Abstract
FLOWERING LOCUS C (FLC) is a central floral repressor for the determination of flowering time in Arabidopsis. FLC expression is reactivated upon fertilization and regulated during seed development to ensure the appropriate floral behavior; however, the molecular mechanism for this process is largely unknown. Here, we report the identification of crucial regulators for FLC reactivation during embryogenesis by analyzing FLC::GUS and endogenous FLC expression. We newly define that the full reactivation of FLC requires a FRIGIDA (FRI)-containing protein complex throughout embryogenesis. Mutations in EARLY FLOWERING 7 (ELF7) and VERNALIZATION INDEPENDENCE4 (VIP4) showed severe defects in the reactivation of FLC transcription, suggesting that both of the genes, Arabidopsis homologs of the members of the yeast RNA polymerase II-associated factor 1 (Paf1) complex, are indispensable for FLC reactivation. actin-related protein 6 (arp6), arabidopsis trithorax 1 (atx1), arabidopsis trithorax-related 7 (atxr7), and atx1 atxr7 double mutants also caused the downregulation of FLC during seed development, but the defects were less severe than those in mutants for the FRI- and Paf1-complexes. These results suggest that the ARP6-containing Swr1-complex and FLC-specific histone methyltransferases, ATX1 and ATXR7, have relatively partial roles in FLC reactivation. In contrast to the roles of the histone modifiers, factors in the DNA methylation pathway and biogenesis of small RNAs are not involved in FLC regulation during reproduction. Taken together, our results demonstrate that adjustment by select FLC activators is critical for the re-establishment of an FLC expression state after fertilization to ensure competence for optimal flowering in the next generation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For plants, the timing of the initiation of flowering has been critical during evolution to maximize reproductive success (Amasino 2010). FLOWERING LOCUS C (FLC), which encodes a MADS box-containing transcription factor, is a central floral repressor in Arabidopsis (Michaels and Amasino 1999). High levels of FLC expression cause late flowering even under inductive long-day conditions, whereas transcriptional repression of FLC promotes the initiation of flowering (Ausin et al. 2004; Lim et al. 2004; Sheldon et al. 2000). FLC and its upstream activator, FRIGIDA (FRI), are the major determinants of the natural variation of flowering time in Arabidopsis: winter annual Arabidopsis plants contain dominant alleles of FLC and FRI. FRI is a plant-specific protein that is required for high levels of FLC expression (Johanson et al. 2000; Michaels and Amasino 1999, 2001). In contrast, rapid-cycling summer annuals contain mutations in either FRI or FLC, and, therefore, FLC expression remains very low (Kim et al. 2009; Shindo et al. 2005; Werner et al. 2005). FRI is a member of a small gene family, and two other members are FRI-LIKE 1 and 2 (FRL1 and FRL2), which are required for high levels of FLC expression. SUPPRESSOR OF FRIGIDA 4 (SUF4), a C2H2-type zinc-finger protein, has been identified as an FRI-interacting protein (Kim et al. 2006). SUF4 binds directly to the FLC promoter and may recruit a protein complex containing FRI to activate FLC expression.
Genetic studies have identified various factors required for a high level of FLC expression (Fig. 1). Most of them, if not all, are components of chromatin-modifying complexes that promote active chromatin (Amasino 2010; Amasino and Michaels 2010). Mutations in genes that encode homologs of the members of yeast RNA polymerase II-associated factor 1 (Paf1) complex have resulted in a failure of elevated FLC expression, even in the FRI-containing winter annuals. EARLY FLOWERING 7 (ELF7, also known as VERNALIZATION INDEPENDENCE 2 [VIP2]), ELF8 (VIP6), VIP4, and VIP5 are members of the Paf1-complex, which has been shown to be required for a high level of FLC expression (He et al. 2004; Kim et al. 2005; Oh et al. 2004) (Fig. 1). In yeast, the Paf1-complex has been found to interact with SET1 and SET2 histone methyltransferases (Belotserkovskaya and Reinberg 2004), and the recruitment of SET1 and SET2 to chromatin has resulted in an increase of methylation in histone H3 lysine 4 and lysine 36 (H3K4 and H3K36), respectively. In Arabidopsis, ARABIDOPSIS TRITHORAX (ATX) 1 through 5, and ARABIDOPSIS TRITHORAX-RELATED (ATXR) 1 through 4, and 7 have a SET domain that is homologous to yeast SET1 and Drosophila Trithorax (Baumbusch et al. 2001). Mutations in ATX1 have been shown to cause decreased FLC expression with reduced tri-methylation of H3K4 (H3K4me3) at the 5′-end of FLC (Pien et al. 2008). In addition, it has been reported that the atx1 atxr7 double mutant showed additive phenotypes in contrast to those of each of the single mutants, and both ATX1 and ATXR7 histone methyltransferase were required for the enrichment of H3K4me3 in FLC chromatin (Amasino 2010) (Fig. 1). EARLY FLOWERING IN SHORT DAYS (EFS, also known as SET DOMAIN GROUP 8 [SDG8]) belongs to the SET2 class that is responsible for H3K36me2 in the promoter region and the first intron of FLC (Xu and Shen 2008; Zhao et al. 2005).
