Abstract
The cDNA for the granule-bound starch synthase (GBSS; ADP-glucose-starch glucosyltransferase, EC 2.4.1.21) of Chlorella kessleri 11 h was isolated and characterized. CkGBSS encodes a 609-amino acid polypeptide (65,627 Da) that includes an N-terminal hydrophobic signal peptide of 55 amino acids. The deduced amino acid sequence of the mature CkGBSS polypeptide shares a greater identity (65%) to that of the GBSS protein of Chlamydomonas reinhardtii, than to those of vascular plant species, but does not have the extra-long C-terminal sequence found in C. reinhardtii. When CO2 concentration was decreased from 3 to 0.04% (air level) in light, the levels of CkGBSS mRNA, CkGBSS protein, and GBSS activity increased. Under this condition, pyrenoid and pyrenoid starch developed, and the relative amount of amylose in starch increased. These observations suggest that low CO2 level up-regulates GBSS biosynthesis at the transcriptional level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Green plants (including vascular plants and green algae) accumulate α-polyglucans in their plastids in the form of starch. Starch consists of two types of α-polyglucans, amylose and amylopectin, which have distinctive structures. Amylose is lightly α-1,6-branched linear α-1,4-polyglucans, while amylopectin has a defined structure composed of tandem linked clusters where linear α-1,4-glucan chains are highly and regularly branched via α-1,6-glucosidic linkages. In vascular plants, amylose accounts for about 15–30% of storage starch.
Starch synthases are enzymes that catalyze the elongation of α-polyglucan chains by transferring the glucose moiety of ADP-glucose to α-polyglucan chains via an α-1,4-glucosidic linkage. Granule-bound starch synthase (GBSS), one of the starch synthases, is considered to be responsible mainly for amylose biosynthesis. A lot of mutants lacking GBSS have been isolated and characterized, e.g., waxy rice (Murata et al. 1965), waxy wheat (Nakamura et al. 1995), and sta2 Chlamydomonas reinhardtii (Delrue et al. 1992). In these mutants, amylose is not synthesized, and therefore, the starch granules are not stained blue–violet with iodine. Vascular plants have two GBSS isoforms, GBSSI which is involved in amylose synthesis in storage tissues (for review see Smith et al. 1997) and GBSSII which is responsible for amylose synthesis in non-storage tissues such as leaves and stems (Denyer et al. 1997; Fujita and Taira 1998; Nakamura et al. 1998; Vrinten and Nakamura 2000; Edwards et al. 2002; Dian et al. 2003).
Unlike vascular plants, algae and cyanobacteria contain a wide variety of storage polyglucans (Manners and Sturgeon 1982; Smith et al. 1997; Ball 1998; Nakamura et al. 2005). In this respect, Chlorophyta (green algae) is an evolutionally primary taxon that accumulates starch including both amylose and amylopectin, similar to vascular plants. The molecular structure of starch and the enzymes involved in starch synthesis, including GBSS, have been investigated intensively in Chlamydomonas (for review see Ball 1998). The GBSS activity in Chlamydomonas starch is an order of magnitude higher than those reported for most vascular plants, and the GBSS cDNA contains an extra C-terminal sequence encoding an 11.4 kDa polypeptide that is absent in vascular plant GBSS (Wattebled et al. 2002).
In our previous studies on the effects of CO2 concentration during growth in microalgae, we investigated the roles of carbonic anhydrase in CO2-concentrating mechanism (CCM) of microalgae and cyanobacteria grown under low-CO2 conditions (Tsuzuki and Miyachi 1989; Fujiwara et al. 1990, 1996; Fukuzawa et al. 1990a, b). In the series of this research, we have also observed the development of pyrenoid and starch around the pyrenoid. To our knowledge, there is no report thus far on the mechanisms of the biogenesis of the pyrenoid starch.
In the present study, we isolated the cDNA for GBSS in Chlorella kessleri 11 h (CkGBSS cDNA), and analyzed its structure and expression pattern as influenced by CO2 concentration. Interestingly, the level of CkGBSS mRNA and the amylose content in the organism increased under low-CO2 condition along with the development of the pyrenoid starch, but were unaffected either in the dark or after the addition of NH +4 , which is known to stimulate starch degradation in Chlorella (Miyachi and Miyachi 1985, 1987). The relationship between GBSS activity under low-CO2 condition and development of the pyrenoid starch is discussed.
Materials and methods
Algal cells and culture conditions
Chlorella kessleri 11 h cells (C-531; IAM Culture Collection, University of Tokyo, Japan) were cultured at 29°C in 0.2×Gamborg’s B5 medium (Gamborg et al. 1968; Nihon Pharmaceutical Co. Ltd., Japan) under continuous illumination at 10 W m−2 by aeration with air containing 3% CO2 to obtain high-CO2 cells. The bubbling gas was changed to ordinary air (0.04% CO2) to obtain low-CO2 cells.
Chlamydomonas reinhardtii CC125 (kindly provided by Kosuke Shimogawara, Teikyo University, School of Medicine, Tokyo, Japan), Scenedesmus obliquus IAM C-538, and Dunaliella tertiolecta IAM C-524 cells (IAM Culture Collection) were cultured at 29°C with constant bubbling of air containing 3% CO2 in 0.3×HSM medium (Sueoka et al. 1967), proteose medium (Starr 1964), and an inorganic medium for Dunaliella (Aizawa and Miyachi 1984), respectively.
