Abstract
We used amplified fragment length polymorphisms (AFLP) to analyze the stability of DNA methylation throughout Arabidopsis development. AFLP can detect genome-wide changes in cytosine methylation produced by DNA demethylation agents, such as 5-azacytidine, or specific mutations at the DDM1 locus. In both cases, cytosine demethylation is associated with a general increase in the presence of amplified fragments. Using this approach, we followed DNA methylation at methylation sensitive restriction sites throughout Arabidopsis development. The results show a progressive DNA methylation trend from cotyledons to vegetative organs to reproductive organs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA methylation is an important mechanism for epigenetic control of gene expression in many eukaryotes (Martienssen and Richards 1995). In mammals, methylation plays a role in regulation of gene expression throughout development. Most epigenetic phenomena in mammals are established in the early embryo (Razin and Riggs 1980) and change throughout development (Razin and Cedar 1994; Yoder et al. 1997). In contrast to the role of methylation in mammals, genome methylation is restricted to the early states of Drosophila melanogaster embryonic development (Lyko et al. 2000), and zebrafish development takes place without variation in DNA methylation (Donald et al. 1999).
DNA methylation is very common in plant genomes. It has frequently been associated with gene silencing, specially in the case of transposable elements (for review see Okamoto and Hirochika 2001). Epigenetic alleles, resulting from DNA methylation at specific loci, have also been described in different species. Hypermethylated alleles of SUP, PAI and AG in Arabidopsis (Bender and Fink 1995; Jacobsen and Meyerowitz 1997; Melquist et al. 1999; Jacobsen et al. 2000) and LCYC in Linaria (Cubas et al. 1999) result in silencing, while hypomethylated alleles of AP3 or FWA in Arabidopsis display ectopic expression (Finnegan et al. 1996; Soppe et al. 2000).
Several reports suggest a role for DNA methylation in the regulation of plant development, although evidence is fragmentary. Significant differences in cytosine methylation have been observed among different organs in species such as tomato (Messeguer et al. 1991), rice (Xiong et al. 1999) or Silene latifolia (Zluvova et al. 2001), and among different developmental phases in Pinus (Fraga et al. 2002) and Prunus (Bitonti et al. 2002). In Arabidopsis, young seedlings were reported to have lower DNA methylation levels than mature leaves (Finnegan et al. 1998a).
Genome-wide demethylation causes abnormal development in Arabidopsis thaliana (Finnegan et al. 1998a, 2000). Depending on the genes affected, reduction in cytosine methylation can result in opposing effects on flowering time. Mutants at the DDM1 (Decrease in DNA Methylation) locus show up to a 70% reduction in their total genomic 5-methylcytosine level and exhibit a late-flowering phenotype after a number of selfing generations (Vongs et al. 1993; Kakutani et al. 1995). The hypomethylated dominant epiallele of FWA is constitutively transcribed and confers late flowering phenotype on plants of Ler and Col ecotypes (Koornneef et al. 1991; Kakutani 1997; Soppe et al. 2000) but not on C24 plants (Genger et al. 2003). Overexpression of antisense DNA methyltransferase MET1 in transgenic Arabidopsis plants causes demethylation, which confers flowering delay in Col transgenic plants (Ronemus et al. 1996) yet early flowering in C24 transgenic plants (Finnegan et al. 1998b). Furthermore, demethylation agents such as 5-azacytidine (5-azaC) as well as vernalization treatments promote earlier flowering associated with reduced levels of DNA methylation (Burn et al. 1993; Finnegan et al. 1998b).
The aim of this work was to analyze methylation stability throughout plant development using an AFLP approach. We showed that this technique is able to detect the effect of demethylation agents or mutations on genome methylation patterns. Using this approach we have followed DNA methylation throughout Arabidopsis development and observed that different organs show different methylation patterns with a trend towards increasing DNA methylation during plant development.
Materials and methods
Plant material and growth conditions
The ecotype Lansberg erecta (Ler) and the fve-1 mutant were provided by M. Koornneef (Wageningen, The Netherlands), and the ddm1 mutant and its wild type genetic background Columbia (Col) were provided by Eric J. Richards (St Louis, Missouri) and Lehle’s Seed (USA), respectively.
