Abstract
Recognition of bacterial effector proteins by plant cells is crucial for plant disease and defense response signaling. The Xanthomonas campestris pv. vesicatoria (Xcv) type III effector protein, AvrBsT, is secreted into plant cells from Xcv strain Bv5-4a. Here, we demonstrate that dexamethasone (DEX): avrBsT overexpression triggers cell death signaling in healthy transgenic Arabidopsis plants. AvrBsT overexpression in Arabidopsis also reduced susceptibility to infection with the obligate biotrophic oomycete Hyaloperonospora arabidopsidis. Overexpression of avrBsT significantly induced some defense-related genes in Arabidopsis leaves. A high-throughput in planta proteomics screen identified TCP-1 chaperonin, SEC7-like guanine nucleotide exchange protein and calmodulin-like protein, which were differentially expressed in DEX:avrBsT-overexpression (OX) Arabidopsis plants during Hp. arabidopsidis infection. Treatment with purified GST-tagged AvrBsT proteins distinctly inhibited the growth and sporulation of Hp. arabidopsidis on Arabdiopsis cotyledons. In contrast, DEX:avrBsT-OX plants exhibited enhanced susceptibility to Pseudomonas syringae pv. tomato (Pst) DC3000 infection. Notably, susceptible cell death and enhanced electrolyte leakage were significantly induced in the Pst-infected leaves of DEX:avrBsT-OX plants. Together, these results suggest that Xcv effector AvrBsT overexpression triggers plant cell death, disease and defense signaling leading to both disease and defense responses to microbial pathogens of different lifestyles.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants have evolved a sensitive and multilayered innate immune system to combat a diverse range of microbial pathogens. Likewise, pathogens have evolved virulence strategies to effectively subvert plant immunity (Jones and Dangl 2006). The induction of plant immunity in response to a bacterial plant pathogen relies on the recognition of bacterial type III effector (T3SE) proteins that are directly injected into the host cell via a type III secretion system (T3SE) (Büttner and He 2009). T3SE have diverse functions and target multiple host pathways for the induction of defense-related genes, the induction of downstream defense response pathways, the modification of specific proteins, and the generation of signaling molecules such as salicylic acid (SA), jasmonic acid (JA) and ethylene (ET) (Stulemeijer and Joosten 2008). T3S effectors, known as avirulence (Avr) proteins, are often associated with the hypersensitive response (HR) in host plants and may lead to effector-triggered immunity (ETI) responses sufficient to prevent pathogen attack.
Xanthomonas campestris pv. vesicatoria (Xcv) causes bacterial spot disease on leaves and fruits of pepper (Capsicum annuum L.) and tomato (Solanum lycopersicum) (Jones et al. 1998, 2004). Most Xcv strains are virulent on either pepper or tomato plants, although some are virulent on both (pepper/tomato group). Infection with Xcv Bv5-4a strain, a tomato group strain, produces a strong HR-like cell death response in pepper, while causing a susceptible disease response in tomato (Kim et al. 2010). The Xcv Bv5-4a strain contains the type III effector protein AvrBsT (Jones et al. 1998; Kim et al. 2010).
AvrBsT is a member of the YopJ/AvrRxv family identified from Xcv (Lewis et al. 2011). This family contains four known members in Xanthomonas: XopJ, AvrXvr4, AvrRxv and AvrBsT (Büttner and Bonas 2010). The YopJ family proteins share a conserved catalytic core, consisting of three key amino acid residues (His, Glu, and Cys) that are identical to the clan CE (C55 family) cystein proteases (Roden et al. 2004; Büttner and Bonas 2010). AvrBsT is required for Xcv fitness and disease symptom development in infected leaves and is responsible for triggering the HR in resistant plants (Büttner and Bonas 2010). AvrBsT alters phospholipid signaling, resulting in defense activation in Arabidopsis (Kirik and Mudgett 2009). Some members of the YopJ/AvrRxv family exhibit weak small ubiquitin-related modifier (SUMO) protease and/or acetyltransferase activity in planta; however, SUMO protease activity of AvrBsT has not been shown (Orth et al. 2000; Szczesny et al. 2010). Genetic studies using SOBER1 mutant Arabidopsis plants have demonstrated that a carboxylesterase functions as a suppressor of AvrBsT-elicited resistance 1 (SOBER1). This leads to the suppression of AvrBsT-elicited HR (Cunnac et al. 2007). However, it is still unknown about the host targets of these enzymes.
Overexpression of type III effector proteins in transgenic plants causes alterations in disease and defense responses during microbial infection. Constitutive expression of the ethanol-inducible Xanthomonas outer protein J (XopJ), a member of the YopJ/AvrRxv family, in transgenic Arabidopsis leaves strongly compromises the callose deposition elicited by a non-pathogenic Pseudomonas syringae pv. tomato DC3000 hrcC mutant (Bartetzko et al. 2009). A stable transgenic Arabidopsis line carrying DEX-inducible avrB exhibits leaf chlorosis (Nimchuk et al. 2000; Eitas et al. 2008). In AvrRpt2-expressing transgenic Arabidopsis plants, there is a loss of the RIN4 protein that interacts with AvrB and AvrRpm1 (Axtell and Staskawicz 2003). This suggests that AvrRpt2 either causes the degradation of RIN4 or inhibits the translation of RIN4 mRNA. Overexpression of Pseudomonas syringae pv. tomato DC3000 T3SE HopF2 in transgenic Arabidopsis plants compromises AvrRpt2-induced ETI and HR (Wilton et al. 2010).
In previous studies, we demonstrated that X. campestris pv. vesicatoria type III effector AvrBsT is differentially recognized by pepper and tomato plants (Kim et al. 2010), and it was hypothesized that pepper plants may contain certain AvrBsT recognition factors that are lacking in tomato plants. To shed further light on the possible functions of Xcv effector AvrBsT, we investigated the disease and defense responses of transgenic Arabidopsis plants expressing avrBsT under a dexamethasone (DEX)-inducible promoter during Hyaloperonospora arabidopsidis (Hpa) and Pseudomonas syringae pv. tomato (Pst) DC3000 and DC3000 (avrRpm1) infection. Notably, the ectopic expression of avrBsT triggered cell death response in transgenic Arabidopsis plants. Two-dimensional (2D) electrophoresis was also used to identify differentially expressed proteins in DEX:avrBsT overexpression (OX) Arabidopsis plants during Hyaloperonospora arabidopsidis infection. In this study, we show that Xcv effector AvrBsT overexpression in heterologous Arabidopsis plants compromises the defense response to the biotrophic oomycete pathogen Hyaloperonospora arabidopsidis while enhancing infection with the hemibiotrophic bacterial pathogen Pseudomonas syringae pv. tomato. Overexpression of avrBsT positively regulated the levels of TCP-1 chaperonin and SEC7-like guanine nucleotide exchange protein, proteins known to play important roles in cell fate determination. SEC7-like guanine nucleotide exchange protein is a putative guanyl-nucleotide exchange factor, and Arabidopsis eds10 mutants exhibit defective embryo development (Pagnussat et al. 2005). The TCP-1 chaperonin or heat shock protein 60 has been implicated in apoptosis and the immune responses of human cells (Kirchhoff et al. 2002; Tsan and Gao 2009). In contrast, avrBsT overexpression negatively regulated the levels of calmodulin-like protein, which was shown previously to be involved in modulating various stress signaling pathways (Luan 2009; Cheong et al. 2010).
