Abstract
We report isolation and transcriptional profiling of rice (Oryza sativa L.) mitogen-activated protein kinase (MAPK), OsSIPK (salicylic acid-induced protein kinase). OsSIPK gene is located on chromosome 6 most probably existing as a single copy in the rice genome, and encodes 398 amino acid polypeptide having the MAPK family signature and phosphorylation activation motif TEY. Steady state mRNA analyses of OsSIPK showed weak constitutive expression in leaves of 2-week-old rice seedlings. A time course (30–120 min) experiment using a variety of elicitors and stresses revealed that the OsSIPK mRNA is strongly induced by jasmonic acid (JA), salicylic acid (SA), ethephon, abscisic acid, cycloheximide (CHX), JA/SA + CHX, cantharidin, okadaic acid, hydrogen peroxide, chitosan, sodium chloride, and cold stress (12°C), but not with wounding by cut, gaseous pollutants ozone, and sulfur dioxide, high temperature, ultraviolet C irradiation, sucrose, and drought. Its transcription was also found to be tissue-specifically regulated, and followed a rhythmic dark induction in leaves. Finally, we showed that the OsSIPK protein is localized to the nucleus. From these results, OsSIPK can be implicated in diverse stimuli-responsive signaling cascades and transcription of certain genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diverse extracellular stimuli are transduced into intracellular responses by an evolutionary conserved mitogen-activated protein kinase (MAPK) cascade(s) in various organisms, ranging from yeast to mammals, and plants (Widmann et al. 1999; Ligterink 2000; Jonak et al. 2002). MAPK (serine/threonine PK) is the last component of the kinase cascade, which is activated by dual phosphorylation of a tripeptide motif (Thr-Xaa-Tyr), located in the activation loop (T-loop). This phosphorylation is mediated by a MAPK kinase (MAPKK) that is in turn activated by phosphorylation by a MAPKK kinase (MAPKKK). Several plant MAPKs have been isolated and characterized, and most of our understanding on the MAPK cascade (MAPKKK-MAPKK-MAPK), which is yet to be established based on in vivo evidence, comes from intensive studies in dicot species (Seo et al. 1995; Mizoguchi et al. 1996; Bögre et al. 1997; Ligterink 2000; Ichimura et al. 2002; Jonak et al. 2002). It is only recently that MAPKs of rice, a monocot cereal crop research model (Goff et al. 2002; Yu et al. 2002), are being cloned and characterized (Agrawal et al. 2002, 2003b, c).
The first rice MAPK, BWMK1 (Accession No. AF177392, hereafter called OsBWMK1 based on the adopted nomenclature for MAPKs), isolated from an indica-type cultivar IR36, was shown to be induced by blast pathogen (Magnaporthe grisea) infection and mechanical wounding (He et al. 1999). Recently, our group reported a MAPK, OsMSRMK2 (multiple stress responsive; EMBL Accession No. AJ486975), induced by jasmonic acid (JA) and diverse environmental cues (Agrawal et al. 2002). Subsequently, two independent groups published three reports on the cloning and characterization of MAPKs, OsMAPK2 (Huang et al. 2002), OsMAPK4 (Fu et al. 2002), and OsBIMK1 (Song and Goodman 2002). Although considerable progress has been made toward isolating rice MAPKs, taking into account the number of MAPKs in the dicot model Arabidopsis (20 MAPKs in its genome) (Ligterink 2000; Ichimura et al. 2002; Jonak et al. 2002), it is tempting to speculate that there are novel MAPKs still waiting to be discovered from rice.
In this study, we present identification of novel MAPK (OsSIPK, EMBL Accession No. AJ535841) showing differential response to wounding by cut, and is responsive not only to salicylic acid (SA) treatment but also to diverse stresses, in leaves of 2-week-old rice (cv. Nipponbare) seedlings. Furthermore, a detailed mRNA expression profile analyses strongly suggest their involvement in rice defense/stress response and development.
Materials and methods
Plant material and treatments
Rice (Oryza sativa L. japonica-type cv. Nipponbare; seeds obtained from the National Institute of Agrobiological Sciences, Tsukuba, Japan) were grown under white fluorescent light (wavelength 390–500 nm, 150 μmol m−2 s−1, 12 h photoperiod) at 25°C and 70% relative humidity, and the middle portions (2 cm long) of fully expanded leaves from 2-week-old seedlings were used for all treatments as described previously, under continuous light or darkness (Hanks et al. 1988; Agrawal et al. 2002; Goff et al. 2002). For circadian/rhythmic study, the seedlings at the 2-week-old stage were left in the growth chamber for a further 3 more days. Leaf segments floated on Milli Q (MQ) water in covered Petri dishes served as a wounding by cut treatment (labeled as CUT in the Figures). Leaf segments placed in open Petri dish without MQ water were used for drought treatment. Ultraviolet light (UV-C, 254 nm) was irradiated from a distance of 15 cm using a Hitachi (Japan) germicidal lamp (Agrawal et al. 2002). In order to study the effect of temperature pots containing 2-week-old rice seedlings were either placed under continuous light (150 μmol m−2 s−1) in growth cabinets at 37°C (high temperature), 12°C (low temperature), and 25°C (control) (Agrawal et al. 2002). Whole seedlings were fumigated with ozone (O3; 0.2 ppm) and sulfur dioxide (SO2; 0.5 ppm) exactly as described previously (Millward et al. 1999; Rakwal et al. 2001). Leaves or leaf segments were sampled at the times indicated in the figures, and immediately frozen at −80°C.
