Abstract
Main conclusion
Phytosterol homeostasis may be maintained in leaves through diversion of intermediates into glycoalkaloid biosynthesis, whereas in tuber flesh, excess intermediates are catalyzed by tuber-specific StLAS - like , resulting in low tuber glycoalkaloids.
Lanosterol synthase (LAS) and cycloartenol synthase (CAS) are phylogenetically related enzymes. Cycloartenol is the accepted precursor leading to cholesterol and phytosterols, and in potato, to steroidal glycoalkaloid (SGA) biosynthesis. LAS was also shown to synthesize some plant sterols, albeit at trace amounts, questioning its role in sterol homeostasis. Presently, a potato LAS-related gene (StLAS-like) was identified and its activity verified in a yeast complementation assay. A transgenic approach with targeted gene expression and metabolic profiling of sterols and SGAs was used. Analyses of StLAS-like transcript levels and StLAS-like-promoter::GUS reporter assays indicated specific expression in tuber flesh tissue. Overexpression of Arabidopsis AtLAS in leaves where the endogenic StLAS-like is not expressed, resulted with increased SGA level and reduced phytosterol level, while in the tuber flesh SGA level was reduced. StLAS-like expression only in tuber flesh may explain the differential accumulation of SGAs in commercial cultivars—low in tubers, high in leaves. In leaves, to maintain phytosterol homeostasis, an excess of intermediates may be diverted into SGA biosynthesis, whereas in tuber flesh these intermediates are catalyzed by tuber-specific StLAS-like instead, resulting in low levels of SGA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
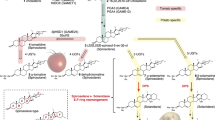
Phytosterols, brassinosteroid hormones, and the steroidal glycoalkaloids (SGAs)—the latter unique to the Solanaceae and the Melanthiaceae—are biosynthesized by diverging pathways of the sterol branch of the mevalonate/isoprenoid pathway (Fig. 1a). Enzyme activities located at biosynthesis branch points, such as cycloartenol synthase (CAS) and lanosterol synthase (LAS), are important for studying precursor channeling in response to growth and environmental stimuli.
a Schematic representation of the mevalonic/isopropenoid pathway and the diverting branches of phytosterol (sitosterol, stigmasterol and campesterol) and steroidal glycoalkaloids (α-solanine and α-chaconine) biosynthetic pathways. Arrows represent enzymatic steps, double arrows represent several enzymatic steps. Only enzymes that are discussed in the current work are indicated in bold along the pathway; the corresponding gene names are italicized. The LAS/CAS branch point is highlighted in grey. HMG1, 3-hydroxy-3-methylglutaryl-CoA reductase; VS1, vetispiradiene/sesquiterpene synthase; SQS1, squalene synthase; SQE1, squalene epoxidase; LAS1, lanosterol synthase; CAS1, cycloartenol synthase; SMT1, sterol C24-methyltransferase type 1; CYP51G1, obtusifoliol 14-α-demethylase; FK, fackel delta(14)-sterol reductase; SMT2, sterol C24-methyltransferase type 2; SSR1/DWF1, Diminuto/Dwarf1 delta(24)-sterol reductase; SSR2, sterol side chain reductase 2; C5-SD, delta(7)-sterol-C5(6)-desaturase; GAME, GLYCOALKALOID METABOLISM; GAME8, a member of the CYP72 subfamily of cytochrome P450s; GAME11, diogenase; GAME4, a member of the 88D subfamily of cytochrome P450; GAME12, transaminase. SGT1, UDP-galactose:solanidine galactosyltransferase; SGT2, UDP-glucose:solanidine glucosyltransferase; SGT3, β-solanine/β-chaconine rhamnosyltransferase. Based on: http://www.genome.jp/kegg-bin/show_pathway?ath00100; Clouse and Sasse 1998; Laule et al. 2003; Benveniste 2004; Itkin et al. 2013; Sawai et al. 2014. b Conversion of 2,3-oxidosqualene by lanosterol synthase (LAS) or cycloartenol synthase (CAS) to lanosterol or cycloartenol, respectively. c Multiple alignment of amino acid residues around the critical region for the reaction specificity of oxidoreductases (based on Suzuki et al. 2006). Arabidopsis CAS1 (At2g07050), S. phureja putative CAS1 (PGSC0003DMB000000420:58827600..58845400), and StCAS isolated by us from S. tuberosum cv. Desirée (accession no. KU313680; upper panel) and Arabidopsis LAS1 (At3g45130), S. phureja putative LAS1 (PGSC0003DMB000000328:43281700..43271000), and StLAS-like isolated by us from Desirée (accession No. KU313679; lower panel). Asterisks denote amino acid substitution in which in Arabidopsis CAS1 altered the catalytic function to LAS1 (Meyer et al. 2002; Segura et al. 2002; Lodeiro et al. 2005; Kolesnikova et al. 2006). Amino acid residues conserved in oxidosqualene cyclases (OSCs) are highlighted in grey; residues conserved in LAS are framed with rectangles. d Functional complementation of yeast mutant GIL77 deficient for ERG7 activity by the OSCs StLAS-like and StCAS from potato and AtLAS from Arabidopsis. GIL77 transformed with ERG7/YHR072W or pYES2 plasmids served as positive and negative controls, respectively. Plate medium contained SC and galactose
The enzyme, 3-hydroxy-3-methylglutaryl coenzyme-A reductase (HMGR), catalyzes the first committed step specific to isoprenoid biosynthesis in the cytosol (Fig. 1a, Suzuki and Muranaka 2007; Nes 2011). Three C5 isoprenoid subunits condense to form farnesyl diphosphate, which is used by terpene synthases to form sesquiterpenoids, and through a tail to tail condensation is used by squalene synthase (SQS1) in the formation of the triterpene squalene. The epoxide at C2 and C3 of squalene, catalyzed by squalene epoxidase (SQE1), facilitates distinct cation-driven cyclization of the C30 hydrocarbon by product-specific oxidosqualene cyclases (OSCs), such as various sesquiterpene synthases, CAS, and LAS (Fig. 1b). These OSCs constitute a significant metabolic branch point that is the initial step of triterpene chemical diversity.
Cycloartenol leads to cholesterol and other sterol (sitosterol, stigmasterol, campesterol) biosynthesis in higher plants (Rahier 2011). Cholesterol is the accepted precursor for SGA biosynthesis (Heftmann 1983; Bergenstråhle et al. 1996; Itkin et al. 2013; Petersson et al. 2013) and is synthesized from cycloartenol via the activity of a sterol side chain reductase 2 (SSR2) (Sawai et al. 2014) and a delta(7)-sterol-C5(6)-desaturase (C5-SD) (Cárdenas et al. 2016). In parallel, a sterol C-24-methyltransferase type 1 (SMT1) directs cycloartenol to phytosterol biosynthesis rather than cholesterol formation (Diener et al. 2000), and SMT1-overexpressing potato plants exhibited reduced cholesterol and SGAs (Arnqvist et al. 2003). The phytosterol biosynthetic pathway has been studied extensively (Benveniste 2004), whereas the biosynthetic pathway from cholesterol to the SGA aglycone has only recently been resolved and includes hydroxylation, oxidation, and transamination steps catalyzed by a series of enzymes encoded by several GLYCOALKALOID METABOLISM (GAME) genes (Fig. 1a) that are organized in metabolic gene clusters in the potato and tomato genomes (Itkin et al. 2013). The final reaction in the biosynthesis of potato SGAs is the glycosylation of the solanidine aglycone by solanidine glycosyltransferases, SGT1, SGT2, and SGT3 (Moehs et al. 1997; McCue et al. 2005, 2007).