Materials and methods
Plant growth conditions
Seeds of wild-type Arabidopsis (Arabidopsis thaliana) ecotype Columbia 0 (Col-0) and DEX:avrBsT overexpression (OX) plants were sown on potting soil mix (peat moss:vermiculite:perlite, 3:1:1, v/v/v). The Arabidopsis plants were cultured in a growth chamber under a 14/10 h (light/dark) light cycle at 60 % humidity and 24 °C. Before sowing on soil, Arabidopsis seeds were vernalized at 4 °C under low light conditions for 2 days.
Generation of avrBsT-OX Arabidopsis
To generate avrBsT-OX Arabidopsis plants, the open reading frame (ORF) of avrBsT was inserted into the XhoI and SpeI sites of pTA7002, a vector containing the dexamethasone (DEX)-inducible promoter. The resulting construct, DEX:avrBsT, was used for Agrobacterium-mediated transformation of Arabidopsis thaliana ecotype Col-0 using a floral dip method, as described by Clough and Bent (1998). Arabidopsis transformants carrying DEX:avrBsT were selected by growing on Murashige–Skoog (MS) medium containing hygromycin (50 μg mL−1). To express avrBsT, transgenic plants were sprayed with 10–20 μM dexamethasone.
Pathogen inoculation and disease rating
Pseudomonas syringae pv. tomato (Pst) DC3000 and DC3000 (avrRpm1) were used to infiltrate the leaves of 5-week-old Arabidopsis plants (Choi and Hwang 2011). To prepare the bacterial inoculum, Pst was grown overnight in yeast nutrient (YN) broth containing kanamycin (50 μg mL−1) and rifampicin (50 μg mL−1) at 28 °C. To monitor bacterial growth [colony forming units (cfu) cm −1] in leaf tissues, the infected leaves were harvested 0 and 3 days after infiltration (105 cfu mL−1).
Hyaloperonospora arabidopsidis isolate Noco2 was maintained on Arabidopsis Col-0 seedling plants to provide a source of fresh inoculum (Hwang and Hwang 2010). A Hp. arabidopsidis suspension of asexual inoculum (5 × 104 conidiosporangia mL−1) was spray-inoculated onto the cotyledons of 10-day-old seedlings. The inoculated plants were covered with a transparent dome to maintain a high relative humidity (80–100 %), followed by incubation in a growth chamber at 17 °C. To determine the asexual sporulation of Hp. arabidopsidis, the number of sporangiophores was counted on both sides of the cotyledons 5 days after inoculation. Visual disease ratings were categorized into four classes based on the number of sporangiophores: 0, 1 to 9, 10 to 19, and over 20 sporangiophores per cotyledon (McDowell et al. 2000).
Reverse-transcription PCR and real-time quantitative PCR
Total RNA from Arabidopsis leaves was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA) following the manufacturer’s instructions. cDNAs were synthesized from total RNA (2 μg) using moloney murine leukemia virus reverse transcriptase (MMLV RT) (Enzynomics, Daejeon, Korea). Amplification of constitutively expressed Actin served as an internal control in the RT-PCR assays. Real-time quantitative PCR was conducted using the Bio-Rad iCycler System with EvaGreen qPCR master mix (ABM, Canada). Expression of defense response genes was normalized to the expression of Actin.
Protein extraction and immunoblotting
Total proteins were extracted by grinding Arabidopsis leaves with the buffer [50 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), pH 7.4, 50 mM NaCl, 10 mM ethylenediamine tetraacetic acid (EDTA), 0.2 % Triton X-100, and 1× proteinase inhibitor cocktail (Roche)] (Hwang and Hwang 2010). After centrifuging the protein extracts at 12,000g for 15 min, the supernatants were used for immunoblotting. Protein extracts were resuspended in sodium dodecyl sulfate (SDS) sample loading buffer, subjected to SDS-PAGE, and transferred onto Hybond-P membranes (GE Healthcare) by wet electroblotting. Proteins were detected using anti-c-Myc antibody (Sigma) diluted to 1:2,000 and anti-rabbit secondary antibody diluted to 1:10,000.
Two-dimensional (2D) electrophoresis
For 2D-electrophoresis (Choi and Hwang 2011), total soluble proteins were extracted from the cotyledons of pathogen-inoculated Arabidopsis seedlings by grinding in extraction buffer [10 % (w/v) trichloroacetic acid and 0.07 % (w/v) dithiothreitol (DTT) in cold acetone (−20 °C)]. Protein extracts (800 μg) suspended in rehydration buffer [9 M urea, 100 mM DTT, 4 % (w/v) CHAPS, 0.5 % (v/v) Bio-lyte 3/10 carrier ampholytes, and 0.002 % bromophenol blue) were applied to an immobilized pH gradient (IPG) strip for in-gel rehydration. Strips were rehydrated at 50 V for 24 h. Isoelectric focusing (IEF) was performed using 24 cm IPG strips (Bio-Rad) with a nonlinear pH 4–7 gradient. IEF was run using a PROTEAN IEF Cell (Bio-Rad, Hercules, CA, USA) at gradient steps of 250 V for 1 h, 500 V for 1 h, 1,000 V for 2 h, and 10,000 V for 4 h, and a final step of 10,000 V toward a total of 90 kVh. After IEF, proteins were separated according to size. IPG strips were incubated in equilibration buffer [50 mM Tris–HCl, pH 8.8, 6 M urea, 30 % (v/v) glycerol, and 2 % (w/v) SDS] containing 1 % (w/v) DTT for 15 min for the first equilibration step, followed by incubation in 4 % (w/v) iodoacetamide for the second step. For 2D-polyacrylamide gel electrophoresis (PAGE), IPG strips were sealed on the top of the 12.5 % 2-dimensional gel using 1 % low-melting agarose in SDS-electrophoresis buffer (25 mM TRIS, 0.2 M glycine, 0.1 % SDS). SDS-PAGE was performed in an Ettan DALTsix electrophoresis unit (GE Healthcare) at 5 W per gel for 1 h, followed by 15 W per gel for 6 h until the bromophenol blue front had reached the end of the gel. Gels were stained with Coomassie blue [0.1 % (w/v) Coomassie Brilliant Blue G 250, 34 % (v/v) methanol, 3 % (v/v) phosphoric acid, and 17 % (w/v) ammonium sulfate]. Coomassie-stained gels were scanned with the UMAX PowerLook 1100XL scanner, and the images were analyzed using ImageMaster 2D Platinum 6.0 (GE Healthcare). The 2D gels were compared and matched, and a quantitative determination of the spot volumes was performed.