Chemicals
Cantharidin (CN), okadaic acid (OA), ±JA, SA, and cycloheximide (CHX) were purchased from Sigma (St. Louis, MO, USA). Abscisic acid (ABA), ethephon (ET, an ethylene generator), hydrogen peroxide (H2O2), the heavy metals, copper (CuSO4), cadmium (CdCl2), and mercury (HgClO3), and fungal elicitor chitosan (CT, water soluble, MW 3,000–30,000) were obtained from Wako Pure Chemicals (Tokyo, Japan). Endothall (EN) was obtained from BIOMOL Research Laboratories Inc. (PA, USA). Stock solutions were prepared as reported previously (Hanks et al. 1988; Agrawal et al. 2002; Goff et al. 2002).
Identification of MAPK-like genes from differentially expressed cDNA library
In our previous study on identification of OsMSRMK2 (Agrawal et al. 2002), few clones having weak signal intensity had been selected on the presumption that some of these clones may be either closely related or novel MAPKs, and that forms the basis of this study. The phagemids (pBluescript SK−) of these positive clones were rescued from phages following the in vivo excision protocol recommended by the manufacturer (Stratagene, La Jolla, CA, USA).
DNA sequencing and sequence analysis
Both strands of the recombinant phagemids were sequenced using a dye-terminator cycle-sequencing kit, and an automated capillary DNA sequencer (Genetic Analyzer ABI 310, PE Applied Biosystems, Foster City, CA, USA). All sequencing data were analyzed using Genetyx software (SDC Software Development, Tokyo, Japan). Alignment and homology of amino acid sequence was done using the MultAlin 5.4.1 (INRA) and CLUSTAL W (1.81) programs against sequences in the GenBank, and EMBL DNA database. The phylogenetic tree was constructed by the NJ (Neighbor–joining) method using the Genetyx program.
Southern analyses
Rice genomic DNA (1 μg) from rice leaves was digested with HindIII, EcoRI, XbaI, BglII, BamHI, or PstI, separated by electrophoresis on a 0.8% agarose gel and blotted onto a nylon membrane (Hybond-N+, Amersham Pharmacia Biotech, Buckinghamshire, UK). The membrane was hybridized with [α-32P]dCTP-labeled (Megaprime DNA labeling system, Amersham) OsSIPK cDNA probe. For this, the OsSIPK-specific probe including the 3′-untranslated region was amplified by performing PCR with the forward primer: 5′-TGGTGTTCTATTTCAGCCTTG-3′ and the reverse primer: 5′-GTTCCAGTCTTACGATCAAC-3′). Hybridized membranes were washed with 2 × SSC and 0.1% (w/v) SDS at 65°C for 1 h, followed by an additional washing with 0.2 × SSC and 0.1% (w/v) SDS at 65°C for 1 h, and exposed to an X-ray film (Kodak, Tokyo, Japan) using two intensifying screens for 2 days at −80°C.
Northern analyses
Total RNA was isolated from rice seedling leaves using the ISOGEN RNA extraction Kit (Nippon Gene, Toyama, Japan), and blotted onto a nylon membrane (Hybond-N+, Amersham, NJ, USA). Northern analyses were carried out as described previously (Agrawal et al. 2002) using the probes mentioned in Southern analyses. The hybridized membranes were washed with 2 × SSC and 0.1% SDS at 65°C for 1 h, and exposed to an X-ray film (Kodak) using two intensifying screens for 2 days at −80°C. Two independent experiments were carried out, and the extracted total RNAs were pooled for the experiment.
Localization and visualization analysis
To make the EGFP :: OsSIPK fusion protein, pK7WGF2 vector was used to clone the OsSIPK gene in frame with the sequence encoding the EGFP protein controlled by a CaMV35S promoter. And for the transient expression assay, the EGFP :: OsSIPK constructs was purified as plasmids using the Qiagen plasmid miniprep kit (Qiagen, Valencia, CA, USA). Plasmid DNAs was coated with tungsten particles and then transformed into the onion epidermal cells by particle bombardment (Bilang and Bogorad 1996). Onion epidermal cells were placed in Petri dishes containing MS media and incubated under dark condition at 28°C for 12 h. DAPI staining was performed to identify the cell nucleus. The epidermal cells expressing the EGFP :: OsSIPK fusion protein was visualized using an Olympus microscope with GFP-optimized ND filter sets (Olympus, Japan). Digital images were collected with an Olympus IX70 fluorescence microscope, an I.CAMSCOPE digital camera (Sometech, Korea), and MicroFiresoftware (USA). The images were further processed with Adobe Photoshop 7.0 software.
Results
Identification of the OsSIPK gene
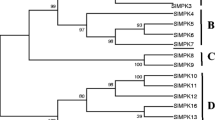
Knowing the importance of MAPKs in signal cascades for plant defense mechanisms against various biotic and abiotic stresses, we cloned few genes related to MAPKs in our previous studies (Agrawal et al. 2002). Of these clones obtained, we identified a novel MAPK gene OsSIPK (EMBL nucleotide sequence database with the Accession No. AJ535841) from JA-treated leaves of japonica-type rice (cv. Nipponbare) seedlings using differential cloning. The OsSIPK cDNA was 1,701-bp-long, contains an open reading frame (ORF) of 1,197 nucleotides, and encoded a protein of 398 amino acid residues with a putative molecular mass of 44858.06 and pI of 5.45 (compute pI/MW tool). The OsSIPK consisted of an ATP-binding region, a MAPK family signature, which indicates that they belong to the MAPK family, a catalytic loop, a phosphorylation motif (TEY motif) in activation loop, and CD domain. The rice genomic database search predicted that rice contains 18 MAPK isoforms. The alignment analysis based on homologies of their amino acid sequences divided them into two groups as shown in Fig. 1. Moreover, the alignment analysis of MAPKs among various species showed that CD domain of OsSIPK, which functions as a binding site of MAPKKs, is evolutionary-conserved as a DxxDE(P)xC motif (inset, Fig. 1). Among MAPKs of rice and other species, OsSIPK (OsMPK1) was grouped into MAPKs containing a TEY motif in activation loop and a CD-domain in C-terminal region, and had high homologies with NtSIPK (87% positive homology; Zhang and Klessig 1997), MsSIMK (85%; Duerr et al. 1993; Jonak et al. 1993) and AtMPK6 (84%; Mizoguchi et al. 1993; Desikan et al. 2001), which are involved in defense mechanism against biotic and abiotic stresses.
