Abstract
Main conclusion
Analysis of stress-associated miRNAs of Glycine max (L.) Merrill reveals wider ramifications of small RNA-mediated (conserved and legume-specific miRNAs) gene regulatory foot prints in molecular adaptive responses.
MicroRNAs (miRNAs) are indispensable components of gene regulatory mechanism of plants. Soybean is a crop of immense commercial potential grown worldwide for its edible oil and soy meal. Intensive research efforts, using the next generation sequencing and bioinformatics techniques, have led to the identification and characterization of numerous small RNAs, especially microRNAs (miRNAs), in soybean. Furthermore, studies have unequivocally demonstrated the significance of miRNAs during the developmental processes and various stresses in soybean. In this review, we summarize the current state of understanding of miRNA-based abiotic and biotic stress responses in soybean. In addition, the molecular insights gained from the stress-related soybean miRNAs have been compared to the miRNAs of other crops, especially legumes, and the core commonalities have been highlighted, though differences among them were not ignored. Nature of response of soybean-derived conserved miRNAs during various stresses was also analyzed to gain deeper insights regarding sRNAome-based defense responses. This review further provides way forward in legume small RNA transcriptomics based on the adaptive responses of soybean and other legume-derived miRNAs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Small non-coding RNAs (sncRNAs) are effectors of RNA-mediated gene silencing and are known to actively participate in a repertoire of plant growth and developmental processes including cellular differentiation, response to environmental stimuli, and defense against invading organisms (Hamilton and Baulcombe 1999; Jones-Rhoades et al. 2006; Mallory and Vaucheret 2006; Brant and Budak 2018). sncRNAs of plants are classified based on their origin, secondary structural features and mode of action on the target RNAs (Meyers et al. 2008). It is well established that small interfering RNAs (siRNAs) and microRNAs (miRNAs) are two major classes of plant ncRNAs (Llave et al. 2002; Bartel 2004; Meyers et al. 2008; Chen 2009). Although siRNAs originate from dsRNAs, they are of diverse nature, namely heterochromatic siRNAs (hc-siRNAs), natural antisense transcript siRNAs (nat-siRNAs) and transacting siRNAs (ta-siRNAs) (Ramesh et al. 2013). However, the actions of siRNAs on the target transcripts vary depending on their size. siRNAs of 24 nt class target heterochromatin region and are involved in RNA-dependent DNA methylation (RdDM), thereby leading to transcriptional gene silencing (TGS). On the other hand, 21 nt siRNAs cleave target mRNAs that show perfect sequence complementarity in a process called post-transcriptional gene silencing (PTGS).
miRNAs have been implicated as effectors of gene expression, especially in the adaptation to biotic and abiotic stresses, which ultimately affect the growth and development of an organism (Lee et al. 1993; Reinhart et al. 2002; Groszhans and Filipowicz 2008; Brant and Budak 2018). miRNAs are generated from imperfect stem-loop RNA structures, that in turn are derived from single-stranded RNA precursors called primary miRNA transcripts (pri-miRNA). Plant miRNA genes are transcribed by the host Pol II to generate pri-miRNAs. pri-miRNA transcripts are further processed into functional miRNA: miRNA* pairs through precursor miRNA (pre-miRNA) due to the concerted activity of many host proteins such as RNAse III, Dicer-like-1 (DCL-1), and DAWDLE. Plant pre-miRNAs are relatively long (~ 90 to 140 bp) and are processed into double-stranded mature miRNA (miRNA: miRNA* pair). Inside the nucleus, DCL-1 interacts with dsRNA-binding proteins (RBPs) such as HYPONASTIC LEAVES1 (HYL1) and the zinc finger protein SERRATE (SE) to process the miRNAs. To improve the stability of the miRNAs, these small RNAs (sRNAs) are methylated by HUA Enhancer 1 (HEN 1), whereas unmethylated sRNAs are uridylated by HEN1 SUPPRESSOR1 (HESO1) (Zhao et al. 2012). In plants, mature miRNAs are exported out of the nucleus by EXPORTIN-like proteins called as HASTY 1 (HST1). In the cytoplasm, the 21 nt long miRNAs are recruited on to the slicers called as RNA-induced silencing complex (RISC) that has argonaute (AGO) as its main component. The RISC then cleaves the cognate mRNA or represses the translation of mRNA based on the nucleotide sequence complementarity. miRNA-mediated translational repression of complementary mRNA occurs in the endoplasmic reticulum of plants (Brodersen et al. 2008). Interestingly, 24 nt long miRNAs (lmiRNAs), discovered in Oryza sativa, have been found to be involved in DNA methylation, suggesting an additional layer of miRNA-mediated transcriptional gene regulation (Wu et al. 2010). Furthermore, imprecisely processed miRNAs cause non-canonical sncRNA biogenesis with profound implications for miRNA expression and target RNA degradation capabilities (Budak and Akpinar 2015).
Plants being sessile have evolved molecular mechanisms to respond to various environmental stimuli such as biotic (including interactions with a symbiotic partner) and abiotic stresses. The phenomenon of RNA silencing and the knowledge of sRNAs have converged in delineating the miRNA-based gene regulatory networks in plants. miRNAs are found throughout the plant kingdom from mosses to angiosperms and few of them are evolutionarily conserved (Axtell et al. 2007). miRNAs are involved in complex regulatory mechanisms that coordinate the plant developmental activities, stress responsiveness, regulation of hormone signaling pathways, maintenance of nutrient homeostasis, symbiosis and regulation of its biogenesis (Carrington and Ambros 2003; Sunkar 2010; Khraiwesh et al. 2012; Budak et al. 2014).
Genomes and transcriptomes of legumes such as Glycine max, Medicago truncatula and Lotus japonicus have been intensively investigated (Mochida et al. 2010; Soares-Cavalcanti et al. 2012). Of these, soybean is a model legume and an economically important crop with an amphidiploid genome. Despite the improved understanding of miRNA-mediated gene regulations in plants, a common stress-responsive miRNA pathway or identification of a conserved set of stress-responsive miRNAs across the plant species to decode miRNA-based functional networks is still incomplete. Additionally, molecular dissection and understanding of the stress-related miRNA networks in soybean could immensely aid the development of improved crop phenotypes.
Plant miRNAs coordinate the expression of transcriptional factors (TFs), suggesting their pre-eminence in programming growth and developmental process (Rhoades et al. 2002; Reyes and Chua 2007; Mitsuda and Ohme-Takagi 2009). In plants, reports of miRNAs responsive to environmental stimuli have revealed upregulation of miRNA 395 during reduced sulphate conditions (Jones-Rhoades and Bartel 2004). Also, miRNAs such as miR395, miR397b, and miR402 have been shown to be involved in stress responsiveness (Phillips et al. 2007). Later miRNA 398a/b and miR408 were identified to be responsive to water-deficit stress in M. truncatula and chickpea (Trindade et al. 2010; Hajyzadeh et al. 2015). The upregulation of miR398a/b and miR408 and downregulation of respective target transcripts (mitochondrial cytochrome oxidase and plantacyanin) disclose a strong molecular connect between copper homeostasis and drought in M. truncatula (Trindade et al. 2010). Similarly, O. sativa-derived miR393 was found to be regulated in response to salinity and alkalinity (Gao et al. 2011). Development of robust next generation sequencing (NGS) platforms and progresses in the field of computational biology have discovered and characterized many stress-responsive miRNAs (Jones-Rhoades and Bartel 2004; Budak et al. 2014; Alptekin et al. 2017). Comparative miRNA expression studies in bread wheat and its wild relative identified candidate miRNAs (miR1435, miR5024, and miR7714) and differentially regulated miRNAs for exploitation and development of drought-tolerant phenotype (Akpinar et al. 2015; Kantar et al. 2011). Abiotic stress-responsive miRNAs of Triticeae members, especially wheat and barley, have helped in identifying conserved regulatory mechanisms so that miRNA:target pairs could be manipulated to develop better crop phenotype (Alptekin et al. 2017). Besides nuclear miRNAs, the function of miRNA variants-isomiRs and organellar miRNAs in stress adaptations are also recognized (Budak et al. 2015a). Conserved miRNAs in model species would pave for rapid exploitation of miRNA-based transcriptomics in delineating stress responses of cultivated crops (Budak and Akpinar 2011).
Importance of small RNAs in general and miRNAs in particular has been well acknowledged because impaired sRNA biogenesis or miRNA-mediated gene regulatory networks cause susceptibility to pathogenic stressors (Ramesh et al. 2014). Further, legume and solanaceous plants-derived miRNAs alter the expression of defense-related NBS-LRR genes and are involved in host’s innate immunity. More than 40 plant-derived miRNA families have been shown to be involved in response to abiotic stresses where in 13 miRNA families play diverse roles in response to salt and drought stresses (Nageshbabu et al. 2013; Carrington and Ambros 2003; Sunkar 2010; Khraiwesh et al. 2012; Budak et al. 2015b; Brant and Budak 2018). Applications of miRNAs in crop genetic modification have not only yielded virus-resistant genotypes (Ramesh et al. 2014) but also various traits of economic importance (Budak et al. 2015b; Zhang and Wang 2016).
Glycine max miRNAome
Preliminary studies of soybean miRNAs were performed using expressed sequence tag (EST) and a genome survey sequence (GSS) approach (Zhang et al. 2005). Expression patterns of G. max precursor miRNAs, deduced from EST databases, have been provided by Dezulian et al. (2006). Thus, EST-GSS approach identified 33 families (69 miRNAs) of soybean miRNAs and five miRNAs in G. soja and G. clandestine (Zhang et al. 2008). G. max-specific miRNAs (gma-miR168, gma-miR393 and gma-miR172) are induced when the roots are colonized by rhizobial partner Bradyrhizobium japonicum during symbiotic nitrogen fixation (SNF) (Subramanian et al. 2008; Wang et al. 2009).