Identification of proteins by MALDI-TOF/MS
The differentially expressed protein spots in healthy and infected plants were excised from the Coomassie-stained 2D gels, digested with trypsin, and identified using MALDI-TOF/MS (Choi and Hwang 2011). The resulting peptide mass, pI, and molecular mass analyzed by Ettan MALDI-TOF/MS (GE Healthcare) were identified via NCBI (http://www.ncbi.nlm.nih.gov/) searching using the Profound database (http://prowl.rockefeller.edu/prowl-cgi/profound.exe) for peptide mass fingerprinting.
Database search for the functional category
The identified Arabidopsis proteins were functionally categorized based on the Munich Information Center for Protein Sequences (MIPS) Arabidopsis thaliana genome database (MatDB, http://mips.gsf.de/proj/funcatDB/search_main_frame.html) (Table 1).
Purification of recombinant AvrBsT protein
The avrBsT coding sequence was cloned into the BamHI and XhoI sites of the pGEX-5X-1 vector (GE Healthcare) to produce AvrBsT fused with a GST tag at the N terminus. The recombinant plasmid was transformed into E. coli strain BL21 for protein expression. Transformed bacteria were incubated in 50 mL Luria–Bertani (LB) medium supplemented with ampicillin (50 μg mL−1) at 37 °C. When the OD600 reached approximately 0.5, protein production was induced by adding 1 mM isopropyl β-d-thiogalactopyranoside (IPTG) to the cultures. This was followed by incubation at room temperature for 8 h to enable protein production. pGEX-5X-1:AvrBsT was purified by incubation with glutathione Sepharose beads for 2 h at room temperature and subsequent elution with 50 mM Tris (pH 8.0). Purified AvrBsT was subjected to 10 % SDS-PAGE and stained with Coomassie Brilliant Blue to confirm sample purity.
Measurement of ion conductivity
For the ion conductivity assay, 8 leaf discs (1 cm in diameter) were collected from the leaves infected with Pst (108 cfu mL−1) and washed in 20 mL of double-distilled water for 30 min (Hwang et al. 2011). The washed leaf discs were transferred into 20 mL of double-distilled water. To quantify the electrolyte leakage from the leaf discs, ion conductivity was measured using a conductivity meter (Model sensION7 HACH, CO).
H2O2 measurement
H2O2 production in leaf tissues was spectrophotometrically determined using the xylenol orange assay. Xylenol orange forms a complex with the Fe3+ produced by the hydroperoxide-based oxidation of Fe2+ (Bindschedler et al. 2001). 1 ml of assay reagent [25 mM FeSO4 and 25 mM (NH4)2SO4 dissolved in 2.5 M H2SO4] was added to 100 mL of solution containing 125 μM xylenol orange and 100 mM sorbitol. To measure the H2O2 quantity, leaf discs were floated for 10 min in 1 mL of distilled water and centrifuged at 5,000g for 10 min. 100 μl of the supernatant was added to 1 mL of xylenol orange reagent. After 30 min of incubation, H2O2 production was determined by measuring the A 560 of the Fe3+-xylenol orange complex.
Staining with trypan blue, DAB and aniline blue
To visualize plant cell death and growth of Hp. arabidopsidis isolate Noco2, healthy and infected plant tissues were stained with lactophenol-trypan blue (10 mL lactic acid, 10 mL glycerol, 10 g phenol and 10 mg trypan blue dissolved in 10 mL distilled water) and destained with chloral hydrate (2.5 g mL−1) (Hwang and Hwang 2010). H2O2 production was monitored by staining the leaves with 1 mg mL−1 3, 3′-diaminobenzidine (DAB) (Sigma) (Hwang and Hwang 2010) and cleared with alcoholic lactophenol. The leaf tissues were mounted in 70 % glycerol for observations with a light microscope. For detection of callose deposition by avrBsT expression, leaves were cleared with alcoholic lactophenol and stained in 0.01 % (w/v) aniline blue in 0.15 M K2HPO4 (pH 9.5). The samples were incubated in the dark for 30 min, mounted on slides, and observed under UV illumination.
Results
The ectopic expression of avrBsT triggers cell death response in transgenic Arabidopsis plants
We previously reported that the Xanthomonas campestris pv. vesicatoria (Xcv) effector AvrBsT is differentially recognized as an avirulence and virulence factor by pepper and tomato plants, respectively (Kim et al. 2010). To investigate whether the Xcv effector AvrBsT induces defense in plant cells, we attempted to generate transgenic Arabidopsis plants expressing avrBsT using the cauliflower mosaic virus promoter (CaMV 35S). However, the 35S:avrBsT transgenic lines were not generated using the CaMV vector, possibly due to the lethal effect of avrBsT during its transformation in plants. Thus, we used a dexamethasone (DEX)-inducible system to generate transgenic plants that overexpressed DEX:avrBsT in Arabidopsis (designated DEX:avrBsT-OX). The transgenic Arabidopsis lines #3, #4 and #5 were selected. Constitutive expression of the DEX:avrBsT transgene in the leaves of these Arabidopsis lines was confirmed by RT-PCR and immunoblot analyses (Fig. 1a, b). The transcriptional and translational ectopic expression of avrBsT was detected in the transgenic lines 24 and 48 h after DEX treatment.
Expression of avrBsT in wild-type (WT) and DEX:avrBsT-OX transgenic Arabidopsis plants treated with dexamethasone (DEX). a RT-PCR analysis of the expression of avrBsT in leaves of WT and DEX:avrBsT-OX plants (lines #3, #4 and #5). Three independent experiments were performed with similar results. h hours after DEX treatment. b Immunological detection of AvrBsT proteins under the control of a DEX-inducible system. Total proteins were extracted 24 and 48 h after treatment with DEX, and cMyc-tagged DEX:AvrBsT was detected by immunoblotting with an anti-cMyc antibody. Three independent experiments were performed with similar results. h hours after DEX treatment
We next investigated whether DEX:avrBsT overexpression triggers cell death phenotype in healthy transgenic Arabidopsis plants. In DEX:avrBsT-OX transgenic plants, avrBsT induction by DEX (10 μM) treatment caused brownish necrotic lesions on the cotyledons; however, wild-type cotyledons were not affected by DEX (Fig. 2a). When stained with trypan blue, weak cell death response was microscopically observed in cotyledon tissues of the transgenic plants 12 h after DEX treatment. However, all the plants non-treated with DEX did not show any cell death phenotype at different plant developmental stages (Supplementary Fig. 1). Reactive oxygen species (ROS) accumulation is often involved in cell death response (Choi et al. 2012). The 3, 3′-diaminobenzidine (DAB) staining was used to detect H2O2 accumulation at the early stage of cell death induction by avrBsT expression. Expectedly, the cotyledons of DEX:avrBsT-OX transgenic lines significantly showed reddish brown regions 12 h after DEX treatment, compared to wild-type plants (Fig. 2a, bottom panel). This indicates that avrBsT overexpression induces H2O2 accumulation in the transgenic plants. We further tested whether avrBsT overexpression also induces cell death response in leaves of different ages (Fig. 2b, c). Significantly chlorotic and necrotic cell death phenotypes were observed in 2- or 5-week-old leaves of DEX:avrBsT-OX transgenic plants 24 h after DEX treatment. However, all the plants non-treated with DEX did not show any cell death phenotype on the leaves (Supplementary Fig. 1). The assessment of cell death levels (scale 0~3) (Fig. 2c) supports avrBsT-triggered cell death response in the transgenic leaves. To quantify the cell death response in DEX-treated transgenic leaves, we examined the time-courses of electrolyte leakages via the measurement of ion conductivity. Electrolyte leakage increased dramatically from the leaves of DEX:avrBsT-OX transgenic plants, compared to the wild-type leaves (Fig. 2c).
