Abstract
The development of an infection in soybean [Glycine max L. cultivar (cv.) Peking] roots by incompatible (I) and compatible (C) populations of soybean cyst nematode (SCN) (Heterodera glycines) was assayed using an Affymetrix® soybean GeneChip®. This time-course microarray analysis, using 37,744 probe sets, measured transcript abundance during I and C. These analyses reveal that infection by individual I and C H. glycines populations influence the transcription of G. max genes differently. A substantial difference in gene expression is present between I and C at 12 h post infection. Thus, G. max can differentiate between I and C nematode populations even before they have begun to select their feeding sites. The microarray analysis identified genes induced earlier in infection during I than C. MA also identified amplitude differences in transcript abundance between I and C reactions. Some of the probe sets measuring increased transcript levels during I represented no apical meristem (NAM) and WRKY transcription factors as well as NBS-LRR kinases. Later during I, heat shock protein (HSPs) probe sets (i.e. HSP90, HSP70, ClpB/HSP101) measured increased transcript abundance. These results demonstrate that G. max roots respond very differently to the different H. glycines races even before their feeding site selection has occurred. The ability of G. max to engage an I reaction, thus, appears to be dependent on the ability of root cells to recognize the different races of H. glycines because these experiments were conducted in the identical G. max genetic background.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The infection of plants by parasitic nematodes is a major agricultural problem that is poorly understood (reviewed in Williamson and Kumar 2006; Klink et al. 2007a). The most prominent of these interactions is Glycine max infection by Heterodera glycines because it accounts for an estimated $460–$818 million in production losses annually in the US (Wrather and Koenning 2006). Resistance loci to H. glycines are present in the germplasm of G. max and those loci have been physically mapped (Cregan et al. 1999; Matthews et al. 2001). However, G. max resistance to H. glycines is limited to varieties that are the poorest in terms of production yield. Thus, identifying genes that could confer resistance are urgently needed. Resistance is also limited to specific nematode populations within those resistant varieties (Riggs and Schmitt 1991; Niblack et al. 2002).
The H. glycines life cycle (Fig. 1) is approximately 1 month in duration (reviewed in Jung and Wyss 1999; Klink et al. 2007a). Well-defined H. glycines populations that accomplish resistant and susceptible reactions are available for examining G. max–H. glycines interactions (Riggs and Schmitt 1991; Niblack et al. 2002). Those histological studies of the resistant and susceptible interactions between G. max roots and H. glycines have been performed (Endo 1964, 1965, 1971, 1991; Endo and Veech 1970; Gipson et al. 1971; Jones and Northcote 1972; Riggs et al. 1973) and demonstrated the anatomical changes that occur in G. max roots during H. glycines invasion. Interestingly, nematodes burrow into the roots of both resistant and susceptible G. max genotypes and subsequently migrate toward the vascular tissue. The nematode selects a cell adjacent to the vascular tissue and pierces it with its stylet to initiate the development of a feeding site. This occurs at approximately 2 days after inoculation. Subsequently, syncytia are established during both resistant and susceptible reactions. The cells adjacent to the feeding site then become metabolically hyperactive (Endo and Veech 1970; Endo 1971). Then, the walls of the cells adjacent to the selected cell begin to dissolve. The infected plant cell incorporates additional cells by fusion events with neighboring cells by 3 days post-infection. Eventually these recruited cells merge to form a syncytium.
Life cycle of H. glycines. The eggs, within cysts, can lie dormant in the soil for years. Eggs hatch as second stage juveniles (J2s) that migrate toward and subsequently burrow into a root toward the root stele. A stylet emerges from the anterior end of the nematode. The nematode pierces a pericycle cell (green cell with nuclei [red circles]) or neighboring root cell, presumably releasing substances that then causes major changes in the physiology of the root cell. Those root cells (yellow) then fuse with neighboring cells (light blue). Ultimately, this process produces a syncytium (orange), a structure that contains approximately 200 merged root cells. The syncytium serves as the H. glycines feeding site. After the establishment of the syncytium, male nematodes feed for several days until the end of the J3 stage. Males meanwhile become sedentary. Subsequently, the males discontinue feeding and molt into vermiform J4. Males burrow toward the female to copulate. In contrast, the female becomes and remains sedentary after the establishment of their feeding site. The female nematodes then expand circumferentially while undergoing J3 and J4 molts. They then mature into feeding adult females. The female ultimately becomes the cyst that encases the eggs. a Cysts that may remain dormant in the soil for years. b Second stage juveniles (purple) hatch and migrate toward the root of G. max. c J2 nematodes burrow into the root and migrate toward the root vasculature and select a cell for feeding site establishment. d J2 nematodes have molted into J3. They undergo a subsequent molt into J4 nematodes. Meanwhile, the female continues to grow circumferentially as it feeds. The male discontinues feeding at the end of its J3 stage. Male and female J4 nematodes become adults and the vermiform male (blue) migrates to the female to copulate. f After ∼30 days, the female with eggs is clearly visible and emerging from the root. Figure adapted from Klink et al. (2007)
Diverse reactions of incompatibility (I resistance) and compatibility (C susceptibility) become evident subsequently. The susceptible response is characterized by various cellular events that are visible at the anatomical level. These changes include hypertrophy of the nuclei and nucleoli, proliferation of cytoplasmic organelles, reduction or dissolution of the vacuole and expansion of the cell as it incorporates adjacent cells (Jones and Dropkin 1975; Endo and Veech 1970; Gipson et al. 1971; Riggs et al. 1973). In contrast, during the resistance response of G. max cultivar ‘Peking’ to H. glycines NL1-RHp, the syncytium both collapses and becomes necrotic. At 4 days after inoculation, cell wall depositions form and there is an increase in lipid globules that occur before necrosis (Riggs et al. 1973). Concomitantly, degeneration of syncytia occurs. ‘Bedford’, a resistant cultivar, has a somewhat different response. In that reaction, nuclei first degrade. After the breakdown of the nuclei, the cytoplasm degrades (Kim et al. 1987). Eventually, roots overcome infection at the site of infection (Endo 1965, 1991; Riggs et al. 1973; Kim et al. 1987).
These observations demonstrate that a dichotomy between I and C infection occurs. Two characterizations of infection can be inferred from these observations (Fig. 2). In one characterization (Fig. 2a), the plant distinguishes between I and C nematodes early during infection before feeding site selection. In this characterization, gene expression specific to I or C occurs throughout the root. Alkharouf et al. (2006) has already demonstrated differential gene expression during C at 6 hpi. This demonstrates that gene expression may not be limited to the syncytium because they have not even begun to form at 6 hpi. A second characterization (Fig. 2b) is that gene expression pertaining to I and C is localized and is limited to the syncytium. Thus, very little gene expression would be expected prior to feeding site selection. At this time it is unclear if important changes pertaining to I or C are expressed throughout the root even before feeding site selection occurs or only within the developing feeding site. Experiments that study I and C gene expression before the selection of feeding sites by the nematodes (i.e. at 12 hpi) would clarify this issue.
Characterization of infection. In one characterization (a), changes in gene expression occur throughout the root, characterizing I (dark blue) and C (red). These changes in gene expression begin early on (by 12 hpi) as I (pink) and C (green) nematodes burrow through the root. These root transcriptional changes that characterize I and C continue on as nematodes establish syncytia by 3 dpi. Syncytia in I (white arrow) or C (black arrow) roots appear similar anatomically at 3 dpi. By 8 dpi, I syncytia (white arrow) collapse while C syncytia (black arrow) continue to develop. In a second characterization (b), differential gene expression is restricted locally to the syncytium during the onset of I or C. At 12 hpi, little change in differential gene expression is present within I (light blue) and C (light blue) roots. By 3 dpi, little or no change has occurred within roots except within the I syncytium (dark blue, white arrow) or C syncytium (in red, black arrow). By 8 dpi, the I syncytium (in black, white arrow) has collapsed while the C syncytium (in red, black arrow) continues to develop
Microarray analysis (MA) of host–pathogen interactions can provide a broad view of genes, pathways and regulatory nodes involved in defense (Tao et al. 2003; Zabala et al. 2006; Wang et al. 2006). The identification of transcriptional events occurring during A. thaliana infection by P. syringae revealed the roles of resistance (R) genes during I and C (Tao et al. 2003). These experiments demonstrated that a large part of the gene expression changes between I and C are quantitative and useful to determine the nature of I and C. Accelerated transcriptional events in I as compared to C have been identified (Lamb et al. 1992). This was confirmed by MA comparing various A. thaliana genetic backgrounds during I and C reactions to P. syringae (Tao et al. 2003). Thus temporal gene expression events may be important to I. The nature of resistance also has been shown to involve a shift from the induction of housekeeping genes to genes involved in the defense response and energy metabolism during A. thaliana infection by an avirulent strain of P. syringae (Scheideler et al. 2001).
Recently these sorts of genomic analyses have been adapted to the study of plant pathogenic nematodes, determining plant gene expression in C reactions at single (Puthoff et al. 2003; Khan et al. 2004) or multiple time points (Jammes et al. 2005; Alkharouf et al. 2006; Ithal et al. 2007). However, these experiments were not designed to account for gene expression occurring during I. I and C reactions of tomato (Solanum lycopersicum) to the root knot nematode (Meloidogyne spp.) have recently been studied. That study used two different S. lycopersicum varieties and two different nematode species (i.e. M. incognita and M. hapla) to achieve I and C, respectively (Schaff et al. 2007). In G. max, syncytia present in both I and C reactions at 3 dpi have recently been isolated by laser capture microdissection (LCM) and used for MA (Klink et al. 2007b). Syncytia that were microdissected from roots undergoing a C reaction at 3 and 8 dpi have also been used for a time-course study (Klink et al. 2007b). At this time, no genomic information at the microarray level exists on the temporal events pertaining to the development or establishment of I to H. glycines in whole root samples.
The differences in anatomy of G. max root cells undergoing I and C reactions imply that a number of different genes are involved in these dissimilar reactions. We chose G. max cv. Peking because it is highly resistant to one H. glycines population (NL1-RHg) and is susceptible to a different H. glycines population (TN8), allowing us to examine changes in global gene expression during the I and C reactions in the same G. max cultivar. This was accomplished by simply changing the H. glycines population and studying gene expression with the Affymetrix® microarray platform. The Affymetrix® GeneChip® technology used for microarray investigations is an established platform for gene identification (Lipshutz et al. 1999). The GeneChip® has been used in A. thaliana to study C reactions to the pathogen P. syringae over time (Wang et al. 2006) and to compare I and C reactions over time in A. thaliana during P. syringae infection (Tao et al. 2003). The Affymetrix® soybean GeneChip® has been used to study a time-course of infection in C (Ithal et al. 2007). That three-time point study (Ithal et al. 2007) mirrored a seven time point study by (Alkharouf et al. 2006). However, all of the time points selected in that analysis (Ithal et al. 2007) were after feeding site selection had occurred. We believed that the GeneChip® could be used to study a time-course comparing I and C. The G. max–H. glycines system is a good candidate system for comparing gene expression occurring during the development of infection. This is due to the obligate nature of H. glycines and the availability of populations that confer I or C reactions in the same G. max cultivar (i.e. Peking) (Niblack et al. 2002). H. glycines is also a relevant, major pathogen of G. max (Wrather and Koenning 2006). Lastly, the tractability for gene expression analyses (Cho et al. 2001; Zhang and Ghabrial 2006) make it a good system because information on genes identified here can be translated directly to improve resistance in an agriculturally relevant plant. We present here a microarray analysis of G. max cv. Peking whole root infection by H. glycines, comparing I and C responses using the Affymetrix® GeneChip®.