Cloning of the CkGBSS cDNA
To obtain a cDNA fragment coding for Chlorella GBSS, we performed nested reverse transcriptional polymerase chain reaction (RT-PCR), using the following pairs of degenerated primers: GBSS-F1 (5′-AAGACCGG(C/T)GG(C/T)CT(C/G)GG(C/T)GA-3′) and GBSS-R1 (5′-GGTGTC(C/G)AC(C/G)AG(A/G)CC(A/G)CCGG-3′); and GBSS-F2 (5′-GG(C/T)AT(C/T)GT(C/G)AACGG(C/T)ATGGA-3′) and GBSS-R2 (5′-(C/G)AG(A/G)CCGCAGGGCTC(A/G)AAGC-3′), all corresponding to the amino acid sequences of conserved domains of GBSS proteins (Cao et al. 1999). An amplified cDNA fragment of 420 bp (named GBSS-Pr) was cloned in pGEM-T Easy (Promega, WI, USA) and sequenced using an ABI PRISM 3100 Genetic Analyzer (Perkin-Elmer Applied Bio-Systems, Warrington, UK). The amino acid sequence deduced from the nucleotide sequence shares high homology with those of GBSS from other green plants. Using the fragment as a probe, about 5×105 plaques of a Chlorella λZAPII cDNA library prepared using cDNA synthesized with an oligo d(T) primer and a mixture of poly (A)+ RNA from high-CO2 cells and air-grown cells were screened using standard procedures (Sambrook and Russell 2001), yielding ten positive clones, all having about 1.6 kb inserts. The cDNA clone with a 1,607 bp insert (named GBSS-1-9) was sequenced on both strands but was found not to include a full-length cDNA. Thus, about 5×105 plaques of Chlorella λZAPII cDNA library, which was constructed with random-primed cDNA synthesized using a mixture of poly (A)+ RNA from high-CO2 cells and air-grown cells as a template, were further screened to obtain the 5′ end of the cDNA. Among the 19 positive clones obtained, the one with the longest insert (2,191 bp; named GBSS-2-9) was fully sequenced on both strands. The complete cDNA obtained (2,664 bp) was submitted to GenBank, and was assigned the accession number AB232549.
RNA-blot hybridization
Ten microgram each of total RNA was electrophoresed in formaldehyde-containing 1% agarose gels (Sambrook and Russell 2001), capillary blotted to nylon membrane (Zeta-probe; Bio-Rad Laboratories, Hercules, CA), and probed with the 32P-labeled cDNA fragment GBSS-Pr. RNA size standard marker II (Wako Chemicals Inc., USA) was used as size markers.
Measurement of starch levels
Cells at the log phase in about 1 ml of a culture were harvested by centrifugation (12,000g, 10 min, 4°C) and suspended in 100% ethanol. After boiling for 30 min, the suspension was centrifuged (15,000g, 10 min, 4°C), and then the pellet was dried up. Starch was extracted by boiling (for 30 min) and sonicated in 1 ml of 0.2 N KOH. The pH of the extract was adjusted to 5.5 with 0.2 ml of 1 N acetic acid. The starch levels were determined using the amyloglucosidase assays (Delrue et al. 1992). Cells were counted using a hemacytometer.
Purification of starch granules
Starch granules were prepared from C. kessleri, C. reinhardtii, S. obliquus, and D. tertiolecta cells according to Delrue et al. (1992) with some modifications. Briefly, cells were harvested and suspended in an appropriate volume of TE buffer [10 mM Tris–HCl (pH 8.0), 1 mM EDTA], and then disrupted by passing through the French Pressure Cell (5500H; Ohtake Works Co. Ltd., Japan) at 1,500 kg cm−2 twice. The crude starch pellet obtained by centrifugation (10,000g, 20 min, 4°C) was resuspended in TE buffer, and then passed twice through a Percoll (Amersham Pharmacia Biotech, Chalfont, UK) gradient (1.2 ml of Percoll per 0.3 ml of crude starch suspension). The purified starch pellet was washed by distilled water.
Molecular-size separation of starch polysaccharides by Sephacryl S-1000SF chromatography
Twenty microgram of the purified starch was washed with ethanol, dried, and suspended in 2 ml of 1 N NaOH. After 30 min at room temperature, 2 ml of distilled water was added to the sample suspension, and then filtrated through a cellulose acetate membrane (pore size, 0.80 μm). Three milliliter of the filtrate was applied onto a Sephacryl S-1000SF (Amersham Pharmacia Biotech) column (2.0 cm diameter; 60 cm long) previously equilibrated with 0.1 N NaOH containing 0.2% (w/v) NaCl. The sample was eluted with the same solution at a flow rate of about 0.14 ml min−1 at room temperature. Fractions were taken at 4 ml intervals. Pullulan standards P-82 (Shodex, Japan) were used as size markers.
