulation of ROS is significantly decreased when Avr3b is transiently produced in N. benthamiana (Dong et al. 2011) (Fig. 1a). The C-terminal of CRN8 is similar to a serine/threonine RD kinase domain, and its autophosphorylation is dependent on an intact catalytic site (van Damme et al. 2012). The functions of Avr3b and CRN8 indicate that plant pathogens can translocate biochemically active kinase effectors inside host cells.
The plant exocyst influences defense responses, and Sec5 is an exocyst subunit (Cole et al. 2005; Hála et al. 2008). The P. infestans RXLR effector AVR1 can interact with Sec5 and regulate callose deposition and PR1 expression (Du et al. 2015) (Fig. 1a). The PcCRN4 effector of P. capsici may inhibit host immunity by suppressing PR1b expression, ROS accumulation, and callose deposition in plants (Mafurah et al. 2015) (Fig. 1a).
Autophagy is a conserved catabolic pathway that sequesters unwanted cytosolic components in newly formed double membrane vesicles (autophagosomes) to direct them to the cell lytic compartment (He et al. 2009). Autophagy has vital roles in plant defense responses. The P. infestans effector PexRD54 can interact with ATG8CL by outcompeting a plant autophagy cargo receptor that would otherwise bind to ATG8CL (Dagdas et al. 2016). By preventing the cargo receptor from interacting with ATG8CL, PexRD54 increases the susceptibility of the plant to an infection by P. infestans. Specifically, autophagy is implicated in the accumulation of defense hormones.
The P. sojae effector Avh238 is a virulence factor that also triggers plant innate immunity, but its target and activity in plants remain unknown (Yang et al. 2017).
Conclusions
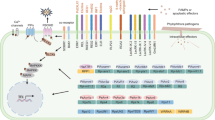
Thousands of effectors represent powerful weapons used by Phytophthora pathogens to promote infections. We herein reviewed the Phytophthora effectors identified in recent decades and described their known targets and modes of action in plants (Table 2, Fig. 2). Although some effectors affect different plant parts, they are ultimately accompanied by changes to SA, callose, and H2O2 contents (Fig. 1). For example, the effects of EPIC2, AVR1, EPI1, EPI10, and PsPSR2 on plants include changes to SA contents, while PsIsc1, PexRD24, and PSE1 directly affect plant hormones, including SA and auxin. Moreover, PsCRN63 and CRN4 influence the accumulation of SA, callose, and H2O2, and PsCRN70 affects the accumulation of SA and H2O2. Additionally, PsCRN115 regulates H2O2 homeostasis, whereas IPI-O affects the accumulation of callose. These results indicate that SA, callose, and H2O2 are important stress-resistance substances in plants, which is consistent with the results of previous studies (Luna et al. 2011; Jwa and Hwang 2017). Furthermore, SA can regulate the metabolism of callose and H2O2 (Nishimura et al. 2003; Dong et al. 2008; Chen et al. 1993), and a correlation between callose metabolism and H2O2 metabolism has been confirmed (Luna et al. 2011).
Proposed model of Phytophthora effectors acting on various plant immune system components. Phytophthora species secrete a large number of effectors acting on various plant immune system components during the infection processes. The effectors may manipulate the phytohormone (e.g., Aux, JA, SA) level or the apoplastic proteases (e.g., P69B, C14, PIP1 and Rcr3), nucleus protein phosphatase (e.g., PP1c), etc. (as indicated with orange arrows). They could also affect the cell signaling through manipulation of the MAPK cascade or the R protein-mediated responses. And, the endoplasmic reticulum (ER) and ER-localized proteins are also the targets of effectors, which may disturb the protein interacting network (e.g., BiPs) or the movement of ER-localized proteins (e.g., NTPs). Furthermore, the effectors could regulate the transcription factors that control the pathogen responses by manipulating the correlated proteasome pathway ligase (e.g., E3) or direct binding their target promoters, and interact with components (e.g., PINP1) that mediate the processing of small RNA or miRNA to affect the production of pathogen responsive siRNAs
However, these physiological phenomena are insufficient to reveal the regulatory roles of effectors related to plant immunity because the functions of effectors in plants are extremely complex. For instance, the P. infestans effector AVR3a interacts with the E3 ligase CMPG1 to manipulate plant immunity, whereas the P. syringae T3SS AvrPtoB exhibits E3 ubiquitin ligase activity, which is required to suppress plant innate immunity (Abramovitch et al. 2006; Rosebrock et al. 2007). In addition, the XopL effector of Xanthomonas campestris pv. vesicatoria can suppress plant defenses in an E3 ubiquitin ligase activity-dependent manner (Singer et al. 2013). Therefore, E3 ubiquitin ligase is a ubiquitous component of plant immune systems, and the effectors of bacteria, fungi and oomycetes affect the E3 ubiquitin ligase-mediated system to promote infections. The MAPK pathway-mediated immune system is influenced by the P. syringae effectors HopAI1 and HopF2 and the Phytophthora effectors PexRD2, Avh331, and PsCRN63 (Zhang et al. 2007; Wang et al. 2010), implying that the effectors of bacteria and oomycetes act on the same plant immune system.