Materials and methods
Plant and nematode procurement
Glycine max and H. glycines were grown at the United States Department of Agriculture Soybean Genomics and Improvement Laboratory, Beltsville, MD, USA. Two populations of H. glycines, NL1-RHg (incompatible I), and TN8 (compatible C), were maintained in the greenhouse using the moisture replacement system (MRS) (Sardanelli and Kenworthy 1997). Seedlings were grown in sterile sand in 20 × 20 × 10 cm flats for a week. The seedlings were then gently removed from the sand and rinsed with sterile water. Seedlings were placed on moistened germination paper (Anchor Paper, St Paul, MN, USA) inside each 20 × 20 × 10 cm flat. Mature female nematodes were harvested from the sand beakers of the MRS 3 days previously, by massaging the roots in water and sieving the solution through nested 850 and 150 μm sieves. Females were purified further by sucrose flotation (Matthews et al. 2003), and the females were crushed gently with a rubber stopper in a 7.5 cm diameter, 250 μm sieve to release the eggs. The eggs flowed through the sieve into a small plastic tray. The debris smaller than the eggs was removed by washing them in a 25 μm mesh sieve. The eggs were placed in a small plastic tray with 1 cm of water. (The tray was covered with plastic wrap and placed on a rotary shaker at 25 rpm. After 3 days, the second stage juvenile nematodes (J2s) were then separated from unhatched eggs by running them through a 41 μm mesh cloth. The J2s were concentrated by centrifugation in an IEC clinical centrifuge for 30 s at 1,720 rpm to 5,000 J2/ml. This represented the inoculum. Two milliliters of inoculum was added directly on the roots for a final concentration of 2,000 J2/root. Control mock-inoculated replicates received the same amount of water. The roots were covered with a moistened sheet of germination paper and placed in a 45 × 50 × 20 cm plastic tray with a 1 cm of water in the bottom to add humidity. A semi-transparent bag was wrapped around the tray, and placed under fluorescent lights of 16/8 h light/dark photoperiod. Identical light intensities were used for each experiment. Infected roots were grown for 12 h post inoculation (hpi), 3 or 8 days post inoculation (dpi). The 12 hpi roots (infected and mock-infected, control) were washed and flash frozen in liquid nitrogen. The other two samples that eventually would become the 3 and 8 dpi samples were also washed to remove the extraneous nematodes that had not yet penetrated the root. This prevented additional nematodes from entering the root. Lateral roots that were maximally infected (as determined later by acid fucshin staining) were harvested and flash frozen in liquid nitrogen. This ensured that we were obtaining the tissue that was the most highly infected with nematodes (Alkharouf et al. 2006). The tissue was ground to a fine powder and stored at −80°C. Total RNA extraction was performed using the method of Mujer et al. (1996), typically yielding 300 μg of total RNA/g of wet tissue. This process was then repeated. Thus, two independent sets of samples, each composed of samples pooled from multiple plants, were generated. For all treatments, at the end of the experiment, representative root samples isolated from each root used in the analysis and used solely to determine the extent of nematode infection was acid fuchsin stained (Byrd et al. 1983). Stereomicroscopy of those acid fuschin-stained roots was performed with a Nikon SMZ 1500 stereomicroscope (Nikon Corporation, Tokyo, Japan). Stereomicroscope images were captured with an Optronics MagnaFire model S99802 CCD camera (Optronics, Goleta, CA, USA). A second set of samples was used for histological observation to confirm nematodes were infecting the roots, selecting feeding sites and creating syncytia (see below).
Histology
Glycine max roots were harvested, cut into 0.5 cm pieces and fixed by vacuum infiltration with in Farmer’s solution composed of 75% ethanol, 25% acetic acid for 12 h at 4°C (Sass 1958; Klink et al. 2005) Farmers solution was removed from the roots using a graded ethanol series (75, 85, 100, 100%), 30 min each, then with 1:1 xylene:ethanol for 30 min, followed by three, 100% xylene incubations (30 min each). Xylene was slowly replaced with Paraplast+® tissue embedding medium (Tyco Healthcare Group LP, Mansfield, MA, USA) in a 58°C oven by infiltrating the roots sequentially in 3:1, 1:1, 1:3 xylene:Paraplast+ in each step for 3 h. Tissue was cast, mounted, and 10 μm serial sections of roots were made using an American Optical 820 microtome (American Optical Co., Buffalo, NY, USA; Klink et al. 2005). Sections were stained with Safranin O in 50% ETOH, counter-stained using Fast Green and then mounted in Permount (Klink et al. 2005).
Microarray analyses
Whole root microarray gene expression analyses were conducted using the GeneChip® Soybean Genome Array (Cat. # 900526; Affymetrix®; Santa Clara, CA, USA) containing 37,744 G. max probe sets (35,611 transcripts). Thus, some redundancy is present. This high-density array consists of an 11-probe pair (25 bp per oligonucleotide) with an 11 μm feature size. It provides multiple independent measurements for each individual transcript. The G. max annotations were kindly provided by Brandon Le and Anhthu Bui of Dr. Robert Goldberg’s lab (University of California-Los Angeles). The annotations were made by comparison to the A. thaliana gene ontology (GO) database (The Gene Ontology Consortium 2004). These annotations are based on the best match from their Blast searches (Altschul et al. 1997). Details of the GeneChip® soybean genome array are available at the Affymetrix® website (http://affymetrix.com/index.affx).
Microarrays were hybridized and scanned at the Laboratory of Molecular Technology, SAIC-Frederick, National Cancer Institute at Frederick, Frederick, MD 21701, USA. Details of the scanning procedure can be found at the Affymetrix website (http://www.affymetrix.com/support/downloads/manuals/expression_s2_manual.pdf).
Bioconductor, a suite of genomics analysis tools (packages) primarily written in the R programming language, was used to analyze the microarray data (Gentleman et al. 2004). R (http://www.r-project.org/) and Bioconductor (http://www.bioconductor.org) are freely available. The Robust Multichip Average (RMA) methodology (Irizarry et al. 2003a, b), as implemented in the Affymetrix® Bioconductor package, was used to extract the expression levels of the probe sets on each chip. The model parameters are estimated using the median polish algorithm (MPA). MPA is a robust procedure that protects against outlier probes. The standard t-test was not used for differential expression analysis because two biological replicates were run for each time point. Instead, a moderated t-statistic was calculated using an Emperical Bayes approach as implemented in the limma Bioconductor package (Gentleman et al. 2004; Smyth 2004). By this method, a more stable estimate of gene-specific variance is calculated by considering the variances of other genes on the microarray. This reduces the possibility of obtaining large variance values. These stable variance estimates were then used to calculate the moderated t-statistic. Subsequently, the moderated t-statistics were used to calculate P-values. Differentially expressed probe sets were any probe set in the test samples (12 hpi, 3 and 8 dpi) having a fold change with absolute value > +1.5 and also having a P-value ≤0.05 as compared to the control (mock inoculated) (Alkharouf et al. 2006). The original data sets, the normalized data sets, statistics and data supplemental to each table and figure are available (http://www.towson.edu/nalkharo/SGMD/SupplementalSites/GmWholeRootIC/).
Results
Histological analysis of I and C responses in the whole root
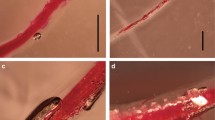
Morphological and anatomical details of I and C disease responses by G. max to H. glycines infection have been published previously (Endo 1964, 1965, 1971, 1991, 1998; Endo and Veech 1970; Gipson et al. 1971; Jones and Northcote 1972; Riggs et al. 1973; reviewed in Jung and Wyss 1999; Klink et al. 2007a). We examined infection and present those data here for comparative purposes to demonstrate the extent and quality of infection in the tissues studied. Our investigation focuses on the events occurring during the first eight days of infection. I or C roots infected for 12 hpi, 3 and 8 dpi were isolated from plants (Fig. 3). We deliberately selected the 12 hpi time point because of our interest in examining whether G. max roots can distinguish between I and C nematodes before feeding site selection occurred. By 12 hpi, I (Fig. 3a) and C (Fig. 3b) nematodes have penetrated the root. I (Fig. 3c) and C (Fig. 3d) nematodes have selected and have begun establishing feeding sites (Fig. 3) by 3 dpi. By 8 dpi, I (Fig. 3e) and C (Fig. 3f) nematodes have established syncytia (Fig. 4) and begin their circumferential expansion that is due to their growth as they feed from syncytia. Roots from those same plants were processed for histological observation to confirm the extent of syncytium formation. Feeding site selection had not begun at 12 hpi (data not presented). Syncytia were developing at 3 dpi during I (Fig. 4a) and C (Fig. 4b) reactions. Syncytia were collapsing at 8 dpi in I (Fig. 4c) but were still developing at 8 dpi in C (Fig. 4d).
Glycine max cv. Peking seedlings were inoculated with I or C H. glycines J2 nematodes. Roots were harvested at 12 hpi, 3 and 8 dpi. a Image of Peking infected with I nematodes at 12 hpi. b Image of Peking infected with C nematodes at 12 hpi. c Image of Peking infected with I nematodes at 3 dpi. Region of syncytium formation is shown (white arrows). Details of developing syncytia are shown in Fig. 4a. d Image of Peking infected with C nematodes at 8 dpi. Region of syncytium formation is shown (white arrows). Details of developing syncytia are shown in Fig. 4b. e Image of Peking infected with I nematodes at 8 dpi. Region of syncytium formation is shown (white arrows). Details of developing syncytia are shown in Fig. 4c. f Image of Peking infected with C nematodes at 8 dpi. Region of syncytium formation is shown (white arrows). Details of developing syncytia are shown in Fig. 4d. (Bar 500 μm)
Glycine max cv. Peking seedlings were inoculated with I or C H. glycines J2 nematodes. Roots were harvested at 12 hpi, 3 and 8 dpi; a 3 dpi Peking infected with I, black arrow syncytium; white arrowhead nematode (bar 50 μm); b 3 dpi Peking infected with C, black arrow syncytium; white arrowhead nematode (bar 50 μm); c 8 dpi Peking infected with I, black arrow syncytium; white arrowhead nematode syncytium (bar 100 μm); d 8 dpi Peking infected with C, black arrow syncytium (bar 100 μm)
Single time point microarray analysis identifies I and C probe sets
Our microarray analysis (MA) examining I and C were performed using the identical G. max genotype (cv. Peking). This eliminates the concern of I and C transcript expression anomalies being caused by using different G. max cultivars. An annotation of the G. max probe sets (Fig. 5) aided various aspects of our analysis.
Probe sets measured induced and suppressed transcript abundance at 12 hpi (defined here as the migration phase), 3 and 8 dpi (Fig. 6a–d). The comparisons were made against a mock-inoculated reference sample. Two classes of time point comparisons were performed after the differentially expressed genes at each time point in the I and C time-series microarrays were identified. These two comparisons are (1) intra-response that analyzes only I or C reactions (Fig. 6) and (2) inter-response that compare I and C reactions (Fig. 7). These analyses focus on two types of differentially expressed probe sets, (1) unique and (2) common. Unique probe sets are defined as those that are differentially expressed and found in only one time point for a particular comparison. Common probe sets are defined as those that are differentially expressed and overlap at multiple time points. Common probe sets can also overlap between I and C reactions, depending on the comparison.