AvrBsT-triggered cell death phenotypes in DEX:avrBsT-OX transgenic Arabidopsis plants. a Visible and microscopic images of cotyledons of 7-day-old plants treated with dexamethasone (DEX, 10 μM). Cotyledons were stained with lactophenol-trypan blue (middle panel) and 3, 3′-diaminobenzidine (DAB) (bottom panel) 12 h after DEX treatment. Bars 200 μm. b Visible images and cell death levels of 2-week-old leaves 24 h after treatment with DEX (10 μM). Images in the boxes were enlarged in the middle panel. The extent of avrBsT-induced cell death was assessed based on four cell death scales (0, <10 %; 1, 10~30 %; 2, 30~80 %; 3, 80~100 %). c Visible images, cell death levels and electrolyte leakage assay of 5-week-old leaves treated with DEX (10 μM). d Quantitative real-time PCR analysis of the expression of avrBsT and defense-related genes in the leaves of wild-type Col-0 and DEX:avrBsT-OX Arabidopsis plants 24 h after DEX treatment. Actin was used as an internal control for normalization. The data (b–d) represent the mean ± standard deviations from three independent experiments. Different letters indicate significant differences, as analyzed by the LSD test (P < 0.05)
Quantitative real-time PCR analysis was used to investigate whether avrBsT overexpression triggers some defense and cell death response genes in the leaves of DEX:avrBsT-OX Arabidopsis plants (Fig. 2d). DEX treatment strongly induced avrBsT expression in the leaves of the transgenic DEX:avrBsT-OX plants, confirming that DEX:avrBsT was expressed in the transgenic Arabidopsis plants. Induction of avrBsT in Arabidopsis by DEX treatment significantly triggered programed cell death (PCD) marker genes PR1 (Reymond and Farmer 1998) and SAG13 (Brodersen et al. 2002). Interestingly, cell death severity and PR1 expression were correlated with the levels of avrBsT expression in the transgenic leaves. Collectively, overexpression of Xcv effector avrBsT could mediate upstream signaling of PCD response in Arabidopsis.
Aniline blue staining assay was used to investigate whether avrBsT overexpression induces callose accumulation in leaves of DEX:avrBsT-OX transgenic Arabidopsis plants after DEX treatment (Fig. 3). Callose, a β-1,3 glucan, is synthesized to form papillae during defense response against pathogen infection (Flors et al. 2005; Choi and Hwang 2011). The ectopic expression of avrBsT stimulated callose deposition in healthy Arabidopsis cells. Callose deposition is generally involved in pathogen-associated molecular pattern (PAMP)-triggered immunity, which is inhibited by bacterial effectors (Debroy et al. 2004; Kim et al. 2009). Therefore, the avrBsT-triggered callose deposition suggests that AvrBsT may positively regulate basal defense in plants.
Induction of callose deposition by the expression of avrBsT in leaves of DEX:avrBsT-OX Arabidopsis plants. Leaf samples were stained with aniline blue 24 h after DEX treatment (10 μM) (upper panel). Bars 100 μm. Callose deposits in the DEX-treated leaves were counted with the ImageJ software (lower panel). The data represent the mean ± standard deviations from three independent experiments. Different letters indicate significant differences, as analyzed by the LSD test (P < 0.05)
Enhanced resistance of DEX:avrBsT-OX Arabidopsis plants to Hyaloperonospora arabidopsidis infection
To investigate the role of the Xcv effector AvrBsT in the plant defense response, wild-type and DEX:avrBsT-OX plants were spray-inoculated with Hyaloperonospora arabidopsidis isolate Noco2, known to be virulent to Arabidopsis Col-0. The sporangiophores began to form on the cotyledons 4 to 5 days after inoculation with Hp. arabidopsidis (5 × 104 conidiosporangia mL−1). The avrBsT overexpression induced by DEX treatment significantly compromised mycelial growth and sporulation on the cotyledons of the DEX:avrBsT-OX plants (Fig. 4a, b). Notably, the DEX:avrBsT-OX plants treated with DEX exhibited a slight cell death phenotype at the site of oomycete infection, as evidenced by trypan blue staining (Fig. 4a). As shown in Fig. 4b, fewer sporangiophores formed on the cotyledons of DEX-treated DEX:avrBsT-OX plants compared to mock-treated DEX:avrBsT-OX plants. The Hp. arabidopsidis infection also promoted H2O2 production in DEX-treated DEX:avrBsT-OX plants compared to mock-treated DEX:avrBsT-OX plants (Fig. 4c). DAB polymerizes instantly and locally upon contact with H2O2 to form reddish-brown polymers. DEX-treated DEX:avrBsT-OX plants inoculated with Hp. arabidopsidis exhibited DAB-stained spots, indicating H2O2 accumulation to high levels. Together, these data indicate that avrBsT overexpression enhances basal defense to infection by the obligate biotrophic oomycete Hp. arabidopsidis.