Whereas cycloartenol leads to phytosterol biosynthesis in higher plants, lanosterol serves a similar function in animals and fungi but has only been sporadically identified in plants (Kolesnikova et al. 2006; Sawai et al. 2006; Suzuki et al. 2006). The genes encoding CAS and LAS are phylogenetically related; sequence alignment of plant LAS1 and CAS1 indicates that LAS emerged from an ancestral CAS in different lineages by divergent evolution (Sawai et al. 2006). The corresponding genes in Arabidopsis (AGI code no. At3g45130 for AtCAS1 and At2g07050 for AtLAS1) encode proteins with two critically different amino acids in their catalytic domain; the LAS1 enzyme has His477Asn and Ile481Val substitutions relative to the CAS1 enzyme (Fig. 1c), changes that specifically increase the biosynthesis of lanosterol relative to cycloartenol (Meyer et al. 2002; Segura et al. 2002; Lodeiro et al. 2005; Kolesnikova et al. 2006; Sawai et al. 2006).
The presence of LAS1 in Arabidopsis raised the possibility that some plant sterols could be formed from lanosterol—radiotracer studies demonstrated the conversion of lanosterol into sitosterol in Arabidopsis leaves, albeit at trace levels; the conversion of lanosterol into other phytosterols probably occurred but was below the limits of detection (Ohyama et al. 2009). It has been suggested that lanosterol-derived metabolites may be involved in plant defense responses (Kolesnikova et al. 2006; Suzuki et al. 2006; Ohyama et al. 2009). Thus, it may be hypothesized that the lanosterol pathway functions, in part, parallel to the CAS pathway, contributing to sitosterol synthesis or to steroidal-derived secondary metabolites such as the SGAs. In support of that, lanosterol has been found in leaves and sprouts of potato (Stankovic et al. 1990) and was shown to be incorporated into the SGAs of Solanum chacoense (Ripperge et al. 1971).
Potato is a suitable model plant to study the CAS/LAS branch point, as downstream diverting pathways synthesize both the primary metabolites of the phytosterols and the secondary metabolites of the SGAs (that are lacking in Arabidopsis), and which may also be co-regulated. The predominant SGAs in the potato, α-chaconine and α-solanine, can be toxic to humans if their content in tubers exceeds 200 mg/kg fresh weight (FW) (Valkonen et al. 1996). Thus, although the SGAs have organoleptic properties that contribute to the potato flavor, their content in the tuber is of concern in human health, which adds more impact to such study.
The above simplified description of the respective pathways requires clarification regarding the involvement of LAS in the metabolism of steroidal metabolites. In the present study, we identified a putative LAS gene from potato (StLAS-like) whose OSC activity was verified in a yeast complementation assay. Analyses of StLAS-like transcript levels and StLAS-like::GUS reporter assays indicated different spatial expression in potato. Overexpression of the Arabidopsis LAS in transgenic potato modified phytosterol and SGA metabolite levels and the transcript levels of the respective biosynthetic genes.
Materials and methods
Plant material
The experiments were carried out with Solanum tuberosum L. cv. Desirée obtained from commercial sources. Plants were grown in 10 L pots in a greenhouse under natural winter/spring conditions (average temperature range of 10–20 °C). Young leaves and stolons were collected during the pre-flowering growth stage. Mature tubers were harvested 80 days after planting and were kept in paper bags to prevent light exposure. Tuber periderm was separated from the flesh using a scalpel. All tissue samples were frozen immediately in liquid nitrogen and kept at −80 °C until use.
Potato transformation
Transformation was performed with leaves from Desirée plantlets that were maintained under tissue culture conditions. Leaves (100–150) were collected from 4 to 5 week old plantlets that were grown on Nitsch medium (Nitsch and Nitsch 1969) (Duchefa Biochemie V.B., Haarlem, The Netherlands). Leaf margins were cut from all sides and the resulting leaf pieces were kept in sterile distilled water until all leaves were ready for co-cultivation with Agrobacterium. The leaf pieces were incubated for 15–20 min in a culture (0.5 A600nm) of Agrobacterium tumefaciens EHA 105 that carried the required plasmid for transformation, blotted dry and incubated for 2 days in the dark on basal medium [BM, Murashige and Skoog (1962) medium with 30-g/L sucrose and 8-g/L agar, pH = 5.8] supplemented with 0.2-mM acetosyringone (Sigma-Aldrich, St. Louis, MO, USA). Two days later, leaf pieces were transferred from the dark to BM supplemented with 5-mg/L α-naphthalene acetic acid (NAA, Duchefa Biochemie), 0.1-mg/L 6-benzyladenine (BA, Sigma-Aldrich), 500-mg/L claforan (Cefotaxime, Duchefa Biochemie) and 50-mg/L kanamycin (Sigma-Aldrich). Leaf pieces were transferred after 10 days to BM supplemented with 0.02-mg/L NAA, 2-mg/L zeatin riboside (Duchefa Biochemie), 0.02-mg/L gibberellic acid (GA3, Duchefa Biochemie), 500-mg/L claforan and 50-mg/L kanamycin, and incubated under a 16-h photoperiod for 6 weeks. Plantlets were transferred to BM supplemented with 500-mg/L claforan and 50-mg/L kanamycin until rooting. Plantlets (30 independent transformation events) were verified as transgenic by PCR with neomycin phosphotransferase gene-specific primers (NPT, Table S1) and genomic DNA extracted from leaves. Expression of the transgenic insert was verified using leaf RNA, the Verso 1-Step RT-PCR Kit ReddyMix (Thermo Scientific, Surrey, UK) and gene-specific primers (qPCR primers, Table S1). At the end of the verification stage, eight to ten independent transgenic plantlets were propagated for further growth and analyses. Five plants that exhibited the highest qPCR signal for the transgenes are presented here. Plants transformed with respective constructs without inserts were used as controls.
In vitro microtuber induction
Microtubers of cv. Desirée were induced according to standard protocol (Donnelly et al. 2003). Single-node stem cuttings were placed on Nitsch medium (Duchefa Biochemie), pH 5.8, containing 8% (w/v) sucrose, 5-mg/L kinetin (Sigma-Aldrich), 2-mg/L ancimidol (Sigma-Aldrich) and 0.8% (w/v) agar (Sigma-Aldrich) in a Petri dish, at 25 °C. Microtuber initiation could be observed after 7 days. The petri dish plates were wrapped with aluminum foil to maintain the stem cuttings and the induced microtubers in the dark.
Isolation of cDNA and promoter region
cDNA isolation and preparation of constructs
Total RNA was extracted from tuber flesh of Desirée or leaves of Arabidopsis thaliana according to Ginzberg et al. (2009). cDNA was synthesized using EZ-First Strand cDNA Synthesis Kit for RT-PCR (Biological Industries, Beit Haemek, Israel). The putative full coding regions of StCAS (GenBank accession KU313680) and the StLAS-like (GenBank accession KU313679) from potato and of AtLAS from Arabidopsis, were isolated using Ex-Taq polymerase (TaKaRa Clontech, Kusatsu, Shiga, Japan) and gene-specific primers (CDS primers, Table S1). cDNA fragments were cloned into pGEM-T Easy (Promega, Madison, WI, USA), sequenced and then cloned into the expression cartridge of the vector pART7 (at XhoI-BamHI site) which comprises the 1.3 kb of the Cauliflower mosaic virus 35S promoter, a multiple-cloning site and the transcriptional termination region of the octopine synthase gene (Gleave 1992). The entire cartridge was removed from pART7 as a NotI fragment and introduced directly into the binary vector pART27 (Gleave 1992). Vectors were introduced into Agrobacterium tumefaciens EHA 105.