Dendrogram of OsSIPK with other MAPK family members. The phylogenetic tree was constructed by NJ (Neighbor–joining) method using the Genetyx program. Predicted functional domain of MAPKs is indicated above the dendrogram. Ser/Thr Kinase domain is indicated in blue box containing sequence of T-loop motif and CD domain needed for MAPK docking interaction is indicated in red. OsMPK1-OsSIPK is marked in red in the dendrogram classified in TEY group. Inset: Conserved domains are analyzed by Rice Genome Automated Annotation System (RiceGAAS; http://ricegaas.dna.affrc.go.jp/) and are highlighted by distinct color; Consensus sequence of the MAPK CD domain and comparison of CD domain of OsSIPK with other MAPKs is shown
Genomic organization and tissue specificity of the OsSIPK gene in rice
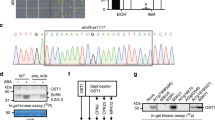
To view the genetic structural features of OsSIPK gene, we confirmed that the OsSIPK gene is located on chromosome 6 and consist of six exons by using the rice genomic database at Knowledge-based Oryza Molecular Biological Encyclopedia (KOME: http://cdna01.dna.affrc.go.jp/cDNA/) (Fig. 2a). The comparative analysis of the exon-intron organization of OsSIPK and its ortholog, AtMPK6, on their genomes showed that the number of exons and the size of their parallel exons between both species are the same and similar, respectively (Fig. 2a). Further, for Southern analysis, rice genomic DNA (1.0 μg) digested with HindIII, EcoRI, XbaI, BglII, BamHI or PstI were prepared, blotted to nylon membrane, and hybridized with a probe derived from 3′-untranslated region of OsSIPK cDNA as shown in Fig. 2b. The result showed that although under low stringency condition (LOW), one additional band with weak intensities is shown in each line with HindIII, XbaI, BglII, or PstI, a single dominant band was detected in each lane under high-stringency condition (HIGH). These data suggest that OsSIPK seems to exist as a single copy in the rice genome.
Exon-intron organization of OsSIPK and its Arabidopsis homolog AtMPK6. The size of genomic DNA was reduced in ratio indicated (a). Genomic DNA is presented by line and boxes describe exons. Start (ATG) and stop (TAG) codons are marked. Southern analysis of OsSIPK. Genomic DNA (1.0 μg) from leaves was digested with HindIII, EcoRI, XbaI, BglII, BamHI, or PstI, separated by electrophoresis on a 0.8% (w/v) agarose gel, and blotted onto nylon membranes (Hybond-N+, Amersham). The membrane was hybridized to the [α-32P]dCTP-labeled OsSIPK cDNA probe under low and high stringency conditions, as described in Materials and methods. M represents the molecular weight marker (λ DNA digested with HindIII) whose fragment size is indicated at the left (b )
Moreover, to observe the tissue-specificity of OsSIPK, we confirmed its transcript level in different tissues, young leaf (L) and shoot (leaf sheath, LS) of 14-day-old seedling, flag (FgL), first leaf (FL), and panicles at different stages of maturity of mature plants (Fig. 3). The results showed that the OsSIPK mRNA is expressed more in leaf sheath than in leaf at the vegetative stage. At maturity, the highest intensity appeared in panicle regardless of its maturation and the lowest signal was detected in flag leaf. These indicate that although OsSIPK is constitutively expressed in all of the tested tissues, its intensity is different depending on the tissue specificity.