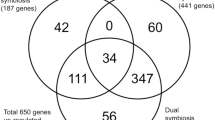
Venn diagrams depicting the numbers of probe sets measuring differential expression in G. max roots during I and C reactions. The size of the circles is not relative to of the quantity of probe sets, respectively. Overlapping areas represent probe sets common between time points. A fold change with absolute value > +1.5 and having a P-value ≤ 0.05 was used for our analyses. Details can be found in the materials section. a Probe sets measuring differentially expressed induced transcript abundance in I. b Probe sets measuring differentially expressed suppressed transcript abundance in I. c Probe sets measuring differentially expressed induced transcript abundance in C. d Probe sets measuring differentially expressed suppressed transcript abundance in C
Venn diagrams depicting the numbers of probe sets measuring differentially expressed transcripts in G. max roots during I and C reactions at their respective time point. The size of the circles is not relative to the quantity of probe sets, respectively. Overlapping areas represent probe sets common to the I and C reaction. A fold change with absolute value > +1.5 and having a P-value ≤ 0.05 cutoff was used for our analyses. Details can be found in the materials section; a 12 hpi induced; b 12 hpi suppressed; c 3 dpi induced; d 3 dpi suppressed; e 8 dpi induced; f 8 dpi suppressed
Our analysis began with the intra-response analysis (Fig. 6a–d). The MA identified probe sets measuring induced transcript abundance for the individual I (Fig. 6a) and C (Fig. 6b) time points. The MA also identified probe sets measuring suppressed transcript abundance for the individual I (Fig. 6c) and C (Fig. 6d) time points.
MA identified a change in the quantity of probe sets at each time point during both I and C (Fig. 6). The single time point MA presented in Fig. 6 was used to identify common probe sets differentially expressed at all time points in a single response. MA identified 339 probe sets measuring induced transcript levels in I at 12 hpi, 3 and 8 dpi (Fig. 6a). Select induced genes common to I at 12 hpi, 3 and 8 dpi (Fig. 6a) are enhanced disease resistance protein 1 (EDS1), intracellular pathogenesis related protein 10 (PR10-like protein), S-adenosyl-l-methionine:carboxyl methyltransferase family protein and hevein-like protein (HEL).
MA identified 740 probe sets measuring suppressed transcript levels in I at 12 hpi, 3 and 8 dpi (Fig. 6b). Among these suppressed genes in I were genes encoding fasciclin-like arabinogalactan-protein (FLA6), multi-copper oxidase type I family protein, peroxidase, lipid transfer protein (LTP) family protein and a pectinesterase family protein.
MA identified 181 probe sets measuring induced transcript levels in C at 12 hpi, 3 and 8 dpi (Fig. 6c). Select induced genes common to C at 12 hpi, 3 and 8 dpi included chitinase, PR10-like protein, germin, cytochrome P450 protein, S-adenosyl-l-methionine and a carboxyl methyltransferase family protein.
MA identified 491 probe sets measuring suppressed transcript levels in C at 12 hpi, 3 and 8 dpi (Fig. 6d). Select suppressed genes common to C at 12 hpi, 3 and 8 dpi included those encoding a glutaredoxin family protein, oleosin, reticulon family protein, 2-oxoglutarate-dependent dioxygenase, zinc finger (CCCH-type) family protein and a chloride channel protein.
We then compared the probe sets measuring induced transcript levels at 12 hpi, 3 and 8 dpi in I (N = 339) (Fig. 6a) with those induced in C at 12 hpi, 3 and 8 dpi (N = 181) (Fig. 6c). MA identified 62 probe sets measuring induced transcript levels at 12 hpi, 3 and 8 dpi that were common to I and C. These probe sets represented genes encoding PR10, several peroxidases, several chalcone synthases, WRKY transcription factors and cytochrome P450. Notably absent in this common set of genes expressed in both I and C were TIR-NBS-LRR protein kinases involved in the defense response (Supplemental Table 1).
The relatively low quantity of induced probe sets common between the 12 hpi, 3 and 8 dpi I and C reactions presented above indicated that most of the probe sets were unique to the I or C response. MA identified 277 probe sets (of the 339 Fig. 6a) measuring induced transcript levels at 12 hpi, 3 and 8 dpi that were unique to I. These probe sets represented several TIR-NBS-LRR protein kinases, extracellular dermal glycoprotein, numerous WRKY transcription factors, cytochrome P450 protein and Kunitz trypsin inhibitor (KTI) (Supplemental Table 2). MA identified 119 probe sets (of the 181 Fig. 6c) measuring induced transcript levels at 12 hpi, 3 and 8 dpi that were unique to C. These probe sets corresponded to genes encoding numerous peroxidases, one WRKY transcription factor and 12-oxophytodienoate reductase (OPR) 3. Again, TIR-NBS-LRR protein kinases were notably absent in C (Supplemental Table 3). The relatively low quantity of probe sets common between I and C and measuring induced differential expression is consistent with shifts from one type (i.e. housekeeping) to another type of gene expression (i.e. defense and energy metabolism) occurring during the course of infection (Scheideler et al. 2001). Complete probe set lists for analyses represented in Figs. 5 and 6 are provided as supplemental data.
Probe sets measuring suppressed transcript levels were the most prevalent in both I (N = 6,377, ∼17% of total probe sets on the microarray) and C (N = 8,061, ∼21.5% of total probe sets on the microarray). The quantity of suppressed probe sets unique to a single time point in I tended to decrease over time (12 hpi, N = 1,660 probe sets; 3 dpi, N = 1,356 probe sets; 8 dpi, N = 804 probe sets). The quantity of suppressed probe sets unique to a single time point in C increased over time (12 hpi, N = 697 probe sets; 3 dpi, N = 1,041 probe sets; 8 dpi, N = 4,048 probe sets). We noted approximately fivefold more suppressed probe sets (N = 4,048) that were unique to 8 dpi in C as compared to I (N = 804). Complete probe set lists for the analyses represented in Figs. 5, 6, 7, 8, 9, 10, 11, 12, 13 are provided as supplemental data.
Single time point MA compares probe sets unique to I and C reactions
Venn diagrams compare the quantity of probe sets that measured differential expression for I and C at 12 hpi, 3 and 8 dpi (Fig. 7a–f). The quantity of induced (Fig. 7a) and suppressed (Fig. 7b) I and C probe sets at 12 hpi are presented. MA identified probe sets measuring induced expression that is unique to I (N = 1,080) and C (N = 899) (Fig. 7a). MA identified probe sets measuring elevated transcript levels of components of the phenylpropanoid pathway in both I and C at 12 hpi (Supplemental Table 4). MA also measured elevated transcript levels of many genes associated with a response by plants to various pathogens. These probe sets are PR1, ACC oxidase, KTI, EDS1, some WRKY transcription factors, one TIR-NBS LRR kinase and some peroxidases. Probe sets measured elevated expression levels for transcripts unique to I at 12 hpi (Fig. 7a, Supplemental Table 5). These probe sets included numerous NBS-LRR and LRR gene family members, numerous kinases, transcription factors related to the no apical meristem (NAM) gene family, a large group of WRKY transcription factors, C2H2, C3H4, CCCH and FYVE-type transcription factors and regulator of chromosome condensation (RCC1). Probe sets measured elevated expression levels for transcripts unique to C at 12 hpi (Fig. 7a, Supplemental Table 6). These probe sets that are unique to C at 12 hpi include cellulose synthase, glucan 1,3-beta-glucosidase precursor, numerous peroxidases, OPR and several disease resistance dirigent proteins. These MA show that substantially more kinase probe sets measure induced transcript levels in I than C at 12 hpi. While numerous WRKY transcription factor probe sets were identified in I, only one probe set unique to C was observed at 12 hpi. Peroxidase probe sets that measured induced transcript abundance were relatively scarce in I as compared to C. The analyses of suppressed probe sets present at 12 hpi in I and C reactions were also identified (Fig. 7b), identifying probe sets unique to I (N = 2,758) or C (N = 1,483). These MA results reveal differences in probe set abundance as a result of infection by I and C nematodes by 12 hpi.
Comparative analyses of induced (Fig. 7c) and suppressed (Fig. 7d) probe sets at 3 dpi are presented. We focus on probe sets measuring induced transcript levels. MA identified ∼13.1 fold more probe sets with induced transcript expression levels present in I (N = 2,030) than C (N = 155) at 3 dpi (Fig. 7c). Many gene families would be expected to be absent at 3 dpi in C because of the relative paucity of probe sets measuring induced transcript levels at that time point. Transcription factor probe sets measuring induced transcript levels at 12 hpi in I were also well represented in I at 3 dpi. Those transcription factors include WRKY, AP2, basic helix–loop–helix, myb and C2H2, C3HC4, CCCH, GATA-type zinc fingers in addition to NAM and scarecrow-like transcription factors. Numerous kinase probe sets measured induced transcript levels in I at 3 dpi. These probe sets include many TIR-NBS-LRR and other LRR receptor-like kinases that are not present in C at 3 dpi (Supplemental Table 7). A substantial amount of probe sets measuring suppressed transcript levels were present in both I and C at 3 dpi (Fig. 7d). MA identified numerous probe sets measuring suppressed transcript levels that are unique to I (N = 3,149) or C (N = 2,290) at 3 dpi (Fig. 7d).
Comparative analyses of induced (Fig. 7e) and suppressed (Fig. 7f) probe sets at 8 dpi are presented. We first focus on probe sets measuring induced transcript levels. MA identified ∼3.1 fold more induced transcripts in I (N = 1,249) than C (N = 409) at 8 dpi (Fig. 7e). Many gene families would be expected to be absent in C at 8 dpi because of the relative scarcity of probe sets measuring induced transcript levels. Many of the kinase and transcription factor probe sets measuring induced transcript levels at 12 hpi and 3 dpi in I, and not well represented in C at any time point, measure induced transcript levels at 8 dpi in I. MA also identified probe sets measuring induced transcript levels at the 8 dpi time point that are not well represented at 12 and 3 dpi I or in any time point in C. MA identified probe sets measuring induced transcript levels of several heat shock proteins (HSPs) and genes involved in energy metabolism at 8 dpi in I (Supplemental Table 8). These probe sets did not measure induced transcript abundance in C and were not well represented in I at 12 hpi and 3 dpi. MA identified a substantial amount of probe sets that are unique to I (N = 1,922) or C (N = 5,209) at 8 dpi and measuring suppressed transcript levels (Fig. 7f).
Single time point MA compares probe sets common to I and C reactions
MA identified probe sets that are common to I and C at 12 hpi (N = 378 Fig. 7a), 3 dpi (N = 349 Fig. 7c), and 8 dpi (N = 390 Fig. 7e) that measured induced transcript levels. These quantities are similar over time. MA identified probe sets that are common to I and C at 12 hpi (N = 420 Fig. 7b), 3 dpi (N = 515 Fig. 7d), and 8 dpi (N = 910 Fig. 7f) measuring suppressed transcript levels. These quantities increase over time, especially between 3 and 8 dpi. These data demonstrate that an underlying core response by G. max to H. glycines infection is present at each time point during I and C. The amplitude of many of the probe sets measuring suppressed transcript levels at 12 hpi is greater in I than C. For example, the probe set measuring the most suppressed transcript level at 12 hpi in I, a peroxidase (AW310249), exhibits a fold suppression of −57.11 (P-value of 0.000058) while in C it exhibits a fold suppression of −1.72184 (P-value of 0.049162). We did not observe probe sets measuring suppressed transcript levels of more than tenfold in C at 12 hpi.