The annotations of StCAS and StLAS-like in version 4.03 of the Phureja DM genome are PGSC0003DMB000000420:58827600..58845400; Sotub04g023080.1.1 and PGSC0003DMB000000328:43281700..43271000; Sotub05g021910.1.1, respectively.
Isolation of the StLAS-like promoter
To isolate the StLAS-like promoter (StLAS-like PRO), we used a reverse primer located at the end of the first exon (Table S1; StLAS-like PRO reverse) together with a forward primer that was designed based on the sequence found upstream of Sotub05g021910.1.1 in version 4.03 of the Phureja DM genome. A promoter segment of 1455 nt (including the first exon) was identified. The exon sequence was removed by PCR amplification using the above forward primer with a reverse primer, StLAS-like PRO/ATG (Table S1) that is located at the first ATG site. The final size of the resulting promoter segment was 1376 nt (GenBank accession KU313681). The annotation in the Phureja DM genome is: PGSC0003DMB000000328: 43283015..43281671. Reporter construct was prepared by cloning the StLAS-like PRO segment into Gateway vector pKGWFS7 according to standard protocols (Karimi et al. 2007).
Gene expression analysis
Yeast complementation assay
Yeast strain GIL77 (gal2 hem3-6 erg7 ura3-167) deficient in LAS activity was used as the host, and maintained on a YPD medium (1% yeast extract, 2% peptone, 2% glucose; Duchefa Biochemie) supplemented with ergosterol (20 µg/mL), 0.5% Tween 80 (v/v) and hemin chloride (13 µg/mL) (Sigma-Aldrich)—named ETH supplements. ERG7/YHR072 W plasmid (http://www.yeastgenome.org/locus/erg7/overview) was used as a complementation positive control. The isolated cDNAs of StLAS-like, StCAS and AtLAS were cloned into pYES2 plasmid (Invitrogen, Carlsbad, CA, USA) fused to galactose-inducible promoter, GAL1.
GIL77 cells were transformed using Li+ transformation method (Ito et al. 1983). The transformants were cultured on agar-based synthetic complete medium without uracil (SC-Ura) [0.17% (w/v) yeast nitrogen base without amino acids, 0.14% (w/v) yeast synthetic dropout media, 0.5% (w/v) ammonium sulfate, tryptophan (10 µg/mL), leucine (40 µg/mL), and histidine (20 µg/mL), 2% glucose; Duchefa Biochemie] with ETH supplements at 28 °C. Colonies were transferred to SC-Ura plates containing 2% galactose instead of glucose, without the ETH supplements. When required, uracil was added (40 µg/mL).
Gene expression analyses by qPCR
Total RNA was extracted from leaves and tuber flesh of greenhouse grown plants. cDNA was synthesized using EZ-First Strand cDNA Synthesis Kit for RT-PCR (Biological Industries) as described above. Taq polymerase (Super-Therm 500 u, cat. no. JMR- 801PCR, JMR Holdings, London, UK) was used for semi-quantitative PCR, and ABsolute™ Blue QPCR SYBR® Green ROX Mix (Thermo Scientific, Surrey, UK) was used for quantitative real-time PCR (qPCR) according to the manufacturers’ protocols. Biological replicates included four to five plants for each transformed line—representing independent transformation events—that were analyzed individually. Values in each sample were normalized to the level of α-chain of the nascent polypeptide-associated complex (α-NAC, Sotub10g027110) as the reference gene (Ginzberg et al. 2009, 2012). Normalized values obtained for the individual biological replicates were used to calculate the average relative expression level of a gene in the transformed line. All primers used are listed in Table S1.
GUS analysis of StLAS-like promoter
To study tissue expression of the StLAS-like gene, we transformed the StLAS-like PRO reporter construct and an empty vector control into Desirée plantlets that were maintained in tissue culture (15 independent transgenics). Cultures of in vitro-induced microtubers were prepared for five selected independent plantlets and were subjected to GUS staining based on Jefferson et al. (1987); staining included leaf, stem and root samples from whole plantlets as well. To analyse putative cis-acting regulatory elements within the StLAS-like PRO, we used the PLACE database (Higo et al. 1999).
Analyses of sterols and SGAs
Metabolite analysis was performed with leaves and tuber flesh of greenhouse grown plants. SGA extraction and analysis were performed as described by Ginzberg et al. (2012) with some modifications. In brief, 0.5 g DW of sample was rehydrated in 5 mL of extraction buffer (0.02 M heptanesulfonic acid in 1% aqueous acetic acid, v/v). The extract was centrifuged at 4000g in a benchtop centrifuge (Sorvall Legend, Thermo Scientific) at 4 °C. The supernatant was applied to a Sep-Pak C18 cartridge (250 mg; Waters, Milford, MA, USA). The SPE cartridge was washed serially with 5 mL each of water, 0.05-M ammonium bicarbonate, 0.025-M ammonium bicarbonate in 50% methanol, and water. The SGAs were eluted with 5-mL methanol-0.1-M hydrochloric acid (80:20, v/v). The SGA analyses were conducted using UPLC-Triple Quadrupole-MS (Waters Xevo TQ MS). Samples were filtered through a Millex-HV Durapore (PVDF) membrane (0.22 µm) before injection into the LC–MS instrument. Separation was performed on a 2.1 × 50 mm i.d., 1.7 µm UPLC BEH C18 column. Chromatographic and MS parameters were as follows: the mobile phase consisted of water (phase A) and 0.1% formic acid in acetonitrile (phase B). The linear gradient program was as follows: 85–65% A over 10 min, 65–5% A over 1 min, held at 5% A over 3 min, then returned to the initial conditions (85% A) after 1 min and held at 85% A over 2 min. The flow rate was 0.3 mL/min and the column temperature was kept at 35 °C. All analyses were performed using the ESI source in positive ion mode with the following settings: capillary voltage 3.2 kV, cone voltage 30 V, desolvation temperature 350 °C, desolvation gas flow 650 L/h, and source temperature 150 °C. Quantitation was performed using MRM acquisition by monitoring the 868/98, 868/398 (RT = 5.1, dwell time of 161 ms for each transition) for α-solanine. The MassLynx software version 4.2 (Waters Inc.) was used to control the instrument and the Targetlynx software was used for calculations. Alpha-solanine (Sigma-Aldrich, Cat. no. 85472) was used as standard and α-chaconine concentration was calculated relative to α-solanine peak area.
Phytosterol extraction was conducted based on a protocol of the Lange group at Washington State University. Frozen plant material (200 mg) was ground to a powder in the presence of liquid nitrogen and analytes were extracted at 75 °C for 60 min with 8-mL chloroform/methanol (2:1, v/v) containing 1.25 mg/L epi-cholesterol (Sigma-Aldrich, Cat. no. C-2882) as an internal standard. Extracts were kept at room temperature for at least 1 h, solvents were evaporated to dryness, and the remaining residues were saponified at 90 °C for 60 min in 4 mL 6% (w/v) KOH in methanol (the saponification in alkaline conditions converts the sterol esters into free sterols, allowing both free sterols and steryl esters to be extracted). Upon cooling to room temperature, 2-mL n-hexanes and 2-mL H2O were added, and the mixture was shaken vigorously for 20 s. Following centrifugation (3000g for 2 min) to separate the phases, the hexane phase was transferred to a 2 mL glass vial, and the aqueous phase was re-extracted with 2 mL n-hexanes, centrifuged as above, and the hexane phase added to the glass vial containing the hexane phase from the first extraction. The combined hexane phases were evaporated to dryness. 50 µL of N-methyl-N-trimethylsilyltrifluoroacetamide (MSTFA) were added to the residue, the sample was shaken vigorously for 20 s, and the mixture transferred to a 2 mL autosampler glass vial with a 100 µL conical glass insert. The reaction mixture was incubated at room temperature in a capped vial for at least 5 min.