Plants contain distinct conserved immune systems, with certain associations existing among the immune system components. Numerous interactions between the effectors and immune system components form a complex cross-linked network that regulates plant stress resistance, but the network has only been preliminarily characterized. The gradual elucidation of the entire network is an important objective for future studies and will greatly aid investigations regarding pathogenesis and plant disease resistance.
Author contribution statement
WW designed the outline of the article, composed the manuscript, and prepared the figures and tables. FJ compiled and reviewed the literature and edited the manuscript.
References
Abramovitch RB, Janjusevic R, Stebbins CE, Martin GB (2006) Type III effector AvrPtoB requires intrinsic E3 ubiquitin ligase activity to suppress plant cell death and immunity. Proc Natl Acad Sci USA 103:2851–2856
Armstrong MR, Whisson SC, Pritchard L, Bos JIB, Venter E, Avrova AO, Rehmany AP, Brooks K, Bohme U, Cherevach I, Hamlin N, White B, Fraser A, Lord A, Quail MA, Churcher C, Hall N, Berriman M, Huang S, Kamoun S, Beynon JL, Birch PRJ (2005) An ancestral oomycete locus contains late blight avirulence gene Avr3a, encoding a protein that is recognized in the host cytoplasm. Proc Natl Acad Sci USA 102:7766–7771
Baulcombe D (2004) RNA silencing in plants. Nature 431:356–363
Beakes GW, Glockling SL, Sekimoto S (2012) The evolutionary phylogeny of the oomycete “fungi”. Protoplasma 249:3–19
Bertolotti A, Zhang Y, Hendershot LM, Harding HP, Ron D (2000) Dynamic interaction of BiP and ER stress transducers in the unfolded-protein response. Nat Cell Biol 2:326–332
Boevink PC, Wang XD, McLellan H, He Q, Naqvi S, Armstrong MR, Zhang W, Hein I, Gilroy EM, Tian ZD, Birch PRJ (2016) A Phytophthora infestans RXLR effector targets plant PP1c isoforms that promote late blight disease. Nat Commun 7:10311
Boller T, He SY (2009) Innate immunity in plants: an arms race between pattern recognition receptors in plants and effectors in microbial pathogens. Science 324:742–744
Bos JIB, Kanneganti TD, Young C, Cakir C, Huitema E, Win J, Armstrong MR, Birch PRJ, Kamoun S (2006) The C-terminal half of Phytophthora infestans RXLR effector AVR3a is sufficient to trigger R3a-mediated hypersensitivity and suppress INF1-induced cell death in Nicotiana benthamiana. Plant J 48:165–176
Bos JIB, Armstrong MR, Gilroy EM, Boevink PC, Hein I, Taylor RM, Tian ZD, Engelhardt S, Vetukuri RR, Harrower B, Dixelius C, Bryan G, Sadanandom A, Whisson SC, Kamoun S, Birch PRJ (2010) Phytophthora infestans effector AVR3a is essential for virulence and manipulates plant immunity by stabilizing host E3 ligase CMPG1. Proc Natl Acad Sci USA 107:9909–9914
Bouwmeester K, de Sain M, Weide R, Gouget A, Klamer S, Canut H, Govers F (2011) The lectin receptor kinase LecRK-I.9 is a novel Phytophthora resistance component and a potential host target for a RXLR effector. PLoS Pathog 7:e1001327
Bozkurt TO, Schornack S, Win J, Shindo T, Ilyas M, Oliva R, Cano LM, Jones AME, Huitema E, Hoorn RALVD, Kamoun S (2011) Phytophthora infestans effector AVRblb2 prevents secretion of a plant immune protease at the haustorial interface. Proc Natl Acad Sci USA 108:20832–20837
Callis J (2014) The ubiquitination machinery of the ubiquitin system. Arabidopsis Book 12:e0174
Champouret N, Bouwmeester K, Rietman H, van der Lee T, Maliepaard C, Maliepaard C, Heupink A, van de Vondervoort PJ, Jacobsen E, Visser RG, van der Vossen EA, Govers F, Vleeshouwers VG (2009) Phytophthora infestans isolates lacking class I ipiO variants are virulent on Rpi-blb1 potato. Mol Plant Microbe Interact 22:1535–1545
Chen ZX, Silva H, Klessig DF (1993) Active oxygen species in the induction of plant systemic acquired resistance by salicylic-acid. Science 262:1883–1886
Chen Y, Liu ZY, Halterman DA (2012) Molecular determinants of resistance activation and suppression by Phytophthora infestans effector IPI-O. Plant Pathog 8:e1002595
Cheng BP, Yu XL, Ma ZC, Dong SM, Dou DL, Wang YC, Zheng XB (2012) Phytophthora sojae effector Avh331 suppresses the plant defence response by disturbing the MAPK signalling pathway. Physiol Mol Plant Pathol 77:1–9