OsSIPK expression in rice tissues. a Samples of leaf (L) and leaf sheath (LS) of young (2-week-old) seedlings, and the flag leaf (FgL) and first leaf (FL), and panicles before heading (PBH), after heading (PAH), and at maturity (pollination stage, PAM) of mature plants were taken and frozen at −80°C. Blots were hybridized to [α-32P]dCTP-labeled OsSIPK cDNA probe and single hybridizing bands are shown. Equal loading (20 μg) was confirmed by staining of membranes with methylene blue, and as a representative a part of ribosomal RNA (rRNA) from one membrane is shown. Northern analyses were carried out as described in Materials and methods. b The relative mRNA level was calculated taking the OsSIPK transcript in the young leaf (L) as 100%
Differential regulation of OsSIPK in response to diverse environmental cues
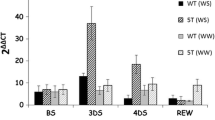
The phylogenic tree of OsSIPK showed that OsSIPK is evolutionarily very close to NtSIPK (Zhang and Klessig 1997), MsSIMK (Duerr et al. 1993; Jonak et al. 1993), and AtMPK6 (Mizoguchi et al. 1993; Desikan et al. 2001), which are implicated in responses to various biotic and abiotic stimuli (Nakagami et al. 2005 and references therein). These results made us examine the responses of OsSIPK to global-signaling molecules such as JA (Reymond and Farmer 1998; Agrawal et al. 2001; Rakwal et al. 2002; Weber 2002), SA (Silverman et al. 1995; Reymond and Farmer 1998; Agrawal et al. 2001), ethylene generator (ET; ethephon) (Ecker 1995; Agrawal et al. 2001), ABA (Grill and Himmelbach 1998; Agrawal et al. 2001) and H2O2 (Grant and Loake 2000; Agrawal et al. 2001), potent protein phosphatase inhibitors (CN and EN) (Millward et al. 1999; Agrawal et al. 2001; Rakwal et al. 2001), fungal elicitor (CT) (Hadwiger 1999; Rakwal et al. 2002), drought, high salt (NaCl), sucrose, heavy metals (copper, cadmium and mercury) and ultraviolet C (UV-C) irradiation in vitro system, and high (37°C) and low (12°C) temperature stresses in vivo system as described previously (Agrawal et al. 2002, 2003b, c). For this, we first observed the changes of OsSIPK transcript level within 15 min because of the intrinsic property of MAPKs, which rapidly and transiently respond to external and internal stimuli (Agrawal et al. 2002). The result showed that the transcript levels of OsSIPK seem not to alter in stressed leaf fragments (data not shown). The unclear early responses of OsSIPK to diverse stressors implored us to further examine its behavior within a delayed time frame, from 30 to 120 min. As shown in Fig. 4, the expression of OsSIPK was not induced by wounding by cut within 120 min. However, it was dramatically induced by JA and SA at 60 min, then maximized at 90 min and maintained till 120 min. Ethylene generator (ET), ABA and H2O2 enhanced its transcript level at 30 min that was early than that seen with JA and SA, and maintained till 120 min. Moreover, a protein synthesis inhibitor, CHX, alone or together with JA or SA, induced its expression within 30 min and increased till 120 min. The treatment of fungal elicitor (CT) and NaCl, phosphatase inhibitors such as CN, EN, and OA, showed that the expression of OsSIPK is increased and reached the maximum at 60 or 90 min. However, drought, UV-C irradiation and sucrose did not give us clear information on OsSIPK expression till 120 min. Furthermore, in vivo experiments showed that the expression of OsSIPK was not enhanced by high (37°C) temperature, O3- and SO2-exposure, whereas low (12°C) temperature increased its transcript level at 30 min that was maintained till 120 min. The results suggest a differential regulation of OsSIPK depending on stressors.
Time course analysis of OsSIPK behavior against diverse environmental factors. Leaf segments were treated with CUT, 100 μM each of JA, SA, ABA, CHX, JA/SA + CHX, protein phosphatases inhibitors [CN, EN, and OA], 1 mM ET, 10 mM H2O2, drought, UV-C, 0.1% CT, 150 mM of NaCl and sucrose (in vitro). For in vivo experiments, the pots containing the 2-week-old seedlings were placed at 37 and 12°C in a controlled growth chamber, or exposed to gaseous pollutants ozone (O3, 0.2 ppm) and sulfur dioxide (SO2, 0.5 ppm) in a controlled fumigation chamber. Treatments were done under continuous light (150 μmol m−2 s−1; or 350 μmol m−2 s−1 for O3 and SO2, CON refers to the clean air control). Equal loading and hybridization was carried out as in Fig. 3
Rhythmically expressed OsSIPK is localized in the nucleus
The previously cloned OsMSRMK2 is weakly and constitutively expressed regardless of light/dark circulation (Agrawal et al. 2002); whereas Fig. 5a showed that the transcription of OsSIPK is repetitively induced at dark period in light (12 h)/dark (12 h) circulation. To know further how light regulates the transcription of OsSIPK, we observed its transcript level under continuous dark condition (Fig. 5b). The result showed that the transcript level of OsSIPK reaches maximum at 12 h and returns to basal level after 24 h.
Rhythmic expression of OsSIPK. Rice seedlings were grown and sampling of leaves was done as described in Materials and methods. Total RNA was isolated at the respective time periods under normal light/dark cycle (a) and under a shift to continuous dark (b) as given above each lane and subjected to Northern analysis as described in Fig. 3. Arrowhead indicates sampling at start of the experiment. Relative abundance of OsSIPK and OsMSRMK2 (a comparative control) mRNAs (in %) according to the signal intensities in panel A/B is given
For the intracellular localization of OsSIPK, a reporter gene encoding GFP was fused to OsSIPK (CaMV 35S-EGFP :: OsSIPK), and then injected into onion (Allium cepa) epidermis cells by using biolistic bombardment as described in Materials and methods. The localization of OsSIPK conjugated with GFP was confirmed in transgenic individual cells by using GFP fluorescence, DAPI staining for nucleus detection, and interference contrast images for whole-cell structures. The results showed that GFP fluorescence signals appear in nucleus in transgenic onion cells containing CaMV 35S-EGFP :: OsSIPK gene, and in nucleus and cytoplasm in control cells containing CaMV 35S-EGFP gene, respectively (Fig. 6). Moreover, the comparative analysis between GFP fluorescence signals in transgenic cells over-expressing EGFP :: OsSIPK and in transgenic onion cells over-expressing EGFP :: CRTintP (calreticulin -interacting protein), which is a marker protein for nucleus localization reconfirmed that OsSIPK is located to nucleus (data not shown). In addition, a prediction program of protein subcellular localization (LOCtree, http://cubic.bioc.columbia.edu/cgi/var/nair/loctree) showed also that OsSIPK might be located in nucleus (Supplementary Fig. 1).