For measurement of the total carbohydrate content by the phenolic sulfuric method (Hodge and Hofreiter 1962), 20 μl of each sample was taken and added successively to 4 μl 0.5 N HCl, 36 μl distilled water, 5 μl 80% (v/v) phenol, and 160 μl sulfuric acid. The absorbance of the mixture was measured at 490 nm.
For determination of the λmax value of the polysaccharide–I2 complex, 400 μl of each sample was mixed with 100 μl 0.5 N HCl and 50 μl 0.1% I2/1% KI, and the absorption spectrum of the resulting solution was obtained in the 450–700 nm range.
SDS-PAGE of starch granule-bound proteins and amino acid sequencing
Starch granule-bound proteins were extracted according to Taira et al. (1995). Briefly, 3–7 mg of purified starch granules was suspended in 80 μl of the sample buffer [50 mM Tris–HCl (pH 6.8), 2% (w/v) SDS, 6% (v/v) 2-mercaptoethanol, 10% (w/v) glycerol] and extracted by boiling for 5 min. The supernatant obtained after centrifugation (10,000g, 20 min, room temperature) was separated by electrophoresis on 10% SDS polyacrylamide gels, followed by staining with Coomasie Brilliant Blue, as previously described (Laemmli 1970).
A starch granule-bound protein in C. kessleri was recovered from the polyacrylamide gel by electroelution, and sequenced using a peptide sequencer (476A; Perkin-Elmer Applied Biosystems, Warrington, UK).
In vitro assays of GBSS and SS activities
Granule-bound starch synthase activity was measured according to Delrue et al. (1992) as follows. About 50 μg of starch granules was incubated at 30°C for 30 min in 100 μl of GBSS reaction buffer [50 mM glygly-NaOH (pH 9.0), 100 mM (NH4)2SO4, 5 mM 2-mercaptoethanol, 5 mM MgCl2, 0.25g l−1 BSA, 3.2 mM ADP-glucose, and 0.75 nmol [U-14C] ADP-glucose (1.2×1010 Bq mmol−1)]. The reaction was stopped by the addition of 2 ml of 70% ethanol. The precipitate was filtered on a glass-fiber filter (Whatmann GF/C), rinsed with 15 ml of 70% ethanol and dried at room temperature. The incorporated 14C was counted in a liquid scintillation counter.
Soluble starch synthase activity was assayed as described by Delrue et al. (1992). Protein concentrations were determined according to the method of Bradford (1976) with BSA as the standard, using the Coomasie Protein Assay Reagent (Pierce Chemicals, TX, USA).
In the assays of GBSS and SS activities, it was confirmed that the incorporation of 14C increased linearly with time (at least up to 30 min) and the amount of starch (at least up to 400 μg wet weight) or protein applied (for the SS assay, crude extracts including SS protein, which were diluted appropriately were used). In both the assays, background labeling when the enzyme had been inactivated by boiling was negligible (it was almost the same as that without the enzyme).
Results and discussion
Isolation and characterization of the CkGBSS cDNA
To prepare the probe for screening CkGBSS cDNA clones, we carried out RT-PCR using pairs of degenerate primers, which were designed based on the conserved amino acid sequence motifs of the GBSSI proteins from rice, wheat, barley, maize, sorghum, pea, and potato (Cao et al. 1999). The amino acid sequence deduced from the nucleotide sequence of an amplified cDNA fragment (420 bp) exhibited high similarity to those of GBSS from other plant species. Using the RT-PCR product (GBSS-Pr) as probe, CkGBSS cDNA clones were isolated from the oligo d(T)-primed and random-primed cDNA libraries of C. kessleri 11 h as is written in Materials and Methods. The complete CkGBSS cDNA obtained (2,664 bp) contains an open reading frame of 1,827 bp encoding a polypeptide of 609 amino acids with a calculated molecular mass of 66 kDa.
Since starch granule-bound proteins are not extracted by conventional methods, the proteins of C. kessleri were extracted by boiling in the presence of 2% SDS and analyzed by SDS-PAGE, as described previously in rice endosperm (Taira et al. 1995). SDS-PAGE of the starch granule-bound proteins in high-CO2 cells of C. kessleri revealed that the purified starch granules contain a single polypeptide of 57 kDa (Fig. 1, lane 1) that was assumed to be GBSS, since the purified starch granules exhibited starch synthase activity (Table 1). N-terminal sequencing of the GBSS polypeptide predicted a transit peptide of 55 amino acids.
In low-CO2 cells of C. kessleri, minor bands of polypeptides, which were considered to be subunits of Rubisco, judging from the apparent Mr and the Western blotting analysis using Rubisco antibody [anti-RbcL global (Rubisco); AgriSera], were also detected (data not shown). This may suggest that Rubisco is actually bound to pyrenoid starch. Alternatively, this may be due to the involvement of pyrenoid components into the pyrenoid starch fraction, since pyrenoid includes a large amount of Rubisco.
In endosperm of cereals such as rice and wheat, SS and starch-branching enzyme (BE) are partly bound to starch granules. In SDS-PAGE of the starch granule-bound proteins of rice endosperm, minor bands of SS and BE with larger molecular weights appeared in addition to a major band of GBSS in the vicinity of 60 kDa, as reported previously (Taira et al. 1995; see also Dian et al. 2003). In contrast, in the green algae including Chlorella, such minor bands with larger molecular weights could not be detected with the extracts obtained from almost the same amounts of starch as that of rice endosperm, suggesting that SS and BE are apparently not bound to starch granules (Fig. 1; Taira et al. 1995).