Chinchilla D, Bauer Z, Regenass M, Boller T, Felix G (2006) The Arabidopsis receptor kinase FLS2 binds flg22 and determines the specificity of flagellin perception. Plant Cell 18:465–476
Cole RA, Synek L, Zarsky V, Fowler JE (2005) SEC8, A subunit of the putative Arabidopsis exocyst complex, facilitates pollen germination and competitive pollen tube growth. Plant Physiol 138:2005–2018
Couto D, Zipfel C (2016a) Regulation of pattern recognition receptor signalling in plants. Immunology 16:537–552
Couto D, Zipfel C (2016b) Regulation of pattern recognition receptor signalling in plants. Immunology 16:537–552
Dagdas YF, Belhaj K, Maqboo A, Chaparro-Garcia A, Pandey P, Petre B (2016) An effector of the Irish potato famine pathogen antagonizes a host autophagy cargo receptor. eLife 5:e10856
Dangl JL, Horvath DM, Staskawicz BJ (2013) Pivoting the plant immune system from dissection to deployment. Science 341:746–751
Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant–pathogen interactions. Nat Rev Genet 11:539–548
Dong X, Hong Z, Chatterjee J, Kim S, Verma DPS (2008) Expression of callose synthase genes and its connection with Npr1 signaling pathway during pathogen infection. Planta 229:87–98
Dong SM, Yin WX, Kong GH, Yang XY, Qutob D, Chen QH, Kale SD, Sui YY, Zhang ZG, Dou DL, Zheng XB, Gijzen M, Tyler BM, Wang YC (2011) Phytophthora sojae avirulence effector Avr3b is a secreted NADH and ADP-ribose Pyrophosphorylase that modulates plant immunity. PLoS Pathog 7:e1002353
Dou D, Zhou JM (2012) Phytopathogen effectors subverting host immunity: different foes, similar battleground. Cell Host Microbe 12:484–495
Du Y, Mpina MH, Birch PRJ, Bouwmeester K, Govers F (2015) Phytophthora infestans RXLR effector AVR1 interacts with exocyst component Sec5 to manipulate plant immunity. Plant Physiol 169:1975–1990
Eitas TK, Dangl JL (2010) NB-LRR proteins: pairs, pieces, perception, partners, and pathways. Curr Opin Plant Biol 13:472–477
Érsek T, Ribeiro OK (2010) An annotated list of new Phytophthora species described post 1996. Acta Phytopathol Hung 45:251–266
Evangelisti E, Govetto B, Minet-Kebdani N, Kuhn ML, Attard A, Ponchet M, Panabieres F, Gourgues M (2013) The Phytophthora parasitica RXLR effector penetration-specific effector 1 favours Arabidopsis thaliana infection by interfering with auxin physiology. New Phytol 199:476–489
Fan GJ, Yang Y, Li TT, Lu WQ, Du Y, Qiang XY, Wen QJ, Shan WX (2018) A Phytophthora capsici RXLR effector targets and inhibits a plant PPIase to suppress endoplasmic reticulum-mediated immunity. Mol Plant 11:1067–1083
Fawke S, Doumane M, Schornack S (2015) Oomycete interactions with plants: infection strategies and resistance principles. Microbiol Mol Biol Rev 79:263–280
Felix G, Duran JD, Volko S, Boller T (1999) Plants have a sensitive perception system for the most conserved domain of bacterial flagellin. Plant J 18:265–276
Fellbrich G, Romanski A, Varet A, Blume B, Brunner F, Engelhardt S, Felix G, Kemmerling B, Krzymowska M, Nürnberger T (2002) NPP1, a Phytophthora-associated trigger of plant defense in parsley and Arabidopsis. Plant J 32:375–390
Galan JE, Lara-Tejero M, Marlovits TC, Wagner S (2014) Bacterial type III secretion systems: specialized nanomachines for protein delivery into target cells. Annu Rev Microbiol 68:415–438
Gaulin E, Dramé N, Lafitte C, Torto-Alalibo T, Martinez Y, Torregrosa C (2006) Cellulose binding domains of a Phytophthora cell wall protein are novel pathogen-associated molecular patterns. Plant Cell 18:1766–1777
Gouget A, Senchou V, Govers F, Sanson A, Barre A, Rouge′ P, Pont-Lezica R, Canut H (2006) Lectin receptor kinases participate in protein-protein interactions to mediate plasma membrane-cell wall adhesions in Arabidopsis. Plant Physiol 140:81–90
Grünwald NJ, Goss EM, Press CM (2008) Phytophthora ramorum: a pathogen with a remarkably wide host-range causing sudden oak death on oaks and ramorum blight on woody ornamentals. Mol Plant Pathol 9:729–740
Haas BJ, Kamoun S, Zody MC, Jiang RHY, Handsaker RE, Cano LM, Grabherr M et al (2009) Genome sequence and analysis of the Irish potato famine pathogen Phytophthora infestans. Nature 461:393–398