Quantification of Hyaloperonospora arabidopsidis infection phenotypes on the cotyledons of wild-type (WT) and DEX:avrBsT-OX Arabidopsis plants (lines #3, #4 and #5). 10-day-old seedlings were spray-inoculated with Hp. arabidopsidis isolate Noco2 (5 × 104 conidiosporangia mL−1) and pathogen development was recorded. a Disease symptoms and cellular responses in the infected cotyledons. Photographs were taken 6 days after inoculation (upper panel). Infected cotyledons were stained with lactophenol-trypan blue 7 days after inoculation to visualize the pathogen mycelium and necrotic plant cells (lower panel). b Sporulation levels of Hp. arabidopsidis on the infected cotyledons. Production of sporangiophores on 50 cotyledons was observed using a stereo-microscope 6 days after inoculation. c Quantification of H2O2 in the leaves 0, 2 and 4 days after inoculation (dpi) with Hp. arabidopsidis. The data (b and c) represent the mean ± standard deviations from three independent experiments. Different letters indicate significant differences from three independent experiments, as analyzed by the LSD test (P < 0.05). dai, days after inoculation
Proteomics analysis of DEX:avrBsT-OX plants infected by Hyaloperonospora arabidopsidis isolate Noco2
To investigate whether avrBsT overexpression regulates the plant proteome during oomycete pathogen infection, 2D-electrophoresis analyses were performed using soluble protein extracts from healthy and Hp. arabidopsidis-infected cotyledons of DEX:avrBsT-OX plants. Coomassie blue staining of the 2D gels revealed some differentially expressed proteins in response to Hp. arabidopsidis infection in the DEX:avrBsT-OX plants (Fig. 5). The differentially expressed protein spots were visualized in 2D gel images, excised and analyzed using MALDI-TOF. Arabidopsis genome sequence analysis identified three proteins (H1, H2 and H3) that were up- or down-regulated (Fig. 6a; Table 1). Protein spots H1 and H2, distinctly upregulated during Hp. arabidopsidis infection, were TCP-1/cpn60 chaperonin family protein (AT5G26360) and SEC7-like guanine nucleotide exchange protein (AT1G01960), respectively. The TCP-1/cpn60 chaperonin family protein also known as heat shock protein 60 has not been extensively studied in Arabidopsis. However, it has been implicated in apoptosis and immune responses in human (Kirchhoff et al. 2002; Tsan and Gao 2009). SEC7-like guanine nucleotide exchange protein was found to be essential for nominal embryo development in Arabidopsis (Pagnussat et al. 2005). Protein spot H3 represented calmodulin-like protein (At2G41100) and was downregulated in DEX:avrBsT-OX plants during infection. Calmodulins are well known to act as signal modulators of stress responses (Luan 2009; Cheong et al. 2010). Real-time quantitative PCR analysis confirmed avrBsT overexpression in leaves of DEX:avrBsT-OX plants by DEX treatment (Fig. 5b). However, Hp. arabidopsidis infection was not effective in enhancing avrBsT overexpression in DEX-treated transgenic plants. H1 (AT5G26360) expression was strongly elevated in both wild-type and DEX:avrBsT-OX plants 3 days after inoculation with Hp. arabidopsidis, but this expression was significantly reduced 5 days after inoculation. H2 (AT1G01960) was significantly induced by avrBsT overexpression and the induction was threefold more in transgenic leaves 3–5 days after Hp. arabidopsidis infection. However, no significant differences in the expression levels of H3 (At2G41100) were detected among all these plants 3 days after inoculation with Hp. arabidopsidis. Interestingly, 5 days after Hp. arabidopsidis infection, H3 (At2G41100) was significantly induced in both wild-type and mock-treated DEX:avrBsT-OX plants, but not in DEX-treated DEX:avrBsT-OX plants. This indicates that the overexpression of avrBsT by DEX treatment suppresses H3 (At2G41100) induction by Hp. arabidopsidis infection. Based on the MIPS Arabidopsis genome database, putative functions of H1, H2, and H3 were identified to be associated with protein folding, metabolism regulation and protein binding, respectively (Table 1). Collectively, these results suggest that avrBsT overexpression in Arabidopsis may differentially regulate diverse proteins of different functions during Hp. arabidopsidis infection.
Two-dimensional electrophoresis of total soluble proteins from the cotyledons of wild-type (healthy) and DEX:avrBsT-OX Arabidopsis plants (line #4) infected with Hyaloperonospora arabidopsidis isolate Noco2. SDS-PAGE gels were stained with Coomassie blue. The circled spots indicate proteins differentially expressed in DEX-treated or mock-treated DEX:avrBsT-OX plants during infection. The spot numbers in the circles correspond to the identifications listed in Table 1
Enlargement of the 2-D gel regions that show the proteins differentially expressed in the cotyledons of mock-treated or DEX-treated DEX:avrBsT-OX plants during Hyaloperonospora arabidopsidis infection. a The histograms show quantitative changes in spot density, as calculated with ImageMaster 2D Platinum 6.0 (GE healthcare), for individual proteins. H1 TCP-1/cpn60 chaperonin family protein (accession no. NP_198008), H2 SEC7-like guanine nucleotide exchange protein (accession no. NP_171698), H3 calmodulin-like protein (accession no. AAN15355). Three independent experiments were performed with similar results. b Real-time quantitative PCR analysis of the expression of avrBsT, H1(AT5G26360), H2 (AT1G01960) and H3 (At2G41100) in wild-type (WT), mock-treated or DEX-treated DEX:avrBsT-OX leaves 0, 3 and 5 days after inoculation with Hp. arabidopsidis. Actin was used as an internal control for normalization of the data. Different letters (a, b, and c) indicate significant differences from three independent experiments, as analyzed by the LSD test (P < 0.05). dai, days after inoculation
Antimicrobial activity of purified AvrBsT against Hyaloperonospora arabidopsidis isolate Noco2
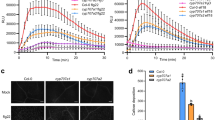
To determine whether AvrBsT has antimicrobial activity against Hp. arabidopsidis, GST-tagged recombinant AvrBsT protein (10 μg mL−1) was applied to Arabidopsis seedlings. These seedlings were subsequently inoculated with Hp. arabidopsidis isolate Noco2. Treatment with 10 μg mL−1 of the purified recombinant AvrBsT protein induced the cell death phenotype in Arabidopsis seedlings (Fig. 7a). However, treatments with lower doses (1–5 μg mL−1) did not cause damage to plant growth (data not shown), but significantly suppressed Hp. arabidopsidis infection and sporangiophore formation on Arabidopsis cotyledons (Fig. 7b, c). Trypan blue staining indicated that treatment with 5 μg mL−1 of the recombinant AvrBsT protein distinctly suppressed sporulation of Hp. arabidopsidis on the cotyledon (Fig. 6b). Hp. arabidopsidis infection on the Arabidopsis seedlings treated with the recombinant AvrBsT protein resulted in lower levels of sporangiophore and spore formation on the cotyledons compared to infection on wild-type plants (Fig. 7c, d). However, treatment with GST (1–5 μg mL−1) alone did not inhibit sporangiophore and spore formation on the cotyledons infected with Hp. arabidopsidis. Collectively, these results indicate that AvrBsT protein possess inhibitory activity against the growth and sporulation of Hp. arabidopsidis and also that the doses of over 10 μg mL−1 AvrBsT exert a toxic effect on Arabidopsis seedlings.