Categorization of probe sets identified in the single time point MA
We further analyzed the probe sets previously identified in the single time point analyses (Fig. 7a–f). Those categorizations (Fig. 8a–f) are the companion to Fig. 7a–f and reveal, on a global scale, the genes experiencing differential expression as a consequence of an I or C reaction. Those analyses revealed many differences between the types of genes expressed during an I or C reaction at different stages of infection. For example, we noted a substantial difference in the number of probe sets measuring increased transcript abundance in I for the following categories; Disease and Defense, Metabolism, Protein Destination and Storage, Signal Transduction and Transcription as compared to C in the 12 hpi samples (Fig. 8a). Differences in the quantities of various groups were also noted in probe sets measuring suppressed transcript abundance at 12 hpi (Fig. 8b). Differences in the quantities of various groups were also noted in probe sets measuring induced (Fig. 8c) and suppressed (Fig. 8d) transcript abundance at 3 dpi. We also observed relatively more Metabolism probe sets in C that measured increased transcript abundance than I in the 8 dpi samples (Fig. 8e). However, we also observed relatively more probe sets in I that measured suppressed transcript abundance than C in the 8 dpi samples (Fig. 8f). The paucity of common probe sets demonstrate the I and C reactions are very different as early as 12 hpi. The annotations of the probe sets permitted us to identify numerous genes common to I and C. Complete probe set lists for analyses represented in Figs. 7 and 8 are provided as supplemental data.
Functional categorization of results obtained in Fig. 7. There are 18 functional categories (A–R). In blue are the compatible unique genes, meaning they are only found in that reaction at that time point. In red are the incompatible unique genes, meaning they are only found in that reaction at that time point. In green are the genes common between I and C. a Probe sets measuring induced transcript abundance at 12 hpi. b Probe sets measuring suppressed transcript abundance at 12 hpi. c Probe sets measuring induced transcript abundance at 3 dpi. d Probe sets measuring suppressed transcript abundance at 3 dpi. e Probe sets measuring induced transcript abundance at 8 dpi. f Probe sets measuring suppressed transcript abundance at 8 dpi
Single time point MA identify accelerated expression during I between time points
Both accelerated gene expression and shifts from the induction of housekeeping genes to the induction of defense and energy metabolism gene expression accounts for changes in gene expression over time. This means that genes important to I experience enhanced transcriptional activity more rapidly (earlier) during I than C. MA identified probe sets measuring induced transcript levels in I at 12 hpi that are not induced in C at 12 hpi, but are induced in C at 72 hpi. Only 32 probe sets fit these criteria (Supplemental Table 9) and include a glycosyl hydrolase family 1 protein, class IV chitinase, KTI, cytochrome P450 family protein, leucine-rich repeat transmembrane protein kinase and WRKY family transcription factor. MA identified probe sets measuring transcript levels that are induced in I at 12 hpi that are not induced in C at 12 hpi or 3 dpi but are induced in C at 8 dpi. MA identified 54 probe sets that fit these criteria (Supplemental Table 10) and include TIR-NBS-LRR gene, SENESCENCE-ASSOCIATED GENE101 (SAG101), GDSL-motif lipase/hydrolase family protein and a WRKY family transcription factor. Our observations indicate that accelerated gene expression is occurring during I. This includes genes involved in the defense response. The quantity of probe sets identified in this analysis, however, is relatively low. This indicates that accelerated gene expression in I as compared to C may not be accounting for the entire disparity of probe sets common between I and C at 12 hpi.
Single time point MA identifies probe sets that are both induced in I and suppressed in C
MA identified that the disparity in probe sets common to I and C at 12 hpi could be due to an extended period of suppressed gene expression in C. MA identified, at each time point, the quantity of probe sets measuring induced transcript levels in I while also measuring suppressed transcript levels in C (Fig. 9a–c). At 12 hpi, MA identified seven probe sets measuring induced transcript levels in I and suppressed levels in C (Fig. 9a). Those probe sets include cytochrome P450 with the remaining being unknown.
Diagram representing numbers of probe sets measuring differential expression at 12 hpi, 3 and 8 dpi. I Incompatible reaction, i induced, C compatible reaction; s suppressed. A fold change with absolute value > +1.5 and having a P-value ≤0.05 cutoff was used for our analyses. a–c Probe sets in the I reaction and measuring induced transcript levels and also found in the C reaction and measuring suppressed transcript levels (I i and C s). d–f Probe sets in the I reaction and measuring suppressed transcript levels and also found in the C reaction and measuring induced transcript levels at 12 hpi (I s and C i). a Probe sets in the I reaction and measuring induced transcript levels and also found in the C reaction and measuring suppressed transcript levels (I i and C s) at 12 hpi. b Probe sets in the I reaction and measuring induced transcript levels and also found in the C reaction and measuring suppressed transcript levels (I i and C s) at 3 dpi. c Probe sets in the I reaction and measuring induced transcript levels and also found in the C reaction and measuring suppressed transcript levels (I i and C s) at 8 dpi. d Probe sets in the I reaction and measuring suppressed transcript levels and also found in the C reaction and measuring induced transcript levels (I s and C i) at 12 hpi. e Probe sets in the I reaction and measuring suppressed transcript levels and also found in the C reaction and measuring induced transcript levels (I s and C i) at 3 dpi. f Probe sets in the I reaction and measuring suppressed transcript levels and also found in the C reaction and measuring induced transcript levels (I s and C i) at 8 dpi
At 3 dpi, MA identified 112 probe sets measuring induced transcript levels in I while being suppressed in C (Fig. 9b). Those probe sets include several NAM transcription factors, zinc finger transcription factors, PAL1, serine carboxypeptidase and extracellular dermal glycoprotein.
At 8 dpi, MA identified 305 probe sets measuring induced transcript levels in I while being suppressed in C (Fig. 9c). Those probe sets include WRKY family transcription factor, zinc finger transcription factor, KTI, GDSL-motif lipase/hydrolase family protein, lipoxygenase (LOX) 1, extracellular dermal glycoprotein and a lectin protein kinase.
The probe sets measuring induced transcript levels at 12 hpi, 3 and 8 dpi in I were then compared to probe sets measuring suppressed transcript levels at 12 hpi, 3 and 8 dpi in C. No transcripts were in common. These results demonstrate that G. max is differentiating between I and C nematode populations at each time point by employing different modes of gene expression as they react to H. glycines infection.
Single time point MA identifies probe sets suppressed in I and induced in C
MA identified probe sets measuring suppressed transcript levels in I and that are also induced in C (Fig. 9d–f). At 12 dpi, MA identified 14 probe sets measuring suppressed transcript levels in I that are also induced in C (Fig. 9d). Those probe sets include expansin, peroxidase, plasma membrane intrinsic protein 1C (PIP1C), germin-like protein (GER) 1, beta-Ig-H3 domain-containing protein and chorismate mutase.
At 3 dpi, MA identified 10 probe sets measuring suppressed transcript levels in I while also being induced in C (Fig. 9e). Those probe sets include 4-coumarateCoA ligase family protein, expansin, LTP1, transketolase and a cytochrome P450.
At 8 dpi, MA identified 35 probe sets measuring suppressed transcript levels in I that are also induced in C (Fig. 9f). Those probe sets include 4-coumarateCoA ligase family protein, peroxidase, expansin, matrix metalloproteinase, matrixin family protein and LTP.
These results demonstrate that G. max can differentiate between I and C nematode populations as early as the migration phase of nematode infection (at 12 hpi). These results also demonstrate that G. max differentiates between I and C nematodes at each time point by employing modes of gene expression that diverge from each other. Complete probe set lists for analyses represented in Fig. 9 are provided as supplemental data.
Time-course MA of the development of I and C reactions
We batched the time-course probe set expression profiles for I and C by their topology (Fig. 10a–h). Differential expression for the time-course of infection is defined here as having a fold change with absolute value > +1.5 and having a P-value ≤ 0.05 at one of the three time points. The probe set must also follow the expression trend imposed by the profile type (Fig. 10a–h). Probe sets both common and unique to I and C were incorporated into the analyses (Fig. 10a–h, respectively). We note that many more profiles are possible if amplitude is taken into consideration between I and C and if common and unique probe sets are presented separately (see supplemental data). We do not present profiles of probe sets that are not differentially expressed in at least one of the three time points. For our analyses, we averaged I and C probe set expression values (averaged amplitude) (Puthoff et al. 2003; Alkharouf et al. 2006) to simplify the presentation of I and C data. For examination of individual expression values please refer to the supplemental data. The quantity of probe sets presented in Profiles 1–8 (Fig. 10a–h, respectively) will not match those obtained for the single time point analysis (Figs. 6, 7, 8, 9) because different criteria were used in these analyses.
Probe sets having similar topologies between I and C over time. Eight differential expression profiles for G. max probe sets identified by MA of whole roots infected with I (in blue) or C (in red). The expression profiles depict trend over time. The time points selected were 12 hpi, 3 and 8 dpi. The probe sets presented are those measuring differential expression for at least one of the three time points per profile. The time point is provided on the x-axis. The y-axis values are the fold change of the test sample versus the control (test/control). The expression profiles are presented as an average value of all probe sets fitting the expression criterion imposed by each profile. The average fold change is obtained by summing all of the fold change values for all of the transcripts and dividing by the number of probe sets present in each profile. The number (N) of probe sets common and unique to I and C is provided in Table 2. The y-axis values are the fold change of the test sample versus the control (test/control). a (Profile 1), probe sets measure induced transcript abundance at the 12 hpi, 3 and 8 dpi time points. b (Profile 2), probe sets measuring transcript abundance that is induced at the 12 hpi and 3 dpi time points and suppressed at the 8 dpi time point. c (Profile 3), probe sets measuring transcript abundance that is induced at the 12 hpi time point and suppressed at the 3 and 8 dpi time points. d (Profile 4), probe sets measuring transcript abundance that is induced at the 12 hpi time point suppressed at the 3 dpi time point and induced at the 8 dpi time point. e (Profile 5), probe sets measuring transcript abundance that is suppressed at the 12 hpi time point and induced at the 3 and 8 dpi time points. f (Profile 6), probe sets measuring transcript abundance that is suppressed at the 12 hpi and 3 dpi time points and induced at the 8 dpi time point. g (Profile 7), probe sets measuring transcript abundance that is suppressed at the 12 hpi time point, induced at the 3 dpi time point and suppressed at the 8 dpi time point. h (Profile 8), probe sets measuring transcript abundance that is suppressed at the 12 hpi, 3 and 8 dpi time points
MA identified probe set profiles present during the time-course of infection [Fig. 10a–h, respectively (and supplemental data)]. In most cases, the averaged I and C probe set expression values are similar over time (Fig. 10a–h). One notable exception was the identification of probe sets measuring more pronounced suppressed transcript abundance at 12 hpi in I as compared to C (Fig. 10h). Probe sets common between I and C for each profile, a subset of those presented in Fig. 10a–h, represented a smaller proportion of those probe sets, respectively (Table 1). A larger proportion of the probe sets composing Fig. 10a–h, thus, were unique to I or C. Select probe set lists that are unique to I [Fig. 10a–h (Profiles 1–8)] are provided (Table 2). These results demonstrate that some I and C probe sets are measuring similar expression profiles over time. However, a substantial proportion of the probe sets (Fig. 10a–h) are measuring transcript abundance found uniquely in I or C. These MA demonstrate that G. max can differentiate between I and C nematode populations during the time-course of infection.
Categorization of probe sets identified in the time-course MA
We further analyzed the probe sets identified in the time-course MA (Fig. 10a–h). Those categorizations (Fig. 11a–h) are the companion to Fig. 10a–h and reveal, on a global scale, the genes experiencing differential expression as a consequence of an I or C reaction for Profiles 1–8. For example, Profile 1 (Fig. 10a) shows a substantial difference in the number of probe sets measuring induced transcript abundance at all time points for genes categorized in the Disease and Defense, Signal Transduction and Transcription bins (Fig. 11a). Those analyses revealed many other differences between the types of genes expressed during an I or C reaction (Fig. 11b–h). Notably, for Profile 7 (Fig. 11g), probe sets measured more Cell Growth and Division and Cell Structure in I than C. Complete probe set lists for analyses represented in Figs. 10 and 11 are provided as supplemental data.