Habib H, Majid K (2007) Plant protease inhibitors: a defense strategy in plants. Biotechnol Mol Biol Rev 2:068–085
Hála M, Cole R, Synek L, Drdová E, Pecenková T, Nordheim A, Lamkemeyer T, Madlung J, Hochholdinger F, Fowler JE, Zársky V (2008) An exocyst complex functions in plant cell growth in Arabidopsis and tobacco. Plant Cell 20:1330–1345
Hausbeck MK, Lamour KH (2004) Phytophthora capsici on vegetable crops: research progress and management challenges. Plant Dis 88:1292–1303
He CC, Bartholomew CR, Zhou WB, Klionsky DJ (2009) Assaying autophagic activity in transgenic GFP-Lc3 and GFP-Gabarap zebrafish embryos. Autophagy 5:520–526
Heese A, Hann DR, Gimenez-Ibanez S, Jones AM, He K, Li J, Schroeder JI, Peck SC, Rathjen JP (2007) The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc Natl Acad Sci USA 104:12217–12222
Hetz C (2012) The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat Rev Mol Cell Biol 13:89–102
Huang J, Yang M, Lu L, Zhang X (2016) Diverse functions of small RNAs in different plant-pathogen communications. Front Microbiol. https://doi.org/10.3389/fmicb.2016.01552
Huang J, Gu LF, Zhang Y, Yan TX, Kong GH, Kong L, Guo BD, Qiu M, Wang Y, Jing MF, Xing WM, Ye WW, Wu Z, Zhang ZG, Zheng XB, Gijzen M, Wang YC, Dong SM (2017) An oomycete plant pathogen reprograms host pre-mRNA splicing to subvert immunity. Nat Commun 8:2051
Jiang RH, Tripathy S, Govers F, Tyler BM (2008) RXLR effector reservoir in two Phytophthora species is dominated by a single rapidly evolving super family with more than 700 members. Proc Natl Acad Sci USA 105:4874–4879
Jing MF, Guo BD, Li HY, Yang B, Wang HN, Kong GH, Zhao Y, Xu HW, Wang Y, Ye WW, Dong SM, Qiao YL, Tyler BM, Ma WB, Wang YC (2016) A Phytophthora sojae effector suppresses endoplasmic reticulum stress-mediated immunity by stabilizing plant binding immunoglobulin Proteins. Nat Commun 7:11685
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Jorda L, Coego A, Conejero V, Vera PA (1999) Genomic cluster containing four differentially regulated subtilisin-like processing protease genes is in tomato plants. J Biol Chem 274:2360–2365
Jwa NS, Hwang BK (2017) Convergent evolution of pathogen effectors toward reactive oxygen species signaling networks in plants. Front Plant Sci 8:01687
Kamoun S, van West P, de Jong AJ, de Groot KE, Vleeshouwers VG, Govers F (1997) A gene encoding a protein elicitor of Phytophthora infestans is down-regulated during infection of potato. Mol Plant Microbe Interact 10:13–20
Kamoun S, van West P, Vleeshouwers VG, de Groot KE, Govers F (1998) Resistance of Nicotiana benthamiana to Phytophthora infestans is mediated by the recognition of the elicitor protein INF1. Plant Cell 10:1413–1426
Kaschani F, Van Der Hoorn RAL (2011) A model of the C14-EPIC complex indicates hot spots for a protease-inhibitor arms race in the oomycete-potato interaction. Plant Signal Behav 6:1109–1112
Kaschani F, Shabab M, Bozkurt T, Shindo T, Schornack S, Gu C, Hoorn RALVD (2010) An effector-targeted protease contributes to defense against Phytophthora infestans and is under diversifying selection in natural hosts. Plant Physiol 154:1794–1804
Kim I, Xu W, Reed JC (2008) Cell death and endoplasmic reticulum stress: disease relevance and therapeutic opportunities. Nat Rev Drug Discov 7:1013–1030
King SRF, McLellan H, Boevink PC, Armstrong MR, Bukharova T, Sukarta O, Win J, Kamoun S, Birch PRJ, Banfield MJ (2014) Phytophthora infestans RXLR Effector PexRD2 interacts with host MAPKKKe to suppress plant immune signaling. Plant Cell 26:1345–1359
Kruger J, Thomas CM, Golstein C, Dixon MS, Smoker M, Tang SJ, Mulder L, Jones JDG (2002) A tomato cysteine protease required for Cf-2-Dependent disease resistance and suppression of autonecrosis. Science 296:744–747
Lamour KH, Mudge J, Gobena D, Hurtado-Gonzales OP, Schmutz J, Kuo A, Miller NA, Rice BJ, Raffaele S, Cano LM, Bharti AK, Donahoo RS, Finley S, Huitema E, Hulvey J, Platt D, Salamov A, Savidor A, Sharma R, Stam R, Storey D, Thines M, Win J, Haas BJ, Dinwiddie DL, Jenkins J, Knight JR, Affourtit JP, Han CS, Chertkov O, Lindquist EA, Detter C, Grigoriev IV, Kamoun S, Kingsmore SF (2012) Genome sequencing and mapping reveal loss of heterozygosity as a mechanism for rapid adaptation in the vegetable pathogen Phytophthora capsici. MPMI 25:1350–1360