Inhibitory effect of GST-tagged AvrBsT protein on the growth of Hyaloperonospora arabidopsidis isolate Noco2 on Arabidopsis seedlings. a Cell death phenotypes induced by purified GST-tagged AvrBsT protein. Photographs were taken 1 day after treatment with purified GST-tagged AvrBsT protein. b Disease symptoms and micrographs of infected cotyledons treated with purified GST-tagged AvrBsT protein. Photographs were taken 5 days after treatment. The empty vector (GST only) was used as a negative control. c Quantification of sporangiophores produced on 50 cotyledons 6 days after inoculation (5 × 104 conidiosporangia mL−1). Bars 0.2 mm. The empty vector (GST only) was used as a negative control. The data represent the mean ± standard deviations. Different letters (a, b, and c) indicate significant differences from three independent experiments, as analyzed by the LSD test (P < 0.05)
Enhanced susceptibility of DEX:avrBsT-OX Arabidopsis plants to Pseudomonas syringae pv. tomato infection
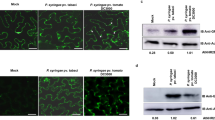
To determine whether avrBsT mediates the basal response to bacterial infection, leaves of DEX:avrBsT-OX Arabidopsis plants were inoculated with Pst DC3000 and Pst DC3000 (avrRpm1) by syringe infiltration. A lower titer of Pst (105 cfu mL−1) was inoculated on the leaves, because avrBsT overexpression is not so effective to inhibit a rapid Pst multiplication in leaves infected with a higher lower titer of Pst (107 cfu mL−1). Unexpectedly, Pst DC3000 and Pst DC3000 (avrRpm1) infection resulted in higher bacterial growth in the leaf tissues of the DEX-treated DEX:avrBsT-OX plants (lines #3, #4 and #5) compared to DEX-treated wild-type plants (Fig. 8a). Next, we investigated whether the overexpression of avrBsT induces electrolyte leakage from the Pst-infected leaves of DEX-treated DEX:avrBsT-OX plants. Both Pst DC3000 and Pst DC3000 (avrRpm1) infection significantly induced electrolyte leakage from the leaf tissues of the DEX:avrBsT-OX plants compared to wild-type leaves (Fig. 8b). However, no significant differences in H2O2 production were observed among wild-type and DEX:avrBsT-OX Arabidopsis plants, regardless of treatment with dexamethasone (DEX), during Pst infection (Fig. 8c). Taken together, the enhanced electrolyte leakages from DEX:avrBsT-OX Arabidopsis leaves infected with avirulent Pst DC3000 (avrRpm1) support the disease-associated cell death response leading to bacterial growth in the transgenic leaves.
Enhanced susceptibility of DEX:avrBsT-OX Arabidopsis plants to Pseudomonas syringae pv. tomato (Pst) infection. Leaves of 5-week-old plants mock-treated or DEX-treated with dexamethasone (DEX) were infiltrated with bacterial suspensions of Pst DC3000 or DC3000 (avrRpm1). a Bacterial growth in leaves inoculated with Pst (105 cfu mL−1). The data represent the mean ± standard deviations. dai days after inoculation. b Electrolyte leakage assay of leaves inoculated with Pst (108 cfu mL−1). The data represent the mean ± standard deviations from three independent experiments. h hours after infiltration. c Quantification of H2O2 produced following Pst inoculation (108 cfu mL−1). The data represent the mean ± standard deviations. Different letters (a, b, and c) indicate significant differences from three independent experiments, as analyzed by the LSD test (P < 0.05). h hours after infiltration
Discussion
Xanthomonas is a large genus of Gram-negative, yellow-pigmented bacteria in plants (Jones et al. 2004). To date, the complete genome sequences of 11 Xanthomonas strains have been determined (Ryan et al. 2011). Four YopJ-like proteins (AvrRxv, AvrBsT, AvrXv4, and XopJ) were identified in X. campestris pv. vesicatoria strains. The Xcv type III effector AvrBsT is the first member of the YopJ family known to suppress effector-triggered plant immunity (Büttner and Bonas 2010), and AvrBsT is necessary for the hypersensitive response (HR) induction in resistant plants (Büttner and Bonas 2010). When Xcv interacts with plants, AvrBsT is translocated into plant cells during infection, triggering defense responses in many plant species (Kirik and Mudgett 2009). In previous studies, we showed that AvrBsT induces cell death in pepper but suppresses defense responses in tomato (Kim et al. 2010).
In this study, AvrBsT was ectopically expressed in the heterologous Arabidopsis system and the dexamethasone (DEX)::avrBsT overexpression (OX) plants were used to investigate the molecular function of AvrBsT in disease and defense responses against microbial pathogens. Induction of avrBsT expression by DEX treatment led to the cell death response, as well as altered defense responses against infection by the obligate biotrophic oomycete Hyaloperonospora arabidopsidis (Hpa) and the hemibiotrophic bacterial pathogen Pseudomonas syringae pv. tomato (Pst). The avrBsT-overexpressing (OX) Arabidopsis plants were less susceptible to Hpa Noco2, but not to Pst DC3000 and DC3000 (avrRpm1). The resistance response of DEX-treated DEX:avrBsT-OX Arabidopsis plants to Hp. arabidopsidis infection was accompanied by a hypersensitive response (HR) and H2O2 accumulation, as determined by trypan blue and DAB staining, respectively. The cell death phenotype in DEX:avrBsT-OX leaves triggered by exogenous dexamethasone application may be a form of basal defense response sufficient to suppress Hp. arabidopsidis infection. These results also suggest that avrBsT overexpression during Hp. arabidopsidis infection enhances the threshold for the activation of defense responses in DEX:avrBsT-OX Arabidopsis plants. The hypersensitive cell death response appears to be an active process to prevent the pathogen from colonizing surrounding tissue. Interestingly, avrBsT overexpression distinctly induced the disease-associated cell death response as well as higher Pst growth in DEX-treated DEX:avrBsT-OX leaves. This finding suggests that the Xcv effector AvrBsT may induce the proliferation of other bacterial pathogens such as Pst, leading to susceptible cell death in plant tissues. In support of these experimental results, overexpression of other bacterial effector proteins such as HopF2 pto and AvrB in Arabidopsis plants has been demonstrated to cause perturbations in host defense responses (Shang et al. 2006; Cui et al. 2010; Wilton et al. 2010). Arabidopsis plants overexpressing HopF2 pto exhibited compromised AvrRpt2-mediated HR (Wilton et al. 2010). Overexpression of AvrB in Arabidopsis plants promoted non-pathogenic P. syringae growth by disrupting the nominal functions of key defense components such as RAR1, HSP90, MPK4 and RIN4 (Shang et al. 2006; Cui et al. 2010). Using 2D-electrophoresis, we were able to identify TCP-1/cpn60 chaperonin family protein and SEC7-like guanine nucleotide exchange protein as proteins upregulated by avrBsT overexpression in Arabidopsis. Both TCP-1/cpn60 chaperonin family protein and SEC7-like guanine nucleotide exchange protein have been implicated in cell fate determination (Kirchhoff et al. 2002; Pagnussat et al. 2005). In contrast, calmodulin-like protein was downregulated by avrBsT overexpression. Calmodulins are generally known to positively modulate stress signaling (Luan 2009; Cheong et al. 2010). The results of this study suggest that these metabolism and protein folding-associated proteins negatively and positively regulate the defense responses against Pst and Hp. arabidopsidis, respectively.
