Abstract
The amino acid sequence of APX4 is similar to other ascorbate peroxidases (APXs), a group of proteins that protect plants from oxidative damage by transferring electrons from ascorbate to detoxify peroxides. In this study, we characterized two apx4 mutant alleles. Translational fusions with GFP indicated APX4 localizes to chloroplasts. Both apx4 mutant alleles formed chlorotic cotyledons with significantly reduced chlorophyll a, chlorophyll b and lutein. Given the homology of APX to ROS-scavenging proteins, this result is consistent with APX4 protecting seedling photosystems from oxidation. The growth of apx4 seedlings was stunted early in seedling development. In addition, APX4 altered seed quality by affecting seed coat formation. While apx4 seed development appeared normal, the seed coat was darker and more permeable than the wild type. In addition, accelerated aging tests showed that apx4 seeds were more sensitive to environmental stress than the wild-type seeds. If APX4 affects seed pigment biosynthesis or reduction, the seed coat color and permeability phenotypes are explained. apx4 mutants had cotyledon chlorosis, increased H2O2 accumulation, and reduced soluble APX activity in seedlings. These results indicate that APX4 is involved in the ROS-scavenging process in chloroplasts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ascorbate peroxidases (APXs) are heme-containing proteins that neutralize peroxides using ascorbate, one of the major antioxidants in plant cells. When superoxides are generated as by-product of photosynthesis or the product of NADPH oxidases, superoxide dismutases (SODs) rapidly convert superoxides into a neutral and relatively stable molecule, H2O2. APXs scavenge H2O2 and neutralize it through the ascorbate–glutathione cycle. In this cycle, another three enzymes, monodehydroascorbate reductase (MDAR/MDHAR), glutathione-dependent dehydroascorbate reductase (DHAR), and glutathione reductase (GR) are involved in regenerating ascorbate back into the cellular antioxidant pool. This ROS-scavenging cycle is active in chloroplasts, mitochondria, the cytosol and peroxisomes (Noctor and Foyer 1998; Asada 1999).
Based on the amino acid sequence similarity to bacterial enzymes, APXs are classified as class I peroxidases along with yeast cytochrome c peroxidase (CCP) and cyanobacterial catalase-peroxidase (CP, Welinder 1992). The crystal structure of a typical APX resembles CP, with 10 α-helixes. APXs have proximal and distal domains, which sandwich the heme-binding region. Amino acid residues in each motif are conserved within APX isoforms. The following are key residues in plant ascorbate peroxidases: (1) Arg38 and His42 are required for catalysis; (2) His163 and His169 bind heme; and (3) Lys30 and Arg172 interact with ascorbate (Henrissat et al. 1990; Schuller et al. 1996; Sharp et al. 2003). Substitution of these residues affects the steady state of catalytic ability or ascorbate utilization of APX (Bursey and Poulos 2000; Celik et al. 2001).
In Arabidopsis, APXs are found in several cellular compartments. APX1 (AT1G07890) and APX2 (AT3G09640) are both cytosolic and protect cells from oxidative stress. Reports show that APX1 is constantly expressed, but is up-regulated by high light, heat, and wounding, while APX2 is only expressed under extreme light stress (Karpinski et al. 1997; Davletova et al. 2005; Maruta et al. 2012). APX3 (AT4G35000) is found in peroxisomes. Increased APX3 activity provided resistance to the oxidative stress present in peroxisome (Wang et al. 1999). The phenotype of apx3 mutants indicates that there is a redundant ROS-scavenging mechanism, which compensates for the loss of apx3 function (Narendra et al. 2006). Thylakoid APX (tAPX, AT1G77490) is in chloroplast membranes with its catalytic domain oriented towards the stroma. The tapx mutant does not show an obvious mutant phenotype, even under oxidative stress (Jespersen et al. 1997; Giacomelli et al. 2007; Kangasjarvi et al. 2008; Kitajima 2008). Stromal APX (sAPX, AT4G08390) has been found in both chloroplasts and mitochondria. During early seedling development, the sapx loss-of-function mutant exhibits chloroplast-bleaching following photo-oxidative stress (Jespersen et al. 1997; Chew et al. 2003; Kangasjarvi et al. 2008).
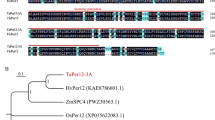
According to amino acid sequence similarity, APX4 (AT4G09010), APX5 (AT4G35970) and APX6 (AT4G32320) are putative Arabidopsis APXs. APX5 is predicted to be microsomal-targeted, but no experimental evidence has been reported (Panchuk et al. 2002). While Arabidopsis APX6 is predicted to be a cytosolic protein (Panchuk et al. 2002), its ortholog in rice (Oryza sativa) was found in chloroplasts and mitochondria, where it interacts with other APXs in these organelles (Lazzarott et al. 2011). Sequence similarity and crystal structure of APX4 reveals this protein resembles other ascorbate peroxidases, but APX4 is missing conservative residues in the catalytic and heme-binding domains (Kieselbach et al. 2000; Teixeira et al. 2004; Granlund et al. 2009; Lunderg et al. 2011). Biochemical evidence indicates that APX4 localizes to the chloroplast lumen (Granlund et al. 2009). Previous researchers reported that APX4 protein expressed in E. coli lacked peroxidase activity and the apx4 mutant showed no obvious phenotype (Granlund et al. 2009). In this study, two mutant alleles, apx4-1 and apx4-2, were characterized. Despite the fact that APX4 lacks ascorbate peroxidase catalytic activity in E. coli, we conclude that APX4 affects seedling growth, seed quality and APX scavenging in plants.