Lee HA, Yeom SI (2015) Plant NB-LRR proteins: tightly regulated sensors in a complex manner. Brief Funct Genom 14:233–242
Li Y, Chen L, Mu J, Zuo J (2013) Lesion simulating disease1 interacts with catalases to regulate hypersensitive cell death in Arabidopsis. Plant Physiol 163:1059–1070
Li Q, Zhang MX, Shen DY, Liu TL, Chen YY, Zhou JM, Dou DL (2016) A Phytophthora sojae effector PsCRN63 forms homo-/hetero-dimers to suppress plant immunity via an inverted association manner. Sci Rep 6:26951
Liu TL, Ye WW, Ru YY, Yang XY, Gu B, Tao K, Lu S, Dong SM, Zheng XB, ShanWX Wang YC, Dou DL (2011) Two host cytoplasmic effectors are required for pathogenesis of Phytophthora sojae by suppression of host defenses. Plant Physiol 155:490–501
Liu TL, Song TQ, Zhang X, Yuan HB, Su LM, Li WL, Xu J, Liu SH, Chen LL, Chen TZ, Zhang MX, Gu LC, Zhang BL, Dou DL (2014) Unconventionally secreted effectors of two filamentous pathogens target plant salicylate biosynthesis. Nat Commun 5:4686
Liu H, Ma X, Yu HQ, Fang DH, Li YP, Wang X, Wang W, Dong Y, Xiao BG (2016) Genomes and virulence difference between two physiological races of Phytophthora nicotianae. GigaScience 5:3. https://doi.org/10.1186/s13742-016-0108-7
Luna E, Pastor V, Robert J, Flors V, Mauch-Mani B, Ton J (2011) Callose deposition: a multifaceted plant defense response. MPMI 24:183–193
Ma Z, Zhu L, Song T, Wang Y, Zhang Q, Xia Y, Qiu M, Lin Y, Li H, Kong L, Fang Y, Ye W, Wang Y, Dong S, Zheng X, Tyler BM, Wang Y (2017) A paralogous decoy protects Phytophthora sojae apoplastic effector PsXEG1 from a host inhibitor. Science 355:710–714
Mafurah JJ, Ma HF, Zhang MX, Xu J, He F, Ye T, Shen DY, Chen YY, Rajput NA, Dou DL, Eugenin EA (2015) A virulence essential CRN effector of Phytophthora capsici suppresses host defense and induces cell death in plant nucleus. PLOS ONE 10:e0127965
McLellan H, Boevink PC, Armstrong MR, Pritchard L, Gomez S, Morales J, Whisson SC, Beynon JL, Birch PRJ (2013) An RxLR effector from Phytophthora infestans prevents re-localisation of two plant NAC transcription factors from the endoplasmic reticulum to the nucleus. PLoS Pathog 9:e1003670
Melech-Bonfil S, Sessa G (2010) Tomato MAPKKKe is a positive regulator of cell-death signaling networks associated with plant immunity. Plant J 64:379–391
Mellersh DG, Heath MC (2001) Plasma membrane-cell wall adhesion is required for expression of plant defense responses during fungal penetration. Plant Cell 13:413–424
Meng X, Zhang S (2013) MAPK cascades in plant disease resistance signaling. Annu Rev Phytopathol 51:245–266
Meng YL, Zhang Q, Ding W, Shan WX (2014) Phytophthora parasitica: a model oomycete plant pathogen. Mycology 5:43–51
Moreno AA, Mukhtar MS, Blanco F, Boatwright JL, Moreno I, Jordan MR, Chen YN, Brandizzi F, Dong XN, Orellana A, Pajerowska-Mukhtar KM (2012) IRE1/bZIP60-mediated unfolded protein response plays distinct roles in plant immunity and abiotic stress responses. PLoS One. https://doi.org/10.1371/journal.pone.0031944
Mosolov VV, Valueva TA (2005) Proteinase inhibitors and their function in plants: a review. Appl Biochem Microbiol 4:227–246
Mukhtar MS, Carvunis AR, Dreze M, Epple P, Steinbrenner J, Moore J, Tasan M, Galli M, Hao T, Nishimura MT, Pevzner SJ, Donovan SE, Ghamsari L, Santhanam B, Romero V, Poulin MM, Gebreab F, Gutierrez BJ, Tam S, Monachello D, Boxem M, Harbort CJ, McDonald N, Gai L, Chen H, He Y, Vandenhaute J, Roth FP, Hill DE, Ecker JR, Vidal M, Beynon J, Braun P, Dangl JL (2011) Independently evolved virulence effectors converge ontohubs in a plant immune system network. Science 333:596–601
Nekrasov V, Li J, Batoux M, Roux M, Chu ZH, Lacombe S, Rougon A, Bittel P, Kiss-Papp M, Chinchilla D, van Esse HP, Jorda L, Schwessinger B, Nicaise V, Thomma BPHJ, Molina A, Jones JDG, Zipfel C (2009) Control of the pattern-recognition receptor EFR by an ER protein complex in plant immunity. EMBO J 28:3428–3438