Purified GST-tagged AvrBsT protein was used to determine whether the AvrBsT protein directly inhibits Hp. arabidopsidis infection in Arabidopsis plants. Treatment with 10 μg mL−1 GST-tagged AvrBsT protein caused the induction of a typical cell death phenotype on Arabidopsis seedlings. This indicates that the AvrBsT protein itself causes cell death in Arabidopsis seedlings. Similarly, transient expression of AvrBsT induces hypersensitive cell death in the leaves of Nicotiana benthamiana and pepper (Kim et al. 2010). Interestingly, cotyledons treated with 1 and 5 μg mL−1 of GST-tagged AvrBsT protein showed significantly reduced Hp. arabidopsidis infection. These results suggest that AvrBsT protein is effective in suppressing the growth and sporulation of Hp. arabidopsidis. Although many bacterial effector proteins have been extensively studied, the research has focused on determining the effector function during plant disease and defense responses (Block et al. 2008). To our knowledge, this study provides the first evidence showing that AvrBsT possess anti-oomycete activity against Hp. arabidopsidis.
Microbial effectors have been proposed to subvert plant immunity in host–pathogen interactions (Göhre and Robatzek 2008; Büttner and Bonas 2010). There is convincing evidence that AvrBsT is a versatile protein involved in the manipulation of plant cell processes such as basal defense for the benefit of the Xanthomonas pathogen (Büttner and Bonas 2010). In tomato plants, AvrBsT suppresses basal defense by functioning as a virulence factor that disrupts early defense signaling (Orth et al. 2000). Our experiments using DEX:avrBsT-OX Arabidopsis plants revealed that avrBsT overexpression in Arabidopsis enhances susceptibility to Pseudomonas syringae pv. tomato (Pst) DC3000 and DC3000 (avrRpm1). DEX-treated DEX:avrBsT-OX Arabidopsis plants exhibited increased Pst growth, as well as a susceptible cell death phenotype. Based on these results, heterologous expression of AvrBsT in Arabidopsis plants enables AvrBsT to function as a virulence factor to promote Pst infection. AvrBsT-mediated disease symptoms on transgenic Arabidopsis leaves may result from the susceptible cell death that occurs commonly in compatible interactions. In compatible plant–microbe interactions, susceptible cell death occurs relatively late in the course of infection (Greenberg 1997; Greenberg and Yao 2004). In pepper plants, the signals triggered by CaHIR1 (Capsicum annuum hypersensitive induced reaction 1) are proposed to lead to susceptible cell death during compatible interactions with Xcv (Choi et al. 2011). Recently, AvrBsT was demonstrated to suppress the HR that is triggered by the effector protein AvrBs1 in resistant pepper plants (Szczesny et al. 2010). However, how AvrBsT suppresses plant immunity to enable the successful colonization of bacterial pathogens in plants is largely unknown.
Th Xcv effector AvrBsT is likely required for cell death, disease and defense responses in Arabidopsis plants. However, the precise molecular mechanisms underlying AvrBsT function remain to be investigated (Stall et al. 2009; Kim et al. 2010). In this study, experimental analyses of AvrBsT function have provided a unique view of Arabidopsis–pathogen interactions. Here, we demonstrate that ectopic expression of avrBsT in Arabidopsis plants not only enhances resistance to the obligate biotrophic oomycete Hp. arabidopsidis, but also significantly suppresses plant immunity to the hemibiotrophic bacterial pathogens Pst DC3000 and DC3000 (avrRpm1). When ectopically expressed in plant cells, the Xcv effector AvrBsT may trigger plant cell death, disease and defense signaling, ultimately leading to disease and defense responses against microbial pathogens of various lifestyles.
Abbreviations
- DAB:
-
3′-diaminobenzidine
- DEX:
-
Dexamethasone
- H2O2 :
-
Hydrogen peroxide
- HR:
-
Hypersensitive response
- OX:
-
Overexpression
- PR:
-
Pathogenesis-related
- Pst :
-
Pseudomonas syringae pv. tomato
- ROS:
-
Reactive oxygen species
- Xcv :
-
Xanthomonas campestris pv. vesicatoria
References
Axtell MJ, Staskawicz BJ (2003) Initiation of RPS2-specified disease resistance in Arabidopsis is coupled to the AvrRpt2-directed elimination of RIN4. Cell 112:369–377
Bartetzko V, Sonnewald S, Vogel F, Hartner K, Stadler R, Hammes UZ, Börnke F (2009) The Xanthomonas campestris pv. vesicatoria type III effector protein XopJ inhibits protein secretion: evidence for interference with cell wall-associated defense responses. Mol Plant–Microbe Interact 22:655–664
Bindschedler LV, Manibayeva F, Gardber SL, Gerrish C, Davies DR, Bolwell GP (2001) Early signaling events in the apoplastic oxidative burst in suspension cultured French bean cells involved cAMP and Ca2+. New Phytol 151:185–194
Block A, Li G, Fu ZQ, Alfano JR (2008) Phytopathogen type III effector weaponry and their plant targets. Curr Opin Plant Biol 11:396–403
Brodersen P, Petersen M, Pike HM, Olszak B, Skov S, Ødum N, Jørgensen LB, Brown RE, Mundy J (2002) Knockout of Arabidopsis ACCELERATED-CELL-DEATH11 encoding a sphingosine transfer protein cause activation of programmed cell death and defense. Genes Dev 16:490–502
Büttner D, Bonas U (2010) Regulation and secretion of Xanthomonas virulence factors. FEMS 34:107–133
Büttner D, He SY (2009) Type III protein secretion in plant pathogenic bacteria. Plant Physiol 150:1656–1664
Cheong YH, Sung SJ, Cho JS, Luan S (2010) Constitutive overexpression of the calcium sensor CBL5 confers osmotic or drought stress tolerance in Arabidopsis. Mol Cells 29:159–165
Choi DS, Hwang BK (2011) Proteomics and functional analyses of pepper abscisic acid-responsive 1 (ABR1), which is involved in cell death and defense signaling. Plant Cell 23:823–842
Choi HW, Kim YJ, Hwang BK (2011) The hypersensitive induced reaction and leucine-rich repeat proteins regulate plant cell death associated with disease and plant immunity. Mol Plant–Microbe Interact 24:68–78
Choi DS, Hwang IS, Hwang BK (2012) Requirement of the cytosolic interaction between PATHOGENESIS-RELATED PROTEIN10 and LEUCINE-RICH REPEAT PROTEIN1 for cell death and defense signaling in pepper. Plant Cell 24:1675–1690
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Cui H, Wang Y, Xue L, Chu J, Yan C, Fu J, Chen M, Innes RW, Zhou JM (2010) Pseudomonas syringae effector protein AvrB perturbs Arabidopsis hormone signaling by activating MAP kinase 4. Cell Host Microbe 7:164–175