Amino acid sequence alignment of GBSS polypeptides
The deduced amino acid sequence of GBSS from C. kessleri (Trebouxiophyceae) was aligned with those of GBSSI from vascular plants and two other green algal species, C. reinhardtii (Chlorophyceae) and Ostreococcus tauri (Prasinophyceae) (Fig. 2). The eight conserved regions present in all plant GBSS (Cao et al. 1999), were identified. Conserved regions 1 and 8 contain the KTGGL motif, which is considered to be the ADP-glucose binding site (Furukawa et al. 1993), and the KTGGL look-alike motif (Edwards et al. 2002), respectively. The deduced amino acid sequence of the mature CkGBSS polypeptide shared higher identity with those of the GBSS from Chlamydomonas (65%) and Ostreococcus (59%) than with the GBSSI and GBSSII sequences of vascular plants (37–53%) (identities were obtained by BLASTP, in which N- and C-terminal ambiguous alignments were excluded). However, CkGBSS does not have such an extra, long C-terminal sequence found in C. reinhardtii (Wattebled et al. 2002). To confirm the difference in the molecular size between the GBSS polypeptides, SDS-PAGE of starch granule-bound proteins in C. reinhardtii (Chlorophyceae, Volvocales) and C. kessleri was performed (Fig. 1, lanes 1 and 2). The size of the mature protein in C. kessleri (57 kDa), as inferred from SDS-PAGE, was expectedly smaller than that in C. reinhardtii (74 kDa). As a reference, starch granule-bound proteins of the green algae D. tertiolecta (Chlorophyceae, Volvocales) and S. obliquus (Chlorophyceae, Sphaeropleales) were also analyzed, and their sizes were inferred from SDS-PAGE as 75 and 57 kDa, respectively (Fig. 1, lanes 3 and 4). These suggest that some species of the Volvocales in the Chlorophyceae might have an extra-long C-terminal sequence.
Amino acid sequence alignment of CkGBSS and GBSSI of other plant species. The sequences were aligned using the CLUSTAL W program, available in DDBJ. Identical and conserved residues are marked with (*) and (.:), respectively. Solid lines and numerals I to VIII indicate conserved regions. The N-terminal amino acids of the mature polypeptides are in bold case. The sequences and their accession numbers are: C. kessleri (AB232549), C. reinhardtii (AF026420), O. tauri (AY570711), potato (X58453), Arabidopsis (AC006424), maize (X03935), and rice (X53694)
A molecular phylogenetic tree of GBSS proteins was constructed by the neighbor-joining method, using the alignment of amino acid sequences trimmed of their ambiguously aligned N- and C-terminal regions (Fig. 3). The tree suggests that the GBSS proteins from green algae form a relatively reliable monophyletic group that could be a sister group of the vascular plant GBSS proteins. The GBSSI and GBSSII proteins in monocots seem to comprise relatively stable monophyletic groups, respectively. These findings suggest that GBSS diverged into GBSSI and GBSSII after vascular plants appeared, probably after monocots and dicots diverged but before the monocot lineage developed.
A molecular phyologenetic tree of GBSS. The tree was constructed by the neighbor-joining method of the CLUSTAL W program, using amino acid sequences with ambiguously aligned N- and C-terminal sequences removed. Stability of monophyletic groups on the obtained tree was estimated with a bootstrap analysis for 10,000 replicates. Bootstrap values are indicated above the internal branches for values greater than 70%. The sequences and their accession numbers, in addition to those in Fig. 2, are as follows: wheat GBSSI (AB019623), wheat GBSSII (AF109395), barley GBSSI (AF486517), barley GBSSII (AF486521), rice GBSSII (AY069940), pea GBSSIa (AJ345045), and pea GBSSIb (X88789)
Effects of CO2 concentration on starch composition and CkGBSS gene expression
To investigate the effects of CO2 concentration on starch composition, starch granules were prepared from cells cultured initially with air containing 3% CO2 (high-CO2 cells) and transferred to the low-CO2 condition (0.04% CO2) or maintained in the high-CO2 condition for 1–2 d. Molecular size separation of starch α-polysaccharides was performed using Sephacryl S-1000SF gel permeation chromatography (Fig. 4). The elution profile demonstrated two starch fractions: one eluted at around 50 ml, and the other around 102 ml, which had molecular weights of ca. 1×107 and 1×105 Da, and polysaccharide-I2 complex λmax of ca. 560 and 610–620 nm, respectively. To confirm that the constituents of the two fractions are amylopectin and amylose, respectively, starch was treated with isoamylase (ISA), which digests α-1,6-glucosidic linkages, and subjected to Sephacryl S-500SF gel permeation chromatography (data not shown). The higher molecular weight fraction disappeared after ISA treatment, and a new peak appeared at less than 5×104 Da, in which the column was over the detectable range for molecular size analysis, indicating that most of the higher molecular weight constituents was amylopectin containing more α-1,6-glucosidic linkages. On the other hand, the lower molecular weight fraction was hardly altered by the treatment, suggesting that the lower molecular weight constituents are mainly amylose. Iodine affinities for amylopectin (λmax of ca. 560 nm) and amylose (λmax of 610–620 nm) were almost identical to those for storage starches of vascular plants and C. reinhardtii (Ball 1998).