Plants were grown either aseptically in Petri dishes containing solidified GM medium (MS, Murashige and Skoog 1962, with 0.7% sucrose) or in pots containing a mixture of peat and vermiculite (3:1, v/v), in a growth chamber at 22°C and 16 h photoperiod conditions.
For 5-azacytidine (5-azaC) treatments, sterile Ler seeds were imbibed on filter paper soaked with different concentrations of fresh 5-azaC (5 μM, 50 μM and 500 μM) or sterile water for control plants. Seeds were transferred daily to new filter paper containing the corresponding fresh 5-azaC solution or sterile water. After 5 days, germinating seeds were transferred to a GM medium 10 days before analysis. For vernalization treatments, fve-1 plants were vernalized by germinating seeds in the dark for 4, 6 or 8 weeks at 4°C, and then transferring the seedlings to long-day photoperiods at 22°C for 1 and 2 weeks. In all cases, total genomic DNA was isolated using the DNeasy kit (QIAGEN).
AFLP analysis
Each sample was simultaneously analyzed using EcoRI +2/HpaII +3 and EcoRI +2/MspI+3 reactions to compare banding patterns, as described by Cervera et al. (2002). The following primer selective nucleotide combinations AC/AAT, AA/AAT, AA/ATC, AT/ATC and AT/ACT, were used to analyze ddm1, 5-azaC and vernalization profiles, while AA/AAT, AA/ATC and AT/ACT were used to study methylation patterns throughout Arabidopsis development. In all experiments, we detected fragments differing in intensity, probably due to different degrees of cytosine methylation in different cells and tissues contained in the analyzed samples. Only clearly scorable fragments were considered and their presence or absence was visually determined by two independent observations. All experiments were repeated twice using different DNA extractions.
Results
AFLPs detect methylation changes in Arabidopsis genomic DNA
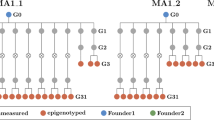
AFLP analysis using double digestion with EcoRI and either HpaII or MspI isoschizomers allowed identification of methylation-sensitive CCGG polymorphisms in the Arabidopsis genome (Cervera et al. 2002). MspI cleaves C5mCGG but not 5mCCGG sequences, whereas HpaII is inactive if one or both cytosines are fully methylated but cleaves the hemimethylated sequence (Korch and Hagblom 1986; McClelland et al. 1994). To test whether this technique can detect genome-wide changes in cytosine methylation, two experiments were performed. First, the total DNA of hypomethylated ddm1 seedlings (Vongs et al. 1993) was compared to the DNA of wild-type Col seedlings. A total of 470 scorable amplified fragments were detected using 5 AFLP primer combinations. Forty percent of them were sensitive to methylation and therefore differentially digested when comparing EcoRI/ HpaII and EcoRI/ MspI patterns. Forty seven percent of the differentially methylated fragments showed the same pattern in ddm1 and Col samples while 53% showed different patterns (differentially methylated polymorphic fragments), as shown in Fig. 1a,b (patterns I and II), confirming that ddm1 mutations strongly affect the methylation state of CCGG sites. Seventy percent of the differentially methylated polymorphic fragments observed between ddm1 and Col samples corresponded to new restriction fragments in the ddm1 background (Fig. 1a,b, pattern I). The other 30% corresponded to fragments present in Col background that disappeared in the demethylated mutant (Fig. 1a,b, pattern II).
Changes in methylation-sensitive polymorphisms associated with ddm1 DNA demethylation. Arrows indicate polymorphic methylation-sensitive fragments and the roman number corresponds to pattern type. a Number and percentage of methylation-sensitive fragments showing a specific pattern when comparing ddm1 mutants and Col wild type plants. b Details of ddm1 and Col AFLP fingerprints
Treatments with 5-azaC cause global demethylation of DNA (Creusot et al. 1982; Jones et al. 1983). Using 5 AFLP primer combinations in Ler plants treated with increasing concentrations of 5-azaC (5 μM, 50 μM, 500 μM) we detected a total of 378 scorable amplified fragments. Thirty eight percent of them were differentially digested when comparing EcoRI/ HpaII and EcoRI/ MspI patterns. Forty-five percent of the differentially digested fragments showed the same patterns among analyzed samples while 55% showed profiles related to the 5-azaC treatment (Fig. 2a,b, patterns I, II). We found a progressive increase of amplified fragments on samples treated with 50 μM to 500 μM of 5-azaC (Fig. 2a,b, pattern I). In fact, 87% of the differences found between control and treated samples were due to the appearance of new amplified fragments in the 5-azaC treated samples (Fig. 2a,b, pattern I). The remaining 13% corresponded to fragments present in Ler wild-type samples that disappeared in the demethylated samples (Fig. 2a,b, pattern II). AFLP profiles resulting from ddm1 and 5-azaC experiments were not comparable since they had been generated on different genetic backgrounds.