Genome-wide analysis of miRNAs, the organization of miRNA families and sequence diversity of mature miRNAs as well as the corresponding target transcript(s) have revealed that gma-miRNAs are primarily intergenic, as in other plant species; however, several intra-genic miRNAs have also been reported (Turner et al. 2012). A potential co-regulation of novel soybean miRNA, gma-new-miR13587 and its parent gene, Glyma05g36870 was documented by Turner et al. (2012), nevertheless, such miRNA:parent gene pairs have not yet been discovered in soybean. Both conserved miRNA families (MIR159, MIR169 and MIR395) and soybean-specific miRNAs (MIR-Seq14) were found to be organized in tandem duplications (Turner et al. 2012). Although duplication of miRNA genes is found in distantly related angiosperms (MIR159 genes are found clustered among soybean, sorghum and maize) (Zhang et al. 2009), the phenomenon is not conserved across the plant kingdom (Arabidopsis encodes unclustered MIR159 genes) (Allen et al. 2007). Genome-wide analysis of tandem duplications revealed that the number and orientation of miRNAs were different in the paralogous genome. Hence, the evolution and diversity of soybean miRNAs are attributable to the genome-wide and localized duplications (Turner et al. 2012). Co-evolution of soybean miRNA genes (MIRs) and their target mRNAs revealed that domestication was a driving factor for the evolution of miRNA gene variants. Besides, factors such as high expression of MIRs and target pairs, duplication status and the number of target mRNAs and flanking genomic regions might have also contributed to miRNA evolution (Liu et al. 2016). In addition, over one half of soybean miRNA-target pairs have undergone purifying selection in the process of domestication and improvement. The process of domestication has increased the genetic similarity among the MIRs and target pairs in cultivated genotypes than in wild relatives (Liu et al. 2016). Promoters and cis-acting elements of soybean-derived miRNAs have been analyzed using in silico tools. Majority of the miRNAs (84%) have upstream promoter sequences, however, 8.7% of miRNA loci were characterized with the downstream promoters (Han et al. 2014). Additionally, hormone-mediated negative feedback mechanism of miRNA regulation in soybean has been identified (Han et al. 2014). Small RNA sequencing and analysis have yielded 638 non-redundant MIRs of soybean (Arikit et al. 2014; Zhao et al. 2015a). However, MIRs that have features of endogenous siRNAs were removed and 454 MIRNAs have been categorized as genuine miRNAs (Zhao et al. 2015b). Genomic distribution of these 454 MIRs revealed that majority of them [213 MIRs (46.9%)] were mapped to unclassified genomic sequences, whereas 162 MIRs (35.7%) were found within the protein-encoding genes (PEGs). Interestingly, 79 MIRs (17.4%) were found among the repetitive sequences especially the transposable elements (Zhao et al. 2015b).
Soybean miRNAome and stressors
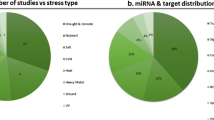
Soybean is exposed to various abiotic stresses such as drought, chilling/freezing, nutrient deficiency/starvation, salinity, heavy metals and biotic stresses such as bacterial, viral and fungal infections and nematode and insect infestations (Miransari 2015; Ramesh et al. 2015). Exploration of stress-responsive miRNAs in soybean has yielded many insights and resulted in identification of miRNAs with regulatory roles in various physiological and molecular processes (Fig. 1; Table 1).
Stress-responsive miRNAs of soybean and their regulatory roles in abiotic and biotic stresses [miRNA-mediated regulation of transcriptional factor (GmNFYA3) during drought stress and legume specific miRNA-based regulation of NBS-LRR loci to produce phased siRNAs (phasiRNAs) during biotic stresses is presented. Further, miRNA-mediated responsiveness during symbiosis that involve promotion of nodulation and repression of defense response pathways are depicted]
miRNAs associated with moisture/water-deficit stress or drought
Among the major stressors of soybean, low soil moisture stress or drought causes adverse impact on the plant’s photosynthetic ability, carbon assimilation, nutrient uptake status and stomatal movement; these affect the overall metabolic process leading to severe yield losses. In a drought-sensitive soybean genotype, a set of miRNAs (miR166-5p, miR169f-3p, miR1513c and miR397ab) are upregulated (Kulcheski et al. 2011), whereas in tolerant genotypes, miR397ab was found to be downregulated under drought. Similarly, miR397 was downregulated during stress in rice and during the drought in peach roots (Eldem et al. 2012), whereas it was upregulated during the drought in Arabidopsis (Zhou et al. 2010). The target of miR397 was found to be a transcript encoding β-fructofuranosidase, a key enzyme involved in starch and sucrose metabolism. Thus, it appears that the expression status of miR397 coordinates carbon fixation and energy supply in plants (Zhou et al. 2010). Dissection of miRNA expressional changes in wheat and its progenitor Aegilops tauschii revealed differential downstream processing of drought-responsive pre-miRNA 5523 in wheat, whereas mature miRNA was observed only in A. tauschii, suggesting the loss of functional miRNAs during domestication (Akpinar and Budak 2016).
In plants, hormonal signaling plays a crucial role in response to drought, wherein miRNAs act as intermediates between stress hormones and transcriptional factors (TFs). Conserved miRNAs such as miR169 were found to accumulate during ABA treatment as well as Rhizobium colonization in P. vulgaris (Arenas-Huertero et al. 2009). In M. truncatula, miR169 was upregulated during Rhizobium root colonization (Combier et al. 2006), whereas in rice, miR169 gene possesses dehydration-responsive element (DRE) (Zhao et al. 2007) suggesting the significance of this particular miRNA during drought and rhizobial colonization processes across the plant kingdom. Contrarily, miR169a and miR196c exhibited downregulation in Arabidopsis, with low abundance of miR169 in P. vulgaris (Arenas-Huertero et al. 2009) and differential regulation of miR169 in wheat documented under drought stress (Akdogan et al. 2016). Induction of miR159a was observed under drought and ABA treatment in Arabidopsis seeds. Further, miR159 directs degradation of MYB TFs such as MYB33 and MYB101 (Reyes and Chua 2007). miR167, which was identified as a negative regulator of phospholipase D (PLD) in Zea mays, was inhibited under the influence of drought and ABA (Wei et al. 2009). Similarly, inhibition of miR169a was observed in Arabidopsis, which resulted in the accumulation of its target nuclear factor Y (NF-Y) transcription factor, NFYA5—a TF that plays an important role in response to many of the environmental stresses (Li et al. 2008). The soybean homolog GmNFYA3 was upregulated during abscisic acid, PEG, salt and cold-induced stresses (Ni et al. 2013). Furthermore, gma-miR169 directs in vivo cleavage of GmNFYA3 which is involved in activation of nuclear-specific transcripts that confer enhanced drought tolerance and induces expression of genes involved in ABA biosynthesis and signaling in Arabidopsis (Ni et al. 2013). Thus, gma-miR169:GmNFYA3 target pair plays a key role in drought stress tolerance in soybean. This view was further corroborated by a genome-wide expression analysis of NF-Y in soybean, demonstrating a prominent role for NF-Y class of TFs in drought responsiveness and in other development related processes (Quach et al. 2015).
miRNAs associated with salinity stress
Growth and development of plants are profoundly impaired when subjected to salinity stress. Under salt stress, soil rhizosphere not only obstructs the ability of the roots to uptake essential nutrients but also interferes in water absorption. Soybean nodules subjected to salt stress showed more than tenfold decrease in the expression of gma-miR159c, gma-miR159b, gma-miR169c and gma-miR319a, b (Dong et al. 2013). In addition, 34 novel miRNAs are repressed, whereas 12 novel miRNAs are induced in the matured root nodules during salt stress (Dong et al. 2013). Analysis of salt-responsive miRNAs identified 770 mRNAs as targets; predominant of them (79) are TFs. Also, the target genes are involved in diverse functions such as Ca2+/calmodulin-dependent protein kinase and ubiquitin-conjugating enzymes, demonstrating the molecular cross-talk upon induction of salt stress in matured root nodules of soybean (Dong et al. 2013). To support this further, miRNA: target pair [miR172c: NNC1 (Nodule Number Control 1)] is involved in modulating the root plasticity during salinity stress (Sahito et al. 2017). Salt stress caused the over-expression of miR172c and the corresponding downregulation of the target gene NNC1 (Nodule Number Control 1); thus knock-down of NNC1 in soybean was found to promote salt stress tolerance (Sahito et al. 2017). Members of miRNA 169 family inhibit NF-YA transcription factor in Oryza sativa (Zhao et al. 2009) and Arabidopsis during drought, whereas in the wheat, miR169 family was found to be differentially regulated upon salt treatment (Eren et al. 2015). In addition, rice-derived miR393a plays an important role in response to salt stress (Gao et al. 2011) as it downregulates mRNA-encoding F-box auxin receptors such as Transport Inhibitor Response 1 (TIR1), AFB2 and AFB3 (Navarro et al. 2006; Xia et al. 2012). Although a gamut of Arabidopsis miRNAs is upregulated under the influence of salt stress, miR398 is down regulated (Liu et al. 2008). Similarly, microarray-based expression profiling of salinity stress-responsive miRNAs of Zea mays resulted in the identification of 27 downregulated miRNA families, whereas miR162, miR168, miR395 and miR474 were upregulated (Ding et al. 2009).