Functional categorization of results obtained in Fig. 10. There are 18 functional categories. In red are the probe-sets present in the I reaction. In red are the probe-sets present in the C reaction. a (Profile 1), functional categorization of Fig. 10a; b (Profile 2), functional categorization of Fig. 10b; c (Profile 3), functional categorization of Fig. 10c; d (Profile 4), functional categorization of Fig. 10d; e (Profile 5), functional categorization of Fig. 10e; f (Profile 6), functional categorization of Fig. 10f; g (Profile 7), functional categorization of Fig. 10g; h (Profile 8), functional categorization of Fig. 10h
Time-course MA identifies contrasting probe set expression between I and C reactions
MA demonstrated that a smaller proportion of probe sets are common between I and C at single time points and during the time-course of infection. Thus, many probe sets are measuring contrasting transcript abundance over time. Numerous comparisons can be made concerning contrasting patterns of transcript abundance occurring between I and C that may be important to I. However, analyzing all of them is well beyond the scope of this paper. To focus our efforts, as for the single time point analysis, we examined accelerated gene expression during I as compared to C (Fig. 12a–c). The same probe sets compose I and C in each of the six individual experiments (Fig. 12a–c).
Contrasting probe set expression patterns (having dissimilar topologies between I and C over time). Six differential expression profiles (a–f) for G. max probe sets identified by MA of whole roots infected with I (in blue) or C (in red). The time points were 12 hpi, 3 and 8 dpi. The expression profiles are presented as the average expression for the sum of probe set measurements for each profile. The number (N) of genes for both I and C G. max transcripts in each profile is provided in the bottom right corner of each figure. The time point is provided on the x-axis. The fold expression is provided on the y-axis. a Probe sets measuring transcript abundance that is induced in I and suppressed in C. b Probe sets measuring transcript abundance that is induced in I at all time points while in C are suppressed at 12 hpi but then become induced at 3 and 8 dpi. c Probe sets measuring transcript abundance that is induced in I at all time points while in C are suppressed at 12 hpi and 3 dpi but then become induced at 8 dpi. d Probe sets measuring transcript abundance that is suppressed at all time points in C and suppressed in I at 12 hpi but then become induced at 3 and 8 dpi. e Probe sets measuring transcript abundance that is suppressed at all time points in C and suppressed in I at 12 hpi and 3 dpi but induced at 8 dpi. f Probe sets measuring transcript abundance that is suppressed in C at 12 hpi and 3 dpi but become induced at 8 dpi. In I, those probe sets measuring transcript abundance that is suppressed at 12 hpi but become induced at 3 and 8 dpi
MA analysis identified 196 probe sets measuring induced transcript abundance at all time points during I while being suppressed at all time points during C (Fig. 12a). These probe sets included TIR-NBS-LRR kinase and other LRR receptor-like kinases, transcription factors including WRKY, NAM, AP2, ERF and centroradialis protein. Notably absent during I were peroxidases.
MA analysis identified 29 probe sets measuring induced transcript levels at all time points during I that during C are suppressed at 12 hpi and subsequently induced at 3 and 8 dpi (Fig. 12b). These probe sets include lectin protein kinase, a WRKY transcription factor, cytochrome P450 family protein, calreticulin, LRR receptor-like kinase and an HSP.
MA analysis identified 12 probe sets measuring induced transcript levels at all time points during I that during C are suppressed at 12 hpi and 3 dpi and subsequently induced at 8 dpi (Fig. 12c). These transcripts include cytochrome P450, WRKY family transcription factor, AP2 domain-containing transcription factor, thaumatin and a LRR receptor-like kinase.
MA also identified probe sets that measure transcript abundance that becomes induced at various time points during I while being suppressed during C at all time points (Fig. 12d, e). MA analysis identified 150 probe sets measuring suppressed transcript levels during I at 12 hpi that subsequently are induced at 3 and 8 dpi. During C, these probe sets measure suppressed transcript levels at all time points (Fig. 12d). These probe sets include LOX1, WRKY, myb, and NAM family transcription factors, F1F0-ATPase inhibitor protein and acid phosphatase.
MA analysis identified 99 probe sets that measure suppressed transcript abundance during I at 12 hpi and 3 dpi that subsequently are induced at 8 dpi. During C, these probe sets measure suppressed transcript levels at all time points (Fig. 12e). These probe sets include RCC1, LOX2, shaggy-related protein kinase, leucine-rich repeat transmembrane protein kinase and HSP7.
MA identified 16 probe sets that, during I, measure suppressed transcript abundance at 12 hpi and subsequently measure induced levels at 3 and 8 dpi during I. In C, these probe sets measure suppressed transcript abundance at 12 hpi and 3 dpi that then become induced at 8 dpi (Fig. 12f). These probe sets include two LRR kinases, protamine, phosphoenolpyruvate carboxylase and an ankyrin repeat family transcript. The MA identified that contrasting gene expression appears to represent a significant amount of the expression occurring during I and C over time. A subset of this contrasting gene expression is accelerated transcript expression occurring during I as compared to C. Some of these genes are known to be involved in defense.
Categorization of contrasting probe set expression between I and C reactions
We further analyzed the probe sets identified in the contrasting probe set expression analyses (Fig. 12a–f). Those categorizations (Fig. 13a–f) are the companion to Fig. 12a–f and reveal, on a global scale, the genes experiencing contrasting expression as a consequence of an I and C reaction. The same probe sets compose I and C in each of the six experiments (Fig. 13a–f). For example, we noted a substantial number of probe sets induced at all time points in I that were suppressed at all time points in C that belong to the Metabolism category (Fig. 13a). While low numbers of probe sets were present in Fig. 13b and c, MA did identify contrasting expression for Energy and Metabolism probe sets that occurs between 12 hpi and 3 dpi in (Fig. 13d). Contrasting gene expression for Signal Transduction and Transcription probe sets was also observed in I, occurring between 3 and 8 dpi (Fig. 13e). Low numbers of Fig. 12f probe sets were available for classification (Fig. 13f), complicating analysis. Complete probe set lists for analyses represented in Figs. 12 and 13 are provided as supplemental data.
Functional categorization of results obtained in Fig. 12. There are 18 functional categories. In blue are the genes common to the I and C reaction. a Functional categorization of Fig. 12a; b functional categorization of Fig. 12b; c functional categorization of Fig. 12c; d functional categorization of Fig. 12d; e functional categorization of Fig. 12e; f functional categorization of Fig. 12e
Discussion
Our study focused on using well-defined nematode populations that yielded bona fide I and C reactions (Niblack et al. 2002) in the same G. max cultivar. We addressed several basic questions concerning the G. max–H. glycines interaction. These questions include: (1) does differential gene expression occur prior to H. glycines feeding site selection? (2) Can G. max distinguish between H. glycines populations before feeding site selection occurs by the nematodes? (3) How does differential expression differ between I and C over time? Our MA evaluated transcript abundance over time, identifying genes both common and unique to I and C and genes experiencing contrasting patterns of expression.
Glycine max exhibits differential expression prior to H. glycines feeding site selection
Damage trails caused by H. glycines as it migrates to its feeding site are clearly evident at the histological level at 12 hpi. These observations indicated that changes in gene expression in G. max probably were occurring as soon as J2 H. glycines entered the root. MA identification of rapid changes in gene expression during pathogen infection has been demonstrated (Scheideler et al. 2001). Those experiments, using A. thaliana, demonstrated that P. syringae infection promoted the differential expression of ∼650 genes within the first 10 min of infection. Our MA identified transcriptional activity present at 12 hpi that was common to I and C. This demonstrated that G. max has the ability to sense the presence of I and C nematode populations during the migration phase. These observations are consistent with Alkharouf et al. (2006) who demonstrated a change in transcriptional activity in C as early as 6 hpi. Our MA identified very little change in the quantity of probe sets measuring induced transcript levels that are common between I and C at individual time points. These data demonstrate that an underlying core response by G. max to H. glycines infection is present at each time point during I and C.
MA indicated that the most highly induced gene common to I and C encodes PR1. PR1 acts downstream of Non expressor of PR genes 1 (NPR1) (Cao et al. 1994) during SA signaling. Other defense-related probe sets identified in our analysis included EDS1. EDS1 works in concert with the known EDS1 interactor PHYTOALEXIN-DEFICIENT4 (PAD4) and SAG101 to mediate resistance in A. thaliana to P. syringae (Feys et al. 2005). Transcript levels of PAL1, encoding an enzyme that is at the beginning of the phenylpropanoid pathway are induced in I and C at 12 hpi. Numerous other genes encoding enzymes of the phenylpropanoid defense pathway are also induced in I and C at 12 hpi. Genes encoding LOX, a component of the jasmonic acid defense pathway, and ACC oxidase, involved in ethylene signaling, are induced in I and C. MA identified a set of gene-for-gene resistance (R) genes induced in both I and C. R genes are essential for defense and confer resistance in plants to other nematode species such as the parasitic nematode H. schachtii (Cai et al. 1997) M. javanica and M. incognita (Milligan et al. 1998) and Globodera rostochiensis (Paal et al. 2004). MA also identified induced transcript levels of the genes encoding members of cell wall modifying dirigent proteins. Dirigent proteins are involved in lignification (Burlat et al. 2001) and their genes have been shown to experience significant increase in transcript abundance upon infection of the white pine weevil (Pissodes strobi Peck), in Sitka spruce [Picea sitchensis (Bong) Carriere; clone FB3-425] (Ralph et al. 2006). Our MA results demonstrate that changes in gene expression in G. max roots occur rapidly after infection by H. glycines. This rapid induction includes genes encoding components of multiple defense pathways. These genes were found to be induced in common between both I and C at 12 hpi. Alkharouf et al. (2006), using cDNA-based microarrays containing >6,500 features, used time points spanning both the migration (6 and 12 hpi) and syncytium establishment phases (1, 2, 4 and 8 dpi) to study C. That analysis demonstrated that hundreds of G. max genes were differentially expressed at 6 hpi during the nematode migration phase. This is the only microarray analysis that evaluated G. max transcript expression during the migration phase of the nematode. Some of the genes identified in that analysis were defense-related and included KTI, germin, peroxidase, phospholipase D, OPR, PR1, phospholipase C, lipoxygenase, WRKY6 transcription factor and calmodulin (Alkharouf et al. 2006). Thus, multiple defense pathways were induced early during the C reaction (Alkharouf et al. 2006). Our MA results confirm that differential expression is occurring during the nematode migration phase and that also demonstrates a core gene expression pattern in G. max roots irrespective of the nematode population used. It is unclear that these induced defense genes are relevant to resistance because they are common to I and C. Our observations show that the same G. max genetic background (cv. Peking) can distinguish between I and C H. glycines populations. Recent experiments of a C reaction in S. lycopersicum cv. Moneymaker infected by the root knot nematode (RKN) (M. incognita) revealed some changes in gene expression by 12 hpi (Schaff et al. 2007). These experiments also compared an I reaction in a different cultivar S. lycopersicum cv. Motelle by infection with another RKN species M. hapla (Schaff et al. 2007). Those experiments did identify some genes (N = 5) in common between I and C. Unfortunately, direct comparisons of gene activity occurring between the I and C reactions at 12 hpi were not presented. However, observations of both SCN and RKN infection processes demonstrate that early transcriptional events pertaining to I and C during the migration phase are occurring and may exhibit aspects common to each.