Materials and methods
Plant material
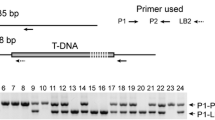
Plants were grown in square 2.5-in. pots with continuous illumination using fluorescent lights (100 μmol m−2 s−1). Arabidopsis thaliana (ecotype Col-0) and two apx4 T-DNA mutant alleles were characterized using ABRC stocks (Alonso et al. 2003). The SAIL_519_E04 and SALK_119726 accessions were named apx4-1 and apx4-2, respectively. Plants were genotyped by polymerase chain reaction (PCR). Two gene-specific primers, F2 (5′-CTGTTCCTTCCTTCACCAAC-3′) and R2 (5′-AGTTTGCTCAGATTGATCCGT-3′), were used to verify the presence of wild-type allele. To identify mutant alleles by PCR, a gene-specific primer (F2 for apx4-1 and R2 for apx4-2) and a T-DNA left border primer (LBb1, 5′-ATTTCGGAACCACCATCAAAC-3′ or Sail_LB, 5′-CATAACCAATCTCGATACACC-3′) were used.
Seed germination, viability, and growth
Seeds were sterilized in either 0.6 or 3 % hypochlorite with 0.01 % Tween-20 for 15 min and washed several times with sterile water. Imbibed seeds (4 °C for 3 days in the dark) were spread on the GM plates. GM plates contain 0.5× MS salts, 1× B5 vitamins, 0.5 % phytoagar, pH 5.8. Sterilized seeds were incubated in a Percival plant growth chamber (Perry, IA, USA) at 22 °C with continuous 50 μmol m−2 s−1 irradiation. Germination was defined as 1 mm radical emergence and recorded daily for a week. For accelerated aging tests, seeds were incubated in a chamber at 41.5 °C with 100 % relative humidity for 3 days and air dried for another 3 days. Aged seeds were sterilized and germinated on GM plates, as described above.
For seedling growth comparison, wild type (Col-0) and two apx4 mutant seeds were grown on GM plates (phytoagar was replaced with 1.5 % agar) for 7 days. Plates were set vertically so the roots grew along the surface of the plates. Images of individual plants were captured. The leaf area was determined using these images, while root length was measured directly. Results were analyzed by ANOVA. Intact and perforated seeds were imbibed in 1 % 2,3,5-triphenyl tetrazolium chloride solution (Sigma, St. Louis, MO, USA) according to Boisson et al. (2001) to determine seed viability and permeability.
Pigment extraction
Fresh weights of 7-day-old plants were measured. Individual plants were ground in liquid nitrogen and pigments dissolved in 1 mL of 100 % acetone for 10 min. Except when samples were vortexed to increase pigment extraction, samples were stored in the dark. The absorbance of the extracts at 662, 645, and 474 nm were used to determine chlorophyll a, chlorophyll b, and lutein levels. Using extinction coefficients for chlorophyll a (82.7 L g−1 cm−1), chlorophyll b (50.7 L g−1 cm−1) and lutein (197.3 L g−1 cm−1), pigment levels were determined (Bulda et al. 2008).
Analysis of APX4 transcripts
Total RNA was isolated from tissues using RNeasy kit (Qiagen, Valencia, CA, USA). Complementary DNA (cDNA) was synthesized using SuperScript II reverse transcriptase (Invitrogen, Grand Island, NY, USA) from 1 μg of total RNA that was primed with 0.5 μg oligo(dT)12–18. APX4 transcripts were amplified using first-strand cDNA and gene-specific primers: to amplify full-length APX4 transcripts F2 and R2 primers were used; to amplify cDNA fragment A (Frag A), F2 and R4 (5′-AGATACGGTTTCACGGCTTC-3′) primers were used; to amplify cDNA fragment B (Frag B), F2 and R5 (5′-CTGCATATGAAATAGGACCTC-3′) primers were used. UBQ10 transcripts were amplified as internal controls using the same first-strand cDNA template and UBQ10 gene-specific primers (5′-GAAAACAATTGGAGGATGGTC-3′ and 5′-GAGACGAGATTTAGAAACCAC-3′).
In situ hybridization
In situ hybridization was performed according to Park et al. (2005). Seven-day-old Arabidopsis seedlings were grown on GM plates with 1 % sucrose, then embedded in paraffin and 10 μm sections mounted on slides. For sense probe synthesis, the following primers were used: 5′-TAATACGACTCACTATAGGGCCTTCACAAAACCAAAACACAC-3′ and 5′-CTTTTGCATCACCCACATTG-3′; for anti-sense probe synthesis, the following primers were used: 5′-TAATACGACTCACTATAGGGCTTTTGCATCACCCACATTG-3′ and 5′-CCTTCACAAAACCAAAACACAC-3′.
GUS staining
To construct APX4::GUS, 1.1 kb of APX4 upstream sequence was PCR amplified and transcriptionally fused to the upstream of the uidA gene start codon, which was in the pHK7 construct (Harikrishna et al. 1996). The APX4::GUS fragment was excised with NotI and inserted into pMLBart (Gleave 1992). This plasmid was moved into Agrobacterium tumefaciens, strain ASE, by electroporation. The T-DNA fragment was mobilized into Arabidopsis plants using the methodology described by Clough and Bent (1998). Plants containing a T-DNA insert were selected by spraying seedlings with a 1,000-fold dilution of BASTA (Finale, AgrEvo, Pikeville, NC, USA). For GUS assays, plant tissues were vacuum infiltrated with substrate solution (1 mM 5-bromo-4-chloro-3-indoyl glucuronide, 50 mM sodium phosphate pH 7.0, 1 % Triton X-100, 1 % DMSO, 10 mM EDTA) and incubated at 37 °C. Seedlings and inflorescences were incubated in substrate for 2 h and 2 days, respectively. Plant pigments were removed by incubating in 70 % ethanol before microscopic examination and image capture.