miRNAs and cold stress responses
Upregulation of cold stress-responsive miRNAs, namely miR393, miR397b, miR402 and miR319c in Arabidopsis were reported (Sunkar and Zhu 2004). Later, many cold stress-responsive miRNAs have been unearthed in Populus (Lu and Huang 2008), Brachypodium (Zhang et al. 2009) and Oryza (Li et al. 2010). Soybean miRNAs responsive to B. japonicum symbiosis were known; however, the effect of low temperature on the expression of miRNAs in soybean root nodules remained largely unexplored (Wang et al. 2009). This led to the identification of nodule-specific, cold-responsive miRNAs of soybean (upregulated: gma-miR397a, gma-miR166u and gma-miR171p and repressed: gma-mi169c, gma-mi159b, gma-miR319a/b and gma-miR5559) (Zhang et al. 2014). The targets of gma-miR166u are basic leucine zipper (bZIP) TF and an HD-ZIP protein. Hence, gma-miR166u acts on these TFs and coordinate the gene expression pathways by turning it “Off and On” when required (Zhang et al. 2014). Further, the target gene for gma-miR171p is GRAS family TF indicating the significance of miRNA-mediated cellular responses during the cold stress. Similarly, 51 chilling-responsive miRNAs have been identified along with 898 miRNA target transcripts that were found to be enriched in red-ox reactions and signaling pathways in vegetable soybean (Xu et al. 2016).
miRNAs associated with nutrient homeostasis
miRNAs have also been identified to play a significant role in the nutrient homeostasis of plants (Pant et al. 2008). A well-characterized miRNA-mediated nutrient uptake system involving miRNA399 and PHOSPHATE2 (PHO2) was known in Arabidopsis (Pant et al. 2008). During phosphate starvation, miRNA399 is upregulated in the roots causing cleavage of PHO2 transcripts, ultimately increasing phosphorus uptake. Once phosphorus uptake is saturated, downregulation of miRNA393 is achieved due to the target mimic activity of Induced by Phosphate Starvation 1 (IPS1) transcript (Pant et al. 2008). Under the limiting condition of copper ions, miRNA398 is upregulated to target CSD1 and CSD2 mRNAs which are involved in the release of copper ions. Besides, other miRNAs have also been found to target transcripts that encode copper-containing proteins such as laccase and plantacyanin (Abdel-Ghany and Pilon 2008). Similarly, the growth of soybean in acidic soils is severely hampered by the high concentration of aluminum ions (Al3+). The molecular mechanism underlying the adaptation to high Al3+ conditions revealed that expression of 30 Glycine soja derived miRNAs are influenced by the aluminum stress. Also, Al3+ phytotoxicity-responsive miRNAs target TFs such as auxin response factor (ARF), MYB transcripts coding for leucine-rich repeat and toll/interleukin-1 receptor-like protein (LRR-TIR) and NB-ARC domain-containing disease resistance protein (Zeng et al. 2012). A study has revealed a set of miRNAs that were differentially regulated during aluminum toxicity stress in soybean (Huang et al. 2017). A deeper understanding of the role of conserved miRNAs during the aluminum stress showed miRNA-mediated root elongation in a tolerant soybean genotype (BX10), whereas miRNAs trigger oxidative stress in a susceptible genotype (BD2) (Huang et al. 2017).
Soybean miRNAome and symbiosis
miRNAs responsive to Bradyrhizobium symbiosis
Early stages of root nodule formation documented upregulation of two miRNAs, viz. miR168 and miR172 and downregulation of miR169 while soybean is infected by B. japonicum (Subramanian et al. 2008). Soybean-derived miRNAs have been associated with the alterations of hormonal signaling pathways by modulating the expression levels of auxin response factors (ARFs) (Subramanian et al. 2008). Similarly, Wang et al. (2009) identified 32 soybean-derived miRNAs, including miR167, miR172, miR396 and miR399 that are involved in the later stage of nodulation and nitrogen fixation. Likewise, target predictions of M. truncatula-derived miRNAs in response to Bradyrhizobium identified TFs, and mRNAs involved in hormone-responsive signaling pathways (EI Yahyaoui et al. 2004). Thus, a common modus operandi of legume-specific miRNAs was emerging in the regulation of host gene expressions during Bradyrhizobium colonization. Furthermore, constitutive expression of soybean miRNAs such as miR482, miR1512 and miR1515 resulted in considerable increase in nodule number, suggesting the direct involvement of these miRNAs in SNF (Li et al. 2010). Interestingly, miR482 represses R-genes linked to disease resistance (Li et al. 2010). Thus, SNF in soybean involves a gene regulatory cascade comprising phytohormone signaling, cell cycle and R genes that aims to repress the host’s antibacterial defense response against rhizobial colonization on one hand and maximizes nitrogen fixation on the other. Nodulation-specific miRNAs have been characterized in soybean. Ectopic overexpression of miR172j improved nodule numbers that were attributed to its inhibitory effect on nodule hemoglobin mediated by APETALA 2 (AP2) TFs (Yan et al. 2013). However, expression of miR160 caused inhibitory effects on soybean nodulation due to its impact on auxin response factors (Turner et al. 2013). Significant changes in the expression level of miR393j-3p corroborated its indispensable role in the process of nodule formation (Yan et al. 2015). Further, miR393j-3p-mediated regulation of Early Nodulin 93 (ENOD93) mRNA is critical for the development of soybean nodule (Yan et al. 2015). Auxin was known to promote the formation of nodules in legumes, however, the precise mechanism behind this action was not known until recently. Cai et al. (2017) showed that soybean-derived miRNA gma-miR393 negatively regulate auxin receptors such as GmTIR1 and GmAFB3. Thus, the spatio-temporal regulation of GmTIR1 and GmAFB3 transcripts by miR393 family significantly affects nodule formation in soybean. Although miRNA-mediated gene expression changes during arbuscular mycorrhiza (AM) symbiosis are being deciphered in diverse crops such as M. truncatula (Devers et al. 2011; Bazin et al. 2013), tomato (Cervantes-Gámez et al. 2016) and maize (Xu et al. 2018), reports in soybean are not available.
Soybean miRNAome and biotic stresses
Phytophthora sojae-responsive miRNAs
Microarray-based profiling of miRNAs in three soybean cultivars (Williams-susceptible; Conrad and Williams-resistant) upon Phytophthora sojae infection identified many miRNA–mRNA pairs. A feedback control kind of network involving soybean miRNAs and protein-coding genes has also been proposed by Guo et al. (2011). Differential regulation of P. sojae-responsive miRNAs was observed in soybean roots (Wong et al. 2014). It was proposed that P. sojae-responsive miRNAs such as gma-miR393 and gma-miR166 are pertinent to the basal defense mechanism against this oomycete infection. This suggestion was further supported by the soybean lines, wherein knockdown of miR393 exhibited greater susceptibility to P. sojae. Furthermore, the genes involved in the isoflavanoid biosynthetic pathway were also downregulated (Wong et al. 2014).
sRNA profiling in P. sojae susceptible soybean cultivar ‘Williams’ and nine near isogenic lines (NILs), each carrying a distinct P. sojae-resistant gene (Rps), deciphered the molecular foot print connecting miRNAs, nucleotide binding site-leucine-rich repeat (NBS-LRR) genes, and phased siRNAs (phasiRNAs) (Zhao et al. 2015a). Eight major soybean-derived miRNA families (miR1510, miR1507, miR2109 miR482/2118, miR5668, miR5376, miR172 and miR5041) targeted 257 NBS-LRR genes (Zhao et al. 2015a). In response to P. sojae infection, G. max miRNAs, viz. miR1510, miR1507, miR2109, miR482/2118 and miR5376 were downregulated in the resistant NILs (Zhao et al. 2015a). Upregulation of phasi-NB-LRRs was also associated with the downregulation of respective phasiRNAs in NILs. Thus, miRNA-NBS-LRR-phasiRNAs interplay was documented during P. sojae infection and disease development (Zhao et al. 2015a). Interestingly, PHAS loci identified in the study were also documented in the vegetative, reproductive parts and nodules of soybean (Arikit et al. 2014), indicating the importance of phasiRNAs not only in biotic stress but also in other biological and developmental processes. Similarly, some of the miRNAs families have already been reported to target NBS-LRR genes in M. truncatula (Zhai et al. 2011). The small RNA atlas of soybean further emphasizes the importance of a molecular connection between the miRNAs and phased siRNAs (phasiRNAs from PHAS loci) (Arikit et al. 2014). Furthermore, the majority of PHAS loci encode NBS-LRR genes implying the importance of miRNA:phasiRNAs interactions in conferring disease resistance in soybean (Arikit et al. 2014). Soybean hairy roots, over-expressing gma-miR1510a/b, is greatly susceptible to P. sojae infection as miR1510 targets and cleaves NBS-LRR class transcript encoded by gene Glyma.16G135500 (Cui et al. 2017).
miRNAs associated with rust pathogen infection
The fungal pathogen Phakopsora pachyrhizi causes devastating Asian soybean rust (ASR). Expression analysis of soybean-derived miRNAs during ASR infection revealed downregulation of miR166a-5p, miR166f, miR169-3p, miR397ab and miR-seq13 in the susceptible genotype (Embrapa 48), whereas in the resistant genotype (PI561356), no differential miRNA expression was observed. miR4415b showed decreased expression in the susceptible genotype upon pathogen infection. Expression of miR4415b remained unchanged in the control and pathogen challenged plants of resistant genotype, whereas the expression levels of miR4415b were still higher than found in the susceptible genotype (Kulcheski et al. 2011).