GC–MS analyses were performed on an Agilent 7890A GC coupled to an Agilent 5975C inert MSD detector. Samples were loaded (injection volume 1 µL) with a LEAP CombiPAL onto a HP-5MS-fused silica column (30 m × 250 µm; 0.25 µm film thickness). The temperatures of the injector and MSD interface were both set to 280 °C. Analytes were separated at a flow rate of 1 mL/min at splitless mode, using He as carrier gas and using a thermal gradient starting at 170 °C (1.5 min), which was ramped first to 280 °C at 37 °C/min and then to 300 °C at 1.5 °C/min, where it was held for 5 min. Eluents were fragmented in electron impact mode with an ionization voltage of 70 eV. To ensure low background signals, we ran a blank injection (followed by a shortened thermal gradient) after each sample run. Prior to sample analyses, and then after every 20 samples, a standard mix was run to evaluate the reproducibility of the analyses. Data were acquired using MSD ChemStation (Revision E.02.) software. Background was subtracted. Analytes were identified based on their mass fragmentation patterns by comparison with those of authentic standards and using the NIST Mass Spectral Search Program (Version 0.8.L). Peak areas were obtained from the total ion chromatogram (TIC) for some detectable peaks with a phytosterol mass fragmentation signature. Raw data were exported to Microsoft Excel and peak areas normalized to tissue mass and internal standard (epi-cholesterol) using Microsoft Access. Sterol standards were obtained from Sigma-Aldrich and included lanosterol (Cat. no. L-5768), cholesterol (Cat. no. C-8667), campesterol (Cat. no. C-5157), and stigmasterol (Cat. no. S-2424). GC–MS mass fragmentation of the tri-methyl-silyl derivatives is given in Table S2.
Statistical analyses
Data are presented as average values of four to five independent transgenic plantlets with ±SE. Data were further analyzed for statistical significance among means by Student’s t test (JMP software, http://www.jmp.com). Significant difference was determined at P < 0.05.
Results
Isolation of potato StLAS-like
To isolate the potato putative LAS cDNAs, we used the tomato CAS coding sequence (EU449280) for a BLAST search against the potato EST library. The resulting potato sequence was used to design primers and to clone the StCAS cDNA and to identify the putative StLAS-like gene. The latter was identified by BLAST search of StCAS putative protein sequences against translated ‘Potato PGSC DM V3 superscaffolds’ database. The homologous sequences were scanned for amino acid substitutions previously shown to alter the catalytic function of CAS1 to LAS1 in Arabidopsis (Fig. 1c). The putative StLAS-like sequence from the database (PGSC0003DMB000000328:43281700..43271000) was used to design primers to isolate the respective coding sequence from cDNA of the commercial potato cultivar Desirée. PCR product was obtained with cDNA generated from tuber flesh but not from leaves.
Although PGSC gene annotation data release ver. 3.4 is lacking in the abovementioned region on the Potato Genome Browser ver. 4.03, the ITAG Solanum tuberosum Group Phureja DM 1–3 gene model Sotub05g02190.1.1 (annotated as CAS but with the characteristic amino acids for LAS) is present at this site; the putative position of StCAS (PGSC0003DMB000000420:58827600..58845400) likewise lacks PGSC gene annotation but the ITAG Solanum tuberosum Group Phureja DM 1-3 gene model Sotub04g23080.1.1 (also annotated as CAS and carrying the signature CAS amino acids) occurs at this site. RNAseq tracks for the exons of Phureja DM1-3 gene model Sotub04g23080.1.1 (CAS) show high expression in all examined tissues of RH, whereas Sotub05g02190.1.1 (LAS) appears to be only weakly expressed. However, the occurrence of similar sequences elsewhere in the potato genome may confound RNAseq alignments.
Alignment of the putative StLAS-like protein from Desirée to Arabidopsis AtLAS and to the potato StCAS shows 67 and 66% homology at the amino acid level. A genetic complementation assay was used to verify the OSC activity of the StLAS-like cDNA. The yeast strain GIL77 deficient in LAS activity was transformed with the pYES2 plasmid harboring the isolated cDNAs of StLAS-like, StCAS, and AtLAS fused to the galactose inducible promoter, GAL1. All three transformant strains grew on the minimal media in the presence of galactose, indicating that the plasmids carried functional enzymes and that StLAS-like activity could complement the ERG7 deficiency (Fig. 1d).
StLAS-like differential expression in tuber flesh
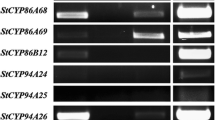
As StLAS-like cDNA could not be isolated from leaves, we examined tissue-specific expression of the StLAS-like gene by PCR using gene-specific primers (qStLAS-like; Table S1) and cDNA prepared from various potato tissues, including: tuber flesh, tuber peel, stolon, root, stem, and leaves. StLAS-like PCR product was only obtained in tuber flesh sample, while its signal was undetectable in leaves and tissues other than tuber flesh (Fig. S1).
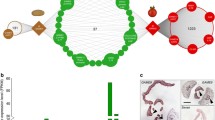
Further expression studies were performed with transgenic potato plantlets grown in tissue culture and carrying a construct of the GUS reporter gene fused to the StLAS-like promoter segment (StLAS-like PRO). These transgenic plantlets were used for in vitro induction of microtubers. StLAS-like PRO expression was detected in microtuber flesh tissue only; no GUS staining was detected in tuber peel, leaf, stem or root tissues (Fig. 2). Some staining was obtained at stem cut edges; however its specificity is uncertain as: (1) no StLAS-like transcripts were detected in stem tissue as described above (Fig. S1); (2) following cutting, stem segments were placed immediately in the staining solution, ruling out a wound response which requires GUS transcription and translation. The gathered data indicate that StLAS-like gene is differentially expressed in potato cultivar Desirée with dominant expression in tuber flesh tissue.
StLAS-like PRO-specific expression in potato tuber flesh as demonstrated with in vitro-induced microtubers. GUS staining assay of leaf, stem, microtuber (cut and un-cut) and root tissues collected from tissue culture-grown transgenic potato plantlets carrying the pKGWFS7 plasmid with the StLAS-like PRO promoter segment, or the plasmid without an insert (Control). Note the bluish coloration under the skin of StLAS-like PRO un-cut microtuber. Figures demonstrate one plantlet out of four independent transgenics for each construct. Bar 2 mm (for all figures)
To better understand StLAS-like promoter function and tissue specificity, we analyzed its sequence for putative cis-acting regulatory elements using the PLACE database (Table S3)—out of 44 elements, 32% were stress-related, 27% were related to tissue-specific expression, and the rest were related to hormone or light signaling. Two additional elements previously reported to play a regulatory role in SGA biosynthesis were also found in the StLAS-like promoter—the G-box CACGTG and the P-box AGCCAACCA. These elements were found in the promoters of SGA biosynthetic genes in tomato and potato that are activated by ethylene response factor (ERF) transcription factors (Cárdenas et al. 2016; Thagun et al. 2016).