Nishimura MT, Stein M, Hou BH, Vogel JP, Edwards H, Somerville SC (2003) Loss of a callose synthase results in salicylic acid-dependent disease resistance. Science 301:969–972
Panstruga R, Dodds PN (2009) Terrific protein traffic: the mystery of effector protein delivery by filamentous plant pathogens. Science 324:748–750
Petrov VD, Van Breusegem F (2012) Hydrogen peroxide: a central hub for information flow in plant cells. AoB Plants 1:pls14
Pieterse CM, Van der Does D, Zamioudis C, Leon-Reyes A, Van Wees SC (2012) Hormonal modulation of plant immunity. Annu Rev Cell Dev Biol 28:489–521
Pitzschke A, Schikora A, Hirt H (2009) MAPK cascade signalling networks in plant defence. Curr Opin Plant Biol 12:421–426
Qiang XY, Zechmann B, Reitz MU, Kogel KH, Schafer P (2012) The mutualistic fungus piriformospora indica colonizes Arabidopsis roots by inducing an endoplasmic reticulum stress-triggered caspase-dependent cell death. Plant Cell 24:794–809
Qiao YL, Lui L, Xiong Q, Flores C, Wong J, Shi JX, Wang XB, Liu XG, Xiang QJ, Jiang SS, Zhang FC, Wang YC, Judelson HS, Chen XM, Ma WB (2013) Oomycete pathogens encode RNA silencing suppressors. Nat Genet 45:330–333
Qiao Y, Shi J, Zhai Y, Hou Y, Ma W (2015) Phytophthora effector targets a novel component of small RNA pathway in plants to promote infection. Proc Natl Acad Sci 112:5850–5855
Qutob D, Kemmerling B, Brunner F, Küfner I, Engelhardt S, Gust AA, Luberacki B, Seitz HU, Stahl D, Rauhut T, Glawischnig E, Schween G, Lacombe B, Watanabe N, Lam E, Schlichting R, Scheel D, Nau K, Dodt G, Hubert D, Gijzen M, Nürnberger T (2006) Phytotoxicity and innate immune responses induced by Nep1-like proteins. Plant Cell 18:3721–3744
Rajput NA, Zhang M, Ru Y, Liu T, Xu J, Liu L, Mafurah JJ, Dou D (2014) Phytophthora sojae effector PsCRN70 suppresses plant defenses in Nicotiana benthamiana. PLoS One 9:e98114
Robert-Seilaniantz A, Grant M, Jones JDG (2011) Hormone crosstalk in plant disease and defense: more than just JASMONATE-SALICYLATE antagonism. Annu Rev Phytopathol 49:317–343
Rodriguez MC, Petersen M, Mundy J (2010) Mitogenactivated protein kinase signaling in plants. Annu Rev Plant Biol 61:621–649
Rose JKC, Ham KS, Darvill AG, Albersheim P (2002) Molecular cloning and characterization of glucanase inhibitor proteins: coevolution of a counterdefense mechanism by plant pathogens. Plant Cell 14:1–17
Rosebrock TR, Zeng LR, Brady JJ, Abramovitch RB, Xiao FM, Martin GB (2007) A bacterial E3 ubiquitin ligase targets a host protein kinase to disrupt plant immunity. Nature 448:370–374
Saijo Y (2010) ER quality control of immune receptors and regulators in plants. Cell Microbiol 12:716–724
Sarkies P, Miska EA (2014) Small RNAs break out: the molecular cell biology of mobile small RNAs. Nat Rev Mol Cell Biol 15:525–535
Saunders DGO, Breen S, Win J, Schornack S, Hein I, Bozkurt TO, Champouret N, Vleeshouwers VGAA, Birch PRJ, Gilroy EM, Kamoun S (2012) Host protein BSL1 associates with Phytophthora infestans RXLR effector AVR2 and the solanum demissum immune receptor R2 to mediate disease resistance. Plant Cell 24:3420–3434
Senchou V, Weide R, Carrascoa A, Bouyssoua H, Pont-Lezicaa R, Govers F, Canut H (2004) High affinity recognition of a Phytophthora protein by Arabidopsis via an RGD motif. Cell Mol Life Sci 61:502–509
Sharpee WC, Dean RA (2016) Form and function of fungal and oomycete effectors. Fungal Biol Rev 30:62–73
Singer AU, Schulze S, Skarina T, Xu X, Cui H, Eschen-Lippold L, Egler M, Srikumar T, Raught B, Lee J, Scheel D, Savchenko A, Bonas U (2013) A pathogen type III effector with a novel E3 ubiquitin ligase architecture. PLoS Pathog 9:e1003121
Song J, Win J, Tian M, Schornack S, Kaschani F, Muhammad I, Song J, Win J, Tian MY, Schornack S, Kaschani F, Ilyas M, van der Hoorn RAL, Kamoun S (2009) Apoplastic effectors secreted by two unrelated eukaryotic plant pathogens target the tomato defense protease Rcr3. Proc Natl Acad Sci USA 106:1654–1659
Song TQ, Ma ZC, Shen DY, Li Q, Li WL, Su LM, Ye TY, Zhang MX, Wang YC, Dou DL (2015) An oomycete CRN effector reprograms expression of plant HSP genes by targeting their promoters. PLOS Pathog 11:e1005348