miRNAs associated with antiviral response
Soybean mosaic virus (SMV) infection (Strain G2) causes downregulation of many defense-related genes during early stages of infection (Babu et al. 2008). Hence, miRNAs have been envisaged to play a greater role in response to SMV infection. Yin et al. (2013) profiled miRNAs from mock-inoculated and SMV-inoculated soybean plants that led to the identification of 52 families of miRNAs (179 miRNAs) during viral infection. Targets of 12 SMV-responsive miRNAs have been validated; miR160, miR393 and miR1510 were shown to be involved in resistance response to SMV infection (Yin et al. 2013). SMV (strains G2 and G7) infected susceptible [Williams 82 (rsv)] and resistant [PI96983 (Rsv1)] genotypes demonstrated that the disease reaction is determined by the interplay of both miRNA- and siRNA-mediated gene silencing systems (Chen et al. 2015). Among the miRNAs, gma-miR168 mediated argonaute 1 (AGO 1) homeostasis was disrupted in Rsv1 genotype upon SMV G7 infection, whereas knock-down of Suppressor of Gene Silencing 3 (SGS3) in Rsv1 plants reduced AGO-1 siRNAs leading to a lessened lethal systemic hypersensitive response (LSHR) (Chen et al. 2015). Similarly, the computational analysis identified that G. max-derived miRNAs exhibit propensity to downregulate DNA viruses infecting soybean, viz. Mungbean yellow mosaic India virus and Mungbean yellow mosaic virus transcripts (Ramesh et al. 2016a, b). Expressional changes of the conserved miRNAs, putatively antiviral miRNAs and their target transcripts were reported in soybean genotypes [JS335 (susceptible) and UPSM534 (resistant)] during MYMIV infection (Ramesh et al. 2017). The expression pattern of soybean-derived miRNAs suggests a greater role of argonaute (AGO) homeostasis and regulatory changes in hormonal signaling pathways in conferring virus resistance. Soybean-derived miRNAs with potential antiviral capability also displayed upregulation during MYMIV infection (Ramesh et al. 2017).
Nematode infestation-responsive miRNAs
Soybean cyst nematode (SCN, Heterodera glycines) responsive miRNAs have been identified (Li et al. 2012a, b). Comparative profiling revealed that miRNAs belonging to 40 families were specific to SCN in soybean. The investigation also revealed 364 known G. max miRNAs and 21 novel candidate miRNAs. Among them, around 101 miRNAs belonging to 40 families were SCN responsive. Interestingly, most of the differentially expressed miRNAs were downregulated during SCN infection (Li et al. 2012a, b). A large scale sRNA sequencing effort of soybean cultivars (KS4607-susceptible, and KS4313N-resistant) during SCN infection identified 60 SCN-responsive miRNAs belonging to 25 different miRNA families (Tian et al. 2017). Some legume-specific miRNAs such as miR1510, miR2109, miR2118, miR4996, and miR1509 were found abundant along with conserved miRNAs during SCN infection.
Conserved miRNAs and stress responsiveness
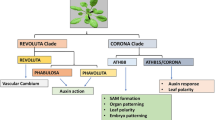
Conserved miRNAs not only share sequence homology but also analogous target characteristic features. Moreover, conserved miRNAs have been evolutionarily selected for orchestrating plant developmental processes by regulating TFs or family of proteins. It was also proposed that conserved miRNAs have acquired supplementary functions in due course of evolution. The phenomenon of conserved miRNAs mediated cross-adaptation has been proposed to account for plant’s capability to concurrently adapt for various biotic and abiotic stresses (Chen et al. 2012). To decode a common molecular pattern of stress responsive miRNAs of soybean, expression status of conserved miRNAs was analyzed (Fig. 2). It is evident that soybean-derived conserved miRNAs form a predominant gene regulatory mechanism countering both abiotic and biotic stresses (Fig. 2). Conserved miRNAs of soybean are generally upregulated during stress except during nitrogen deficiency, phosphorous starvation, rust and MYMIV infection (Fig. 2). Genotypic or varietal differences in the expression of conserved miRNAs of soybean were observed during various stresses (Fig. 2). However, it is unclear why some conserved miRNAs of soybean cultivar are differently regulated under similar stress conditions. Since conserved miRNAs are preserved for their protective function against stresses, considerable variations in their expression status warrant thorough investigation. Also, the advents of robust gene expression profiling systems or ectopic expression techniques have unearthed many non-conserved miRNAs with a potential role in the gene regulatory mechanisms. Brassicaceae-specific non-conserved miRNA, miR163 targets PXMT1and FAMT genes of Arabidopsis involved in secondary metabolite synthesis (Ng et al. 2011), whereas miR400 confers heat tolerance by targeting the target gene PPR (Yan et al. 2012). On the other hand, solanaceous crop-specific miRNAs such as miR482 (Shivaprasad et al. 2012), miR6019 and miR6020 (Li et al. 2012a, b), target NBS-LRR genes which determine pathogen resistance. Thus, it is pertinent to explore the functions of the novel or non-conserved miRNAs of soybean and enhance the miRNA repository of soybean to gain a deeper understanding of molecular stress adaptation strategies. Similarly, most of the legume-specific miRNAs (Fig. 2) are upregulated during abiotic stresses such as drought, salinity and chilling injury and biotic stresses such as Heterodera and MYMIV infection. Interestingly, nutrient toxicity or starvation, symbiosis and fungal or oomycete infections such as rust disease and Phytophthora, respectively, cause downregulation of legume-specific miRNAs. SNF serves as an excellent link between nutrient toxicity and stress and fungal infection as nitrogen fixation in legumes entails both the nutrient supply and pathogen infection process. Hence, a greater understanding of SNF in the small RNA interface might resolve the molecular basis of miRNA downregulation during these stresses.
Expression profile of conserved and legume-specific miRNAs of soybean [color codes: red—downregulated miRNAs, green—upregulated miRNAs, yellow—differentially regulated; a—downregulated in resistant cultivar and upregulated in a susceptible cultivar; b—differentially regulated (up and down); c—variable expression in resistant cultivar; d—downregulated in susceptible cultivar and upregulated in the resistant genotype]
Perspectives and concluding remarks
A decade after the discovery of sncRNAs and their role in RNA silencing of C. elegans (lin-4), plant miRNAs were identified (Reinhart et al. 2002). It is abundantly clear that discovery of plant miRNAs has led to a better understanding of complex gene regulatory mechanisms including molecular events associated with stress tolerance. Technological advances such as miRNA array platform have helped effortless miRNAs profiling in various plant species and delineate the stress-induced gene regulatory networks (Jia et al. 2010). EST-based homology analysis identified 262 candidate miRNAs belonging (143 miRNA families) in faba bean, suggesting the utility of in silico tools in characterizing sRNAome of economically important legumes even when genome sequences were not publicly available (Koptekin and Aktas 2016). The advent of robust and sensitive NGS platforms has helped in defining even very low copy number miRNAs but having a potential role in plant stress. Contrary to the established notion, that conserved miRNAs play a profound role in stress responsiveness, 13 non-conserved miRNAs and seven novel miRNAs are aluminum stress responsive in a wild-type soybean (Glycine soja) (Zeng et al. 2012). Both conserved (gma-miR156b/GmSPL9a) and species-specific (gma-miR4413b/GmPPR) miRNA-target pairs have been implicated in the development of floral organs of soybean and with a potential application for cytoplasmic male sterility (CMS) (Ding et al. 2019). Furthermore, Brassicaceae and Solanaceous-specific miRNAs have been implicated in secondary metabolism and biotic stress, respectively, suggesting yet unravelled features of sRNA-mediated gene regulation.
Despite the importance of miRNAs in gene regulatory responses to biotic stressors, the involvement of soybean-derived miRNAs in agriculturally important trait such as insect infestation is lacking. However, insect herbivory induced miRNA expressional changes are not uncommon as both species-specific and conserved miRNAs were unearthed in Cucumis melo (Sattar et al. 2012) and Chrysanthemum (Xia et al. 2015). Hence, profiling of soybean miRNAs during Aphis glycines infestation might provide sRNA biomarkers associated with the insect resistance. Although transcriptome analysis of soybean roots subjected to flooding stress was made (Nanjo et al. 2011), no studies have been conducted to ascertain the importance of soybean-derived miRNAs in flooding tolerance. Similarly, a comprehensive understanding of the role of legume and/or soybean-derived miRNAs during SNF is relevant to develop legume genotypes tolerant to environmental stresses. Molecular mechanism underlying miR172-mediated enhancement of soybean nodulation (Yan et al. 2013) has led to the development of synthetic miRNA peptides (miPEPS)-based crop production approach. Application of synthetic miPEP172c in soybean mimicked the effects of ectopic over-expression of gma-miR172c (Couzigou et al. 2016). The miR172 of L. japonicus has been shown to regulate AP2 (APETALA2-type) TFs (Holt et al. 2015). The participation of conserved miRNAs, such as miR172, in SNF suggests that these miRNAs might have evolved from non-symbiotic contexts such as core growth and developmental processes but have attained the requisite functional diversification during the course of evolution. Central regulatory roles of miRNAs in SNF, and the nexus of miRNA-NBS-LRR genes-phasiRNAs divulge that legume sRNA transcriptomics is an intriguing area of research. Identification of three new DCLs in M. truncatula and alternative splicing of MtDCL1 mRNA provide greater insights to legume sRNA transcriptomics (Tworak et al. 2016). Upregulation of MtDCL2b and MtDCL4 in nodules and flg22 treatment further suggests the shared gene regulatory networks of miRNAs in controlling SNF and pathogen-induced immunity (Tworak et al. 2016). Similarly, miRNA–phasiRNA mediated gene regulation has gained much attention, especially in legumes, because some protein-coding genes such as NBS-LRRs have been shown to generate miRNA-triggered phasiRNAs (Zhai et al. 2011; Shivaprasad et al. 2012; Li et al. 2012a, b; Fei et al. 2013). Drought-responsive legume miRNA, miR1514a targets two NAC TFs in Phaseolus vulgaris leading to production of phasiRNAs from a NAC transcript (Phvul.010g120700), suggesting the significance of miRNA–phasiRNA based gene regulatory networks in abiotic stress (Sosa-Valencia et al. 2017). Complementing the findings in Phaseolus vulgaris, M. truncatula-derived miRNA (Mtr-miR1514a) has been shown to be involved in targeting NAC TF and generation of phasiRNAs. Thus, elucidation of the functional significance of such conserved phenomenon in legumes will help in identifying universal biomarkers for engineering drought-tolerant legume crops. Discovery of 60 phasiRNA loci in chickpea (Srivastava et al. 2015), 125 loci (among them 47 were shown to be triggered by miRNAs) in Phaseolus vulgaris (Formey et al. 2015) and their targets provide a comprehensive resource for comparative analysis in legumes to decipher the miRNA–phasiRNA nexus.