Cunnac S, Wilson A, Nuwer J, Kirik A, Baranage G, Mudgett MB (2007) A conserved carboxylesterase is a SUPPRESSOR OF AVRBST-ELICITED RESISTANCE in Arabidopsis. Plant Cell 19:688–705
DebRoy S, Thilmony R, Kwack YB, Nomura K, He SY (2004) A family of conserved bacterial effectors inhibits salicylic acid-mediated basal immunity and promotes disease necrosis in plants. Proc Natl Acad Sci USA 101:9927–9932
Eitas TK, Nimchuk ZL, Dangl JL (2008) Arabidopsis TAO1 is a TIR-NB-LRR protein that contributes to disease resistance induced by the Pseudomonas syringae effector AvrB. Proc Natl Acad Sci USA 105:6475–6480
Flors V, Ton J, Jakab G, Mauch-Mani B (2005) Abscisic acid and callose: team players in defence against pathogens? J Phytopathol 153:377–383
Göhre V, Robatzek S (2008) Breaking the barriers: microbial effector molecules subvert plant immunity. Annu Rev Phytopathol 46:180–215
Greenberg JT (1997) Programmed cell death in plant–pathogen interactions. Annu Rev Plant Physiol Plant Mol Biol 48:525–545
Greenberg JT, Yao N (2004) The role and regulation of programmed cell death in plant-pathogen interactions. Cell Microbiol 6:201–211
Hwang IS, Hwang BK (2010) The pepper 9-lipoxygenase gene CaLOX1 functions in defense and cell death responses to microbial pathogens. Plant Physiol 152:948–967
Hwang IS, An SH, Hwang BK (2011) Pepper asparagine synthetase 1 (CaAS1) is required for plant nitrogen assimilation and defense responses to microbial pathogens. Plant J 67:749–762
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Jones JB, Stall RE, Bouzar H (1998) Diversity among Xanthomonads pathogenic on pepper and tomato. Annu Rev Phytopathol 36:41–58
Jones JB, Lacy GH, Bouzar H, Stall RE, Schaad NW (2004) Reclassification of the Xanthomonads associated with bacterial spot disease of tomato and pepper. Syst Appl Micobiol 27:755–762
Kim J-G, Li X, Roden JA, Taylor KW, Aakre CD, Su B, Lalonde S, Kiril A, Chen Y, Baranage G, McLane H, Martin GB, Mudgett MB (2009) Xanthomonas T3S effector XopN suppresses PAMP-triggered immunity and interacts with a tomato atypical receptor-like kinase and TFT1. Plant Cell 21:1305–1323
Kim NK, Choi HW, Hwang BK (2010) Xanthomonas campestris pv. vesicatoria effector AvrBsT induces cell death in pepper, but suppresses defense responses in tomato. Mol Plant–Microbe Interact 23:1069–1082
Kirchhoff SR, Gupta S, Knowlton AA (2002) Cytosolic heat shock protein 60, apoptosis, and myocardial injury. Circulation 105:2899–2904
Kirik A, Mudgett MB (2009) SOBER1 phospholipase activity suppresses phosphatidic acid accumulation and plant immunity in response to bacterial effector AvrBsT. Proc Natl Acad Sci USA 106:20532–20537
Lewis JD, Lee A, Ma W, Zhou H, Guttman DS, Desveaux D (2011) The YopJ superfamily in plant-associated bacteria. Mol Plant Pathol 12:928–937
Luan S (2009) The CBL-CIPK network in plant calcium signaling. Trends Plant Sci 14:37–42
McDowell JM, Cuzick A, Can C, Beynon J, Dangl JL, Holub EB (2000) Downy mildew (Peronospora parasitica) resistance genes in Arabidopsis vary in functional requirements for NDR1, EDS1, NPR1 and salicylic acid accumulation. Plant J 22:23–529
Nimchuk Z, Marois E, Kjemtrup S, Leister RT, Katagiri F, Dangl JL (2000) Eukaryotic fatty acylation drives plasma membrane targeting and enhances function of several type III effector proteins from Pseudomonas syringae. Cell 101:353–363
Orth K, Xu Z, Mudgett MB, Bao ZQ, Palmer LE, Bliska JB, Mangel WF, Staskawicz B, Dixon JE (2000) Disruption of signaling by Yersinia effector YopJ, a ubiquitin-like protein protease. Science 290:1594–1597
Pagnussat GC, Yu H-J, Ngo QA, Rajani S, Mayalagu S, Johnson CS, Capron A, Xie L-F, Ye D, Sundaresan V (2005) Genetic and molecular identification of genes required for female gametophyte development and function in Arabidopsis. Development 132:603–614
Reymond P, Farmer EE (1998) Jasmonate and salicylate as global signals for defense gene expression. Curr Opin Plant Biol 1:404–411
Roden J, Belt B, Ross J, Tachibana T, Bargas J, Mudgett MB (2004) Genetic screen to isolate type III effectors translocated into plant cells during Xanthomonas campestris pv. vesicatoria infection. Proc Natl Acad Sci USA 101:16624–16629
Ryan RP, Vorhölter FJ, Potnis N, Jones JB, Van Sluys MA, Bogdanove AJ, Dow JM (2011) Pathogenomics of Xanthomonas:understanding bacterium–plant interactions. Nat Rev Microbiol 9:344–355
Shang Y, Li X, Cui H, He P, Thilmony R, Chintamanani S, Zwiesler-Vollick J, Gopalan S, Tang X, Zhou JM (2006) RAR1, a central player in plant immunity, is targeted by Pseudomonas syringae effector AvrB. Proc Natl Acad Sci USA 103:19200–19205
Stall RE, Jones JB, Minsavage GV (2009) Durability of resistance in tomato and pepper to Xanthomonads causing bacterial spot. Annu Rev Phytopathol 47:265–284
Stulemeijer IJ, Joosten MH (2008) Post-translational modification of host proteins in pathogen-triggered defence signaling in plant. Mol Plant Pathol 9:545–560
Szczesny R, Buttner D, Escolar L, Schulze S, Seiferth A, Bonas U (2010) Suppression of the AvrBs1-specific hypersensitive response by the YopJ effector homolog AvrBsT from Xanthomonas depends on a SNF1-related kinase. New Phytol 187:1058–1074
Tsan M-F, Gao B (2009) Heat shock proteins and immune system. J Leukoc Biol 85:905–910
Wilton M, Subramaniam R, Elmore J, Felsensteiner C, Coaker G, Desveaux D (2010) The type III effector HopF2Pto targets Arabidopsis RIN4 protein to promote Pseudomonas syringae virulence. Proc Natl Acad Sci USA 107:2349–2354
Acknowledgments
This work was supported by the Next Generation BioGreen21 Program (Plant Molecular Breeding Center; Grant No. PJ008027), Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
I. S. Hwang and N. H. Kim contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2012_1672_MOESM1_ESM.pdf
Supplementary Fig. 1. Growth phenotypes of wild-type (Col-0) and DEX:avrBsT-OX transgenic Arabidopsis plants (lines #3, #4 and #5) at 1-, 2- and 5-week-old plant developmental stages. All the plants non-treated with dexamethasone (DEX) did not show any cell death phenotype on the leaves (PDF 60 kb)
Rights and permissions
About this article
Cite this article
Hwang, I.S., Kim, N.H., Choi, D.S. et al. Overexpression of Xanthomonas campestris pv. vesicatoria effector AvrBsT in Arabidopsis triggers plant cell death, disease and defense responses. Planta 236, 1191–1204 (2012). https://doi.org/10.1007/s00425-012-1672-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-012-1672-4