Changes in methylation-sensitive polymorphisms associated with 5-azaC DNA demethylation. Arrows indicate polymorphic methylation-sensitive fragments and the roman number corresponds to pattern type. a Number and percentage of methylation-sensitive fragments showing a specific pattern comparing Ler plants untreated or treated with different concentrations of 5-azaC. b Details of AFLP profiles obtained for Ler plants treated with 5-azaC
Finally, a prolonged exposure to low non freezing temperatures, also known as vernalization, has also been proposed as a cause for genomic DNA demethylation (Finnegan et al. 1998b). We compared the total DNA of fve-1 seedlings vernalized for 4, 6 or 8 weeks, to DNA from non-vernalized fve-1 plants showing the same developmental stage as the previous ones, using 5 AFLP primer combinations. The results did not show any consistent DNA methylation changes either immediately,or after 4, 6 or 8 weeks of treatment or 1 or 2 weeks after resuming growth at room temperature.
DNA methylation changes throughout Arabidopsis development
The sensitivity of AFLP in the analysis of the status of DNA methylation at genome level was such that we used this technique to study the existence of methylation changes associated with Arabidopsis development. With this purpose we characterized the patterns of DNA methylation in cotyledons (C), first pair of rosette leaves (1L), mature rosette leaves (ML), cauline leaves (CL), flower buds (B) and open flowers (F) of Ler, using three AFLP primer combinations. A total of 257 scorable amplified fragments were detected, and 103 (40%) of these were differentially digested when comparing EcoRI/ HpaII and EcoRI/ MspI patterns. Thirty one of the differentially digested fragments (30%) showed distinct organ-specific patterns (Fig. 3) which could be grouped into five different classes (I-V). Class I grouped 8 fragments (26%) specifically present in cotyledons but absent in the rest of the analyzed organs, while class II included 1 fragment (3%) with the opposite pattern, absent in cotyledons, but present in the rest of the organs. Class III included 15 fragments (48%) present in cotyledons, and at least the first pair of rosette leaves, but disappearing gradually in the rest of the vegetative or reproductive organs. Class IV grouped 3 fragments (10%) present only in reproductive organs and class V, 4 fragments (13%) absent in the embryonic and first vegetative organs, appearing in later vegetative organs and disappearing in reproductive organs. Therefore, out of 31 methylation-sensitive organ-specific polymorphisms, 23 (74%) were initially present in cotyledons and disappeared gradually during later development of the plant (pattern types I and III of Fig. 3). Only 4 (13%) of the methylation-sensitive organ-specific polymorphisms were restricted to vegetative or reproductive organs (pattern types II and IV of Fig. 3), and 4 were found in vegetative organs but not in cotyledons and reproductive organs (pattern type V of Fig. 3). Specific representative fragments of these patterns were cloned and sequenced previously (Cervera et al. 2002). None of them showed internal HpaII/ MspI restriction sites.