Schematic diagram for the regulatory pathways of FLC activation. a Four major genetic pathways are known to be involved in FLC activation. FRI-complex binds to the FLC promoter and mediates transcriptional activation. FRI, FRL1, SUF4, FES1, and FLX have been isolated as components of FRI-complex. Loss-of-function mutations of FRI and FRL1 cause significant decrease of FLC transcription in vegetative tissues. FRI-complex is also known to recruit Swr1-complex to FLC region. Swr1-complex is involved in incorporation of histone variant H2A.Z in various organisms and loss-of-biochemical function of Swr1-complex results in the partial reduction of FLC expression in vegetative tissues. PIE1, ARP6, AtSWC2, and AtSWC6 have been isolated as components of Swr1-complex in Arabidopsis. EFS, ELF7, VIP3, VIP4, VIP5, and VIP6 comprise the Paf1-complex in Arabidopsis. Paf1-complex associates with transcriptional machineries and maintains the elongation of transcriptional process by deposition of H3K4 methylation. Loss-of-Paf1-complex function induces the significant reduction of FLC expression in vegetative tissues. ATX1 and ATXR7 have been isolated as Arabidopsis homolog of TRITHORAX family that mediates H3K4 methylation. Both ATX1 and ATXR7 function redundantly and atx1 atxr7 double mutant shows partial reduction of FLC in vegetative tissues. Protein components examined in this study are marked with white circles or white rounded boxes within each protein complex. b Simple diagram of Arabidopsis reproduction. Arabidopsis produces female gametophyte within the ovule and male gametophyte, pollen, for reproduction. After double fertilization, embryo and endosperm are developed. Based on morphology of embryo, seed development was divided into globular, heart, torpedo, and mature stages in this study. Most of the FLC regulatory mechanisms have been studied in seedling stage after germination. By contrast, our study has focused on the roles of various FLC activators during reproduction
In Arabidopsis, PHOTOPERIOD-INDEPENDENT EARLY FLOWERING 1 (PIE1) (Noh and Amasino 2003), ACTIN-RELATED PROTEIN 6/ SUPPRESSOR OF FRIGIDA 3/EARLY IN SHORT DAYS 1 (ARP6/ SUF3/ESD1) (Choi et al. 2005; Deal et al. 2005; Martin-Trillo et al. 2006), and AtSWC6/SERRATED LEAVES AND EARLY FLOWERING (SEF) (Choi et al. 2007; March-Diaz et al. 2007) have been isolated as homologs of members of the yeast Swr1 complex (Fig. 1). These Arabidopsis members are involved in the exchange of histone H2A to H2A.Z, and it has been reported these are required for full FLC expression (Choi et al. 2007).
Vernalization, a prolonged exposure to cold, establishes the repressive epigenetic histone marker H3K27me3 at FLC chromatin. This silencing of FLC expression is stably maintained mitotically, even under warm conditions (Gendall et al. 2001; Sung and Amasino 2004), but this “memory of winter” is not transmitted to the next generation and has to be reset during reproduction for the re-establishment of vernalization. Many studies have focused on identifying the FLC regulators and their regulatory mechanisms during the vegetative stage, but FLC resetting and its mechanisms during the reproductive stage remain largely unknown.
Recently, Sheldon et al. and our group have reported that, regardless of the transcriptional activity in the adult plants, FLC::GUS expression was not detected in male or female gametophytes, but the reactivation of both parental alleles occurred after fertilization in the embryo, not in the endosperm (Choi et al. 2009; Sheldon et al. 2008). We also have examined the possibility that these FLC regulators are involved in the FLC resetting mechanism.
Here, we describe the defects in the reactivation of FLC in various mutants for FLC regulators that are required for elevating FLC expression during the vegetative stage. Our results suggest that some specific activators of FLC, rather than canonical mechanisms, play important roles in the reactivation of FLC during embryogenesis.
Materials and methods
Plant material and growth conditions
Most of the plants used in our experiment have an active FRI allele from SF-2 (Lee et al. 1994) by a genetic cross. In this article, we called Col-0 having an active FRI allele the wild type. fri refers to the inactive allele of FRI from Col-0. All of the plants used in this study are in the Col-0 background, except for met1-6 in the Col-gl, and, cmt3-7 and ago4-1 in Ler, and drm1 drm2 in the Ws background. Plants were grown under previously described conditions (Choi et al. 2009).
Histochemical GUS imaging
The FLC::GUS reference line (Michaels et al. 2005) used in this paper was a gift from S. Michaels and various mutant plants containing FLC::GUS transgene were generated by genetic crosses between mutant plants and FLC::GUS plant. Methods for GUS staining, fixing tissue, and microscopy are as described previously (Choi et al. 2009).
Analysis of gene expression
Total RNA extraction from developing seeds was performed using seeds independently harvested according to the stage of embryo development. The Qiagen RNeasy Plant Mini Kit (Qiagen) was used for RNA extraction from seeds. Genomic DNA was eliminated on-column with a Qiagen RNase-free DNase set. A total of 0.5–1 μg of RNA was reverse transcribed using M-MLV reverse transcriptase (Ambion) and an oligo-dT primer. PCR amplification was performed using gene-specific primers: TUB2, 5′-ATCGATTCCGTTCTCGATGT-3′ and 5′-ATCCAGTTCCTCCTCCCAAC-3′; FLC, 5′-GAAGAGAACCAGGTTTTGGCTA-3′ and 5′-TTTGTCCAGCAGGTGACATC-3′.
Real-time qPCR
Real-time qPCR was performed as described previously (Choi et al. 2009). Reactions for all of the templates were performed in duplicate or triplicate. The threshold cycle (Ct) was determined using the CFX Manager Software (Bio-Rad). Gene-specific transcripts were quantified using the ddCt method (ddCt = Ctgene of interest – CtTUB2) (Livak and Schmittgen 2001). Real-time SYBR-green dissociation curves showed one species of amplicon for each primer combination.
Results
Effect of the FRI family on high levels of FLC reactivation during embryogenesis
FLC expression is known to require not only FRI but also other FRI family members to be competent for vernalization (Michaels et al. 2004; Schlappi 2006) (Fig. 1). To explore the function of the FRI family in FLC reactivation, we compared the expression pattern of FLC::GUS in fri and in frl1 mutants to the expression in the wild type during the reproductive stage. In the FLC::GUS construct (Michaels et al. 2005), GUS was inserted into the sixth exon of a 16-kb genomic clone containing 5.4-kb upstream of the FLC start site and 5-kb downstream of the stop codon. We introduced the FLC::GUS transgene into fri or frl1 mutants by genetic crosses and analyzed the GUS expression pattern in the self-fertilized F2 progenies.
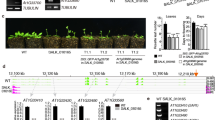
In the wild type, FLC::GUS activity was not detected in the female or male gametophyte, except in the sporophytic tissues of the ovule (Fig. 2a, b), which are of diploid maternal origin (Choi et al. 2009). After fertilization, FLC::GUS was expressed only in embryos, and a high level of GUS expression was maintained throughout embryogenesis (Fig. 2c–f). In fri or frl1 mutant plants, FLC::GUS was not expressed in either of the gametophytes, as in wild type (Fig. 2g, h, m, n), suggesting that FRI and FRL1 do not function in the regulation of FLC expression during gametogenesis. The sporophytic expression of GUS in the tissues surrounding the embryo sac disappeared (Fig. 2g, m), indicating a decrease of FLC expression in the vegetative tissues of the mutants. After fertilization, GUS was detected in the young embryos of both fri and frl1 mutants (Fig. 2i, o). As the embryo matured, however, GUS activity gradually decreased from the cotyledons of the embryo (Fig. 2j, k, p, q), and, eventually, GUS expression was largely restricted to the vascular regions of both mutants (Fig. 2l, r).