The large quantity of differentially expressed genes unique to I and C suggest that contrasting gene expression may be important during I and C during the course of H. glycines infection. These genes that are exhibiting contrasting expression may represent the lagging gene expression identified by Lamb et al. (1992). Our MA indicated that G. max differentiated between I and C nematodes, which supports the observations in A. thaliana (Tao et al. 2003) and G. max (Zabala et al. 2006) that these plants could differentiate between I and C strains of P. syringae. Several studies have demonstrated that gene expression during a disease response changes over time during P. syringae infection of A. thaliana and this can be seen as a shift from housekeeping genes to defense genes and energy metabolism used to fuel the defense response (Scheideler et al. 2001). Transcript expression during a disease response also lags over time in C as compared to I during P. syringae infection of A. thaliana, presumably providing a temporal advantage to the host when defending against the pathogen (Tao et al. 2003). Our observations of fluctuations in gene expression occurring at least over the first three days of infection during I in G. max contrast with observations made during RKN infection of S. lycopersicum (Schaff et al. 2007) as we observed both changes in expression occurring over time and differences in expression chronology between the I and C reactions.
We compared the quantity of induced and suppressed transcript levels for I and C at 12 hpi, 3 and 8 dpi. MA identified very few induced genes at 3 dpi in C as compared to I. MA also identified a contrasting trend in transcript expression occurring between 3 and 8 dpi for I and C. The quantity of suppressed genes in I at 3 dpi had decreased at 8 dpi. The quantity of suppressed genes in C at 3 dpi had increased substantially by 8 dpi. These expression trends demonstrate that G. max roots respond differently during I and C. The observation that relatively few transcripts are common between I and C as compared to the quantity unique to I or C demonstrate that G. max perceives the two populations of H. glycines differently during infection. These observations are consistent with I and C infection of S. lycopersicum with M. hapla and M. incognita, respectively (Schaff et al. 2007). These transcripts that are exhibiting contrasting gene expression may represent the lagging gene expression found to be important during I (Lamb et al. 1992; Tao et al. 2003) that represent shifts from one type of gene expression to another during the course of infection (Scheideler et al. 2001). That was clear from our functional categorizations of those differentially expressed probe sets.
Glycine max experiences transcript expression unique to I during H. glycines migration phase
We sought to determine if there were genes expressed uniquely during I or C at 12 hpi. This would provide an estimate as to whether G. max could sense I and C nematodes differently before they had established feeding sites. Our MA identified a relatively small quantity of probe sets common between I and C as compared to those unique to I and C that were measuring suppressed transcript levels. Many of these probe sets measured amplitude differences between I and C. These observations demonstrate that G. max can sense the presence of individual populations of H. glycines at 12 hpi. This population-specific response by G. max to infection by H. glycines occurs early during infection, demonstrating that the population-specific sensing process by G. max begins as the nematodes are burrowing through the root. This is consistent with the ability of A. thaliana to distinguish rapidly (within 9 h of infection) between I and C strains of P. syringae (Tao et al. 2003). Similar observations were also made concerning the differential expression of genes encoding components of the phenylpropanoid pathway in G. max during infection with I and C strains of P. syringae (Zabala et al. 2006). These results are consistent with Zou et al. (2005) who demonstrated that approximately 10% of G. max leaf genes are differentially expressed during P. syringae infection during I versus C.
Our gene classifications permitted the identification of quantitative differences for Disease and Defense, Metabolism, Secondary Metabolism, Signal Transduction and Transcription probe sets occurring as early as 12 hpi between I and C reactions. These results may indicate that G. max is fortifying root cells in ways that drive I later during feeding site establishment phase of the H. glycines lifecycle. This fortification may involve the large number of WRKY genes that are induced in I reactions at both 12 hpi and 3 dpi. WRKY genes are important to defense. There was a substantial difference in the expression of genes encoding WRKY transcription factors between I and C. We identified 107 members of the WRKY family of transcription factors for G. max on the Affymetrix® microarrays. Analysis of these WRKY probe sets revealed complex expression both during the course of infection and between I and C nematode populations beginning at 12 hpi. MA identified a large quantity of genes encoding WRKY transcription factors that were uniquely expressed in I at 12 hpi. WRKY transcription factors are important in defense (Wang et al. 2006; Zheng et al. 2006; Liu et al. 2006; Ryu et al. 2006; Ulker et al. 2007). A WRKY6 homolog was shown to be induced during H. glycines infection of G. max (Alkharouf et al. 2006). WRKY genes have been shown to be rapidly induced during I in pepper (Capsicum annum) (Oh et al. 2006). Numerous other observations also point toward the involvement of WRKY genes in plant defense (Zheng et al. 2006; Ryu et al. 2006; Xu et al. 2006; Journot-Catalino et al. 2006; Shen et al. 2007).
R genes work in concert with WRKY genes during the defense response. We observed a substantial number of NBS-LRR (R) genes induced during I. Gene for gene resistance is essential for I by recognizing pathogen avirulence (Avr) proteins (Hammond-Kosak and Jones 1997; Dangl and Jones 2001) and are encoded by NBS-LRR (R) genes. These R genes exhibit a substantial difference in transcript abundance between I and C during G. max infection by H. glycines. We identified 68 NBS-LRR-like genes on the microarrays and many of those genes were induced in I by 12 hpi. The induction of R gene expression that was unique to I is consistent with their involvement in resistance to parasitic nematodes (Cai et al. 1997; Milligan et al. 1998; Hwang et al. 2000, 2003; Paal et al. 2004). However, it was unclear whether a functional link exists between these WRKY and R genes. Another class of immune receptors is the membrane-resident pattern recognition receptors (PRRs) (Jones and Dangl 2006). These proteins recognize pathogen proteins (i.e. bacterial flagellin or chitin) referred to as pathogen-associated molecular patterns (PAMPs) (Jones and Dangl 2006). The PRRs are involved in impeding pathogen growth. R gene and PRR pathways are linked and involve WRKY transcription factors that de-repress basal defense that is triggered by PAMPs (Shen et al. 2007). In those experiments, the powdery mildew Blumeria graminis AVR A10 protein works through the barley mildew A (MLA) R locus (MLA10) to associate with WRKY transcription factors. WRKY then represses PAMP-triggered plant basal defense. This repression caused by WRKY is interfered with by MLA and thus ensures that basal resistance activity is maintained (Shen et al. 2007). It is not clear whether functional homologs exist in all agricultural plants. However, the induction of WRKY and R gene expression during I as G. max overcomes infection by H. glycines is consistent with such a defense role.
Glycine max experiences transcript expression unique to C during H. glycines migration phase
An intriguing observation was the large quantity of genes encoding peroxidases that were induced specifically in C by 12 hpi, but were not induced in I. Peroxidases are involved in the hypersensitive response (HR). Scheideler et al. (2001) demonstrated in A. thaliana infected with P. syringae that most of the HR response was complete by 24 hpi. However, in our analysis, some peroxidases were also found to be common to I and C, but many more were found uniquely in C at 12 hpi. Our MA identified only one peroxidase gene that was induced that was unique to I at 12 hpi. Thus, while the induction of WRKY and R gene mediated defense pathways are prominent in I at 12 hpi, a large quantity of peroxidase genes are induced in C at 12 hpi. Our analysis was not designed to account for gene expression prior to 12 hpi. Thus, it is possible that peroxidases are induced in I prior to 12 hpi, representing accelerated or shifts in gene expression.
Glycine max experiences transcript expression unique to I at 3 dpi
There are substantial differences in the quantity of genes induced in I as compared to C at 3 dpi. Our MA revealed a large quantity of defense probe sets at 3 dpi in I that includes several peroxidases. Numerous transcription factor probe sets, including the aforementioned WRKY transcription factors, are well represented in I at 3 dpi.
The NAM family of transcription factors is particularly well represented in I at 3 dpi. NAM belongs to a family of transcription factors first identified in Petunia (Souer et al. 1996) and is required for pattern formation in embryos and flowers. Its expression pattern in meristems and organ boundaries was consistent with mutant phenotypes lacking apical meristems or having aberrant floral organ quantity (Souer et al. 1996). NAM is a member of a family of transcription factors including NAM, ATAF, and CUP-SHAPED COTYLEDON (CUC) and referred to as NAC (Vroeman et al. 2003). A. thaliana has at least 105 NAC genes while Oryza sativa has at least 75 (Olsen et al. 2005). The NAC family gene, ATAF (Collinge and Boller 2001), is induced by wounding and P. syringae infection. The O. sativa NAC gene, OsNAC6, is responsive to wounding and JA treatment (Ohnishi et al. 2005). NAC1 is involved in lateral root emergence (Guo et al. 2005; He et al. 2005) and has also been shown to be involved in secondary cell wall thickening (Kubo et al. 2005; Mitsuda et al. 2005, 2007). Ectopic expression of SND1, a NAC homolog, induces the deposition of secondary wall material in cells that normally do not have secondary wall thickenings (Zhong et al. 2006). This is consistent with the cell wall thickenings observed in syncytia and cells surrounding the syncytium as it collapses during I (Endo 1965, 1991; Riggs et al. 1973; Kim et al. 1987). Our MA also demonstrated that NAC probe sets are measuring suppressed transcriptional activity at 3 dpi in C. This is consistent with our previous observations of reduced transcript levels for NAC at 24 hpi during C (Alkharouf et al. 2006).
Glycine max experiences transcript expression unique to I at 8 dpi
By 8 dpi, G. max is undergoing substantial differences in gene expression between I and C. MA identified induced genes in I at 8 dpi that encoded LOX, HSP and genes involved in energy metabolism that were not present in C. These genes were induced to some extent at 3 dpi in I, but are more highly induced in I at 8 dpi. One of these HSPs was HSP90. The HSP90 chaperone complex is crucial for R gene mediated plant defense and other aspects of development (reviewed in Sangster and Queitsch 2005). The induction of HSPs also coincides with the impairment of mitochondrial function (Kuzmin et al. 2004). The induction of HSPs in mutants impaired for mitochondrial function, however, does not involve Ca+2 or ROS-dependent pathways (Karpova et al. 2002). Our MA identified probe sets measuring induced transcript levels of ClpB/HSP101 at 8 dpi. ClpB/HSP101 is crucial during recovery after severe cellular damage (Queitsch et al. 2000). Our MA also identified HSP70 probe sets measuring induced transcript levels at 8 dpi. HSP70 is important in preventing mitochondria from inducing both caspase-dependent and independent apoptosis (Saleh et al. 2000; Ravagnan et al. 2001). These results may put into context why, in Nicotiana sylvestris, nad7 mutants that delete NAD7 (Pla et al. 1995), and leading to an impaired mitochondrial function (Gutierres et al. 1997), experience a concomitant increase in resistance to viral infection and tolerance to O3 stresses (Dutilleul et al. 2003). Altered HSP70 expression affects root development (Sung and Guy 2003) whereby overexpression of HSP70 in A. thaliana results in shorter roots that are more branched (Sung and Guy 2003). Thus, the overexpression of HSP70 can be attributed to a root phenotype. The identification of these and several other HSPs by our MA are consistent with their role during recovery after cellular damage. It may also explain why HSP genes are induced at 8 dpi in I and not in C.