Identification and characterization of novel miRNA-based biomarkers would not only help in defining stress regulatory networks but also to develop new molecular tools to impart stress tolerance in plants. The greatest challenge in this arena of research is to assign unambiguous functions to the stress-responsive miRNAs (Ni et al. 2012). Molecular tools such as miRNA arrest, target mimicry and decoy miRNAs are most valuable in deciphering the functions of target transcripts. The loss-of-function analysis of miRNAs using the powerful clustered regularly interspaced short palindromic repeats and CRISPR associated protein 9 (CRISPR–Cas9) based nuclease systems not only provided novel means of miRNA modulation but also insights into the regulatory roles of miRNAs (Zhou et al. 2017). Transgenic expression of candidate miRNAs and their effects on the target genes are crucial for utilization of these miRNA-based biomarkers in crop improvement. Towards this direction, soybean miRNA functional network (miRFN) on a system-wide level is an important addition in defining soybean sncRNA transcriptomics (http://nclab.hit.edu.cn/SoymiRNet) (Xu et al. 2014). Also, miRNAs have been proposed as a potential molecular marker (Fu et al. 2013). The applicability of miRNA-microsatellite (miRNA-SSRs) markers developed from M. truncatula was studied in other legume crops including soybean, wherein 77.5% of the 169 primer pairs showed cross-transferability implying its appropriateness for crop improvement programs (Min et al. 2017).
Genome wide survey of the evolution of MIRs and target genes of soybean during the process of domestication and crop improvement programs have identified that MIRs have high evolutionary rates than miRNA targets. Also, soybean MIRs and miRNA targets showing high expression levels, gene/genome duplications and multiple partners display a little nucleotide divergence. Moreover, it was proposed that the process of domestication and crop improvement has increased similarities among most of the miRNA-target pairs in cultivated genotypes of soybean compared to their counterparts in wild genotypes (Liu et al. 2016). Thus, understanding co-evolution of MIRs (miRNA genes) and their target genes is an important area of research that draws the attention of the biologists.
To further complement the research in plant sRNAs, it is relevant to examine long non-coding RNAs (lncRNAs). Plant lncRNAs define various biological processes such as in response to cold (Swiezewski et al. 2009) and other stresses (Xin et al. 2011; Zhang and Chen 2013; Shuai et al. 2014; Wang et al. 2015; Chen et al. 2018). In particular, soybean-derived lncRNA, ENOD40, had been shown to be involved in nodule organogenesis and development (Yang et al.1993) and its orthologs have been characterized in M. truncatula and Medicago sativa (Crespi et al. 1994). Differential expression analysis of wheat genes and associated lncRNAs has provided a comprehensive transcriptome tool for developing drought-tolerant wheat genotypes (Cagirici et al. 2017a). An interaction network involving wheat stem sawfly (WSS) derived miRNAs, lncRNAs and mRNAs was developed to ascertain typical transcriptome changes of pest that weakens the defense response of wheat (Cagirici et al. 2017b). Interestingly, wheat target mRNAs that are likely to be affected by the WSS-derived miRNAs are involved in the defense mechanism of wheat against insect attacks. Since the lncRNAs act as target mimics of miRNAs (Shuai et al. 2014; Wang et al. 2015; Cagirici et al. 2017a, b), the molecular interaction of lncRNAs and miRNAs and the underlying molecular intricacies are required to be unravelled.
Author contribution statement
All authors drafted various segments of the manuscript. All authors read and approved the final version of the manuscript.
Abbreviations
- AGO:
-
Argonaute
- AM:
-
Arbuscular mycorrhiza
- AP2:
-
APETALA 2
- ARF:
-
Auxin response factor
- ASR:
-
Asian soybean rust
- DCL-1:
-
Dicer-like-1
- DRE:
-
Dehydration responsive element
- ENOD93:
-
Early nodulin 93
- GSS:
-
Genome survey sequence
- hc-siRNAs:
-
Heterochromatic siRNAs
- HEN 1:
-
HUA enhancer 1
- HESO1:
-
HEN1 SUPPRESSOR1
- HST1:
-
HASTY 1
- HYL1:
-
HYPONASTIC LEAVES1
- miRNAs:
-
MicroRNAs
- nat-siRNAs:
-
Natural antisense transcript siRNAs
- NGS:
-
Next generation sequencing
- PEGs:
-
Protein encoding genes
- PTGS:
-
Post transcriptional gene silencing
- RBPs:
-
dsRNA-binding proteins
- RdDM:
-
RNA-dependent DNA methylation
- RISC:
-
RNA-induced silencing complex
- SCN:
-
Soybean cyst nematode
- SE:
-
SERRATE
- siRNAs:
-
Small interfering RNAs
- SMV:
-
Soybean mosaic virus
- sncRNAs:
-
Small non-coding RNAs
- SNF:
-
Symbiotic nitrogen fixation
- TFs:
-
Transcriptional factors
- TGS:
-
Transcriptional gene silencing
References
Abdel-Ghany SE, Pilon M (2008) MicroRNA-mediated systemic downregulation of copper protein expression in response to low copper availability in Arabidopsis. J Biol Chem 283:15932–15945
Akdogan G, Tufekci ED, Uranbey S, Unver T (2016) miRNA-based drought regulation in wheat. Funct Integr Genom 16(3):221–233
Akpinar BA, Budak H (2016) Dissecting miRNAs in wheat D genome progenitor, Aegilops tauschii. Front Plant Sci 7:606
Akpinar BA, Kantar M, Budak H (2015) Root precursors of microRNAs in wild emmer and modern wheats show major differences in response to drought stress. Funct Integr Genom 15(5):587–598
Allen RS, Li J, Stahle MI, Dubroue A, Gubler F, Millar AA (2007) Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc Natl Acad Sci USA 104:16371–16376
Alptekin B, Langridge P, Budak H (2017) Abiotic stress miRNomes in the Triticeae. Funct Integr Genom 17(2–3):145–170
Arenas-Huertero C, Pérez B, Rabanal F, Blanco-Melo D, De la Rosa C, Estrada-Navarrete G, Sanchez F, Covarrubias AA, Reyes JL (2009) Conserved and novel miRNAs in the legume Phaseolus vulgaris in response to stress. Plant Mol Biol 70:385–401
Arikit S, Xia R, Kakrana A, Huang K, Zhai J, Yan Z, Valdés-López O, Prince S, Musket TA, Nguyen HT, Stacey G, Meyers BC (2014) An atlas of soybean small RNAs identifies phased siRNAs from hundreds of coding genes. Plant Cell 26:4584–4601
Axtell MJ, Snyder JA, Bartell DP (2007) Common functions for diverse small RNAs of land plants. Plant Cell 19:1750–1769
Babu M, Gagarinova AG, Brandle JE, Wang A (2008) Association of the transcriptional response of soybean plants with Soybean Mosaic Virus systemic infection. J Gen Virol 89:1069–1080
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism and function. Cell 116:281–297
Bazin J, Khan GA, Combier JP, Bustos-Sanmamed P, Debernardi JM, Rodriguez R, Sorin C, Palatnik J, Hartmann C, Crespi M, Lelandais-Brière C (2013) miR396 affects mycorrhization and root meristem activity in the legume Medicago truncatula. Plant J 74(6):920–934
Brant EJ, Budak H (2018) Plant small non-coding RNAs and their roles in biotic stresses. Front Plant Sci 9:1038. https://doi.org/10.3389/fpls.2018.01038
Brodersen P, Sakvarelidze-Achard L, Bruun-Rasmussen M, Dunoyer P, Yamamoto YY, Sieburth L, Voinnet O (2008) Widespread translational inhibition by plant miRNAs and siRNAs. Science 320:1185–1190
Budak H, Akpinar A (2011) Dehydration stress-responsive miRNA in Brachypodium distachyon: evident by genome-wide screening of microRNAs expression. Omics J Integr Biol 15(11):791–799
Budak H, Akpinar BA (2015) Plant miRNAs: biogenesis, organization and origins. Funct Integr Genom 15(5):523–531
Budak H, Khan Z, Kantar M (2014) History and current status of wheat miRNAs using next-generation sequencing and their roles in development and stress. Brief Funct Genom 14(3):189–198
Budak H, Kantar M, Bulut R, Akpinar BA (2015a) Stress responsive miRNAs and isomiRs in cereals. Plant Sci 235:1–13
Budak H, Hussain B, Khan Z, Ozturk NZ, Ullah N (2015b) From genetics to functional genomics: improvement in drought signaling and tolerance in wheat. Front. Plant Sci 6:1012. https://doi.org/10.3389/fpls.2015.01012