Overexpression of Arabidopsis AtLAS in potato leaves and tubers
The role of LAS in SGA and phytosterol metabolism was studied in greenhouse-grown potato plants overexpressing the well-characterized LAS from Arabidopsis (AtLASoe-plants). First, the effect of AtLAS overexpression was monitored in leaves, where LAS activity is not naturally expressed.
AtLAS transcript accumulated to a high level in the leaves of AtLASoe-plants (Fig. 3a). As a result, the relative transcript levels of HMG1, SQS1, and CAS that provide intermediates to the phytosterol and SGA pathways were reduced, the latter two significantly (Fig. 3a). In accordance, the levels of cycloartenol and the phytosterols campesterol and sitosterol were reduced, the latter two significantly (Fig. 3b), albeit the expression levels of SMT1, CYP51G and SSR1 were not changed (Fig. 3a). In contrast, transcript levels of SGA-related genes SSR2, GAME4, and SGT2 were upregulated (the first two significantly), and the respective SGA metabolite levels were significantly increased (Fig. 3c). Overall, overexpression of AtLAS in potato leaves resulted in sterol reduction and an increased abundance of SGA-related transcripts and metabolites.
SGA- and phytosterol-related gene expression and metabolic profiling in leaf of greenhouse-grown AtLAS overexpressing plants (AtLASoe-plants) and empty vector control plants (Control). a Transcript levels were determined by qPCR and normalized with the reference gene NAC. Tested genes included HMG1 and SQS1 (key enzymes of the mevalonic/isoprenoid pathway); LAS and CAS; SSR2, GAME8a, GAME4, SGT1 and SGT2 (SGA-related biosynthetic genes); SMT1, CYP51G and SSR1 of the phytosterol biosynthetic pathway. b GC–MS profiling of phytosterols. Metabolite identification was based on their mass fragmentation patterns by comparison with those of authentic standards (lanosterol, cholesterol, campesterol, and stigmasterol) or those in the database. The value of ‘total sterols’ bar sums the levels of the metabolites downstream from the lanosterol/cycloartenol step. c UPLC profiling of α-solanine and α-chaconine and their total level. Values represent an average of five AtLASoe-plants and four control plants with SE bars. Data were analyzed for statistical significance among means by Student’s t test. Significant differences, marked with an asterisk, were determined at P < 0.05
Similar analyses were performed in tuber flesh of AtLASoe-plants, bearing in mind that both the endogenous StLAS-like and the transgenic AtLAS were expressed in that tissue (Fig. 4a). SQS1 transcript abundance was greater in AtLASoe-tubers compared to control plants, likewise the SGT1 transcript, albeit not significantly (Fig. 4a). HMG1 and GAME4 transcript abundance were below detection level in tubers of both AtLASoe and control plants. No change was detected in transcript level of CAS, SMT1, and SGT2. No significant change was obtained for sitosterol and other phytosterols level, except for increased accumulation of campesterol (TMT derivative) and reduced level for lanosterol (Fig. 4b). Interestingly, SGA levels were significantly reduced compared to control, as α-chaconine could not be detected (Fig. 4c). Overall, overexpression of AtLAS in potato tuber did not change total sterols and sitosterol, but reduced the level of SGA metabolites.
SGA- and phytosterol-related gene expression and metabolic profiling in tuber flesh of greenhouse-grown AtLASoe-plants and empty vector control plants (Control). a Relative transcript levels were determined by qPCR as described for Fig. 3. b GC–MS profiling of phytosterols as described for Fig. 3. c UPLC profiling of α-solanine and α-chaconine and their total levels. Values represent an average of five AtLASoe-plants and four control plants with SE bars. Data were analyzed for statistical significance among means by Student’s t test. Significant differences, marked with an asterisk, were determined at P < 0.05
The opposing trends in AtLASoe-plants, where SGA levels were increased in leaves (where LAS activity is not naturally expressed) while reduced in tuber flesh may suggest different regulating mechanism for SGA biosynthesis in these plant organs: in leaves, sterol intermediates are diverted into the SGA pathway, while in tuber flesh, they are catalyzed by LAS activity, hence fewer precursors are diverted into SGA biosynthesis.
Discussion
The potato StLAS-like
The present study was designed to examine the involvement of LAS activity in secondary metabolism of potato SGAs. The LAS gene is presumed to have originated by duplication of the CAS gene (Sawai et al. 2006) which encodes a key enzyme of phytosterol and SGA synthesis in plants. Previous reports suggested that lanosterol participates in part, albeit minimally, in phytosterol production as well as in synthesis of secondary metabolites that contribute to the fitness of the plant to biotic and abiotic stress (Kolesnikova et al. 2006; Suzuki et al. 2006; Ohyama et al. 2009). Yet the function of LAS activity in plants remained an enigma.
The potato LAS-like gene was identified and isolated based on amino acid substitutions relative to CAS sequence that were shown in Arabidopsis to biosynthesize lanosterol instead of cycloartenol; i.e., the AtCAS1 gene could be converted to a functional AtLAS gene with nonsynonymous mutations changing His477Asn and Ile481Val (Fig. 1c) (Lodeiro et al. 2005). The ability of StLAS-like (and StCAS) to complement the erg7 deficiency of the yeast mutant GIL77 demonstrated the isolation of potato oxidosqualene cyclases with protosteryl cation activity (CAS/LAS) rather than with dammarenyl cation activity (Fig. 1d). The erg7 deficiency can be chemically complemented by cycloartenol as well as lanosterol, both of which are converted into membrane sterols essential for yeast viability (Nes et al. 1993).
The StLAS-like gene is differentially expressed in potato with dominant expression in tuber flesh tissue as indicated by exclusive GUS activity in flesh of microtubers transformed with the StLAS-like promoter fused to the reporter gene (Fig. 2), and undetectable levels of StLAS-like cDNA in leaves and tissues other than tuber flesh (Fig. S1). In accordance, the StLAS-like promoter bears several cis-regulatory elements that have been found to direct tissue-specific expression and may support its localized expression (Table S3). With relevance to the subject in question, two additional elements previously reported to play a regulatory role in SGA biosynthesis were also found in the StLAS-like promoter—the G-box CACGTG and the P-box AGCCAACCA. These elements were found in the promoters of SGA biosynthetic genes in tomato and potato that are activated by members of the ethylene response factor (ERF) transcription factors (Cárdenas et al. 2016; Thagun et al. 2016). The P-box was required for gene activation by JRE4, a jasmonate-responsive ERF transcription factor, and found in the promoter region of GAME4 and the sterol reductase gene DWF5 (Thagun et al. 2016). GAME9, the potato ortholog of JRE4, was also shown to regulate chaconine and solanine biosynthesis, either directly or in cooperation with the MYC2 transcription factor that binds to a G-box element detected in the putative promoters of several GAME and putative cholesterol genes, including GAME4 and C5-SD (Cárdenas et al. 2016). CAS, SSR2 and C5-SD, were upregulated in potato overexpressing GAME9, indicating its action on genes downstream to 2,3-oxidosqualene (Cárdenas et al. 2016). Hence, the present finding of similar cis-elements in the StLAS-like promoter suggests its co-regulation with sterol and SGA pathways. Other elements within the promoter functioning in light, stress response, and hormone signaling further support the suggestion of LAS involvement in the secondary metabolism of plant defense response (Kolesnikova et al. 2006; Suzuki et al. 2006; Ohyama et al. 2009). Similar defense-related cis-elements were identified within the OSCs promoter from Solanaceae including Withania somnifera and tomato that have also been shown to be expressed in a tissue-specific manner (Wang et al. 2011; Dhar et al. 2014).