Typical Sephacryl S-1000SF chromatograms of starch from high-CO2 cells and air-adapted cells. Starch granules from high-CO2 cells (X) and the cells transferred to the low-CO2 condition for 1 d (closed circles) and 2 d (closed triangles) or kept in the high-CO2 condition for 1 d (open circles) and 2 d (open triangles) were analyzed. Relative A595 value of the polysaccharide–I2 complex of each fraction was plotted. Identical trends were observed in another experiment
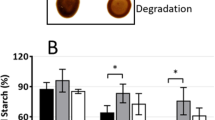
Comparison of the starch composition of C. kessleri 11 h showed that the relative amount of amylose (80–120 ml of the eluate) in starch of 2 d-air-adapted cells (28%) was higher than in high-CO2 cells (9–14%; Fig. 4), suggesting that amylose synthesis increased in the low-CO2 treatment that also induced the development of both the pyrenoid and the pyrenoid starch (data not shown, also see Miyachi et al. 1986). The relative amount of amylose in starch of the 2 d-air-adapted cells (28%) was relatively high, even in comparison with those of vascular plants and C. reinhardtii (15–30%; Ball 1998).
Because CO2 concentration affected the intracellular amylose content, the expression of GBSS, which is considered to be mainly responsible for amylose synthesis, was investigated. Northern hybridization demonstrated that the level of CkGBSS mRNA rapidly increased and peaked within 4 h and then decreased gradually, when high-CO2 cells at the log phase were transferred to the low-CO2 condition (Fig. 5a). This expression pattern was similar to those of genes known to be involved in the CCM in C. reinhardtii, e.g., CAH1, a gene for carbonic anhydrase (Fujiwara et al. 1990).
a, b Northern blot analysis of total RNA from C. kessleri cells. In each lane, 10 μg of total RNA was electrophoresed in a denaturing agarose gel, blotted to membranes, and probed with the 32P-labeled cDNA fragment. a Air induction of CkGBSS transcript in Chlorella cells under continuous illumination. High-CO2 cells (lane 1) were transferred to the low-CO2 condition for 4 (lane 2), 8 (lane 3), 12 (lane 4), and 24 h (lane 5) or kept in the high-CO2 condition for 24 h (lane 6) in light. b Effect of light, NH +4 -addition and Fe-starvation on CkGBSS transcript abundance in high-CO2 cells. High-CO2 cells were kept in the high-CO2 condition for 4 h in light (lane 1) and in dark (lane 4) and in the presence of light and 0.4 mM NH +4 (lane 2) or transferred to the low-CO2 condition for 4 h in light (lane 3). High-CO2 cells were centrifuged and transferred into the fresh Fe-replete medium (normal 0.2× Gamborg’s B5 medium; lane 5) or the Fe-starved medium in the high-CO2 condition for 4 h in light (lane 6). The molecular size was calibrated with the RNA size marker (Wako Chemicals). Identical trends were observed in another experiment
CkGBSS gene expression was also investigated under several starch-accumulating and starch-degrading conditions as a reference (Fig. 5b). Under Fe-starvation, a starch-accumulating condition, the level of CkGBSS mRNA was hardly altered for at least 4 h, suggesting that GBSS biosynthesis is not stimulated, unlike in the low-CO2 condition. CkGBSS transcript abundance was barely affected by high-CO2 treatment either in the presence of NH +4 (0.4 mM) or by dark treatment for at least 4 h, in which starch was degraded (Miyachi and Miyachi 1985, 1987).
To further investigate the effects of CO2 concentration on the level of CkGBSS activity, we determined the intracellular amounts and activity of GBSS using 15 h-air-adapted cells (Fig. 6 and Table 1). The Chlorella cells are fully adapted to the low-CO2 concentration after 15 h: it has been shown that the rate of photosynthetic fixation in the presence of a rate-limiting concentration of CO2 increases to cause two to threefold enhancement after about 5 h, and thereafter, it remains almost constant (Hogetsu and Miyachi 1979). The starch content per cell in the 15 h-air-adapted cells was 3.8-fold higher than that in the high-CO2 cells (Table 1). Since the relative amount of amylose in starch did not significantly increase a day after the transfer to the low-CO2 condition but increased drastically by the end of the second day (Fig. 4), it was considered that amylose and amylopectin increased almost at comparable rates during the early half of the first day, and then amylose synthesis exceeded that of amylopectin on the second day. SDS-PAGE of starch granule-bound proteins demonstrated that the amount of GBSS per starch amount was more than twofold higher in the 15 h-air-treated cells than that in high-CO2 cells (Fig. 6). Similarly, the activity of GBSS per starch amount in the 15 h-air-adapted cells was 2.7-fold higher than that in high-CO2 cells (Table 1). These suggest that GBSS biosynthesis is enhanced by low-CO2 concentration at the transcriptional level, when pyrenoid starch develops (in addition to stroma starch, well-developed pyrenoid starch has been observed under this condition by transmission electron microscopy, Miyachi et al. 1986). On the other hand, the activity of SS per protein amount in low-CO2 cells was almost the same as that in high-CO2 cells. These suggest the possibility that, among starch synthases, GBSS may play some important roles in pyrenoid starch synthesis under low-CO2 conditions.