FLC::GUS and FLC mRNA expression in fri family mutants. FLC::GUS expression in the wild type ovule (a), stamen (b), developing embryos at the globular stage (c), heart stage (d), torpedo to walking stick stage (e) and mature stage (f). FLC::GUS expression in the fri mutant ovule (g), stamen (h), developing embryos at the globular stage (i), heart stage (j), torpedo to walking stick stage (k) and mature stage (l). FLC::GUS expression in the frl1 mutant ovule (m), stamen (n), developing embryos at the globular stage (o), heart stage (p), torpedo to walking stick stage (q) and mature stage (r). s Real time qRT-PCR analysis of FLC mRNA expression in the developing seeds of wild type, fri and frl1 mutants. t Comparison of FLC mRNA expression by qRT-PCR in the fri mutant with vernalization-treated wild type FRI plants in seeds at globular stage. Values in (s) and (t) are normalized to an internal reference TUBULIN-β-CHAIN2 (TUB2) gene and plotted relative to the expression of FLC in the fri globular seeds, which was set to 1.0. Value represents the average of triplicate measurements ± standard error. Scale bars: ovules, 50 μm; stamens, 100 μm; seeds, 100 μm
We also performed real-time quantitative RT-PCR (qRT-PCR) to examine the effect of the FRI family on the endogenous FLC expression. Seeds at the globular stage show higher FLC expression than the heart stage, but the FLC expression gradually increased as the seed matured (Fig. 2s, blue bar). In fri or frl1 mutants, the overall level of FLC mRNA was significantly reduced in both of the mutants (Fig. 2s, red and green bar, respectively). According to our previous report, the sporophytic GUS expression of the ovules correlated with the vegetative expression of FLC, and it retained a considerable amount of expression in the maternal tissues of globular seeds in the wild type (Choi et al. 2009). In contrast, vegetative FLC expression has been reported to be reduced in fri and frl1 (Michaels et al. 2004), and this likely led to the lack of sporophytic GUS expression in the ovules of both mutants (Fig. 2g, m). Through vernalization, the vegetative FLC expression in a FRI plant has been shown to be reduced to a level similar to that in a fri plant, even in the presence of FRI-complex genes (Michaels et al. 2004). Therefore, we directly compared the level of FLC mRNA in fri plants to that in vernalized FRI plants with seeds at the globular stage. The FLC expression of vernalized FRI plants was approximately eightfold higher than that of fri plants (Fig. 2t). This clearly demonstrates that the reactivation of FLC was impaired in fri-complex mutants from the early stages of seed development. Taken together, our results suggest that the FRI and FRL1 are essential for the full activation of FLC during embryogenesis.
Effects of Swr1- and Paf1-complexes on FLC reactivation during embryogenesis
ARP6 encodes nuclear ACTIN-RELATED PROTEIN 6, a homolog of a component of the yeast ATP-dependent chromatin remodeling Swrl-complex (Choi et al. 2005) (Fig. 1). VIP4 and ELF7 encode plant relatives of yeast LEO1 and PAF1, respectively, which are components of the Paf1-complex (He et al. 2004; Zhang and van Nocker 2002) (Fig. 1); mutations in these genes have been shown to not only induce early flowering but also result in pleiotropic phenotypes (Dennis and Peacock 2007). To investigate the potential roles of ARP6, VIP4, and ELF7 on FLC reactivation, the FLC::GUS transgene was introduced into arp6, vip4, and elf7 mutants by genetic crosses, and the histochemical GUS staining patterns were analyzed in the F2 progenies. Similar to the fri and frl1 mutant plants, FLC::GUS expression was detected in neither ovule sporophytic tissues (Fig. 3e, i, m) nor gametophytes (Fig. 3e, f, i, j, m, n). The lack of GUS signal in the female and male gametophyte indicates that ARP6, VIP4, and ELF7 do not function in FLC expression in gamete cells.
FLC::GUS and FLC mRNA expression in arp6, vip4 and elf7 mutants. FLC::GUS expression in the wild-type ovule (a), stamen (b), developing embryos at the globular stage (c), and mature stage (d). FLC::GUS expression in arp6 mutant ovule (e), stamen (f), developing embryos at the globular to heart stage (g), and mature stage (h). FLC::GUS expression in the vip4 mutant ovule (i), stamen (j), developing embryos at the globular to heart stage (k), and mature stage (l). FLC::GUS expression in the elf7 mutant ovule (m), stamen (n), developing embryos at the globular to heart stage (o), and mature stage (p). The arrows in (l) and (p) show weak expression in the shoot apical meristem region of vip4 and elf7 mutant embryos, respectively, and the insets show enlarged images of FLC::GUS expression. q Real-time qRT-PCR analysis of FLC mRNA expression in the developing seeds of wild type and the fri, vip4 and arp6 mutants. Values are normalized to an internal reference TUB2 gene and plotted relative to the expression of FLC in fri globular seeds, which was set to 1.0. Value represents the average of duplicate measurements ± standard error. Scale bars ovules, 50 μm; stamens, 100 μm; seeds, 100 μm
After fertilization, FLC::GUS expression in the arp6 mutant was detected from young embryos, but showed at a reduced level in the mature embryo (Fig. 3g, h) than that in the wild-type mature embryo (Fig 3c, d). We previously observed that FLC::GUS activity was significantly reduced in pie1 mutant (Choi et al. 2009). Both PIE1 and ARP6 are components of the same Swr1-complex (Fig. 1), and the mutations in pie1 and arp6 resulted in the reduction of FLC::GUS expression.
In contrast, FLC::GUS activity was not detected in any of the developmental stages of vip4 and elf7 mutant embryos (Fig. 3k, l, o, p), except for a slight expression in the shoot apical meristem of mature embryos in each mutant (arrows in Fig. 3l, p). This result indicates that VIP4 and ELF7 have critical roles in the reactivation, as well as the maintenance, of FLC expression during embryogenesis and further suggests the important function of histone modification on FLC chromatin by the Arabidopsis Paf1-complex during reproduction. In addition, vip4 and elf7 mutants also showed an occasional ectopic GUS expression in the endosperm or seed coat (Supplementary Fig. S1a, b). The seeds of many vip4 and elf7 mutants consistently exhibited abnormal morphology, with a smaller and rounder shape than wild-type seeds at the same stages (Fig. S1), suggesting that the observed ectopic GUS expression might be due to pleiotropism in these mutants.