Cellular localization of EGFP :: OsSIPK in onion epidermal cells. Cells were bombarded with constructs carrying EGFP or EGFP :: OsSIPK as described in Materials and methods. To confirm the nuclear localization of the GFP fluorescence by OsSIPK, the positions of nuclei (in blue) were visualized by DAPI staining. Merge image confirms the GFP and DAPI staining results. Results demonstrate that OsSIPK is localized to the nucleus in the bombarded cells
Discussion
Characteristics of the nuclear localized OsSIPK
The MAPK cascade is a crucial component of signal transduction involved in growth, development, and responses to endogenous and environmental cues in all eukaryotes. In plants, several MAPKs activated by a variety of signal molecules (JA, SA, ET, and ABA) and stressors such as pathogen attack, wounding, temperature shift, drought, salt, metal, ozone, UV-C irradiation, and reactive oxygen species (ROS) have been identified (Treisman 1996). The computational analysis of complete rice genome database revealed the existence of 18 rice MAPKs (unpublished observations; see also Liu and Xue 2007). Of rice MAPKs, we cloned several rice MAPKs and reported their transcriptional kinetics by diverse environmental stressors and signal molecules (Agrawal et al. 2002, 2003b, c). Subsequently, we cloned OsSIPK with high amino acid homologies to NtSIPK, AtMPK6, and MsSIMK, which are involved in the response to multiple stressors (Nakagami et al. 2005 and references therein), and revealed its location on chromosome 6 and its existence with one copy number. Particularly, considering that among rice MAPK genes, the number of exon in each gene is very various, from 2 to 12 (Liu and Xue 2007), the comparative analysis of genomic structures of OsSIPK and AtMPK6 indicates that the evolutional distance of their genes may be very close. Moreover, LOCtree program, a prediction program of protein subcellular localization, predicted that OsSIPK and AtMPK6 might be located in nucleus (Supplementary Fig. 1). Actually, Schweighofer et al. (2007) showed that AtMPK6 exists in the nucleus and in the cytoplasm in Arabidopsis protoplast. The results suggest that an ortholog of NtSIPK, AtMPK6, and MsSIMK in rice may be OsSIPK, which is implicated in a multistressor-responsive MAPK cascade functionally.
Tissue specificity and rhythmic expression of OsSIPK
Lieberherr et al. (2005) reported that the transcription and the expression of OsMAPK6, which is identical to OsSIPK, were not induced, but its activity was strongly enhanced within 30 min in sphingolipid elicitor-treated rice cell culture system, and that the expression of pathogen-induced phenylalanine ammonia-lyase (PAL) is regulated by OsMAPK6 pathway. The results could not be matched with the transcript profile of OsSIPK in fungal elicitor-treated rice leaf fragment. However, in Fig. 3 we showed that the transcription efficiency of OsSIPK in green tissues, including leaf (L), flag leaf (FgL), and first leaf (FL), was lower than in leaf sheath (LS) and panicles. Particularly, the transcript levels of OsSIPK during light/dark rhythm was enhanced in the dark and returned to basal level in light, repetitively. Under continuous darkness, the repetitive up- and down-regulation of its transcription disappeared and its expression was transiently enhanced at between 12 and 18 h. The results indicate that the transcription of OsSIPK is differentially regulated depending on tissue specificity and light condition. Thus, the conditions such as tissue and light may induce the different transcription pattern of OsMAPK6 (in cell culture) and OsSIPK.
Transcriptional kinetics of OsSIPK by various stimuli
MAPKs, the last participants of MAPK cascade, in plants, participate in signal perception and transfer and then induce rapidly and correctly necessary information for adaptation from extracellular intimidation during their lifetime (Zhang and Klessig 2001; Nakagami et al. 2005). Thus, most of MAPKs has been reported to be involved in the early response to various stimuli (Zhang and Klessig 2001; Agrawal et al. 2003a; Nakagami et al. 2005). We also reported that diverse stressors induce the dramatic increase of the transcript levels of OsMSRMK2 and OsBWMK1 in leaves within 15 min (Agrawal et al. 2002, 2003b). However, we found that 15 min after treatments by various stressors no significant induction of OsSIPK was visible (data not shown). Further experiments clearly indicated that the transcription of OsSIPK is maximally activated at 60 and 90 min by most of the tested stressors. These results demonstrate that OsSIPK may participate in relatively late response than those of OsMSRMK2 and OsBWMK1.
Moreover, considering the effect of each stressor on the transcription of OsSIPK, signal molecules such as JA, SA, ET and ABA, and CHX, which inhibit translation elongation and assembly of polysome, enhance the accumulation of its mRNA. These effects are similar to those on other OsMAPKs (Agrawal et al. 2002, 2003b, c). Particularly, the CHX effect demonstrates that the transcription of the MAPKs is inhibited by the de novo synthesized negative regulator(s). Further investigation by using three phosphatase inhibitors such as CN, EN, and OA revealed that the dephosphorylation of certain molecule(s) activate the expression of OsSIPK. As previously reported (Agrawal et al. 2002, 2003b, c), the result suggests that the phosphorylation/dephosphorylation of certain signal molecule by kinase/phosphatase regulates the transcription of OsSIPK. In addition, H2O2, one of major ROS, enhances the expression of OsSIPK as shown in Fig. 4. Recently, it has been reported that H2O2 inhibits the activity of Arabidopsis tyrosine phosphatase (AtPTP1) by the oxidation of the cysteine in its catalytic domain and thus regulates the activity of MAPK (Xu et al. 1998; Gupta and Luan 2003). High salt (NaCl), fungal elicitor (CT) and cold stresses also up-regulate the transcription of OsSIPK, similar to those of AtMPK6, OsMSRMK2, and OsMSRMK3 (Nakagami et al. 2005 and references therein). However, although OsSIPK is assigned as a multiple stress-responsive MAPK as mentioned above, it is not induced by drought (one of the most extreme changes in the abiotic environment), sucrose, air pollution, high temperature, and UV-C irradiation. The results suggest considerable and fundamental differences among the MAPKs in sensing the stressors.