Cagirici HB, Alptekin B, Budak H (2017a) RNA sequencing and co-expressed long non-coding RNA in modern and wild wheats. Sci Rep 7(1):10670
Cagirici HB, Biyiklioglu S, Budak H (2017b) Assembly and annotation of transcriptome provided evidence of miRNA mobility between wheat and wheat stem sawfly. Front Plant Sci 8:1653
Cai Z, Wang Y, Zhu L, Tian Y, Chen L, Sun Z, Ullah I, Li X (2017) GmTIR1/GmAFB3-based auxin perception regulated by miR393 modulates soybean nodulation. New Phytol 215:672–686
Carrington JC, Ambros V (2003) Role of microRNAs in plant and animal development. Science 301:336–338
Cervantes-Gámez RG, Bueno-Ibarra MA, Cruz-Mendívil A, Calderón-Vázquez CL, Ramírez-Douriet CM, Maldonado-Mendoza IE, Villalobos-López MÁ, Valdez-Ortíz Á, López-Meyer M (2016) Arbuscular mycorrhizal symbiosis-induced expression changes in Solanum lycopersicum leaves revealed by RNA-seq analysis. Plant Mol Bio Rep 34(1):89–102
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 25:21–44
Chen L, Wang T, Zhao M, Tian Q, Zhang WH (2012) Identification of aluminum responsive microRNAs in Medicago truncatula by genome-wide high-throughput sequencing. Planta 235:375–386
Chen H, Zhang L, Yu K, Wang A (2015) Pathogenesis of soybean mosaic virus in soybean carrying Rsv1 gene is associated with miRNA and siRNA pathways, and breakdown of AGO1 homeostasis. Virology 476:395–404
Chen H, Arsovski AA, Yu K, Wang A (2016) Genome-wide investigation using sRNA-Seq, degradome-Seq and transcriptome-Seq reveals regulatory networks of microRNAs and their target genes in soybean during soybean mosaic virus infection. PLoS One 11:e0150582
Chen L, Shi S, Jiang N, Khanzada H, Wassan GM, Zhu C, Peng X, Xu J, Chen Y, Yu Q, He X (2018) Genome-wide analysis of long non-coding RNAs affecting roots development at an early stage in the rice response to cadmium stress. BMC Genom 19(1):460
Combier JP, Frugier F, de Billy F, Boualem A, El-Yahyaoui F, Moreau S, Vernie T, Ott T, Gamas P, Crespi M, Niebel A (2006) MtHAP2-1 is a key transcriptional regulator of symbiotic nodule development regulated by microRNA169 in Medicago truncatula. Genes Dev 20:3084–3088
Couzigou JM, André O, Guillotin B, Alexandre M, Combier JP (2016) Use of microRNA-encoded peptide miPEP172c to stimulate nodulation in soybean. New Phytol 211:379–381
Crespi MD, Jurkevitch E, Poiret M, d’Aubenton-Carafa Y, Petrovics G, Kondorosi E, Kondorosi A (1994) enod40, a gene expressed during nodule organogenesis, codes for a non-translatable RNA involved in plant growth. EMBO J 13:5099–5112
Cui X, Yan Q, Gan S, Xue D, Dou D, Guo N, Xing H (2017) Overexpression of gma-miR1510a/b suppresses the expression of a NB-LRR domain gene and reduces resistance to Phytophthora sojae. Gene 20(621):32–39
Devers EA, Branscheid A, May P, Krajinski F (2011) Stars and symbiosis: microRNA-and microRNA*-mediated transcript cleavage involved in arbuscular mycorrhizal symbiosis. Plant Physiol 156:111
Dezulian T, Remmert M, Palatnik JF, Weigel D, Huson DH (2006) Identification of plant microRNA homologs. Bioinformatics 22:359–360
Ding D, Zhang L, Wang H, Liu Z, Zhang Z, Zheng Y (2009) Differential expression of miRNAs in response to salt stress in maize roots. Ann Bot 103:29–38
Ding X, Zhang H, Ruan H, Li Y, Chen L, Wang T, Jin L, Li X, Yang S, Gai J (2019) Exploration of miRNA-mediated fertility regulation network of cytoplasmic male sterility during flower bud development in soybean. 3 Biotech 9:22. https://doi.org/10.1007/s13205-018-1543-1
Dong Z, Shi L, Chen L, Wang Y, Cai Z, Wang Y, Jin J, Li X (2013) Identification and dynamic regulation of microRNAs involved in salt stress responses in functional soybean nodules by high-throughput sequencing. Int J Mol Sci 14:2717–2738
EI Yahyaoui F, Kuster H, Amor BB, Hohnjec N, Puhler A, Becker A, Gouzy J, Vernie T, Gough C, Niebel A, Godiard L, Gamas P (2004) Expression profiling in Medicago truncatula identifies more than 750 genes differentially expressed during nodulation, including many potential regulators of the symbiotic program. Plant Physiol 136:3159–3176
Eldem V, Akcay UC, Ozhuner E, Bakır Y, Uranbey S, Unver T (2012) Genome-wide identification of miRNAs responsive to drought in peach (Prunus persica) by high-throughput deep sequencing. PLoS One 7(12):e50298
Eren H, Pekmezci MY, Okay S, Turktas M, Inal B, Ilhan E, Atak M, Erayman M, Unver T (2015) Hexaploid wheat (Triticum aestivum) root miRNome analysis in response to salt stress. Ann Appl Biol 167(2):208–216
Fang X, Zhao Y, Ma Q, Huang Y, Wang P, Zhang J, Nian H, Yang C (2013) Identification and comparative analysis of cadmium tolerance-associated miRNAs and their targets in two soybean genotypes. PLoS One 8:e81471
Fei Q, Xia R, Meyers BC (2013) Phased, secondary, small interfering RNAs in posttranscriptional regulatory networks. Plant Cell 25:2400–2415
Formey D, Iñiguez LP, Peláez P, Li YF, Sunkar R, Sánchez F, Reyes JL, Hernández G (2015) Genome-wide identification of the Phaseolus vulgaris sRNAome using small RNA and degradome sequencing. BMC Genom 16:423
Fu D, Ma B, Mason AS, Xiao M, Wei L, An Z (2013) Micro RNA-based molecular markers: a novel PCR-based genotyping technique in Brassica species. Plant Breed 132(4):375–381
Gao P, Bai X, Yang L, Lv D, Pan X, Li Y, Cai H, Ji W, Chen Q, Zhu Y (2011) osa-MIR393: a salinity- and alkaline stress related microRNA gene. Mol Biol Rep 38:237–242
Groszhans H, Filipowicz W (2008) Molecular biology: the expanding world of small RNAs. Nature 451:414–416
Guo N, Ye WW, Wu XL, Shen DY, Wang YC, Xing H, Dou DL (2011) Microarray profiling reveals microRNAs involving soybean resistance to Phytophthora sojae. Genome 54(11):954–958
Hajyzadeh M, Turktas M, Khawar KM, Unver T (2015) miR408 over-expression causes increased drought tolerance in chickpea. Gene 555(2):186–193
Hamilton AJ, Baulcombe DC (1999) A species of small antisense RNA in post-transcriptional gene silencing in plants. Science 286:950–952
Han Y, Zheng HU, Zheng D, Gao Y (2014) Analysis of promoters of microRNAs from a Glycine max degradome library. J Zhejiang Univ Sci B (Biomed Biotechnol) 15:125–132
Holt DB, Gupta V, Meyer D, Abel NB, Andersen SU, Stougaard J, Markmann K (2015) Micro RNA 172 (miR172) signals epidermal infection and is expressed in cells primed for bacterial invasion in Lotus japonicus roots and nodules. New Phytopathol 208:241–256
Huang SC, Lu GH, Tang CY, Ji YJ, Tan GS, Hu DQ, Cheng J, Wang GH, Qi JL, Yang YH (2017) Identification and comparative analysis of aluminum-induced microRNAs conferring plant tolerance to aluminum stress in soybean. Biol Plant 62:97–108
Jia X, Mendu V, Tang G (2010) An array platform for identification of stress-responsive microRNAs in plants. Methods Mol Biol 639:253–269
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14(6):787–799
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAs and their regulatory roles in plants. Ann Rev Plant Biol 57:19–53
Kantar M, Lucas SJ, Budak H (2011) miRNA expression patterns of Triticum dicoccoides in response to shock drought stress. Planta 233(3):471–484
Khraiwesh B, Zhu JK, Zhu J (2012) Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim Biophys Acta 1819:137–148
Koptekin D, Aktas LY (2016) Identification of conserved miRNAs and their target genes in faba bean by EST based homology analysis. Eur J Sci Technol 5:1–6
Kulcheski FR, de Oliveira LF, Molina LG, Almerão MP, Rodrigues FA, Marcolino J, Barbosa JF, Stolf-Moreira R, Nepomuceno AL, Marcelino-Guimarães FC, Abdelnoor RV (2011) Identification of novel soybean microRNAs involved in abiotic and biotic stresses. BMC Genom 12:307
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854
Li WX, Oono Y, Zhu J, He XJ, Wu JM, Iida K, Lu XY, Cui X, Jin H, Zhu JK (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and post transcriptionally to promote drought resistance. Plant Cell 20:2238–2251