Localization of GFP translational fusion proteins
A binary plasmid was created that contained the APX4 coding sequence (upstream) translationally fused to the GFP coding sequence (downstream). The APX coding sequence was PCR amplified using these primers: GAATTCTGAAATGGGAGGAGTGTC and GAATTCTTAGCTTGAGTTTGCTCAG. This PCR product was cloned into the EcoRI sites of pBH10 (Hauser et al. 2000). The 35S::APX4-GFP fragment was excised with NotI and inserted into pMLBart (Gleave 1992). Following the transformation method described earlier, this 35S::APX4-GFP construct and the 35S::DsRed-KSRM construct (Cheng et al. 2006) were co-transformed into Col-0 Arabidopsis plants. Transformants containing both transgenes were fixed and examined by confocal microscopy. Plant tissues were submerged in fixative solution containing 4 % paraldehyde, 0.5 % glutaraldehyde, 1 % DMSO, 3 mM DTT, 10 mM Hepes (pH 6.4), 2 mM EDTA, 5.6 mM KCl, and 10 mM MgCl2 and a weak vacuum applied. Samples were washed three times in 10 mM Hepes (pH 6.4), 2 mM EDTA, 5.6 mM KCl, and 10 mM MgCl2 and mounted with 10 % glycerol in 20 mM Tris–HCl (pH 9). GFP and DsRed fluorescence was detected using an Olympus IX81-DSU spinning disk confocal microscope (Center Valley, PA, USA) and images were processed with SlideBook software (3i, Denver, CO, USA). GFP was excited with a 472/30-nm filter and emission detected with a 520/35-nm filter. The filter set used for DsRed excited samples at 562/40 nm and detected emission at 624/40 nm. Chlorophyll autofluorescence was detected using a Cy5 filter (excitation 650/13 nm and emission 684/24 nm). Identical exposure times were used to capture images from transgenic and wild-type plants. Using these images, autofluorescence can be differentiated from GFP and DsRed signals. No detectable GFP and DsRed fluorescence was observed in wild-type plants.
APX activity assay
Zymographs were used to visualize active APX complexes on 10 % native polyacrylamide gel according to Mittler and Zilinskas (1993). Previous researchers observed that chloroplast APX activity was eliminated unless proteins were extracted in the presence of ascorbate (Mittler and Zilinskas 1991; Kitajima 2008). Tissue from 11-day-old seedlings grown on GM plates with 1 % sucrose was frozen in N2(l) and ground to a powder. Total protein was extracted in freshly made buffer that contained 50 mM sodium phosphate buffer (pH 7.0), 5 mM ascorbate, 2 mM EDTA, and 1 mM PMSF. The gel and loading buffer contained 2 and 4.5 mM ascorbate, respectively. Ascorbate altered the rate of acrylamide polymerization so the gel was pre-run for 30 min to allow ascorbate from the running buffer to penetrate. Before the samples were loaded, fresh running buffer was added. Ascorbate peroxidase activity was measured in the native gel with assay buffer, which contained 50 mM potassium phosphate (pH 7.0), 0.5 mM ascorbate, 1 mM hydrogen peroxide, and 0.1 mM EDTA.
The same assay buffer was used to spectrophotometrically measure APX activity in total protein extracts. APX activity was defined as the consumption of ascorbate per microgram total protein per minute, assuming an absorption coefficient of 2.8 mM−1 cm−1 at 290 nm.
H2O2 detection
Amplex Red (Invitrogen) readily penetrates cells and interacts with H2O2 (Ashtamker et al. 2007). Seedlings were immersed in the Amplex Red solution for 10 min in darkness following by a 10-s rinse with sterile water. Fluorescence images were captured with an MZFL dissecting microscope (Leica, Buffalo Grove, IL, USA) using a 546/10-nm narrow-band excitation filter and 590-nm long-pass emission filter.
Results
Chlorotic apx4 cotyledons
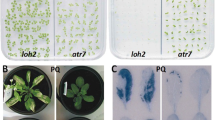
Chlorotic cotyledons were found in two different apx4 mutant alleles, designated as apx4-1 and apx4-2 (Fig. 1e, f). The apx4-1 mutant has a T-DNA insertion in the sixth intron and the apx4-2 mutant has a T-DNA insertion in the last exon (Fig. 1m). In both apx4 alleles, the chlorotic-cotyledon phenotype was incompletely penetrant or had variable expressivity (Fig. 1b, c). This phenotype was observed in 66 and 80 % of the apx4-1 and apx4-2 seedlings, respectively. Seedlings with severely chlorotic cotyledons often died when sown in soil, but they survived when grown on GM plates containing sucrose. This indicates that apx4 seedlings acquired nutrients from the media that they were unable to synthesize in sufficient amounts when grown on soil.