Taguchi F, Shimizu R, Inagaki Y, Toyoda K, Shiraishi T, Ichinose Y (2003) Posttranslational modification of flagellin determines the specificity of HR induction. Plant Cell Physiol 44:342–349
Thaler JS, Humphrey PT, Whiteman NK (2012) Evolution of jasmonate and salicylate signal crosstalk. Trends Plant Sci 17:260–270
Thines M, Lebeda A, Burdon JJ, Thrall P, Jege MJ (2014) Phylogeny and evolution of plant pathogenic oomycetes-a global overview. Eur J Plant Pathol 138:431–447
Tian M, Huitema E, Da Cunha L, Torto-Alalibo T, Kamoun S (2004) A Kazal-like extracellular serine protease inhibitor from Phytophthora infestans targets the tomato pathogenesis-related protease P69B. J Biol Chem 279:26370–26377
Tian M, Benedetti B, Kamoun S (2005) A Second Kazal-like protease inhibitor from Phytophthora infestans inhibits and interacts with the apoplastic pathogenesis-related protease P69B of tomato. Plant Physiol 138:1785–1793
Tian M, Win J, Song J, van der Hoorn R, van der Knaap E, Kamoun S (2007) A Phytophthora infestans cystatin-like protein targets a novel tomato papain-like apoplastic protease. Plant Physiol 143:364–377
Tornero P, Conejero V, Vera P (1997) Identification of a new pathogen-induced member of the subtilisin-like processing protease family from plants. J Biol Chem 272:14412–14419
Tyler BM (2007) Phytophthora sojae: root rot pathogen of soybean and model oomycete. Mol Plant Pathol 8:1–8
Tyler BM, Tripathy S, Zhang XM, Dehal P, Jiang RHY, Aerts A, Arredondo FD, Baxter L, Bensasson D, Beynon JL, Chapman J, Damasceno CMB, Dorrance AE, Dou DL, Dickerman AW, Dubchak IL, Garbelotto M, Gijzen M, Gordon SG, Govers F, Grunwald NJ, Huang W, Ivors KL, Jones RW, Kamoun S, Krampis K, Lamour Lee MK, McDonald WH, Medina M, Nordberg EK, Maclean DJ, Ospina-Giraldo MD, Morris PF, Phuntumart V, Putnam NH, Rash S, Rose JKC, Sakihama Y, Salamov AA, Savidor A, Scheuring CF, Smith BM, Sobral BWS, Terry A, Torto-Alalibo TA, Win J, Xu ZY, Zhang HB, Grigoriev IV, Rokhsar DS, Boore JL (2006) Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science 313:1261–1266
van Damme M, Bozkurt TO, Cakir C, Schornack S, Sklenar J, Jones AME, Kamoun S (2012) The Irish potato famine pathogen Phytophthora infestans translocates the CRN8 kinase into host plant cells. PLoS Pathog 8:e1002875
Vance V, Vaucheret H (2001) RNA silencing in plants-defense and counter defense. Science 292:2277–2280
Vetukuri RR, Whisson SC, Grenville-Briggs LJ (2017) Phytophthora infestans effector Pi14054 is a novel candidate suppressor of host silencing mechanisms. Eur J Plant Pathol 149:771–777
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136:669–687
Wang YJ, Li JF, Hou SG, Wang XW, Li YA, Ren DT, Chen S, Tang XY, Zhou JM (2010) A Pseudomonas syringae ADP-ribosyltransferase inhibits Arabidopsis mitogen-activated protein kinase kinases. Plant Cell 22:2033–2044
Wang XD, Boevink P, McLellan H, Armstrong M, Bukharova T, Qin ZW, Birch PRJ (2015) A host KHRNA-binding protein is a susceptibility factor targeted by an RXLR effector to promote late blight disease. Mol Plant 8:1385–1395
Wang SM, Boevink PC, Welsh L, Zhang RF, Whisson SC, Birch PRJ (2017) Delivery of cytoplasmic and apoplastic effectors from Phytophthora infestans haustoria by distinct secretion pathways. New Phytol 216:205–215
Wang SM, Welsh L, Thorpe P, Whisson SC, Boevinkb PC, Birch PRJ (2018) The Phytophthora infestans haustorium is a site for secretion of diverse classes of infection-associated proteins. mBio. https://doi.org/10.1128/mBio.01216-18
Wang SM, McLellan H, Bukharova T, He Q, Murphy F, Shi JY, Sun SH, van Weymers P, Ren YJ, Thilliez G, Wang HX, Chen XW, Engelhardt S, Vleeshouwers V, Gilroy EM, Whisson SC, Hein I, Wang XD, Tian ZD, Birch PRJ, Boevink PC (2019) Phytophthora infestans RXLR effectors act in concert at diverse subcellular locations to enhance host colonization. J Exp Bot 70:343–356
Whisson SC, Boevink PC, Moleleki L, Avrova AO, Morales JG, Gilroy EM, Armstrong MR, Grouffaud S, van West P, Chapman S, Hein I, Toth IK, Pritchard L, Birch PRJ (2007) A translocation signal for delivery of oomycete effector proteins into host plant cells. Nature 450:115–118