Overexpression of Arabidopsis AtLAS in potato
The well-characterized Arabidopsis AtLAS was overexpressed in greenhouse-grown transgenic potato plants to elucidate its involvement in phytosterol and SGA biosynthesis. Several representative genes were chosen to monitor SGA and phytosterol-specific pathways based on their position at critical points in sterol metabolism—SSR2 and GAME4 for SGA formation and SMT1 and SSR1 for phytosterol biosynthesis. SSR2, a sterol side chain reductase, exhibiting Δ24(25) reductase activity that converts cycloartenol to cycloartanol, was recently shown to be committed to cholesterol and SGA biosynthesis (Sawai et al. 2014). SSR2 is a homolog of the SSR1/DWF1 gene that is involved in the downstream biosynthetic steps of the phytosterols, sitosterol, and campesterol. Hence, SSR2 and SSR1 activities can differentiate between the two pathways (Fig. 1a). SMT1 converts cycloartenol, the same substrate used by SSR2, to 24-methylene cycloartenol, which is an intermediate of C-24 alkylsterol (campesterol and sitosterol) biosynthesis (Diener et al. 2000). Accordingly, overexpression of a soybean SMT1 in potato plants increased the metabolic flux of cycloartenol into alkylated sterols at the expense of cholesterol and SGAs (Arnqvist et al. 2003). Finally, GAME4 has a central role in SGA biosynthesis as a C24 oxidase that provides an intermediate for the transaminase activity essential for potato SGAs (Itkin et al. 2013). Silencing of GAME4 in potato dramatically reduced the levels of α-solanine/α-chaconine in both leaves and tubers (Itkin et al. 2013). Therefore, the pairs SSR2-GAME4 and SMT1-SSR1 can be considered as the main determinants for phytosterol/SGA pathway diversion.
First, the effect of AtLAS overexpression was monitored in leaves, where LAS activity is not naturally expressed. Leaves of AtLASoe-plants exhibited reduced expression of HMG1, SQS1, and CAS and concomitantly, reduced accumulation of cycloartenol, campesterol, and sitosterol (albeit not all significantly, Fig. 3a, b). In contrast, the SGA biosynthetic pathway was upregulated as evident by increased transcript levels of SGA-related key genes SSR2 and GAME4, and increased SGA metabolite levels. We suggest that AtLAS activity in leaves resulted in downregulation of sterol synthesis, such that intermediates in the pathway were diverted into the SGA branch to maintain sterol homeostasis. The LAS effect may be due to synthesis of end-product sterol via the lanosterol pathway (i.e., sitosterol, Ohyama et al. 2009), feedback-inhibiting key regulatory gene expression and enzymatic activities in the phytosterol pathway to maintain sterol homeostasis—and this, as one mechanism out of several options to regulate sterol level (Fig. 3; Nes and Venkatramesh 1999; Holmberg et al. 2002).
In tuber flesh of AtLASoe-plants, high LAS expression resulted in the opposite, i.e., SGA metabolite level was dramatically reduced while no significant change was observed in total sterols and sitosterol (Fig. 4). We suggest that high LAS expression in AtLASoe-tubers (the endogenous StLAS-like plus the transgenic AtLAS), competes with CAS for precursors; thus, to maintain sterol homeostasis, fewer precursors are diverted into SGA biosynthesis, hence reducing their level. The opposing effects of LAS overexpression in AtLASoe-leaves and tuber flesh may result from disparity in cycloartenol levels in these tissues, high in leaves, low in tuber flesh (around 120 and 2 µg/gFW, respectively). In the former, removal of intermediates by the transgenic AtLAS may be less critical.
Modeling the involvement of potato StLAS-like in phytosterol and SGA biosynthesis in leaves and tubers
LAS from plants has mainly been characterized in Arabidopsis; however, in potato the presence of a parallel biosynthetic pathway for SGA metabolites, in addition to the sterol and brassinosteroid routes, may add to the complexity by which the respective pathways are coordinated. Furthermore, in cultivated potato, SGAs accumulate in leaves while low/nil in tuber flesh, suggesting differential tissue regulation, which further challenges the characterization of the respective regulatory networks. In addition, unlike the ubiquitous expression of CAS1 throughout plant development, the expression of LAS1 is different in each plant tissue as was shown here and in Arabidopsis (Suzuki et al. 2006), and may affect sterol homeostasis differently in each plant tissue.
Although the general biosynthetic enzymes of plant sterols have been defined, the arrangement and regulation are not fully understood (Suzuki and Muranaka 2007; Vranová et al. 2012; Cárdenas et al. 2015). The cycloartenol–lanosterol bifurcation in sterol biosynthesis is not absolute as shown in feeding experiments of isotope-labeled acetic acid, mevalonate, cycloartenol or lanosterol (Baisted et al. 1968; Hewlins et al. 1969; Ohyama et al. 2009). Sterol synthesis is modulated, in part, by the mevalonate pathway through feedback-regulation of end-products. The concentration of intermediates such as Δ5-sterols or cycloartenol may regulate HMGR activity (Holmberg et al. 2002). SQS1 activity may also be critical for controlling carbon flow to sterol end-product (Zook and Kuć 1991) and co-regulation of HMG1 and SQS1 (Yoshioka et al. 1999; Wentzinger et al. 2002; Ginzberg et al. 2012). Downstream in the pathway, sitosterol has been shown to downregulate the activity of SMT1 in sunflower, exhibiting an additional feedback regulatory point at the first step in the phytosterol specific pathway (Janssen and Nes 1992). Free sterols are the active form, and esterification of sterols and their compartmentalization in lipid bodies have been suggested as additional means to overcome sterol overproduction and maintain sterol homeostasis (Gondet et al. 1994; Schaller et al. 1995). Sterol glycosides and acylated sterol glycosides were also reported to be major fractions of potato tuber sterols (Galliard 1968; Catz et al. 1985).
Although the phytosterol pathway seems to be tightly regulated for constitutive levels despite the perturbations of the pathway made in this study, diverting extra intermediates to a parallel pathway of functionally distinct importance might be an alternative means of regulation or a mechanism that allows some tolerance of excess intermediates without the necessity to up- or downregulate the abovementioned critical enzymes. As StLAS-like is not expressed in leaves, we suggest that extra intermediates towards the CAS1/SMT1 pathway are diverted via SSR2 into cholesterol and SGA biosynthesis (Fig. 5a). In support of this, overexpression of HMG1 or SQS1 in potato resulted in increased levels of SGAs (Ginzberg et al. 2012). Accordingly, in AtLASoe-leaves, atypical LAS-like activity may disrupt the balance by increasing the concentration of sterol intermediates to the system, resulting in downregulation of the HMG1/SQS1/CAS1 upper part of the pathway and diversion of intermediates to the SGA pathway via SSR2, thereby increasing SGA levels (Fig. 3).