Mutant phenotypes of two apx4 mutant alleles. a–h Arabidopsis plants were grown on GM plates with 1 % sucrose. Compared to 11-day-old wild-type seedlings (a, d), apx4-1 (b, e) and apx4-2 (c, f) exhibited chlorotic cotyledons. g Heteroallelic apx4-1/apx4-2 seedlings showed chlorotic cotyledons, indicating that these mutants are allelic. h The chlorotic-cotyledon phenotype reverted when apx4-1 plants contained the APX4 transgene. i The color of wild-type seeds (left) was lighter than apx4-1 (center), and apx4-2 (right) seeds. Intact (j, k) and perforated (l) seeds were incubated in 1 % tetrazolium solution for 1 day (j) or 2 days (k, l). m The structure of APX4, including the T-DNA insertion sites and primer locations. Boxes show the putative peroxidase domain. Arrows reveal primer locations. Solid boxes exons, open boxes UTRs, striped boxes 3′ UTR of APX4 and an adjacent gene, lines introns or intergenic regions. Size bars are 2.5 mm (d–h) or 1 mm (i–l)
Two findings show that the chlorosis phenotype was caused by mutation of the APX4 locus. First, the heteroallelic apx4-1/apx4-2 plants displayed a chlorotic-cotyledon phenotype, which was indistinguishable from the parental phenotype (Fig. 1g). This eliminates the possibility that a second-site mutation caused the phenotype. Secondly, introducing an intact APX4 gene into the apx4-1 mutant background complemented the chlorotic-cotyledon phenotype (Fig. 1h). This indicates that the genomic fragment containing APX4 was sufficient to compensate for the mutated APX4 locus and eliminate the cotyledon phenotype.
Photosynthetic pigment levels and seedling growth were reduced in apx4 mutants
Seedlings of apx4 mutants were separated into two groups, green cotyledons (GC) and chlorotic cotyledons (CC), and their pigments were extracted. Data revealed that chlorophyll a, chlorophyll b and lutein contents in both apx4 mutant alleles with chlorotic cotyledons were significantly reduced (Fig. 2). Surprisingly, apx4-1 seedlings without apparent chlorotic cotyledons had significantly less chlorophyll and lutein than the wild type. The apx4-2 seedlings with green cotyledons had lower, but not significantly different, levels. The data revealed that cotyledon chlorosis occurred in both groups of seedlings, which shows that the apx4 mutants exhibited variable expressivity and not poor penetrance.
Chlorophyll a, chlorophyll b and lutein levels were lower in apx4 mutants. Pigments were extracted from seedlings with chlorotic cotyledons (CC) and green cotyledons (GC). Significantly decreases (P < 0.01) from the wild-type controls (WT) are indicated (asterisk). Plotted values are mean ± SE (n = 5–19)
Because the biomass of a single Arabidopsis seedling is difficult to measure accurately (Paul-Victor et al. 2010), seedling growth was evaluated by measuring leaf area (including the cotyledons) and root length. Seedlings were grown on GM plates that were oriented vertically so the roots would grow across the surface. In 7-day-old seedlings, the root lengths and leaf area of both apx4 alleles were significantly lower than controls (Fig. 3).
Decreased germination of apx4 seeds
The seed color of apx4 mutants was darker than wild-type seeds (Fig. 1i). Because changes in seed coat color are associated with other seed property modifications (Koornneef 1981; Debeaujon et al. 2001), seed quality and seed vigor were evaluated in apx4 alleles. Only 20 % of the apx4-1 seeds germinated after sterilization in 3 % hypochlorite, but this germination behavior was maternally controlled (Fig. 4a), indicating this phenotype was determined by seed coat properties. When 0.6 % hypochlorite was used for sterilization, freshly harvested wild-type seeds completely germinated in 2 days, whereas apx4 seeds did this in 4 days (Fig. 4b). Germination rates of apx4 significantly decreased after accelerated aging (Fig. 4b) as well as natural aging (Fig. 4c). These results indicate apx4 seeds were less tolerant of harsh environmental conditions, probably due to changes in the apx4 seed coat. In order to determine whether apx4 mutant seeds were viable, wild type and mutant seeds were evaluated to determine if they could oxidize tetrazolium chloride. Once tetrazolium penetrates the seed coat, a metabolically active embryo oxidizes this colorless dye, forming a red pigment. Normally tetrazolium cannot penetrate the Arabidopsis seed coat, so seeds need to be perforated to carry out this test. Results showed that apx4 seeds are metabolically active, but the seed coat of apx4 mutant alleles was more permeable than the wild type since tetrazolium penetrated the seed without perforation (Fig. 1j–l).
The germination of apx4 seeds was lower than the wild-type seeds due to an altered seed coat. a Seeds of wild type (WT), apx4-1 mutants, and apx4-1/+ heterozygotes from reciprocal crosses were sterilized in 3 % hypochlorite and germination rates were plotted. b Control (N.A.) and accelerated aged (A.A.) seeds were sterilized in 0.6 % hypochlorite and the germination was measured. c The germination of wild-type and apx4-2 seeds stored at room temperature for various time periods was tested. Seeds were sterilized in 0.6 % hypochlorite. Values are mean ± SE of three or four biological replicates. Some error bars are smaller than their point
Expression and sub-cellular localization of APX4
As revealed by in situ hybridization, APX4 transcripts were found in leaf mesophyll and phloem (Fig. 5b). β-Glucuronidase (GUS) staining indicated that the APX4 promoter is most active in the cotyledons and first true leaf (Fig. 5c). These results correlate nicely with the posted expression of this gene (Winter et al. 2007). In reproductive organs, APX4 was expressed in sepals (especially in vasculature) and the abscission area at the base of sepals (Fig. 5d, e). Examination of the dissected edge of the carpel wall revealed intense APX4::GUS staining (Fig. 5f), indicating that wounding induced the expression of this gene. According to RT-PCR results, APX4 is expressed in many tissues throughout the aboveground plant, but not in roots (Fig. 6a).