Changes in methylation-sensitive polymorphisms throughout Arabidopsis development. Arrows indicate polymorphic methylation-sensitive fragments and the roman number corresponds to pattern type. a Number and percentage of methylation-sensitive fragments showing specific patterns in different Ler organs: C (cotyledons), 1L (first pair of rosette leaves), ML (mature rosette leaves), CL (cauline leaves), B (flower buds) and F (open flowers). b Details of AFLP fingerprint in different Ler organs
Discussion
We previously used AFLP analysis combining double digestion with EcoRI and either HpaII or MspI isoschizomers to identify anonymous CCGG sites in which cytosines are differentially methylated (Cervera et al. 2002). This technique was applied here to follow genome-wide methylation changes in ddm1 seedlings and in seedlings treated with 5-azaC. In both analyses, 60% of the scorable amplified fragments showed similar digestibility in HpaII and MspI assays, representing unmethylated CCGG restriction sites. The other 40% corresponded to CCGG restriction sites in which differential cytosine methylation resulted in differential digestion by the two isoschizomers. In our experiments, ddm1 mutations or 5-azaC treatments caused over 50% of the differentially digested restriction fragments (differentially methylated polymorphic fragments). Comparison of these differentially methylated polymorphic fragments between ddm1 or 5-azaC samples and their wild type genetic background Col or Ler, respectively, indicated that a reduction in DNA methylation in ddm1 samples and 5-azaC treated plants was associated with a general increase in the number of amplified fragments. A total of 70% of the differentially methylated polymorphic fragments were only observed in ddm1 plants, which could be associated with the demethylation of CCGG restriction sites in this mutant (Fig. 1a,b, pattern I). This result is in agreement with the 5-methylcytosine (5 mC) reduction observed in ddm1 mutants. Vongs et al. (1993) reported that ddm1 affects global levels of DNA methylation resulting in a 70% reduction of 5 mC in TaqI sites (TCGA), measured by thin-layer chromatography. Furthermore, using reversed-phase HPLC, Ronemus et al. (1996) estimated that ddm1 mutation caused a 75% reduction in 5 mC content. In the same way, 5-azaC treatments also yielded a large proportion of new restriction fragments (87%) among the differentially methylated polymorphic fragments, which could also be associated with demethylation of CCGG restriction sites (Fig. 2a,b, pattern I). There are no data for the effect of 5-azaC on Arabidopsis 5 mC content. However, Nicotiana cells treated with 100 μM 5-azaC showed a reduction in the total 5 mC content ranging from 37% to 55%, as determined by HPLC (Burn et al. 1993).
Vernalization treatments in Arabidopsis C24 ecotype resulted in a 15% decrease in total DNA methylation in TaqI sites, measured by thin-layer chromatography (Finnegan et al. 1998b). We have not detected DNA methylation changes related to vernalization, indicating that this treatment does not cause such a general effect on DNA methylation as 5-azaC treatments do. However, we cannot exclude the possibility that vernalization results in DNA methylation changes at specific loci. Nevertheless, the silencing of the flowering repressor FLC caused by vernalization, has recently been shown to be related to changes in histone methylation at the FLC chromatin (Bastow et al. 2004) and not to involve DNA methylation (Finnegan et al. 2004).
Aerial plant development takes place post-embryonically as a result of cell proliferation and differentiation at the apical meristem. Transitions between different developmental phases involve changes in the pattern of cellular differentiation and organ formation at the apical meristem which are genetically regulated. DNA methylation could play a role in plant development as a mechanism to epigenetically maintain developmental decisions in proliferating cells. Our results concerning methylation changes associated with plant development showed that 40% of the scorable amplified fragments corresponded to differentially methylated CCGG restriction sites. Among them, 30% showed different methylation pattern during Arabidopsis development. Cotyledons showed the highest number of differentially methylated HpaII/ MspI amplified fragments (74%, pattern types I and III of Fig. 3), the intensity and number of which decreased gradually along post-embryonic differentiation of vegetative and reproductive organs. Only 26% of differentially methylated HpaII/ MspI amplified fragments were restricted to vegetative or reproductive organs (pattern types II, IV and V of Fig. 3). Either new methylation events or demethylation and digestion of internal restriction sites giving rise to fragments of smaller size could be the origin of the disappearance of amplified fragments along with development. Given the association between cytosine methylation and the number of amplified fragments, we think methylation could be a major cause for the disappearance of amplified fragments. Furthermore, most amplified methylation-sensitive polymorphic fragments lack internal 5′-CCGG restriction sites (Cervera et al. 2002), which supports this hypothesis. Hence, Arabidopsis cotyledons contain a higher number of demethylated HpaII/ MspI restriction sites than post-embrionically developed vegetative and reproductive organs.