We further confirmed the endogenous FLC expression by qRT-PCR in arp6, vip4, and elf7 mutant seeds. In the arp6 mutants, FLC mRNA was expressed at very low levels in globular and heart seeds (Fig 3q, orange bar). From torpedo to mature stage, FLC expression increased but still showed considerably lower than that in the wild type (Fig 3q, orange and blue bar). Therefore, arp6 mutation causes down-regulation of FLC expression throughout embryogenesis as in the pie1 mutant. Mutations in vip4 and elf7 resulted in very low levels of FLC mRNA expression throughout embryogenesis (Fig. 3q, green and purple bar). The expression levels in the mature seeds of both mutants were even lower than in the fri mutant, demonstrating that a lack of the Paf1-complex causes severe defects and failed to induce FLC reactivation in the early stages of embryogenesis, which could not be overcome during the later stages.
Effects of H3K4 methyltransferases in FLC reactivation during embryogenesis
SET domain-containing ATX1 and ATXR7 have been characterized as Arabidopsis H3K4 methyltransferases that play roles in activating FLC (Pien et al. 2008; Tamada et al. 2009) (Fig. 1). To evaluate the potential roles of histone methyltransferases during the reproductive stage, we analyzed GUS expression in atx1 and atxr7 mutants. Each single mutant showed the same GUS expression pattern as FRI FLC::GUS plants (Fig. 4a–l). These results indicate that ATX1 and ATXR7 did not affect FLC::GUS expression during reproduction, but it is still possible that ATX1 and ATXR7 might have functional redundancy during embryogenesis. Therefore, we generated atx1 atxr7 double mutants harboring the FLC::GUS transgene. However, the atx1 atxr7 double mutants also did not show the altered GUS expression pattern (Fig. 4m–p), indicating that H3K4 methylation by ATX1 and ATXR7 did not have an essential role in FLC::GUS transgene reactivation during reproduction. This is in contrast to the previous reports that atx1 and atx1 atxr7 mutations resulted in a significant decrease in H3K4 methylation and a concomitant reduction of FLC expression, causing an early flowering phenotype similar to the fri background (Pien et al. 2008; Tamada et al. 2009). We further investigated FLC::GUS expression in F3 seedlings after harvesting seeds. However, the simultaneous absence of ATX1 and ATXR7 did not alter the GUS-staining pattern (Supplementary Fig. S2).
FLC::GUS and FLC mRNA expression in H3K4 methyltransferase mutants. FLC::GUS expression in the wild-type ovule (a), stamen (b), developing embryos at the globular stage (c), and mature stage (d). FLC::GUS expression in the atx1 mutant ovule (e), stamen (f), developing embryos at the globular stage (g), and mature stage (h). FLC::GUS expression in the atxr7 mutant ovule (i), stamen (j), developing embryos at the globular stage (k), and mature stage (l). FLC::GUS expression in the atx1 atxr7 double mutant ovule (m), stamen (n), developing embryos at the globular stage (o), and mature stage (p). q Real-time qRT-PCR analysis of FLC mRNA expression in the developing seeds of wild type and the fri, atx1, and atxr7 mutants and the atx1 atxr7 double mutant. Values are normalized to an internal reference TUB2 gene and plotted relative to the expression of FLC in fri globular seeds, which was set to 1.0. Value represents the average of duplicate measurements ± standard error. Scale bars ovules, 50 μm; stamens, 100 μm; seeds, 100 μm
We also performed real-time qRT-PCR for endogenous FLC expression. In atx1, FLC mRNA was dramatically decreased at the globular stage (Fig. 4q, green bar) but still higher than that in fri seeds (Fig. 4q, red bar). From the globular to mature stages, atx1 plants showed a gradual increase in FLC mRNA, and atx1 mature seeds showed a high FLC expression, with a level similar to FRI. In contrast, the absence of ATXR7 had weaker effect than the atx1 mutation (Fig. 4q, purple bar).
In globular seeds of atx1 atxr7mutants, FLC expression was highly decreased, similar to that observed in atx1 mutant seeds (Fig. 4q, orange bar). FLC expression in the mature seeds of the double mutant was much lower than that in FRI or the single mutants of atx1 and atxr7, but was still higher than that in fri. This suggests that ATX1 and ATXR7 H3K4 methyltransferases have functional redundancy on FLC chromatin and are required for full reactivation of FLC during seed development, although their impacts are weaker than those of the FRI-complex and Paf1-complex.
Effect of DNA methylation on FLC::GUS reactivation during reproduction
Regardless of the epigenetic state in adult plants, FLC expression is repressed in gametophytes (Choi et al. 2009) (Sheldon et al. 2008). We have previously proposed the possibility that the silencing of FLC in gametophytes might be established by the canonical process of epigenetic reprogramming rather than by specific FLC repressors (Choi et al. 2009). To examine this possibility, we performed FLC::GUS staining in various mutants related to DNA methylation. We chose mutants for the following factors: DNA METHYLTRANSFERASE 1 (MET1), a CG maintenance methyltransferase; CHROMOMETHYALTRANSFERASE 3 (CMT3), a CNG methyltransferase; DOMAINS REARRANGED METHYLTRANSFERASE 1 and 2 (DRM1 DRM2), two de novo methyltransferases; and DECREASED DNA METHYLATION1 (DDM1), a SWI1/SNF2 chromatin remodeling ATPase. It has been reported that global DNA hypomethylation occurs in ddm1 mutant plants (Jeddeloh et al. 1999). We also chose RPD3-like HISTONE DEACETYLASE 6 (HDA6) as a candidate because in the hda6 mutant, FLC expression is up-regulated (Wu et al. 2008), and symmetric DNA methylation is reduced at RNA dependent DNA Methylation (RdDM)-silenced promoters (Aufsatz et al. 2007; Wu et al. 2008). We introduced FLC::GUS and FRI into met1 fri, cmt3 fri, drm1 drm2 fri, ddm1 fri, and hda6 fri, and analyzed the GUS expression pattern in segregating F2 populations.