The activation of OsMAPK6 was reported to be regulated by OsRac1, which plays crucial roles in rice defense mechanisms, including production of ROS, cell death, phytoalexin production, and induction of transcription of defense-related genes (Lieberherr et al. 2005; Kawasaki et al. 1999; Ono et al. 2001; Wong et al. 2004). In addition, long-lasting activation of SIPK and delayed activation of WIPK are implicated in hypersensitive-response (HR) cell death in tobacco mosaic virus (TMV)-infected or elicitor-treated tobacco plants (Zhang and Klessig 2001). In Fig. 6 it was shown that the OsSIPK protein is localized into the nucleus. The results prove that OsSIPK, which responds to various stressors (Fig. 4), may interact with the active form of OsRac1, and then might be involved in the transcription of HR-related genes.
In conclusion, cloning of OsSIPK, a detailed expression analysis at the transcript level and its subcellular localization analysis revealed that OsSIPK gene is involved in “late” response to diverse stimuli, suppressed by light signal, and localized into nucleus, which will help provide crucial cues for understanding and defining the OsSIPK pathway in rice.
Abbreviations
- ABA:
-
Abscisic acid
- AtMPK6:
-
Arabidopsis thaliana MAPK6
- AtMPK6 :
-
Arabidopsis thaliana MAPK6 gene
- AtPTP1:
-
Arabidopsis thaliana tyrosine phosphatase 1
- CHX:
-
Cycloheximide
- CRTintP:
-
Calreticulin-interacting protein
- CT:
-
Chitosan
- EN:
-
Endothall
- ET:
-
Ethephon
- FgL:
-
Flag leaf
- FL:
-
First leaf
- HR:
-
Hypersensitive-response
- JA:
-
Jasmonic acid
- L:
-
Leaf
- LS:
-
Leaf sheath
- MAPK:
-
Mitogen-activated protein kinase
- MAPKK:
-
MAPK kinase
- MAPKKK:
-
MAPKK kinase
- MsSIMK:
-
Medicago sativa salt-induced MAPK
- MQ:
-
Milli Q
- NtSIPK:
-
Nicotiana tabacum salicylic acid-induced protein kinase
- OA:
-
Okadaic acid
- OsBIMK1 :
-
Oryza sativa benzothiadiazole-induced MAPK1 gene
- OsBWMK1 :
-
Oryza sativa blast and wound-induced MAPK1 gene
- OsMAPK2 :
-
Oryza sativa MAPK2 gene
- OsMAPK4 :
-
Oryza sativa MAPK4 gene
- OsMAPK6:
-
Oryza sativa MAPK6
- OsMAPK6 :
-
Oryza sativa MAPK6 gene
- OsMSRMK2 :
-
Oryza sativa mutiple stress responsive MAPK2 gene
- OsMSRMK3 :
-
Oryza sativa mutiple stress responsive MAPK3 gene
- OsSIPK:
-
Oryza sativa salicylic acid-induced protein kinase
- OsSIPK :
-
Oryza sativa salicylic acid-induced protein kinase gene
- PAH:
-
Panicles after heading
- PAL:
-
Phenylalanine ammonia-lyase
- PBH:
-
Panicles before heading
- ROS:
-
Reactive oxygen species
- SA:
-
Salicylic acid
- TMV:
-
Tobacco mosaic virus
- UV-C:
-
Ultraviolet C
- WIPK:
-
Wound-inducible protein kinase
References
Agrawal GK, Rakwal R, Jwa NS, Agrawal VP (2001) Signaling molecules and blast pathogen attack activates rice OsPR1a and OsPR1b genes: a model illustrating components participating during defense/stress response. Plant Physiol Biochem 39:1095–1103
Agrawal GK, Rakwal R, Iwahashi H (2002) Isolation of novel rice (Oryza sativa L.) multiple stress responsive MAP kinase gene, OsMSRMK2, whose mRNA accumulates rapidly in response to environmental cues. Biochem Biophys Res Commun 294:1009–1016
Agrawal GK, Iwahashi H, Rakwal R (2003a) Rice MAPKs. Biochem Biophys Res Commun 302:171–180
Agrawal GK, Tamogami S, Iwahashi H, Agrawal VP, Rakwal R (2003b) Transient regulation of jasmonic acid-inducible rice MAP kinase gene (OsBWMK1) by diverse biotic and abiotic stresses. Plant Physiol Biochem 41:355–361
Agrawal GK, Agrawal SK, Shibato J, Iwahashi H, Rakwal R (2003c) Novel rice MAP kinase OsMSRMK3 and OsWJUMK1 involved in encountering diverse environmental stresses and developmental regulation. Biochem Biophys Res Commun 300:775–783
Bilang R, Bogorad L (1996) Light-dependent developmental control of rbcS gene expression in epidermal cells of maize leaves. Plant Mol Biol 31:831–841
Bögre L, Ligterink W, Meskiene I, Baker P, Heberle-Bors E, Hirt H (1997) Wounding induces the rapid and transient activation of a specific MAP kinase pathway. Plant Cell 9:75–83
Desikan R, Hancock JT, Ichimura K, Shinozaki K, Neill SJ (2001) Harpin induces activation of the Arabidopsis mitogen-activated protein kinases AtMPK4 and AtMPK6. Plant Physiol 126:1579–1587
Duerr B, Gawienowski M, Ropp T, Jacobs T (1993) MsERK1: a mitogen-activated protein kinase from a flowering plant. Plant Cell 5:87–96