Li T, Li H, Zhang YX, Liu JY (2010) Identification and analysis of seven H2O2-responsive miRNAs and 32 new miRNAs in the seedlings of rice (Oryza sativa L. ssp. indica). Nucleic Acids Res 39:2821–2833
Li H, Dong Y, Yin H, Wang N, Yang J, Liu X, Wang Y, Wu J, Li X (2011) Characterization of the stress associated microRNAs in Glycine max by deep sequencing. BMC Plant Biol 11:1–12
Li F, Pignatta D, Bendix C, Brunkard JO, Cohn MM, Tung J, Sun H, Kumar P, Baker B (2012a) MicroRNA regulation of plant innate immune receptors. Proc Nat Acad Sci USA 109:1790–1795
Li X, Wang X, Zhang S, Liu D, Duan Y, Dong W (2012b) Identification of soybean microRNAs involved in soybean cyst nematode infection by deep sequencing. PLoS One 7:e39650
Liu HH, Tian X, Li YJ, Wu CA, Zheng CC (2008) Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 14:836–843
Liu T, Fang C, Ma Y, Shen Y, Li C, Li Q, Wang M, Liu S, Zhang J, Zhou Z, Yang R (2016) Global investigation of the co-evolution of MIRNA genes and microRNA targets during soybean domestication. Plant J 85:396–409
Llave C, Kasschau KD, Rector MA, Carrington JC (2002) Endogenous and silencing-associated small RNAs in plants. Plant Cell 14:1605–1619
Lu XY, Huang XL (2008) Plant miRNAs and abiotic stress responses. Biochem Biophys Res Commun 368:458–462
Mallory AC, Vaucheret H (2006) Functions of microRNAs and related small RNAs in plants. Nat Gen 38:S31–S36
Meyers BC, Axtell MJ, Bartel B, Bartel DP, Baulcombe D, Bowman JL, Cao X, Carrington JC, Chen X, Green PJ, Griffiths-Jones S, Jacobsen SE, Mallory AC, Martienssen RA, Poethig RS, Qi Y, Vaucheret H, Voinnet O, Watanabe Y, Weigel D, Zhu JK (2008) Criteria for annotation of plant microRNAs. Plant Cell 20:3186–3190
Min X, Zhang Z, Liu Y, Wei X, Liu Z, Wang Y, Liu W (2017) Genome-wide development of microRNA-based SSR markers in Medicago truncatula with their transferability analysis and utilization in related legume species. Int J Mol Sci 18(11):2440
Miransari M (2015) Abiotic and biotic stresses in soybean production. Soybean Prod 1:4. https://doi.org/10.1016/b978-0-12-801536-0.00014-1(ISBN: 978-0-12-801536-0)
Mitsuda N, Ohme-Takagi M (2009) Functional analysis of transcription factors in Arabidopsis. Plant Cell Physiol 50:1232–1248
Mochida K, Yoshida T, Sakurai T, Yamaguchi-Shinozaki K, Shinozaki K, Tran LSP (2010) LegumeTFDB: an integrative database of Glycine max, Lotus japonicus and Medicago truncatula transcription factors. Bioinformatics 26(2):290–291
Nageshbabu R, Usha Jyothi MN, Sharadamma N (2013) Expression of miRNAs confers enhanced tolerance to drought and salt stress in finger millet (Eleusine coracana). J Stress Physiol Biochem 9:220–231
Nanjo Y, Maruyama K, Yasue H, Yamaguchi-Shinozaki K, Shinozaki K, Komatsu S (2011) Transcriptional responses to flooding stress in roots including hypocotyl of soybean seedlings. Plant Mol Biol 77:129–144
Navarro L, Dunoyer P, Jay F, Arnold B, Dharmasiri N, Estelle M, Voinnet O (2006) A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312:436–439
Ng DW, Zhang C, Miller M, Palmer G, Whiteley M, Tholl D, Chen ZJ (2011) cis- and trans-regulation of miR163 and target genes confers natural variation of secondary metabolites in two Arabidopsis species and their allopolyploids. Plant Cell 23:1729–1740
Ni Z, Hu Z, Jiang Q, Zhang H (2012) Over-expression of gma-MIR394a confers tolerance to drought in transgenic Arabidopsis thaliana. Biochem Biophys Res Commun 427:330–335
Ni Z, Hu Z, Jiang Q, Zhang H (2013) GmNFYA3, a target gene of miR169, is a positive regulator of plant tolerance to drought stress. Plant Mol Biol 82:113–129
Pant BD, Buhtz A, Kehr J, Scheible WR (2008) microRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J 53:731–738
Phillips JR, Dalmay T, Bartels D (2007) The role of small RNAs in abiotic stress. FEBS Lett 581:3592–3597
Quach TN, Nguyen HT, Valliyodan B, Joshi T, Xu D, Nguyen HT (2015) Genome-wide expression analysis of soybean NF-Y genes reveals potential function in development and drought response. Mol Genet Genom 290:1095–1115
Ramesh SV, Admane N, Husain SM (2013) Small RNAs landscape (sRNAome) of soybean [Glycine max (L.)]: biogenesis, vital functions and potential applications. Plant Knowl J 2:24–37
Ramesh SV, Ratnaparkhe MB, Kumawat G, Gupta GK, Husain SM (2014) Plant miRNAome and antiviral resistance: a retrospective view and prospective challenges. Virus Genes 48:1–14
Ramesh SV, Ratnaparkhe MB, Husain SM, Bhatia VS (2015) Viral micro RNA transcriptomics (miRNAomics). Transcriptomics 3:108
Ramesh SV, Chouhan BS, Kumar G, Praveen S (2016) Soybean derived miRNAs and Mungbean yellow mosaic India virus resistance. In: 16th biennial conference of the molecular and cellular biology of the soybean conference, Columbus, OH, August 7–10, 2016
Ramesh SV, Gupta GK, Husain SM (2016b) Soybean (Glycine max) miRNAs display proclivity to repress Begomovirus genomes. Curr Sci 110:424–428
Ramesh SV, Chouhan BS, Gaurav K, Praveen S, Chand S (2017) Expression dynamics of Glycine max (L.) Merrill derived microRNAs (miRNAs) and their targets during Mungbean yellow mosaic India virus (MYMIV) infection. Physiol Mol Plant Pathol 100:13–22
Reinhart BJ, Weinstein EG, Rhoades MW, Bartel B, Bartel DP (2002) MicroRNAs in plants. Genes Dev 16:1616–1626
Reyes JL, Chua NH (2007) ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant J 49:592–606
Rhoades M, Reinhart B, Lim L, Burge C, Bartel B, Bartel D (2002) Prediction of plant microRNA targets. Cell 110:513–520
Sahito ZA, Wang L, Sun Z, Yan Q, Zhang X, Jiang Q, Ullah I, Tong Y, Li X (2017) The miR172c-NNC1 module modulates root plastic development in response to salt in soybean. BMC Plant Biol 17(1):229
Sattar S, Song Y, Anstead JA, Sunkar R, Thompson GA (2012) Cucumis melo microRNA expression profile during aphid herbivory in a resistant and susceptible interaction. Mol Plant Microbe Interact 25:839–848
Shivaprasad PV, Chen HM, Patel K, Bond DM, Santos BA, Baulcombe DC (2012) A microRNA superfamily regulates nucleotide binding site-leucine-rich repeats and other mRNAs. Plant Cell 24:859–874
Shuai P, Liang D, Tang S, Zhang Z, Ye CY, Su Y, Xia X, Yin W (2014) Genome-wide identification and functional prediction of novel and drought-responsive lincRNAs in Populus trichocarpa. J Exp Bot 65(17):4975–4983
Soares-Cavalcanti NM, Belarmino LC, Kido EA, Pandolfi V, Marcelino-Guimarães FC, Rodrigues FA, Pereira GA, Benko-Iseppon AM (2012) Overall picture of expressed Heat Shock Factors in Glycine max, Lotus japonicus and Medicago truncatula. Genet Mol Biol 35(1):247–259
Sosa-Valencia G, Palomar M, Covarrubias AA, Reyes JL (2017) The legume miR1514a modulates a NAC transcription factor transcript to trigger phasiRNA formation in response to drought. J Exp Bot 68:2013–2026
Srivastava S, Zheng Y, Kudapa H, Jagadeeswaran G, Hivrale V, Varshney RK, Sunkar R (2015) High throughput sequencing of small RNA component of leaves and inflorescence revealed conserved and novel miRNAs as well as phasiRNA loci in chickpea. Plant Sci 235:46–57
Subramanian S, Fu Y, Sunkar R, Barbazuk WB, Zhu JK, Yu O (2008) Novel and nodulation-regulated microRNAs in soybean roots. BMC Genom 9:160
Sunkar R (2010) MicroRNAs with macro-effects on plant stress responses. Sem Cell Dev Biol 21:805–811
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16:2001–2019
Swiezewski S, Liu F, Magusin A, Dean C (2009) Cold-induced silencing by long antisense transcripts of an Arabidopsis Polycomb target. Nature 462(7274):799
Tian B, Wang S, Todd TC, Johnson CD, Tang G, Trick HN (2017) Genome-wide identification of soybean microRNA responsive to soybean cyst nematodes infection by deep sequencing. BMC Genom 18:572
Trindade I, Capitao C, Dalmay T, Fevereiro MP, Santos DM (2010) miR398 and miR408 are upregulated in response to water deficit in Medicago truncatula. Planta 231:705–716