Except for the met1mutant, all the mutant plants showed GUS expression in the sporophytic tissues of the ovules (Fig. 5a, i, m, q, u). The absence of GUS expression in met1 ovules (Fig. 5e) might be a result of the reduction of FLC expression in plants with hypomethylation, as previously reported (Finnegan et al. 1998; Genger et al. 2003; Jean Finnegan et al. 2005). However, the down-regulation of FLC is not associated with changes in DNA methylation at the FLC locus. Instead, antisense MET1 transgenic plants have shown decreases in H3/H4 acetylation and H3K4 methylation at FLC the promoter (Jean Finnegan et al. 2005). To compare the DNA methylation states in the FLC region in the wild type and the met1 mutant, we performed PCR amplification after McrBC treatment, which specifically digests methylated DNA (Supplementary Fig. S3). In our results, none of the regulatory sequences in the 9-kb region covering the FLC locus were methylated in wild-type rosette leaves. Moreover, the methylation level was unchanged in the met1-6 null mutant, confirming the indirect role of MET1 on FLC expression (Jean Finnegan et al. 2005). None of the mutants showed any GUS expression in the embryo sacs or pollen grains that contain the gametophytes (Fig. 5e, f, i, j, m, n, q, r, u, v), suggesting that canonical DNA methylation did not play a role in the gametogenesis-specific FLC repression. Rather, low CG DNA methylation can repress FLC expression in the sporophyte without alteration in the DNA methylation at the FLC locus.
FLC::GUS expression in mutants of the DNA methylation pathway. FLC::GUS expression in the wild-type ovule (a), stamen (b), developing embryos at the globular to heart stage (c), and mature stage (d). FLC::GUS expression in met1 mutant ovule (e), stamen (f), developing embryos at the globular to hear stage (g), and mature stage (h). FLC::GUS expression in the cmt3 mutant ovule (i), stamen (j), developing embryos at the globular to heart stage (k), and mature stage (l). FLC::GUS expression in the drm1 drm2 mutant ovule (m), stamen (n), developing embryos at the globular to heart stage (o), and mature stage (p). FLC::GUS expression in the ddm1 mutant ovule (q), stamen (r), developing embryos at the globular to heart stage (s), and mature stage (t). FLC::GUS expression in the ddm1 mutant ovule (u), stamen (v), developing embryos at the globular to heart stage (w), and mature stage (x). Scale bars ovules, 50 μm; stamens, 100 μm; seeds, 100 μm
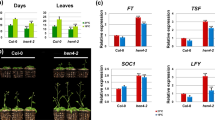
After fertilization, mutations in met1, cmt3, ddm1, and hda6 showed almost the same FLC::GUS pattern as that in FRI (Fig. 5c, d, g, h, k, l, o, p, s, t, w, x), indicating that MET1, CMT3, DDM1, and HDA6 are not relevant for FLC reactivation or full activation. Interestingly, FLC::GUS was not reactivated in the drm1 drm2 double mutant, and the lack of expression was found throughout embryogenesis (Fig. 5o, p). This raised the possibility that FLC cannot be reactivated in drm1 drm2 mutant. However, endogenous FLC level in drm1 drm2 double mutant by qRT-PCR was slightly decreased but still showed considerable amount compared to other activator mutants during entire embryogenesis (Supplementary Fig. S4a).We reasoned that if the drm1 drm2 double mutant prevented FLC expression, the progenies from these mutants would show decreased expression of endogenous FLC and cause early flowering even if they have active FRI and FLC alleles. Consistent with GUS data in seeds, FLC::GUS was not expressed in 10 DAG seedlings (Supplementary Fig. S4b). However, the F3 progeny from the drm1 drm2 mutants did not show an early flowering phenotype (data not shown). We performed semi-quantitative RT-PCR in the seedling of F3 progenies 10 days after germination (DAG) to check the FLC mRNA levels. In the drm1 drm2 mutant, the endogenous mRNA level of FLC was not altered, and only GUS mRNA expression decreased compared to that in the FRI wild type (Supplementary Fig. S4c). Therefore, the lack of FLC::GUS reactivation in drm1 drm2 may be limited to the FLC::GUS transgene.
We also performed GUS staining in ddm1 fri and hda6 fri mutants that contains an inactive fri allele to check the effect of DNA methylation and histone acetylation on FLC::GUS expression in the absence of the FRI-complex. Both of the mutants showed same GUS expression patterns during reproductive stage as that in fri FLC::GUS plants (Supplementary Fig. S5). In addition, the flowering time of the ddm1 fri, drm1 drm2 fri, and cmt3 fri mutants was not accelerated compared to that of fri plants (Supplementary Fig. S6). Taken together, our results suggest that DNA methylation does not play a role in FLC reprogramming during the reproductive stage, regardless of the activity of the FRI-complex.
Effect of the siRNA pathway on FLC expression during reproduction
DICER-LIKE 3 (DCL3), RNA-DEPENDENT RNA POLYMERASE 2 (RDR2), and NUCLEAR RNA POLYMERASE IVa (NRPD1a) produce 24-nt small RNAs complementary to the 3′ end of FLC, and a lack of these small RNAs have caused the up-regulation of FLC mRNA levels (Swiezewski et al. 2007). It is, thus, possible that these 24-nt small RNAs might regulate FLC reprogramming during the reproductive stage. To examine this possibility, we crossed rdr1 fri, dcl3 fri, and ago4 (ARGONAUTE 4) fri mutants with FRI FLC::GUS plants, and the FLG::GUS patterns were analyzed in the F2 population.
None of the siRNA mutants showed gamete-specific FLC::GUS expression (Fig. 6e, f, i, j, m, n), indicating that the siRNA pathway was not relevant to the repression of FLC during gametogenesis. All of the siRNA mutants showed the same GUS expression pattern as the control FLC::GUS plants during embryogenesis (Fig. 6g, h, k, l, o, p). siRNA mutants in an inactive fri background also showed the same GUS expression pattern as the control fri FLC::GUS plants (Supplementary Fig. S7). Our data suggest that the siRNA pathway does not participate in the reprogramming of FLC during the reproductive stage, regardless of the activity of the FRI-complex.
FLC::GUS expression in mutants of the siRNA pathway. FLC::GUS expression in the wild-type ovule (a), stamen (b), developing embryos at the globular to heart stage (c), and torpedo stage (d). FLC::GUS expression in the rdr2 mutant ovule (e), stamen (f), developing embryos at the globular to heart stage (g), and torpedo stage (h). FLC::GUS expression in the dcl3 mutant ovule (i), stamen (j), developing embryos at the globular to heart stage (k), and torpedo stage (l). FLC::GUS expression in the ago4 mutant ovule (m), stamen (n), developing embryos at the globular to heart stage (o), and torpedo stage (p). Scale bars ovules, 50 μm; stamens, 100 μm; seeds, 100 μm
To examine the effect of miRNA on FLC resetting, we generated plants containing the FLC::GUS transgene in the dcl1 mutant with an active FRI allele. We observed the same sporophytic GUS expression in the ovules of dcl1 mutants and no expression in gamete cells, suggesting that the miRNA pathway was not involved in FLC repression during gametogenesis. Unfortunately, the dcl1 mutant plant showed severe defects in floral organs (data not shown), and its integument did not develop (Supplementary Fig. S8), therefore, it was impossible for us to analyze the GUS expression during embryogenesis due to the lack of viable seeds after fertilization.