Ecker JR (1995) The ethylene signal transduction pathway in plants. Science 268:667–675
Fu S-F, Chou W-C, Huang D-D, Huang H-J (2002) Transcriptional regulation of a rice mitogen-activated protein kinase gene, OsMAPK4, in response to environmental stresses. Plant Cell Physiol 43:958–963
Grant JJ, Loake GJ (2000) Role of reactive oxygen intermediates and cognate redox signaling in disease resistance. Plant Physiol 124:21–30
Goff SA, Ricke D, Lan TH, Presting G, Wang R, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H, Hadley D, Hutchison D, Martin C, Katagiri F, Lange BM, Moughamer T, Xia Y, Budworth P, Zhong J, Miguel T, Paszkowski U, Zhang S, Colbert M, Sun WL, Chen L, Cooper B, Park S, Wood TC, Mao L, Quail P, Wing R, Dean R, Yu Y, Zharkikh A, Shen R, Sahasrabudhe S, Thomas A, Cannings R, Gutin A, Pruss D, Reid J, Tavtigian S, Mitchell J, Eldredge G, Scholl T, Miller RM, Bhatnagar S, Adey N, Rubano T, Tusneem N, Robinson R, Feldhaus J, Macalma T, Oliphant A, Briggs S (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296:92–100
Grill E, Himmelbach A (1998) ABA signal transduction. Curr Opin Plant Biol 1:412–418
Gupta R, Luan S (2003) Redox control of protein tyrosine phosphatases and mitogen-activated protein kinases in plants. Plant Physiol 132:1149–1152
Hadwiger LA (1999) Host-parasite interactions: elicitation of defense responses in plants with chitosan. In: Jolles P, Muzzarelli RAAA (eds) Chitin and chitinases. Birkhauser- Verlag, Basel, Switzerland, pp 185–200
Hanks SK, Quinn AM, Hunter T (1988) The protein kinase family: conserved features and deduced phylogeny of the catalytic domains. Science 241:42–52
He C, Fong SHT, Yang D, Wang G-L (1999) BWMK1, a novel MAP kinase induced by fungal infection and mechanical wounding in rice. Mol Plant Microbe Interact 12:1064–1073
Huang H-J, Fu S-F, Tai Y-H, Chou W-C, Huang D-D (2002) Expression of Oryza sativa MAP kinase gene is developmentally regulated and stress-responsive. Physiol Plant 114:572–580
Ichimura K, Shinozaki K, Tena G, Sheen J, Henry Y Champion A, Kreis M, Zhang S, Hirt H, Wilson C, Heberle-Bors E, Ellis BE, Morris PC, Innes RW, Ecker JR, Scheel D, Klessig DF, Machida Y, Mundy J, Ohashi Y, Walker JC (2002) Mitogen-activated protein kinase cascades in plants: a new nomenclature. Trends Plant Sci 7:301–308
Jonak C, Páy A, Bögre L, Hirt H, Heberle-Bors E (1993) The plant homolog of MAP kinase is expressed in a cell cycle-dependent and organ specific manner. Plant J 3:611–617
Jonak C, Okresz L, Bögre L, Hirt H (2002) Complexity, cross talk and integration of plant MAP kinase signaling. Curr Opin Plant Biol 5:415–424
Kawasaki T, Henmi K, Ono E, Hatakeyama S, Iwano M, Satoh H, Shimamoto K (1999) The small GTP-binding protein rac is a regulator of cell death in plants. Proc Natl Acad Sci USA 96:10922–10926
Lieberherr D, Thao NP, Nakashima A, Umemura K, Kawasaki T, Shimamoto K (2005) A sphingolipid elicitor-inducible mitogen-activated protein kinase is regulated by the small GTPase OsRac1 and heterotrimeric G-protein in rice. Plant Physiol 138:1644–1652
Ligterink W (2000) MAP kinases in plant signal transduction: how many, and what for. In: Hirt H (ed) MAP kinases in plant signal transduction. Springer, Berlin, pp 11–27
Liu Q, Xue Q (2007) Computational identification and phylogenetic analysis of the MAPK gene family in Oryza sativa. Plant Physiol Biochem 45:6–14
Millward TA, Zolnierowicz S, Hemmings BA (1999) Regulation of protein kinase cascades by protein phosphatase 2A. Trends Biochem Sci 24:186–191
Mizoguchi T, Hayashida N, Yamaguch-Shinozaki K, Kamada H, Shinozaki K (1993) ATMPKs: a gene family of plant MAP kinase in Arabidopsis thaliana. FEBS Lett 336:440–444
Mizoguchi T, Irie K, Hirayama T, Hayashida N, Yamaguchi-Shinozaki K, Matsumoto K, Shinozaki K (1996) A gene encoding a mitogen-activated protein kinase kinase kinase is induced simultaneously with genes for a mitogen-activated protein kinase and S6 ribosomal protein kinase by touch, cold, and water stress in Arabidopsis thaliana. Proc Natl Acad Sci USA 93:765–769
Nakagami H, Pitzschke A, Hirt H (2005) Emerging MAP kinase pathways in plant stress signaling. Trends Plant Sci 10:339–346