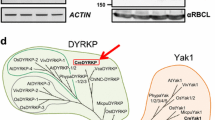
SDS-PAGE pattern of proteins bound to starch granules of high-CO2 cells (lanes 1, 3) and 15 h-air-adapted cells (lanes 2, 4) in C. kessleri (lanes 1, 2) and C. reinhardtii (lanes 3, 4). In each lane, one-fourth volume of protein extract from 3 mg starch was electrophoresed. The apparent Mr of the polypeptides were determined using protein markers from Bio-Rad Laboratories (Precision Protein Standards). Identical trends were observed in another experiment
As a reference, a similar experiment was performed in C. reinhardtii, which is also known to develop the pyrenoid when CO2 level is low (Kuchitsu et al. 1988). In C. reinhardtii, total starch content per cell either increased or decreased under the low-CO2 condition depending on the starch content, because the content of stroma starch decreased and that of pyrenoid starch increased under this condition (Kuchitsu et al. 1988, see also Table 1). On the other hand, in C. kessleri 11 h, total starch content always increased under the low-CO2 condition, irrespective of starch content (a typical datum shown in Table 1), suggesting that stroma starch content is relatively low in C. kessleri. GBSS activity increased during the air adaptation in both C. kessleri and C. reinhardtii, although the ratio of GBSS activity in air-adapted cells to that in high-CO2 cells was higher in C. kessleri under the experimental conditions employed in this study. Recently, CO2-responsive genes have been identified based on the expression profiling, using a cDNA macroarray of Chlamydomonas (Miura et al. 2004). Among 51 low-CO2 induced genes, Sta2 coding for GBSS has been found. Our data further demonstrated that the increased transcripts are actually reflected by GBSS activity in Chlamydomonas and in Chlorella.
The induction of the CCM is considered to be correlated with the formation of the pyrenoid starch (Ramazanov et al. 1994). In the present study, the expression pattern of CkGBSS was similar to those of the CCM-related genes in C. reinhardtii (Fig. 5a; e.g., Fujiwara et al. 1990), suggesting that GBSS might participate in the regulational network of the CCM-related genes. The macroarray experiments of Chlamydomonas showed that Sta2 encoding GBSS is controlled by a master regulator of CCM-related genes, Ccm1 (Miura et al. 2004). These results indicate that GBSS is probably one of the CCM-related genes which are regulated through putative low-CO2 signal transduction pathway, and that the product is possibly involved in the development of pyrenoid starch and CCM in green microalgae including Chlorella and Chlamydomonas.
In vascular plants, GBSSI is regulated by circadian rhythm (Mérida et al. 1999; Wang et al. 2001). The regulation of GBSSII has also been investigated in rice leaves, and it has been suggested that in light or under nitrogen deficiency, the accumulation of sugars induces the expression of starch-synthesis genes including GBSSII to convert the sugar to starch (Dian et al. 2003). On the other hand, in C. kessleri, the higher usage of carbon skeleton in the dark or after the addition of NH +4 (Miyachi and Miyachi 1985, 1987) did not seem to repress CkGBSS gene expression (Fig. 5b). Furthermore, in the low-CO2 condition in which CkGBSS expression was stimulated, rates of photosynthesis and sugar synthesis appeared to be rather lower than those in the high-CO2 condition (Fig. 5a, b). Thus, the regulatory mechanism for CkGBSS expression in the unicellular alga C. kessleri seems to be different from those in the vascular plants such as rice, which have storage- and non-storage tissues. In vascular plants, leaf transitory starch is synthesized during the day and mobilized at night to supply the carbon requirements of the plant. To maintain such a complex system, the expression mechanism of genes for starch synthases might have evolved to be regulated by circadian rhythm and in a strictly tissue-specific manner.
The levels of GBSS activity per starch amount in C. reinhardtii were almost the same as those in C. kessleri (Table 1). Wattebled et al. (2002) reported that C. reinhardtii GBSS has an activity per starch amount that is one order of magnitude higher than those reported for most vascular plants. They suggested that the difference in GBSS activities can be attributed to the difference in starch structure, granule size distribution, and/or to a more active GBSS protein per se, and that the extra C-terminal 11.4 kDa peptide seems to be required for full activity of GBSS in C. reinhardtii. Since the amounts of GBSS per starch amount were almost the same for C. reinhardtii and C. kessleri (Fig. 6), the specific activities per protein amount were not substantially different between the two species at least under the cultivation conditions used in this study. Therefore, if the specific activities obtained in the green algae under our experimental conditions would also be higher than those in vascular plants, the difference might be due to the difference in starch structure and granule size distribution.