Suggested model for the involvement of potato StLAS-like in phytosterol and SGA biosynthetic pathways. In both leaf and tuber flesh, the CAS/SMT1 pathway (thick arrow) is dominant over the SGA branch. a In the leaf, where StLAS-like is not expressed, phytosterol homeostasis is maintained by diverting excess intermediates to SGA synthesis. b In the tuber flesh, to maintain phytosterol homeostasis extra intermediates are catalyzed by the endogenous StLAS-like activity, resulting with reduced SGA levels
In contrast to leaves, in the tuber flesh endogenous, LAS-like activity may play a role in regulating phytosterol homeostasis—extra phytosterol intermediates may be catalyzed by StLAS-like activity in addition or instead of being diverted to the SGA pathway (Fig. 5b). Accordingly, in AtLASoe-tubers—with endogenous StLAS-like and the transgenic AtLAS—SGA level was reduced to maintain phytosterol homeostasis as high AtLAS activity was competing for precursors: in that case, SQS1 was upregulated probably to maintain the required phytosterol level (Fig. 4).
The characterization of GAME9/JRE4 transcription factor that controls not only other GAME genes of the core SGA pathway but also biosynthetic genes involved in cholesterol biosynthesis (Cárdenas et al. 2016; Thagun et al. 2016) supports our postulation of feedback mechanisms and coordinated expression of genes localized to branch points of the respective pathways that influence the flux between SGAs, phytosterols, and other plant defense compounds.
Conclusion
The gathering results imply that in tuber flesh, StLAS-like activity and possibly the lanosterol pathway dominate the SGA pathway, resulting in low/nil levels of SGAs typical for commercial cultivars. In leaves, the absence of StLAS-like activity facilitates the diversion of intermediates into the SGA pathway, resulting in significantly increased levels compared to tuber flesh.
Accession numbers
Sequence data from this article can be found in the GenBank/EMBL databases under the following accession numbers: StCAS, KU313680; StLAS-like, KU313679; StLAS-like promoter KU313681.
Author contribution statement
AK performed most of the experiments; EF provided technical assistance to AK; MW performed the metabolic analyses; ZT conducted greenhouse experiments; RV conducted genomic analysis and complemented the writing; IG conceived the project and wrote the article with contributions of all the authors.
Abbreviations
- CAS:
-
Cycloartenol synthase
- GAME:
-
Glycoalkaloid metabolism
- HMG:
-
3-Hydroxy-3-methylglutaryl coenzyme-A reductase
- LAS:
-
Lanosterol synthase
- SGA:
-
Steroidal glycoalkaloid(s)
- SGT:
-
Solanidine glycosyltransferases
- SMT:
-
C-24-methyltransferase type
- SQS:
-
Squalene synthase
- SSR:
-
Sterol side chain reductase
References
Arnqvist L, Dutta PC, Jonsson L, Sitbon F (2003) Reduction of cholesterol and glycoalkaloid levels in transgenic potato plants by overexpression of a type 1 sterol methyltransferase cDNA. Plant Physiol 131:1792–1799
Baisted DJ, Gardner RL, McReynolds LA (1968) Phytosterol biosynthesis from lanosterol in Euphorbia peplus. Phytochemistry 7:945–949
Benveniste P (2004) Biosynthesis and accumulation of sterols. Annu Rev Plant Biol 55:429–457
Bergenstråhle A, Borga P, Jonsson MV (1996) Sterol composition and synthesis in potato tuber discs in relation to glycoalkaloid synthesis. Phytochemistry 41:155–161
Cárdenas PD, Sonawane PD, Heinig U, Bocobza SE, Burdman S, Aharoni A (2015) The bitter side of the nightshades: genomics drives discovery in Solanaceae steroidal alkaloid metabolism. Phytochemistry 113:24–32
Cárdenas PD, Sonawane PD, Pollier J, Vanden Bossche R, Dewangan V, Weithorn E, Tal L, Meir S, Rogachev I, Malitsky S, Giri AP, Goossens A, Burdman S, Aharoni A (2016) GAME9 regulates the biosynthesis of steroidal alkaloids and upstream isoprenoids in the plant mevalonate pathway. Nat Commun 7:10654
Catz DS, Tandecartz JS, Cardini CE (1985) UDP-glucose: sterol glucosyltransferase and a steryl glucoside acyltransferase activity in amyloplast membranes from potato tubers. J Exp Bot 36:602–609
Clouse SD, Sasse JM (1998) Brassinosteroids: essential regulators of plant growth and development. Annu Rev Plant Physiol Plant Mol Biol 49:427–451
Dhar N, Rana S, Razdan S, Bhat WW, Hussain A, Dhar RS, Vaishnavi S, Hamid A, Vishwakarma R, Lattoo SK (2014) Cloning and functional characterization of three branch point oxidosqualene cyclases from Withania somnifera (L.) Dunal. J Biol Chem 289:17249–17267
Diener AC, Li H, W-x Zhou, Whoriskey WJ, Nes WD, Fink GR (2000) STEROL METHYLTRANSFERASE 1 controls the level of cholesterol in plants. Plant Cell 12:853–870
Donnelly DJ, Coleman WK, Coleman SE (2003) Potato microtuber production and performance: a review. Am J Potato Res 80:103–115
Galliard T (1968) Aspects of lipid metabolism in higher plants—I. Identification and quantitative determination of the lipids in potato tubers. Phytochemistry 7:1907–1914
Ginzberg I, Barel G, Ophir R, Tzin E, Tanami Z, Muddarangappa T, de Jong W, Fogelman E (2009) Transcriptomic profiling of heat-stress response in potato periderm. J Exp Bot 60:4411–4421
Ginzberg I, Thippeswamy M, Fogelman E, Demirel U, Mweetwa A, Tokuhisa J, Veilleux R (2012) Induction of potato steroidal glycoalkaloid biosynthetic pathway by overexpression of cDNA encoding primary metabolism HMG-CoA reductase and squalene synthase. Planta 235:1341–1353
Gleave AP (1992) A versatile binary vector system with a T-DNA organizational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20:1203–1207
Gondet L, Bronner R, Benveniste P (1994) Regulation of sterol content in membranes by subcellular compartmentation of steryl-esters accumulating in a sterol-overproducing tobacco mutant. Plant Physiol 105:509–518
Heftmann E (1983) Biogenesis of steroids in Solanaceae. Phytochemistry 22:1843–1860
Hewlins MJE, Ehrhardt JD, Hirth L, Ourisson G (1969) The conversion of [14C]cycloartenol and [14C]lanosterol into phytosterols by cultures of Nicotiana tabacum. Eur J Biochem 8:184–188
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300
Holmberg N, Harker M, Gibbard CL, Wallace AD, Clayton JC, Rawlins S, Hellyer A, Safford R (2002) Sterol C-24 methyltransferase type 1 controls the flux of carbon into sterol biosynthesis in tobacco seed. Plant Physiol 130:303–311
Itkin M, Heinig U, Tzfadia O, Bhide AJ, Shinde B, Cardenas PD, Bocobza SE, Unger T, Malitsky S, Finkers R, Tikunov Y, Bovy A, Chikate Y, Singh P, Rogachev I, Beekwilder J, Giri AP, Aharoni A (2013) Biosynthesis of antinutritional alkaloids in solanaceous crops is mediated by clustered genes. Science 341:175–179
Ito H, Fukuda Y, Murata K, Kimura A (1983) Transformation of intact yeast cells treated with alkali cations. J Bacteriol 153:163–168