Ono E, Wong HL, Kawasaki T, Hasegawa M, Kodama O, Shimamoto K (2001) Essential role of the small GTPase Rac in disease resistance of rice. Proc Natl Acad Sci USA 98:759–764
Rakwal R, Shii K, Agrawal GK, Yonekura M (2001) Protein phosphatase inhibitors activate defense responses in rice (Oryza sativa) leaves. Physiol Plant 111:151–157
Rakwal R, Tamogami S, Agrawal GK, Iwahashi H (2002) Octadecanoid signaling components “burst” in rice (Oryza sativa L.) seedling leaves upon wounding by cut and treatment with fungal elicitor chitosan. Biochem Biophys Res Commun 295:1041–1045
Reymond P, Farmer EE (1998) Jasmonate and salicylate as global signals for defense gene expression. Curr Opin Plant Biol 1:404–411
Schweighofer A, Kazanaviciute V, Scheikl E, Teige M, Doczi R, Hirt H, Schwanninger M, Kant M, Schuurink R, Mauch F, Buchala A, Cardinale F, Meskiene I (2007) The PP2C-type phosphatase AP2C1, which negatively regulates MPK4 and MPK6, modulates innate immunity, jasmonic acid, and ethylene levels in Arabidopsis. Plant Cell 19:2213–2224
Seo S, Okamoto M, Seto H, Ishizuka K, Sano H, Ohashi Y (1995) Tobacco MAP kinase: a possible mediator in wound signal transduction pathways. Science 270:1988–1992
Silverman P, Seskar M, Kanter D, Schweizer P, Metraux JP, Raskin I (1995) Salicylic acid in rice biosynthesis, conjugation and possible role. Plant Physiol 18:633–639
Song F, Goodman RM (2002) OsBIMK1, a rice MAP kinase gene involved in disease resistance responses. Planta 215:997–1005
Treisman R (1996) Regulation of transcription by MAP kinase cascades. Curr Opin Cell Biol 8:205–215
Weber H (2002) Fatty acid-derived signals in plants. Trends Plant Sci 7:217–224
Widmann JW, Gibson S, Jarpe MB, Johnson GL (1999) Mitogen-activated protein kinase: conservation of a three-kinase module from yeast to human. Physiol Rev 79:143–180
Wong HL, Sakamoto T, Kawasaki T, Umemura K, Shimamoto K (2004) Down-regulation of metallothionein, a reactive oxygen scavenger, by the small GTPase OsRac1 in rice. Plant Physiol 135:1447–1456
Xu Q, Fu HH, Gupta R, Luan S (1998) Molecular characterization of tyrosine-specific protein phosphatase encoded by a stress-responsive gene in Arabidopsis. Plant Cell 10:849–857
Yu J, Hu S, Wang J, Wong GK, Li S, Liu B, Deng Y, Dai L, Zhou Y, Zhang X, Cao M, Liu J, Sun J, Tang J, Chen Y, Huang X, Lin W, Ye C, Tong W, Cong L, Geng J, Han Y, Li L, Li W, Hu G, Huang X, Li W, Li J, Liu Z, Li L, Liu J, Qi Q, Liu J, Li L, Li T, Wang X, Lu H, Wu T, Zhu M, Ni P, Han H, Dong W, Ren X, Feng X, Cui P, Li X, Wang H, Xu X, Zhai W, Xu Z, Zhang J, He S, Zhang J, Xu J, Zhang K, Zheng X, Dong J, Zeng W, Tao L, Ye J, Tan J, Ren X, Chen X, He J, Liu D, Tian W, Tian C, Xia H, Bao Q, Li G, Gao H, Cao T, Wang J, Zhao W, Li P, Chen W, Wang X, Zhang Y, Hu J, Wang J, Liu S, Yang J, Zhang G, Xiong Y, Li Z, Mao L, Zhou C, Zhu Z, Chen R, Hao B, Zheng W, Chen S, Guo W, Li G, Liu S, Tao M, Wang J, Zhu L, Yuan L, Yang H (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296:79–92
Zhang S, Klessig DF (1997) Salicylic acid activates a 48-kD MAP kinase in tobacco. Plant Cell 9:809–824
Zhang S, Klessig DF (2001) MAPK cascades in plant defense signaling. Trends Plant Sci 6:520–527
Acknowledgments
The study was funded in part by a grant from the Plant Signaling Network Research Center, Korea Science and Engineering Foundation. The study was also carried out with the support of “On-Site Cooperative Agriculture Research Project (No. 20070301080003), RDA, Republic of Korea (NSJ), and by grant number CG2130 from the Crop Functional Genomics Center of the 21st Century Frontier R&D Program, which is funded by Ministry of Science and Technology of the Republic of Korea. We appreciate a kind gift of rice seeds of the cv. Nipponbare from Dr. Hirohiko Hirochika, Director, Molecular Genetics Department, National Institute of Agrobiological Sciences, Tsukuba, Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Mi-Ok Lee and Kyoungwon Cho contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, MO., Cho, K., Kim, SH. et al. Novel rice OsSIPK is a multiple stress responsive MAPK family member showing rhythmic expression at mRNA level. Planta 227, 981–990 (2008). https://doi.org/10.1007/s00425-007-0672-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-007-0672-2