In summary, cDNA for the GBSS of C. kessleri (CkGBSS cDNA) was isolated and characterized. The level of CkGBSS mRNA increased when CO2 concentration was lowered from 3 to 0.04% (air level) and the pyrenoid and the starch surrounding it developed actively (Miyachi et al. 1986). It was revealed that the biosynthesis of GBSS, which is considered to be mainly responsible for amylose synthesis, is regulated at the transcription level by changes in environmental CO2 concentration.
Abbreviations
- BE:
-
starch-branching enzyme
- CCM:
-
CO2-concentrating mechanism
- DP:
-
degree of polymerization
- GBSS:
-
granule-bound starch synthase
- High-CO2 cells:
-
cells cultured with air containing 3% CO2
- ISA:
-
isoamylase
- RT-PCR:
-
reverse transcriptional polymerase chain reaction
- Rubisco:
-
ribulose 1,5-bisphosphate carboxylase/oxygenase
- SS:
-
soluble starch synthase
References
Aizawa K, Miyachi S (1984) Carbonic anhydrase located on cell surface increases the affinity for inorganic carbon in photosynthesis of Dunaliella tertiolecta. FEBS Lett 173:41–44
Ball SG (1998) Regulation of starch biosynthesis. In: Rochaix J-D, Goldschmidt-Clermont M, Merchant S (eds) The molecular biology of chloroplasts and mitochondria in Chlamydomonas. Kluwer Academic Publishers, Dordrecht, pp 549–567
Bradford MM (1976) A rapid sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cao H, Imparl-Radosevich J, Guan H, Keeling PL, James MG, Myers AM (1999) Identification of the soluble starch synthase activities of maize endosperm. Plant Physiol 120:205–215
Delrue B, Fontaine T, Routier F, Decq A, Wieruszeski JM, van den Koornhuyse N, Maddelein ML, Fournet B, Ball S (1992) Waxy Chlamydomonas reinhardtii: monocellular algal mutants defective in amylose biosynthesis and granule-bound starch synthase activity accumulate a structurally modified amylopectin. J Bacteriol 174:3612–3620
Denyer K, Barber LM, Edwards EA, Smith AM, Wang TL (1997) Two isoforms of the GBSSI class of granule-bound starch synthase are differentially expressed in the pea plant (Pisum sativum L.). Plant Cell Environ 20:1566–1572
Dian W, Jiang H, Chen Q, Liu F, Wu P (2003) Cloning and characterization of the granule-bound starch synthase II gene in rice: gene expression is regulated by the nitrogen level, sugar and circadian rhythm. Planta 218:261–268
Edwards A, Vincken JP, Suurs LCJM, Visser RGF, Zeeman S, Smith A, Martin C (2002) Discrete forms of amylose are synthesized by isoforms of GBSSI in pea. Plant Cell 14:1767–1785
Fujita N, Taira T (1998) A 56-kDa protein is a novel granule-bound starch synthase existing in the pericarps, aleurone layers of immature seed in diploid wheat (Triticum monococcum L.). Planta 207:125–132
Fujiwara S, Fukuzara H, Tachiki A, Miyachi S (1990) Structure and differential expression of two genes encoding carbonic anhydrase in Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 87:9779–9783
Fujiwara S, Ishida N, Tsuzuki M (1996) Circadian expression of the carbonic anhydrase gene, Cah1, in Chlamydomonas reinhardtii. Plant Mol Biol 32:745–749
Fukuzawa H, Fujiwara S, Tachiki A, Miyachi S (1990a) Nucleotide sequences of two genes CAH1 and CAH2 which encode carbonic anhydrase polypeptides in Chlamydomonas reinhardtii. Nucleic Acids Res 18:6441–6442
Fukuzawa H, Fujiwara S, Yamamoto Y, Dionisio-Sese ML, Miyachi S (1990b) cDNA cloning, sequence, and expression of carbonic anhydrase in Chlamydomonas reinhardtii: regulation by environmental CO2 concentration. Proc Natl Acad Sci USA 87:4383–4387
Furukawa K, Tagaya M, Tanizawa K, Fukui T (1993) Role of the conserved Lys-X-Gly-Gly sequence at the ADP-glucose-binding site in Escherichia coli glycogen synthase. J Biol Chem 268:23837–23842
Gamborg OL, Miller RA, Ojima K (1968) Nutrient requirements of suspension cultures of soybean root cells. Exp Cell Res 50:151–158
Hodge JE, Hofreiter BT (1962) Determination of reducing sugars and carbohydrates. In: Whistler RL, Wolfrom ML (eds) Methods in carbohydrate chemistry, vol 1. Yew York Academic Press, New York, pp 380–394
Hogetsu D, Miyachi S (1979) Role of carbonic anhydrase in photosynthetic CO2 fixation in Chlorella. Plant Cell Physiol 20:747–756
Kuchitsu K, Tsuzuki M, Miyachi S (1988) Changes of starch localization within the chloroplast induced by changes in CO2 concentration during growth of Chlamydomonas reinhardtii: regulation of pyrenoid starch and stroma starch. Plant Cell Physiol 29:1269–1278
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Manners DJ, Sturgeon RJ (1982) Reserve carbohydrates of algae, fungi, and lichens. In: Loewus FA, Tanner W (eds) Encyclopedia of plant physiology, vol 13A, Plant carbohydrates I, Intracellular carbohydrates. Springer, Berlin Heidelberg, pp 472–514