Expression of APX4. Using control sense (a) and APX4 anti-sense (b) RNA probes, in situ hybridizations detected APX4 transcripts in leaf primordia and phloem. c In seedlings containing the APX4::GUS transgene, expression was most abundant in the cotyledon (c) and first leaf (1). d, e In flowers, GUS was found in sepals, anthers, the style, and the petal abscission zone. f In siliques, GUS was present in developing testa, but absent in pigmented seed coats. In plants transformed with APX4-GFP and DsRed-KSRM constructs (g–j), APX4 fluorescence (green, g) co-localized with chlorophyll fluorescence (red, h). i KSRM localized to peroxisomes (blue). j Fluorescent signals were merged. ct cortex, e epidermis, lp leaf primordium, p phloem, s stoma, x xylem. Bar 1 mm (a, b, d–f), 5 mm (c), 10 μm (g–j)
Detection of APX4 transcripts by RT-PCR. a APX4 transcripts were present in siliques (s), rosette leaves (rl), cauline leaves (cl), stage-12 flowers (fl) and pistils (p), but not in roots (r). b No full-length APX4 transcripts were detected in apx4 mutants. c However, truncated transcripts were observed in apx4 seedlings. UBQ10 was used as internal control
APX4-GFP translational fusion protein localized to the chloroplasts (Fig. 5g–j). The chloroplasts in guard cells have a stronger GFP signal than chloroplasts in mesophyll cells. Although APX4 has a SKL motif at its C-terminal and was predicted to be present in peroxisomes (Panchuk et al. 2002), our APX4-GFP translational fusion proteins did not co-localize with the DsRed-KSRM marker protein, which is targeted to peroxisomes. This GFP localization corroborates previous biochemical data showing that APX4 is present in the chloroplast lumen (Granlund et al. 2009).
The apx4 mutants are null alleles
According to the RT-PCR results, no full-length APX4 transcripts were found in either apx4 mutant alleles (Fig. 6b). Two other reverse primers, R4 and R5, amplified partial APX4 cDNA products (Fig. 1m). APX4 fragment A, amplified with F2 and R4 primers, was present in wild-type Arabidopsis, but not in the two apx4 mutants (Fig. 6c). The truncated APX4 cDNA fragment B, amplified with F2 and R5, was found in both apx4 alleles, but the amount was greatly reduced in apx4-2 seedlings (Fig. 6c). If the truncated APX4 transcripts (fragment B) were to be stably translated, the truncated protein would only contain a third of the amino acid sequence of a mature APX4 protein. While the apx4-2 allele is predicted to encode nearly the entire coding sequence of this gene (Fig. 1m), the T-DNA insert disrupted stable transcript accumulation for this allele (Fig. 6c). Data indicate that both apx4-1 and apx4-2 are null alleles.
apx4 mutants have increased H2O2 levels and lower APX activity
To investigate whether APX4 is involved in the ROS-scavenging metabolism, H2O2 levels and soluble APX activity in wild-type plants and apx4 mutants were measured. We observed large H2O2 accumulation in 5-day-old apx4 seedlings, which were higher than the baseline levels in wild-type plants (Fig. 7a). Zymographs were used to visualize active APX complexes on native polyacrylamide gels. No detergent was used on these extracts, so membrane-embedded APX likely were not extracted. Examination of Rubisco extraction efficiency with and without detergent reveals that soluble proteins in organelles were efficiently extracted. In the zymograph, the prominent signal consisted of a doublet of bands, while four minor bands displayed differing migration rates (Fig. 7c). In apx4 alleles, two bands showed diminished intensity when compared to controls (see arrows in Fig. 7c). The kinetic assay measured APX activity by the consumption of ascorbate in an APX-dependent fashion, confirming that the decrease of soluble APX activity in apx4 alleles were significant (P < 0.01).
Mutation of apx4 leads to H2O2 accumulation and reduction of soluble APX activity in seedlings. a In wild-type (left), apx4-1 (right) and apx4-2 (bottom) seedlings, H2O2 levels correlate with fluorescence intensity. b Photograph of the seedlings shown in a. c APX zymograph. Arrows identify bands in wild-type controls, which are missing in mutants. d Kinetics assays showed significantly lower APX activity in 11-day-old apx4 seedlings
Discussion
Analysis of the apx4 mutant lesion
The chlorotic-cotyledon phenotype was distinctive in apx4 mutants, but it did not appear in all apx4 seedlings. Data presented in the results shows that cotyledon chlorosis resulted from recessive mutations in the APX4 locus. In apx4 cotyledons, 66–80 % of the plants exhibited a visible phenotype. Analysis of pigment accumulation in apx4 alleles revealed the range of phenotypes due to varied expressivity—even the apx4 seedlings without obvious chlorosis, exhibited significantly lower pigment levels than the wild-type seedlings.
APX4 regulates seed coat formation and affects seed vigor
Before encountering favorable conditions for germination, the seed coat maintains seed longevity by forming a selective permeable barrier to oxygen, water, and pathogens (Mohamed-Yasseen et al. 1994). Tetrazolium tests and increased sensitivity to hypochlorite indicate that apx4 seed coats were more permeable than the wild-type seeds (Figs. 1j, k, 4b). This change in apx4 seed coat properties not only affected seed permeability, but also influenced seed protection, quality and longevity. After natural aging for 2 years, the germination of apx4 seeds was reduced significantly, possibly due to gradual oxidation of the embryo caused by changes in the permeability of the seed coat (Fig. 4c). Similarly, apx4 seeds are less viable under harsh environmental conditions, such as high humidity and temperature (Fig. 4b).