Discussion
Here, we present evidence that diverse FLC activators play critical roles in the FLC reactivation process during embryogenesis. Mutants of factors in the Paf1-complex and FRI-complex display a significant failure in the full reactivation of FLC from young to mature seed development, but arp6 shows a weaker defect than mutants for the FRI- and Paf1-complexes. ATX1 and ATXR7 have redundant roles in FLC reactivation, but their effects are weaker than mutants for the FRI- and Paf1-complexes. Our results provide clear evidence that the genetic and epigenetic states for appropriate FLC expression in germinated growing plants are already established during embryogenesis by diverse FLC regulators.
Function of FLC activators in embryogenesis
FLC expression requires various complexes for its activation (Fig 1). The biological function of these complexes has been well studied in the vegetative stage. FRI interacts with FRL1, SUF4, FRIGIDA ESSENTIAL1 (FES1), and FLC EXPRESSOR (FLX), and these proteins comprise the FRI-complex and specifically induce FLC expression (Choi et al. 2011). The FRI-complex recruits the chromatin modification factor, the SWR1-complex and a general transcription factor, a TAF14 homolog, for the transcriptional activation of FLC (Choi et al. 2011). Similar with the effects in the vegetative tissue, mutations in the FRI-complex components resulted in a significant reduction of the expression level of FLC during embryogenesis (Fig. 2s), and the reactivated FLC level in fri globular seeds was much lower than that in vernalized FRI seeds (Fig. 2t). This indicated that the FRI-complex components have critical roles in reactivating FLC expression in early embryogenesis, and the defect caused by its deficiency is maintained until seed maturation.
In Arabidopsis, it has been reported that mutants of the Paf1-complex show an early flowering phenotype caused by the reduced expression of FLC and also show low levels of H3K4 methylation on FLC chromatin (He et al. 2004; Oh et al. 2004). This suggests that the Arabidopsis Paf1-complex regulates FLC by not only transcriptional machinery but also histone modification. The vip4 and elf7 mutants almost failed to reactivate FLC throughout embryogenesis (Fig. 3q). Because both the FRI-complex and Paf1-complex are bifunctional in the transcriptional process and the histone modification of FLC chromatin, it is possible that those complexes play the most critical roles in FLC reactivation in our experiments.
In contrast to the FRI- and Paf1-complex, the arp6 mutant showed a partial decrease of FLC expression during seed development (Fig. 3q). ARP6 is a component of the Swr1-complex, which is required for the incorporation of histone variant H2A.Z (Choi et al. 2007; Deal et al. 2007) (Fig. 1). Previously, we have shown that FLC::GUS expression was partially decreased in the pie1 mutant (Choi et al. 2009); thus, it seems that the Swr1-complex has a partial effect in FLC reactivation. Given that the Swr1-complex is recruited by the FRI-complex (Choi et al. 2011), it is reasonable to think that the less severe defect by the arp6 mutation on FLC reactivation was due to a hierarchy of complexes. Interestingly, the atx1 atxr7 double mutant also showed a partial defect in activating FLC during embryogenesis (Fig. 4q). It has been reported that FRI recruited WDR5a to the FLC locus and that WDR5a interacted with ATX1 in vitro (Jiang et al. 2009). Therefore, ATX1 and ATXR7 might be factors recruited by the FRI-complex, similar to the Swr1-complex.
Conclusively, our results provide evidence that the FLC activators that regulate vegetative FLC expression already function during embryogenesis. FLC reactivation is established by the cooperation of diverse regulators upon fertilization and is maintained during reproduction, which is a prerequisite for the flowering competence of the ensuing generation.
Possible mechanism for FLC reprogramming during the reproduction process
In our previous study, FLC expression was found to be repressed in gametophytes regardless of the epigenetic state in adult plants (Choi et al. 2009). We also showed that an autonomous-pathway that represses FLC expression in the absence of FRI is not involved in the repression of FLC in gametophyte.
DNA methylation and siRNA are generally associated with gene silencing. CG, CNG and CNN methylation are abundant in the plant genome and function not only in the transcriptional silencing of transposons and repetitive sequences, but also in gene imprinting (Bender 2004; Chan et al. 2005). DNA methylation is maintained by DNA methyltransferase, such as MET1 and CMT3. In addition, DRM2 mediates de novo DNA methylation. The siRNA pathway recruits DRM2 to its target region and causes RdDM (Feng et al. 2010). Because DNA methylation and siRNA, by themselves or working together, participate in gene and transposon silencing during reproduction, we suspected these processes as candidates for the reprogramming of FLC expression in the gametophyte. Although key regulators of these mechanisms were used in our experiment, none of them showed a de-regulation of FLC both in gametogenesis and in embryogenesis (Figs. 5, 6). Therefore, neither DNA methylation nor siRNA mechanisms function in FLC reprogramming during reproductive process, which supports our conclusion that histone-mediated FLC regulation might be the basal mechanism of FLC reactivation.
In contrast, dynamic chromatin exchange occurs in the primordial germ cells of mice (Hajkova et al. 2008). The erasure of the histone marker and the exchange of histone variants might be involved in this global chromatin dynamic, and this is thought to mediate genomic reprogramming (Hajkova et al. 2008). Similar with this, it has recently been reported that a limited number of H3.3 variants are dominantly present in both female and male germ cells and are actively removed from zygote chromatin in plants (Ingouff et al. 2007, 2010). This raises the possibility of global reprogramming of the epigenetic state by exchange of histone variants during plant reproduction, but this hypothesis requires further studied. Given that FLC is not expressed in gametophytes, regardless of the epigenetic state of the adult plants, but is reactivated after fertilization, it is tempting to speculate that the FLC epigenetic markers of adult plants are erased and reset sometime during gametogenesis by global histone exchange or by a histone demethylation process. However, the erasure of repressive markers seems insufficient to activate FLC without transacting activators. It is intriguing that FLC and many activators of FLC start to be expressed in the embryos after fertilization during the reproductive phase. It will be interesting to see whether FLC can be reactivated during gametogenesis if its activators are expressed in the gamete cells. If so, it will prove the evidence that the epigenetic state of FLC in the gametophyte is receptive to activation once activators are induced.