A core G. max response occurs irrespective of the nematode population over time
Experiments using microarrays to investigate plant gene expression during nematode infection have typically focused on C reactions (Puthoff et al. 2003; Alkharouf et al. 2004, 2006; Jammes et al. 2005; Ithal et al. 2007). In our experiments, we analyzed both the I and C reactions using microarrays and we analyzed these reactions that were occurring over time. Our MA identified a core set of probe sets that are measuring differential expression over time that are common between I and C. A large proportion of the genes we identified as differentially expressed in both I and C reactions were suppressed throughout the time points we selected for our analysis. This demonstrates that G. max responds to the presence of H. glycines by suppressing the transcription of many genes, irrespective of whether or not I or C is occurring. It is unclear if some of these genes (like PR1) contribute to I, because they are highly induced to similar levels during both I and C. Many genes were also suppressed in C reactions in A. thaliana infected with M. incognita (Jammes et al. 2005). A panel of genes was suppressed in C reactions during a C reaction in G. max cv. Williams (Ithal et al. 2007). Unlike our analysis, few genes appeared to be common between I and C reactions in S. lycopersicum cv. Motelle and cv. Moneymaker, respectively (Schaff et al. 2007).
Our MA identified differences in gene transcript abundance as measured by the probe sets on the microarrays. These differences in transcript abundance are present as early as 12 hpi (i.e. Fig. 10h). These observations occurring between I and C are similar to Lamb et al. (1992) and Tao et al. (2003). However, induction of gene expression does not appear to occur for genes encoding members of all defense pathways during H. glycines infection of G. max. Our results indicate that the use of multiple populations of nematodes to achieve I or C may be useful in terms of better understanding fundamental aspects of nematode pathogenicity in plants. This is because it is possible to identify differentially expressed genes that are common to I and C in roots infected with different populations of nematodes. The commonalties may represent the core response by the plant regardless of the nematode population. Some commonalities were recently observed during a time-course of I and C infection of S. lycopersicum by M. hapla and M. incognita, respectively (Schaff et al. 2007). Conversely, by using multiple nematode populations like our analyses have presented, it may be possible to identify genes that actually underlie I and C and make then distinct. Such observations were recently made by using different RKN species to achieve I and C reactions in different S. lycopersicum cultivars (Schaff et al. 2007).
Unique aspects of transcript expression in G. max over time during I and C
Our observations demonstrate that there is a core response by G. max roots during I and C reactions to H. glycines. Our results also indicate that important changes in gene expression are occurring before the nematode has had a chance to select their feeding site. Thus, G. max appears to be engaging unique I and C transcriptional programs to cope with I and C nematodes as early as their migration phase and before feeding site selection has occurred. Thus, G. max can distinguish between nematode populations before syncytium establishment occurs. It appears that much of the gene expression between whole I and C roots are different throughout infection. However, there are additional differences in gene expression occurring locally within the syncytia as they develop. These changes in gene expression make syncytia different from the surrounding I and C tissue (reviewed by Gheysen and Fenoll 2002; Klink et al. 2005). Experiments on isolated syncytium samples are clarifying these issues, especially from the standpoint of cell fusion events that are central to the formation of the syncytium Klink et al. 2007b). However, it is unclear what affect the homogenization of various cell types (i.e. cortex and pericycle) has on the gene expression during the formation of the syncytium.
Abbreviations
- EST:
-
Expressed sequence tag
- hpi:
-
Hours post inoculation
- dpi:
-
Days post inoculation
- SCN:
-
Soybean cyst nematode
- J2:
-
Second stage juvenile
- cv.:
-
Cultivar
- I:
-
Incompatible
- C:
-
Compatible
- NAM:
-
No apical meristem
- MA:
-
Microarray analysis
- CUC:
-
CUP-SHAPED COTYLEDON
- NAC:
-
NAM-ATAF-CUC
- HSP:
-
Heat shock protein
- MRS:
-
Moisture replacement system
- KTI:
-
Kunitz trypsin inhibitor
- OPR:
-
12-Oxophytodienoate reductase
- EDS1:
-
Enhanced disease susceptibility 1
- SAG101:
-
SENESCENCE-ASSOCIATED GENE101
- LOX:
-
Lipoxygenase
- NPR1:
-
Non expressor of PR genes 1
- LTP:
-
Lipid transfer protein
- PAD4:
-
PHYTOALEXIN-DEFICIENT4
- PIP1C:
-
Plasma membrane intrinsic protein 1C
- GER:
-
Germin-like protein
- HEL:
-
Hevein-like protein
- FLA6:
-
Fasciclin-like arabinogalactan-protein
- MPA:
-
Median polish algorithm
- RCC1:
-
Regulator of chromosome condensation
- PRR:
-
Pattern recognition receptors
- PAMP:
-
Pathogen-associated molecular pattern
- RKN:
-
Root knot nematode
References
Alkharouf N, Matthews BF (2004) SGMD: the soybean genomics and microarray database. Nucleic Acids Res 32:D398–D400
Alkharouf NW, Klink VP, Chouikha IB, Beard HS, MacDonald MH, Meyer S, Knap HT, Khan R, Matthews BF (2006) Timecourse microarray analyses reveals global changes in gene expression of susceptible Glycine max (soybean) roots during infection by Heterodera glycines (soybean cyst nematode). Planta 224:838–852
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Burlat V, Kwon M, Davin LB, Lewis NG (2001) Dirigent proteins and dirigent sites in lignifying tissues. Phytochemistry 57:883–897
Byrd DW Jr, Kirkpatrick T, Barker KR (1983) An improved technique for clearing and staining plant tissue for detection of nematodes. J Nematol 15:142–143
Cai D, Kleine M, Kifle S, Harloff HJ, Sandal NN, Marcker KA, Klein-Lankhorst RM, Salentijn EM, Lange W, Stiekema WJ, Wyss U, Grundler FM, Jung C (1997) Positional cloning of a gene for nematode resistance in sugar beet. Science 275:832–834
Cao H, Bowling SA, Gordon AS, Dong X (1994) Characterization of an Arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance. Plant Cell 6:1583–1592
Cho HJ, Farrand SK, Noel GR, Widholm JM (2001) High-efficiency induction of soybean hairy roots and propagation of the soybean cyst nematode. Planta 210:195–204
Collinge M, Boller T (2001) Differential induction of two potato genes, Stprx2 and StNAC, in response to infection by Phytophthora infestans and to wounding. Plant Mol Biol 46:521–529
Cregan PB, Mudge J, Fickus EW, Marek LF, Danesh D, Denny R, Shoemaker RC, Matthews BF, Jarvik T, Young ND (1999) Targeted isolation of simple sequence repeat markers through the use of bacterial artificial chromosomes. Theor Appl Genet 98:919–928
Dangl JL, Jones JD (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826–833
Dutilleul C, Garmier M, Noctor G, Mathieu C, Chetrit P, Foyer CH, de Paepe R (2003) Leaf mitochondria modulate whole cell redox homeostasis, set antioxidant capacity, and determine stress resistance through altered signaling and diurnal regulation. Plant Cell 15:1212–1226
Endo BY (1964) Penetration and development of Heterodera glycines in soybean roots and related and related anatomical changes. Phytopathology 54:79–88
Endo BY (1965) Histological responses of resistant and susceptible soybean varieties, and backcross progeny to entry development of Heterodera glycines. Phytopathology 55:375–381
Endo B (1971) Synthesis of nucleic acids at infection sites of soybean roots parasitized by Heterodera glycines. Phytopathology 61:395–399
Endo BY (1991) Ultrastructure of initial responses of resistant and susceptible soybean roots to infection by Heterodera glycines. Rev Nematol 14:73–94
Endo BY (1998) Atlas on ultrastructure of infective juveniles of the soybean cyst nematode Heterodera glycines. Agricultura handbook number 711, 224 pp
Endo BY, Veech JA (1970) Morphology and histochemistry of soybean roots infected with Heterodera glycines. Phytopathology 60:1493–1498
Feys BJ, Wiermer M, Bhat RA, Moisan LJ, Medina-Escobar N, Neu C, Cabral A, Parker JE (2005) Arabidopsis SENESCENCE-ASSOCIATED GENE101 stabilizes and signals within an ENHANCED DISEASE SUSCEPTIBILITY1 complex in plant innate immunity. Plant Cell 17:2601–2613
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Li FLC, Maeschler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JYH, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Gheysen G, Fenoll C (2002) Gene expression in nematode feeding sites. Annu Rev Phytopathol 40:191–219
Gipson I, Kim KS, Riggs RD (1971) An ultrastructural study of syncytium development in soybean roots infected with Heterodera glycines. Phytopathology 61:347–353
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for Arabidopsis lateral root development. Plant Cell 17:1376–1386
Gutierres S, Sabar M, Lelandais C, Chetrit P, Diolez P, Degand H, Boutry M, Vedel F, de Kouchkovsky Y, De Paepe R (1997) Lack of mitochondrial and nuclear-encoded subunits of complex I and alteration of the respiratory chain in Nicotiana sylvestris mitochondrial deletion mutants. Proc Natl Acad Sci USA 94:3436–3441
Hammond-Kosack KE, Jones JD (1997) Plant disease resistance genes. Annu Rev Plant Physiol Plant Mol Biol 48:575–607
He XJ, Mu RL, Cao WH, Zhang ZG, Zhang JS, Chen SY (2005) AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development. Plant J 44:903–916
Hwang CF, Bhakta AV, Truesdell GM, Pudlo WM, Williamson VM (2000) Evidence for a role of the N terminus and leucine-rich repeat region of the Mi gene product in regulation of localized cell death. Plant Cell 12:1319–1329
Hwang CF, Williamson VM (2003) Leucine-rich repeat-mediated intramolecular interactions in nematode recognition and cell death signaling by the tomato resistance protein Mi. Plant J 34:585–593
Irizarry R, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP (2003a) Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31:e15
Irizarry R, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003b) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Ithal N, Recknor J, Nettleston D, Hearne L, Maier T, Baum TJ, Mitchum MG (2007) Parallel genome-wide expression profiling of host and pathogen during soybean cyst nematode infection of soybean. Mol Plant Microbe Int 20:293–305
Jammes F, Lecomte P, de Almeida-Engler J, Bitton F, Martin-Magniette ML, Renou JP, Abad P, Favery B (2005) Genome-wide expression profiling of the host response to root-knot nematode infection in Arabidopsis. Plant J 44:447–458
Jones MGK, Northcote DH (1972) Nematode-induced syncytium-a multinucleate transfer cell. J Cell Sci 10:789–809
Jones MGK, Dropkin VH (1975) Scanning electron microscopy of syncytial transfer cells induced in roots by cyst-nematode. Physiol Plant Pathol 7:259–263
Jones JD, Dangl JL (2006) The plant immune system. Nature 444:323–329
Journot-Catalino N, Somssich IE, Roby D, Kroj T (2006) The transcription factors WRKY11 and WRKY17 act as negative regulators of basal resistance in Arabidopsis thaliana. Plant Cell 18:3289–3302