The seed coat develops from maternal sporophyte tissues: the inner and outer integuments. Germination of apx4 heterozygous seeds exhibited maternal effects (Fig. 4a), indicating that the apx4 seed coat affects germination efficiency. During seed maturation, polymerization of phenolic compounds in the integuments colors the seed coat and establishes a selectively permeable barrier for water and other metabolites (Debeaujon et al. 2000; Moise et al. 2005). In Arabidopsis seeds, colorless proanthocyanidins are synthesized in the inner integument. The color of the seed coat develops after oxidative enzymes, such as peroxidases and laccases, alter phenolic compounds and start the free-radical polymerization of these monomers (Debeaujon et al. 2003; Liang et al. 2006). Mutants that affect seed coat pigmentation sometimes affect seed vigor due to the change in seed coat permeability (Debeaujon et al. 2000). The seed coat of apx4 mutants not only increased seed permeability, but also displayed darker pigmentation (Fig. 1i). Low levels of APX4 expression were detected during seed maturation, but not observed after seed pigmentation (Fig. 5f). Thus, APX4 may affect the seed coat by changing the biosynthesis or oxidation of phenolic precursors in this structure as it develops.
APX4 is needed during seedling photosystem development
In seedlings, apx4 alleles displayed lower growth rates (Fig. 3). Reduced pigment levels in apx4 mutants (Fig. 2) would lead to less light absorption by the photosystems, which would cause reduced biomass accumulation and growth. In 7-day-old apx4 seedlings, the total leaf area was 25 % lower and root length was 20 % shorter than the wild type (Fig. 3). Although APX4 was not expressed in roots, decreased root growth probably resulted from less available photosynthate for root growth. These apx4 phenotypes were only observed in seedlings. The differences in growth between apx4 mutants and wild-type plants become less apparent as plants aged.
Chlorophyll a, b, and lutein are photosynthetic pigments found in light-harvesting complexes (Peter and Thornber 1991). Lower photosynthetic pigment levels could result from lower synthesis or higher degradation rates. Chlorophyll biosynthesis mutants not only showed more yellowing or chlorosis, but also altered chloroplast development (Wu et al. 2007; Lepisto et al. 2009). Biosynthesis mutants would specifically affect chlorophyll or xanthophyll biosynthesis. More likely, APX4 protects seedling photosystems and/or decreases the rate of pigment degradation since, on a percent basis, apx4 exhibited similar reductions in both lutein and chlorophyll levels. sAPX scavenges hydrogen peroxides and protects chloroplasts from photo-oxidative stress in the stroma. Loss-of-function sapx mutant exhibited a seedling-bleaching phenotype under photo-oxidative stress (Chew et al. 2003).
Possible roles for APX4 in seedlings
In apx4 seedlings, we observed increased H2O2 accumulation and reduced soluble APX activity (Fig. 7). APX4 previously was hypothesized to be a non-functioning ascorbate peroxidase because this protein lacks critical residues in the catalytic, heme-binding and ascorbate-binding motifs (Kieselbach et al. 2000; Teixeira et al. 2004; Granlund et al. 2009). We propose that APX4 modulates ROS scavenging through one of two alternative mechanisms.
Since some APXs dimerize (Mittler and Zilinskas 1991), APX4 may bind to and stabilize another APX. However, our yeast two-hybrid analysis failed to identify a direct interaction between APX4 and another APX. While APX4 may have indirect interactions with other APXs, no other Arabidopsis APX has been reported to localize to the chloroplast lumen. These two observations indicate that the above molecular function for APX4 is improbable.
Biochemical data showed that APX4 associates with the water-splitting complex or another PSII subunit (Granlund et al. 2009), so we propose that APX4 may be involved in protecting the photosystems within the thylakoid lumen. Six amino acids that are invariant in active APX proteins, which participate in heme binding, ascorbate binding, or catalysis, differ in APX4 so it is unlikely this protein has canonical APX activity. Granlund et al. (2009) observed that (1) APX4 binds heme, but does so with much lower affinity than other APXs and (2) two APX4 masses were detected in the lumen, which differed by 0.5 kDa. A simple hypothesis, consistent with all data, is that the heme molecule in APX4 proteins scavenges high-energy electrons formed by PSII or the water-splitting complex. Because heme is bound at lower affinity in APX4, oxidized heme might be shuttled from the APX4 protein complex in PSII, thereby reducing the mass of the protein by 616 Daltons, and allowing a fresh heme to enter. This reduction in mass closely matches the 500 Da change reported by Granlund et al. (2009). Depending on the interactions with nearby amino acids, and oxidation and spin of the chelated iron atom, heme can interact with singlet oxygen, superoxides, and peroxides (Jones et al. 2002). Co-expression analysis with AraNet (Lee et al. 2010) identified three genes; two heme biosynthetic genes and ROC4, a locus involved in the repair of photo-damaged PSII (Cai et al. 2008). Coupled expression of heme, ROC4, and APX4 supports this proposed function. Experiments with inhibitors of photosynthetic electron transport and repair are underway in apx4 cotyledons and leaves to test the hypothesis that APX4 associates with PSII in the chloroplast lumen and detoxifies energetic electrons.