In this study, we addressed the re-activation of FLC during embryogenesis in mutants for various epigenetic regulators. Our results contribute to the knowledge of the processes that mediate FLC reprogramming for the induction of the appropriate time for flowering.
Abbreviations
- Paf1:
-
RNA polymerase II-associated factor 1
- H3K4me3:
-
Trimethylation at histone H3 Lysine4
- DAG:
-
Days after germination
- qRT-PCR:
-
Quatitative RT-PCR
- siRNA:
-
Short interfering RNA
- miRNA:
-
Micro RNA
- RdDM:
-
RNA dependent DNA Methylation
References
Amasino R (2010) Seasonal and developmental timing of flowering. Plant J 61:1001–1013
Amasino RM, Michaels SD (2010) The timing of flowering. Plant Physiol 154:516–520
Aufsatz W, Stoiber T, Rakic B, Naumann K (2007) Arabidopsis histone deacetylase 6: a green link to RNA silencing. Oncogene 26:5477–5488
Ausin I, Alonso-Blanco C, Jarillo JA, Ruiz-Garcia L, Martinez-Zapater JM (2004) Regulation of flowering time by FVE, a retinoblastoma-associated protein. Nat Genet 36:162–166
Baumbusch LO, Thorstensen T, Krauss V, Fischer A, Naumann K, Assalkhou R, Schulz I, Reuter G, Aalen RB (2001) The Arabidopsis thaliana genome contains at least 29 active genes encoding SET domain proteins that can be assigned to four evolutionarily conserved classes. Nucleic Acids Res 29:4319–4333
Belotserkovskaya R, Reinberg D (2004) Facts about FACT and transcript elongation through chromatin. Curr Opin Genet Dev 14:139–146
Bender J (2004) DNA methylation and epigenetics. Annu Rev Plant Biol 55:41–68
Chan SW, Henderson IR, Jacobsen SE (2005) Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat Rev Genet 6:351–360
Choi J, Hyun Y, Kang MJ, In Yun H, Yun JY, Lister C, Dean C, Amasino RM, Noh B, Noh YS, Choi Y (2009) Resetting and regulation of Flowering Locus C expression during Arabidopsis reproductive development. Plant J 57:918–931
Choi K, Kim J, Hwang HJ, Kim S, Park C, Kim SY, Lee I (2011) The FRIGIDA complex activates transcription of FLC, a strong flowering repressor in Arabidopsis, by recruiting chromatin modification factors. Plant Cell 23:289–303
Choi K, Kim S, Kim SY, Kim M, Hyun Y, Lee H, Choe S, Kim SG, Michaels S, Lee I (2005) SUPPRESSOR OF FRIGIDA3 encodes a nuclear ACTIN-RELATED PROTEIN6 required for floral repression in Arabidopsis. Plant Cell 17:2647–2660
Choi K, Park C, Lee J, Oh M, Noh B, Lee I (2007) Arabidopsis homologs of components of the SWR1 complex regulate flowering and plant development. Development 134:1931–1941
Deal RB, Kandasamy MK, McKinney EC, Meagher RB (2005) The nuclear actin-related protein ARP6 is a pleiotropic developmental regulator required for the maintenance of FLOWERING LOCUS C expression and repression of flowering in Arabidopsis. Plant Cell 17:2633–2646
Deal RB, Topp CN, McKinney EC, Meagher RB (2007) Repression of flowering in Arabidopsis requires activation of FLOWERING LOCUS C expression by the histone variant H2A.Z. Plant Cell 19:74–83
Dennis ES, Peacock WJ (2007) Epigenetic regulation of flowering. Curr Opin Plant Biol 10:520–527
Feng S, Jacobsen SE, Reik W (2010) Epigenetic reprogramming in plant and animal development. Science 330:622–627
Finnegan EJ, Genger RK, Kovac K, Peacock WJ, Dennis ES (1998) DNA methylation and the promotion of flowering by vernalization. Proc Natl Acad Sci USA 95:5824–5829
Gendall AR, Levy YY, Wilson A, Dean C (2001) The VERNALIZATION 2 gene mediates the epigenetic regulation of vernalization in Arabidopsis. Cell 107:525–535
Genger RK, Peacock WJ, Dennis ES, Finnegan EJ (2003) Opposing effects of reduced DNA methylation on flowering time in Arabidopsis thaliana. Planta 216:461–466
Hajkova P, Ancelin K, Waldmann T, Lacoste N, Lange UC, Cesari F, Lee C, Almouzni G, Schneider R, Surani MA (2008) Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 452:877–881
He Y, Doyle MR, Amasino RM (2004) PAF1-complex-mediated histone methylation of FLOWERING LOCUS C chromatin is required for the vernalization-responsive, winter-annual habit in Arabidopsis. Genes Dev 18:2774–2784
Ingouff M, Hamamura Y, Gourgues M, Higashiyama T, Berger F (2007) Distinct dynamics of HISTONE3 variants between the two fertilization products in plants. Curr Biol 17:1032–1037
Ingouff M, Rademacher S, Holec S, Soljic L, Xin N, Readshaw A, Foo SH, Lahouze B, Sprunck S, Berger F (2010) Zygotic resetting of the HISTONE 3 variant repertoire participates in epigenetic reprogramming in Arabidopsis. Curr Biol 20:2137–2143
Jean Finnegan E, Kovac KA, Jaligot E, Sheldon CC, James Peacock W, Dennis ES (2005) The downregulation of FLOWERING LOCUS C (FLC) expression in plants with low levels of DNA methylation and by vernalization occurs by distinct mechanisms. Plant J 44:420–432
Jeddeloh JA, Stokes TL, Richards EJ (1999) Maintenance of genomic methylation requires a SWI2/SNF2-like protein. Nat Genet 22:94–97
Johanson U, West J, Lister C, Michaels S, Amasino R, Dean C (2000) Molecular analysis of FRIGIDA, a major determinant of natural variation in Arabidopsis flowering time. Science 290:344–347
Kim DH, Doyle MR, Sung S, Amasino RM (2009) Vernalization: winter and the timing of flowering in plants. Annu Rev Cell Dev Biol 25:277–299
Kim S, Choi K, Park C, Hwang HJ, Lee I (2006) SUPPRESSOR OF FRIGIDA4, encoding a C2H2-Type zinc finger protein, represses flowering by transcriptional activation of Arabidopsis FLOWERING LOCUS C. Plant Cell 18:2985–2998