Janssen GG, Nes WD (1992) Structural requirements for transformation of substrates by the S-adenosyl-l-methionine: Δ24(25)-sterol methyltransferase. Inhibition by analogs of the transition state coordinate. J Biol Chem 267:25856–25863
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Karimi M, Depicker A, Hilson P (2007) Recombinational cloning with plant gateway vectors. Plant Physiol 145:1144–1154
Kolesnikova MD, Xiong QB, Lodeiro S, Hua L, Matsuda SPT (2006) Lanosterol biosynthesis in plants. Arch Biochem Biophys 447:87–95
Laule O, Furholz A, Chang HS, Zhu T, Wang X, Heifetz PB, Gruissem W, Lange BM (2003) Crosstalk between cytosolic and plastidial pathways of isoprenoid biosynthesis in Arabidopsis thaliana. Proc Natl Acad Sci USA 100:6866–6871
Lodeiro S, Schulz-Gasch T, Matsuda SPT (2005) Enzyme redesign: two mutations cooperate to convert cycloartenol synthase into an accurate lanosterol synthase. J Am Chem Soc 127:14132–14133
McCue KF, Shepherd LVT, Allen PV, Maccree MM, Rockhold DR, Corsini DL, Davies HV, Belknap WR (2005) Metabolic compensation of steroidal glycoalkaloid biosynthesis in transgenic potato tubers: using reverse genetics to confirm the in vivo enzyme function of a steroidal alkaloid galactosyltransferase. Plant Sci 168:267–273
McCue KF, Allen PV, Shepherd LVT, Blake A, Maccree MM, Rockhold DR, Novy RG, Stewart D, Davies HV, Belknap WR (2007) Potato glycosterol rhamnosyltransferase, the terminal step in triose side-chain biosynthesis. Phytochemistry 68:327–334
Meyer MM, Xu R, Matsuda SPT (2002) Directed evolution to generate cycloartenol synthase mutants that produce lanosterol. Org Lett 4:1395–1398
Moehs CP, Allen PV, Friedman M, Belknap WR (1997) Cloning and expression of solanidine UDP-glucose glucosyltransferase from potato. Plant J 11:227–236
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiologia plantarum 15(3):473–497
Nes WD (2011) Biosynthesis of cholesterol and other sterols. Chem Rev 111:6423–6451
Nes WD, Venkatramesh M (1999) Enzymology of phytosterol transformations. Crit Rev Biochem Mol Biol 34:81–93
Nes WD, Janssen GG, Crumley FG, Kalinowska M, Akihisa T (1993) The structural requirements of sterols for membrane function in Saccharomyces cerevisiae. Arch Biochem Biophys 300:724–733
Nitsch JP, Nitsch C (1969) Haploid plants from pollen grains. Science 163(3862):85–87. doi:10.1126/science.163.3862.85
Ohyama K, Suzuki M, Kikuchi J, Saito K, Muranaka T (2009) Dual biosynthetic pathways to phytosterol via cycloartenol and lanosterol in Arabidopsis. Proc Natl Acad Sci USA 106:725–730
Petersson EV, Nahar N, Dahlin P, Broberg A, Tröger R, Dutta PC, Jonsson L, Sitbon F (2013) Conversion of exogenous cholesterol into glycoalkaloids in potato shoots, using two methods for sterol solubilisation. PLoS One 8:e82955
Rahier A (2011) Dissecting the sterol C-4 demethylation process in higher plants. From structures and genes to catalytic mechanism. Steroids 76:340–352
Ripperge H, Moritz W, Schreibe K (1971) Solanum alkaloids: biosynthesis of Solanum alkaloids from cycloartenol or lanosterol. Phytochemistry 10:2699–2704
Sawai S, Akashi T, Sakurai N, Suzuki H, Shibata D, S-i Ayabe, Aoki T (2006) Plant lanosterol synthase: divergence of the sterol and triterpene biosynthetic pathways in eukaryotes. Plant Cell Physiol 47:673–677
Sawai S, Ohyama K, Yasumoto S, Seki H, Sakuma T, Yamamoto T, Takebayashi Y, Kojima M, Sakakibara H, Aoki T, Muranaka T, Saito K, Umemoto N (2014) Sterol side chain reductase 2 is a key enzyme in the biosynthesis of cholesterol, the common precursor of toxic steroidal glycoalkaloids in potato. Plant Cell 26:3763–3774
Schaller H, Grausem B, Benveniste P, Chye ML, Tan CT, Song YH, Chua NH (1995) Expression of the Hevea brasiliensis (H.B.K.) Müll. Arg. 3-hydroxy-3-methylglutaryl coenzyme A reductase 1 in tobacco results in sterol overproduction. Plant Physiol 109:761–770
Segura MJR, Lodeiro S, Meyer MM, Patel AJ, Matsuda SPT (2002) Directed evolution experiments reveal mutations at cycloartenol synthase residue His477 that dramatically alter catalysis. Org Lett 4:4459–4462
Stankovic M, Stojanovic O, Kobilarov N (1990) Unsaponifiable lipids from haulm and tuber sprouts of potato (Solanum tuberosum L). Potato Res 33:459–464
Suzuki M, Muranaka T (2007) Molecular genetics of plant sterol backbone synthesis. Lipids 42:47–54
Suzuki M, Xiang T, Ohyama K, Seki H, Saito K, Muranaka T, Hayashi H, Katsube Y, Kushiro T, Shibuya M, Ebizuka Y (2006) Lanosterol synthase in dicotyledonous plants. Plant Cell Physiol 47:565–571
Thagun C, Imanishi S, Kudo T, Nakabayashi R, Ohyama K, Mori T, Kawamoto K, Nakamura Y, Katayama M, Nonaka S, Matsukura C, Yano K, Ezura H, Saito K, Hashimoto T, Shoji T (2016) Jasmonate-responsive ERF transcription factors regulate steroidal glycoalkaloid biosynthesis in tomato. Plant Cell Physiol 57:961–975
Valkonen JPT, Keskitalo M, Vasara T, Pietilä L (1996) Potato glycoalkaloids: a burden or a blessing? Crit Rev Plant Sci 15:1–20
Vranová E, Coman D, Gruissem W (2012) Structure and dynamics of the isoprenoid pathway network. Mol Plant 5:318–333
Wang Z, Guhling O, Yao R, Li F, Yeats TH, Rose JKC, Jetter R (2011) Two oxidosqualene cyclases responsible for biosynthesis of tomato fruit cuticular triterpenoids. Plant Physiol 155:540–552
Wentzinger LF, Bach TJ, Hartmann M-A (2002) Inhibition of squalene synthase and squalene epoxidase in tobacco cells triggers an up-regulation of 3-hydroxy-3-methylglutaryl coenzyme A reductase. Plant Physiol 130:334–346
Yoshioka H, Yamada N, Doke N (1999) cDNA cloning of sesquiterpene cyclase and squalene synthase, and expression of the genes in potato tuber infected with Phytophthora infestans. Plant Cell Physiol 40:993–998
Zook MN, Kuć JA (1991) The use of a sterol inhibitor to investigate changes in the rate of synthesis of 2,3-oxidosqualene in elicitor-treated potato tuber tissue. Physiol Mol Plant Pathol 39:391–401
Acknowledgements
Yeast strain GIL77 and the ERG7/YHR072W plasmid were kindly obtained from Dr. Efraim Lewinsohn, Newe Ya’ar Research Center, Israel. We would like to thank Dr. Amir Sherman, Volcani Center, Israel, for his help with establishing the yeast complementation assay. The research was partially supported by Research Grant no. IS-4134-08 from BARD, the United States-Israel Binational Agricultural Research and Development Fund.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, A., Fogelman, E., Weissberg, M. et al. Lanosterol synthase-like is involved with differential accumulation of steroidal glycoalkaloids in potato. Planta 246, 1189–1202 (2017). https://doi.org/10.1007/s00425-017-2763-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2763-z