These results are consistent with data from tomato (Messeguer et al. 1991) and white campion (Silene latifolia) (Zluvova et al. 2001). In both species, 5 mC content was lower in cotyledons than in vegetative organs. Similarly, Pinus radiata juvenile phase was characterised by a lower content of 5 mC (30–35%) than mature phase (60%). The same behaviour was observed in Prunus persica, where adult meristems showed a higher 5 mC content than juvenile ones (Bitonti et al. 2002). Altogether, these results suggest the existence of a progressive DNA methylation along with plant development, in a similar way to what has been described throughout the embryonic development of mammals (Goto and Monk 1998; Hsieh 2000). Whether these methylation changes have a role in maintaining developmental states at the plant meristem, or, as has been shown in maize (Lund et al. 1995), reflect the output of a genetic mechanism silencing transposable elements and other repeated sequences, remains to be fully understood. The use of AFLP in combination with other genome wide analytical tools such as DNA microarrays in fully sequenced genomes, will allow a complete characterization of the developmental dynamics of DNA methylation at sequence-specific sites.
Abbreviations
- AFLP :
-
Amplified fragment length polymorphism
- 5-azaC :
-
5-azacytidine
- 5-mC :
-
5-methylcytosine
References
Bastow R, Mylne JS, Lister C, Lippman Z, Martienssen RA, Dean C (2004) Vernalization requires epigenetic silencing of FLC by histone methylation. Nature 427:164–167
Bender J, Fink GR (1995) Epigenetic control of an endogenous gene family is revealed by a novel blue fluorescent mutant of Arabidopsis. Cell 83:725–734
Bitonti MB, Cozza R, Chiappetta A, Giannino D, Castiglione MR, Dewitte W, Mariotti D, Van Onckelen H, Innocenti AM (2002) Distinct nuclear organization, DNA methylation pattern and cytokinin distribution mark juvenile, juvenile-like and adult vegetative apical meristems in peach (Prunus persica (L.) Batsch). J Exp Bot 53:1047–1054
Burn JE, Bagnall DJ, Metzger JD, Dennis ES, Peacock WJ (1993) DNA methylation, vernalization, and the initiation of flowering. Proc Natl Acad Sci USA 90:287–291
Cervera MT, Ruiz-García L, Martínez-Zapater JM (2002) Analysis of DNA methylation-sensitive AFLP markers. Mol Genet Genomics 268:543–552
Creusot F, Acs G, Christman JK (1982) Inhibition of DNA methyltransferase and induction of Friend erythroleukemia cell differentiation by 5-azacytidine and 5-aza-2′-deoxycytidine. J Biol Chem 257:2041–2048
Cubas P, Vincent C, Coen E (1999) An epigenetic mutation responsible for natural variation in floral symmetry. Nature 401:157–161
Donald M, Clark VH, Bird A (1999) Absence of genome-wide changes in DNA methylation during development of the zebrafish. Nat Genet 23:139–140
Finnegan EJ, Peacock WJ, Dennis ES (1996) Reduced DNA methylation in Arabidopsis thaliana results in abnormal plant development. Proc Natl Acad Sci USA 93:8449–8454
Finnegan EJ, Genger RK, Peacock WJ, Dennis ES (1998a) DNA methylation in plants. Annu Rev Plant Physiol Plant Mol Biol 49:223–247
Finnegan EJ, Genger RK, Kovac K, Peacock WJ, Dennis ES (1998b) DNA methylation and the promotion of flowering by vernalization. Proc Natl Acad Sci USA 95:5824–5829
Finnegan EJ, Peacock WJ, Dennis ES (2000) DNA methylation, a key regulator of plant development and other processes. Curr Opin Genet Dev 10:217–223
Finnegan EJ, Sheldon CC, Jardinaud F, Peacock WJ, Dennis ES (2004) A cluster of Arabidopsis genes with a coordinate response to an environmental stimulus. Current Biol 14:911–916
Fraga M, Cañal MJ, Rodriguez R (2002) Phase-change related epigenetic and physiological changes in Pinus radiata D. Don. Planta 215:672–678
Genger RK, Peacock WJ, Dennis ES, Finnegan EJ (2003) Opposing effects of reduced DNA methylation on flowering time in Arabidopsis thaliana. Planta 216:461–466