Kim SY, He Y, Jacob Y, Noh YS, Michaels S, Amasino R (2005) Establishment of the vernalization-responsive, winter-annual habit in Arabidopsis requires a putative histone H3 methyl transferase. Plant Cell 17:3301–3310
Lee I, Aukerman MJ, Gore SL, Lohman KN, Michaels SD, Weaver LM, John MC, Feldmann KA, Amasino RM (1994) Isolation of LUMINIDEPENDENS: a gene involved in the control of flowering time in Arabidopsis. Plant Cell 6:75–83
Lim MH, Kim J, Kim YS, Chung KS, Seo YH, Lee I, Hong CB, Kim HJ, Park CM (2004) A new Arabidopsis gene, FLK, encodes an RNA binding protein with K homology motifs and regulates flowering time via FLOWERING LOCUS C. Plant Cell 16:731–740
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408
March-Diaz R, Garcia-Dominguez M, Florencio FJ, Reyes JC (2007) SEF, a new protein required for flowering repression in Arabidopsis, interacts with PIE1 and ARP6. Plant Physiol 143:893–901
Martin-Trillo M, Lazaro A, Poethig RS, Gomez-Mena C, Pineiro MA, Martinez-Zapater JM, Jarillo JA (2006) EARLY IN SHORT DAYS 1 (ESD1) encodes ACTIN-RELATED PROTEIN 6 (AtARP6), a putative component of chromatin remodelling complexes that positively regulates FLC accumulation in Arabidopsis. Development 133:1241–1252
Michaels SD, Amasino RM (1999) FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11:949–956
Michaels SD, Amasino RM (2001) Loss of FLOWERING LOCUS C activity eliminates the late-flowering phenotype of FRIGIDA and autonomous pathway mutations but not responsiveness to vernalization. Plant Cell 13:935–941
Michaels SD, Bezerra IC, Amasino RM (2004) FRIGIDA-related genes are required for the winter-annual habit in Arabidopsis. Proc Natl Acad Sci USA 101:3281–3285
Michaels SD, Himelblau E, Kim SY, Schomburg FM, Amasino RM (2005) Integration of flowering signals in winter-annual Arabidopsis. Plant Physiol 137:149–156
Noh YS, Amasino RM (2003) PIE1, an ISWI family gene, is required for FLC activation and floral repression in Arabidopsis. Plant Cell 15:1671–1682
Oh S, Zhang H, Ludwig P, van Nocker S (2004) A mechanism related to the yeast transcriptional regulator Paf1c is required for expression of the Arabidopsis FLC/MAF MADS box gene family. Plant Cell 16:2940–2953
Pien S, Fleury D, Mylne JS, Crevillen P, Inze D, Avramova Z, Dean C, Grossniklaus U (2008) ARABIDOPSIS TRITHORAX1 dynamically regulates FLOWERING LOCUS C activation via histone 3 lysine 4 trimethylation. Plant Cell 20:580–588
Schlappi MR (2006) FRIGIDA LIKE 2 is a functional allele in Landsberg erecta and compensates for a nonsense allele of FRIGIDA LIKE 1. Plant Physiol 142:1728–1738
Sheldon CC, Hills MJ, Lister C, Dean C, Dennis ES, Peacock WJ (2008) Resetting of FLOWERING LOCUS C expression after epigenetic repression by vernalization. Proc Natl Acad Sci USA 105:2214–2219
Sheldon CC, Rouse DT, Finnegan EJ, Peacock WJ, Dennis ES (2000) The molecular basis of vernalization: the central role of FLOWERING LOCUS C (FLC). Proc Natl Acad Sci USA 97:3753–3758
Shindo C, Aranzana MJ, Lister C, Baxter C, Nicholls C, Nordborg M, Dean C (2005) Role of FRIGIDA and FLOWERING LOCUS C in determining variation in flowering time of Arabidopsis. Plant Physiol 138:1163–1173
Sung S, Amasino RM (2004) Vernalization in Arabidopsis thaliana is mediated by the PHD finger protein VIN3. Nature 427:159–164
Swiezewski S, Crevillen P, Liu F, Ecker JR, Jerzmanowski A, Dean C (2007) Small RNA-mediated chromatin silencing directed to the 3′ region of the Arabidopsis gene encoding the developmental regulator, FLC. Proc Natl Acad Sci USA 104:3633–3638
Tamada Y, Yun JY, Woo SC, Amasino RM (2009) ARABIDOPSIS TRITHORAX-RELATED7 is required for methylation of lysine 4 of histone H3 and for transcriptional activation of FLOWERING LOCUS C. Plant Cell 21:3257–3269
Werner JD, Borevitz JO, Uhlenhaut NH, Ecker JR, Chory J, Weigel D (2005) FRIGIDA-independent variation in flowering time of natural Arabidopsis thaliana accessions. Genetics 170:1197–1207
Wu K, Zhang L, Zhou C, Yu CW, Chaikam V (2008) HDA6 is required for jasmonate response, senescence and flowering in Arabidopsis. J Exp Bot 59:225–234
Xu L, Shen WH (2008) Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr Biol 18:1966–1971
Zhang H, van Nocker S (2002) The VERNALIZATION INDEPENDENCE 4 gene encodes a novel regulator of FLOWERING LOCUS C. Plant J 31:663–673
Zhao Z, Yu Y, Meyer D, Wu C, Shen WH (2005) Prevention of early flowering by expression of FLOWERING LOCUS C requires methylation of histone H3 K36. Nat Cell Biol 7:1256–1260
Acknowledgments
We are grateful to S. Michaels for providing the FLC::GUS seeds. We thank R. Amasino, R. Fischer, E. Richard, J. Carrington and Arabidopsis Biological Resource Center (ABRC) for providing mutant seeds. This work was supported by grants from Brain Korea 21 program to H. Yun and Y. Hyun, and from the Korea Research Foundation (KRF-2008-314-C00359) to Y. Choi. This work was also supported by National Research Fund Grant 2009-0079227 from the Ministry of Education, Science and Technology Mid-Career Researcher Program Y. Choi.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yun, H., Hyun, Y., Kang, MJ. et al. Identification of regulators required for the reactivation of FLOWERING LOCUS C during Arabidopsis reproduction. Planta 234, 1237–1250 (2011). https://doi.org/10.1007/s00425-011-1484-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-011-1484-y