Jung C, Wyss U (1999) New approaches to control plant parasitic nematodes. Appl Microbiol Biotechnol 51:439–446
Karpova OV, Kuzmin EV, Elthon TE, Newton KJ (2002) Differential expression of alternative oxidase genes in maize mitochondrial mutants. Plant Cell 14:3271–3284
Khan R, Alkharouf N, Beard HS, MacDonald M, Chouikha I, Meyer S, Grefenstette J, Knap H, Matthews BF (2004) Resistance mechanisms in soybean: gene expression profile at an early stage of soybean cyst nematode invasion. J Nematol 36:241–248
Kim YH, Riggs RD, Kim KS (1987) Structural changes associated with resistance of soybean to Heterodera glycines. J Nematol 19:177–187
Klink VP, MacDonald M, Alkharouf N, Matthews BF (2005) Laser capture microdissection (LCM) and expression analyses of Glycine max (soybean) syncytium containing root regions formed by the plant pathogen Heterodera glycines (soybean cyst nematode). Plant Mol Biol 59:969–983
Klink VP, Overall CC, Matthews BF (2007a) Developing a systems biology approach to study disease progression caused by Heterodera glycines in Glycine max. Gene Regulat Syst Biol 2:17–33
Klink VP, Overall CC, Alkharouf N, MacDonald M, Matthews BF (2007b) Laser capture microdissection (LCM) and comparative microarray expression analysis of syncytial cells isolated from incompatible and compatible soybean (Glycine max) roots infected by the soybean cyst nematode (Heterodera glycines) (in press)
Kubo M, Udagawa M, Nishikubo N, Horiguchi G, Yamaguchi M, Ito J, Mimura T, Fukuda H, Demura T (2005) Transcription switches for protoxylem and metaxylem vessel formation. Genes Dev 19:1855–1860
Kuzmin EV, Karpova OV, Elthon TE, Newton KJ (2004) Mitochondrial respiratory deficiencies signal up-regulation of genes for heat shock proteins. J Biol Chem 279:20672–20677
Lamb CJ, Ryals JA, Ward ER, Dixon RA (1992) Emerging strategies for enhancing crop resistance to microbial pathogens. Biotechnology 10:1436–1445
Lipshutz RJ, Fodor SP, Gingeras TR, Lockhart DJ (1999) High density synthetic oligonucleotide arrays. Nat Genet (1 suppl):20–24
Liu X, Bai X, Wang X, Chu C (2006) OsWRKY71, a rice transcription factor, is involved in rice defense response. J Plant Physiol (in press)
Matthews BF, Devine TE, Weisemann JM, Beard HS, Lewers KS, MacDonald MH, Park Y-B, Maiti R, Lin J-J, Kuo J, Pedroni MJ, Cregan PB, Saunders JA (2001) Incorporation of sequenced cDNA and genomic markers into the soybean genetic map. Crop Sci 41:516–521
Matthews B, MacDonald MH, Thai VK, Tucker ML (2003) Molecular characterization of argenine kinase in the soybean cyst nematode (Heterodera glycines). J Nematol 35:252–258
Milligan SB, Bodeau J, Yaghoobi J, Kaloshian I, Zabel P, Williamson VM (1998) The root knot nematode resistance gene Mi from tomato is a member of the leucine zipper, nucleotide binding, leucine-rich repeat family of plant genes. Plant Cell 10:1307–1319
Mitsuda N, Seki M, Shinozaki K, Ohme-Takagi M (2005) The NAC transcription factors NST1 and NST2 of Arabidopsis regulate secondary wall thickenings and are required for anther dehiscence. Plant Cell 17:2993–3006
Mitsuda N, Iwase A, Yamamoto H, Yoshida M, Seki M, Shinozaki K, Ohme-Takagi M (2007) NAC transcription factors, NST1 and NST3, are key regulators of the formation of secondary walls in woody tissues of Arabidopsis. Plant Cell (in press)
Mujer CV, Andrews DL, Manhart JR, Pierce SK, Rumpho ME (1996) Chloroplast genes are expressed during intracellular symbiotic association of Vaucheria litorea plastids with the sea slug Elysia chlorotica. Proc Natl Acad Sci USA 93:12333–12338
Niblack TL, Arelli PR, Noel GR, Opperman CH, Orf JH, Schmitt DP, Shannon JG, Tylka GL (2002) A revised classification scheme for genetically diverse populations of Heterodera glycines. J Nematol 34:279–288
Oh SK, Yi SY, Yu SH, Moon JS, Park JM, Choi D (2006) CaWRKY2, a chili pepper transcription factor, is rapidly induced by incompatible plant pathogens. Mol Cells 22:58–64
Ohnishi T, Sugahara S, Yamada T, Kikuchi K, Yoshiba Y, Hirano HY, Tsutsumi N (2005) OsNAC6, a member of the NAC gene family, is induced by various stresses in rice. Genes Genet Syst 80:135–139
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci.10:79–87
Paal J, Henselewski H, Muth J, Meksem K, Menendez CM, Salamini F, Ballvora A, Gebhardt C (2004) Molecular cloning of the potato Gro1-4 gene conferring resistance to pathotype Ro1 of the root cyst nematode Globodera rostochiensis, based on a candidate gene approach. Plant J 38:285–297
Pla M, Mathieu C, De Paepe R, Chetrit P, Vedel F (1995) Deletion of the last two exons of the mitochondrial nad7 gene results in lack of the NAD7 polypeptide in a Nicotiana sylvestris CMS mutant. Mol Gen Genet 248:79–88
Puthoff DP, Nettleton D, Rodermel SR, Baum TJ (2003) Arabidopsis gene expression changes during cyst nematode parasitism revealed by statistical analyses of microarray expression profiles. Plant J 33:911–921
Queitsch C, Hong SW, Vierling E, Lindquist S (2000) Heat shock protein 101 plays a crucial role in thermotolerance in Arabidopsis. Plant Cell 12:479–492
Ralph S, Park JY, Bohlmann J, Mansfield SD (2006) Dirigent proteins in conifer defense: gene discovery, phylogeny, and differential wound- and insect-induced expression of a family of DIR and DIR-like genes in spruce (Picea spp.). Plant Mol Biol 60:21–40
Ravagnan L, Gurbuxani S, Susin SA, Maisse C, Daugas E, Zamzami N, Mak T, Jaattela M, Penninger JM, Garrido C, Kroemer G (2001) Heat-shock protein 70 antagonizes apoptosis-inducing factor. Nat Cell Biol 3:839–843
Riggs RD, Schmitt DP (1991) Optimization of the Heterodera glycines race test procedure. J Nematol 23:149–154
Riggs RD, Kim KS, Gipson I (1973) Ultrastructural changes in Peking soybeans infected with Heterodera glycines. Phytopathology 63:76–84
Ryu HS, Han M, Lee SK, Cho JI, Ryoo N, Heu S, Lee YH, Bhoo SH, Wang GL, Hahn TR, Jeon JS (2006) A comprehensive expression analysis of the WRKY gene superfamily in rice plants during defense response. Plant Cell Rep 25:836–847
Saleh A, Srinivasula SM, Balkir L, Robbins PD, Alnemri ES (2000) Negative regulation of the Apaf-1 apoptosome by Hsp70. Nat Cell Biol 2:476–483
Sangster TA, Queitsch C (2005) The HSP90 chaperone complex, an emerging force in plant development and phenotypic plasticity. Curr Opin Plant Biol 8:86–892
Sardanelli S, Kenworthy WJ (1997) Soil moisture control and direct seeding for bioassay of Heterodera glycines on Soybean. J Nematol (Suppl) 29:625–634
Sass JE (1958) Botanical microtechnique. Iowa State College Press, Ames
Schaff JE, Nielsen DM, Smith CP, Scholl EH, Bird DMck (2007). Comprehensive transcriptome profiling in tomato reveals a role for glycosyltransferase in Mi-mediated nematode resistance. Plant Physiol (in press)
Scheideler M, Schlaich NL, Fellenberg K, Beissbarth T, Hauser NC, Vingron M, Slusarenko AJ, Hoheisel JD (2001) Monitoring the switch from housekeeping to pathogen defense metabolism in Arabidopsis thaliana using cDNA arrays. J Biol Chem 277:10555–10561
Shen QH, Saijo Y, Mauch S, Biskup C, Bieri S, Keller B, Seki H, Ulker B, Somssich IE, Schulze-Lefert P (2007) Nuclear activity of MLA immune receptors links isolate-specific and basal disease-resistance responses. Science 315:1098–1103
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3:Article3
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85:159–170
Sung DY, Guy CL (2003) Physiological and molecular assessment of altered expression of Hsc70-1 in Arabidopsis. Evidence for pleiotropic consequences. Plant Physiol 132:979–987
Tao Y, Xie Z, Chen W, Glazebrook J, Chang HS, Han B, Zhu T, Zou G, Katagiri F (2003) Quantitative nature of Arabidopsis responses during C and I interactions with the bacterial pathogen Pseudomonas syringae. Plant Cell 15:317–330
The Gene Ontology Consortium (2004) The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res 32:D258–D261
Ulker B, Shahid Mukhtar M, Somssich IE (2007) The WRKY70 transcription factor of Arabidopsis influences both the plant senescence and defense signaling pathways. Planta (in press)
Vroemen CW, Mordhorst AP, Albrecht C, Kwaaitaal MA, de Vries SC (2003) The CUP-SHAPED COTYLEDON3 gene is required for boundary and shoot meristem formation in Arabidopsis. Plant Cell 15:1563–1577
Wang D, Amornsiripanitch N, Dong X (2006) A genomic approach to identify regulatory nodes in the transcriptional network of systemic acquired resistance in plants. PLoS Pathog 2:e123
Williamson VM, Kumar A (2006) Nematode resistance in plants: the battle underground. Trends Genet 22:396–403
Wrather JA, Koenning SR (2006) Estimates of disease effects on soybean yields in the United States 2003–2005. J Nematol 38:173180
Xu X, Chen C, Fan B, Chen Z (2006) Physical and functional interactions between pathogen-induced Arabidopsis WRKY18, WRKY40, and WRKY60 transcription factors. Plant Cell 18:1310–1326
Zabala G, Zou J, Tuteja J, Gonzalez DO, Clough SJ, Vodkin LO (2006) Transcriptome changes in the phenylpropanoid pathway of Glycine max in response to Pseudomonas syringae infection. BMC Plant Biol 6:26
Zhang C, Ghabrial SA (2006) Development of Bean pod mottle virus-based vectors for stable protein expression and sequence-specific virus-induced gene silencing in soybean. Virology 344:401–411
Zheng Z, Qamar SA, Chen Z, Mengiste T (2006) Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J 48:592–605
Zhong R, Demura T, Ye ZH (2006) SND1, a NAC domain transcription factor, is a key regulator of secondary wall synthesis in fibers of Arabidopsis. Plant Cell 18:3158–3170
Zou J, Rodriguez-Zas S, Aldea M, Li M, Zhu J, Gonzalez DO, Vodkin LO, DeLucia E, Clough SJ (2005) Expression profiling soybean response to Pseudomonas syringae reveals new defense-related genes and rapid HR-specific downregulation of photosynthesis. Mol Plant Microbe Interact 18:1161–1174
Acknowledgments
The authors thank Dr. David Munroe and Nicole Lum at the Laboratory of Molecular Technology, SAIC-Frederick, National Cancer Institute at Frederick, Frederick, MD 21701, USA for the Affymetrix® array hybridizations and data acquisition. All data, raw and normalized, are stored in the Soybean Genomics and Microarray Database (Alkharouf et al. 2004), available through the web site (http://www.towson.edu/nalkharo/SGMD/SupplementalSites/GmWholeRootIC/). The authors greatly appreciate the continued support provided by the United Soybean Board under grant 5214. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the United States Department of Agriculture. The authors greatly appreciate careful reading and critical comments of the manuscript provided by Dr. Leslie Wanner, United States Department of Agriculture. The authors thank Veronica Martins for careful editing of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Vincent P. Klink and Christopher C. Overall have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Klink, V.P., Overall, C.C., Alkharouf, N.W. et al. A time-course comparative microarray analysis of an incompatible and compatible response by Glycine max (soybean) to Heterodera glycines (soybean cyst nematode) infection. Planta 226, 1423–1447 (2007). https://doi.org/10.1007/s00425-007-0581-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-007-0581-4