Goto T, Monk M (1998) Regulation of X-chromosome inactivation in development in mice and humans. Microbiol Mol Biol Reviews 62:362–378
Hsieh CL (2000) Dynamics of DNA methylation pattern. Curr Opin Genet Dev 10:224–228
Jacobsen SE, Meyerowitz EM (1997) Hypermethylated SUPERMAN epigenetic alleles in Arabidopsis. Science 277:1100–1103
Jacobsen SE, Sakai H, Finnegan EJ, Cao X, Meyerowitz EM (2000) Ectopic hypermethylation of flower-specific genes in Arabidopsis. Curr Biol 10:179–186
Jones PA, Taylor SM, Wilson VL (1983) Inhibition of DNA methylation by 5-azacytidine. Recent Res Cancer Res 84:202–211
Kakutani T (1997) Genetic characterization of late-flowering traits induced by DNA hypomethylation mutation in Arabidopsis thaliana. Plant J 12:1447–1451
Kakutani T, Jeddeloh JA, Richards EJ (1995) Characterization of an Arabidopsis thaliana DNA hypomethylation mutant. Nucleic Acids Res 23:130–137
Koornneef M, Hanhart CJ, van der Veen JH (1991) A genetic and physiological analysis of late flowering mutants in Arabidopsis thaliana. Mol Gen Genet 229:57–66
Korch C, Hagblom P (1986) In-vivo-modified gonococcal plasmid pJD1. A model system for analysis of restriction enzyme sensitivity to DNA modifications. Eur J Biochem 161:519–524
Lund G, Messing J, Viotti A (1995) Endosperm-specific demethylation and activation of specific alleles of alpha-tubulin genes of Zea mays L. Mol Gen Genet 246:716–722
Lyko F, Ramsahoye BH, Jainisch R (2000) DNA methylation in Drosophila melanogaster. Nature 408:538–540
Martienssen RA, Richards EJ (1995) DNA methylation in eukaryotes. Curr Opin Genet Dev 5:234–242
McClelland M, Nelson M, Raschke E (1994) Effect of site-specific modification on restriction endonucleases and DNA modification methyltransferases. Nucleic Acids Res 22:3640–3659
Melquist S, Luff B, Bender J (1999) Arabidopsis PAI gene arrangements, cytosine methylation and expression. Genetics 153:401–413
Messeguer R, Ganal MW, Steffens JC, Tanksley SD (1991) Characterization of the level, target sites and inheritance of cytosine methylation in tomato nuclear DNA. Plant Mol Biol 16:753–770
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassay with tobacco tissue cultures. Physiol Plant 15:473–479
Okamoto H, Hirochika H (2001) Silencing of transposable elements in plants. Trends Plant Sci 6:527–534
Razin A, Cedar H (1994) DNA methylation and genomic imprinting. Cell 77:473–476
Razin A, Riggs AD (1980) DNA methylation and gene regulation. Science 210:604–610
Ronemus MJ, Galbiati M, Ticknor C, Chen J, Dellaporta SL (1996) Demethylation-induced developmental pleiotropy in Arabidopsis. Science 273:654–657
Soppe W, Jacobsen SE, Alonso-Blanco C, Jackson JP, Kakutani T, Koornneef M, Peeters AJ (2000) The late flowering phenotype of fwa mutants is caused by gain-of-function epigenetic alleles of a homeodomain gene. Mol Cell 6:791–802
Vongs A, Kakutani T, Martienssen RA, Richards EJ (1993) Arabidopsis thaliana DNA methylation mutants. Science 260:1926–1928
Xiong LZ, Xu CG, Saghai Maroof MA, Zhang Q (1999) Patterns of cytosine methylation in an elite rice hybrid and its parental lines, detected by a methylation-sensitive amplification polymorphism technique. Mol Gen Genet 261:439–446
Yoder JA, Soman NS, Verdine GL, Bestor TH (1997) DNA (cytosine-5)-methyltransferases in mouse cells and tissues. Studies with a mechanism-based probe. J Mol Biol 270:385–395
Zluvova J, Janousek B, Vyskot B (2001) Immunohistochemical study of DNA methylation dynamics during plant development. J Exp Bot 52:2265–2273
Acknowledgements
L. Ruiz-Garcia and M.-T. Cervera contributed equally to this work. M.-T. C. was funded by a MEC contract. This research was funded by MCYT Grant# BIO2001-3891-CO2-01.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ruiz-García, L., Cervera, M.T. & Martínez-Zapater, J.M. DNA methylation increases throughout Arabidopsis development. Planta 222, 301–306 (2005). https://doi.org/10.1007/s00425-005-1524-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-005-1524-6