Whisson SC, Boevink PC, Wang SM, Birch PRJ (2016) The cell biology of late blight disease. Curr Opin Microbiol 34:127–135
Win J, Kamoun S (2007) Adaptive evolution has targeted the C-terminal domain of the RXLR effectors of plant pathogenic oomycetes. Plant Cell 19:2349–2369
Win J, Krasileva KV, Kamoun S, Shirasu K, Staskawicz BJ, Banfield MJ (2012) Sequence divergent RXLR effectors share a structural fold conserved across plant pathogenic oomycete species. PLoS Pathog 8:e1002400
Xiong Q, Ye WW, Choi D, Wong J, Qiao YL, Tao K, Wang YC, Ma WB (2014) Phytophthora suppressor of RNA silencing 2 is a conserved RxLR effector that promotes infection in soybean and Arabidopsis thaliana. MPMI 27:1379–1389
Yang ZT, Wang MJ, Sun L, Lu SJ, Bi DL, Sun L, Song ZT, Zhang SS, Zhou SF, Liu JX (2014a) The membrane-associated transcription factor NAC089 controls ER-stress-induced programmed cell death in plants. PLoS Genet 10:e1004243
Yang ZT, Lu SJ, Wang MJ, Bi DL, Sun L, Zhou SF, Song ZT, Liu JX (2014b) A plasma membrane-tethered transcription factor, NAC062/ANAC062/NTL6, mediates the unfolded protein response in Arabidopsis. Plant J 79:1033–1043
Yang B, Wang QQ, Jing MF, Guo BD, Wu JW, Wang HN, Wang Y, Lin L, Wang Y, Ye W, Dong S, Wang Y (2017) Distinct regions of the Phytophthora essential effector Avh238 determine its function in cell death activation and plant immunity suppression. New Phytol 214:361–375
Yoshioka H, Asai S, Yoshioka M, Kobayashi M (2009) Molecular mechanisms of generation for nitric oxide and reactive oxygen species, and role of the radical burst in plant immunity. Mol Cells 28:321e9
Yu XL, Tang JL, Wang QQ, Ye WW, Tao K, Duan SY, Lu CC, Yang XY, Dong SM, Zheng XB, Wang YC (2012) The RxLR effector Avh241 from Phytophthora sojae requires plasma membrane localization to induce plant cell death. New Phytol 196:247–260
Zadoks JC (2008) The potato murrain on the European continent and the revolutions of 1848. Potato Res 51:5–45
Zhang J, Shao F, Li Y, Cui HT, Chen LJ, Li HT, Zou Y, Long CZ, Lan LF, Chai JJ, Chen S, Tang XY, Zhou JM (2007) A Pseudomonas syringae effector inactivates MAPKs to suppress PAMP-Induced immunity in plants. Cell Host Microbe 1:175–185
Zhang MX, Li Q, Liu TL, Liu L, Shen DY, Zhu Y, Liu PH, Zhou JM, Dou DL (2015) Two cytoplasmic effectors of Phytophthora sojae regulate plant cell death via interactions with plant catalases. Plant Physiol 167:164–175
Zhang W, Corwin JA, Copeland D, Feusier J, Eshbaugh R, Chen F, Atwell S, Kliebenstein DJ (2017) Plastic transcriptomes stabilize immunity to pathogen diversity: the jasmonic acid and salicylic acid networks within the Arabidopsis/Botrytis Pathosystem. Plant Cell 29:2727–2752
Zheng XZ, McLellan HZ, Fraiture MZ, Liu XY, Boevink PC, Gilroy EM, Chen Y, Kandel K, Sessa G, Birch PRJ, Brunner F (2014) Functionally redundant RXLR effectors from Phytophthora infestans act at different steps to suppress early flg22-triggered immunity. PLoS Pathog 10:e1004057
Zhou BJ, Zeng LR (2017) Conventional and unconventional ubiquitination in plant immunity. Mol Plant Pathol 18:1313–1330
Zhu JK, Shi J, Singh U, Wyatt SE, Bressan RA, Hasegawa PM, Carpita NC (1993) Enrichment of vitronectin- and fibronectin-like proteins in NaCl adapted plant cells and evidence for their involvement in plasma membrane-cell wall adhesion. Plant J 3:637–646
Zipfel C, Kunze G, Chinchilla D, Caniard A, Jones JD, Boller T, Felix G (2006) Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium mediated transformation. Cell 125:749–760
Acknowledgements
This work was supported by the Fundamental Research Funds for the Chinese Academy of Agricultural Sciences (Grant no. 1610232016018), the Agricultural Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences (Grant no. ASTIP-TRIC04), and the China National Tobacco Corp. Yunnan Science and Technology Project: Construction of Tobacco Genome Breeding Platform (Grant no. 2017YN05). We declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, W., Jiao, F. Effectors of Phytophthora pathogens are powerful weapons for manipulating host immunity. Planta 250, 413–425 (2019). https://doi.org/10.1007/s00425-019-03219-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-019-03219-x