Abbreviations
- APX:
-
Ascorbate peroxidase
- CC:
-
Chlorotic cotyledons
- CCP:
-
Cytochrome c peroxidase
- GC:
-
Green cotyledons
- GFP:
-
Green fluorescent protein
- GR:
-
Glutathione reductase
- GUS:
-
β-Glucuronidase
- H2O2 :
-
Hydrogen peroxide
- MDAR/MDHAR:
-
Monodehydroascorbate reductase
- PCR:
-
Polymerase chain reaction
- SOD:
-
Superoxide dismutase
References
Alonso JM, Stepanova AN, Leisse TJ et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Asada K (1999) The water–water cycle in chloroplasts: scavenging of active oxygens and dissipation of excess photons. Annu Rev Plant Physiol Plant Mol Biol 50:601–639
Ashtamker C, Kiss V, Sagi M, Davydov O, Fluhr R (2007) Diverse subcellular locations of cryptogein-induced ROS production in tobacco BY-2 cells. Plant Physiol 143:1817–1826
Boisson M, Gomord V, Audran C, Berger N, Dubreucq B, Granier F, Lerouge P, Faye L, Caboche M, Lepiniec L (2001) Arabidopsis glucosidase I mutants reveal a critical role of N-glycan trimming in seed development. EMBO J 20:1010–1019
Bulda OV, Rassadina VV, Alekseichuk HN, Laman NA (2008) Spectrophotometric measurement of carotenes, xanthophylls, and chlorophylls in extracts from plant seeds. Russ J Plant Physiol 55:544–551
Bursey EH, Poulos TL (2000) Two substrate binding sites in ascorbate peroxidase: the role of arginine 172. Biochemistry 39:7374–7379
Cai W, Ma J, Guo J, Zhang L (2008) Function of ROC4 in the efficient repair of photodamaged photosystem II in Arabidopsis. Photochem Photobiol 84:1343–1348
Celik A, Cullis PM, Sutcliffe MJ, Sangar R, Raven EL (2001) Engineering the active site of ascorbate peroxidase. Eur J Biochem 268:78–85
Cheng N-H, Liu JZ, Brock A, Nelson RS, Hirschi KD (2006) AtGRXcp, an Arabidopsis chloroplastic glutaredoxin, is critical for protection against protein oxidative damage. J Biol Chem 281:26280–26288
Chew O, Whelan J, Millar AH (2003) Molecular definition of the ascorbate-glutathione cycle in Arabidopsis mitochondria reveals dual targeting of antioxidant defenses in plants. J Biol Chem 278:46869–46877
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Davletova S, Rizhsky L, Liang H, Shengqiang Z, Oliver DJ, Coutu J, Shulaev V, Schlauch K, Mittler R (2005) Cytosolic ascorbate peroxidase 1 is a central component of the reactive oxygen gene network of Arabidopsis. Plant Cell 17:268–281
Debeaujon I, Leon-Kloosterziel KM, Koornneef M (2000) Influence of the testa on seed dormancy, germination, and longevity in Arabidopsis. Plant Physiol 122:403–413
Debeaujon I, Peeters AJM, Leon-Kloosterziel KM, Koornneef M (2001) The TRANSPARENT TESTA12 gene of Arabidopsis encodes a multidrug secondary transporter-like protein required for flavonoid sequestration in vacuoles of the seed coat endothelium. Plant Cell 13:853–871
Debeaujon I, Nesi N, Perez P, Devic M, Grandjean O, Caboche M, Lepiniec L (2003) Proanthocyanidin-accumulating cells in Arabidopsis testa: regulation of differentiation and role in seed development. Plant Cell 15:2514–2531
Giacomelli L, Masi A, Ripoll DR (2007) Arabidopsis thaliana deficient in two chloroplast ascorbate peroxidases shows accelerated light-induced necrosis when levels of cellular ascorbate are low. Plant Mol Biol 65:627–644
Gleave AP (1992) A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20:1203–1207
Granlund I, Storm P, Schubert M, Garcia-Cerdan JG, Funk C, Schroder WP (2009) The TL29 protein is lumen located, associated with PSII and not an ascorbate peroxidase. Plant Cell Physiol 50:1898–1910
Harikrishna K, Jampates-Beale R, Milligan SB, Gasser CS (1996) An endochitinase gene expressed at high levels in the transmitting tissue of tomatoes. Plant Mol Biol 30:899–911
Hauser BA, He JQ, Park SO, Gasser CS (2000) TSO1 is a novel protein that modulates cytokinesis and cell expansion in Arabidopsis. Development 127:2219–2226
Henrissat B, Saloheimo M, Lavaitte S, Knowles JKC (1990) Structural homology among the peroxidase enzyme family revealed by hydrophobic cluster analysis. Proteins 8:251–257
Jespersen HM, Kjaersgard IVH, Ostergaard L, Welinder KG (1997) From sequence analysis of three novel ascorbate peroxidases from Arabidopsis thaliana to structure, function and evolution of seven types of ascorbate peroxidase. Biochem J 326:305–310
Jones DK, Patel N, Raven EL (2002) Redox control in heme proteins: electrostatic substitution in the active site of leghemoglobin. Arch Biochem Biophys 400:111–117
Kangasjarvi S, Lepisto A, Hannikainen K, Piippo M, Luomala E-M, Aro E-M, Rintamaki E (2008) Diverse roles for chloroplast stromal and thylakoid-bound ascorbate peroxidases in plant stress responses. Biochem J 412:275–285
Karpinski S, Escobar C, Karpinska B, Creissen G, Mullineaux PM (1997) Photosynthetic electron transport regulates the expression of cytosolic ascorbate peroxidase genes in Arabidopsis during excess light stress. Plant Cell 9:627–640
Kieselbach T, Bystedt M, Hynds P, Robinson C, Schroder WP (2000) A peroxidase homologue and novel plastocyanin located by proteomics to the Arabidopsis chloroplast thylakoid lumen. FEBS Lett 480:271–276