Mérida A, Rodríguez-Galán JM, Vincent C, Romero JM (1999) Expression of the granule-bound starch synthase I (Waxy) gene from snapdragon is developmentally and circadian clock regulated. Plant Physiol 120:401–409
Miura K, Yamano T, Yoshioka S, Kohinata T, Inoue Y, Taniguchi F, Asamizu E, Nakamura Y, Tabata S, Yamato KT, Ohyama K, Fukuzawa H (2004) Expression profiling-based identification of CO2-responsive genes regulated by CCM1 controlling a carbon-concentrating mechanism in Chlamydomonas reinhardtii. Plant Physiol 135:1595–1607
Miyachi S, Miyachi S (1985) Ammonia induces starch degradation in Chlorella cells. Plant Cell Physiol 26:245–252
Miyachi S, Miyachi S (1987) Some biochemical changes related to starch breakdown induced by blue light illumination and by addition of ammonia to Chlorella cells. Plant Cell Physiol 28:309–314
Miyachi S, Tsuzuki M, Maruyama I, Gangar M, Miyachi S, Matsushima H (1986) Effects of CO2 concentration during growth on the intracellular structure of Chlorella and Scenedesmus (Chlorophyta). J Phycol 22:313–319
Murata T, Sugiyama T, Akazawa T (1965) Enzymatic mechanism of starch synthesis in glutinous rice grains. Biochem Biophys Res Commun 18:371–376
Nakamura T, Yamamori M, Hirano H, Hidaka S, Nagamine T (1995) Production of waxy (amylose-free) wheats. Mol Gen Genet 248:253–259
Nakamura T, Vrinten P, Hayakawa K, Ikeda J (1998) Characterization of a granule-bound starch synthase isoform found in the pericarp of wheat. Plant Physiol 118:451–459
Nakamura Y, Takahashi J, Sakurai A, Inaba Y, Suzuki E, Nihei S, Fujiwara S, Tsuzuki M, Miyashita H, Ikemoto H, Kawachi M, Sekiguchi H, Kurano N (2005) Some cyanobacteria synthesize semi-amylopectin type α-polyglucans instead of glycogen. Plant Cell Physiol 46:539–545
Ramazanov Z, Rawat M, Henk MC, Mason CB, Matthews SW (1994) The induction of the CO2-concentrating mechanism is correlated with the formation of the starch sheath around the pyrenoid of Chlamydomonas reinhardtii. Planta 195:210–216
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. 3rd edn. Cold Spring Harbor Labortory Press, New York
Smith AM, Denyer K, Martin C (1997) The synthesis of the starch granule. Annu Rev Plant Physiol Plant Mol Biol 48:67–87
Starr RC (1964) The culture collection of algae at Indiana University. Am J Bot 51:1013–1044
Sueoka N, Chiang KS, Kates JR (1967) Deoxyribonucleic acid replication in meiosis of Chlamydomonas reinhardtii I. Isotopic transfer experiments with a strain producing eight zoospores. J Mol Biol 25:44–67
Taira T, Fujita N, Takaoka K, Uematsu M, Wadano A, Kozaki S, Okabe S (1995) Variation in the primary structure of waxy proteins (granule-bound starch synthase) in diploid cereals. Biochem Genet 33:269–281
Tsuzuki M, Miyachi S (1989) The function of carbonic anhydrase in aquatic photosynthesis. Aquat Bot 34:85–104
Vrinten PL, Nakamura T (2000) Wheat granule-bound starch synthase I and II are encoded by separate genes that are expressed in different tissues. Plant Physiol 122:255–263
Wang SJ, Yeh KW, Tsai CY (2001) Regulation of starch granule-bound starch synthase I gene expression by circadian clock and sucrose in the source tissue of sweet potato. Plant Sci 161:635–644
Wattebled F, Buléon A, Bouchet B, Ral J-P, Liénard L, Delvallé D, Binderup K, Dauvillée D, Ball S, d’Hulst C (2002) Granule-bound starch synthase I: a major enzyme involved in the biogenesis of B-crystallites in starch granules. Eur J Biochem 269:3810–3820
Acknowledgments
We are grateful to Drs. Naoko Fujita and Perigio B. Francisco Jr. (Akita Prefectural University, Japan) for helpful comments and for correcting the English version. This work was supported by Grants-in-Aid from the Ministry of Education, Science, Sports and Culture, Japan (13640657, 13740463, and 13874112) and the Promotion and Mutual Aid Corporation for Private Schools.
Author information
Authors and Affiliations
Corresponding author
Additional information
The nucleotide sequence reported in this paper has been submitted to the DDBJ/EMBL/GenBank databases and was assigned the accession number AB232549. Yasunori Oyama, Asako Izumo, Shoko Fujiwara contributed equally to this work.
Rights and permissions
About this article
Cite this article
Oyama, Y., Izumo, A., Fujiwara, S. et al. Granule-bound starch synthase cDNA in Chlorella kessleri 11 h: cloning and regulation of expression by CO2 concentration. Planta 224, 646–654 (2006). https://doi.org/10.1007/s00425-006-0239-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-006-0239-7