Turner M, Yu O, Subramanian S (2012) Genome organization and characteristics of soybean microRNAs. BMC Genom 13:169
Turner M, Nizampatnam NR, Baron M, Coppin S, Damodaran S, Adhikari S, Arunachalam SP, Yu O, Subramanian S (2013) Ectopic expression of miR160 results in auxin hypersensitivity, cytokinin hyposensitivity, and inhibition of symbiotic nodule development in soybean. Plant Physiol 162:2042–2055
Tworak A, Urbanowicz A, Podkowinski J, Kurzynska-Kokorniak A, Koralewska N, Figlerowicz M (2016) Six Medicago truncatula Dicer-like protein genes are expressed in plant cells and upregulated in nodules. Plant Cell Rep 35:1043–1052
Wang Y, Li P, Cao X, Wang X, Zhang A, Li X (2009) Identification and expression analysis of miRNAs from nitrogen-fixing soybean nodules. Biochem Biophys Res Commun 378:799–803
Wang Y, Zhang C, Hao Q, Sha A, Zhou R, Zhou X, Yuan L (2013) Elucidation of miRNAs-mediated responses to low Nitrogen stress by deep sequencing of two soybean genotypes. PLoS One 8:e67423
Wang J, Yu W, Yang Y, Li X, Chen T, Liu T, Ma N, Yang X, Liu R, Zhang B (2015) Genome-wide analysis of tomato long non-coding RNAs and identification as endogenous target mimic for microRNA in response to TYLCV infection. Sci Rep 5:16946
Wei L, Zhang D, Xiang F, Zhang Z (2009) Differentially expressed miRNAs potentially involved in the regulation of defense mechanism to drought stress in maize seedlings. Int J Plant Sci 170:979–989
Wong J, Gao L, Yang Y, Zhai J, Arikit S, Yu Y, Duan S, Chan V, Xiong Q, Yan J, Li S, Liu R, Wang Y, Tang G, Meyers BC, Chen X, Ma W (2014) Roles of small RNAs in soybean defense against Phytophthora sojae infection. Plant J 79:928–940
Wu L, Zhou H, Zhang Q, Zhang J, Ni F, Liu C, Qi Y (2010) DNA methylation mediated by a microRNA pathway. Mol Cell 38:465–475
Xia K, Wang R, Ou X, Fang Z, Tian C, Duan J, Wang Y, Zhang M (2012) OsTIR1 and OsAFB2 downregulation via OsmiR393 over-expression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS One 7:1–10
Xia X, Shao Y, Jiang J, Du X, Sheng L, Chen F, Fang W, Guan Z, Chen S (2015) MicroRNA expression profile during Aphid feeding in Chrysanthemum (Chrysanthemum morifolium). PLoS One 10:e0143720
Xin M, Wang Y, Yao Y, Song N, Hu Z, Qin D, Xie C, Peng H, Ni Z, Sun Q (2011) Identification and characterization of wheat long non-protein coding RNAs responsive to powdery mildew infection and heat stress by using microarray analysis and SBS sequencing. BMC Plant Biol 11(1):61
Xu MY, Zhang L, Li WW, Hu X, Wang MB, Fan YL, Zhang CY, Wang L (2014) Stress-induced early flowering is mediated by miR169 in Arabidopsis thaliana. J Exp Bot 65:89–101
Xu S, Liu N, Mao W, Hu Q, Wang G, Gong Y (2016) Identification of chilling-responsive microRNAs and their targets in vegetable soybean (Glycine max L.). Sci Rep 6:26619
Xu Y, Zhu S, Liu F, Wang W, Wang X, Han G, Cheng B (2018) Identification of Arbuscular Mycorrhiza fungi responsive microRNAs and their regulatory network in maize. Int J Mol Sci 19:3201
Yan K, Liu P, Wu CA, Yang GD, Xu R, Guo QH, Haung JG, Zheng CC (2012) Stress-induced alternative splicing provides a mechanism for the regulation of microRNA processing in Arabidopsis thaliana. Mol Cell 48:521–531
Yan Z, Hossain MS, Wang J, Valdés-López O, Liang Y, Libault M, Qiu L, Stacey G (2013) miR172 regulates soybean nodulation. Mol Plant Microbe Interact 26(12):1371–1377
Yan Z, Hossain MS, Arikit S, Valdés-López O, Zhai J, Wang J, Libault M, Ji T, Qiu L, Meyers BC, Stacey G (2015) Identification of microRNAs and their mRNA targets during soybean nodule development: functional analysis of the role of miR393j-3p in soybean nodulation. New Phytol 207:748–759
Yang WC, Katinakis P, Hendriks P, Smolders A, de Vries F, Spee J, van Kammen A, Bisseling T, Franssen H (1993) Characterization of GmENOD40, a gene showing novel patterns of cell-specific expression during soybean nodule development. Plant J 3:573–585
Yin X, Wang J, Cheng H, Wang X, Yu D (2013) Detection and evolutionary analysis of soybean miRNAs responsive to Soybean Mosaic Virus. Planta 237:1213–1225
Zeng HQ, Zhu YY, Huang SQ, Yang ZM (2010) Analysis of phosphorus-deficient responsive miRNAs and cis-elements from soybean (Glycine max L.). J Plant Physiol 167:1289–1297
Zeng QY, Yang CY, Ma QB, Li XP, Dong WW, Nian H (2012) Identification of wild soybean miRNAs and their target genes responsive to aluminum stress. BMC Plant Biol 12:1
Zhai J, Jeong DH, De Paoli E, Park S, Rosen BD, Li Y, González AJ, Yan Z, Kitto SL, Grusak MA, Jackson S (2011) MicroRNAs as master regulators of the plant NB-LRR defense gene family via the production of phased, trans-acting siRNAs. Genes Dev 25:2540–2553
Zhang YC, Chen YQ (2013) Long noncoding RNAs: new regulators in plant development. Biochem Biophys Res Commun 436:111–114
Zhang B, Wang Q (2016) MicroRNA, a new target for engineering new crop varieties. Bioengineered 7:7–10
Zhang BH, Pan XP, Wang QL, Cobb GP, Anderson TA (2005) Identification and characterization of new plant microRNAs using EST analysis. Cell Res 15:336–360
Zhang Z, Wei L, Zou X, Tao Y, Liu Z, Zheng Y (2008) Submergence-responsive MicroRNAs are potentially involved in the regulation of morphological and metabolic adaptations in maize root cells. Ann Bot 102:509–519
Zhang L, Chia JM, Kumari S, Stein JC, Liu Z, Narechania A, Maher CA, Guill K, McMullen MD, Ware D (2009) A genome-wide characterization of microRNA genes in maize. PLoS Genet 5:e1000716
Zhang S, Wang Y, Li K, Zou Y, Chen L, Li X (2014) Identification of cold-responsive miRNAs and their target genes in nitrogen-fixing nodules of soybean. Int J Mol Sci 15:13596–13614
Zhao BT, Liang RQ, Ge LF, Li W, Xiao HS, Lin HX, Ruan KC, Jin YX (2007) Identification of drought-induced microRNAs in rice. Biochem Biophys Res Commun 354:585–590
Zhao B, Ge L, Liang R, Li W, Ruan K, Lin H, Jin Y (2009) Members of miR-169 family are induced by high salinity and transiently inhibit the NF-YA transcription factor. BMC Mol Biol 10:29
Zhao Y, Yu Y, Zhai J, Ramachandran V, Dinh TT, Meyers BC, Mo B, Chen X (2012) The Arabidopsis nucleotidyl transferase HESO1 uridylates unmethylated small RNAs to trigger their degradation. Curr Biol 22:689–694
Zhao M, Cai C, Zhai J, Lin F, Li L, Shreve J, Thimmapuram J, Hughes TJ, Meyers BC, Ma J (2015a) Coordination of microRNAs, phasiRNAs, and NBS-LRR Genes in response to a plant pathogen: insights from analyses of a set of soybean Rps gene near-isogenic lines. Plant Genome 8:1
Zhao M, Meyers BC, Cai C, Xu W, Ma J (2015b) Evolutionary patterns and co-evolutionary consequences of MIRNA genes and microRNA targets triggered by multiple mechanisms of genomic duplications in soybean. Plant Cell 27:546–562
Zheng Y, Hivrale V, Zhang X, Valliyodan B, Lelandais-Brière C, Farmer AD, May GD, Crespi M, Nguyen HT, Sunkar R (2016) Small RNA profiles in soybean primary root tips under water deficit. BMC Syst Biol 10:126
Zhou L, Liu Y, Liu Z, Kong D, Duan M, Luo L (2010) Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J Exp Bot 61:4157–4168
Zhou J, Deng K, Cheng Y, Zhong Z, Tian L, Tang X, Tang A, Zheng X, Zhang T, Qi Y, Zhang Y (2017) CRISPR-Cas9 based genome editing reveals new insights into microRNA function and regulation in rice. Front Plant Sci 8:1598
Acknowledgements
This study was funded by Indian Council of Agricultural Research (ICAR)-Indian Institute of Soybean Research (ICAR-IISR) sponsored project (Grant no.: 1.24/12).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that there are no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ramesh, S.V., Govindasamy, V., Rajesh, M.K. et al. Stress-responsive miRNAome of Glycine max (L.) Merrill: molecular insights and way forward. Planta 249, 1267–1284 (2019). https://doi.org/10.1007/s00425-019-03114-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-019-03114-5