Kitajima S (2008) Hydrogen peroxide-mediated inactivation of two chloroplastic peroxidases, ascorbate peroxidase and 2-Cys peroxiredoxin. J Photochem Photobiol 84:1404–1409
Koornneef M (1981) The complex syndrome of ttg mutants. Arabidopsis Inf Serv 18:45–51
Lazzarott F, Teixeira FK, Rosa SB, Dunand C, Fernandes CL, Fontenele AdeV, Silvera JAG, Verli H, Margis R, Margis-Pinheiro M (2011) Ascorbate peroxidase-related (APx-R) is a new heme-containing protein functionally associated with ascorbate peroxidase but evolutionarily divergent. New Phytol 191:234–250
Lee I, Ambaru B, Thakkar P, Marcotte EM, Rhee SY (2010) Rational association of genes with traits using a genome-scale gene network for Arabidopsis thaliana. Nature Biotech 28:149–156
Lepisto A, Kangasjarvi S, Luomala E-M, Brader G, Sipari N, Keranen M, Keinanen M, Rintamaki E (2009) Chloroplast NADPH-thioredoxin reductase interacts with photoperiodic development in Arabidopsis. Plant Physiol 149:1261–1276
Liang M, Davis E, Gardner D, Gai X, Wu Y (2006) Involvement of AtLAC15 in lignin synthesis in seeds and in root elongation of Arabidopsis. Planta 224:1185–1196
Lunderg E, Storm P, Schroder WP, Funk C (2011) Crystal structure of the TL29 protein from Arabidopsis thaliana: an APX homolog without peroxidase activity. J Struct Biol 176:24–31
Maruta T, Inoue T, Noshi M, Tamoi M, Yabuta Y, Yoshimura K, Ishikawa T, Shigeoka S (2012) Cytosolic ascorbate peroxidase 1 protects organelles against oxidative stress by wounding- and jasmonate-induced H2O2 in Arabidopsis plants. Biochim Biophys Acta 1820:1901–1907
Mittler R, Zilinskas BA (1991) Purification and characterization of pea cytosolic ascorbate peroxidase. Plant Physiol 97:962–968
Mittler R, Zilinskas BA (1993) Detection of ascorbate peroxidase activity in native gels by inhibition of the ascorbate-dependent reduction of nitroblue tetrazolium. Anal Biochem 212:540–546
Mohamed-Yasseen Y, Barringer SA, Splittstoesser WE, Costanza S (1994) The role of seed coats in seed viability. Bot Rev 60:426–439
Moise JA, Han S, Gudynaite-Savitch L, Johnson DA, Miki BLA (2005) Seed coats: structure, development, composition, and biotechnology. In Vitro Cell Dev Biol 41:620–644
Narendra S, Venkataramani S, Shen G, Wang J, Pasapula V, Lin Y, Kornyeyev D, Holaday AS, Zhang H (2006) The Arabidopsis ascorbate peroxidase 3 is a peroxisomal membrane-bound antioxidant enzyme and is dispensable for Arabidopsis growth and development. J Exp Bot 57:3033–3042
Noctor G, Foyer CH (1998) Ascorbate and glutathione: keeping active oxygen under control. Annu Rev Plant Physiol Plant Mol Biol 49:249–279
Panchuk II, Volkov RA, Schöffl F (2002) Heat stress- and heat shock transcription factor-dependent expression and activity of ascorbate peroxidase in Arabidopsis. Plant Physiol 129:838–853
Park SO, Zheng Z, Oppenheimer DG, Hauser BA (2005) The PRETTY FEW SEEDS2 gene encodes an Arabidopsis homeodomain protein that regulates ovule development. Development 132:841–849
Paul-Victor C, Zust T, Rees M, Kliebenstein DJ, Turnbull LA (2010) A new method for measuring relative growth rate can uncover the cost of defensive compounds in Arabidopsis thaliana. New Phytol 187:1102–1111
Peter GF, Thornber JP (1991) Biochemical composition and organization of higher plant photosystem II light-harvesting pigment-proteins. J Biol Chem 266:16745–16754
Schuller DJ, Ban N, van Huystee RB, McPherson A, Poulos TL (1996) The crystal structure of peanut peroxidase. Structure 4:311–321
Sharp KH, Mewies M, Moody PCE, Raven EL (2003) Crystal structure of the ascorbate peroxidase–ascorbate complex. Nat Struct Mol Biol 10:303–307
Teixeira FK, Menezes-Benavente L, Margis R, Margis-Pinheiro M (2004) Analysis of the molecular evolutionary history of the ascorbate peroxidase gene family: inferences from the rice genome. J Mol Evol 59:761–770
Wang J, Zhang H, Allen RD (1999) Overexpression of an Arabidopsis peroxisomal ascorbate peroxidase gene in tobacco increases protection against oxidative stress. Plant Cell Physiol 40:725–732
Welinder KG (1992) Superfamily of plant, fungal and bacterial peroxidases. Curr Opin Struct Biol 2:388–393
Winter D, Vinegar B, Nahal H, Ammar R, Wilson GV, Provart NJ (2007) An “electronic fluorescent pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS 1(2):e718
Wu Z, Zhang X, He B, Diao L, Sheng S, Wang J, Guo X, Su N, Wang L, Jiang L, Wang C, Zhai H, Wan J (2007) A chlorophyll-deficient rice mutant with impaired chlorophyllide esterification in chlorophyll biosynthesis. Plant Physiol 145:29–40
Acknowledgments
We thank Dr. Ning-Hui Cheng in Baylor College of Medicine for the gift of the DsRed-KSRM construct and reviewers for constructive comments. This research was supported by the National Research Initiative of the USDA Cooperative State Research, Education, and Extension Service (grant number 2008-35100-19244).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, YY., Hecker, A.G. & Hauser, B.A. The APX4 locus regulates seed vigor and seedling growth in Arabidopsis thaliana . Planta 239, 909–919 (2014). https://doi.org/10.1007/s00425-014-2025-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-014-2025-2